Identification of the Anti-Infective Aborycin Biosynthetic Gene Cluster from Deep-Sea-Derived Streptomyces sp. SCSIO ZS0098 Enables Production in a Heterologous Host
Abstract
:1. Introduction
2. Results
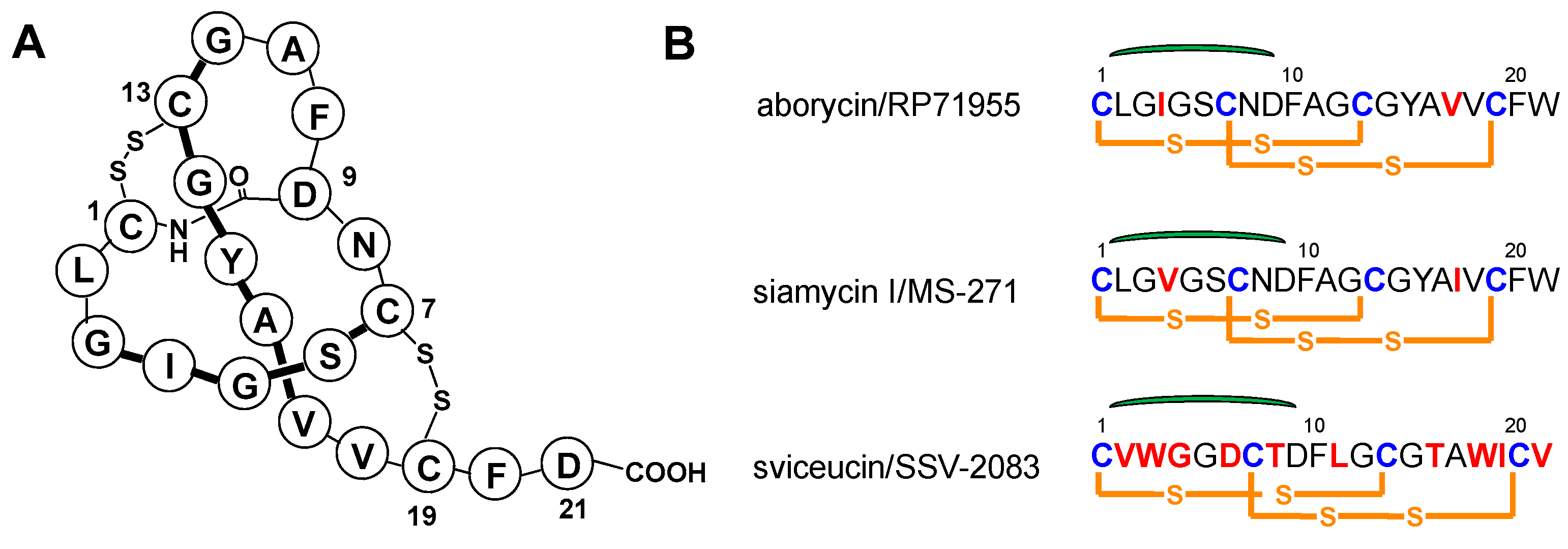
2.1. Characterization of Aborycin
2.2. Bioinformatics Analysis of Streptomyces sp. SCSIO ZS0098
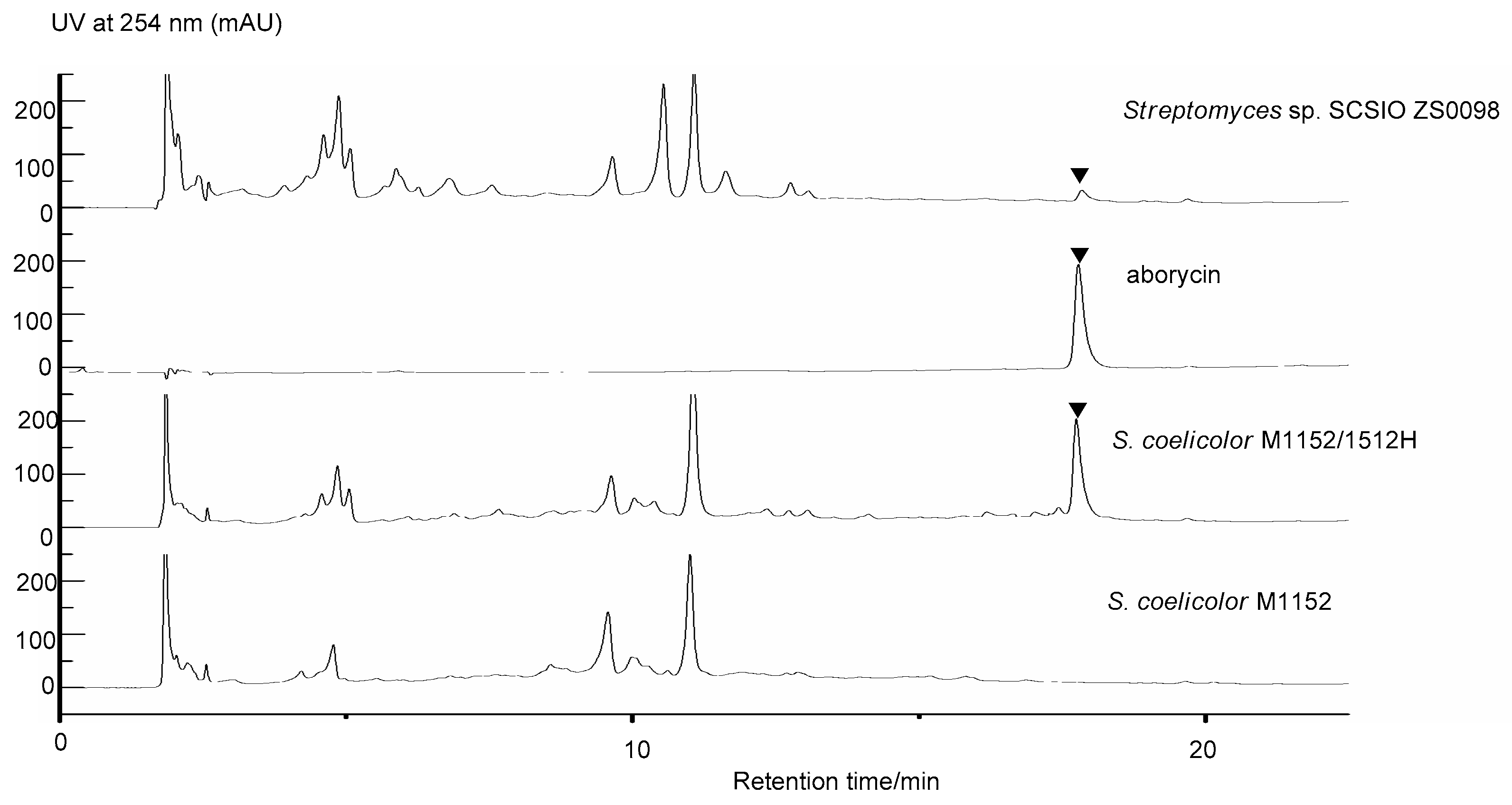
2.3. Heterologous Expression of the Candidate Aborycin Gene Cluster in S. coelicolor M1152
2.4. Antibacterial Activity of the Aborycin
3. Discussion
4. Materials and Methods
4.1. General Experimental Procedures
4.2. Nucleotide Sequence Accession Number
4.3. Bacterial Strains, Plasmids
4.4. Whole Genome Scanning and Bioinformatics Analysis
4.5. Genomic Library Construction and Screening
4.6. Construction of the Aborycin Gene Cluster Heterologous Expression Strains
4.7. Metabolite Analyses of WT and Recombinant Heterologous abo Cluster Expression Strains
4.8. Aborycin Isolation
4.9. Antibacterial Activities Assay
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Helynck, G.; Dubertret, C.; Mayaux, J.F.; Leboul, J. Isolation of RP 71955, a new anti-HIV-1 peptide secondary metabolite. J. Antibiot. 1993, 11, 1756–1757. [Google Scholar] [CrossRef]
- Potterat, O.; Stephan, H.; Metzger, J.W.; Gnau, V.; Zähner, H.; Jung, G. Aborycin–a tricyclic 21-peptide antibiotic isolated from Streptomyces griseoflavus. Eur. J. Org. Chem. 1994, 7, 741–743. [Google Scholar]
- Frechet, D.; Guitton, J.D.; Herman, F.; Faucher, D.; Helynck, G.; Monegier du Sorbier, B.; Vuilhorgne, M. Solution structure of RP 71955, a new 21 amino acid tricyclic peptide active against HIV-1 virus. Biochemistry 1994, 1, 42–50. [Google Scholar] [CrossRef]
- Maksimov, M.O.; Pan, S.J.; James Link, A. Lasso peptides: Structure, function, biosynthesis, and engineering. Nat. Prod. Rep. 2012, 9, 996–1006. [Google Scholar] [CrossRef] [PubMed]
