Genome-Wide Identification and Characterization of SODs in Zhikong Scallop Reveals Gene Expansion and Regulation Divergence after Toxic Dinoflagellate Exposure
Abstract
1. Introduction
2. Results and Discussion
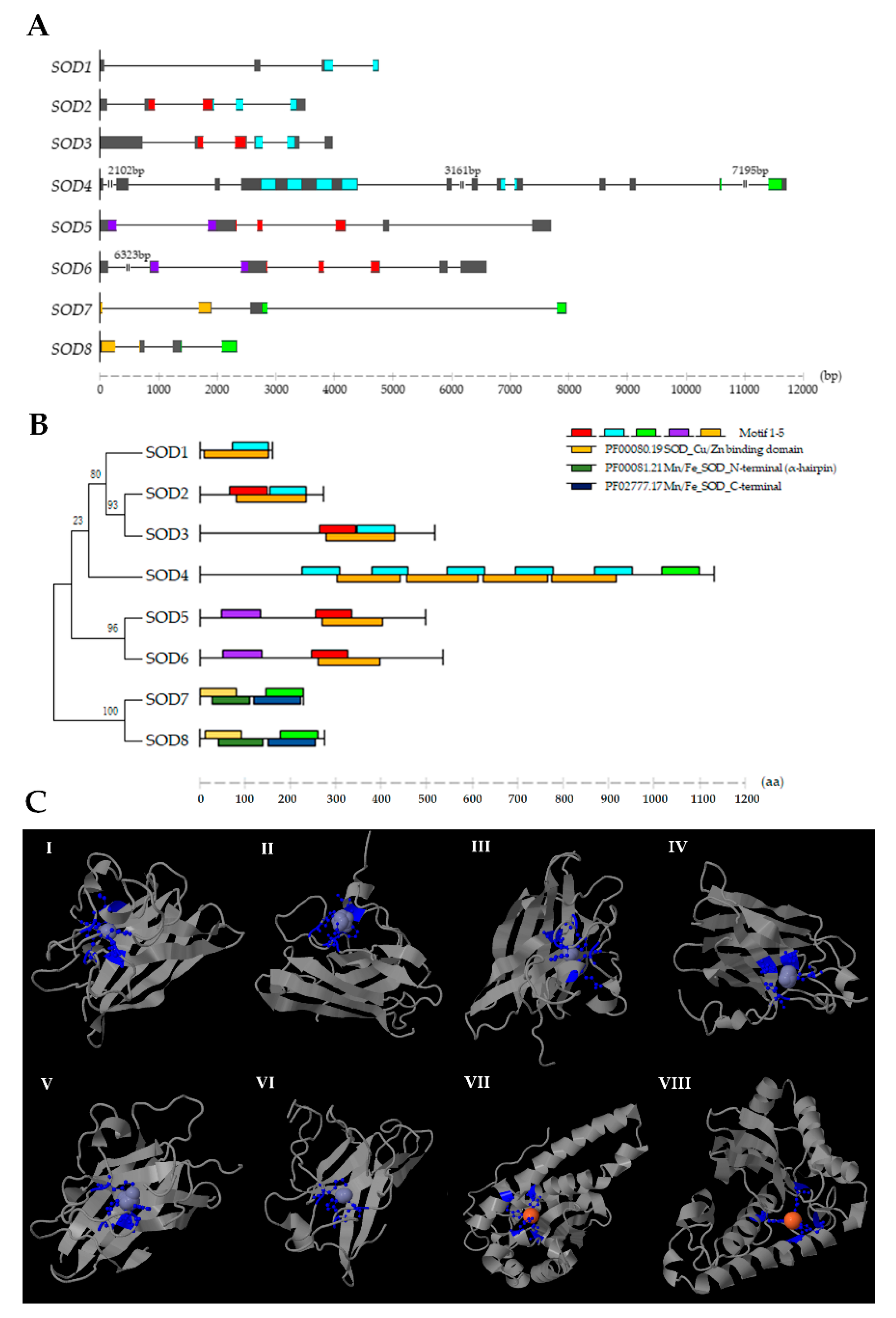
2.1. Identification of SOD Genes in C. farreri
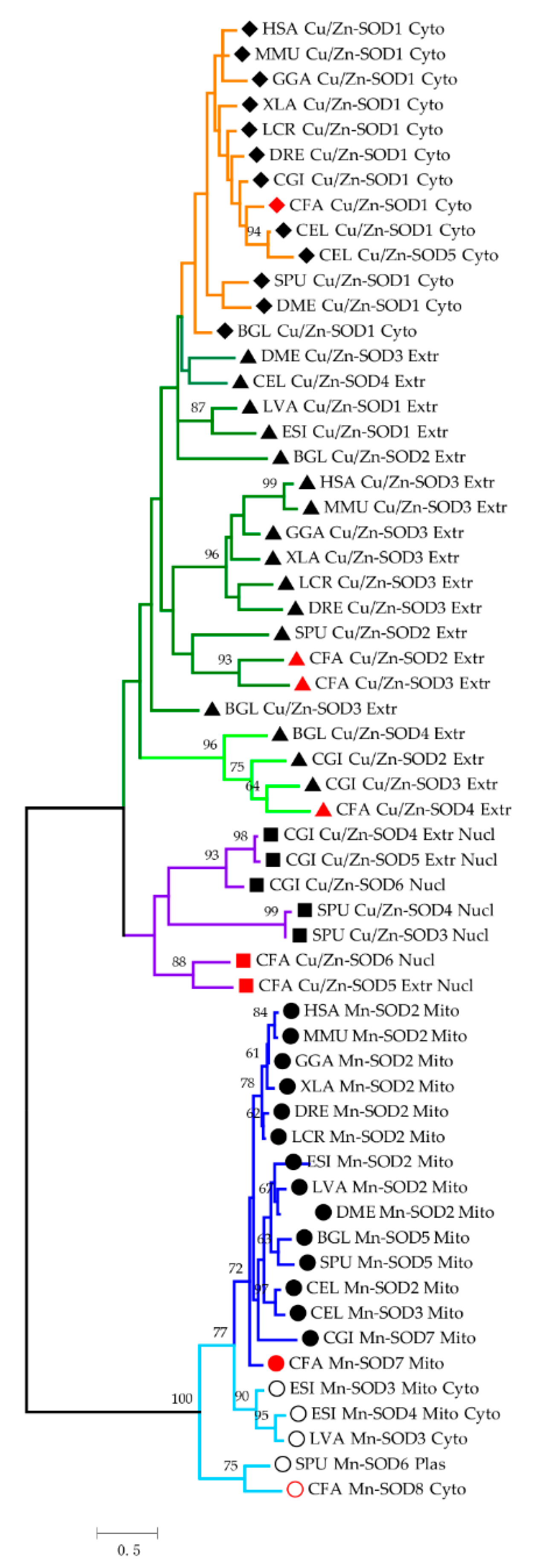
2.2. Phylogenetic Analysis of SODs
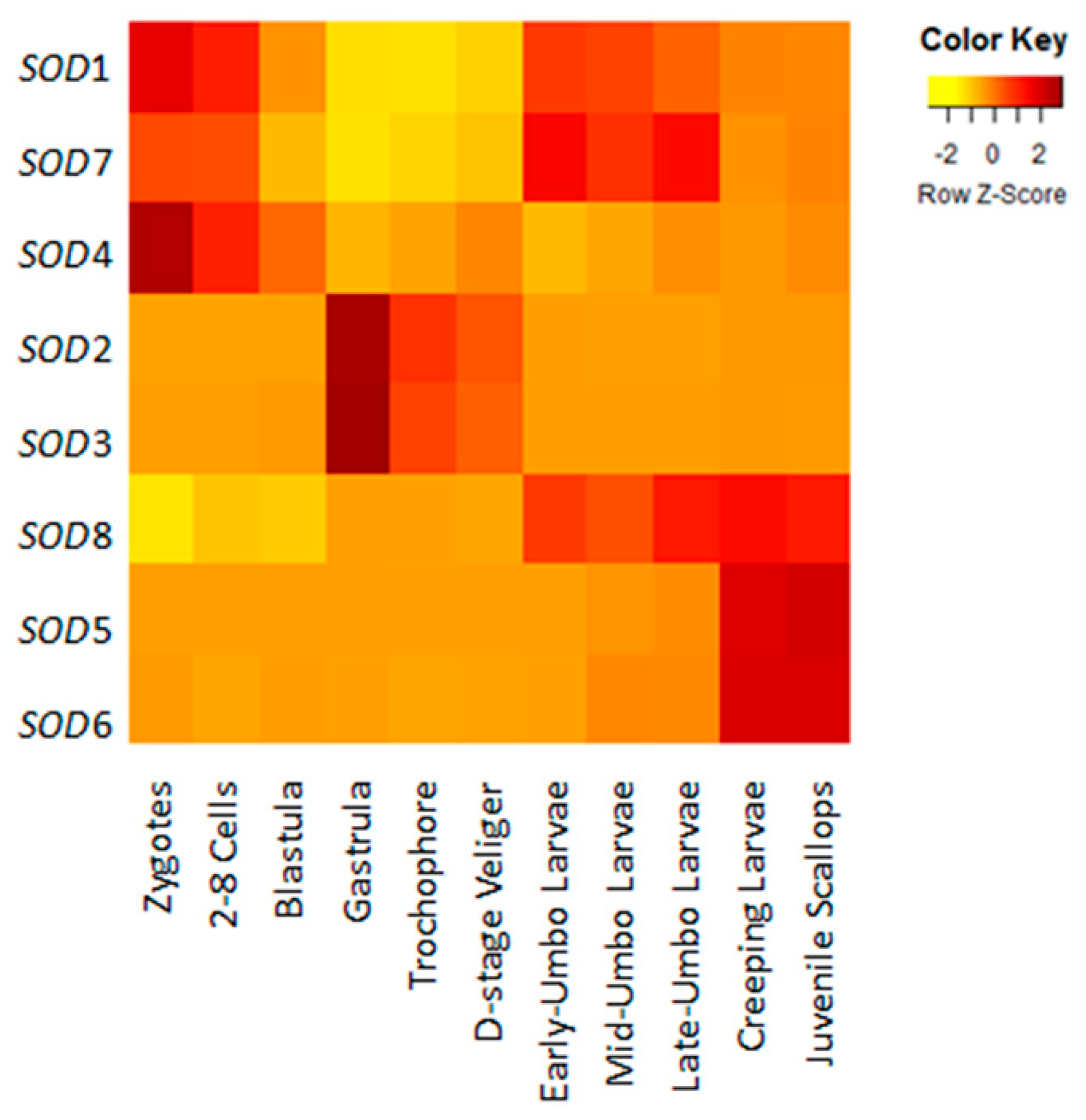
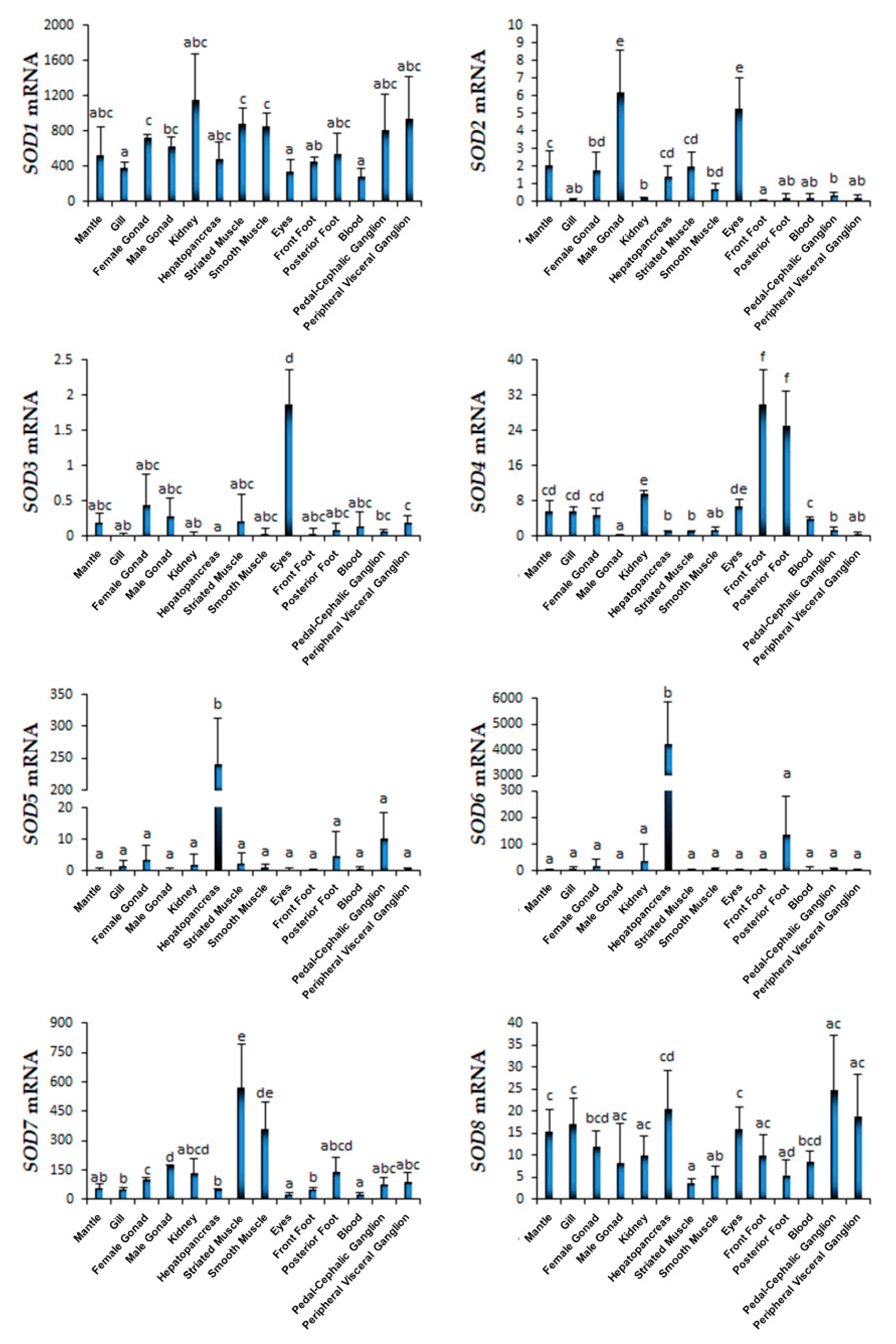
2.3. Spatiotemporal Expression Profiles of Scallop SODs During Development and in Adult Organs/Tissues
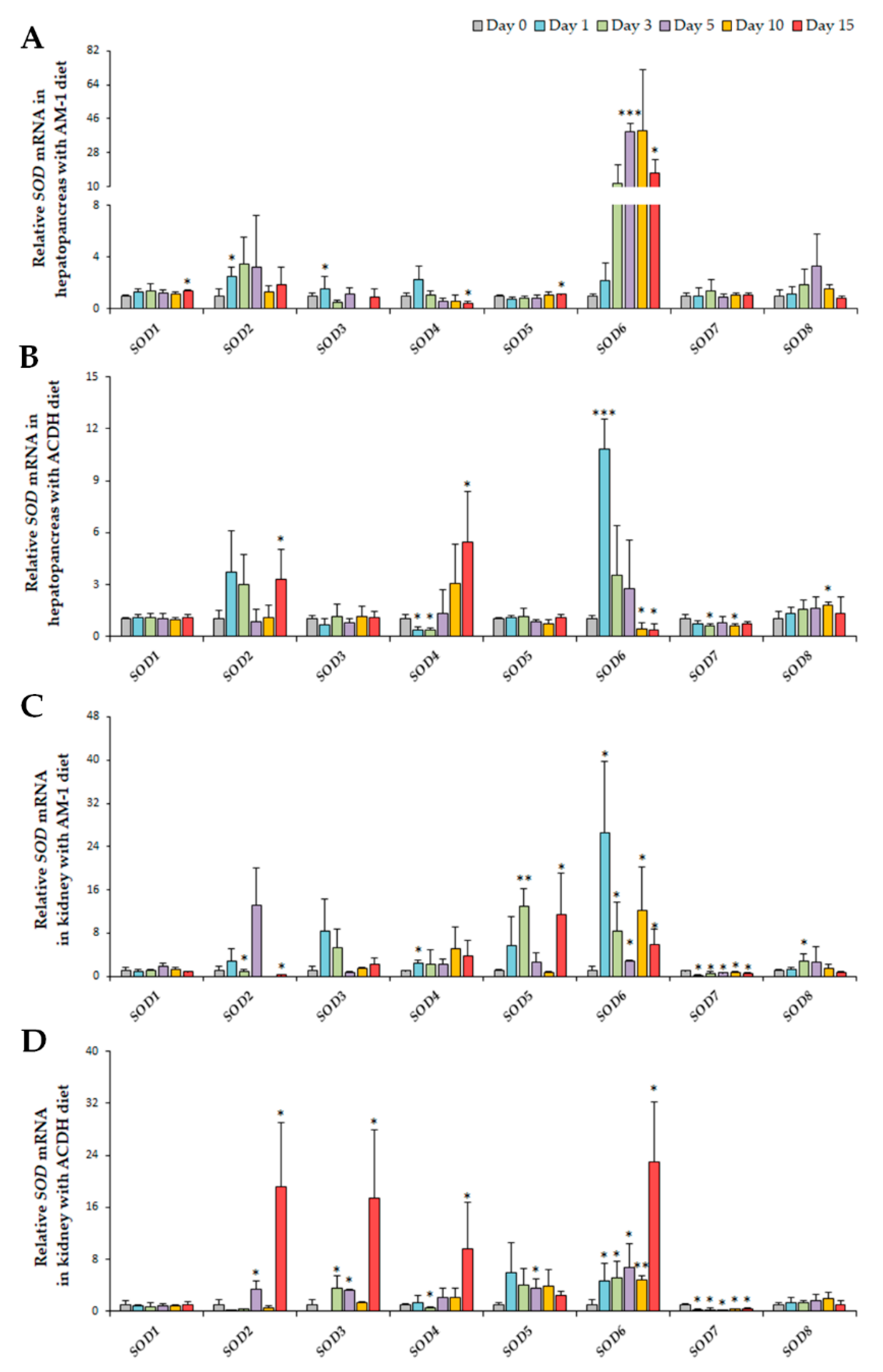
2.4. Diversified Expression Regulation of SODs in Response to PSTs Producers
3. Conclusions
4. Materials and Methods
4.1. Genome-Wide Identification and Sequence Analysis of SOD Genes in C. farreri
4.2. Multiple Alignment and Phylogenetic Analysis
4.3. Expression Profiles of SOD Genes During C. farreri Development and in Adult Organs/Tissues
4.4. Expression Response of SOD Genes to A. minutum and A. catenella Exposure
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Nathan, C.; Cunningham-Bussel, A. Beyond oxidative stress: An immunologist’s guide to reactive oxygen species. Nat. Rev. Immunol. 2013, 13, 349–361. [Google Scholar] [CrossRef] [PubMed]
- Van Raamsdonk, J.M.; Hekimi, S. Superoxide dismutase is dispensable for normal animal lifespan. Proc. Natl. Acad. Sci. USA 2012, 109, 5785–5790. [Google Scholar] [CrossRef] [PubMed]
- Zelko, I.N.; Mariani, T.J.; Folz, R.J. Superoxide dismutase multigene family: A comparison of the CuZn-SOD (SOD1), Mn-SOD (SOD2), and EC-SOD (SOD3) gene structures, evolution, and expression. Free Radic. Biol. Med. 2002, 33, 337–349. [Google Scholar] [CrossRef]
- Benzie, I.F.F. Evolution of antioxidant defence mechanisms. Eur. J. Nutr. 2000, 39, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Culotta, V.C.; Yang, M.; O’Halloran, T.V. Activation of superoxide dismutases: Putting the metal to the pedal. BBA Mol. Cell Res. 2006, 1763, 747–758. [Google Scholar] [CrossRef] [PubMed]
- Scandalios, J.G. Oxygen stress and superoxide dismutases. Plant Physiol. 1993, 101, 7–12. [Google Scholar] [CrossRef] [PubMed]
- Fantel, A.G.; Person, R.E. Involvement of mitochondria and other free radical sources in normal and abnormal fetal development. Ann. N. Y. Acad. Sci. 2002, 959, 424–433. [Google Scholar] [CrossRef]
- Barnham, K.J.; Masters, C.L.; Bush, A.I. Neurodegenerative diseases and oxidative stress. Nat. Rev. Drug Discov. 2004, 3, 205–214. [Google Scholar] [CrossRef]
