Abstract
The microorganisms that evolved at low temperatures express cold-adapted enzymes endowed with unique catalytic properties in comparison to their mesophilic homologues, i.e., higher catalytic efficiency, improved flexibility, and lower thermal stability. Cold environments are therefore an attractive research area for the discovery of enzymes to be used for investigational and industrial applications in which such properties are desirable. In this work, we will review the literature on cold-adapted enzymes specifically focusing on those discovered in the bioprospecting of polar marine environments, so far largely neglected because of their limited accessibility. We will discuss their existing or proposed biotechnological applications within the framework of the more general applications of cold-adapted enzymes.
1. Introduction
The economic progress in the “Era of Biotech” demands new biocatalysts for a wide range of applications, from therapy to industrial manufacturing. Particularly, biocatalysis is an attractive alternative to chemical synthesis, as biocatalysts are biodegradable and non-toxic, they originate from renewable sources, they have high selectivity and they provide products in high yields [1]. Up to 40% of the industrially relevant chemical reactions that require organic solvents harmful to the environment could be substituted by enzymatic catalysis by 2030 [2]. Many industrial processes already benefit from the use of enzymes (e.g., proteases, amylases, cellulases, carboxymethylcellulases, xylanases) in the production of pharmaceuticals, foods, beverages, confectionery, paper, as well as in textile and leather processing and waste-water treatment. Most of these enzymes are microbial in origin because they are relatively more thermostable than their counterparts from plants and animals [3]. Their market is expected to reach $7.0 billion by 2023 from $5.5 billion in 2018, with an annual growth rate of 4.9% for the period 2018–2023 [4]. In this market, industrial detergents will reach $10.8 billion by 2022, with an annual growth rate of 4.2% for the period 2017–2022 [4], whereas the global market for enzymes in the food and beverages industries is expected to grow from $1.8 billion in 2017 to nearly $2.2 billion by 2022, with an annual growth rate of 4.6% from 2017 to 2022 [4].
The increasing demand for new or better biocatalysts with different substrate selectivity, chiral selectivity, stability and activity at various pH and temperatures is likely to be met through the investigation of microorganisms, the largest reservoir of biological activities in Nature. Marine microorganisms are particularly promising in this respect and it is predicted [5] that the global market for marine biotechnology, estimated at $4.1 billion in 2015, may reach $4.8 billion by 2020 and $6.4 billion by 2025, with the identification of new marine-derived enzymes [6]. The bioprospecting of marine environments, i.e., the systematic search for new bioactive compounds or biomolecules, has been traditionally focused on temperate/tropical latitudes [7], and polar marine environments have been mostly neglected, mainly due to their limited accessibility. These habitats include not only seawater but also sediments and sea ice, where internal fluids remain liquid in winter. Although both polar regions contain freezing seas, their microorganisms have different evolutionary histories due to their different geography and land-sea distribution [8]. The northern polar region is characterized by extensive, shallow shelves of landmasses that surround a partially land-locked ocean [9], whereas the Antarctic region consists in a dynamic open ocean that surrounds the continent [10,11]. The opening of the Drake Passage between Tierra del Fuego and the Antarctic Peninsula 23.5–32.5 million years ago was the key event for the development of the Antarctic Circumpolar Current (ACC), partially responsible for cooling of the Antarctic waters. The Antarctic Polar Front, the northern boundary of the ACC, promoted the isolation of the Antarctic marine fauna [10,11]. In both polar regions, the constantly low temperatures have driven the evolution of cold-adapted microorganisms, which are classified as psychrophilic and psychrotolerant according to the physiological features of their growth [12]. In this review, the general term “cold-adapted” will be used to indicate “psychrophile” organisms indigenous to cold environments [13].
In view of recent breakthroughs in sampling methodologies, sequencing, and bioinformatics, polar marine environments have recently come into the scientific spotlight—also because of growing concerns about their role in global climate change dynamics. An increasing number of works demonstrated their biological diversity [14,15,16,17,18,19], which includes bacteria, archaea, yeasts, fungi and algae [20,21]. Among polar microorganisms, those defined as ‘cold-adapted’ are those that thrive in permanently cold environments, even at subzero temperatures in super-cooled liquid water, and that evolved the physiological and biochemical capability to survive and reproduce under these extreme conditions [22]. With a better knowledge of marine microorganisms of polar regions, previously inaccessible bio-products, both in terms of new bioactive metabolites and proteins/enzymes with potential commercial applications, have been described [23,24]. Their biotechnological use can take many forms, from agricultural production to industrial processes, food chemistry [25], synthetic biology, and biomedical uses [26,27]. It is expected that, in the near future, more cold-adapted bacteria and their enzymes will find their way to biotechnological applications.
In this review, we will describe recent findings in enzymes isolated in either Arctic or Antarctic regions, some of which are of potential biotechnological interest. Unlike previous reviews, we particularly focused on enzymes isolated from marine polar environments. Cold-adapted enzymes from non-marine environments (i.e., from soil or lakes) or from non-polar sources (i.e., deep sea) will be reported to suggest potential future applications for enzymes isolated from marine polar microorganisms or to highlight structure-function relationships common to all cold-adapted enzymes, regardless of their origin.
2. Methods for Enzyme Discovery and Engineering
The identification of enzymes from cultured or uncultured marine microorganisms have been carried out through several ‘-omics’ techniques. Additionally, natural enzymes have been optimized through enzyme engineering to improve their biotechnological properties (Figure 1).
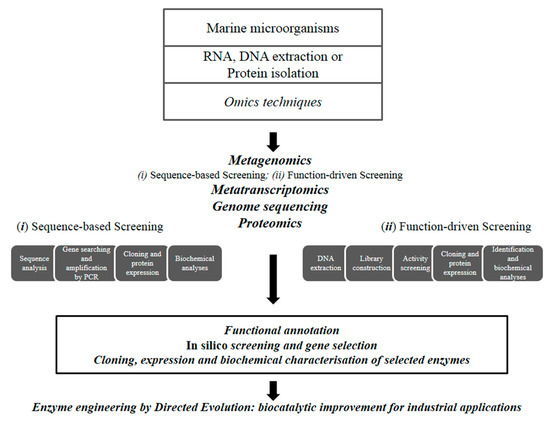
Figure 1.
Workflow for the discovery of novel industrial enzymes by omics technologies.
Omics techniques. The overall knowledge on polar microorganisms has recently increased through the application of “omic” approaches (environmental shotgun sequencing, metatranscriptomics, proteomics) that have revealed the peculiar properties of their communities [28].
High-throughput metagenomic screening approaches, using both sequence-based and function-driven screenings, are contributing to the identification of a large number of enzymes, with most industrial enzymes having being identified by traditional functional screening of (meta)genomic libraries [29,30,31]. Recently, rapid technological developments in bioinformatics have revolutionized the exploration of the microbial diversity for the discovery of enzymes with commercial impact [32]. The sequence-based analysis of their (meta)genomic DNA by high-throughput sequencing methods followed by in silico analysis was used to predict enzyme function [33] allowing an increase in the rate of discovery. Sequence-based metagenomic approaches represents a powerful tool by avoiding the intensive, expensive, and time-consuming lab work associated to classic screening by reducing the number of targets to be functionally tested. Nevertheless, the latter approach is only limited to the discovery of genes with high similarities to already deposited sequences, making the discovery of new enzymatic functions difficult [34].
Some recent EU FP7-funded projects (e.g., MACUMBA and PHARMASEA) made valuable efforts in exploiting the potential of microorganisms (mostly bacteria and microalgae) for pharmaceutical, cosmeceutical, and nutraceutical applications [35]. In order to explore marine biodiversity in several environments, including extreme habitats, a pan-European consortium was instituted in 2015 [36] for the creation of one of the largest collections of marine enzymes, with a focus on those of interest for the industrial market [32]. At present, this collection contains about 1000 enzymes, of which around 94% are available in ready-to-use expression systems and 32% have been completely characterized [2,32,37]. However, the publication of gene/genome sequences is faster than the identification of new protein functions, the most important challenge in modern biology due to the lack of experimental evidence to support functional annotation of a large fraction of genes and proteins [38,39]. Indeed, there is a gap between the numbers of biological activities predicted in silico and the number of new enzymes described experimentally, with the proportion of the latter ones approaching 0% [38,39], despite successful approaches of enzyme screen programmes through metagenomic approaches [40].
In addition to genomic screenings, recent technological developments in the complementary fields of transcriptomics have developed tools suitable for further discoveries. Indeed, advances within the field of RNA sequencing [41] have allowed analysis of whole (meta)transcriptomes and made them available for the generation of high-quality protein sequence databases, facilitating enzyme identification. The functional annotation of selected genomes and transcriptomes is relevant to infer associated biological functions and application of proteomics studies in the discovery of bioactive proteins.
As for proteomics approaches, the most commonly used method for enzyme discovery is bottom-up shotgun proteomics [42], in which proteins are enzymatically digested and the obtained peptides are analysed either by liquid chromatography coupled to electrospray ionization (ESI–LC–MS/MS) or by matrix-assisted laser desorption/ionization followed by time of flight mass spectrometry (MALDI–TOF–MS). A further direct method for enzyme discovery is activity-based proteomics, which takes advantage of enzyme class-specific substrates for the identification of groups of enzymes [43,44].
Enzyme Engineering
The ultimate value of biotechnology is the production and delivery of molecules or processes of interest for many applications; the emerging requirement for biotechnological products is to be energy saving, thus reducing environmental impacts [45]. Therefore, enzymes from cold-adapted organisms are important because their properties meet the ongoing efforts and needs to decrease energy consumption [46].
The ability of enzymes to catalyze a diverse set of reactions with exquisite specificity makes these proteins essential for reactions useful to humankind. Due their structural and functional complexity, the use of natural enzymes in biotechnology has been limited over the past decades. In fact, natural enzymes generally require modification for industrial use [47], given the improvements in catalytic efficiency, stereo-selectivity, and stability that can be reached through combinatorial engineering strategies [48]. Recent advances in science and technology have led to activity improvements for native and non-native substrates, synthesis of new chemistries and functional characterization of promising novel enzymes.
In the past, enzyme-based industrial processes were designed around the limitations of natural enzymes discovered in nature; today, enzymes can be engineered to fit the process specifications through enzyme engineering [1]. Enzyme engineering can be pursued by either rational site-directed mutagenesis based on currently available sequence and 3D structures deposited in the RCSB Protein Data Bank, or by directed evolution. Directed evolution is a key technology at the base of the development of new industrial biocatalysts [49,50,51,52] by random amino acid residues changes in the enzyme, largely carried out using PCR-based methods, followed by selection or screening of resulting libraries for variants with improved enzymatic properties. Changes in enzyme properties usually require simultaneous multiple amino-acid substitutions, creating exponentially more variants for testing. However, modern high-throughput screening methods, such as fluorescence-activated cell sorting [53,54,55] allow the screening of tens of millions of variants in a short time. Moreover, the best approach to create multiple mutations is to limit the choices by statistical or bioinformatic methods, such as the statistical correlation approach based on the ProSAR (protein structure-activity relationship) algorithm [56], to identify if a particular substitution is beneficial or not.
Furthermore, the development of synthetic biology methodology, through which large DNA sequences can be synthesized de novo, is increasing the ability for designing and engineering enzymes with new functions. Moreover, the possibility to synthesize whole-gene synthesis can also be used to build high-quality DNA libraries [1].
3. Cold-Adapted Enzymes and Their Biotechnological Applications
The bioprospecting of microorganisms living in extreme environmental conditions has already led to the discovery of several enzymes endowed with catalytic properties potentially useful for biotechnological applications [57], including some isolated from marine polar regions [23]. Moreover, the systematic investigation of the relationship between the structure and function of these enzymes—in comparison with their mesophilic homologs—has allowed recognising specific structural features associated with cold adaptation, which can be applied to the design of non-natural enzymes endowed with high activity at low/room temperatures. New catalytic activities are expected to arise from the Enzyme Function Initiative (EFI), which was recently established to address the challenge of assigning reliable functions to enzymes discovered in bacterial genome projects [58]. The main aim of the initiative is the computation-based prediction of substrate specificity for functional assignment of unknown enzymes.