- Feng, Z.; Ogasawara, Y.; Nomura, S.; Dairi, T. Biosynthetic gene cluster of a d-tryptophan-containing lasso peptide, MS-271. ChemBioChem 2018, 19, 2045–2048. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Ducasse, R.; Zirah, S.; Blond, A.; Goulard, C.; Lescop, E.; Rebuffat, S. Characterization of sviceucin from Streptomyces provides insight into enzyme exchangeability and disulfide bond formation in lasso peptides. ACS Chem. Biol. 2015, 11, 2641–2649. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Qin, X.; Liu, J.; Gui, C.; Wang, B.; Li, J.; Ju, J. Deciphering the biosynthetic origin of L-allo-isoleucine. J. Am. Chem. Soc. 2015, 1, 408–415. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Chen, Q.; Song, Y.; Huang, H.; Li, J.; Ma, J.; Li, Q.; Ju, J. Deciphering the sugar biosynthetic pathway and tailoring steps of nucleoside antibiotic A201A unveils a GDP-L-galactose mutase. Proc. Natl. Acad. Sci. USA 2017, 19, 4948–4953. [Google Scholar] [CrossRef] [PubMed]
- Ongley, S.E.; Bian, X.; Neilan, B.A.; Müller, R. Recent advances in the heterologous expression of microbial natural product biosynthetic pathways. Nat. Prod. Rep. 2013, 8, 1121–1138. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Huang, H.; Chen, Q.; Luo, M.; Sun, A.; Song, Y.; Ju, J. Identification of the grincamycin gene cluster unveils divergent roles for GcnQ in different hosts, tailoring the L-rhodinose moiety. Org. Lett. 2013, 15, 3254–3257. [Google Scholar] [CrossRef] [PubMed]
- Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically; Approved Standard, 8th ed.; Clinical and Laboratory Standards Institute: Villanova, PA, USA, 2009.
- Knappe, T.A.; Linne, U.; Robbel, L.; Marahiel, M.A. Insights into the biosynthesis and stability of the lasso peptide capistruin. Chem. Biol. 2009, 12, 1290–1298. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zirah, S.; Rebuffat, S. Lasso Peptides: Bacterial Strategies to Make and Maintain Bioactive Entangled Scaffolds; Springer: Berlin, Germany, 2014. [Google Scholar]
