Natural Products from Cyanobacteria: Focus on Beneficial Activities
Abstract
1. Introduction
2. Methods for Dataset Construction
3. Taxonomy of the Producing Strains
4. Chemical Diversity and Bioactivity of Natural Products from Cyanobacteria
5. Beneficial Activities of Natural Products Produced by Cyanobacteria
5.1. Antimicrobial Activity
5.1.1. Antibacterial Activity
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Eucapsitrione | Anthraquinone derivative | -Antibacterial -No antifungal -Cytotoxic | Eucapsis sp. UTEX 1519 | [74] |
| Kulolide-like analogs | Depsipeptide | -Antibacterial -No antifungal -Antiprotozoal -Lethal -Cytotoxic -VGSC (Voltage Gate Sodium Channel) activation | Lyngbya majuscula, Rivularia sp., Moorea producens, Okeania sp., Symploca hydnoides, Oscillatoria margaritifera | [26,75,80,81,82,83,84,85,86,87,88,89,90] |
| Abietic acids | Terpene | -Antibacterial -No lethality -No antialgal | Plectonema radiosum LEGE 06105, Nostoc sp. LEGE 06077 and LEGE 07365, Chroococcidiopsis sp. LEGE 06174, Synechocystis sp. LEGE 06079, Synechocystis salina LEGE 06099, Leptolyngbya ectocarpi LEGE 11425, Nodosilinea sp. LEGE 13457, Nodosilinea nodulosa LEGE 07084 | [91] |
| Hapalindole-like | Alkaloid | -Antibacterial -Antifungal -Antialgal -Cytotoxic -Insecticidal -Lethal activity -Reverse multidrug resistance (MDR) -VGSC modulator | Hapalosiphon fontinalis, Westiellopsis sp., Fischerella musicola, Hapalosiphon welwitschii, Westiella intricata, Fischerella ambigua, Hapalosiphon delicatulus, Hapalosiphon hibernicus, Westiellopsis prolifica, Fischerella sp., Hapalosiphon laingii | [92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117] |
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Cyanobacterin | Lactone derivative | -Antialgal -Anti-cyanobacterial -Growth inhibition | Scytonema hofmanni UTEX 2349, Nostoc linckia CALU 892 | [118,119,120,121] |
| Fischerellins | Polyketide | -Antialgal -Anti-cyanobacterial -Antifungal -Lethal -Growth inhibition | Fischerella musicola, Fischerella sp., Fischerella ambigua, Fischerella tesserantii | [122,123,124,125] |
| Westiellamide-like analogs | Peptide | -Antialgal -Anti-cyanobacterial -No antifungal -Lethal activity -Cytotoxic | Westiellopsis prolifica EN-3-1, Nostoc sp. 31, Stigonema dendroideum IA-45-3, Oscillatoria raoi TAU IL-76-1-2, Nostoc spongiaeforme var. tenue str. Carmeli | [126,127,128,129,130,131] |
| Ambigols | Alkaloid | -Antialgal -Antibacterial -Antifungal -Antiprotozoal -Lethal activity -Cytotoxicity -Enzyme inhibition | Fischerella ambigua 108b | [132,133] |
| Schizotrin-like analogs | Peptide | -Antialgal -Antibacterial -Antifungal -Antiprotozoal -Lethal activity -Cytotoxicity | Schizothrix sp. TAU IL-82-2, Lyngbya sp. 36.91, Phormidium sp. LEGE 05292, Tychonema sp. CCAP 1462/13 | [134,135,136,137,138,139,140,141] |
5.1.2. Antialgal Activity
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Hassallidins | Glycolipopeptide | -Antifungal -No antibacterial activity | Hassalia sp. B02-07, Anabaena sp. (SYKE 748A, 90y1998, 90M3, 299B, 258, SYKE763A, 0TU33S16, 0TU43S8, 1TU33S8, 1TU35S12, 1TU44S9, 1TU44S16, SYKE971/6, NIVA-CYA269/2, NIVA-CYA269/6, XPORK5C, XSPORK7B, XSPORK36B, XSPORK14D, BECID19), Anabaena cylindrica Bio33 Cylindrospermopsis raciborskii (ATC-9502 & CS-505), Aphanizomenon gracile Heaney/Camb 1986 140 1/1, Nostoc sp. (159 & 113.5), Tolypothrix sp. PCC 9009 Planktothix serta PCC 8927 | [144,145,146,147,148] |
| Lyngbyabellins | Depsipeptide | -Antifungal -No antibacterial activity -Lethal activity -Cytotoxic | Lyngbya majuscula Lyngbya sp., Lyngbya bouillonii Moorea bouillonii | [149,150,151,152,153,154,155,156] |
| Micro-guanidines | Guanidine derivative | -Antifungal -No cytotoxicity -No protease inhibition | Microcystis sp. TAU IL-306, Microcystis aeruginosa TAU IL-374 | [157,158,159] |
| Majusculamides | Lipopeptide | -Antifungal -Cytotoxic -Immunosuppressive activity -Actin filaments disrupting -Anti-settlement activity | Lyngbya majuscula, Lyngbya polychroa | [160,161,162,163,164,165,166] |
5.1.3. Antifungal Activity
5.1.4. Antiviral Activity
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Aplysiatoxins | Alkaloid | -Antiviral -Dermatitis and swimmer’s itch agents -Cytotoxic | Lyngbya majuscula, Schizothrix calcicola, Oscillatoria nigro-viridis, Trichodesmium erythaeum | [51,168,170,171,172,173] |
| Cyanovirin-N | Protein | -Antiviral -No cytotoxicity -Stop fusion and transmission of HIV-1 virus | Nostoc ellipsosporum Cyanothece sp. | [174,175,176] |
| Calcium spirulan | Polysaccharide | -Antiviral -No cytotoxicity -Low anticoagulant activity | Arthrospira platensis | [67,177,178] |
5.1.5. Antiprotozoal Activity (Against Malaria, Leishmaniosis, Chagas Disease)
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Companeramides | Depsipeptide | -Antiprotozoal -No significant cytotoxicity | Leptolyngbya sp. or «Hyalidium» | [29] |
| Hoshinolactam | Lactam | -Antiprotozoal -No cytotoxicity | Oscillatoria sp. | [182] |
| Dolastatins | Peptide | -Antiprotozoal -Lethal -Cytotoxic | Lyngbya majuscula, Symploca hydnoides, Lyngbya sp., Symploca sp. VP642, Lyngbya-Schizothrix assemblage | [30,32,75,183,184,185,186,187,188,189] |
5.2. Potential Anticancer Activity
5.2.1. Cytotoxic Activity
5.2.2. Protease Inhibitory Activity
5.2.3. Histone Deacetylase Inhibitors
5.3. Anti-Inflammatory and Antioxidant Activity
5.3.1. Anti-Inflammatory Activity
5.3.2. Antioxidant Activity
5.4. Other Metabolites with Potential Beneficial Properties
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Whitton, B.A.; Potts, M. Ecology of Cyanobacteria II: Their Diversity in Space and Time; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2012; pp. 1–13. [Google Scholar]
- Kurmayer, R.; Deng, L.; Entfellner, E. Role of toxic and bioactive secondary metabolites in colonization and bloom formation by filamentous cyanobacteria Planktothrix. Harmful Algae 2016, 54, 69–86. [Google Scholar] [CrossRef]
- Mazard, S.; Penesyan, A.; Ostrowski, M.; Paulsen, I.T.; Egan, S. Tiny Microbes with a Big Impact: The Role of Cyanobacteria and Their Metabolites in Shaping Our Future. Mar. Drugs 2016, 14, 97. [Google Scholar] [CrossRef] [PubMed]
- Buratti, F.M.; Manganelli, M.; Vichi, S.; Stefanelli, M.; Scardala, S.; Testai, E.; Funari, E. Cyanotoxins: Producing organisms, occurrence, toxicity, mechanism of action and human health toxicological risk evaluation. Arch. Toxicol. 2017, 91, 1049–1130. [Google Scholar] [CrossRef] [PubMed]
- Humbert, J.-F.; Törökné, A. New Tools for the Monitoring of Cyanobacteria in Freshwater Ecosystems. In Handbook of Cyanobacterial Monitoring and Cyanotoxin Analysis; John Wiley & Sons, Ltd.: Chichester, UK, 2017; pp. 84–88. [Google Scholar]
- Salmaso, N.; Bernard, C.; Humbert, J.-F.; Akçaalan, R.; Albay, M.; Ballot, A.; Catherine, A.; Fastner, J.; Häggqvist, K.; Horecká, M.; et al. Basic Guide to Detection and Monitoring of Potentially Toxic Cyanobacteria. In Handbook of Cyanobacterial Monitoring and Cyanotoxin Analysis; John Wiley & Sons, Ltd.: Chichester, UK, 2017; pp. 46–69. ISBN 0-7803-8560-8. [Google Scholar]
- Paerl, H.W.; Otten, T.G. Harmful Cyanobacterial Blooms: Causes, Consequences, and Controls. Microb. Ecol. 2013, 65, 995–1010. [Google Scholar] [CrossRef]
- Zanchett, G.; Oliveira-Filho, E.C. Cyanobacteria and cyanotoxins: From impacts on aquatic ecosystems and human health to anticarcinogenic effects. Toxins 2013, 5, 1896–1917. [Google Scholar] [CrossRef] [PubMed]
- Tidgewell, K.; Clark, B.R.; Gerwick, W.H. The natural products chemistry of cyanobacteria. In Comprehensive Natural Products II Chemistry and Biology; Elsevier: Oxford, UK, 2010; Volume 2, pp. 141–188. [Google Scholar]
- Ali Shah, S.A.; Akhter, N.; Auckloo, B.N.; Khan, I.; Lu, Y.; Wang, K.; Wu, B.; Guo, Y.W. Structural diversity, biological properties and applications of natural products from cyanobacteria. A review. Mar. Drugs 2017, 15, 354. [Google Scholar] [CrossRef] [PubMed]
- Mi, Y.; Zhang, J.; He, S.; Yan, X. New peptides isolated from marine cyanobacteria, an overview over the past decade. Mar. Drugs 2017, 15, 132. [Google Scholar] [CrossRef]
- Wang, M.; Zhang, J.; He, S.; Yan, X. A review study on macrolides isolated from cyanobacteria. Mar. Drugs 2017, 15, 126. [Google Scholar] [CrossRef] [PubMed]
- Chlipala, G.E.; Mo, S.; Orjala, J. Chemodiversity in freshwater and terrestrial cyanobacteria—A source for drug discovery. Curr. Drug Targets 2011, 12, 1654–1673. [Google Scholar] [CrossRef]
- Sivonen, K.; Leikoski, N.; Fewer, D.P.; Jokela, J. Cyanobactins-ribosomal cyclic peptides produced by cyanobacteria. Appl. Microbiol. Biotechnol. 2010, 86, 1213–1225. [Google Scholar] [CrossRef]
- Dittmann, E.; Gugger, M.; Sivonen, K.; Fewer, D.P. Natural Product Biosynthetic Diversity and Comparative Genomics of the Cyanobacteria. Trends Microbiol. 2015, 23, 642–652. [Google Scholar] [CrossRef] [PubMed]
- Pattanaik, B.; Lindberg, P. Terpenoids and Their Biosynthesis in Cyanobacteria. Life 2015, 5, 269–293. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Parihar, P.; Singh, M.; Bajguz, A.; Kumar, J.; Singh, S.; Singh, V.P.; Prasad, S.M. Uncovering potential applications of cyanobacteria and algal metabolites in biology, agriculture and medicine: Current status and future prospects. Front. Microbiol. 2017, 8, 515. [Google Scholar] [CrossRef] [PubMed]
- Singh, J.S.; Kumar, A.; Rai, A.N.; Singh, D.P. Cyanobacteria: A precious bio-resource in agriculture, ecosystem, and environmental sustainability. Front. Microbiol. 2016, 7, 529. [Google Scholar] [CrossRef]
- Gerwick, W.H.; Fenner, A.M. Drug Discovery from Marine Microbes. Microb. Ecol. 2013, 65, 800–806. [Google Scholar] [CrossRef] [PubMed]
- Shishido, T.K.; Humisto, A.; Jokela, J.; Liu, L.; Wahlsten, M.; Tamrakar, A.; Fewer, D.P.; Permi, P.; Andreote, A.P.D.; Fiore, M.F.; et al. Antifungal compounds from cyanobacteria. Mar. Drugs 2015, 13, 2124–2140. [Google Scholar] [CrossRef] [PubMed]
- Vijayakumar, S.; Menakha, M. Pharmaceutical applications of cyanobacteria-A review. J. Acute Med. 2015, 5, 15–23. [Google Scholar] [CrossRef]
- Ciferri, O.; Tiboni, O. The Biochemistry and Industrial Potential of Spirulina. Annu. Rev. Microbiol. 1985, 39, 503–526. [Google Scholar] [CrossRef]
- Abdulqader, G.; Barsanti, L.; Tredici, M.R. Harvest of Arthrospira platensis from Lake Kossorom (Chad) and its household usage among the Kanembu. J. Appl. Phycol. 2000, 12, 493–498. [Google Scholar] [CrossRef]
- Lau, N.-S.; Matsui, M.; Abdullah, A.A.-A. Cyanobacteria: Photoautotrophic Microbial Factories for the Sustainable Synthesis of Industrial Products. BioMed Res. Int. 2015, 2015, 754934. [Google Scholar] [CrossRef]
- Janssen, E.M.-L. Cyanobacterial peptides beyond microcystins—A review on co-occurrence, toxicity, and challenges for risk assessment. Water Res. 2019, 151, 488–499. [Google Scholar] [CrossRef]
- Boudreau, P.D.; Byrum, T.; Liu, W.T.; Dorrestein, P.C.; Gerwick, W.H. Viequeamide A, a cytotoxic member of the kulolide superfamily of cyclic depsipeptides from a marine button cyanobacterium. J. Nat. Prod. 2012, 75, 1560–1570. [Google Scholar] [CrossRef]
- Komarek, J.; Kastovsky, J.; Mares, J.; Johansen, J.R. Taxonomic classification of cyanoprokaryotes (cyanobacterial genera) 2014, using a polyphasic approach. Preslia 2014, 86, 295–335. [Google Scholar]
- Soares, A.R.; Engene, N.; Gunasekera, S.P.; Sneed, J.M.; Paul, V.J. Carriebowlinol, an Antimicrobial Tetrahydroquinolinol from an Assemblage of Marine Cyanobacteria Containing a Novel Taxon. J. Nat. Prod. 2014, 78, 534–538. [Google Scholar] [CrossRef]
- Vining, O.B.; Medina, R.A.; Mitchell, E.A.; Videau, P.; Li, D.; Serrill, J.D.; Kelly, J.X.; Gerwick, W.H.; Proteau, P.J.; Ishmael, J.E.; et al. Depsipeptide companeramides from a panamanian marine cyanobacterium associated with the coibamide producer. J. Nat. Prod. 2015, 78, 413–420. [Google Scholar] [CrossRef]
- Nogle, L.M.; Williamson, R.T.; Gerwick, W.H. Somamides A and B, two new depsipeptide analogues of dolastatin 13 from a Fijian cyanobacterial assemblage of Lyngbya majuscula and Schizothrix species. J. Nat. Prod. 2001, 64, 716–719. [Google Scholar] [CrossRef]
- Iwasaki, A.; Sumimoto, S.; Ohno, O.; Suda, S.; Suenaga, K. Kurahamide, a cyclic depsipeptide analog of dolastatin 13 from a marine cyanobacterial assemblage of Lyngbya sp. Bull. Chem. Soc. Jpn. 2014, 87, 609–613. [Google Scholar] [CrossRef]
- Harrigan, G.G.; Yoshida, W.Y.; Moore, R.E.; Nagle, D.G.; Park, P.U.; Biggs, J.; Paul, V.J.; Mooberry, S.L.; Corbett, T.H.; Valeriote, F.A. Isolation, structure determination, and biological activity of dolastatin 12 and Lyngbyastatin 1 from Lyngbya majuscula/Schizothrix calcicola cyanobacterial assemblages. J. Nat. Prod. 1998, 61, 1221–1225. [Google Scholar] [CrossRef]
- Pereira, A.; Cao, Z.; Murray, T.F.; Gerwick, W.H. Hoiamide A, a Sodium Channel Activator of Unusual Architecture from a Consortium of Two Papua New Guinea Cyanobacteria. Chem. Biol. 2009, 16, 893–906. [Google Scholar] [CrossRef]
- Graber, M.A.; Gerwick, W.H. Kalkipyrone, a toxic gamma-pyrone from an assemblage of the marine cyanobacteria Lyngbya majuscula and Tolypothrix sp. J. Nat. Prod. 1998, 61, 677–680. [Google Scholar] [CrossRef]
- Pereira, A.R.; Cao, Z.; Engene, N.; Soria-Mercado, I.E.; Murray, T.F.; Gerwick, W.H. Palmyrolide A, an unusually stabilized neuroactive macrolide from Palmyra Atoll cyanobacteria. Org. Lett. 2010, 12, 4490–4493. [Google Scholar] [CrossRef]
- Pereira, A.R.; Etzbach, L.; Engene, N.; Müller, R.; Gerwick, W.H. Molluscicidal metabolites from an assemblage of Palmyra Atoll cyanobacteria. J. Nat. Prod. 2011, 74, 1175–1181. [Google Scholar] [CrossRef]
- Nogle, L.M.; Gerwick, W.H. Somocystinamide A, a Novel Cytotoxic Disulfide Dimer from a Fijian Marine Cyanobacterial Mixed Assemblage. Org. Lett. 2002, 4, 1095–1098. [Google Scholar] [CrossRef]
- Williamson, R.T.; Singh, I.P.; Gerwick, W.H. Taveuniamides: New chlorinated toxins from a mixed assemblage of marine cyanobacteria. Tetrahedron 2004, 60, 7025–7033. [Google Scholar] [CrossRef]
- Harrigan, G.G.; Luesch, H.; Yoshida, W.Y.; Moore, R.E.; Nagle, D.G.; Biggs, J.; Park, P.U.; Paul, V.J. Tumonoic acids, novel metabolites from a cyanobacterial assemblage of Lyngbya majuscula and Schizothrix calcicola. J. Nat. Prod. 1999, 62, 464–467. [Google Scholar] [CrossRef]
- Sitachitta, N.; Williamson, R.T.; Gerwick, W.H. Yanucamides A and B, two new depsipeptides from an assemblage of the marine cyanobacteria Lyngbya majuscula and Schizothrix species. J. Nat. Prod. 2000, 63, 197–200. [Google Scholar] [CrossRef]
- Osborne, N.J.; Shaw, G.R.; Webb, P.M. Health effects of recreational exposure to Moreton Bay, Australia waters during a Lyngbya majuscula bloom. Environ. Int. 2007, 33, 309–314. [Google Scholar] [CrossRef]
- Osborne, N.J.; Shaw, G.R. Dermatitis associated with exposure to a marine cyanobacterium during recreational water exposure. BMC Dermatol. 2008, 8, 5. [Google Scholar] [CrossRef]