3.1. Applications of Cold-Adapted Enzymes
The ultimate value of biotechnology is the production and delivery of molecules or processes of interest for many applications; the emerging requirement for biotechnological products is to be energy saving, thus reducing environmental impacts. Therefore, enzymes from cold-adapted organisms are important because their properties meet the ongoing efforts and needs to decrease energy consumption.
The ability of enzymes to catalyze a diverse set of reactions with exquisite specificity makes these proteins essential for biochemical studies, as well as promising catalysts for reactions useful to humankind. Due to their structural and functional complexity, the use of natural enzymes in biotechnology has been limited over the past decades. In fact, natural enzymes generally require modification for industrial use, given the improvements in catalytic efficiency, stereo-selectivity, and stability that can be reached through combinatorial engineering strategies. Recent advances in science and technology have led to activity improvements for native and non-native substrates, synthesis of new chemistries and functional characterization of promising novel enzymes.
Protein engineering technologies make it possible to develop enzymes that are highly active on non-natural targets, in the presence of organic solvents, and even enzymes for chemical transformations not found in nature. Application of these advanced engineering technologies to the creation of biocatalysts typically starts with an integrated approach with iterative cycles of DNA mutation and selection to create—by directed evolution—enzymes with novel functions for biotechnological approaches [59]. Due to our enhanced understanding of how these enzymes work in extreme environments, we can look forward to increasingly use computational tools, such as Artificial Intelligence, and to design enzymes with new structures and functions [60].
Cold-adapted enzymes have been suggested as biotechnological tools for several reasons:
- (1)
- They are cost-effective, e.g., lower amounts are required, due to higher catalytic efficiency at low temperature;
- (2)
- They can catalyze reactions at temperatures where competitive, undesirable chemical reactions are slowed down. This property is particularly relevant in the food industry, where deterioration and loss of thermolabile nutrients can occur at room temperature;
- (3)
- They catalyze the desired reactions at temperatures where bacterial contamination is reduced. There is a number of advantages in working at lower temperature (around 10–15 °C) than those currently used for large-scale industrial production;
- (4)
- Most cold-adapted enzymes can be inactivated by moderate heat due to their thermolability, avoiding chemical-based inactivation. A striking application of this property has been described for the design of live vaccines. Mesophilic pathogens were engineered for production of thermolabile homologs of essential enzymes, making them temperature-sensitive (TS). The engineered strains are inactivated at mammalian body temperatures, thus losing pathogenicity, but retaining their entire antigenic repertoire. Duplantis and colleagues [61] were able to entirely shift the lifestyle of Francisella (F. novicida), responsible for tularaemia disease in mice, by substituting its genes encoding essential enzymes with those identified in an Arctic bacterium. The authors applied the same approach to Salmonella enterica and the Gram-positive Mycobacterium [62]. The TS S. enterica strains were shown to be safe in research or diagnostic laboratories but were still capable of stimulating a protective immune response [63]. The TS strain of M. tuberculosis could be a safe research or diagnostic strain that is incapable of causing serious disease in humans while being identical to wild-type M. tuberculosis except for the TS phenotype [62].
The isolation of cold-adapted enzymes from natural sources is mostly unfeasible and their recombinant expression through standard protocols is hindered by the relatively high growth temperature of the commonly used expression hosts, including E. coli. However, several recombinant expression systems have been devized for the production of cold-adapted enzymes [64], mostly aimed at the stabilization and solubilization of heat-sensitive proteins. Development of low-temperature expression system has been attempted in the psychrophilic Arthrobacter sp. isolated from a Greenland glacier [65]. An alternative strategy is to use the engineered Arctic Express Cells which co-express cold-adapted homologues of the E. coli GroELS chaperonines, Cpn60 and Cpn10, from O. antarctica [66]. This system allows protein processing at very low temperatures (4–12 °C), potentially increasing the yield of active, soluble recombinant protein.
3.2. Structural Features of Cold-Adapted Enzymes
A basic requirement of evolutionary cold adaptation is that the enzyme should afford metabolic rates comparable to those of mesophilic organisms at its working temperature. Indeed, the temperature-dependence of the catalyzed reaction constants is described by the Arrhenius Equation (1)
where k is the reaction rate, E is the activation energy of the reaction, R is the gas constant, T is temperature and A is a collision frequency factor. It shows that a decrease of 10 °C brings about a decrease of 2–3 fold in the reaction rate. Therefore, a mesophilic enzyme with an optimal working temperature around 37 °C is expected to work 16–80 times slower when brought to 0 °C [67]. Increases in enzyme expression in psychrophilic microorganisms to compensate for the slower reactivity have been observed only in a small number of cases. Rather, the most common evolutionary adaptation consists in the evolution of enzyme homologs endowed with increased catalytic efficiency, the kcat/KM ratio, either by an increase of kcat—particularly for enzymes working at saturating concentrations of substrate—or by a decrease in KM—particularly for enzymes working at sub-saturating concentrations of substrate. kcat is almost invariably increased in cold-adapted enzymes, up to 10-fold. The trade-off between kcat and KM in cold-adapted enzymes has been comprehensively reviewed [68].
k = Ae−E/(RT)
From a structural point of view, it is widely accepted that these kinetic parameters are associated with the increased flexibility of these enzymes [69,70]. This dynamics adaptation allows for the conformational rearrangements required for catalysis at low temperatures, making it possible for cold-adapted enzymes to maintain the same conformational space accessible to their mesophilic and thermophilic homologs at higher temperatures. Indeed, for the latter enzymes, the kinetic energy associated with low temperatures would be insufficient to overcome the kinetic barriers required for the conformational rearrangements associated with catalysis. The turnover number, or kcat, is a good index of enzyme flexibility, as it reflects the rate of transition between all conformational states involved in the catalytic cycle.
Multiple structural features are usually associated with increased flexibility, involving both the active site and the peripheral regions. Commonly observed structural features are: Decreased core hydrophobicity, fewer prolyl residues in loops, increased hydrophobicity of the surface, more glycyl residues, lower arginyl/lysyl ratio, weaker interactions between subunits and domains, longer loops, decreased secondary-structure content, more prolyl residues in α-helices, fewer and weaker metal-binding sites, fewer disulfide bridges, fewer electrostatic interactions, reduced oligomerization and increase in conformational entropy of the unfolded state [71,72]. Many of these structural adaptations are summarized in Table 1. The crystal structures of 11 proteins isolated from the Antarctic marine oil-degrading bacterium Oleispira antarctica (α/β hydrolase, phosphodiesterase, transaldolase, isochorismatase, amidohydrolase, fumarylacetoacetate isomerase/hydrolase, 2-keto-3-deoxy-6-phosphogluconate aldolase, phoshonoacetaldehyde hydrolase, inorganic pyrophosphatase, and protein with unknown function) revealed that the most dominant structural feature is an increase in surface hydrophobicity and negative charge, with higher Glu+Asp/Arg+Lys ratio compared to their mesophilic counterparts [73].

Table 1.
Structural adaptations in cold-adapted enzymes and their effects on protein structure.
In other instances, however, a small number of specific residues was recognized as solely responsible for the cold adaptation of enzymes. In the β-galactosidase from the polyextremophilic Antarctic archaeon Halorubrum lacusprofundi, the mutation of only six amino-acid residues in comparison to its mesophilic homologs resulted in altered temperature activity profiles, suggesting that a small number of mutations can indeed account for cold adaptation [71]. In other proteins, a limited number of regions were associated to cold adaptation: the characterization of chimeric derivatives of monomeric isocitrate dehydrogenases of Colwellia maris and Pseudomonas psychrophila—cold adapted and mesophilic, respectively—led to the identification of two regions responsible for their different thermal properties [78]. The relative contribution of local and global protein flexibility in determining cold adaptation was assessed by performing molecular dynamics simulations on cytosolic malate dehydrogenases orthologous from marine mollusks adapted to different temperatures [79]. A convergent evolution trend was observed, with a significant negative correlation between the adaptation temperature of the organism and overall protein flexibility, with regions involved in ligand binding and catalysis showing the largest fluctuation differences [79].
Generally, the higher flexibility at low temperatures brings about low stability at higher temperatures. However, thermolability, although common [70], is not universal in cold-adapted enzymes. Three superoxide dismutases from the Antarctic psychrophilic ciliate Euplotes focardii were recently showed to exhibit a melting temperature of around 50–70 °C, suggesting a combination of cold adaptation and relative tolerance to high temperatures [80]. GroEL and thioredoxin from the Antarctic bacterium Pseudoalteromonas haloplanktis TAC125 were shown to be only marginally less stable than their Escherichia coli homologs [81].
It should also be highlighted that cold adaptation does not only consist in altered kinetic parameters of individual enzymes. For instance, sequencing and functional analysis of the genome the Antarctic marine oil-degrading bacterium O. antarctica revealed an array of alkane monooxygenases, osmoprotectants, siderophores and micronutrient-scavenging pathways [73] associated to cold adaptation.
3.3. Examples of Biotechnological Applications of Polar Enzymes
According to the type of catalyzed reaction, enzymes are classified into six main classes, i.e., oxidoreductases, transferases, hydrolases, lyases, isomerases and ligases, as reported in Table 2. The proposed biotechnological applications of cold-adapted enzymes belonging to several of these classes have been extensively reviewed [23,25,82,83,84,85].

Table 2.
Polar-active enzymes isolated from Antarctic and Arctic marine polar environments.
Currently, approximately 65% of more than 3000 enzymes with an industrial application are hydrolases, used in the detergent, textile, paper, and food industries [167,191]. Hydrolases, classified as Class 3 (EC 3) by the Nomenclature Committee of the International Union of Biochemistry and Molecular Biology (NC-IUBMB), catalyze the hydrolysis of chemical bonds. Herein, we focus on some recent examples of cold-adapted enzymes, particularly hydrolases, isolated from marine polar microorganisms. More examples of microbial enzymes isolated in Arctic or Antarctic marine polar regions are reported in Table 2. To our knowledge, this is the first table including only cold-active enzymes isolated from marine polar microorganisms. Indeed, several recent reviews do not distinguish between polar and non-polar enzymes or between marine and non-marine origin. The latter distinction is crucial to guide bioprospecting in search for novel activities.
3.3.1. Glycoside Hydrolases
Glycoside hydrolases (GH; EC 3.2.1.), or glycosidases, hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety (Scheme 1a), overall processing a wide variety of complex substrates [192]. They are extremely common in nature, with more than 100 GH families currently recognized and grouped in the Carbohydrate-Active Enzymes database (CAZy), which provides a continuously updated list of families [193]. Their function ranges from the degradation of complex carbohydrates for their metabolic use (cellulases, amylases), to the defense against pathogens (lysozyme, chitosanase), to the synthesis of the oligosaccharide groups of glycoproteins. Their biotechnological applications are manifold, especially in the food processing industry, where enzyme activity at low temperatures is desirable to slow down spoilage and loss of nutrients. For this reason, cold-adapted hydrolases are usually in need [194].
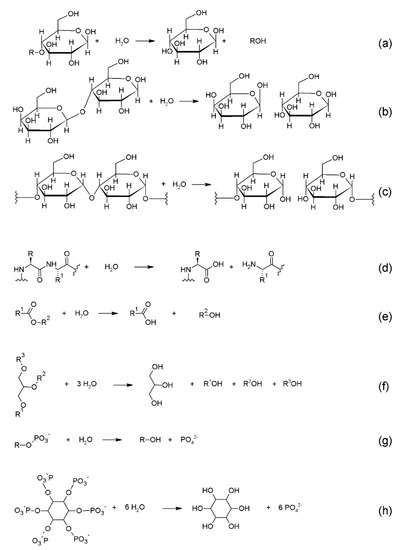
Scheme 1.