- Keskin, I.; Birve, A.; Berdynski, M.; Hjertkvist, K.; Rofougaran, R.; Nilsson, T.K.; Glass, J.D.; Marklund, S.L.; Andersen, P.M. Comprehensive analysis to explain reduced or increased SOD1 enzymatic activity in ALS patients and their relatives. Amyotroph. Lat. Scler. Front. Degener. 2017, 18, 457–463. [Google Scholar] [CrossRef]
- Silverman, J.M.; Fernando, S.M.; Grad, L.I.; Hill, A.F.; Turner, B.J.; Yerbury, J.J.; Cashman, N.R. Disease Mechanisms in ALS: Misfolded SOD1 transferred through exosome-dependent and exosome-independent pathways. Cell Mol. Neurobiol. 2016, 36, 377–381. [Google Scholar] [CrossRef]
- McCord, J.M.; Edeas, M.A. SOD, oxidative stress and human pathologies: A brief history and a future vision. Biomed. Pharmacother. 2005, 59, 139–142. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.W.; Wang, Q.H.; Zhang, J.L.; Li, S.; Wang, X.L.; Xu, S.W. Effects of oxidative stress on immunosuppression induced by selenium deficiency in chickens. Biol. Trace Elem. Res. 2012, 149, 352–361. [Google Scholar] [CrossRef] [PubMed]
- Che, M.X.; Wang, R.; Li, X.X.; Wang, H.Y.; Zheng, X.F.S. Expanding roles of superoxide dismutases in cell regulation and cancer. Drug Discov. Today 2016, 21, 143–149. [Google Scholar] [CrossRef] [PubMed]
- Robbins, D.; Zhao, Y.F. Manganese superoxide dismutase in cancer prevention. Antioxid. Redox Sign. 2014, 20, 1628–1645. [Google Scholar] [CrossRef] [PubMed]
- Zhen, H.; Wen, M.; Yang, Y.; Can, Z.; Hui, G.; Li, X.; Deli, L. Toxic effects of HgCl2 on activities of SOD, AchE and relative expression of SOD, AChE, CYP1A1 of zebrafish. Ecotoxicology 2014, 23, 1842–1845. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.J.; Li, Y.L.; Zhang, L.B.; Zucker, I.H.; Gao, L.; Zimmerman, M.C.; Wang, W. Endogenous reactive oxygen species modulates voltage-gated sodium channels in dorsal root ganglia of rats. J. Appl. Physiol. 2011, 110, 1439–1447. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, A.; Ray, D.; Shrivastava, A.N.; Sarker, S. Molecular biomarkers: Their significance and application in marine pollution monitoring. Ecotoxicology 2006, 15, 333–340. [Google Scholar] [CrossRef] [PubMed]
- Philipp, E.E.R.; Wessels, W.; Gruber, H.; Strahl, J.; Wagner, A.E.; Ernst, I.M.A.; Rimbach, G.; Kraemer, L.; Schreiber, S.; Abele, D.; et al. Gene expression and physiological changes of different populations of the long-lived bivalve Arctica islandica under low oxygen conditions. PLoS ONE 2012, 7. [Google Scholar] [CrossRef]