- Engene, N.; Cameron Coates, R.; Gerwick, W.H. 16S Rrna gene heterogeneity in the filamentous marine cyanobacterial genus Lyngbya. J. Phycol. 2010, 46, 591–601. [Google Scholar] [CrossRef]
- Engene, N.; Rottacker, E.C.; Kaštovský, J.; Byrum, T.; Choi, H.; Ellisman, M.H.; Komárek, J.; Gerwick, W.H. Moorea producens gen. nov., sp. nov. and Moorea bouillonii comb. nov., tropical marine cyanobacteria rich in bioactive secondary metabolites. Int. J. Syst. Evol. Microbiol. 2012, 62, 1171–1178. [Google Scholar] [CrossRef]
- Engene, N.; Paul, V.J.; Byrum, T.; Gerwick, W.H.; Thor, A.; Ellisman, M.H. Five chemically rich species of tropical marine cyanobacteria of the genus Okeania gen. nov. (Oscillatoriales, Cyanoprokaryota). J. Phycol. 2013, 49, 1095–1106. [Google Scholar] [CrossRef]
- Komarek, J.; Zapomelova, E.; Smarda, J.; Kopecky, J.; Rejmankova, E.; Woodhouse, J.; Neilan, B.A.; Komarkova, J. Polyphasic evaluation of Limnoraphis robusta, a water-bloom forming cyanobacterium from Lake Atitlan, Guatemala, with a description of Limnoraphis gen. nov. Fottea 2013, 13, 39–52. [Google Scholar] [CrossRef]
- Mcgregor, G.B.; Sendall, B.C. Phylogeny and toxicology of Lyngbya wollei (Cyanobacteria, Oscillatoriales) from north-eastern Australia, with a description of Microseira gen. nov. J. Phycol. 2015, 51, 109–119. [Google Scholar] [CrossRef]
- Engene, N.; Choi, H.; Esquenazi, E.; Rottacker, E.C.; Ellisman, M.H.; Dorrestein, P.C.; Gerwick, W.H. Underestimated biodiversity as a major explanation for the perceived rich secondary metabolite capacity of the cyanobacterial genus Lyngbya. Environ. Microbiol. 2011, 13, 1601–1610. [Google Scholar] [CrossRef]
- Shih, P.M.; Wu, D.; Latifi, A.; Axen, S.D.; Fewer, D.P.; Talla, E.; Calteau, A.; Cai, F.; Tandeau de Marsac, N.; Rippka, R.; et al. Improving the coverage of the cyanobacterial phylum using diversity-driven genome sequencing. Proc. Natl. Acad. Sci. USA 2013, 110, 1053–1058. [Google Scholar] [CrossRef]
- Médigue, C.; Calteau, A.; Cruveiller, S.; Gachet, M.; Gautreau, G.; Josso, A.; Lajus, A.; Langlois, J.; Pereira, H.; Planel, R.; et al. MicroScope—An integrated resource for community expertise of gene functions and comparative analysis of microbial genomic and metabolic data. Brief. Bioinform. 2017. [Google Scholar] [CrossRef]
- Moore, R.E.; Blackman, A.J.; Cheuk, C.E.; Mynderse, J.S.; Matsumoto, G.K.; Clardy, J.; Woodard, R.W.; Craig, J.C. Absolute Stereochemistries of the Aplysiatoxins and Oscillatoxin A. J. Org. Chem. 1984, 49, 2484–2489. [Google Scholar] [CrossRef]
- Cardellina, J.H.; Marner, F.-J.; Moore, R.E. Seaweed dermatitis: Structure of Lyngbyatoxin A. Science 1979, 204, 193–195. [Google Scholar] [CrossRef]
- Ohtani, I.; Moore, R.E.; Runnegar, M.T.C. Cylindrospermopsin: A Potent Hepatotoxin from the Blue-Green Alga Cylindrospermopsis raciborskii. J. Am. Chem. Soc. 1992, 114, 7941–7942. [Google Scholar] [CrossRef]
- MacKintosh, C.; Beattie, K.A.; Klumpp, S.; Cohen, P.; Codd, G.A. Cyanobacterial microcystin-LR is a potent and specific inhibitor of protein phosphatases 1 and 2A from both mammals and higher plants. FEBS Lett. 1990, 264, 187–192. [Google Scholar] [CrossRef]
- Jokela, J.; Heinilä, L.M.P.; Shishido, T.K.; Wahlsten, M.; Fewer, D.P.; Fiore, M.F.; Wang, H.; Haapaniemi, E.; Permi, P.; Sivonen, K. Production of high amounts of hepatotoxin nodularin and new protease inhibitors pseudospumigins by the brazilian benthic Nostoc sp. CENA543. Front. Microbiol. 2017, 8, 1963. [Google Scholar] [CrossRef] [PubMed]
- Stahl, W.; Sies, H. Antioxidant activity of carotenoids. Mol. Asp. Med. 2003, 24, 345–351. [Google Scholar] [CrossRef]
- Lanfer-Marquez, U.M.; Barros, R.M.C.; Sinnecker, P. Antioxidant activity of chlorophylls and their derivatives. Food Res. Int. 2005, 38, 885–891. [Google Scholar] [CrossRef]
- Jain, S.; Prajapat, G.; Abrar, M.; Ledwani, L.; Singh, A.; Agrawal, A. Cyanobacteria as efficient producers of mycosporine-like amino acids. J. Basic Microbiol. 2017, 57, 715–727. [Google Scholar] [CrossRef]
- Romay, C.; Armesto, J.; Remirez, D.; González, R.; Ledon, N.; García, I. Antioxidant and anti-inflammatory properties of C-phycocyanin from blue-green algae. Inflamm. Res. 1998, 47, 36–41. [Google Scholar] [CrossRef] [PubMed]
- Balunas, M.J.; Grosso, M.F.; Villa, F.A.; Engene, N.; McPhail, K.L.; Tidgewell, K.; Pineda, L.M.; Gerwick, L.; Spadafora, C.; Kyle, D.E.; et al. Coibacins A-D, antileishmanial marine cyanobacterial polyketides with intriguing biosynthetic origins. Org. Lett. 2012, 14, 3878–3881. [Google Scholar] [CrossRef]
- Choi, H.; Mascuch, S.J.; Villa, F.A.; Byrum, T.; Teasdale, M.E.; Smith, J.E.; Preskitt, L.B.; Rowley, D.C.; Gerwick, L.; Gerwick, W.H. Honaucins A-C, potent inhibitors of inflammation and bacterial quorum sensing: Synthetic derivatives and structure-activity relationships. Chem. Biol. 2012, 19, 589–598. [Google Scholar] [CrossRef] [PubMed]
- Kapuścik, A.; Hrouzek, P.; Kuzma, M.; Bártová, S.; Novák, P.; Jokela, J.; Pflüger, M.; Eger, A.; Hundsberger, H.; Kopecký, J. Novel Aeruginosin-865 from Nostoc sp. as a Potent Anti-inflammatory Agent. ChemBioChem 2013, 14, 2329–2337. [Google Scholar] [CrossRef]
- Shaala, L.A.; Youssef, D.T.A.; McPhail, K.L.; Elbandy, M. Malyngamide 4, a new lipopeptide from the Red Sea marine cyanobacterium Moorea producens (formerly Lyngbya majuscula). Phytochem. Lett. 2013, 6, 183–188. [Google Scholar] [CrossRef]
- Romay, C.; Gonzalez, R.; Ledon, N.; Remirez, D.; Rimbau, V. C-Phycocyanin: A Biliprotein with Antioxidant, Anti-Inflammatory and Neuroprotective Effects. Curr. Protein Pept. Sci. 2003, 4, 207–216. [Google Scholar] [CrossRef]
- Rastogi, R.P.; Sonani, R.R.; Madamwar, D. Cyanobacterial Sunscreen Scytonemin: Role in Photoprotection and Biomedical Research. Appl. Biochem. Biotechnol. 2015, 176, 1551–1563. [Google Scholar] [CrossRef]
- Prinsep, M.R.; Thomson, R.A.; West, M.L.; Wylie, B.L. Tolypodiol, an antiinflammatory diterpenoid from the cyanobacterium Tolypothrix nodosa. J. Nat. Prod. 1996, 59, 786–788. [Google Scholar] [CrossRef]
- Hayashi, T.; Hayashi, K.; Maeda, M.; Kojima, I. Calcium spirulan, an inhibitor of enveloped virus replication, from a blue-green alga Spirulina platensis. J. Nat. Prod. 1996, 59, 83–87. [Google Scholar] [CrossRef]
- Entzeroth, M.; Moore, R.E.; Niemczura, W.P.; Patterson, G.M.L.; Shoolery, J.N. O-Acetyl-O-butyryl-O-carbamoyl-O,O-dimethyl-.alpha.-cyclodextrins from the cyanophyte Tolypothrix byssoidea. J. Org. Chem. 1986, 51, 5307–5310. [Google Scholar] [CrossRef]
- Thammana, S.; Suzuki, H.; Lobkovsky, E.; Clardy, J.; Shimizu, Y. Isolation and structure assignment of an iminotetrasaccharide from a cultured filamentous cyanobacterium Anabaena sp. J. Nat. Prod. 2006, 69, 365–368. [Google Scholar] [CrossRef]
- Sueyoshi, K.; Kaneda, M.; Sumimoto, S.; Oishi, S.; Fujii, N.; Suenaga, K.; Teruya, T. Odoamide, a cytotoxic cyclodepsipeptide from the marine cyanobacterium Okeania sp. Tetrahedron 2016, 72, 5472–5478. [Google Scholar] [CrossRef]
- Mo, S.; Krunic, A.; Pegan, S.D.; Franzblau, S.G.; Orjala, J. An antimicrobial guanidine-bearing sesterterpene from the cultured cyanobacterium Scytonema sp. J. Nat. Prod. 2009, 72, 2043–2045. [Google Scholar] [CrossRef]
- Sung, S.Y.; Sin, L.T.; Tee, T.T.; Bee, S.T.; Rahmat, A.R.; Rahman, W.A.W.A.; Tan, A.C.; Vikhraman, M. Antimicrobial agents for food packaging applications. Trends Food Sci. Technol. 2013, 33, 110–123. [Google Scholar] [CrossRef]
- Abushelaibi, A.A.; Al Shamsi, M.S.; Afifi, H.S. Use of antimicrobial agents in food processing systems. Recent Pat. Food. Nutr. Agric. 2012, 4, 2–7. [Google Scholar] [CrossRef]
- Sturdy, M.; Krunic, A.; Cho, S.; Franzblau, S.; Orjala, J. Eucapsitrione, an anti- mycobacterium tuberculosis anthraquinone derivative from the cultured freshwater cyanobacterium Eucapsis sp. J. Nat. Prod. 2010, 73, 1441–1443. [Google Scholar] [CrossRef]
- Nogle, L.M.; Gerwick, W.H. Isolation of four new cyclic depsipeptides, antanapeptins A-D, and dolastatin 16 from a Madagascan collection of Lyngbya majuscula. J. Nat. Prod. 2002, 65, 21–24. [Google Scholar] [CrossRef]
- Kosalec, I.; Kremer, D.; Locatelli, M.; Epifano, F.; Genovese, S.; Carlucci, G.; Randić, M.; Zovko Končić, M. Anthraquinone profile, antioxidant and antimicrobial activity of bark extracts of Rhamnus alaternus, R. fallax, R. intermedia and R. pumila. Food Chem. 2013, 136, 335–341. [Google Scholar] [CrossRef]
- Yang, E.J.; Kim, S.H.; Lee, K.Y.; Song, K.S. Neuroprotective and anti-neuroinflammatory activities of anthraquinones isolated from Photorhabdus temperata culture broth. J. Microbiol. Biotechnol. 2018, 28, 12–21. [Google Scholar] [CrossRef]
- Park, J.G.; Kim, S.C.; Kim, Y.H.; Yang, W.S.; Kim, Y.; Hong, S.; Kim, K.H.; Yoo, B.C.; Kim, S.H.; Kim, J.H.; et al. Anti-Inflammatory and Antinociceptive Activities of Anthraquinone-2-Carboxylic Acid. Mediat. Inflamm. 2016, 2016, 1903849. [Google Scholar] [CrossRef]
- Yen, G.C.; Duh, P.D.; Chuang, D.Y. Antioxidant activity of anthraquinones and anthrone. Food Chem. 2000, 70, 437–441. [Google Scholar] [CrossRef]
- Reese, M.T.; Gulavita, N.K.; Nakao, Y.; Hamann, M.T.; Yoshida, W.Y.; Coval, S.J.; Scheuer, P.J. Kulolide: A Cytotoxic Depsipeptide from a Cephalaspidean. J. Am. Chem. Soc. 1996, 7863, 11081–11084. [Google Scholar] [CrossRef]
- Bunyajetpong, S.; Yoshida, W.Y.; Sitachitta, N.; Kaya, K. Trungapeptins A-C, cyclodepsipeptides from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2006, 69, 1539–1542. [Google Scholar] [CrossRef]
- Tripathi, A.; Puddick, J.; Prinsep, M.R.; Lee, P.P.F.; Tan, L.T. Hantupeptins B and C, cytotoxic cyclodepsipeptides from the marine cyanobacterium Lyngbya majuscula. Phytochemistry 2010, 71, 307–311. [Google Scholar] [CrossRef]
- Malloy, K.L. Structure Elucidation of Biomedically Relevant Marine Cyanobacterial Natural Products. Ph.D. Thesis, UC San Diego, San Diego, CA, USA, 2011. [Google Scholar]
- Mevers, E.; Liu, W.T.; Engene, N.; Mohimani, H.; Byrum, T.; Pevzner, P.A.; Dorrestein, P.C.; Spadafora, C.; Gerwick, W.H. Cytotoxic veraguamides, alkynyl bromide-containing cyclic depsipeptides from the marine cyanobacterium cf. Oscillatoria margaritifera. J. Nat. Prod. 2011, 74, 928–936. [Google Scholar] [CrossRef]
- Montaser, R.; Paul, V.J.; Luesch, H. Pitipeptolides C-F, antimycobacterial cyclodepsipeptides from the marine cyanobacterium Lyngbya majuscula from Guam. Phytochemistry 2011, 72, 2068–2074. [Google Scholar] [CrossRef]
- Salvador, L.A.; Biggs, J.S.; Paul, V.J.; Luesch, H. Veraguamides A-G, cyclic hexadepsipeptides from a dolastatin 16-producing cyanobacterium Symploca cf. hydnoides from Guam. J. Nat. Prod. 2011, 74, 917–927. [Google Scholar] [CrossRef]
- Almaliti, J.; Malloy, K.L.; Glukhov, E.; Spadafora, C.; Gutiérrez, M.; Gerwick, W.H. Dudawalamides A-D, Antiparasitic Cyclic Depsipeptides from the Marine Cyanobacterium Moorea producens. J. Nat. Prod. 2017, 80, 1827–1836. [Google Scholar] [CrossRef]
- Iwasaki, A.; Shiota, I.; Sumimoto, S.; Matsubara, T.; Sato, T.; Suenaga, K. Kohamamides A, B, and C, Cyclic Depsipeptides from an Okeania sp. Marine Cyanobacterium. J. Nat. Prod. 2017, 80, 1948–1952. [Google Scholar] [CrossRef]
- Luesch, H.; Pangilinan, R.; Yoshida, W.Y.; Moore, R.E.; Paul, V.J. Pitipeptolides A and B, new cyclodepsipeptides from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2001, 64, 304–307. [Google Scholar] [CrossRef]
- Medina, R.A. Biologically Active Cyclic Depsipeptides from Marine Cyanobacteria. Ph.D. Thesis, Oregon State University, Corvallis, OR, USA, 2009; 160p. [Google Scholar]
- Costa, M.S.; Rego, A.; Ramos, V.; Afonso, T.B.; Freitas, S.; Preto, M.; Lopes, V.; Vasconcelos, V.; Magalhães, C.; Leaõ, P.N. The conifer biomarkers dehydroabietic and abietic acids are widespread in Cyanobacteria. Sci. Rep. 2016, 6, 23436. [Google Scholar] [CrossRef] [PubMed]
- Acuña, U.M.; Zi, J.; Orjala, J.; Carcache de Blanco, E.J. Ambiguine I Isonitrile from Fischerella ambigua Induces Caspase-Independent Cell Death in MCF-7 Hormone Dependent Breast Cancer Cells. Int. J. Cancer Res. 2015, 49, 1655–1662. [Google Scholar]
- Becher, P.G.; Jüttner, F. Insecticidal compounds of the biofilm-forming cyanobacterium Fischerella sp. (ATCC 43239). Environ. Toxicol. 2005, 20, 363–372. [Google Scholar] [CrossRef]
- Becher, P.G.; Keller, S.; Jung, G.; Süssmuth, R.D.; Jüttner, F. Insecticidal activity of 12-epi-hapalindole J isonitrile. Phytochemistry 2007, 68, 2493–2497. [Google Scholar] [CrossRef]
- Cagide, E.; Becher, P.G.; Louzao, M.C.; Espiña, B.; Vieytes, M.R.; Jüttner, F.; Botana, L.M. Hapalindoles from the cyanobacterium Fischerella: Potential sodium channel modulators. Chem. Res. Toxicol. 2014, 27, 1696–1706. [Google Scholar] [CrossRef]
- Etchegaray, A.; Rabello, E.; Dieckmann, R.; Moon, D.H.; Fiore, M.F.; Von Döhren, H.; Tsai, S.M.; Neilan, B.A. Algicide production by the filamentous cyanobacterium Fischerella sp. CENA 19. J. Appl. Phycol. 2004, 16, 237–243. [Google Scholar] [CrossRef]
- Hillwig, M.L.; Zhu, Q.; Liu, X. Biosynthesis of Ambiguine Indole Alkaloids in Cyanobacterium Fischerella ambigua. ACS Chem. Biol. 2014, 9, 372–377. [Google Scholar] [CrossRef]
- Hillwig, M.L.; Fuhrman, H.A.; Ittiamornkul, K.; Sevco, T.J.; Kwak, D.H.; Liu, X. Identification and characterization of a welwitindolinone alkaloid biosynthetic gene cluster in the stigonematalean cyanobacterium Hapalosiphon welwitschii. ChemBioChem 2014, 15, 665–669. [Google Scholar] [CrossRef]
- Huber, U.; Moore, R.E.; Patterson, G.M.L. Isolation of a nitrile-containing indole alkaloid from the terrestrial blue-green alga Hapalosiphon delicatulus. J. Nat. Prod. 1998, 61, 1304–1306. [Google Scholar] [CrossRef]
- Kim, H.; Lantvit, D.; Hwang, C.H.; Kroll, D.J.; Swanson, S.M.; Franzblau, S.G.; Orjala, J. Indole alkaloids from two cultured cyanobacteria, Westiellopsis sp. and Fischerella muscicola. Bioorg. Med. Chem. 2012, 20, 5290–5295. [Google Scholar] [CrossRef]
- Kim, H.; Krunic, A.; Lantvit, D.; Shen, Q.; Kroll, D.J.; Swanson, S.M.; Orjala, J. Nitrile-containing fischerindoles from the cultured cyanobacterium Fischerella sp. Tetrahedron 2012, 68, 3205–3209. [Google Scholar] [CrossRef]
- Klein, D.; Daloze, D.; Braekman, J.C.; Hoffmann, L.; Demoulin, V. New hapalindoles from the cyanophyte Hapalosiphon laingii. J. Nat. Prod. 1995, 58, 1781–1785. [Google Scholar] [CrossRef]
- Koodkaew, I.; Sunohara, Y.; Matsuyama, S.; Matsumoto, H. Isolation of ambiguine D isonitrile from Hapalosiphon sp. and characterization of its phytotoxic activity. Plant Growth Regul. 2012, 68, 141–150. [Google Scholar] [CrossRef]
- Micallef, M.L.; Sharma, D.; Bunn, B.M.; Gerwick, L.; Viswanathan, R.; Moffitt, M.C. Comparative analysis of hapalindole, ambiguine and welwitindolinone gene clusters and reconstitution of indole-isonitrile biosynthesis from cyanobacteria. BMC Microbiol. 2014, 14, 213. [Google Scholar] [CrossRef]