Principal enzymatic activities of cold-adapted enzymes investigated for biotechnological applications. (a) Generic reaction catalyzed by glycoside hydrolases, with a glucose derivative as substrate. (b) Reaction catalyzed by lactases, with conversion of lactose to galactose and glucose. (c) Reaction catalyzed by amylases on glucose polymers, with glucose or maltooligosaccharides as final products. (d) Generic reaction catalyzed by proteases, which catalyze the hydrolysis of peptide bonds of proteins or peptides. (e) Generic reaction catalyzed by esterases, which hydrolyse ester bonds. (f) Reaction catalyzed by lipases, with the hydrolysis of fatty acids from acyl glycerol. In this case, the full hydrolysis of a triglyceride to glycerol is represented. (g) Generic reaction catalyzed by phosphatases with the hydrolysis of phosphate monoesters. (h) Reaction catalyzed by phytases. In this case, the full hydrolysis to phosphate ions and inositol is represented.
Lactases or β-galactosidases (EC 3.2.1.23) (Scheme 1b) offer a clear example in this respect: with more than 70% of the world population suffering from lactose intolerance, the enzymatic removal of lactose from milk and dairy products using β-galactosidases is now common. The enzyme catalysing the hydrolysis of lactose into its constituent monosaccharides, glucose, and galactose, has drawn the interest of many research groups and enterprizes due to nutritional (lactose intolerance) and technological (crystallization), challenges associated with lactose [195]. The isolation of several cold-adapted β-galactosidases has been reported. Among them, some are from Antarctic organisms, e.g., H. lacusprofundi, isolated from the hypersaline Deep Lake [71,196], Arthrobacter species from soil samples [197,198], and marine Pseudoalteromonas species [86,87,88,199]. Other enzymes were isolated from marine organisms from the Arctic region, including those from Enterobacter ludwigii [91] and Alkalilactibacillus ikkense [92]. A cold-active β-galactosidase from the Antarctic marine bacterium P. haloplanktis TAE 79 was applied to lactose hydrolysis at low temperatures during milk storage [88] and was proposed for the production of tagatose, a natural monosaccharide with low caloric value and glycemic index [200]. A cold-active lactase isolated from the Antarctic marine P. haloplanktis LMG P-19143 was patented [89] and is also produced in large quantities by Nutrilab NV (Bekkevoort, Belgium). Yeasts are important sources for β-galactosidase production. Guehomyces pullulans 17-1, isolated from the Antarctic sea sediment, synthesize both extracellular and cell-bound β-galactosidases [90]. A 1,3-α-3,6-anhydro-L-galactosidase—an enzyme involved in the final steps of the agarolytic pathway and used in the industrial processing of agar—was identified in the agarolytic marine bacterium Gayadomonas joobiniege G7 and was shown to be active between 7.0 and 15 C [201].
Amylases (Scheme 1c) are also interesting enzymes for industrial processes, in detergent formulations and in food industry for beer and wine fermentation and for the preparation of bread and fruit juice. They hydrolyse starch to maltose, maltotriose, glucose monomers and limit dextrins. They can be classified in exoamylases (β-amylase EC 3.2.1.2; glucoamylase, EC 3.2.1.3; α-glucosidase, EC 3.2.1.20) endoamylases (α-amylase, EC 3.2.1.1) and debranching (pullulanases, EC 3.2.1.41; isoamylases, EC 3.2.1.68; dextrinases, EC 3.2.1.142) according to the specificity of the hydrolysis reaction they catalyze [194]. In the detergent industry, cold-active amylases not isolated from polar regions are already available on the market [167], i.e., Stainzyme® (active at temperatures between 30 and 70 °C) and Stainzyme® Plus (active below 20 °C) released by Novozymes and Preferenz™ S100 (active at 16 °C) released by DuPont Industrial Biosciences. In the textile industry, Genencor-DuPont developed Optisize® COOL, using a cold-active amylase for the desizing of woven fabrics at low temperatures. Other cold-active, detergent-stable α-amylases, not isolated from polar regions, were identified from Bacillus cereus GA6 [202] and from the marine bacterium Zunongwangia profunda [203]. The latter was isolated from surface seawater in the costal area of Fujian, China, but shows a cold-adapted and salt-tolerant α-amylase activity. An α-amylase produced by the psychrophilic yeast Glaciozyma antarctica PI12 was recently characterized [94].
Cold-adapted enzymes from marine polar regions could successfully be employed in these applications, and some of them are already under investigation or in use. A cold-adapted α-amylases from the Antarctic sea-ice bacterium Pseudoalteromonas sp. M175 exhibited resistance towards all the tested commercial detergents and was shown to improve their stain removal efficiency [93]. Heat-labile α-amylases displaying high activity at low temperature isolated from bacteria of Antarctic seawater were structurally studied [96,97].
Pullulanases (EC 3.2.1.41), debranching enzymes that hydrolyse α-1,6-and α-1,4-linkages in pullulan, starch, amylopectin and various related oligosaccharides are interesting enzymes in starch processing. There are different pullulanase groups according to their substrate specificities and reaction products. Pul13A is a type-I pullulanase isolated from Arctic seawater strain Shewanella arctica in Spitsbergen, Norway. It is able to hydrolase α-1,6-glycosidic bonds in pullulan to produce maltotriose at low temperature, displaying low thermostability at elevated temperatures. It represents a potential candidate for industrial applications such as starch degradation for ethanol-based biofuel production [143].
Although poorly studied, cold-active xylanases (EC 3.2.1.8) are also interesting in the food industry for bread making, as they convert the insoluble hemicellulose of dough into soluble sugars, thus yielding soft and elastic bread. Three cold-adapted xylanases from psychrophilic bacteria were shown to improve dough properties and bread volume with respect to mesophilic orthologues [204]. The xylanase pXyl from P. haloplanktis, isolated from soil samples [205], was very efficient in improving the dough properties and bread volume due to its high activity at low temperature [205]. The psychrophilic enzyme is now sold by Puratos (Grand-Bigard, Belgium). Antarctic marine fungi from marine sponges, which are the dominant macroinvertebrates in many benthic communities, are considered new and promising sources of cold-active xylanases. Cladosporium sp. isolated from marine sponges collected in King George Island, Antarctica, showed high xylanase activity at low temperature and very low thermal stability [99]. Also Flavobacterium frigidarium sp. nov., an aerobic psychrophilic and halotolerant bacterium isolated from marine sediment of shallow waters surrounding Adelaide Island in Antarctica, exhibited xylanolytic activity [100].
Cellulases, used in detergents for color and brightness care, have been classified according to their activity into endo- (EC 3.2.1.4) and exo-cellulases with exo-β-1,4-glucan cellobiohydrolase (EC 3.2.1.91) and β-glucosidase (EC 3.2.1.21). P. haloplanktis is able to convert the cellulose into an immediate nutritive compound for plants by hydrolysis. P. haloplanktis secretes a multi-modular endocellulase composed N-terminal catalytic module, a linker region and the cellulose-binding module. Structural adaptations to cellulose hydrolysis at low temperatures have been identified in the catalytic module and unusually long linker region by using both X-ray diffraction and small angle X-ray scattering methods [145].
Leucosporidium antarcticum strain 171, widespread in the cold marine waters below the Antarctic ice, was isolated from a seawater sample collected at a depth of 100 m in Admiralty Bay, King George Island. The yeast was found to produce cold-adapted invertases (β-D-fructofuranoside fructohydrolases, EC 3.2.1.26) and glucosidases (α-D-glucoside glucohydrolases, EC 3.2.1.20). Their synthesis may facilitate assimilation of β-fructofuranosides and α-glucopyranosides in cold environments where nutrient availability is fluctuating [144].
Chitin, one of the most abundant organic compounds in nature, is a structural polysaccharide composed of N-acetylglucosamine (GlcNAc) residues. Chitobiases (EC 3.2.1.29) found in bacteria, fungi, and eukaryotes, hydrolyse chitobiose to GlcNAc, which is further converted to glucosamine by a deacetylase. A cell-bound chitobiase was isolated from the marine psychrophile Arthrobacter sp. TAD20 collected along the Antarctic ice. The cold-adapted chitobiase overexpressed in E. coli displayed four functionally independent domains: (i) The catalytic domain, (ii) the galactose-binding domain, and (iii) the immunoglobulin-like domain followed by (iv) the cell-wall anchorage signal. The enzyme exhibited features that are typical of cold-adapted enzymes, with unusually low-Km and high-kcat values, with improved flexibility around the active site for efficient activity at low temperatures [146].
3.3.2. Proteases
Proteases (EC 3.4) catalyze the hydrolysis of peptide bonds (Scheme 1d). This catalytic activity has evolved multiple times, yielding enzymes with different catalytic mechanisms [206]. These enzymes are classified as exo-peptidases (EC 3.4.11-19), when they cleave the terminal amino acid, and as endo-peptidases (EC 3.4.21-99), when the peptide bonds they hydrolysed is internal. Proteases have several industrial applications, particularly in the food industry and as components of laundry detergents [207,208]. Both thermophilic [209] and cold-adapted homologs [210] are of biotechnological interest, depending on the intended working temperature of the process. To obtain cold-active proteases, two approaches have been pursed: mutagenesis of mesophilic enzymes or identification and expression of naturally occurring psychrophilic proteases. The former approach was applied to subtilisins from Bacillus species, mutated to increase their activity at low temperature. Particularly, the subtilisin savinase from Bacillus was mutated to produce a thermostable variant with increased activity at low temperature [211]. The latter approach has also been investigated.
Commercially exploited cold-adapted proteases include a protease from Bacillus amyloliquefaciens—a non-polar soil bacterium—commercialized as Purafect Prime LTM, which is active at 20 °C, and the protease ProperaseTM from B. alcalophilus, which combines activity at high pH and low temperature and ExcellaseTM. They are commercialized by Genencor (Genencor International Inc., Palo Alto, CA, USA). A cold-adapted extracellular aspartic protease was isolated from the yeast Sporobolomyces roseus isolated from an underground water sample drawn from the disused silver and lead mine “Luiza” (Zabrze, Poland) and was proposed as biocatalyst in the food industry, particularly in cheese and soy-sauce production, meat tenderization, and as bread additive [212]. It was also proposed in the production of antioxidant peptides from dairy and animal proteins, as hydrolysis of beef casein by this enzyme yielded peptides endowed with antioxidant activity [212].
Bioprospecting of cold-adapted bacteria from the polar regions could further increase the variety of cold-active proteases for industrial use. Several recent examples of such enzymes have been reported. Particularly, Antarctic fungi capable of producing proteases have been recently reviewed [85]. Two cold-adapted subtilisins from Antarctic marine Bacillus TA39 [102] and TA41 [103] showed high activity at low temperatures but also limited stability. A mutant with unaltered activity at low temperatures but higher stability was obtained through directed evolution [101]. A subtilisin-like cold-adapted serine peptidase was isolated from the Antarctic bacterium Lysobacter sp. A03, and is both active at low temperatures and resistant to higher temperatures [113].
A protease from the Antarctic bacterium Janthinobacterium lividum obtained from the Polar BioCenter (Korea Polar Research Institute) [213], recombinantly expressed and purified, was reported. Among the fungal proteases identified from polar marine samples, the extracellular subtilase lap2, from the Antarctic yeast Leucosporidium antarcticum 171, exhibited a low optimal temperature (25 °C) poor thermal stability, and high catalytic efficiency in the temperature range 0–25 °C [109].