- Maldonado-Aguayo, W.; Lafarga-De la Cruz, F.; Gallardo-Escarate, C. Identification and expression of antioxidant and immune defense genes in the surf clam Mesodesma donacium challenged with Vibrio anguillarum. Mar. Genomics 2015, 19, 65–73. [Google Scholar] [CrossRef]
- Carregosa, V.; Velez, C.; Soares, A.M.V.M.; Figueira, E.; Freitas, R. Physiological and biochemical responses of three Veneridae clams exposed to salinity changes. Comp. Biochem. Phys. B 2014, 177, 1–9. [Google Scholar] [CrossRef]
- Yu, J.H.; Song, J.H.; Choi, M.C.; Park, S.W. Effects of water temperature change on immune function in surf clams, Mactra veneriformis (Bivalvia: Mactridae). J. Invertebr. Pathol. 2009, 102, 30–35. [Google Scholar] [CrossRef] [PubMed]
- Anderson, D.M.; Cembella, A.D.; Hallegraeff, G.M. Progress in understanding harmful algal blooms: Paradigm shifts and new technologies for research, monitoring, and management. Annu. Rev. Mar. Sci. 2012, 4, 143–176. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.H.; Wu, J.Y. Occurrence and potential risks of harmful algal blooms in the East China Sea. Sci. Total Environ. 2009, 407, 4012–4021. [Google Scholar] [CrossRef]
- Shumway, S.E. A review of the effects of algal blooms on shellfish and aquaculture. J. World Aquacult. Soc. 1990, 21, 65–104. [Google Scholar] [CrossRef]
- Li, Y.L.; Sun, X.Q.; Hu, X.L.; Xun, X.G.; Zhang, J.B.; Guo, X.M.; Jiao, W.Q.; Zhang, L.L.; Liu, W.Z.; Wang, J.; et al. Scallop genome reveals molecular adaptations to semi-sessile life and neurotoxins. Nat. Commun. 2017, 8. [Google Scholar] [CrossRef]
- Whittle, K.; Gallacher, S. Marine toxins. Brit. Med. Bull. 2000, 56, 236–253. [Google Scholar] [CrossRef]
- Tran, D.; Haberkorn, H.; Soudant, P.; Ciret, P.; Massabuau, J.C. Behavioral responses of Crassostrea gigas exposed to the harmful algae Alexandrium minutum. Aquaculture 2010, 298, 338–345. [Google Scholar] [CrossRef]
- Manfrin, C.; De Moro, G.; Torboli, V.; Venier, P.; Pallavicini, A.; Gerdol, M. Physiological and molecular responses of bivalves to toxic dinoflagellates. ISJ Invertebr. Surviv. J. 2012, 9, 184–199. [Google Scholar]
- Bricelj, V.M.; Ford, S.E.; Lambert, C.; Barbou, A.; Paillard, C. Effects of toxic Alexandrium tamarense on behavior, hemocyte responses and development of brown ring disease in Manila clams. Mar. Ecol. Prog. Ser. 2011, 430, 35–48. [Google Scholar] [CrossRef][Green Version]