- Micallef, M.L.; D’Agostino, P.M.; Sharma, D.; Viswanathan, R.; Moffitt, M.C. Genome mining for natural product biosynthetic gene clusters in the Subsection V cyanobacteria. BMC Genom. 2015, 16, 669. [Google Scholar] [CrossRef]
- Mo, S.; Krunic, A.; Chlipala, G.; Orjala, J. Antimicrobial ambiguine isonitriles from the cyanobacterium Fischerella ambigua. J. Nat. Prod. 2009, 72, 894–899. [Google Scholar] [CrossRef]
- Mo, S.; Krunic, A.; Santarsiero, B.D.; Franzblau, S.G.; Orjala, J. Hapalindole-related alkaloids from the cultured cyanobacterium Fischerella ambigua. Phytochemistry 2010, 71, 2116–2123. [Google Scholar] [CrossRef]
- Moore, R.E.; Cheuk, C.; Patterson, G.M.L. Hapalindoles: New Alkaloids from the Blue-Green Alga Hapalosiphon fontinalis. J. Am. Chem. Soc. 1984, 106, 6456–6457. [Google Scholar] [CrossRef]
- Moore, R.E.; Yang, X.Q.G.; Patterson, G.M.L. Fontonamide and Anhydrohapaloxindole A, Two New Alkaloids from the Blue-Green Alga Hapalosiphon fontinalis. J. Org. Chem. 1987, 52, 3773–3777. [Google Scholar] [CrossRef]
- Moore, R.E.; Cheuk, C.; Yang, X.Q.G.; Patterson, G.M.L.; Bonjouklian, R.; Smitka, T.A.; Mvnderse, J.S.; Foster, R.S.; Jones, N.D.; Swartzendruber, J.K.; et al. Hapalindoles, Antibacterial and Antimycotic Alkaloids from the Cyanophyte Hapalosiphon fontinalis. J. Org. Chem. 1987, 52, 1036–1043. [Google Scholar] [CrossRef]
- Moore, R.E.; Yang, X.Q.G.; Patterson, G.M.L.; Bonjouklian, R.; Smitka, T.A. Hapalonamides and other oxidized hapalindoles from Hapalosiphon fontinalis. Phytochemistry 1989, 28, 1565–1567. [Google Scholar] [CrossRef]
- Park, A.; Moore, R.E.; Patterson, G.M. Fischerindole L, a new isonitrile from the terrestrial blue-green alga Fischerella muscicola. Tetrahedron Lett. 1992, 33, 3257–3260. [Google Scholar] [CrossRef]
- Raveh, A.; Carmeli, S. Antimicrobial ambiguines from the cyanobacterium Fischerella sp. collected in Israel. J. Nat. Prod. 2007, 70, 196–201. [Google Scholar] [CrossRef]
- Schwartz, R.E.; Hirsch, C.F.; Pettibone, D.J.; Zink, D.L.; Springer, J.P. Unusual Cyclopropane-Containing Hapalindolinones from a Cultured Cyanobacterium. J. Org. Chem. 1987, 52, 3704–3706. [Google Scholar] [CrossRef]
- Smitka, T.A.; Bonjouklian, R.; Doolin, L.; Jones, N.D.; Deeter, J.B.; Yoshida, W.Y.; Prinsep, M.R.; Moore, R.E.; Patterson, G.M.L. Ambiguine Isonitriles, Fungicidal Hapalindole-Type Alkaloids from Three Genera of Blue-Green Algae Belonging to the Stigonemataceae. J. Org. Chem. 1992, 57, 857–861. [Google Scholar] [CrossRef]
- Stratmann, K.; Moore, R.E.; Patterson, G.M.L.; Bonjouklian, R.; Deeter, J.B.; Shaffer, S.; Smitka, T.A.; Smith, C.D. Welwitindolinones, Unusual Alkaloids from the Blue-Green Algae Hapalosiphon welwitschii and Westiella intricata. Relationship to Fischerindoles and Hapalinodoles. J. Am. Chem. Soc. 1994, 116, 9935–9942. [Google Scholar] [CrossRef]
- Walton, K.; Gantar, M.; Gibbs, P.D.L.; Schmale, M.C.; Berry, J.P. Indole alkaloids from Fischerella inhibit vertebrate development in the zebrafish (Danio rerio) embryo model. Toxins 2014, 6, 3568–3581. [Google Scholar] [CrossRef]
- Mason, C.P.; Edwards, K.R.; Carlson, R.E.; Pignatello, J.; Gleason, F.K.; Wood, J.M. Isolation of Chlorine-Containing Antibiotic from the Freshwater Cyanobacterium Scytonema hofmanni. Science 1982, 215, 400–402. [Google Scholar] [CrossRef]
- Gromov, B.V.; Vepritskiy, A.A.; Titova, N.N.; Mamkayeva, K.A.; Alexandrova, O.V. Production of the antibiotic cyanobacterin LU-1 by Nostoc linckia CALU 892 (cyanobacterium). J. Appl. Phycol. 1991, 3, 55–59. [Google Scholar] [CrossRef]
- Gleason, F.K.; Case, D.E. Activity of the natural algicide, cyanobacterin, on angiosperms. Plant Physiol. 1986, 80, 834–837. [Google Scholar] [CrossRef]
- Ishibashi, F.; Park, S.; Kusano, T.; Kuwano, K. Synthesis and algicidal activity of (+)-cyanobacterin and its stereoisomer. Biosci. Biotechnol. Biochem. 2005, 69, 391–396. [Google Scholar] [CrossRef][Green Version]
- Gross, E.M.; Wolk, C.P.; Jüttner, F. Fischerellin, a new allelochemical from the freshwater cyanobacterium Fischerella muscicola. J. Phycol. 1991, 27, 686–692. [Google Scholar] [CrossRef]
- Hagmann, L.; Jüttner, F. Fischerellin A, a novel photosystem-II-inhibiting allelochemical of the cyanobacterium Fischerella muscicola with antifungal and herbicidal activity. Tetrahedron Lett. 1996, 37, 6539–6542. [Google Scholar] [CrossRef]
- Papke, U.; Gross, E.M.; Francke, W. Isolation, identification and determination of the absolute configuration of Fischerellin B. A new algicide from the freshwater cyanobacterium Fischerella muscicola (Thuret). Tetrahedron Lett. 1997, 38, 379–382. [Google Scholar] [CrossRef]
- Srivastava, V.C.; Manderson, G.J.; Bhamidimarri, R. Inhibitory metabolites production by the cyanobacterium Fischerella muscicola. Microbiol. Res. 1999, 153, 309–317. [Google Scholar] [CrossRef]
- Ogino, J.; Moore, R.E.; Patterson, G.M.L.; Smith, C.D. Dendroamides, new cyclic hexapeptides from a blue-green alga. Multidrug- resistance reversing activity of dendroamide A. J. Nat. Prod. 1996, 59, 581–586. [Google Scholar] [CrossRef]
- Todorova, A.K.; Jüttner, F.; Linden, A.; Plüiss, T.; von Philipsborn, W. Nostocyclamide: A New Macrocyclic, Thiazole-Containing Allelochemical from Nostoc sp. 31 (Cyanobacteria). J. Org. Chem. 1995, 60, 7891–7895. [Google Scholar] [CrossRef]
- Prinsep, M.R.; Moore, R.E.; Levine, I.A.; Patterson, G.M.L. Westiellamide, a bistratamide-related cyclic peptide from the blue-green alga Westiellopsis prolifica. J. Nat. Prod. 1992, 55, 140–142. [Google Scholar] [CrossRef]
- Admi, V.; Afek, U.; Carmeli, S. Raocyclamides A and B, novel cyclic hexapeptides isolated from the cyanobacterium Oscillatoria raoi. J. Nat. Prod. 1996, 59, 396–399. [Google Scholar] [CrossRef]
- Banker, R.; Carmeli, S. Tenuecyclamides A−D, Cyclic Hexapeptides from the Cyanobacterium Nostoc spongiaeforme var. tenue. J. Nat. Prod. 1998, 61, 1248–1251. [Google Scholar] [CrossRef]
- Jüttner, F.; Todorova, A.K.; Walch, N.; Von Philipsborn, W. Nostocyclamide M: A cyanobacterial cyclic peptide with allelopathic activity from Nostoc 31. Phytochemistry 2001, 57, 613–619. [Google Scholar] [CrossRef]
- Falch, B.S.; König, G.M.; Wright, A.D.; Sticher, O.; Röegger, H.; Bernardinelli, G. Ambigol A and B: New Biologically Active Polychlorinated Aromatic Compounds from the Terrestrial Blue-Green Alga Fischerella ambigua. J. Org. Chem. 1993, 58, 6570–6575. [Google Scholar] [CrossRef]
- Wright, A.D.; Papendorf, O.; König, G.M. Ambigol C and 2, 4 dichlorobenzoic acid, natural products produced by terrestrial cyanobacterium Fischerella ambigua. J. Nat. Prod. 2005, 68, 459–461. [Google Scholar] [CrossRef]
- An, T.; Kumar, T.K.S.; Wang, M.; Liu, L.; Lay, J.O.; Liyanage, R.; Berry, J.; Gantar, M.; Marks, V.; Gawley, R.E.; et al. Structures of pahayokolides A and B, cyclic peptides from a Lyngbya sp. J. Nat. Prod. 2007, 70, 730–735. [Google Scholar] [CrossRef]
- Berry, J.P.; Gantar, M.; Gawley, R.E.; Wang, M.; Rein, K.S. Pharmacology and toxicology of pahayokolide A, a bioactive metabolite from a freshwater species of Lyngbya isolated from the Florida Everglades. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2004, 139, 231–238. [Google Scholar] [CrossRef]
- Dias, F.; Antunes, J.T.; Ribeiro, T.; Azevedo, J.; Vasconcelos, V.; Leão, P.N. Cyanobacterial allelochemicals but not cyanobacterial cells markedly reduce microbial community diversity. Front. Microbiol. 2017, 8, 1495. [Google Scholar] [CrossRef]
- Leao, P.N.; Pereira, A.R.; Liu, W.-T.; Ng, J.; Pevzner, P.A.; Dorrestein, P.C.; Konig, G.M.; Vasconcelos, V.M.; Gerwick, W.H. Synergistic allelochemicals from a freshwater cyanobacterium. Proc. Natl. Acad. Sci. USA 2010, 107, 11183–11188. [Google Scholar] [CrossRef]
- Mehner, C.; Müller, D.; Krick, A.; Kehraus, S.; Löser, R.; Gütschow, M.; Maier, A.; Fiebig, H.H.; Brun, R.; König, G.M. A novel β-amino acid in cytotoxic peptides from the cyanobacterium Tychonema sp. Eur. J. Org. Chem. 2008, 2008, 1732–1739. [Google Scholar] [CrossRef]
- Pergament, I.; Carmeli, S. Schizotrin A; a Nwel Antimicrobial Cyclic Peptide from a Cyanobacterium. Tetrahedron Lett. 1994, 35, 8473–8476. [Google Scholar] [CrossRef]
- Ribeiro, T.; Lemos, F.; Preto, M.; Azevedo, J.; Sousa, M.L.; Leão, P.N.; Campos, A.; Linder, S.; Vitorino, R.; Vasconcelos, V.; et al. Cytotoxicity of portoamides in human cancer cells and analysis of the molecular mechanisms of action. PLoS ONE 2017, 12, e0188817. [Google Scholar] [CrossRef] [PubMed]
- Zainuddin, E.N.; Jansen, R.; Nimtz, M.; Wray, V.; Preisitsch, M.; Lalk, M.; Mundt, S. Lyngbyazothrins A-D, antimicrobial cyclic undecapeptides from the cultured cyanobacterium Lyngbya sp. J. Nat. Prod. 2009, 72, 1373–1378. [Google Scholar] [CrossRef]
- Foster, M.P.; Concepción, G.P.; Caraan, G.B.; Ireland, C.M. Bistratamides C and D. Two New Oxazole-Containing Cyclic Hexapeptides Isolated from a Philippine Lissoclinum bistratum Ascidian. J. Org. Chem. 1992, 57, 6671–6675. [Google Scholar] [CrossRef]
- Ramsay, E.E.; Hogg, P.J.; Dilda, P.J. Mitochondrial metabolism inhibitors for cancer therapy. Pharm. Res. 2011, 28, 2731–2744. [Google Scholar] [CrossRef]
- Bui, T.H.; Wray, V.; Nimtz, M.; Fossen, T.; Preisitsch, M.; Schröder, G.; Wende, K.; Heiden, S.E.; Mundt, S. Balticidins A-D, antifungal hassallidin-like lipopeptides from the Baltic Sea cyanobacterium Anabaena cylindrica Bio33. J. Nat. Prod. 2014, 77, 1287–1296. [Google Scholar] [CrossRef]
- Neuhof, T.; Schmieder, P.; Seibold, M.; Preussel, K.; Von Dö Hren, H. Hassallidin B—Second antifungal member of the Hassallidin family. Bioorg. Med. Chem. Lett. 2006, 16, 4220–4222. [Google Scholar] [CrossRef]
- Neuhof, T.; Schmieder, P.; Preussel, K.; Dieckmann, R.; Pham, H.; Bartl, F.; Von Döhren, H. Hassallidin A, a Glycosylated Lipopeptide with Antifungal Activity from the Cyanobacterium Hassallia sp. J. Nat. Prod. 2005, 68, 695–700. [Google Scholar] [CrossRef]
- Pancrace, C.; Jokela, J.; Sassoon, N.; Ganneau, C.; Desnos-Ollivier, M.; Wahlsten, M.; Humisto, A.; Calteau, A.; Bay, S.; Fewer, D.P.; et al. Rearranged Biosynthetic Gene Cluster and Synthesis of Hassallidin in Planktothrix serta PCC 8927. ACS Chem. Biol. 2017, 12, 1796–1804. [Google Scholar] [CrossRef] [PubMed]
- Vestola, J.; Shishido, T.K.; Jokela, J.; Fewer, D.P.; Aitio, O.; Permi, P.; Wahlsten, M.; Wang, H.; Rouhiainen, L.; Sivonen, K. Hassallidins, antifungal glycolipopeptides, are widespread among cyanobacteria and are the end-product of a nonribosomal pathway. Proc. Natl. Acad. Sci. USA 2014, 111, E1909–E1917. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.; Mevers, E.; Byrum, T.; Valeriote, F.A.; Gerwick, W.H. Lyngbyabellins K-N from two Palmyra atoll collections of the marine cyanobacterium Moorea bouillonii. Eur. J. Org. Chem. 2012, 2012, 5141–5150. [Google Scholar] [CrossRef] [PubMed]
- Han, B.; McPhail, K.L.; Gross, H.; Goeger, D.E.; Mooberry, S.L.; Gerwick, W.H. Isolation and structure of five Lyngbyabellin derivatives from a Papua New Guinea collection of the marine cyanobacterium Lyngbya majuscula. Tetrahedron 2005, 61, 11723–11729. [Google Scholar] [CrossRef]
- Luesch, H.; Yoshida, W.Y.; Moore, R.E.; Paul, V.J.; Mooberry, S.L. Isolation, structure determination, and biological activity of Lyngbyabellin A from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2000, 63, 611–615. [Google Scholar] [CrossRef] [PubMed]
- Marquez, B.L.; Watts, K.S.; Yokochi, A.; Roberts, M.A.; Verdier-Pinard, P.; Jimenez, J.I.; Hamel, E.; Scheuer, P.J.; Gerwick, W.H. Structure and absolute stereochemistry of hectochlorin, a potent stimulator of actin assembly. J. Nat. Prod. 2002, 65, 866–871. [Google Scholar] [CrossRef]
- Milligan, K.E.; Marquez, B.L.; Williamson, R.T.; Gerwick, W.H. Lyngbyabellin B, a toxic and antifungal secondary metabolite from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2000, 63, 1440–1443. [Google Scholar] [CrossRef]
- Ramaswamy, A.V.; Sorrels, C.M.; Gerwick, W.H. Cloning and Biochemical Characterization of the Hectochlorin Biosynthetic Gene Cluster from the Marine Cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2007, 70, 1977–1986. [Google Scholar] [CrossRef]
- Williams, P.G.; Luesch, H.; Yoshida, W.Y.; Moore, R.E.; Paul, V.J. Continuing studies on the cyanobacterium Lyngbya sp.: Isolation and structure determination of 15-norlyngbyapeptin A and Lyngbyabellin D. J. Nat. Prod. 2003, 66, 595–598. [Google Scholar] [CrossRef]
- Matthew, S.; Salvador, L.A.; Schupp, P.J.; Paul, V.J.; Luesch, H. Cytotoxic halogenated macrolides and modified peptides from the apratoxin-producing marine cyanobacterium Lyngbya bouillonii from Guam. J. Nat. Prod. 2010, 73, 1544–1552. [Google Scholar] [CrossRef]
- Adiv, S.; Carmeli, S. Protease inhibitors from Microcystis aeruginosa bloom material collected from the Dalton reservoir, israel. J. Nat. Prod. 2013, 76, 2307–2315. [Google Scholar] [CrossRef]
- Gesner-Apter, S.; Carmeli, S. Three novel metabolites from a bloom of the cyanobacterium Microcystis sp. Tetrahedron 2008, 64, 6628–6634. [Google Scholar] [CrossRef]
- Lifshits, M.; Carmeli, S. Metabolites of Microcystis aeruginosa bloom material from Lake Kinneret, Israel. J. Nat. Prod. 2012, 75, 209–219. [Google Scholar] [CrossRef]
- Marner, F.J.; Moore, R.E.; Hirotsu, K.; Clardy, J. Majusculamides A and B, Two Epimeric Lipodipeptides from Lyngbya majuscula Gomont. J. Org. Chem. 1977, 42, 2815–2819. [Google Scholar] [CrossRef]
- Carter, D.C.; Moore, R.E.; Mynderse, J.S.; Niemczura, W.P.; Todd, J.S. Structure of Majusculamide C, a Cyclic Depsipeptide from Lyngbya majuscula. J. Org. Chem. 1984, 49, 236–241. [Google Scholar] [CrossRef]
- Moore, R.E.; Entzeroth, M. Majusculamide D and deoxymajusculamide D, two cytotoxins from Lyngbya majuscula. Phytochemistry 1988, 27, 3101–3103. [Google Scholar] [CrossRef]
- Koehn, F.E.; Longley, R.E.; Reed, J.K. Microcolins a and b, new immunosuppressive peptides from the blue-green alga Lyngbya majuscula. J. Nat. Prod. 1992, 55, 613–619. [Google Scholar] [CrossRef]
- Meickle, T.; Matthew, S.; Ross, C.; Luesch, H.; Paul, V. Bioassay-guided isolation and identification of desacetylmicrocolin B from Lyngbya cf. polychroa. Planta Med. 2009, 75, 1427–1430. [Google Scholar] [CrossRef]
- Simmons, T.L.; Nogle, L.M.; Media, J.; Valeriote, F.A.; Mooberry, S.L.; Gerwick, W.H. Desmethoxymajusculamide C, a cyanobacterial depsipeptide with potent cytotoxicity in both cyclic and ring-opened forms. J. Nat. Prod. 2009, 72, 1011–1016. [Google Scholar] [CrossRef]
- Tan, L.T.; Goh, B.P.L.; Tripathi, A.; Lim, M.G.; Dickinson, G.H.; Lee, S.S.C.; Teo, S.L.M. Natural antifoulants from the marine cyanobacterium Lyngbya majuscula. Biofouling 2010, 26, 685–695. [Google Scholar] [CrossRef]
- World Health Organisation (WHO). Number of Deaths Due to HIV/AIDS. Available online: https://www.who.int/gho/hiv/epidemic_status/deaths_text/en/ (accessed on 9 April 2019).