- Li, Q.; Song, Y.; Qin, X.; Zhang, X.; Sun, A.; Ju, J. Identification of the biosynthetic gene cluster for the anti-infective desotamides and production of a new analogue in a heterologous host. J. Nat. Prod. 2015, 4, 944–948. [Google Scholar] [CrossRef] [PubMed]
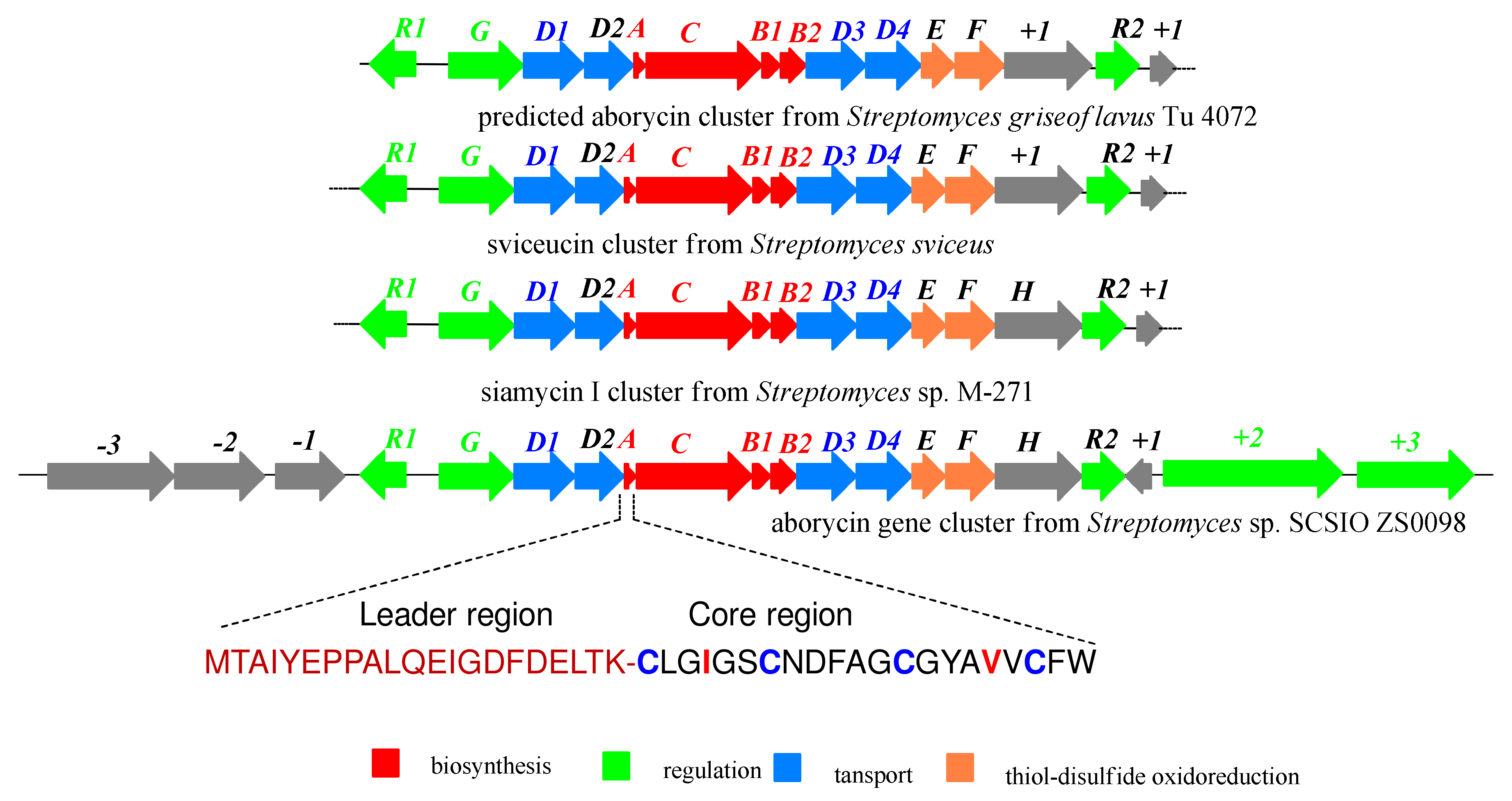
| ORF | Size a | Closest Protein Similarity | ID/SI (%) | Accession Numbers |
|---|---|---|---|---|
| orf(-3) | 688 | AAA-like domain protein putative ATPase | 34/50 | ALJ40332 |
| orf(-2) | 466 | nuclease | 26/44 | WP-045436186 |
| orf(-1) | 356 | D12 class N6 adenine-specific DNA methyltransferase | 44/55 | ACU76144 |
| aboR1 | 233 | helix-turn-helix transcriptional regulator | 99/99 | WP-053077004 |
| aboG | 369 | integral membrane sensor signal transduction histidine kinase | 72/81 | KPI21044 |
| aboD1 | 313 | multidrug ABC transporter ATP-binding protein | 74/82 | WP-059005878 |
| aboD2 | 250 | ABC transporter integral membrane protein | 44/61 | EDY58504 |
| aboA | 42 | aborycin family tricyclic lasso peptide | 98/100 | KPI21041 |
| aboC | 603 | lasso peptide isopeptide bond-forming cyclase | 99/99 | WP_114873958.1 |
| aboB1 | 87 | lasso peptide biosynthesis PqqD family chaperone | 100/100 | WP-037640857.1 |
| aboB2 | 141 | lasso peptide biosynthesis B2 protein | 99/99 | WP-048459500.1 |
| aboD3 | 321 | ABC transporter ATP-binding protein | 75/85 | KUN45102 |
| aboD4 | 289 | ABC transporter | 98/98 | WP-048459502 |
| aboE | 157 | DoxX family protein | 56/69 | KWV32827 |
| aboF | 241 | disulfide bond formation protein DsbA | 98/99 | WP-102640382.1 |
| aboH | 448 | poly-gamma-glutamate biosynthesis protein | 46/60 | WP-038038039 |
| aboR2 | 225 | DNA-binding response regulator | 99/99 | WP-048459506 |