- Haberkorn, H.; Lambert, C.; Le Goic, N.; Gueguen, M.; Moal, J.; Palacios, E.; Lassus, P.; Soudant, P. Effects of Alexandrium minutum exposure upon physiological and hematological variables of diploid and triploid oysters, Crassostrea gigas. Aquat. Toxicol. 2010, 97, 96–108. [Google Scholar] [CrossRef]
- Cao, R.W.; Wang, D.; Wei, Q.Y.; Wang, Q.; Yang, D.L.; Liu, H.; Dong, Z.J.; Zhang, X.L.; Zhang, Q.Q.; Zhao, J.M. Integrative biomarker assessment of the influence of saxitoxin on marine bivalves: A comparative study of the two bivalve species oysters, Crassostrea gigas, and scallops, Chlamys farreri. Front. Physiol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.B.; Ma, F.F.; Fan, H.; Li, A.F. Effects of feeding Alexandrium tamarense, a paralytic shellfish toxin producer, on antioxidant enzymes in scallops (Patinopecten yessoensis) and mussels (Mytilus galloprovincialis). Aquaculture 2013, 396, 76–81. [Google Scholar] [CrossRef]
- Estrada, N.; Romero, M.D.; Campa-Cordova, A.; Luna, A.; Ascencio, F. Effects of the toxic dinoflagellate, Gymnodinium catenatum on hydrolytic and antioxidant enzymes, in tissues of the giant lions-paw scallop Nodipecten subnodosus. Comp. Biochem. Phys. C 2007, 146, 502–510. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Ma, J.; Cao, J.; Wang, Q.; Yu, R.; Thomas, K.; Quilliam, M.A. Analysis of paralytic shellfish toxins and their metabolites in shellfish from the North Yellow Sea of China. Food Addit. Contam. A 2012, 29, 1455–1464. [Google Scholar] [CrossRef]
- Ebersbach, G.; Galli, E.; Moller-Jensen, J.; Lowe, J.; Gerdes, K. Novel coiled-coil cell division factor ZapB stimulates Z ring assembly and cell division. Mol. Microbiol. 2008, 68, 720–735. [Google Scholar] [CrossRef]
- Bao, Y.B.; Li, L.; Zhang, G.F. The manganese superoxide dismutase gene in bay scallop Argopecten irradians: Cloning, 3D modelling and mRNA expression. Fish Shellfish Immun. 2008, 25, 425–432. [Google Scholar] [CrossRef]
- Brouwer, M.; Brouwer, T.H.; Grater, W.; Brown-Peterson, N. Replacement of a cytosolic copper/zinc superoxide dismutase by a novel cytosolic manganese superoxide dismutase in crustaceans that use copper (haemocyanin) for oxygen transport. Biochem. J. 2003, 374, 219–228. [Google Scholar] [CrossRef]
- Brouwer, M.; Brouwer, T.H.; Grater, W.; Enghild, J.J.; Thogersen, I.B. The paradigm that all oxygen-respiring eukaryotes have cytosolic CuZn-superoxide dismutase and that Mn-superoxide dismutase is localized to the mitochondria does not apply to a large group of marine arthropods. Biochemistry 1997, 36, 13381–13388. [Google Scholar] [CrossRef]