- Gupta, D.K.; Kaur, P.; Leong, S.T.; Tan, L.T.; Prinsep, M.R.; Chu, J.J.H. Anti-Chikungunya viral activities of aplysiatoxin-related compounds from the marine cyanobacterium Trichodesmium erythraeum. Mar. Drugs 2014, 12, 115–127. [Google Scholar] [CrossRef] [PubMed]
- Chlipala, G.E.; Sturdy, M.; Krunic, A.; Lantvit, D.D.; Shen, Q.; Porter, K.; Swanson, S.M.; Orjala, J. Cylindrocyclophanes with proteasome inhibitory activity from the Cyanobacterium Nostoc sp. J. Nat. Prod. 2010, 73, 1529–1537. [Google Scholar] [CrossRef]
- Han, B.N.; Liang, T.T.; Keen, L.J.; Fan, T.T.; Zhang, X.D.; Xu, L.; Zhao, Q.; Wang, S.P.; Lin, H.W. Two Marine Cyanobacterial Aplysiatoxin Polyketides, Neo-debromoaplysiatoxin A and B, with K+Channel Inhibition Activity. Org. Lett. 2018, 20, 578–581. [Google Scholar] [CrossRef] [PubMed]
- Rastogi, R.P.; Madamwar, D.; Incharoensakdi, A. Bloom Dynamics of Cyanobacteria and Their Toxins: Environmental Health Impacts and Mitigation Strategies. Front. Microbiol. 2015, 6, 1254. [Google Scholar] [CrossRef]
- Chlipala, G.E.; Tri, P.H.; Van Hung, N.; Krunic, A.; Shim, S.H.; Soejarto, D.D.; Orjala, J. Nhatrangins A and B, aplysiatoxin-related metabolites from the marine cyanobacterium Lyngbya majuscula from Vietnam. J. Nat. Prod. 2010, 73, 784–787. [Google Scholar] [CrossRef] [PubMed]
- Mynderse, J.S.; Moore, R.E.; Kashiwagi, M.; Norton, T.R. Antileukemia activity in the Oscillatoriaceae: Isolation of debromoaplysiatoxin from Lyngbya. Science 1977, 196, 538–540. [Google Scholar] [CrossRef]
- Boyd, M.R.; Gustafson, K.R.; McMahon, J.B.; Shoemaker, R.H.; O’Keefe, B.R.; Mori, T.; Gulakowski, R.J.; Wu, L.; Rivera, M.I.; Laurencot, C.M.; et al. Discovery of cyanovirin-N, a novel human immunodeficiency virus- inactivating protein that binds viral surface envelope glycoprotein gp120: Potential applications to microbicide development. Antimicrob. Agents Chemother. 1997, 41, 1521–1530. [Google Scholar] [CrossRef]
- Dey, B.; Lerner, D.L.; Lusso, P.; Boyd, M.R.; Elder, J.H.; Berger, E.A. Multiple antiviral activities of cyanovirin-N: Blocking of human immunodeficiency virus type 1 gp120 interaction with CD4 and coreceptor and inhibition of diverse enveloped viruses. J. Virol. 2000, 74, 4562–4569. [Google Scholar] [CrossRef]
- Matei, E.; Basu, R.; Furey, W.; Shi, J.; Calnan, C.; Aiken, C.; Gronenborn, A.M. Structure and glycan binding of a new cyanovirin-N homolog. J. Biol. Chem. 2016, 291, 18967–18976. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, K.; Hayashi, T.; Kojima, I. A Natural Sulfated Polysaccharide, Calcium Spirulan, Isolated from Spirulina platensis: In vitro and ex Vivo Evaluation of Anti-Herpes simplex Virus and Anti-Human Immunodeficiency Virus Activities. AIDS Res. Hum. Retrovir. 1996, 12, 1463–1471. [Google Scholar] [CrossRef]
- Mader, J.; Gallo, A.; Schommartz, T.; Handke, W.; Nagel, C.H.; Günther, P.; Brune, W.; Reich, K. Calcium spirulan derived from Spirulina platensis inhibits Herpes simplex virus 1 attachment to human keratinocytes and protects against herpes labialis. J. Allergy Clin. Immunol. 2016, 137, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Wijesekara, I.; Pangestuti, R.; Kim, S.K. Biological activities and potential health benefits of sulfated polysaccharides derived from marine algae. Carbohydr. Polym. 2011, 84, 14–21. [Google Scholar] [CrossRef]
- Baba, M.; Pauwels, R.; Balzarini, J.; Arnout, J.; Desmyter, J.; De Clercq, E. Mechanism of inhibitory effect of dextran sulfate and heparin on replication of human immunodeficiency virus in vitro. Proc. Natl. Acad. Sci. USA 1988, 85, 6132–6136. [Google Scholar] [CrossRef]
- World Health Organisation. Malaria. Available online: https://www.who.int/news-room/fact-sheets/detail/malaria (accessed on 10 April 2019).
- Ogawa, H.; Iwasaki, A.; Sumimoto, S.; Iwatsuki, M.; Ishiyama, A.; Hokari, R.; Otoguro, K.; Omura, S.; Suenaga, K. Isolation and Total Synthesis of Hoshinolactam, an Antitrypanosomal Lactam from a Marine Cyanobacterium. Org. Lett. 2017, 19, 890–893. [Google Scholar] [CrossRef] [PubMed]
- Davies-Coleman, M.T.; Dzeha, T.M.; Gray, C.A.; Hess, S.; Pannell, L.K.; Hendricks, D.T.; Arendse, C.E. Isolation of homodolastatin 16, a new cyclic depsipeptide from a Kenyan collection of Lyngbya majuscula. J. Nat. Prod. 2003, 66, 712–715. [Google Scholar] [CrossRef]
- Fennell, B.J.; Carolan, S.; Pettit, G.R.; Bell, A. Effects of the antimitotic natural product dolastatin 10, and related peptides, on the human malarial parasite Plasmodium falciparum. J. Antimicrob. Chemother. 2003, 51, 833–841. [Google Scholar] [CrossRef]
- Harrigan, G.G.; Luesch, H.; Yoshida, W.Y.; Moore, R.E.; Nagle, D.G.; Paul, V.J.; Mooberry, S.L.; Corbett, T.H.; Valeriote, F.A. Symplostatin 1: A Dolastatin 10 Analogue from the Marine Cyanobacterium Symploca hydnoides. J. Nat. Prod. 1998, 61, 1075–1077. [Google Scholar] [CrossRef]
- Harrigan, G.G.; Luesch, H.; Yoshida, W.Y.; Moore, R.E.; Nagle, D.G.; Paul, V.J. Symplostatin 2: A dolastatin 13 analogue from the marine cyanobacterium Symploca hydnoides. J. Nat. Prod. 1999, 62, 655–658. [Google Scholar] [CrossRef] [PubMed]
- Luesch, H.; Moore, R.E.; Paul, V.J.; Mooberry, S.L.; Corbett, T.H. Isolation of dolastatin 10 from the marine cyanobacterium Symploca species VP642 and total stereochemistry and biological evaluation of its analogue symplostatin 1. J. Nat. Prod. 2001, 64, 907–910. [Google Scholar] [CrossRef] [PubMed]
- Luesch, H.; Yoshida, W.Y.; Moore, R.E.; Paul, V.J.; Mooberry, S.L.; Corbett, T.H. Symplostatin 3, a new dolastatin 10 analogue from the marine cyanobacterium Symploca sp. VP452. J. Nat. Prod. 2002, 65, 16–20. [Google Scholar] [CrossRef]
- Mitchell, S.S.; Faulkner, D.J.; Rubins, K.; Bushman, F.D. Dolastatin 3 and two novel cyclic peptides from a palauan collection of Lyngbya majuscula. J. Nat. Prod. 2000, 63, 279–282. [Google Scholar] [CrossRef]
- Pettit, G.R. The Dolastatins. Prog. Chem. Org. Nat. Prod. 1997, 70, 2–79. [Google Scholar]
- Flahive, E.; Srirangam, J. The Dolastatins: Novel Antitumor Agents from Dolabella auricularia. In Anticancer Agents from Natural Products; CRC Press: Boca Raton, FL, USA, 2005; p. 600. [Google Scholar]
- Bai, R.; Pettit, G.R.; Hamel, E. Structure-activity studies with chiral isomers and with segments of the antimitotic marine peptide dolastatin 10. Biochem. Pharmacol. 1990, 40, 1859–1864. [Google Scholar] [CrossRef]
- World Health Organisation. Cancer. Available online: https://www.who.int/news-room/fact-sheets/detail/cancer (accessed on 10 April 2019).
- Stewart, B.W.; Wild, C.P. (Eds.) World Cancer Report 2014; The International Agency for Research on Cancer: Lyon, France, 2014. [Google Scholar]
- Mirsalis, J.C.; Schindler-Horvat, J.; Hill, J.R.; Tomaszewski, J.E.; Donohue, S.J.; Tyson, C.A. Toxicity of dolastatin 10 in mice, rats and dogs and its clinical relevance. Cancer Chemother. Pharmacol. 1999, 44, 395–402. [Google Scholar] [CrossRef]
- Francisco, J.A.; Cerveny, C.G.; Meyer, D.L.; Mixan, B.J.; Klussman, K.; Chace, D.F.; Rejniak, S.X.; Gordon, K.A.; DeBlanc, R.; Toki, B.E.; et al. cAC10-vcMMAE, an anti-CD30-monomethyl auristatin E conjugate with potent and selective antitumor activity. Blood 2003, 102, 1458–1465. [Google Scholar] [CrossRef]
- Rose, A.A.N.; Biondini, M.; Curiel, R.; Siegel, P.M. Targeting GPNMB with glembatumumab vedotin: Current developments and future opportunities for the treatment of cancer. Pharmacol. Ther. 2017, 179, 127–141. [Google Scholar] [CrossRef]
- Barchi, J.J.; Norton, T.R.; Furusawa, E.; Patterson, G.M.L.; Moore, R.E. Identification of a cytotoxin from Tolypothrix byssoidea as tubercidin. Phytochemistry 1983, 22, 2851–2852. [Google Scholar] [CrossRef]
- Stewart, J.B.; Bornemann, V.; Chen, J.L.; Moore, R.E.; Caplan, F.R.; Karuso, H.; Larsen, L.K.; Patterson, G.M.L. Cytotoxic, fungicidal nucleosides from blue green algae belonging to the Scytonemataceae. J. Antibiot. (Tokyo) 1988, 61, 1048–1056. [Google Scholar] [CrossRef]
- Mooberry, S.L.; Stratman, K.; Moore, R.E. Tubercidin stabilizes microtubules against vinblastine-induced depolymerization, a taxol-like effect. Cancer Lett. 1995, 96, 261–266. [Google Scholar] [CrossRef]
- Altmann, K.-H. Microtubule-stabilizing agents: A growing class of important anticancer drugs. Curr. Opin. Chem. Biol. 2001, 5, 424–431. [Google Scholar] [CrossRef]
- Han, B.; Gross, H.; Goeger, D.E.; Mooberry, S.L.; Gerwick, W.H. Aurilides B and C, cancer cell toxins from a Papua New Guinea collection of the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2006, 69, 572–575. [Google Scholar] [CrossRef]
- Tripathi, A.; Puddick, J.; Prinsep, M.R.; Rottmann, M.; Tan, L.T. Lagunamides A and B: Cytotoxic and antimalarial cyclodepsipeptides from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2010, 73, 1810–1814. [Google Scholar] [CrossRef]
- Tripathi, A.; Puddick, J.; Prinsep, M.R.; Rottmann, M.; Chan, K.P.; Chen, D.Y.K.; Tan, L.T. Lagunamide C, a cytotoxic cyclodepsipeptide from the marine cyanobacterium Lyngbya majuscula. Phytochemistry 2011, 72, 2369–2375. [Google Scholar] [CrossRef]
- Williams, P.G.; Yoshida, W.Y.; Quon, M.K.; Moore, R.E.; Paul, V.J. The Structure of Palau’amide, a Potent Cytotoxin from a Species of the Marine Cyanobacterium Lyngbya. J. Nat. Prod. 2003, 66, 1545–1549. [Google Scholar] [CrossRef]
- Andrianasolo, E.H.; Gross, H.; Goeger, D.; Musafija-Girt, M.; McPhail, K.; Leal, R.M.; Mooberry, S.L.; Gerwick, W.H. Isolation of swinholide A and related glycosylated derivatives from two field collections of marine cyanobacteria. Org. Lett. 2005, 7, 1375–1378. [Google Scholar] [CrossRef]
- Humisto, A.; Jokela, J.; Liu, L.; Wahlsten, M.; Wang, H.; Permi, P.; Machado, J.P.; Antunes, A.; Fewer, D.P.; Sivonen, K. The swinholide biosynthesis gene cluster from a terrestrial cyanobacterium, Nostoc sp. strain UHCC 0450. Appl. Environ. Microbiol. 2018, 84, e02321-17. [Google Scholar] [CrossRef]
- Tao, Y.; Li, P.; Zhang, D.; Glukhov, E.; Gerwick, L.; Zhang, C.; Murray, T.F.; Gerwick, W.H. Samholides, Swinholide-Related Metabolites from a Marine Cyanobacterium cf. Phormidium sp. J. Org. Chem. 2018, 83, 3034–3046. [Google Scholar] [CrossRef]
- Oftedal, L.; Myhren, L.; Jokela, J.; Gausdal, G.; Sivonen, K.; Doskeland, S.O.; Herfindal, L. The lipopeptide toxins anabaenolysin A and B target biological membranes in a cholesterol-dependent manner. Biochim. Biophys. Acta Biomembr. 2012, 1818, 3000–3009. [Google Scholar] [CrossRef]
- Jokela, J.; Oftedal, L.; Herfindal, L.; Permi, P.; Wahlsten, M.; Døskeland, S.O.; Sivonen, K. Anabaenolysins, Novel Cytolytic Lipopeptides from Benthic Anabaena Cyanobacteria. PLoS ONE 2012, 7, e41222. [Google Scholar] [CrossRef]