The extracellular protease released by Rhodotorula mucilaginosa L7 strain was isolated from an Antarctic marine alga and was shown to be stable in the presence of high concentrations of NaCl [106]. A new cold-adapted protease with a potential biotechnological application, largely represented amongst Antarctic bacterial genus (Pseudoalteromonas sp., Marinobacter sp., Psychrobacter sp., Polaribacter sp.), exhibits a higher catalytic efficiency at lower temperatures compared to its mesophilic counterpart [107]. The extracellular cold-active protease from the marine psychrophilic bacterium Pseudoalteromonas sp. NJ276, for its catalytic activity and broad substrate specificities, has a potential application in low-temperature food processing, in particular in the preservation of milk at low temperature [108]. A secreted a cold-active serine protease identified in Colwellia sp. NJ341, isolated from Antarctic sea-ice, showing a 30% of activity at 0 °C and a better thermostability than other cold-active proteases, represents a good candidate for industrial applications, particularly in processes where there may be a risk of microbial contamination or a temperature instability of reactants or products [104].
An aminopeptidase produced by the marine psychrophile Colwellia psychrerythraea strain 34H was structurally investigated, displaying structural features (fewer proline residues, fewer ion pairs, and lower hydrophobic residue content) related to the cold adaptation and able to increase the flexibility for activity in the cold [111].
Besides their industrial applications, proteases have also long been used for therapeutic applications, particularly in the treatment of cardiovascular diseases, sepsis, digestive disorders, inflammation, cystic fibrosis, retinal disorders, and psoriasis [214]. The proteases so far approved by the US FDA derive from mesophilic organisms. However, the cold-adapted ones are endowed with a higher catalytic efficiency [215] and might, therefore, be of great interest. Indeed, they evolved to be efficient at low temperatures and, if thermostable enough, they would further increase their reaction rates at 37 °C, the intended temperatures for therapeutic applications. To overcome their thermal instability, mutational studies have been conducted [83].
As for general applications in molecular biology, proteinase, an endopeptidase from an Arctic marine microbial source, was developed by ArcticZymes [118] for the digestion of chromatin, thus releasing naked DNA. As it is thermolable, it can be inactivated at temperatures compatible with RNA integrity and DNA as double strands.
3.3.3. Lipases and Esterases
Esterases (EC 3.1.1) hydrolyse ester bonds (Scheme 1e). Lipases in particular (EC 3.1.1.3) are glycerol-ester hydrolases that catalyze the hydrolysis of triglycerides to fatty acids and glycerol (Scheme 1f). They are widely used in organic synthesis and as laundry detergents [216]. Although thermophilic homologs might appear more promising for industrial applications for their higher stability under harsh conditions, cold-adapted homologs are also actively investigated for their high activity at low temperatures [217]. Moreover, lipases are promising tools for the preparation of chiral molecules in the pharmaceutical industry, where low working temperatures are desirable to reduce the rate of non-catalytic side reactions. Particularly, the predictable enantiopreference of lipases allows the determination of the absolute configuration of secondary alcohols using the lipase-catalyzed kinetic resolution and the use of cold-active lipases in organic solvents is excellent for the preparation of single-isomer chiral drugs, and the enantioselective or regioselective preparation of alcohol and amine intermediates in the synthesis of pharmaceuticals [218].
There are already examples of lipases identified in organisms isolated from the polar regions which are currently on the market. Lipozyme® CALB is a non-specific lipase from Candida antarctica which was successfully used in the resolution of racemic alcohols, amines, and acids, and in the preparation of optically active compounds from meso substrates, as well as for the regio-selective catalysis in the selective acylation of different carbohydrates [219]. Lipases from C. antarctica catalyze hydrolysis on several substrates and are thermostable, a surprising property considering their origin. CALB is currently commercialized by Novozyme (Bagsværd, Denmark). The lipase from C. antarctica is also available in an immobilized form and commercialized as Novozym 435 (Novozyme, Bagsværd, Denmark).
More cold-adapted lipases were recently discovered in Arctic and Antarctic microorganisms. A lipase from the psychrotolerant yeast Rhodotorula sp. Y-23 isolated in the Nella Lake, East Antarctica, exhibited the highest Vmax at 15 °C and high compatibility with commercial detergents, which brought about an increase in activity, making it a potential candidate in detergent formulation active at low temperatures [220]. The Gram-positive psychrotrophic Arctic bacterium Arthrobacter gangotriensis, obtained from an soil sample, was recently shown to exhibit lipolytic activity associated with a lipase that can work in a wide pH range and that exhibits good stability [221]. The bacterium Bacillus pumilus ArcL5 with lipolytic activity was isolated from the Chukchi Sea within the Arctic Ocean; the lipase BpL5, was recombinantly expressed in E. coli and was shown to retain 85% of its activity at 5 °C. Two mutants active against tricaprylin were also characterized [119].
Other lipases were identified in Antarctic soils [222]. The lipase LipG7 from the Antarctic filamentous fungus Geomyces sp. P7 was unusually thermostable and was proposed as an enantioselective biocatalyst [223].
Other cold-active lipases were identified in the Antarctic marine P. haloplanktis TAC125 [120], in the Arctic marine bacterium C. psychrerythraea 34H [121], Psychrobacter sp. of Antarctic marine origin [123,124,126] and Antarctic deep-sea water strain Moritella sp.2-5-10-1 [127,128].
Esterases (EC 3.1.1.1), hydrolases that catalyze the cleavage of simple esters such as triglycerides with short chains of fatty acids (less than eight carbon atoms) (Scheme 1e) [194], are interesting enzymes in organic synthesis and cheese ripening processes [137]. Cold-adapted esterases were identified in Arctic marine bacteria Pseudoalteromonas arctica, [131] and Thalassospira sp. [132] and the Antarctic marine bacteria O. antarctica [133] and P. haloplanktis TAC125 [134,135]. Moreover, a cold-adapted esterase was identified from Pseudoalteromonas sp. strain 643A, isolated from the alimentary tract of Antarctic krill Euphasia superba Dana. It displayed 20–50% of maximum activity at 0–20 °C, and the optimal temperature was close to 35 °C. The optimal pH for enzyme activity was around 8.0; however, it was stable between pH 9 and 11.5 [136].
3.3.4. Phosphatases
Alkaline phosphatases (EC 3.1.3.1), which catalyze the hydrolysis of phosphate monoesters (Scheme 1g), have an important application in molecular biology for the dephosphorylation of DNA linearized at the 5′ end to avoid its re-circularization during cloning processes. A recombinant alkaline phosphatase isolated from the Antarctic bacterial strain TAB5 [147,148] was developed by New England Biolabs [149]. The crystal structure of a cold-active alkaline phosphatase from a psychrophile, Shewanella sp. isolated from the intestine of an Antarctic shellfish, revealed local flexibility, responsible for the high catalytic efficiency at low temperatures [150].
Phytic acid is the phosphate ester of inositol. The phosphate groups in this form are not available for absorption by animals, with the partial exception of ruminants, where the hydrolysis is carried out by the rumen microbiota. Phytases (EC 3.1.3.) have been used for decades to enrich animal food of absorbable phosphate groups, both for non-ruminant livestock and fish [224] (Scheme 1h). Thermophilic phytases have been suggested for industrial use [225], but cold-adapted homologs were also proposed, especially in aquaculture, where low temperatures might be limiting for the activity of the mesophilic orthologues. In this view, the purification and characterization of novel cold-adapted phytases from the Antarctic marine R. mucilaginosa strain JMUY14 [130] and Pseudomonas sp. strain JPK1 isolated from soil were reported [226].
3.3.5. Other Hydrolases
A cold-adapted uracil-DNA glycosylase (EC 3.2.2.27 and EC 3.2.2.28) of Antarctic marine source was released by New England Biolabs as thermolabile uracil-DNA glycosylase [154]. This hydrolase, isolated from a psychrophilic marine bacterium, catalyzes the release of free uracil from uracil-containing single-stranded or double-stranded DNA; it is sensitive to heat and can be inactivated at temperatures above 50 °C.
Cryonase, a cold-active nuclease, isolated from an Antarctic marine psychrophile, Shewanella sp. strain Ac10 and patented by Awazu and colleagues [227] was developed by Takara-Clontech [151]. It is a recombinant endonuclease that can digest all types of DNA and RNA (single-stranded, double-stranded, linear or circularized) at low temperatures, frequently used during DNA digestion of samples in the presence of heat-labile proteins. A 3′–5′ exonuclease specific for single-stranded DNA and derived from an Arctic marine bacterium was released by ArcticZymes [152].
Antimicrobial enzymes endowed with bactericidal or bacteriostatic properties are an emerging strategy to combat pathogens, particularly by degrading their DNA, polysaccharides, and proteins, or by interfering with biofilm formation, or by catalysing reactions which result in the production of antimicrobial compounds [228]. For example, N-acyl homoserine lactones are signaling molecules involved in bacterial quorum sensing and their hydrolysis can reduce growth and biofilm formation of bacterial pathogens. A cold-adapted N-acylhomoserine lactonase (EC 3.1.1.81) was recently identified in Antarctic Planococcus sp. isolated from a soil sample collected on Lagoon Island (Antarctica) and was shown to attenuate the pathogenicity of Pectobacterium carotovorum, a plant pathogen that causes soft-rot disease [229]. As the enzyme was shown to be thermolabile at the human body temperature, it was suggested as a safe antimicrobial agent in the treatment of crops, as it would be inactivated upon ingestion.
3.3.6. Other Enzymes
Besides hydrolases, other enzymes have been isolated in marine microorganisms from the polar regions and, for some of them, a biotechnological application has been proposed.
Recently, few enzymes showing signatures of cold adaptation in their activity and structure have been isolated and fully characterized from the Arctic marine bacterium C. psychrerythraea strain 34H and also included a phenylalanine hydroxylase (PAH), an oxidoreductase, with high-catalytic efficiency at 10 °C, high thermostability and low affinity for substrate, probably due to enhanced flexibility of the active site [76]. PAH (EC 1.14.16.1) catalyzes the conversion of L-Phe to L-Tyr by para-hydroxylation of the aromatic side-chain and recently there has been an increased number of studies on the structure and function of PAH from bacteria and lower eukaryote organisms. Almost all characterized bacterial PAHs are monomeric and display a fold similar to the catalytic domain of mammalian PAHs.
Many oxidoreductases that catalyze redox reactions of industrial interest have been described in cold-adapted bacteria. The crystal structure of a Fe-superoxide dismutase, Fe-SOD (EC 1.15.1.1) isolated from the marine P. haloplanktis, revelead that cold-adapted enzyme displays high catalysis at low temperature increasing the flexibility of its active site without modifying the overall structure [161]. SODs have been used in the pharmaceutical and cosmetic industries, food, agricultural, and chemical industries [169]. Antioxidant defense is an important component of evolutionary adaptations in the cold to face increased levels of reactive oxygen species (ROS). The cold waters of the polar regions promote the formation of ROS and would be expected to lead to enhanced ROS damage of DNA and membrane lipid peroxidation in polar species [72]. P. haloplanktis TAC125 copes with increased O2 solubility by deleting entire metabolic pathways that generate ROS as side products [230]. In contrast, the Arctic bacterium C. psychrerythraea has developed an enhanced antioxidant capacity, owing to the presence of three copies of catalase genes, as well as two superoxide-dismutase genes, one of which codes for a nickel-containing superoxide-dismutase, never reported before in proteobacteria [231].
A recombinant form of glutathione reductase (EC 1.8.1.7) from the Antarctic C. psychrerythraea, was recently characterized [168]. Since the cold-adapted enzyme displays activity also at moderate temperatures when overexpressed in E. coli, it may have some potential for industrial applications to protect cells and tissues from oxidative stress. Malato dehydrogenase (MDH) (EC 1.1.1.37) was purified from the Antarctic marine bacterium Flavobacterium frigidimaris KUC-1 and, among cold-adapted MDHs, was shown to be the most thermolabile and cold-active [158]. Thioredoxin and thioredoxin reductase (EC 1.8.1.9) from P. haloplanktis TAC125 were obtained as recombinant proteins. Both proteins exhibit activity at 10 °C [170]. The recombinat cold-adapted peroxiredoxin (EC 1.11.1.15) was biochemically characterized from the Antarctic marine psychrophilic bacterium Psychrobacter sp. ANT206, which had an optimum growth temperature of 10–12 °C. These enzymes may be relevant for many applications in food and medicine, mostly for their ability to protect super-coiled DNA from oxidative stress [172].