- Gomez-Anduro, G.A.; Barillas-Mury, C.V.; Peregrino-Uriarte, A.B.; Gupta, L.; Gollas-Galvan, T.; Hernandez-Lopez, J.; Yepiz-Plascencia, G. The cytosolic manganese superoxide dismutase from the shrimp Litopenaeus vannamei: Molecular cloning and expression. Dev. Comp. Immunol. 2006, 30, 893–900. [Google Scholar] [CrossRef]
- Wang, M.Q.; Wang, L.L.; Yi, Q.L.; Gai, Y.L.; Song, L.H. Molecular cloning and characterization of a cytoplasmic manganese superoxide dismutase and a mitochondrial manganese superoxide dismutase from Chinese mitten crab Eriocheir sinensis. Fish Shellfish Immun. 2015, 47, 407–417. [Google Scholar] [CrossRef]
- Rodriguez-Trelles, F.; Tarrio, R.; Ayala, F.J. Erratic overdispersion of three molecular clocks: GPDH, SOD, and XDH. Proc. Natl. Acad. Sci. USA 2001, 98, 11405–11410. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.W.; Doolittle, R.F. A comparison of evolutionary rates of the two major kinds of superoxide-dismutase. J. Mol. Evol. 1992, 34, 175–184. [Google Scholar] [CrossRef] [PubMed]
- Dandapat, J.; Chainy, G.B.N.; Rao, K.J. Lipid peroxidation and antioxidant defence status during larval development and metamorphosis of giant prawn, Macrobrachium rosenbergii. Comp. Biochem. Phys. C 2003, 135, 221–233. [Google Scholar] [CrossRef]
- Kalaimani, N.; Chakravarthy, N.; Shanmugham, R.; Thirunavukkarasu, A.R.; Alavandi, S.V.; Santiago, T.C. Anti-oxidant status in embryonic, post-hatch and larval stages of Asian seabass (Lates calcarifer). Fish Physiol. Biochem. 2008, 34, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Menon, J.; Roman, R. Oxidative stress, tissue remodeling and regression during amphibian metamorphosis. Comp. Biochem. Phys. C 2007, 145, 625–631. [Google Scholar] [CrossRef]
- Hilliker, A.J.; Duyf, B.; Evans, D.; Phillips, J.P. Urate-null rosy mutants of Drosophila-melanogaster are hypersensitive to oxygen stress. Proc. Natl. Acad. Sci. USA 1992, 89, 4343–4347. [Google Scholar] [CrossRef]
- Dhage, P.A.; Kamble, L.K.; Bhargava, S.Y. Localization and distribution of superoxide dismutase-1 in the neural tube morphogenesis of chick embryo. Int. J. Dev. Neurosci. 2017, 56, 1–9. [Google Scholar] [CrossRef]
- Nonogaki, T.; Noda, Y.; Narimoto, K.; Umaoka, Y.; Mori, T. Effects of superoxide dismutase on mouse in vitro fertilization and embryo culture system. J. Assist. Reprod. Genet. 1992, 9, 274–280. [Google Scholar] [CrossRef]