- Rao, J.Y.; Li, N. Microfilament Actin Remodeling as a Potential Target for Cancer Drug Development. Curr. Cancer Drug Targets 2004, 4, 345–354. [Google Scholar] [CrossRef]
- Suenaga, K.; Mutou, T.; Shibata, T.; Itoh, T.; Fujita, T.; Takada, N.; Hayamizu, K.; Takagi, M.; Irifune, T.; Kigoshi, H.; et al. Aurilide, a cytotoxic depsipeptide from the sea hare Dolabella auricularia: Isolation, structure determination, synthesis, and biological activity. Tetrahedron 2004, 60, 8509–8527. [Google Scholar] [CrossRef]
- Nakao, Y.; Yoshida, W.Y.; Takada, Y.; Kimura, J.; Yang, L.; Mooberry, S.L.; Scheuer, P.J. Kulokekahilide-2, a cytotoxic depsipeptide from a cephalaspidean mollusk Philinopsis speciosa. J. Nat. Prod. 2004, 67, 1332–1340. [Google Scholar] [CrossRef]
- Carmeli, S.; Kashman, Y. Structure of swinholide-a, a new macrolide from the marine sponge Theonella swinhoei. Tetrahedron Lett. 1985, 26, 511–514. [Google Scholar] [CrossRef]
- Bubb, M.R.; Spector, I.; Bershadsky, A.D.; Korn, E.D. Swinholide A is a microfilament disrupting marine toxin that stabilizes actin dimers and severs actin filaments. J. Biol. Chem. 1995, 270, 3463–3466. [Google Scholar] [CrossRef]
- Golakoti, T.; Ohtani, I.; Patterson, G.M.L.; Moore, R.E.; Corbett, T.H.; Valeriote, F.A.; Demchik, L. Total Structures of Cryptophycins, Potent Antitumor Depsipeptides from the Blue-Green Alga Nostoc sp. Strain GSV 224. J. Am. Chem. Soc. 1994, 116, 4729–4737. [Google Scholar]
- Golakoti, T.; Ogino, J.; Heltzel, C.E.; Husebo, T.L.; Jensen, C.M.; Larsen, L.K.; Patterson, G.M.L.; Moore, R.E.; Mooberry, S.L.; Corbett, T.H.; et al. Structure Determination, Conformational Analysis, Chemical Stability Studies, and Antitumor Evaluation of the Cryptophycins. Isolation of 18 New Analogs from Nostoc sp. Strain GSV 224. J. Am. Chem. Soc. 1995, 117, 12030–12049. [Google Scholar] [CrossRef]
- Subbaraju, G.V.; Golakoti, T.; Patterson, G.M.L.; Moore, R.E. Three new cryptophycins from Nostoc sp. GSV 224. J. Nat. Prod. 1997, 60, 302–305. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.D.; Zhang, X.; Mooberry, S.L.; Patterson, G.M.; Moore, R.E. Cryptophycin: A new antimicrotubule agent active against drug-resistant cells. Cancer Res. 1994, 54, 3779–3784. [Google Scholar]
- Stratmann, K.; Burgoyne, D.L.; Moore, R.E.; Patterson, G.M.L.; Smith, C.D. Hapalosin, a Cyanobacterial Cyclic Depsipeptide with Multidrug-Resistance Reversing Activity. J. Org. Chem. 1994, 59, 7219–7226. [Google Scholar] [CrossRef]
- Schmidt, E.W.; Nelson, J.T.; Rasko, D.A.; Sudek, S.; Eisen, J.A.; Haygood, M.G.; Ravel, J. Patellamide A and C biosynthesis by a microcin-like pathway in Prochloron didemni, the cyanobacterial symbiont of Lissoclinum patella. Proc. Natl. Acad. Sci. USA 2005, 102, 7315–7320. [Google Scholar] [CrossRef] [PubMed]
- Williams, A.B.; Jacobs, R.S. A marine natural product, patellamide D, reverses multidrug resistance in a human leukemic cell line. Cancer Lett. 1993, 71, 97–102. [Google Scholar] [CrossRef]
- Prinsep, M.R.; Caplan, F.R.; Moore, R.E.; Patterson, G.M.L.; Smith, C.D. Tolyporphin, a Novel Multidrug Resistance Reversing Agent from the Blue-Green Alga Tolypothrix nodosa. J. Am. Chem. Soc. 1992, 114, 385–387. [Google Scholar] [CrossRef]
- Prinsep, M.R.; Patterson, G.M.L.; Larsen, L.K.; Smith, C.D. Further tolyporphins from the Blue-Green alga Tolypothrix nodosa. Tetrahedron 1995, 51, 10523–10530. [Google Scholar] [CrossRef]
- Nanayakkara, A.K.; Follit, C.A.; Chen, G.; Williams, N.S.; Vogel, P.D.; Wise, J.G. Targeted inhibitors of P-glycoprotein increase chemotherapeutic-induced mortality of multidrug resistant tumor cells. Sci. Rep. 2018, 8, 967. [Google Scholar] [CrossRef]
- Ilies, M.A.; Supuran, C.T.; Scozzafava, A. Therapeutic applications of serine protease inhibitors. Expert Opin. Ther. Pat. 2002, 12, 1181–1214. [Google Scholar] [CrossRef]
- Drag, M.; Salvesen, G.S. Emerging principles in protease-based drug discovery. Nat. Rev. Drug Discov. 2010, 9, 690–701. [Google Scholar] [CrossRef] [PubMed]
- Dey, T.; Kalita, J.; Weldon, S.; Taggart, C. Proteases and Their Inhibitors in Chronic Obstructive Pulmonary Disease. J. Clin. Med. 2018, 7, 244. [Google Scholar] [CrossRef]
- Guay, C.; Laviolette, M.; Tremblay, G.M. Targeting serine proteases in asthma. Curr. Top. Med. Chem. 2006, 6, 393–402. [Google Scholar] [CrossRef]
- Williams, M.R.; Nakatsuji, T.; Sanford, J.A.; Vrbanac, A.F.; Gallo, R.L. Staphylococcus aureus Induces Increased Serine Protease Activity in Keratinocytes. J. Investig. Dermatol. 2017, 137, 377–384. [Google Scholar] [CrossRef]
- Lee, H.; Ren, J.; Nocadello, S.; Rice, A.J.; Ojeda, I.; Light, S.; Minasov, G.; Vargas, J.; Nagarathnam, D.; Anderson, W.F.; et al. Identification of novel small molecule inhibitors against NS2B/NS3 serine protease from Zika virus. Antivir. Res. 2017, 139, 49–58. [Google Scholar] [CrossRef]
- Adams, J. The development of proteasome inhibitors as anticancer drugs. Cancer Cell 2004, 5, 417–421. [Google Scholar] [CrossRef]
- Koblinski, J.E.; Ahram, M.; Sloane, B.F. Unraveling the role of proteases in cancer. Clin. Chim. Acta 2000, 291, 113–135. [Google Scholar] [CrossRef]
- Kos, J.; Mitrović, A.; Mirković, B. The current stage of cathepsin B inhibitors as potential anticancer agents. Future Med. Chem. 2014, 6, 1355–1371. [Google Scholar] [CrossRef]
- Bian, B.; Mongrain, S.; Cagnol, S.; Langlois, M.-J.; Boulanger, J.; Bernatchez, G.; Carrier, J.C.; Boudreau, F.; Rivard, N. Cathepsin B promotes colorectal tumorigenesis, cell invasion, and metastasis. Mol. Carcinog. 2016, 55, 671–687. [Google Scholar] [CrossRef] [PubMed]
- Garcia, M.; Platet, N.; Liaudet, E.; Laurent, V.; Derocq, D.; Brouillet, J.P.; Rochefort, H. Biological and clinical significance of cathepsin D in breast cancer metastasis. Stem Cells 1996, 14, 642–650. [Google Scholar] [CrossRef]
- Lankelma, J.M.; Voorend, D.M.; Barwari, T.; Koetsveld, J.; Van der Spek, A.H.; De Porto, A.P.N.A.; Van Rooijen, G.; Van Noorden, C.J.F. Cathepsin L, target in cancer treatment? Life Sci. 2010, 86, 225–233. [Google Scholar] [CrossRef]
- Leto, G.; Sepporta, M.V.; Crescimanno, M.; Flandina, C.; Tumminello, F.M. Cathepsin L in metastatic bone disease: Therapeutic implications. Biol. Chem. 2010, 391, 655–664. [Google Scholar] [CrossRef]
- Fujii, K.; Sivonen, K.; Adachi, K.; Noguchi, K.; Sano, H.; Hirayama, K.; Suzuki, M.; Harada, K. Comparative study of toxic and non-toxic cyanobacterial products: Novel peptides from toxic Nodularia spumigena AV1. Tetrahedron Lett. 1997, 38, 5525–5528. [Google Scholar] [CrossRef]
- Anas, A.R.J.; Kisugi, T.; Umezawa, T.; Matsuda, F.; Campitelli, M.R.; Quinn, R.J.; Okino, T. Thrombin Inhibitors from the Freshwater Cyanobacterium Anabaena compacta. J. Nat. Prod. 2012, 75, 1546–1552. [Google Scholar] [CrossRef]
- Fewer, D.P.; Jokela, J.; Rouhiainen, L.; Wahlsten, M.; Koskenniemi, K.; Stal, L.J.; Sivonen, K. The non-ribosomal assembly and frequent occurrence of the protease inhibitors spumigins in the bloom-forming cyanobacterium Nodularia spumigena. Mol. Microbiol. 2009, 73, 924–937. [Google Scholar] [CrossRef]
- Gondi, C.S.; Rao, J.S. Cathepsin B as a cancer target. Expert Opin. Ther. Targets 2013, 17, 281–291. [Google Scholar] [CrossRef]
- Adiv, S.; Ahronov-Nadborny, R.; Carmeli, S. New aeruginazoles, a group of thiazole-containing cyclic peptides from Microcystis aeruginosa blooms. Tetrahedron 2012, 68, 1376–1383. [Google Scholar] [CrossRef]
- Al-Awadhi, F.H.; Salvador, L.A.; Law, B.K.; Paul, V.J.; Luesch, H. Kempopeptin C, a novel marine-derived serine protease inhibitor targeting invasive breast cancer. Mar. Drugs 2017, 15, 290. [Google Scholar] [CrossRef]
- Banker, R.; Carmeli, S. Inhibitors of serine proteases from a waterbloom of the cyanobacterium Microcystis sp. Tetrahedron 1999, 55, 10835–10844. [Google Scholar] [CrossRef]
- Beresovsky, D.; Hadas, O.; Livne, A.; Sukenik, A.; Kaplan, A.; Carmeli, S. Toxins and Biologically Active Secondary Metabolites of Microcystis sp. isolated from Lake Kinneret. Isr. J. Chem. 2006, 46, 79–87. [Google Scholar] [CrossRef]
- Blom, J.F.; Bister, B.; Bischoff, D.; Nicholson, G.; Jung, G.; Süssmuth, R.D.; Jüttner, F. Oscillapeptin J, a new grazer toxin of the freshwater cyanobacterium Planktothrix rubescens. J. Nat. Prod. 2003, 66, 431–434. [Google Scholar] [CrossRef]
- Bonjouklian, R.; Smitka, T.A.; Hunt, A.H.; Occolowitz, J.L.; Perun, T.J.; Doolin, L.; Stevenson, S.; Knauss, L.; Wijayaratne, R.; Szewczyk, S.; et al. A90720A, a serine protease inhibitor isolated from a terrestrial blue-green alga Microchaete loktakensis. Tetrahedron 1996, 52, 395–404. [Google Scholar] [CrossRef]
- Choi, H.; Oh, S.K.; Yih, W.; Chin, J.; Kang, H.; Rho, J.-R. Cyanopeptoline CB071: A cyclic depsipeptide isolated from the freshwater cyanobacterium Aphanocapsa sp. Chem. Pharm. Bull. (Tokyo) 2008, 56, 1191–1193. [Google Scholar] [CrossRef]
- Elkobi-Peer, S.; Carmeli, S. New prenylated aeruginosin, microphycin, anabaenopeptin and micropeptin analogues from a Microcystis bloom material collected in Kibbutz Kfar Blum, Israel. Mar. Drugs 2015, 13, 2347–2375. [Google Scholar] [CrossRef]
- Fujii, K.; Sivonen, K.; Naganawa, E.; Harada, K. ichi Non-toxic peptides from toxic cyanobacteria, Oscillatoria agardhii. Tetrahedron 2000, 56, 725–733. [Google Scholar] [CrossRef]
- Gallegos, D.A.; Saurí, J.; Cohen, R.D.; Wan, X.; Videau, P.; Vallota-Eastman, A.O.; Shaala, L.A.; Youssef, D.T.A.; Williamson, R.T.; Martin, G.E.; et al. Jizanpeptins, Cyanobacterial Protease Inhibitors from a Symploca sp. Cyanobacterium Collected in the Red Sea. J. Nat. Prod. 2018, 81, 1417–1425. [Google Scholar] [CrossRef] [PubMed]
- Grach-Pogrebinsky, O.; Sedmak, B.; Carmeli, S. Protease inhibitors from a Slovenian Lake Bled toxic waterbloom of the cyanobacterium Planktothrix rubescens. Tetrahedron 2003, 59, 8329–8336. [Google Scholar] [CrossRef]
- Gunasekera, S.P.; Miller, M.W.; Kwan, J.C.; Luesch, H.; Paul, V.J. Molassamide, a depsipeptide serine protease inhibitor from the marine cyanobacterium Dichothrix utahensis. J. Nat. Prod. 2010, 73, 459–462. [Google Scholar] [CrossRef]
- Harada, K.I.; Mayumi, T.; Shimada, T.; Fujii, K.; Kondo, F.; Park, H.D.; Watanabe, M.F. Co-production of microcystins and aeruginopeptins by natural cyanobacterial bloom. Environ. Toxicol. 2001, 16, 298–305. [Google Scholar] [CrossRef] [PubMed]