| orf(+1) | 116 | Membrane protein involved in colicin uptake | 51/59 | CEL20868 |
| orf(+2) | 986 | SARP family transcriptional regulator | 92/93 | WP-052183254 |
| orf(+3) | 632 | transcriptional regulator | 99/99 | WP-048459508 |
| Pathogens | MIC (μg/mL) | ||||
|---|---|---|---|---|---|
| Aborycin | Ampicillin a | Kanamycin a | Ciprofloxacin a | Polymyxin B a | |
| Staphylococcus aureus ATCC 29213 | 8.0 | 16.0 | 2.0 | b NT | NT |
| MRSA/methicillin-resistant Staphylococcus aureus | 8.0 | >128 | >128 | NT | NT |
| MRSE/methicillin-resistant Staphylococcus epidermidis | 128 | >128 | >128 | NT | NT |
| Staphylococcus aureus Sau 29213 | 8.0 | 32.0 | 4.0 | NT | NT |
| Staphylococcus aureus Sau 1862 | 16 | >128 | 128.0 | NT | NT |
| Staphylococcus aureus Sau 669 | 32 | >128 | 1.0 | NT | NT |
| Staphylococcus aureus Sau 991 | 16 | >128 | 32.0 | NT | NT |
| Staphylococcus aureus 16339 | 16 | >128 | 2.0 | NT | NT |
| Staphylococcus aureus 6917 | 16 | >128 | >128 | NT | NT |
| Staphylococcus aureus 16162 | 64 | >128 | >128 | NT | NT |
| Staphylococcus aureus 718306 | 16.0 | >128 | 2.0 | NT | NT |
| Staphylococcus aureus 745524 | 16.0 | >128 | >128 | NT | NT |
| Staphylococcus aureus MRSA GDE4P037P | >128 | >128 | >128 | NT | NT |
| Staphylococcus aureus (cfr) GDQ6P012P | 64 | >128 | 4.0 | NT | NT |
| Staphylococcus cohnii DKG4 | >128 | >128 | >128 | NT | NT |
| Staphylococcus simulans AKA1 | >128 | >128 | >128 | NT | NT |
| Vibrio alginolyticus XSBZ14 | >128 | >128 | 4.0 | NT | NT |
| Enterococcus faecalis ATCC 29212 | 8.0 | >128 | >128 | NT | NT |
| Enterococcus gallinarum 5F52C | 0.5 | 8.0 | 8.0 | NT | NT |
| Clostridium perfringens FSKP20 | >128 | >128 | >128 | NT | NT |
| Micrococcus luteus | >128 | 2.0 | 0.5 | NT | NT |
| Bacillus thuringiensis | 2.0 | >128 | 64.0 | NT | NT |
| Escherichia coli 16369 | >128 | NT | NT | 32.0 | 4.0 |