A cold-adapted leucine dehydrogenase (EC 1.4.1.9), a NAD+-dependent oxidoreductase, with unique substrate specificity was cloned from the Antarctic marine psychrotrophic bacterium Pseudoalteromonas sp. ANT178 and was shown to retain 40% of its maximal activity at 0 °C. Being the key enzyme in the enzymatic conversion of L-leucine and other branched chain L-amino acids in the corresponding α-keto acid, it was suggested as biocatalyst in the pharmaceutical industry [157].
Aminotransferases (EC 2.6.1.), belonging to the class of transferases, have also been isolated from marine polar microorganisms, although no industrial application has been suggested so far. The aspartate aminotransferase, a ubiquitous transaminase enzyme that catalyzes the conversion of aspartate and α-ketoglutarate to oxaloacetate and glutamate, was isolated from the marine psychrophilic bacterium P. haloplanktis TAC125 and characterized from a structural and functional point of view [173]. It was shown to be thermolable, being inactivated at 50 °C. Its optimal working temperature is 10 °C lower than its E. coli homolog.
Photolyases (EC 4.1.99.3) were identified in the genera Pseudomonas, Hymenobacter and Sphingomonas, isolated in fresh waters in Antarctica (King George Island, Fildes Peninsula). Present in all living forms, except placental mammals and some marsupials, they are resistant to UV-radiations, being able to reverse DNA lesions (cyclobutane pyrimidine dimers and pyrimidine photoproducts [232] and, as UV radiations are known to be linked to skin cancer, they may have some potential in the cosmetic and pharmaceutical industries.
Cold-active γ-carbonic anhydrases (EC 4.2.1.1), able to catalyze the physiologic reaction of CO2 hydration to bicarbonate and protons, were isolated from the psychrophilic marine bacteria C. psychrerythraea [178] and P. haloplanktis [179] cloned and characterized.
A cold-active DNA ligase from the psychrophile P. haloplanktis TAE72, isolated from Antarctic seawater at the Dumont d’Urville Antarctic Station, displays activity at temperatures as low as 4 °C [177]. DNA ligases (EC 6.5.1.) are enzymes involved in DNA replication, DNA recombination and DNA repair. They are commonly used in molecular biology to catalyze the formation of a phosphodiester bond between adjacent 5′-phosphoryl and 3′-hydroxyl groups in double stranded DNA. Currently, DNA ligases on the market, such as the recombinant versions of bacteriophage-derived DNA ligases, T4 and T7 ligases, and E. coli DNA ligases, are enzymes active at temperatures above 15 °C, where residual nucleases may interfere with the ligation process [167]. The possibility to use psychrophilic DNA ligases active at low temperature may guarantee high specific activity at low temperatures, thus avoiding unwanted reactions, carrying out the reaction in shorter times with respect to mesophilic ligases.
The crystal structure of the sedoheptulose 7-phosphate isomerase from the marine psychrophilic organism C. psychrerythraea 34H was determined. The Arctic bacterium produces extracellular polysaccharide substances to cope with cold and the isomerase is essential for producing D-glycero-D-mannoheptose 7-phosphate, a key mediator in the lipopolysaccharide biosynthetic pathway [184]. Other polar enzymes belonging to the class of isomerase are triose phosphate isomerases (EC 5.3.1.1), enzymes involved in the glycolytic pathway, identified in the Antarctic marine bacteria Pseudomonas sp. π9 [185], isolated from sea-ice and Moraxella sp. TA137 [186], isolated from the intestine of fish caught in Terre d’Adelie in Antarctica.
4. Perspectives and Conclusions
Marine microorganisms, whose immense genetic and biochemical diversity is only beginning to be appreciated, may become a rich source of novel enzyme activities in the near future. The marine bioprospecting of polar regions has begun only relatively recently but has already yielded success stories. Taking into account the evidence that the total number of species, and thus most likely biochemical diversity in the oceans, is higher than on land, there is a good reason to believe that a larger number of marine natural products from low-temperature environments may reach different sectors of biotechnology in the near future. For those reasons, a large number of research and development programmes in oceans are in progress worldwide. Although the genomic, functional and physiological knowledge of individual organisms is necessary to understand how enzymes and products work, more efficient strategies are needed to overcome the limits of the cultivation of microbes, especially those living in extreme environments such as the polar ones. The specific properties of polar marine microorganisms that make them unique and biotechnologically interesting are also responsible for their resistance to handling and cultivation. The most significant challenges are (i) the current lack of ability to produce extreme compounds (including enzymes) on large scale; (ii) the need to generate stronger synergies among marine scientists and industries at earlier levels to compensate the chronic underfunding of basic research and before large-scale production; (iii) the need to develop or improve technology transfer pathways for data and to secure access to fair and equitable benefit sharing of marine genetic resources. Metagenomics, genome engineering and systems biology will be fundamental to efficiently produce larger quantities of known and novel bioresources.
In the race to find new products for biotechnology, although there are good reasons to be optimistic, diverse strategies need to be adequately supported and funded in the academic context. The future perspectives require active collaboration between academia and industry at the beginning to support the research thus reducing the time lapse from the discovery to the industrial applications.
Author Contributions
All authors gave an important contribution in the consideration of the available bibliographic information for the review and in the preparation of the final version of the manuscript. S.B. and C.V. developed the original idea and organized all the materials.
Funding
This study was financially supported by the Italian National Programme for Antarctic Research (PNRA) (2016/AZ1.06-Project PNRA16_00043 and 2016/AZ1.20-Project PNRA16_00128). It was carried out in the framework of the SCAR Programme “Antarctic Thresholds–Ecosystem Resilience and Adaptation” (AnT-ERA).
Acknowledgments
We thank the Reviewers for the time and effort they have spent in reviewing the manuscript.
Conflicts of Interest
The authors declare no conflict of interest
References
- Bornscheuer, U.T.; Huisman, G.W.; Kazlauskas, R.J.; Lutz, S.; Moore, J.C.; Robins, K. Engineering the third wave of biocatalysis. Nature 2012, 485, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Martinez, M.; Coscolin, C.; Santiago, G.; Chow, J.; Stogios, P.J.; Bargiela, R.; Gertler, C.; Navarro-Fernandez, J.; Bollinger, A.; Thies, S.; et al. Determinants and Prediction of Esterase Substrate Promiscuity Patterns. ACS Chem. Biol. 2018, 13, 225–234. [Google Scholar] [CrossRef] [PubMed]
- Raveendran, S.; Parameswaran, B.; Ummalyma, S.B.; Abraham, A.; Mathew, A.K.; Madhavan, A.; Rebello, S.; Pandey, A. Applications of Microbial Enzymes in Food Industry. Food Technol. Biotechnol. 2018, 56, 16–30. [Google Scholar] [CrossRef] [PubMed]
- BCC. Global Markets for Enzymes in Industrial Applications; BCC Publishing: Wellesley, MA, USA, 2018. [Google Scholar]
- SmithersGroup. The Future of Marine Biotechnology for Industrial Applications to 2025; SmithersGroup: Akron, OH, USA, 2015. [Google Scholar]
- Di Donato, P.; Buono, A.; Poli, A.; Finore, I.; Abbamondi, G.R.; Nicolaus, B.; Lama, L. Exploring Marine Environments for the Identification of Extremophiles and Their Enzymes for Sustainable and Green Bioprocesses. Sustainability 2019, 11, 149. [Google Scholar] [CrossRef]
- Gerwick, W.H.; Moore, B.S. Lessons from the past and charting the future of marine natural products drug discovery and chemical biology. Chem. Biol. 2012, 19, 85–98. [Google Scholar] [CrossRef] [PubMed]
- Verde, C.; Giordano, D.; Bellas, C.M.; di Prisco, G.; Anesio, A.M. Chapter Four—Polar Marine Microorganisms and Climate Change. Adv. Microb. Physiol. 2016, 69, 187–215. [Google Scholar]
- Jakobsson, M. Hypsometry and volume of the Arctic Ocean and its constituent seas. Geochem. Geophys. Geosyst. 2002, 3, 1–18. [Google Scholar] [CrossRef]
- Scher, H.D.; Whittaker, J.M.; Williams, S.E.; Latimer, J.C.; Kordesch, W.E.; Delaney, M.L. Onset of Antarctic Circumpolar Current 30 million years ago as Tasmanian Gateway aligned with westerlies. Nature 2015, 523, 580–583. [Google Scholar] [CrossRef]
- Maldonado, A.; Bohoyo, F.; Galindo-Zaldívar, J.; Hernández-Molina, F.J.; Lobo, F.J.; Lodolo, E.; Martos, Y.M.; Pérez, L.F.; Schreider, A.A.; Somoza, L. A model of oceanic development by ridge jumping: Opening of the Scotia Sea. Glob. Planet. Chang. 2014, 123, 152–173. [Google Scholar] [CrossRef]
- Russell, N.J. Molecular adaptations in psychrophilic bacteria: Potential for biotechnological applications. Adv. Biochem. Eng. Biotechnol. 1998, 61, 1–21. [Google Scholar]
- Cavicchioli, R. On the concept of a psychrophile. ISME J. 2015, 10, 793. [Google Scholar] [CrossRef] [PubMed]
- Lebar, M.D.; Heimbegner, J.L.; Baker, B.J. Cold-water marine natural products. Nat. Prod. Rep. 2007, 24, 774–797. [Google Scholar] [CrossRef]
- Wilson, Z.E.; Brimble, M.A. Molecules derived from the extremes of life. Nat. Prod. Rep. 2009, 26, 44–71. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.T.; Lu, X.L.; Liu, X.Y.; Gao, Y.; Hu, B.; Jiao, B.H.; Zheng, H. Bioactive natural products from the antarctic and arctic organisms. Mini Rev. Med. Chem. 2013, 13, 617–626. [Google Scholar] [CrossRef] [PubMed]
- Skropeta, D.; Wei, L. Recent advances in deep-sea natural products. Nat. Prod. Rep. 2014, 31, 999–1025. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Li, Y.L.; Zhao, F.C. Secondary Metabolites from Polar Organisms. Mar. Drugs 2017, 15, 28. [Google Scholar] [CrossRef]
- Nunez-Montero, K.; Barrientos, L. Advances in Antarctic Research for Antimicrobial Discovery: A Comprehensive Narrative Review of Bacteria from Antarctic Environments as Potential Sources of Novel Antibiotic Compounds Against Human Pathogens and Microorganisms of Industrial Importance. Antibiotics 2018, 7, 90. [Google Scholar] [CrossRef]
- Murray, A.E.; Grzymski, J.J. Diversity and genomics of Antarctic marine micro-organisms. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2007, 362, 2259–2271. [Google Scholar] [CrossRef]
- Margesin, R.; Miteva, V. Diversity and ecology of psychrophilic microorganisms. Res. Microbiol. 2011, 162, 346–361. [Google Scholar] [CrossRef]
- Casanueva, A.; Tuffin, M.; Cary, C.; Cowan, D.A. Molecular adaptations to psychrophily: The impact of ‘omic’ technologies. Trends Microbiol. 2010, 18, 374–381. [Google Scholar] [CrossRef]
- Cavicchioli, R.; Siddiqui, K.S.; Andrews, D.; Sowers, K.R. Low-temperature extremophiles and their applications. Curr. Opin. Biotechnol. 2002, 13, 253–261. [Google Scholar] [CrossRef]
- de Pascale, D.; De Santi, C.; Fu, J.; Landfald, B. The microbial diversity of Polar environments is a fertile ground for bioprospecting. Mar. Genom. 2012, 8, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Joseph, B.; Kumar, V.; Ramteke, P.W. Chapter 47—Psychrophilic Enzymes: Potential Biocatalysts for Food Processing. Enzym. Food Biotechnol. 2019, 817–825. [Google Scholar] [CrossRef]
- Dhamankar, H.; Prather, K.L. Microbial chemical factories: Recent advances in pathway engineering for synthesis of value added chemicals. Curr. Opin. Struct. Biol. 2011, 21, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Weber, W.; Fussenegger, M. Emerging biomedical applications of synthetic biology. Nat. Rev. Genet. 2011, 13, 21–35. [Google Scholar] [CrossRef]
- Cavicchioli, R. Microbial ecology of Antarctic aquatic systems. Nat. Rev. Microbiol. 2015, 13, 691–706. [Google Scholar] [CrossRef] [PubMed]
- Uchiyama, T.; Miyazaki, K. Functional metagenomics for enzyme discovery: Challenges to efficient screening. Curr. Opin. Biotechnol. 2009, 20, 616–622. [Google Scholar] [CrossRef]
- Ferrer, M.; Beloqui, A.; Timmis, K.N.; Golyshin, P.N. Metagenomics for mining new genetic resources of microbial communities. J. Mol. Microbiol. Biotechnol. 2009, 16, 109–123. [Google Scholar] [CrossRef]
- Pena-Garcia, C.; Martinez-Martinez, M.; Reyes-Duarte, D.; Ferrer, M. High Throughput Screening of Esterases, Lipases and Phospholipases in Mutant and Metagenomic Libraries: A Review. Comb. Chem. High Throughput Screen. 2016, 19, 605–615. [Google Scholar] [CrossRef]
- Ferrer, M.; Mendez-Garcia, C.; Bargiela, R.; Chow, J.; Alonso, S.; Garcia-Moyano, A.; Bjerga, G.E.K.; Steen, I.H.; Schwabe, T.; Blom, C.; et al. Decoding the ocean’s microbiological secrets for marine enzyme biodiscovery. FEMS Microbiol. Lett. 2019, 366. [Google Scholar] [CrossRef]
- Zallot, R.; Oberg, N.O.; Gerlt, J.A. ‘Democratized’ genomic enzymology web tools for functional assignment. Curr. Opin. Chem. Biol. 2018, 47, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Behrens, G.A.; Hummel, A.; Padhi, S.K.; Schätzle, S.; Bornscheuer, U.T. Discovery and Protein Engineering of Biocatalysts for Organic Synthesis. Adv. Synth. Catal. 2011, 353, 1615–4150. [Google Scholar] [CrossRef]
- Lauritano, C.; Ianora, A. Overview of Recent EU-Funded Projects. In Grand Challenges in Marine Biotechnology; Springer: Berlin, Germany, 2018; pp. 425–449. [Google Scholar]
- Available online: http://www.inmare-h2020.eu/ (accessed on 15 July 2019).