- Ishida, K.; Matsuda, H.; Murakami, M.; Yamaguchi, K. Micropeptins 478-A and -B, plasmin inhibitors from the cyanobacterium Microcystis aeruginosa. J. Nat. Prod. 1997, 60, 184–187. [Google Scholar] [CrossRef]
- Ishida, K.; Matsuda, H.; Murakami, M. Micropeptins 88-A to 88-F, chymotrypsin inhibitors from the cyanobacterium Microcystis aeruginosa (NIES-88). Tetrahedron 1998, 54, 5545–5556. [Google Scholar] [CrossRef]
- Ishida, K.; Murakami, M.; Matsuda, H.; Yamaguchi, K. Micropeptin 90, a plasmin and trypsin inhibitor from the blue-green alga Microcystis aeruginosa (NIES-90). Tetrahedron Lett. 1995, 36, 3535–3538. [Google Scholar] [CrossRef]
- Itou, Y.; Ishida, K.; Shin, H.J.; Murakami, M. Oscillapeptins A to F, serine protease inhibitors from the three strains of Oscillatoria agardhii. Tetrahedron 1999, 55, 6871–6882. [Google Scholar] [CrossRef]
- Jakobi, C.; Oberer, L.; Quiquerez, C.; König, W.A.; Weckesser, J. Cyanopeptolin S, a sulfate-containing depsipeptide from a water bloom of Microcystis sp. FEMS Microbiol. Lett. 1995, 129, 129–133. [Google Scholar]
- Kang, H.S.; Krunic, A.; Orjala, J. Stigonemapeptin, an Ahp-containing depsipeptide with elastase inhibitory activity from the bloom-forming freshwater cyanobacterium Stigonema sp. J. Nat. Prod. 2012, 75, 807–811. [Google Scholar] [CrossRef]
- Kaya, K.; Sano, T.; Beattie, K.A.; Codd, G.A. Nostocyclin, a novel 3-amino-6-hydroxy-2-piperidone-containing cyclic depsipeptide from the cyanobacterium Nostoc sp. Tetrahedron Lett. 1996, 37, 6725–6728. [Google Scholar] [CrossRef]
- Kodani, S.; Suzuki, S.; Ishida, K.; Murakami, M. Five new cyanobacterial peptides from water bloom materials of Lake Teganuma (Japan). FEMS Microbiol. Lett. 1999, 178, 343–348. [Google Scholar] [CrossRef]
- Kwan, J.C.; Taori, K.; Paul, V.J.; Luesch, H. Lyngbyastatins 8-10, elastase inhibitors with cyclic depsipeptide scaffolds isolated from the marine cyanobacterium Lyngbya semiplena. Mar. Drugs 2009, 7, 528–538. [Google Scholar] [CrossRef] [PubMed]
- Lifshits, M.; Zafrir-Ilan, E.; Raveh, A.; Carmeli, S. Protease inhibitors from three fishpond water blooms of Microcystis spp. Tetrahedron 2011, 67, 4017–4024. [Google Scholar] [CrossRef]
- Linington, R.G.; Edwards, D.J.; Shuman, C.F.; McPhail, K.L.; Matainaho, T.; Gerwick, W.H. Symplocamide A, a potent cytotoxin and chymotrypsin inhibitor from the marine cyanobacterium Symploca sp. J. Nat. Prod. 2008, 71, 22–27. [Google Scholar] [CrossRef] [PubMed]
- Lodin-Friedman, A.; Carmeli, S. Metabolites from Microcystis aeruginosa bloom material collected at a water reservoir near Kibbutz Hafetz Haim, Israel. J. Nat. Prod. 2013, 76, 1196–1200. [Google Scholar] [CrossRef]
- Martin, C.; Oberer, L.; Buschdtt, M.; Weckesser, J. Cyanopeptolins, new depsipeptides from the cyanobacterium Microcystis sp. PCC 7806. J. Antibiot. 1993, 46, 1550–1556. [Google Scholar] [CrossRef]
- Matern, U.; Oberer, L.; Falchetto, R.A.; Erhard, M.; König, W.A.; Herdman, M.; Weckesser, J. Scyptolin A and B, cyclic depsipeptides from axenic cultures of Scytonema hofmanni PCC 7110. Phytochemistry 2001, 58, 1087–1095. [Google Scholar] [CrossRef]
- Matern, U.; Oberer, L.; Erhard, M.; Herdmand, M.; Weckesser, J. Hofmannolin, a cyanopeptolin from Scytonema hofmanni PCC 7110. Phytochemistry 2003, 64, 1061–1067. [Google Scholar] [CrossRef]
- Matthew, S.; Ross, C.; Paul, V.J.; Luesch, H. Pompanopeptins A and B, new cyclic peptides from the marine cyanobacterium Lyngbya confervoides. Tetrahedron 2008, 64, 4081–4089. [Google Scholar] [CrossRef]
- Taori, K.; Matthew, S.; Rocca, J.R.; Paul, V.J.; Luesch, H. Lyngbyastatins 5-7, potent elastase inhibitors from Floridian marine cyanobacteria, Lyngbya spp. J. Nat. Prod. 2007, 70, 1593–1600. [Google Scholar] [CrossRef] [PubMed]
- Mehner, C.; Müller, D.; Kehraus, S.; Hautmann, S.; Gütschow, M.; König, G.M. New Peptolides from the Cyanobacterium Nostoc insulare as Selective and Potent Inhibitors of Human Leukocyte Elastase. ChemBioChem 2008, 9, 2692–2703. [Google Scholar] [CrossRef] [PubMed]
- Murakami, M.; Kodani, S.; Ishida, K.; Matsuda, H.; Yamaguchi, K. Micropeptin 103, a chymotrypsin inhibitor from the cyanobacterium Microcystis viridis (NIES-103). Tetrahedron Lett. 1997, 38, 3035–3038. [Google Scholar] [CrossRef]
- Okano, T.; Sano, T.; Kaya, K. Micropeptin T-20, a novel phosphate-containing cyclic depsipeptide from the cyanobacterium Microcystis aeruginosa. Tetrahedron Lett. 1999, 40, 2379–2382. [Google Scholar] [CrossRef]
- Okino, T.; Murakami, M.; Haraguchi, R.; Munekata, H.; Matsuda, H.; Yamaguchi, K. Micropeptins A and B, plasmin and trypsin inhibitors from the blue-green alga Microcystis aeruginosa. Tetrahedron Lett. 1993, 34, 8131–8134. [Google Scholar] [CrossRef]
- Okino, T.; Qi, S.; Matsuda, H.; Murakami, M.; Yamaguchi, K. Nostopeptins A and B, elastase inhibitors from the cyanobacterium Nostoc minutum. J. Nat. Prod. 1997, 60, 158–161. [Google Scholar] [CrossRef]
- Okumura, H.S.; Philmus, B.; Portmann, C.; Hemscheidt, T.K. Homotyrosine-containing cyanopeptolins 880 and 960 and anabaenopeptins 908 and 915 from Planktothrix agardhii CYA 126/8. J. Nat. Prod. 2009, 72, 172–176. [Google Scholar] [CrossRef]
- Ploutno, A.; Carmeli, S. Modified peptides from a water bloom of the cyanobacterium Nostoc sp. Tetrahedron 2002, 58, 9949–9957. [Google Scholar] [CrossRef]
- Ploutno, A.; Shoshan, M.; Carmeli, S. Three novel protease inhibitors from a natural bloom of the cyanobacterium Microcystis aeruginosa. J. Nat. Prod. 2002, 65, 973–978. [Google Scholar] [CrossRef]
- Reshef, V.; Carmeli, S. Protease inhibitors from a water bloom of the cyanobacterium Microcystis aeruginosa. Tetrahedron 2001, 57, 2885–2894. [Google Scholar] [CrossRef]
- Sano, T.; Kaya, K. Oscillamide-Y, a Chymotrypsin Inhibitor from Toxic Oscillatoria agardhii. Tetrahedron Lett. 1995, 36, 5933–5936. [Google Scholar] [CrossRef]
- Sano, T.; Kaya, K. Oscillapeptin G, a tyrosinase inhibitor from toxic Oscillatoria agardhii. J. Nat. Prod. 1996, 59, 90–92. [Google Scholar] [CrossRef] [PubMed]
- Shin, H.J.; Murakami, M.; Matsuda, H.; Ishida, K.; Yamaguchi, K. Oscillapeptin, an elastase and chymotrypsin inhibitor from the cyanobacterium Oscillatoria agardhii (NIES-204). Tetrahedron Lett. 1995, 36, 5235–5238. [Google Scholar] [CrossRef]
- Stolze, S.C.; Deu, E.; Kaschani, F.; Li, N.; Florea, B.I.; Richau, K.H.; Colby, T.; Van Der Hoorn, R.A.L.; Overkleeft, H.S.; Bogyo, M.; et al. The antimalarial natural product symplostatin 4 is a nanomolar inhibitor of the food vacuole falcipains. Chem. Biol. 2012, 19, 1546–1555. [Google Scholar] [CrossRef]
- Taori, K.; Paul, V.J.; Luesch, H. Kempopeptins A and B, serine protease inhibitors with different selectivity profiles from a marine cyanobacterium, Lyngbya sp. J. Nat. Prod. 2008, 71, 1625–1629. [Google Scholar] [CrossRef]
- Thorskov Bladt, T.; Kalifa-Aviv, S.; Ostenfeld Larsen, T.; Carmeli, S. Micropeptins from Microcystis sp. collected in Kabul Reservoir, Israel. Tetrahedron 2014, 70, 936–943. [Google Scholar] [CrossRef]
- Tsukamoto, S.; Painuly, P.; Young, K.A.; Yang, X.; Shimizu, Y.; Cornell, L. Microcystilide A: A Novel Cell-Differentiation-Promoting Depsipeptide from Microcystis aeruginosa NO-15-1840. J. Am. Chem. Soc. 1993, 115, 11046–11047. [Google Scholar] [CrossRef]
- Vegman, M.; Carmeli, S. Eight micropeptins from a Microcystis spp. bloom collected from a fishpond near Kibbutz Lehavot HaBashan, Israel. Tetrahedron 2013, 69, 10108–10115. [Google Scholar] [CrossRef]
- Von Elert, E.; Oberer, L.; Merkel, P.; Huhn, T.; Blom, J.F. Cyanopeptolin 954, a chlorine-containing chymotrypsin inhibitor of Microcystis aeruginosa NIVA Cya 43. J. Nat. Prod. 2005, 68, 1324–1327. [Google Scholar] [CrossRef]
- Zafrir-Ilan, E.; Carmeli, S. Eight novel serine proteases inhibitors from a water bloom of the cyanobacterium Microcystis sp. Tetrahedron 2010, 66, 9194–9202. [Google Scholar] [CrossRef]
- Fujii, K.; Sivonen, K.; Nakano, T.; Harada, K.I. Structural elucidation of cyanobacterial peptides encoded by peptide synthetase gene in Anabaena species. Tetrahedron 2002, 58, 6863–6871. [Google Scholar] [CrossRef]
- Pereira, A.R.; Kale, A.J.; Fenley, A.T.; Byrum, T.; Debonsi, H.M.; Gilson, M.K.; Valeriote, F.A.; Moore, B.S.; Gerwick, W.H. The Carmaphycins: New Proteasome Inhibitors Exhibiting an α,β-Epoxyketone Warhead from a Marine Cyanobacterium. ChemBioChem 2012, 13, 810–817. [Google Scholar] [CrossRef] [PubMed]
- LaMonte, G.M.; Almaliti, J.; Bibo-Verdugo, B.; Keller, L.; Zou, B.Y.; Yang, J.; Antonova-Koch, Y.; Orjuela-Sanchez, P.; Boyle, C.A.; Vigil, E.; et al. Development of a Potent Inhibitor of the Plasmodium Proteasome with Reduced Mammalian Toxicity. J. Med. Chem. 2017, 60, 6721–6732. [Google Scholar] [CrossRef] [PubMed]
- Hirota, M.; Ohmuraya, M.; Baba, H. The role of trypsin, trypsin inhibitor, and trypsin receptor in the onset and aggravation of pancreatitis. J. Gastroenterol. 2006, 41, 832–836. [Google Scholar] [CrossRef] [PubMed]
- Hilpert, K.; Ackermann, J.; Banner, D.W.; Gast, A.; Gubernator, K.; Hadvary, P.; Labler, L.; Mueller, K.; Schmid, G. Design and Synthesis of Potent and Highly Selective Thrombin Inhibitors. J. Med. Chem. 1994, 37, 3889–3901. [Google Scholar] [CrossRef] [PubMed]
- Bui, H.T.N.; Jansen, R.; Pham, H.T.L.; Mundt, S. Carbamidocyclophanes A-E, chlorinated paracyclophanes with cytotoxic and antibiotic activity from the Vietnamese cyanobacterium Nostoc sp. J. Nat. Prod. 2007, 70, 499–503. [Google Scholar] [CrossRef]
- Chen, J.L.; Moore, R.E.; Patterson, G.M.L. Structures of Nostocyclophanes A-D. J. Org. Chem. 1991, 56, 4360–4364. [Google Scholar] [CrossRef]
- Kang, H.S.; Santarsiero, B.D.; Kim, H.; Krunic, A.; Shen, Q.; Swanson, S.M.; Chai, H.; Kinghorn, A.D.; Orjala, J. Merocyclophanes A and B, antiproliferative cyclophanes from the cultured terrestrial Cyanobacterium Nostoc sp. Phytochemistry 2012, 79, 109–115. [Google Scholar] [CrossRef]
- Luo, S.; Kang, H.S.; Krunic, A.; Chlipala, G.E.; Cai, G.; Chen, W.L.; Franzblau, S.G.; Swanson, S.M.; Orjala, J. Carbamidocyclophanes F and G with anti-Mycobacterium tuberculosis activity from the cultured freshwater cyanobacterium Nostoc sp. Tetrahedron Lett. 2014, 55, 686–689. [Google Scholar] [CrossRef]
- Moore, B.S.; Chen, J.-L.; Patterson, G.M.L.; Moore, R.E. Structures of Cylindrocyclophanes A-F. Tetrahedron 1992, 48, 3001–3006. [Google Scholar] [CrossRef]
- Nakamura, H.; Hamer, H.A.; Sirasani, G.; Balskus, E.P. Cylindrocyclophane biosynthesis involves functionalization of an unactivated carbon center. J. Am. Chem. Soc. 2012, 134, 18518–18521. [Google Scholar] [CrossRef]