| Escherichia coli 16447 | >128 | NT | NT | 0.25 | 4.0 |
| Escherichia coli 737720 | >128 | NT | NT | 8.0 | 4.0 |
| Escherichia coli 16351 | >128 | NT | NT | 64.0 | 4.0 |
| Escherichia coli ATCC 13124 | >128 | NT | NT | 32.0 | >128 |
| Escherichia coli (E11) | >128 | NT | NT | 8.0 | 2.0 |
| Acinetobacter baumannii ATCC 19606 | 128.0 | NT | NT | 8.0 | 4.0 |
| Acinetobacter baumannii 15122 | >128 | NT | NT | 32.0 | 4.0 |
| Acinetobacter baumannii 15407 | >128 | NT | NT | 32.0 | 4.0 |
| Acinetobacter baumannii 15199 | >128 | NT | NT | 32.0 | 4.0 |
| Acinetobacter baumannii 14892 | >128 | NT | NT | 16.0 | 4.0 |
| Klebsiella pneumoniae ATCC 13883 | >128 | NT | NT | 0.25 | 4.0 |
| Klebsiella pneumoniae 15580 | >128 | NT | NT | 4.0 | 2.0 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shao, M.; Ma, J.; Li, Q.; Ju, J. Identification of the Anti-Infective Aborycin Biosynthetic Gene Cluster from Deep-Sea-Derived Streptomyces sp. SCSIO ZS0098 Enables Production in a Heterologous Host. Mar. Drugs 2019, 17, 127. https://doi.org/10.3390/md17020127
Shao M, Ma J, Li Q, Ju J. Identification of the Anti-Infective Aborycin Biosynthetic Gene Cluster from Deep-Sea-Derived Streptomyces sp. SCSIO ZS0098 Enables Production in a Heterologous Host. Marine Drugs. 2019; 17(2):127. https://doi.org/10.3390/md17020127
Chicago/Turabian StyleShao, Mingwei, Juying Ma, Qinglian Li, and Jianhua Ju. 2019. "Identification of the Anti-Infective Aborycin Biosynthetic Gene Cluster from Deep-Sea-Derived Streptomyces sp. SCSIO ZS0098 Enables Production in a Heterologous Host" Marine Drugs 17, no. 2: 127. https://doi.org/10.3390/md17020127
APA StyleShao, M., Ma, J., Li, Q., & Ju, J. (2019). Identification of the Anti-Infective Aborycin Biosynthetic Gene Cluster from Deep-Sea-Derived Streptomyces sp. SCSIO ZS0098 Enables Production in a Heterologous Host. Marine Drugs, 17(2), 127. https://doi.org/10.3390/md17020127