- Behndig, A.; Karlsson, K.; Brannstrom, T.; Sentman, M.L.; Marklund, S.L. Corneal endothelial integrity in mice lacking extracellular superoxide dismutase. Investig. Ophthalmol. Vis. Sci. 2001, 42, 2784–2788. [Google Scholar] [PubMed]
- Behndig, A.; Karlsson, K.; Johansson, B.O.; Brannstrom, T.; Marklund, S.L. Superoxide dismutase isoenzymes in the normal and diseased human cornea. Investig. Ophthalmol. Vis. Sci. 2001, 42, 2293–2296. [Google Scholar] [PubMed]
- Mruk, D.; Cheng, C.H.; Cheng, Y.H.; Mo, M.Y.; Grima, J.; Silvestrini, B.; Lee, W.M.; Cheng, C.Y. Rat testicular extracellular superoxide dismutase: Its purification, cellular distribution, and regulation. Biol. Reprod. 1998, 59, 298–308. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Li, S.G.; Huang, J.L.; Liu, Y.J.; Jia, G.C.; Xie, L.P.; Zhang, R.Q. Extensible byssus of Pinctada fucata: Ca2+-stabilized nanocavities and a thrombospondin-1 protein. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Lustgarten, M.S.; Jang, Y.C.; Liu, Y.H.; Muller, F.L.; Qi, W.B.; Steinhelper, M.; Brooks, S.V.; Larkin, L.; Shimizu, T.; Shirasawa, T.; et al. Conditional knockout of Mn-SOD targeted to type IIB skeletal muscle fibers increases oxidative stress and is sufficient to alter aerobic exercise capacity. Am. J. Physiol. Cell Physiol. 2009, 297, C1520–C1532. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef]
- Yu, C.S.; Chen, Y.C.; Lu, C.H.; Hwang, J.K. Prediction of protein subcellular localization. Proteins 2006, 64, 643–651. [Google Scholar] [CrossRef]
- Chou, K.C.; Shen, H.B. A new method for predicting the subcellular localization of eukaryotic proteins with both single and multiple sites: Euk-mPLoc 2.0. PLoS ONE 2010, 5. [Google Scholar] [CrossRef]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef]
- Kosugi, S.; Hasebe, M.; Matsumura, N.; Takashima, H.; Miyamoto-Sato, E.; Tomita, M.; Yanagawa, H. Six classes of nuclear localization signals specific to different binding grooves of importin alpha. J. Biol. Chem. 2009, 284, 478–485. [Google Scholar] [CrossRef]
- Fukasawa, Y.; Tsuji, J.; Fu, S.C.; Tomii, K.; Horton, P.; Imai, K. MitoFates: Improved prediction of mitochondrial targeting sequences and their cleavage sites. Mol. Cell Proteomics 2015, 14, 1113–1126. [Google Scholar] [CrossRef]
- Kelley, L.A.; Sternberg, M.J.E. Protein structure prediction on the Web: A case study using the Phyre server. Nat. Protoc. 2009, 4, 363–371. [Google Scholar] [CrossRef]
- Wass, M.N.; Kelley, L.A.; Sternberg, M.J.E. 3DLigandSite: Predicting ligand-binding sites using similar structures. Nucleic Acids Res. 2010, 38, W469–W473. [Google Scholar] [CrossRef] [PubMed]