- Preisitsch, M.; Harmrolfs, K.; Pham, H.T.; Heiden, S.E.; Füssel, A.; Wiesner, C.; Pretsch, A.; Swiatecka-Hagenbruch, M.; Niedermeyer, T.H.; Müller, R.; et al. Anti-MRSA-acting carbamidocyclophanes H-L from the Vietnamese cyanobacterium Nostoc sp. CAVN2. J. Antibiot. (Tokyo) 2015, 68, 165–177. [Google Scholar] [CrossRef] [PubMed]
- Golakoti, T.; Yoshida, W.Y.; Chaganty, S.; Moore, R.E. Isolation and Structure Determination of Nostocyclopeptides A1 and A2 from the Terrestrial Cyanobacterium Nostoc sp. ATCC53789. J. Nat. Prod. 2001, 64, 54–59. [Google Scholar] [CrossRef]
- Krunic, A.; Vallat, A.; Mo, S.; Lantvit, D.D.; Swanson, S.M.; Orjala, J. Scytonemides A and B, cyclic peptides with 20S proteasome inhibitory activity from the cultured cyanobacterium Scytonema hofmanii. J. Nat. Prod. 2010, 73, 1927–1932. [Google Scholar] [CrossRef]
- Shim, S.H.; Chlipala, G.; Orjala, J. Isolation and structure determination of a proteasome inhibitory metabolite from a culture of Scytonema hofmanni. J. Microbiol. Biotechnol. 2008, 18, 1655–1658. [Google Scholar]
- Kobayashi, A.; Kaiiyama, S.; Inawaka, K.; Kanzaki, H.; Kawazu, K. Nostodione A, a Novel Mitotic Spindle Poison* from a Blue-Green Alga Nostoc commune. Z. Nat. C 1994, 49, 464–470. [Google Scholar] [CrossRef]
- Almaliti, J.; Miller, B.; Pietraszkiewicz, H.; Glukhov, E.; Naman, C.B.; Kline, T.; Hanson, J.; Li, X.; Zhou, S.; Valeriote, F.A.; et al. Exploration of the carmaphycins as payloads in antibody drug conjugate anticancer agents. Eur. J. Med. Chem. 2018, 161, 416–432. [Google Scholar] [CrossRef] [PubMed]
- Salvador-Reyes, L.A.; Luesch, H. Biological targets and mechanisms of action of natural products from marine cyanobacteria. Nat. Prod. Rep. 2015, 32, 478–503. [Google Scholar] [CrossRef] [PubMed]
- Bolden, J.E.; Peart, M.J.; Johnstone, R.W. Anticancer activities of histone deacetylase inhibitors. Nat. Rev. Drug Discov. 2006, 5, 769–784. [Google Scholar] [CrossRef] [PubMed]
- Taori, K.; Paul, V.J.; Luesch, H. Structure and activity of largazole, a potent antiproliferative agent from the Floridian marine cyanobacterium Symploca sp. J. Am. Chem. Soc. 2008, 130, 1806–1807. [Google Scholar] [CrossRef]
- Liu, Y.; Salvador, L.A.; Byeon, S.; Ying, Y.; Kwan, J.C.; Law, B.K.; Hong, J.; Luesch, H. Anticolon Cancer Activity of Largazole, a Marine-Derived Tunable Histone Deacetylase Inhibitor. J. Pharmacol. Exp. Ther. 2010, 335, 351–361. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.C.; Wen, Z.S.; Qiu, Y.T.; Chen, X.Q.; Chen, H.B.; Wei, M.M.; Liu, Z.; Jiang, S.; Zhou, G.B. Largazole arrests cell cycle at g1 phase and triggers proteasomal degradation of E2F1 in lung cancer cells. ACS Med. Chem. Lett. 2013, 4, 921–926. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Jiang, S.; Chen, J.; Ren, X.; Jin, J.; Su, S.B. Largazole, an inhibitor of class I histone deacetylases, attenuates inflammatory corneal neovascularization. Eur. J. Pharmacol. 2014, 740, 619–626. [Google Scholar] [CrossRef] [PubMed]
- Poli, G.; Di Fabio, R.; Ferrante, L.; Summa, V.; Botta, M. Largazole Analogues as Histone Deacetylase Inhibitors and Anticancer Agents: An Overview of Structure–Activity Relationships. ChemMedChem 2017, 12, 1917–1926. [Google Scholar] [CrossRef]
- Ungermannova, D.; Parker, S.J.; Nasveschuk, C.G.; Wang, W.; Quade, B.; Zhang, G.; Kuchta, R.D.; Phillips, A.J.; Liu, X. Largazole and its derivatives selectively inhibit ubiquitin activating enzyme (E1). PLoS ONE 2012, 7, e29208. [Google Scholar] [CrossRef] [PubMed]
- Pavlik, C.M.; Wong, C.Y.B.; Ononye, S.; Lopez, D.D.; Engene, N.; McPhail, K.L.; Gerwick, W.H.; Balunas, M.J. Santacruzamate A, a potent and selective histone deacetylase inhibitor from the Panamanian marine cyanobacterium cf. Symploca sp. J. Nat. Prod. 2013, 76, 2026–2033. [Google Scholar] [CrossRef]
- Hanke, T.; Merk, D.; Steinhilber, D.; Geisslinger, G.; Schubert-Zsilavecz, M. Small molecules with anti-inflammatory properties in clinical development. Pharmacol. Ther. 2016, 157, 163–187. [Google Scholar] [CrossRef]
- Ridker, P.M.; Lüscher, T.F. Anti-inflammatory therapies for cardiovascular disease. Eur. Heart J. 2014, 35, 1782–1791. [Google Scholar] [CrossRef]
- Rocha, N.P.; De Miranda, A.S.; Teixeira, A.L. Insights into neuroinflammation in Parkinson’s disease: From biomarkers to anti-inflammatory based therapies. BioMed Res. Int. 2015, 2015, 628192. [Google Scholar] [CrossRef]
- Ishida, K.; Okita, Y.; Matsuda, H.; Okino, T.; Murakami, M. Aeruginosins, protease inhibitors from the cyanobacterium Microcystis aeruginosa. Tetrahedron 1999, 55, 10971–10988. [Google Scholar] [CrossRef]
- Ishida, K.; Christiansen, G.; Yoshida, W.Y.; Kurmayer, R.; Welker, M.; Valls, N.; Bonjoch, J.; Hertweck, C.; Börner, T.; Hemscheidt, T.; et al. Biosynthesis and Structure of Aeruginoside 126A and 126B, Cyanobacterial Peptide Glycosides Bearing a 2-Carboxy-6-Hydroxyoctahydroindole Moiety. Chem. Biol. 2007, 14, 565–576. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, H.; Okino, T.; Murakami, M.; Yamaguchi, K. Aeruginosins 102-A and B, New Thrombin Inhibitors from the Cyanobacterium Microcystis viridis (NIES-102). Tetrahedron 1996, 52, 14501–14506. [Google Scholar] [CrossRef]
- Benedetti, S.; Benvenuti, F.; Pagliarani, S.; Francogli, S.; Scoglio, S.; Canestrari, F. Antioxidant properties of a novel phycocyanin extract from the blue-green alga Aphanizomenon flos-aquae. Life Sci. 2004, 75, 2353–2362. [Google Scholar] [CrossRef] [PubMed]
- Kuddus, M.; Singh, P.; Thomas, G.; Al-Hazimi, A. Recent developments in production and biotechnological applications of c-phycocyanin. BioMed Res. Int. 2013, 2013, 742859. [Google Scholar] [CrossRef]
- Patel, A.; Mishra, S.; Ghosh, P.K. Antioxidant potential of C-phycocyanin isolated from cyanobacterial species Lyngbya, Phormidium and Spirulina spp. Indian J. Biochem. Biophys. 2006, 43, 25–31. [Google Scholar] [PubMed]
- Garcia-Pichel, F.; Castenholz, R.W. Characterization and biological implications of scytonemin, a cyanobacterial sheath pigment. J. Phycol. 1991, 27, 395–409. [Google Scholar] [CrossRef]
- Proteau, P.J.; Gerwick, W.H.; Garcia-Pichel, F.; Castenholz, R. The structure of scytonemin, an ultraviolet sunscreen pigment from the sheaths of cyanobacteria. Experientia 1993, 49, 825–829. [Google Scholar] [CrossRef]
- Soule, T.; Stout, V.; Swingley, W.D.; Meeks, J.C.; Garcia-Pichel, F. Molecular genetics and genomic analysis of scytonemin biosynthesis in Nostoc punctiforme ATCC 29133. J. Bacteriol. 2007, 189, 4465–4472. [Google Scholar] [CrossRef]
- Stevenson, C.S.; Capper, E.A.; Roshak, A.K.; Marquez, B.; Grace, K.; Gerwick, W.H.; Jacobs, R.S.; Marshall, L.A. Scytonemin-a marine natural product inhibitor of kinases key in hyperproliferative inflammatory diseases. Inflamm. Res. 2002, 51, 112–114. [Google Scholar] [CrossRef]
- Stevenson, C.S.; Capper, E.A.; Roshak, A.M.Y.K.; Marquez, B.; Eichman, C.; Jackson, J.R.; Mattern, M.; Gerwick, W.H.; Jacobs, R.S.; Marshall, L.A. The identification and characterization of the marine natural product scytonemin as a novel antiproliferative pharmacophore. J. Pharmacol. Exp. Ther. 2002, 303, 858–866. [Google Scholar] [CrossRef] [PubMed]
- Malloy, K.L.; Villa, F.A.; Engene, N.; Matainaho, T.; Gerwick, L.; Gerwick, W.H. Malyngamide 2, an oxidized lipopeptide with nitric oxide inhibiting activity from a Papua New Guinea marine cyanobacterium. J. Nat. Prod. 2011, 74, 95–98. [Google Scholar] [CrossRef] [PubMed]
- Padrines, M.; Wolf, M.; Walz, A.; Baggiolini, M. Interleukin-8 processing by neutrophil elastase, cathepsin G and proteinase-3. FEBS Lett. 1994, 352, 231–235. [Google Scholar] [CrossRef]
- Wiedow, O.; Meyer-Hoffert, U. Neutrophil serine proteases: Potential key regulators of cell signalling during inflammation. J. Intern. Med. 2005, 257, 319–328. [Google Scholar] [CrossRef] [PubMed]
- Pham, C.T.N. Neutrophil serine proteases: Specific regulators of inflammation. Nat. Rev. Immunol. 2006, 6, 541–550. [Google Scholar] [CrossRef]
- Dubois, R.N.; Abramson, S.B.; Crofford, L.; Gupta, R.A.; Simon, L.S.; Van De Putte, L.B.; Lipsky, P.E. Cyclooxygenase in biology and disease. FASEB J. 1998, 12, 1063–1073. [Google Scholar] [CrossRef]
- Patrignani, P.; Patrono, C. Cyclooxygenase inhibitors: From pharmacology to clinical read-outs. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2015, 1851, 422–432. [Google Scholar] [CrossRef] [PubMed]
- Vane, J.R.; Botting, R.M. Anti-inflammatory drugs and their mechanism of action. Inflamm. Res. 1998, 47, 78–87. [Google Scholar] [CrossRef]
- Uttara, B.; Singh, A.V.; Zamboni, P.; Mahajan, R.T. Oxidative stress and neurodegenerative diseases: A review of upstream and downstream antioxidant therapeutic options. Curr. Neuropharmacol. 2009, 7, 65–74. [Google Scholar] [CrossRef]
- Kim, G.H.; Kim, J.E.; Rhie, S.J.; Yoon, S. The Role of Oxidative Stress in Neurodegenerative Diseases. Exp. Neurobiol. 2015, 24, 325. [Google Scholar] [CrossRef]
- Rani, V.; Deep, G.; Singh, R.K.; Palle, K.; Yadav, U.C.S. Oxidative stress and metabolic disorders: Pathogenesis and therapeutic strategies. Life Sci. 2016, 148, 183–193. [Google Scholar] [CrossRef]
- Baradaran, A.; Nasri, H.; Rafieian-Kopaei, M. Oxidative stress and hypertension: Possibility of hypertension therapy with antioxidants. J. Res. Med. Sci. 2014, 19, 358–367. [Google Scholar] [PubMed]
- Li, S.; Tan, H.Y.; Wang, N.; Zhang, Z.J.; Lao, L.; Wong, C.W.; Feng, Y. The role of oxidative stress and antioxidants in liver diseases. Int. J. Mol. Sci. 2015, 16, 26087–26124. [Google Scholar] [CrossRef]
- Siti, H.N.; Kamisah, Y.; Kamsiah, J. The role of oxidative stress, antioxidants and vascular inflammation in cardiovascular disease (a review). Vascul. Pharmacol. 2015, 71, 40–56. [Google Scholar] [CrossRef] [PubMed]
- Wada, N.; Sakamoto, T.; Matsugo, S. Multiple roles of photosynthetic and sunscreen pigments in cyanobacteria focusing on the oxidative stress. Metabolites 2013, 3, 463–483. [Google Scholar] [CrossRef]
- Hirschberg, J.; Chamovitz, D. Carotenoids in Cyanobacteria BT—The Molecular Biology of Cyanobacteria. In The Molecular Biology of Cyanobacteria; Springer: Dordrecht, The Netherlands, 1994; pp. 559–579. ISBN 978-94-011-0227-8. [Google Scholar]
- Balskus, E.P.; Walsh, C.T. The genetic and molecular basis for sunscreen biosynthesis in cyanobacteria. Science 2010, 329, 1653–1656. [Google Scholar] [CrossRef] [PubMed]
- Waditee-Sirisattha, R.; Kageyama, H.; Sopun, W.; Tanaka, Y.; Takabe, T. Identification and upregulation of biosynthetic genes required for accumulation of Mycosporine-2-glycine under salt stress conditions in the halotolerant cyanobacterium Aphanothece halophytica. Appl. Environ. Microbiol. 2014, 80, 1763–1769. [Google Scholar] [CrossRef] [PubMed]