- Popovic, A.; Hai, T.; Tchigvintsev, A.; Hajighasemi, M.; Nocek, B.; Khusnutdinova, A.N.; Brown, G.; Glinos, J.; Flick, R.; Skarina, T.; et al. Activity screening of environmental metagenomic libraries reveals novel carboxylesterase families. Sci. Rep. 2017, 7, 44103. [Google Scholar] [CrossRef] [PubMed]
- Bastard, K.; Smith, A.A.; Vergne-Vaxelaire, C.; Perret, A.; Zaparucha, A.; De Melo-Minardi, R.; Mariage, A.; Boutard, M.; Debard, A.; Lechaplais, C.; et al. Revealing the hidden functional diversity of an enzyme family. Nat. Chem. Biol. 2014, 10, 42–49. [Google Scholar] [CrossRef] [PubMed]
- Chistoserdova, L. Is metagenomics resolving identification of functions in microbial communities? Microb. Biotechnol. 2014, 7, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, M.; Martinez-Martinez, M.; Bargiela, R.; Streit, W.R.; Golyshina, O.V.; Golyshin, P.N. Estimating the success of enzyme bioprospecting through metagenomics: Current status and future trends. Microb. Biotechnol. 2016, 9, 22–34. [Google Scholar] [CrossRef] [PubMed]
- McGettigan, P.A. Transcriptomics in the RNA-seq era. Curr. Opin. Chem. Biol. 2013, 17, 4–11. [Google Scholar] [CrossRef] [PubMed]
- Sturmberger, L.; Wallace, P.W.; Glieder, A.; Birner-Gruenberger, R. Synergism of proteomics and mRNA sequencing for enzyme discovery. J. Biotechnol. 2016, 235, 132–138. [Google Scholar] [CrossRef] [PubMed]
- Cravatt, B.F.; Wright, A.T.; Kozarich, J.W. Activity-based protein profiling: From enzyme chemistry to proteomic chemistry. Annu. Rev. Biochem. 2008, 77, 383–414. [Google Scholar] [CrossRef]
- Schittmayer, M.; Birner-Gruenberger, R. Lipolytic proteomics. Mass Spectrom. Rev. 2012, 31, 570–582. [Google Scholar] [CrossRef]
- Perfumo, A.; Banat, I.M.; Marchant, R. Going Green and Cold: Biosurfactants from Low-Temperature Environments to Biotechnology Applications. Trends Biotechnol. 2018, 36, 277–289. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.Q.; Jiang, X.R. Next generation industrial biotechnology based on extremophilic bacteria. Curr. Opin. Biotechnol. 2018, 50, 94–100. [Google Scholar] [CrossRef] [PubMed]
- Longwell, C.K.; Labanieh, L.; Cochran, J.R. High-throughput screening technologies for enzyme engineering. Curr. Opin. Biotechnol. 2017, 48, 196–202. [Google Scholar] [CrossRef] [PubMed]
- Lalonde, J. Highly engineered biocatalysts for efficient small molecule pharmaceutical synthesis. Curr. Opin. Biotechnol. 2016, 42, 152–158. [Google Scholar] [CrossRef] [PubMed]
- Porter, J.L.; Rusli, R.A.; Ollis, D.L. Directed Evolution of Enzymes for Industrial Biocatalysis. ChemBioChem 2016, 17, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Tyzack, J.D.; Furnham, N.; Sillitoe, I.; Orengo, C.M.; Thornton, J.M. Understanding enzyme function evolution from a computational perspective. Curr. Opin. Struct. Biol. 2017, 47, 131–139. [Google Scholar] [CrossRef] [PubMed]
- Acevedo-Rocha, C.G.; Gamble, C.G.; Lonsdale, R.; Li, A.; Nett, N.; Hoebenreich, S.; Lingnau, J.B.; Wirtz, C.; Fares, C.; Hinrichs, H.; et al. P450-Catalyzed Regio- and Diastereoselective Steroid Hydroxylation: Efficient Directed Evolution Enabled by Mutability Landscaping. ACS Catal. 2018, 8, 3395–3410. [Google Scholar] [CrossRef]
- Agostini, F.; Völler, J.-S.; Koksch, B.; Acevedo-Rocha, C.G.; Kubyshkin, V.; Budisa, N. Biocatalysis with Unnatural Amino Acids: Enzymology Meets Xenobiology. Angew. Chem. Int. Ed. 2017, 56, 9680–9703. [Google Scholar] [CrossRef]
- Bernath, K.; Hai, M.; Mastrobattista, E.; Griffiths, A.D.; Magdassi, S.; Tawfik, D.S. In vitro compartmentalization by double emulsions: Sorting and gene enrichment by fluorescence activated cell sorting. Anal. Biochem. 2004, 325, 151–157. [Google Scholar] [CrossRef]
- Becker, S.; Hobenreich, H.; Vogel, A.; Knorr, J.; Wilhelm, S.; Rosenau, F.; Jaeger, K.E.; Reetz, M.T.; Kolmar, H. Single-cell high-throughput screening to identify enantioselective hydrolytic enzymes. Angew. Chem. Int. Ed. Engl. 2008, 47, 5085–5088. [Google Scholar] [CrossRef]
- Fernandez-Alvaro, E.; Snajdrova, R.; Jochens, H.; Davids, T.; Bottcher, D.; Bornscheuer, U.T. A combination of in vivo selection and cell sorting for the identification of enantioselective biocatalysts. Angew. Chem. Int. Ed. Engl. 2011, 50, 8584–8587. [Google Scholar] [CrossRef]
- Fox, R.J.; Davis, S.C.; Mundorff, E.C.; Newman, L.M.; Gavrilovic, V.; Ma, S.K.; Chung, L.M.; Ching, C.; Tam, S.; Muley, S.; et al. Improving catalytic function by ProSAR-driven enzyme evolution. Nat. Biotechnol. 2007, 25, 338–344. [Google Scholar] [CrossRef] [PubMed]
- Coker, J.A. Extremophiles and biotechnology: Current uses and prospects. F1000Res 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Gerlt, J.A.; Allen, K.N.; Almo, S.C.; Armstrong, R.N.; Babbitt, P.C.; Cronan, J.E.; Dunaway-Mariano, D.; Imker, H.J.; Jacobson, M.P.; Minor, W.; et al. The Enzyme Function Initiative. Biochemistry 2011, 50, 9950–9962. [Google Scholar] [CrossRef] [PubMed]
- Devine, P.N.; Howard, R.M.; Kumar, R.; Thompson, M.P.; Truppo, M.D.; Turner, N.J. Extending the application of biocatalysis to meet the challenges of drug development. Nat. Rev. Chem. 2018, 2, 409–421. [Google Scholar] [CrossRef]
- Liszka, M.J.; Clark, M.E.; Schneider, E.; Clark, D.S. Nature versus nurture: Developing enzymes that function under extreme conditions. Annu. Rev. Chem. Biomol. Eng. 2012, 3, 77–102. [Google Scholar] [CrossRef] [PubMed]
- Duplantis, B.N.; Osusky, M.; Schmerk, C.L.; Ross, D.R.; Bosio, C.M.; Nano, F.E. Essential genes from Arctic bacteria used to construct stable, temperature-sensitive bacterial vaccines. Proc. Natl. Acad. Sci. USA 2010, 107, 13456–13460. [Google Scholar] [CrossRef]
- Pinto, C.T.; Nano, F.E. Stable, temperature-sensitive recombinant strain of Mycobacterium smegmatis generated through the substitution of a psychrophilic ligA gene. FEMS Microbiol. Lett. 2015, 362, fnv152. [Google Scholar] [CrossRef] [PubMed]
- Duplantis, B.N.; Puckett, S.M.; Rosey, E.L.; Ameiss, K.A.; Hartman, A.D.; Pearce, S.C.; Nano, F.E. Temperature-Sensitive Salmonella enterica Serovar Enteritidis PT13a Expressing Essential Proteins of Psychrophilic Bacteria. Appl. Environ. Microbiol. 2015, 81, 6757–6766. [Google Scholar] [CrossRef]
- Santiago, M.; Ramirez-Sarmiento, C.A.; Zamora, R.A.; Parra, L.P. Discovery, Molecular Mechanisms, and Industrial Applications of Cold-Active Enzymes. Front. Microbiol. 2016, 7, 1408. [Google Scholar] [CrossRef]
- Miteva, V.; Lantz, S.; Brenchley, J. Characterization of a cryptic plasmid from a Greenland ice core Arthrobacter isolate and construction of a shuttle vector that replicates in psychrophilic high G+C Gram-positive recipients. Extremophiles 2008, 12, 441–449. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, M.; Chernikova, T.N.; Yakimov, M.M.; Golyshin, P.N.; Timmis, K.N. Chaperonins govern growth of Escherichia coli at low temperatures. Nat. Biotechnol. 2003, 21, 1266–1267. [Google Scholar] [CrossRef] [PubMed]
- Georlette, D.; Blaise, V.; Collins, T.; D’Amico, S.; Gratia, E.; Hoyoux, A.; Marx, J.C.; Sonan, G.; Feller, G.; Gerday, C. Some like it cold: Biocatalysis at low temperatures. FEMS Microbiol. Rev. 2004, 28, 25–42. [Google Scholar] [CrossRef] [PubMed]
- Feller, G. Psychrophilic enzymes: From folding to function and biotechnology. Scientifica 2013, 2013, 512840. [Google Scholar] [CrossRef] [PubMed]
- Feller, G.; Gerday, C. Psychrophilic enzymes: Hot topics in cold adaptation. Nat. Rev. Microbiol. 2003, 1, 200–208. [Google Scholar] [CrossRef]
- Siddiqui, K.S.; Cavicchioli, R. Cold-adapted enzymes. Annu. Rev. Biochem. 2006, 75, 403–433. [Google Scholar] [CrossRef] [PubMed]
- Laye, V.J.; Karan, R.; Kim, J.M.; Pecher, W.T.; DasSarma, P.; DasSarma, S. Key amino acid residues conferring enhanced enzyme activity at cold temperatures in an Antarctic polyextremophilic beta-galactosidase. Proc. Natl. Acad. Sci. USA 2017, 114, 12530–12535. [Google Scholar] [CrossRef] [PubMed]
- Giordano, D.; Coppola, D.; Russo, R.; Tinajero-Trejo, M.; di Prisco, G.; Lauro, F.; Ascenzi, P.; Verde, C. The globins of cold-adapted Pseudoalteromonas haloplanktis TAC125: From the structure to the physiological functions. Adv. Microb. Physiol. 2013, 63, 329–389. [Google Scholar] [CrossRef]
- Kube, M.; Chernikova, T.N.; Al-Ramahi, Y.; Beloqui, A.; Lopez-Cortez, N.; Guazzaroni, M.E.; Heipieper, H.J.; Klages, S.; Kotsyurbenko, O.R.; Langer, I.; et al. Genome sequence and functional genomic analysis of the oil-degrading bacterium Oleispira antarctica. Nat. Commun. 2013, 4, 2156. [Google Scholar] [CrossRef]
- D’Amico, S.; Claverie, P.; Collins, T.; Georlette, D.; Gratia, E.; Hoyoux, A.; Meuwis, M.A.; Feller, G.; Gerday, C. Molecular basis of cold adaptation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2002, 357, 917–925. [Google Scholar] [CrossRef]