- Ferre, F.; Clote, P. DiANNA 1.1: An extension of the DiANNA web server for ternary cysteine classification. Nucleic Acids Res. 2006, 34, W182–W185. [Google Scholar] [CrossRef] [PubMed]
- Carugo, O.; Djinovic-Carugo, K. A proteomic Ramachandran plot (PRplot). Amino Acids 2013, 44, 781–790. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Hu, B.Y.; Li, M.L.; Yu, X.H.; Xun, X.G.; Lu, W.; Li, X.; Li, Y.J.; Lou, J.R.; Wang, S.; Zhang, L.L.; et al. Diverse expression regulation of Hsp70 genes in scallops after exposure to toxic Alexandrium dinoflagellates. Chemosphere 2019, 234, 62–69. [Google Scholar] [CrossRef]
- Xun, X.; Cheng, J.; Wang, J.; Li, Y.; Li, X.; Li, M.; Lou, J.; Kong, Y.; Bao, Z.; Hu, X. Solute carriers in scallop genome: Gene expansion and expression regulation after exposure to toxic dinoflagellate. Chemosphere 2019, 241, 124968. [Google Scholar] [CrossRef]
- Hwang, D.F.; Lu, Y.H. Influence of environmental and nutritional factors on growth, toxicity, and toxin profile of dinoflagellate Alexandrium minutum. Toxicon 2000, 38, 1491–1503. [Google Scholar] [CrossRef]
- Navarro, J.M.; Munoz, M.G.; Contreras, A.M. Temperature as a factor regulating growth and toxin content in the dinofiagellate Alexandrium catenella. Harmful Algae 2006, 5, 762–769. [Google Scholar] [CrossRef]
| Gene Characteristics | Protein Characteristics | Sub-Cellular Localization Prediction | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Gene Name | NCBI ID | Genomic Position | ORF (bp) | Length (aa) | MW (kDa) | pI | Instability Index | Aliphatic Index | GRAVY | CELLO v.2.5 | Euk-mPLoc 2.0 | WoLF PSORT II | Final |
| Cu/Zn-SOD1 | MK374365 | 50981.1: 5129-13060(+) | 462 | 153 | 15615.35 | 6.1 | 7.98 | 75.16 | −0.292 | Cyto | Cyto | Cyto | Cyto |
| Cu/Zn-SOD2 | MK374366 | 9607.44: 811894-816438(+) | 855 | 284 | 30094.36 | 5.33 | 32.34 | 67.89 | −0.341 | Extr | Chloro | Extr | Extr |
| Cu/Zn-SOD3 | MK374367 | 11157.31: 577292-581362(−) | 1533 | 510 | 54202.24 | 9.2 | 34.37 | 44.73 | −0.712 | Extr | Extr | Extr | Extr |
| Cu/Zn-SOD4 | MK374368 | 39551.24: 361068-385243(−) | 3348 | 1115 | 120753.05 | 8.82 | 37.55 | 76.36 | −0.254 | Extr | Extr | Nucl | Extr |
| Cu/Zn-SOD5 | MK374369 | 62345.9: 206833-214902(−) | 1455 | 484 | 55240.85 | 5.57 | 35.29 | 58.99 | −1.26 | Nucl | Extr | Nucl | Nucl |
| Cu/Zn-SOD6 | MK374370 | 10699.65: 1421918-1435004(+) | 1572 | 523 | 58628.12 | 5.58 | 36.05 | 53.31 | −1.283 | Nucl | Chloro | Nucl | Nucl |