- Kosourov, S.; Murukesan, G.; Jokela, J.; Allahverdiyeva, Y. Carotenoid biosynthesis in calothrix sp. 336/3: Composition of carotenoids on full medium, during diazotrophic growth and after long-term H2photoproduction. Plant Cell Physiol. 2016, 57, 2269–2282. [Google Scholar] [CrossRef] [PubMed]
- Al-Awadhi, F.H.; Law, B.K.; Paul, V.J.; Luesch, H. Grassystatins D-F, Potent Aspartic Protease Inhibitors from Marine Cyanobacteria as Potential Antimetastatic Agents Targeting Invasive Breast Cancer. J. Nat. Prod. 2017, 80, 2969–2986. [Google Scholar] [CrossRef]
- Al-Awadhi, F.H.; Ratnayake, R.; Paul, V.J.; Luesch, H. Tasiamide F, a potent inhibitor of cathepsins D and E from a marine cyanobacterium. Bioorg. Med. Chem. 2016, 24, 3276–3282. [Google Scholar] [CrossRef]
- Kwan, J.C.; Eksioglu, E.A.; Liu, C.; Paul, V.J.; Luesch, H. Grassystatins A-C from marine cyanobacteria, potent cathepsin E inhibitors that reduce antigen presentation. J. Med. Chem. 2009, 52, 5732–5747. [Google Scholar] [CrossRef] [PubMed]
- Molinski, T.F.; Reynolds, K.A.; Morinaka, B.I. Symplocin A, a linear peptide from the bahamian cyanobacterium Symploca sp. configurational analysis of N,N-dimethylamino acids by chiral-phase HPLC of naphthacyl esters. J. Nat. Prod. 2012, 75, 425–431. [Google Scholar] [CrossRef]
- Williams, P.G.; Yoshida, W.Y.; Moore, R.E.; Paul, V.J. Tasiamide, a cytotoxic peptide from the marine cyanobacterium Symploca sp. J. Nat. Prod. 2002, 65, 1336–1339. [Google Scholar] [CrossRef]
- Williams, P.G.; Yoshida, W.Y.; Moore, R.E.; Paul, V.J. The isolation and structure elucidation of tasiamide B, a 4-amino-3-hydroxy-5-phenylpentanoic acid containing peptide from the marine cyanobacterium Symploca sp. J. Nat. Prod. 2003, 66, 1006–1009. [Google Scholar] [CrossRef] [PubMed]
- Mevers, E.; Haeckl, F.P.J.; Boudreau, P.D.; Byrum, T.; Dorrestein, P.C.; Valeriote, F.A.; Gerwick, W.H. Lipopeptides from the tropical marine cyanobacterium Symploca sp. J. Nat. Prod. 2014, 77, 969–975. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, K.; Okamoto, K.; Tsukuba, T. Cathepsin E: An Aspartic Protease with Diverse Functions and Biomedical Implications. In Encyclopedia of Cell Biology; Academic Press: Cambridge, MA, USA, 2015; Volume 1, pp. 681–690. ISBN 9780123944474. [Google Scholar]
- Vassar, R.; Bennett, B.D.; Babu-Khan, S.; Kahn, S.; Mendiaz, E.A.; Denis, P.; Teplow, D.B.; Ross, S.; Amarante, P.; Loeloff, R.; et al. β-Secretase cleavage of Alzheimer’s amyloid precursor protein by the transmembrane aspartic protease BACE. Science 1999, 286, 735–741. [Google Scholar] [CrossRef] [PubMed]
- Sitachitta, N.; Gerwick, W.H. Grenadadiene and grenadamide, cyclopropyl-containing fatty acid metabolites from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 1998, 61, 681–684. [Google Scholar] [CrossRef] [PubMed]
- Han, B.; McPhail, K.L.; Ligresti, A.; Di Marzo, V.; Gerwick, W.H. Semiplenamides A-G, Fatty Acid Amides from a Papua New Guinea Collection of the Marine Cyanobacterium Lyngbya semiplena. J. Nat. Prod. 2003, 66, 1364–1368. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, M.; Pereira, A.R.; Debonsi, H.M.; Ligresti, A.; Di Marzo, V.; Gerwick, W.H. Cannabinomimetic lipid from a marine cyanobacterium. J. Nat. Prod. 2011, 74, 2313–2317. [Google Scholar] [CrossRef] [PubMed]
- Mevers, E.; Matainaho, T.; Allara’, M.; Di Marzo, V.; Gerwick, W.H. Mooreamide A: A cannabinomimetic lipid from the marine cyanobacterium Moorea bouillonii. Lipids 2014, 49, 1127–1132. [Google Scholar] [CrossRef]
- Kleigrewe, K.; Almaliti, J.; Tian, I.Y.; Kinnel, R.B.; Korobeynikov, A.; Monroe, E.A.; Duggan, B.M.; Di Marzo, V.; Sherman, D.H.; Dorrestein, P.C.; et al. Combining Mass Spectrometric Metabolic Profiling with Genomic Analysis: A Powerful Approach for Discovering Natural Products from Cyanobacteria. J. Nat. Prod. 2015, 78, 1671–1682. [Google Scholar] [CrossRef] [PubMed]
- Aizpurua-Olaizola, O.; Elezgarai, I.; Rico-Barrio, I.; Zarandona, I.; Etxebarria, N.; Usobiaga, A. Targeting the endocannabinoid system: Future therapeutic strategies. Drug Discov. Today 2017, 22, 105–110. [Google Scholar] [CrossRef] [PubMed]
- Patel, S.; Hill, M.N.; Cheer, J.F.; Wotjak, C.T.; Holmes, A. The endocannabinoid system as a target for novel anxiolytic drugs. Neurosci. Biobehav. Rev. 2017, 76, 56–66. [Google Scholar] [CrossRef] [PubMed]
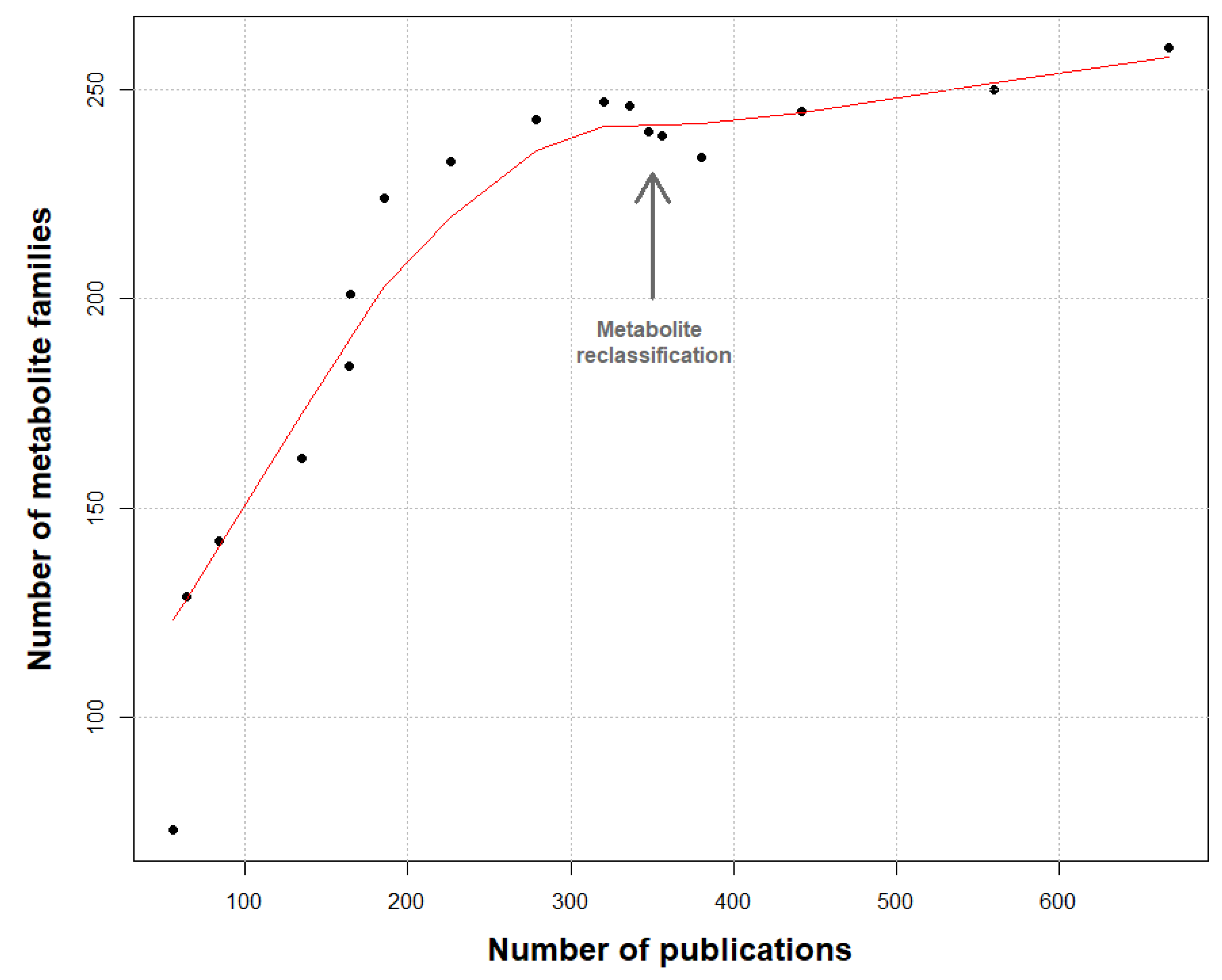
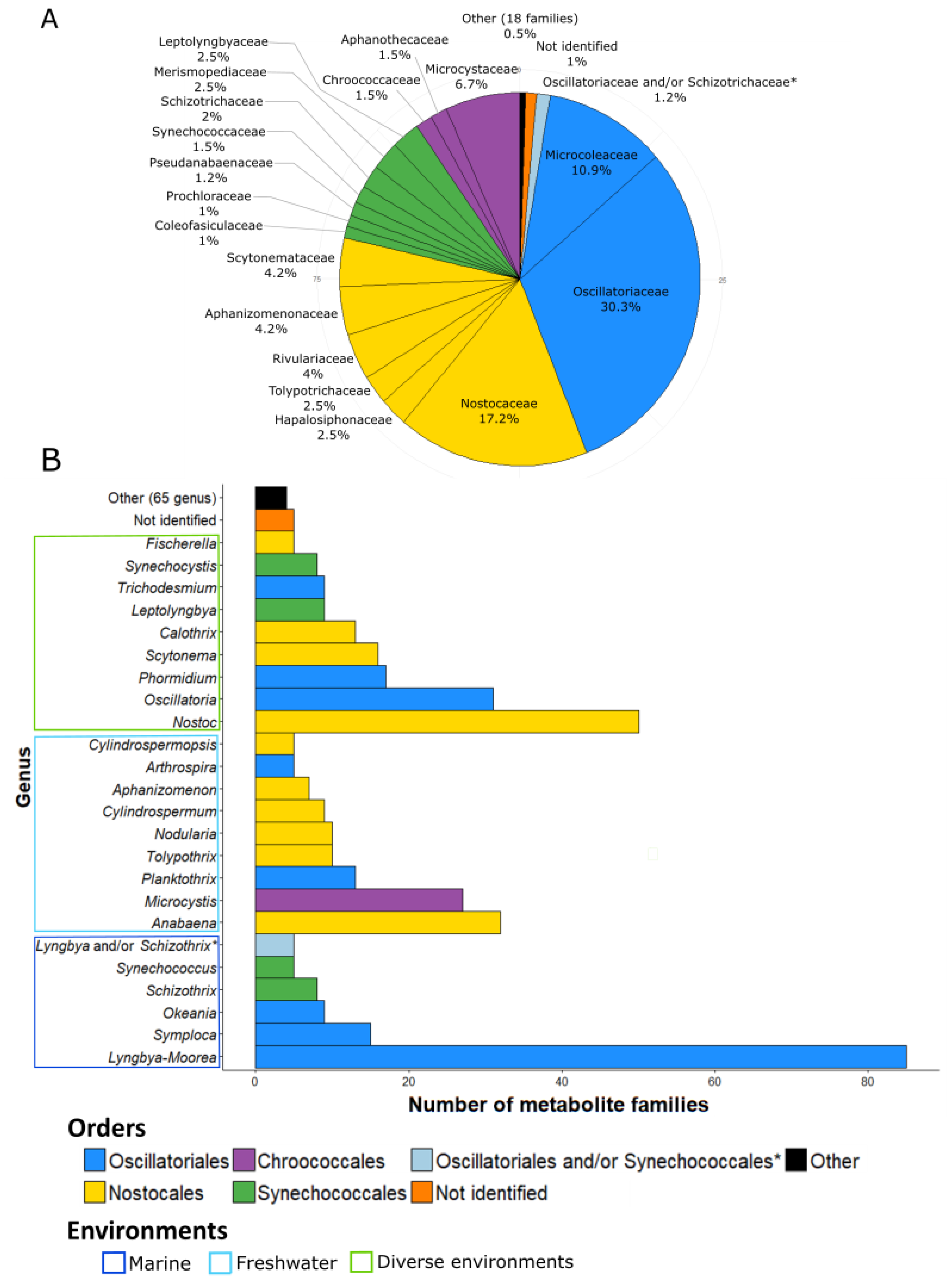
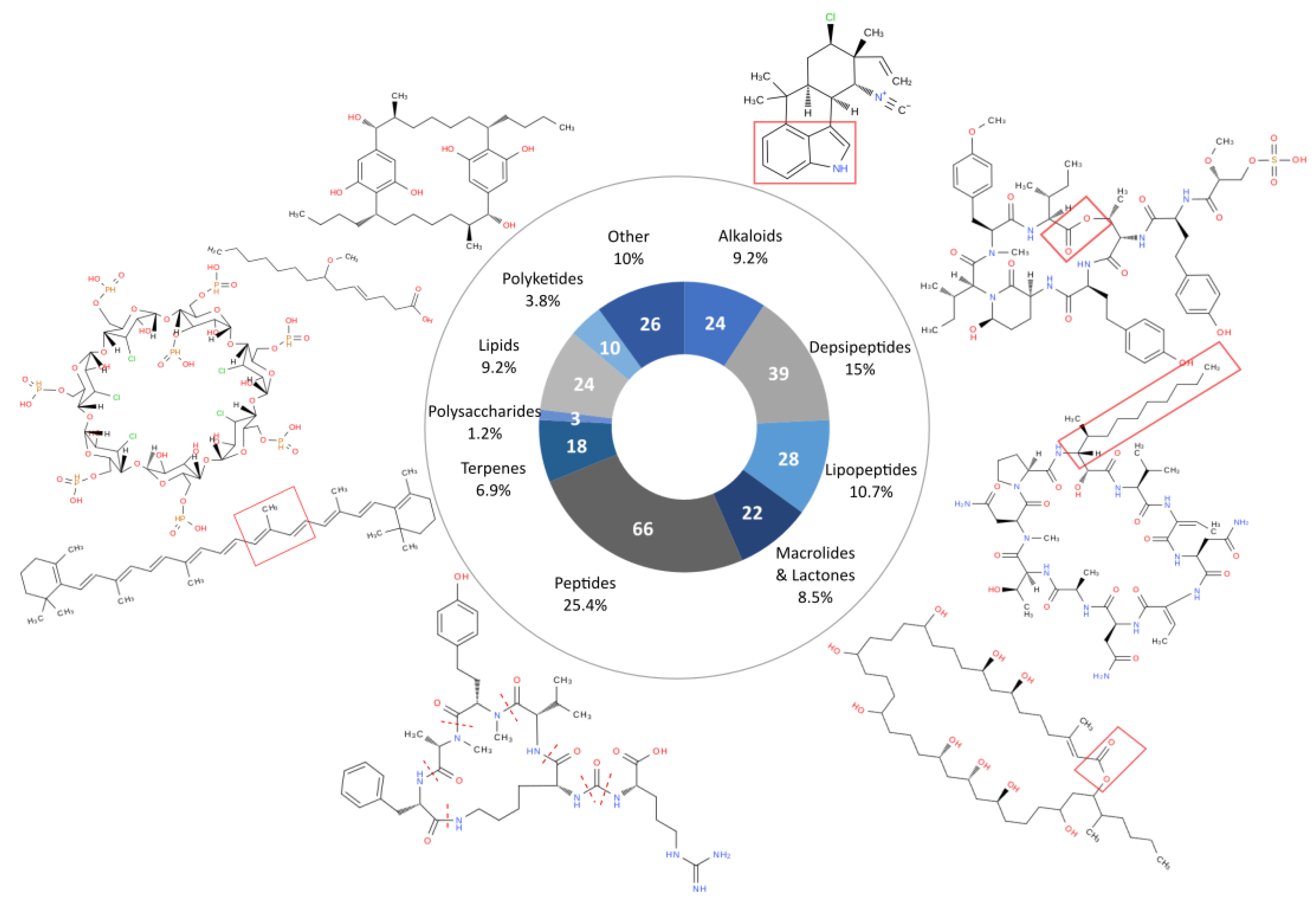
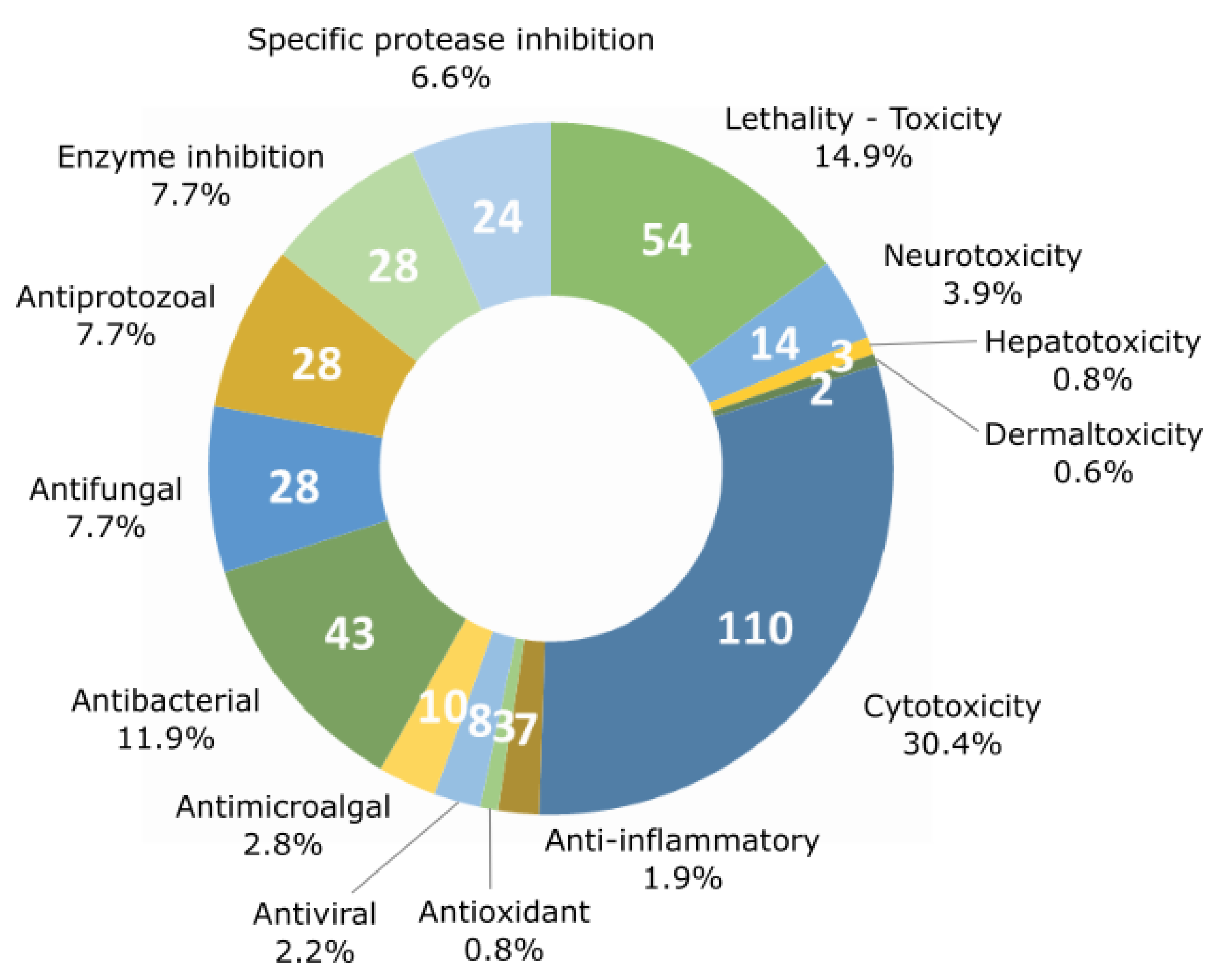
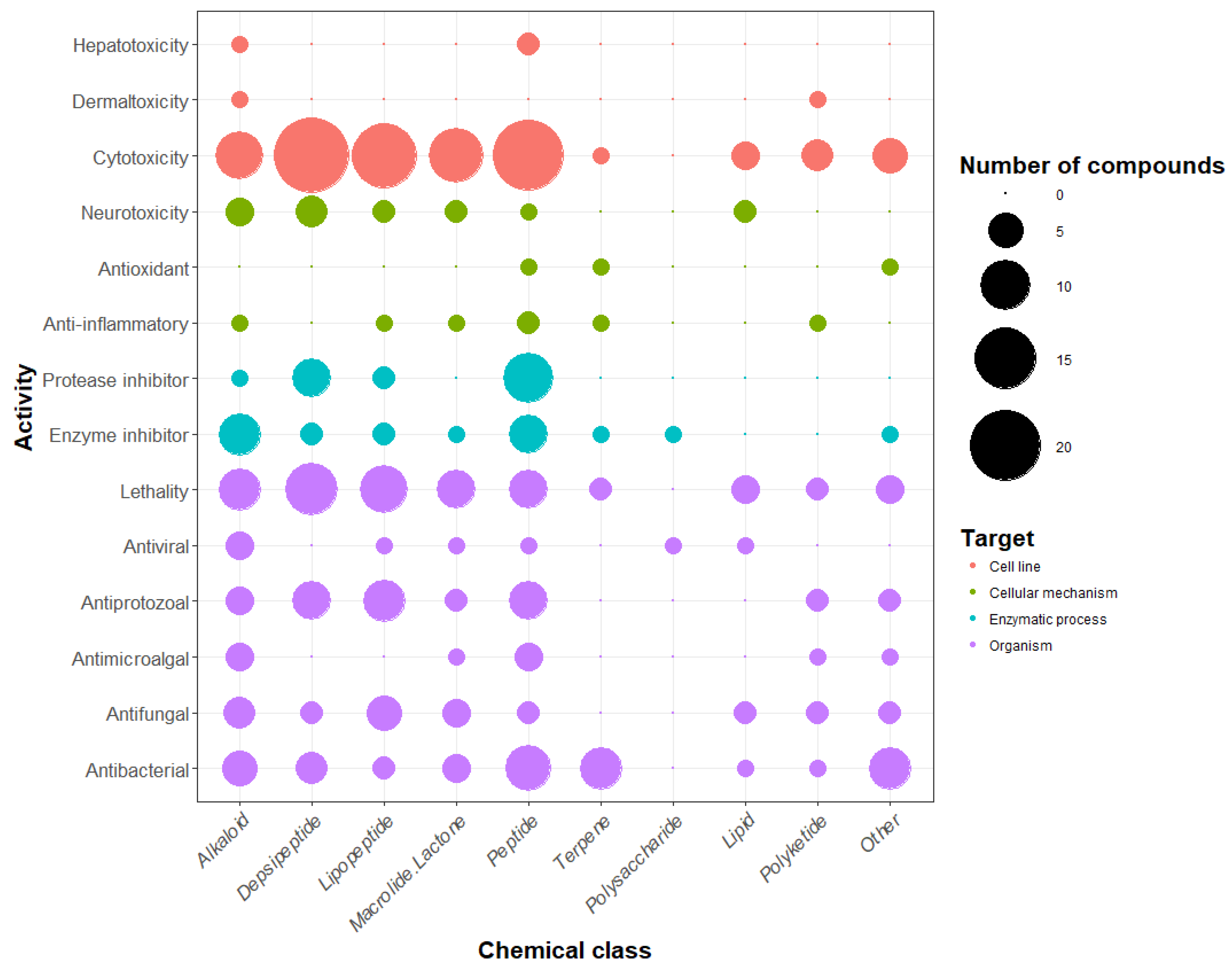
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Tubercidin | Nucleoside | -Cytotoxic -Microtubule stabilizer | Tolypothrix byssoidea H-6-2, Scytonema saleyeriense var. indica CV-14-1, Plectonema radiosum DF-6-1, Tolypothrix distorta BL-11-2 | [198,199,200] |
| Aurilides | Depsipeptide | -Cytotoxic -Lethal activity -Anti-swarming -Antiprotozoal -Induce loss of microfilament network | Lyngbya majuscula, Okeania sp., Lyngbya sp. | [70,202,203,204,205] |
| Swinholide-type | Macrolide | -Cytotoxic -Actin microfilament disruption | Symploca sp., Geitlerinema sp., Nostoc sp. UHCC0451, Phormidium sp. | [206,207,208] |
| Anabaenolysins | Lipopeptide | -Cytotoxic -Antifungal -Hemolytic activity -Ability to permeabilize cell membranes | Anabaena sp. XPORK 15F, Anabaena sp. XSPORK 27C | [209,210] |
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Spumigins | Peptide | -Proteases inhibitory activity | Nodularia spumigena AV1 & CCY 9414, Anabaena compacta NIES-835 | [239,240,241] |
| Cyanopeptolin-like | Depsipeptide | -Protease inhibitory activity -Other enzyme inhibition -Cytotoxic -Lethal -Antibacterial -Antifungal -Antiprotozoal | Microcystis sp., Microcystis aeruginosa, Aphanocapsa sp.; Microchaete loktahensis, Planktothrix agardhii, Scytonema hofmanni, Lyngbya sp., Lyngbya confervoides, Lyngbya spp., Lyngbya semiplena, Microcystis viridis, Dichothrix utahensis, Nostoc sp., Nostoc minutum, Planktothrix rubescens, Lyngbya majuscula-Schizothrix sp. (Assemblage), Stigonema sp., Symploca sp., Symploca hydnoides, Nostoc insulare | [30,31,159,185,186,188,243,244,245,246,247,248,249,250,251,252,253,254,255,256,257,258,259,260,261,262,263,264,265,266,267,268,269,270,271,272,273,274,275,276,277,278,279,280,281,282,283,284,285,286,287,288,289,290,291,292] |
| Carmaphycins | Peptide | -Protease inhibition -Cytotoxic -Antiprotozoal | Symploca sp. WHG NAC15/Dec/08–5 | [293,294] |
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Largazole | Depsipeptide | -Histone deacetylases inhibitor -Cytotoxic -Other enzyme inhibition -Pro-drug | Symploca sp. | [311,312,313,314,315,316] |
| Santacruzamate A | Carboxylic acid derived | -Histone deacetylases inhibitor -Cytotoxic | Symploca sp. PAC-19-FEB-10-1 | [317] |
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Aeruginosins | Peptide | -Anti-inflammatory activity -Protease inhibitor -No cytotoxicity | Microcystis aeruginosa NIES-98, NIES-298, NIES-101, NIES-89. Microcystis viridis NIES-102 Planktothrix agardhii CYA 126/8. Nodularia spumigena CCY9414. Nostoc sp. Lukesova 30/93 | [62,321,322,323] |
| Phycocyanin | Peptide | -Anti-inflammatory -Antioxidant -Specific inhibitor of COX-2 -No lethality | All | [59,64,324,325,326] |
| Scytonemin | Alkaloid | -Anti-inflammatory -Enzyme inhibition -No cytotoxicity | Stigonema sp., Nostoc punctiforme, Anabaena variabilis, Anabaena ambigua, Aphanocapsa/Synechocystis sp. (assembly), Aulosira fertilissima, Calothrix sp., Calothrix parietina, Calothrix crustacea, Chlorogloeopsis sp., Chroococcidiopsis sp., Chroococcus sp.; Cylindrospermum sp., Diplocolon sp., Entophysalis granulos, Gloeocapsa sp., Hapalosiphon sp., Hapalosiphon fontinalis; Lyngbya sp., Lyngbya aestuarii, Nostoc parmelioides, Nostoc commune, Nostoc microscopium, Nostoc pruniforme, Phormidium sp., Pleurocapsa sp., Rivularia atra, Rivularia sp., Schizothrix sp., Scytonema sp., Tolyothrix sp., Tolypothrix tenni, Westiellopsis prolifica, Scytonema hoffmani | [65,327,328,329,330,331] |
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Carotenoids | Terpenoid | -Antioxidant -Sunscreen | All | [56,345,346] |
| Chlorophylls | Chlorin | -Photosynthesis -Antioxidant -Pro-oxidant (sensitizer for singlet oxygen production) | All | [57,345] |
| MAAs | Cyclohexenone linked with an amino acid | -Antioxidant -Sunscreens | Synechocystis sp. PCC 6803, Gloeocapsa sp. CU-2556, Aphanothece halophytica, Gloeocapsa sp., Euhalothece sp., Microcystis aeruginosa, Arthrospira sp. CU2556, Lyngbya sp. CU2555, Leptolyngbya sp., Phormidium sp., Lyngbya cf. aestuarii, Microcoleus chthonoplastes, Microcoleus sp., Oscillatoria spongelidae, Trichodesmium spp., Anabaena sp., Anabaena doliolum, Anabaena variabilis PCC 7937, Nostoc sp., Nostoc commune var. Vaucher, Nostoc commune, Scytonema sp., Nostoc punctiforme ATCC 29133, Nostoc sp. HKAR-2 and HKAR-6, Nodularia baltica, Nodularia harveyana, Nodularia spumigena, Aphanizomenon flos-aquae, Chlorogloeopsis PCC 6912 | [58,345,347,348] |
| Phycocyanin | Peptide | -Anti-inflammatory -Antioxidant -Specific inhibitor of COX-2 -No lethality | All | [59,64,324,325,326] |
| Molecule Family | Chemical Classes | Activity | Producing Organisms | References |
|---|---|---|---|---|
| Grassystatins-Tasiamides | Depsipeptide | -Protease inhibitory activity -Cytotoxic -Reduce antigen presentation in dendritic cells | Lyngbya confervoides, Symploca sp., Symploca sp. NHI304, Lyngbya sp. NIH399 | [350,351,352,353,354,355,356] |
| Columbamides | Acyl amide | -CB1 and CB2 ligands | Moorea bouillonii PNG05-198 | [363] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Demay, J.; Bernard, C.; Reinhardt, A.; Marie, B. Natural Products from Cyanobacteria: Focus on Beneficial Activities. Mar. Drugs 2019, 17, 320. https://doi.org/10.3390/md17060320
Demay J, Bernard C, Reinhardt A, Marie B. Natural Products from Cyanobacteria: Focus on Beneficial Activities. Marine Drugs. 2019; 17(6):320. https://doi.org/10.3390/md17060320
Chicago/Turabian StyleDemay, Justine, Cécile Bernard, Anita Reinhardt, and Benjamin Marie. 2019. "Natural Products from Cyanobacteria: Focus on Beneficial Activities" Marine Drugs 17, no. 6: 320. https://doi.org/10.3390/md17060320
APA StyleDemay, J., Bernard, C., Reinhardt, A., & Marie, B. (2019). Natural Products from Cyanobacteria: Focus on Beneficial Activities. Marine Drugs, 17(6), 320. https://doi.org/10.3390/md17060320