- D’Amico, S.; Collins, T.; Marx, J.C.; Feller, G.; Gerday, C. Psychrophilic microorganisms: Challenges for life. EMBO Rep. 2006, 7, 385–389. [Google Scholar] [CrossRef] [PubMed]
- Leiros, H.K.; Pey, A.L.; Innselset, M.; Moe, E.; Leiros, I.; Steen, I.H.; Martinez, A. Structure of phenylalanine hydroxylase from Colwellia psychrerythraea 34H, a monomeric cold active enzyme with local flexibility around the active site and high overall stability. J. Biol. Chem. 2007, 282, 21973–21986. [Google Scholar] [CrossRef] [PubMed]
- Sonan, G.K.; Receveur-Brechot, V.; Duez, C.; Aghajari, N.; Czjzek, M.; Haser, R.; Gerday, C. The linker region plays a key role in the adaptation to cold of the cellulase from an Antarctic bacterium. Biochem. J. 2007, 407, 293–302. [Google Scholar] [CrossRef] [PubMed]
- Mouri, Y.; Takada, Y. Contribution of Three Different Regions of Isocitrate Dehydrogenases from Psychrophilic and Psychrotolerant Bacteria to Their Thermal Properties. Curr. Microbiol. 2018, 75, 1523–1529. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.W.; Liao, M.L.; Meng, X.L.; Somero, G.N. Structural flexibility and protein adaptation to temperature: Molecular dynamics analysis of malate dehydrogenases of marine molluscs. Proc. Natl. Acad. Sci. USA 2018, 115, 1274–1279. [Google Scholar] [CrossRef] [PubMed]
- Pischedda, A.; Ramasamy, K.P.; Mangiagalli, M.; Chiappori, F.; Milanesi, L.; Miceli, C.; Pucciarelli, S.; Lotti, M. Antarctic marine ciliates under stress: Superoxide dismutases from the psychrophilic Euplotes focardii are cold-active yet heat tolerant enzymes. Sci. Rep. 2018, 8, 14721. [Google Scholar] [CrossRef] [PubMed]
- Tosco, A.; Birolo, L.; Madonna, S.; Lolli, G.; Sannia, G.; Marino, G. GroEL from the psychrophilic bacterium Pseudoalteromonas haloplanktis TAC 125: Molecular characterization and gene cloning. Extremophiles 2003, 7, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Joshi, S.; Satyanarayana, T. Biotechnology of cold-active proteases. Biology 2013, 2, 755–783. [Google Scholar] [CrossRef] [PubMed]
- Marx, J.C.; Collins, T.; D’Amico, S.; Feller, G.; Gerday, C. Cold-adapted enzymes from marine Antarctic microorganisms. Mar. Biotechnol. 2007, 9, 293–304. [Google Scholar] [CrossRef]
- Cavicchioli, R.; Charlton, T.; Ertan, H.; Mohd Omar, S.; Siddiqui, K.S.; Williams, T.J. Biotechnological uses of enzymes from psychrophiles. Microb. Biotechnol. 2011, 4, 449–460. [Google Scholar] [CrossRef]
- Duarte, A.W.F.; Dos Santos, J.A.; Vianna, M.V.; Vieira, J.M.F.; Mallagutti, V.H.; Inforsato, F.J.; Wentzel, L.C.P.; Lario, L.D.; Rodrigues, A.; Pagnocca, F.C.; et al. Cold-adapted enzymes produced by fungi from terrestrial and marine Antarctic environments. Crit. Rev. Biotechnol. 2018, 38, 600–619. [Google Scholar] [CrossRef] [PubMed]
- Cieslinski, H.; Kur, J.; Bialkowska, A.; Baran, I.; Makowski, K.; Turkiewicz, M. Cloning, expression, and purification of a recombinant cold-adapted beta-galactosidase from antarctic bacterium Pseudoalteromonas sp. 22b. Protein Expr. Purif. 2005, 39, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Turkiewicz, M.; Kur, J.; Bialkowska, A.; Cieslinski, H.; Kalinowska, H.; Bielecki, S. Antarctic marine bacterium Pseudoalteromonas sp. 22b as a source of cold-adapted beta-galactosidase. Biomol. Eng. 2003, 20, 317–324. [Google Scholar] [CrossRef]
- Hoyoux, A.; Jennes, I.; Dubois, P.; Genicot, S.; Dubail, F.; Francois, J.M.; Baise, E.; Feller, G.; Gerday, C. Cold-adapted beta-galactosidase from the Antarctic psychrophile Pseudoalteromonas haloplanktis. Appl. Environ. Microbiol. 2001, 67, 1529–1535. [Google Scholar] [CrossRef] [PubMed]
- Gerday, C.; Hoyoux, A.; Marie Francois, J.M.; Dubois, P.; Baise, E.; Jennes, I.; Genicot, S. Cold-Active Beta Galactosidase, the Process for its Preparation and the Use Thereof. U.S. Patent WO2001004276A1, 9 July 1999. [Google Scholar]
- Song, C.; Chi, Z.; Li, J.; Wang, X. beta-Galactosidase production by the psychrotolerant yeast Guehomyces pullulans 17-1 isolated from sea sediment in Antarctica and lactose hydrolysis. Bioprocess. Biosyst. Eng. 2010, 33, 1025–1031. [Google Scholar] [CrossRef] [PubMed]
- Alikkunju, A.P.; Sainjan, N.; Silvester, R.; Joseph, A.; Rahiman, M.; Antony, A.C.; Kumaran, R.C.; Hatha, M. Screening and Characterization of Cold-Active beta-Galactosidase Producing Psychrotrophic Enterobacter ludwigii from the Sediments of Arctic Fjord. Appl. Biochem. Biotechnol. 2016, 180, 477–490. [Google Scholar] [CrossRef]
- Schmidt, M.; Stougaard, P. Identification, cloning and expression of a cold-active beta-galactosidase from a novel Arctic bacterium, Alkalilactibacillus ikkense. Environ. Technol. 2010, 31, 1107–1114. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Kan, G.; Ren, X.; Yu, G.; Shi, C.; Xie, Q.; Wen, H.; Betenbaugh, M. Molecular Cloning and Characterization of a Novel alpha-Amylase from Antarctic Sea Ice Bacterium Pseudoalteromonas sp. M175 and Its Primary Application in Detergent. BioMed Res. Int. 2018, 2018, 3258383. [Google Scholar] [CrossRef] [PubMed]
- Ramli, A.N.; Azhar, M.A.; Shamsir, M.S.; Rabu, A.; Murad, A.M.; Mahadi, N.M.; Illias, R.M. Sequence and structural investigation of a novel psychrophilic alpha-amylase from Glaciozyma antarctica PI12 for cold-adaptation analysis. J. Mol. Model. 2013, 19, 3369–3383. [Google Scholar] [CrossRef] [PubMed]
- Vardhan Reddy, P.V.; Shiva Nageswara Rao, S.S.; Pratibha, M.S.; Sailaja, B.; Kavya, B.; Manorama, R.R.; Singh, S.M.; Radha Srinivas, T.N.; Shivaji, S. Bacterial diversity and bioprospecting for cold-active enzymes from culturable bacteria associated with sediment from a melt water stream of Midtre Lov´enbreen glacier, an Arctic glacier. Res. Microbiol. 2009, 160, 538–546. [Google Scholar] [CrossRef] [PubMed]
- Chessa, J.P.; Feller, G.; Gerday, C. Purification and characterization of the heat-labile alpha-amylase secreted by the psychrophilic bacterium TAC 240B. Can. J. Microbiol. 1999, 45, 452–457. [Google Scholar] [CrossRef] [PubMed]
- Feller, G.; D’Amico, D.; Gerday, C. Thermodynamic stability of a cold-active alpha-amylase from the Antarctic bacterium Alteromonas haloplanctis. Biochemistry 1999, 38, 4613–4619. [Google Scholar] [CrossRef] [PubMed]
- Kuddus, M.; Roohi, J.M.A.; Ramteke, P.W. An Overview of Cold-active Microbial α-amylase: Adaptation Strategies and Biotechnological Potentials. Biotechnol. Biotechnol. Equip. 2011, 10, 246–258. [Google Scholar]
- Del-Cid, A.; Ubilla, P.; Ravanal, M.C.; Medina, E.; Vaca, I.; Levican, G.; Eyzaguirre, J.; Chavez, R. Cold-active xylanase produced by fungi associated with Antarctic marine sponges. Appl. Biochem. Biotechnol. 2014, 172, 524–532. [Google Scholar] [CrossRef] [PubMed]
- Humphry, D.R.; George, A.; Black, G.W.; Cummings, S.P. Flavobacterium frigidarium sp. nov., an aerobic, psychrophilic, xylanolytic and laminarinolytic bacterium from Antarctica. Int. J. Syst. Evol. Microbiol. 2001, 51, 1235–1243. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, K.; Wintrode, P.L.; Grayling, R.A.; Rubingh, D.N.; Arnold, F.H. Directed evolution study of temperature adaptation in a psychrophilic enzyme. J. Mol. Biol. 2000, 297, 1015–1026. [Google Scholar] [CrossRef] [PubMed]
- Narinx, E.; Davail, S.; Feller, G.; Gerday, C. Nucleotide and derived amino acid sequence of the subtilisin from the antarctic psychrotroph Bacillus TA39. Biochim. Biophys. Acta 1992, 1131, 111–113. [Google Scholar] [CrossRef]
- Davail, S.; Feller, G.; Narinx, E.; Gerday, C. Cold adaptation of proteins. Purification, characterization, and sequence of the heat-labile subtilisin from the antarctic psychrophile Bacillus TA41. J. Biol. Chem. 1994, 269, 17448–17453. [Google Scholar]
- Wang, Q.F.; Miao, J.L.; Hou, Y.H.; Ding, Y.; Wang, G.D.; Li, G.Y. Purification and characterization of an extracellular cold-active serine protease from the psychrophilic bacterium Colwellia sp. NJ341. Biotechnol. Lett. 2005, 27, 1195–1198. [Google Scholar] [CrossRef]
- Kulakova, L.; Galkin, A.; Kurihara, T.; Yoshimura, T.; Esaki, N. Cold-active serine alkaline protease from the psychrotrophic bacterium Shewanella strain ac10: Gene cloning and enzyme purification and characterization. Appl. Environ. Microbiol. 1999, 65, 611–617. [Google Scholar]
- Lario, L.D.; Chaud, L.; Almeida, M.D.G.; Converti, A.; Duraes Sette, L.; Pessoa, A., Jr. Production, purification, and characterization of an extracellular acid protease from the marine Antarctic yeast Rhodotorula mucilaginosa L7. Fungal Biol. 2015, 119, 1129–1136. [Google Scholar] [CrossRef] [PubMed]