| Mn-SOD7 | MK374371 | 62487.8: 111489-119434(+) | 681 | 226 | 25020.53 | 6.44 | 38.82 | 88.45 | −0.16 | Mito | Mito | Mito | Mito |
| Mn-SOD8 | MK374372 | 20055.26: 792736-795083(+) | 789 | 262 | 30332.28 | 4.94 | 41.32 (un) | 87.06 | −0.346 | Cyto | Chloro | Cyto | Cyto |
| Motif | Width (aa) | Consensus Sequence | Pfam Domain |
|---|---|---|---|
| 1 | 80 | EHEL[HY]A[HY]CECM[PF]N[KM][TQ]TPDIA[GY]KI[DR][GF]H[QI]ELTGEPHEDEVSIVYNL[PR]NLKPDTEHGIHIHEYGD[VM]GRCC[DY]SLGPHYNPTHKS | sod_Cu/Zn PF00080.19 |
| 2 | 80 | R[DN]E[GM][GY]E[LE]RHYGDLGN[VM]RQDGKGVV[KM][TW]D[IF]VDKLLPL[SR]GP[TR]SV[LI]GRS[VM]VIHYD[NR]DD[LM]G[KR]GGN[VA][VM]S[LY][TQ]TGNAGT[RP]LAC[AC]VIAR | sod_Cu/Zn PF00080.19 |
| 3 | 80 | G[GC]WGWLSRNPIS[NK]RPLVASKPT[NY]RPLQPT[DA][GE]LQPIFGIDVWEHAYY[LI][QK]YKNIRPKYVK[DR]WWNIVNW[DR][GK]V[VA]QF[GD]HW[AW][KY][GR]PC | Sod_Mn/Fe_C PF02777.17 |
| 4 | 85 | CCCFAIPEFDPKPENM[DK][LM]KWFGALVALALLNPN[PE]G[DK][GQ]RRKRQRRSTAVVDEIAELKAKVEKLEEQVNKLEEKDLGVHIHIGND[HY]H | ZapB PF06005.11 |
| 5 | 81 | WLLAFCYFIKVMPKKM[GP][AY]E[GF]IFDQRLN[HY]VLP[DK]LPYDFKDLEPFIDEEIMRIH[HY]L[GK]HHAAYVKK[LM]NIAEEKWAEDMEVKNV[NM] | Sod_Mn/Fe_N PF00081.21 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lian, S.; Zhao, L.; Xun, X.; Lou, J.; Li, M.; Li, X.; Wang, S.; Zhang, L.; Hu, X.; Bao, Z. Genome-Wide Identification and Characterization of SODs in Zhikong Scallop Reveals Gene Expansion and Regulation Divergence after Toxic Dinoflagellate Exposure. Mar. Drugs 2019, 17, 700. https://doi.org/10.3390/md17120700
Lian S, Zhao L, Xun X, Lou J, Li M, Li X, Wang S, Zhang L, Hu X, Bao Z. Genome-Wide Identification and Characterization of SODs in Zhikong Scallop Reveals Gene Expansion and Regulation Divergence after Toxic Dinoflagellate Exposure. Marine Drugs. 2019; 17(12):700. https://doi.org/10.3390/md17120700
Chicago/Turabian StyleLian, Shanshan, Liang Zhao, Xiaogang Xun, Jiarun Lou, Moli Li, Xu Li, Shi Wang, Lingling Zhang, Xiaoli Hu, and Zhenmin Bao. 2019. "Genome-Wide Identification and Characterization of SODs in Zhikong Scallop Reveals Gene Expansion and Regulation Divergence after Toxic Dinoflagellate Exposure" Marine Drugs 17, no. 12: 700. https://doi.org/10.3390/md17120700
APA StyleLian, S., Zhao, L., Xun, X., Lou, J., Li, M., Li, X., Wang, S., Zhang, L., Hu, X., & Bao, Z. (2019). Genome-Wide Identification and Characterization of SODs in Zhikong Scallop Reveals Gene Expansion and Regulation Divergence after Toxic Dinoflagellate Exposure. Marine Drugs, 17(12), 700. https://doi.org/10.3390/md17120700