- Acevedo, J.P.; Rodriguez, V.; Saavedra, M.; Munoz, M.; Salazar, O.; Asenjo, J.A.; Andrews, B.A. Cloning, expression and decoding of the cold adaptation of a new widely represented thermolabile subtilisin-like protease. J. Appl. Microbiol. 2013, 114, 352–363. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.-F.; Hou, Y.-H.; Xu, Z.; Miao, J.-L.; Li, G.-Y. Purification and properties of an extracellular cold-active protease from the psychrophilic bacterium Pseudoalteromonas sp. NJ276. Biochem. Eng. J. 2008, 38, 362–368. [Google Scholar] [CrossRef]
- Turkiewicz, M.; Pazgier, M.; Kalinowska, H.; Bielecki, S. A cold-adapted extracellular serine proteinase of the yeast Leucosporidium antarcticum. Extremophiles 2003, 7, 435–442. [Google Scholar] [CrossRef] [PubMed]
- de Pascale, D.; Giuliani, M.; De Santi, C.; Bergamasco, N.; Amoresano, A.; Carpentieri, A.; Parrilli, E.; Tutino, M.L. PhAP protease from Pseudoalteromonas haloplanktis TAC125: Gene cloning, recombinant production in E. coli and enzyme characterization. Polar Sci. 2010, 4, 285–294. [Google Scholar] [CrossRef]
- Huston, A.L.; Methe, B.; Deming, J.W. Purification, characterization, and sequencing of an extracellular cold-active aminopeptidase produced by marine psychrophile Colwellia psychrerythraea strain 34H. Appl. Environ. Microbiol. 2004, 70, 3321–3328. [Google Scholar] [CrossRef] [PubMed]
- Huston, A.L.; Haeggstrom, J.Z.; Feller, G. Cold adaptation of enzymes: Structural, kinetic and microcalorimetric characterizations of an aminopeptidase from the Arctic psychrophile Colwellia psychrerythraea and of human leukotriene A(4) hydrolase. Biochim. Biophys. Acta 2008, 1784, 1865–1872. [Google Scholar] [CrossRef]
- Pereira, J.Q.; Ambrosini, A.; Passaglia, L.M.P.; Brandelli, A. A new cold-adapted serine peptidase from Antarctic Lysobacter sp. A03: Insights about enzyme activity at low temperatures. Int. J. Biol. Macromol. 2017, 103, 854–862. [Google Scholar] [CrossRef]
- Larsen, A.N.; Moe, E.; Helland, R.; Gjellesvik, D.R.; Willassen, N.P. Characterization of a recombinantly expressed proteinase K-like enzyme from a psychrotrophic Serratia sp. FEBS J. 2006, 273, 47–60. [Google Scholar] [CrossRef]
- Xie, B.B.; Bian, F.; Chen, X.L.; He, H.L.; Guo, J.; Gao, X.; Zeng, Y.X.; Chen, B.; Zhou, B.C.; Zhang, Y.Z. Cold adaptation of zinc metalloproteases in the thermolysin family from deep sea and arctic sea ice bacteria revealed by catalytic and structural properties and molecular dynamics: New insights into relationship between conformational flexibility and hydrogen bonding. J. Biol. Chem. 2009, 284, 9257–9269. [Google Scholar] [CrossRef]
- Turkiewicz, M.; Gromek, E.; Kalinowska, H.; Zielińska, M. Biosynthesis and properties of an extracellular metalloprotease from the Antarctic marine bacterium Sphingomonas paucimobilis. J. Biotechnol. 1999, 70, 53–60. [Google Scholar] [CrossRef]
- Denner, E.B.; Mark, B.; Busse, H.J.; Turkiewicz, M.; Lubitz, W. Psychrobacter proteolyticus sp. nov., a psychrotrophic, halotolerant bacterium isolated from the Antarctic krill Euphausia superba Dana, excreting a cold-adapted metalloprotease. Syst. Appl. Microbiol. 2001, 24, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://arcticzymes.com/technology/proteinase/ (accessed on 15 July 2019).
- Wi, A.R.; Jeon, S.J.; Kim, S.; Park, H.J.; Kim, D.; Han, S.J.; Yim, J.H.; Kim, H.W. Characterization and a point mutational approach of a psychrophilic lipase from an arctic bacterium, Bacillus pumilus. Biotechnol. Lett. 2014, 36, 1295–1302. [Google Scholar] [CrossRef] [PubMed]
- de Pascale, D.; Cusano, A.M.; Autore, F.; Parrilli, E.; di Prisco, G.; Marino, G.; Tutino, M.L. The cold-active Lip1 lipase from the Antarctic bacterium Pseudoalteromonas haloplanktis TAC125 is a member of a new bacterial lipolytic enzyme family. Extremophiles 2008, 12, 311–323. [Google Scholar] [CrossRef] [PubMed]
- Do, H.; Lee, J.H.; Kwon, M.H.; Song, H.E.; An, J.Y.; Eom, S.H.; Lee, S.G.; Kim, H.J. Purification, characterization and preliminary X-ray diffraction analysis of a cold-active lipase (CpsLip) from the psychrophilic bacterium Colwellia psychrerythraea 34H. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2013, 69, 920–924. [Google Scholar] [CrossRef] [PubMed]
- Irgens, R.L.; Gosink, J.J.; Staley, J.T. Polaromonas vacuolata gen. nov., sp. nov., a psychrophilic, marine, gas vacuolate bacterium from Antarctica. Int. J. Syst. Bacteriol. 1996, 46, 822–826. [Google Scholar] [CrossRef][Green Version]
- Parra, L.P.; Reyes, F.; Acevedo, J.P.; Salazar, O.; Andrews, B.A.; Asenjo, J.A. Cloning and fusion expression of a cold-active lipase from marine Antarctic origin. Enzym. Microb. Technol. 2008, 42, 371–377. [Google Scholar] [CrossRef]
- Xuezheng, L.; Shuoshuo, C.; Guoying, X.; Shuai, W.; Ning, D.; Jihong, S. Cloning and heterologous expression of two cold-active lipases from the Antarctic bacterium Psychrobacter sp. G. Polar Res. 2010, 29, 421–429. [Google Scholar] [CrossRef]
- Parra, L.P.; Espina, G.; Devia, J.; Salazar, O.; Andrews, B.; Asenjo, J.A. Identification of lipase encoding genes from Antarctic seawater bacteria using degenerate primers: Expression of a cold-active lipase with high specific activity. Enzym. Microb. Technol. 2015, 68, 56–61. [Google Scholar] [CrossRef]
- Feller, G.; Thiry, M.; Arpigny, J.L.; Gerday, C. Cloning and expression in Escherichia coli of three lipase-encoding genes from the psychrotrophic antarctic strain Moraxella TA144. Gene 1991, 102, 111–115. [Google Scholar] [CrossRef]
- Zhang, J.; Lin, S.; Zeng, R. Cloning, expression, and characterization of a cold-adapted lipase gene from an antarctic deep-sea psychrotrophic bacterium, Psychrobacter sp 7195. J. Microbiol. Biotechnol. 2007, 17, 604–610. [Google Scholar] [PubMed]
- Yang, X.; Lin, X.; Fan, T.; Bian, J.; Huang, X. Cloning and expression of lipP, a gene encoding a cold-adapted lipase from Moritella sp.2-5-10-1. Curr. Microbiol. 2008, 56, 194–198. [Google Scholar] [CrossRef]
- Lo Giudice, A.; Michaud, L.; de Pascale, D.; De Domenico, M.; di Prisco, G.; Fani, R.; Bruni, V. Lipolytic activity of Antarctic cold-adapted marine bacteria (Terra Nova Bay, Ross Sea). J. Appl. Microbiol. 2006, 101, 1039–1048. [Google Scholar] [CrossRef] [PubMed]
- Yu, P.; Wang, X.T.; Liu, J.W. Purification and characterization of a novel cold-adapted phytase from Rhodotorula mucilaginosa strain JMUY14 isolated from Antarctic. J. Basic Microbiol. 2015, 55, 1029–1039. [Google Scholar] [CrossRef] [PubMed]
- Al Khudary, R.; Venkatachalam, R.; Katzer, M.; Elleuche, S.; Antranikian, G. A cold-adapted esterase of a novel marine isolate, Pseudoalteromonas arctica: Gene cloning, enzyme purification and characterization. Extremophiles 2010, 14, 273–285. [Google Scholar] [CrossRef] [PubMed]
- De Santi, C.; Leiros, H.K.; Di Scala, A.; de Pascale, D.; Altermark, B.; Willassen, N.P. Biochemical characterization and structural analysis of a new cold-active and salt-tolerant esterase from the marine bacterium Thalassospira sp. Extremophiles 2016, 20, 323–336. [Google Scholar] [CrossRef]
- Lemak, S.; Tchigvintsev, A.; Petit, P.; Flick, R.; Singer, A.U.; Brown, G.; Evdokimova, E.; Egorova, O.; Gonzalez, C.F.; Chernikova, T.N.; et al. Structure and activity of the cold-active and anion-activated carboxyl esterase OLEI01171 from the oil-degrading marine bacterium Oleispira antarctica. Biochem. J. 2012, 445, 193–203. [Google Scholar] [CrossRef]
- D’Auria, S.; Aurilia, V.; Marabotti, A.; Gonnelli, M.; Strambini, G. Structure and dynamics of cold-adapted enzymes as investigated by phosphorescence spectroscopy and molecular dynamics studies. 2. The case of an esterase from Pseudoalteromonas haloplanktis. J. Phys. Chem. B 2009, 113, 13171–13178. [Google Scholar] [CrossRef]
- Aurilia, V.; Parracino, A.; Saviano, M.; Rossi, M.; D’Auria, S. The psychrophilic bacterium Pseudoalteromonas halosplanktis TAC125 possesses a gene coding for a cold-adapted feruloyl esterase activity that shares homology with esterase enzymes from gamma-proteobacteria and yeast. Gene 2007, 397, 51–57. [Google Scholar] [CrossRef]
- Cieslinski, H.; Bialkowska, A.M.; Dlugolecka, A.; Daroch, M.; Tkaczuk, K.L.; Kalinowska, H.; Kur, J.; Turkiewicz, M. A cold-adapted esterase from psychrotrophic Pseudoalteromas sp. strain 643A. Arch. Microbiol. 2007, 188, 27–36. [Google Scholar] [CrossRef]
- De Santi, C.; Altermark, B.; Pierechod, M.M.; Ambrosino, L.; de Pascale, D.; Willassen, N.P. Characterization of a cold-active and salt tolerant esterase identified by functional screening of Arctic metagenomic libraries. BMC Biochem. 2016, 17, 1. [Google Scholar] [CrossRef] [PubMed]
- Jeon, J.H.; Kim, J.T.; Kang, S.G.; Lee, J.H.; Kim, S.J. Characterization and its potential application of two esterases derived from the arctic sediment metagenome. Mar. Biotechnol. 2009, 11, 307–316. [Google Scholar] [CrossRef]
- Kang, J.H.; Woo, J.H.; Kang, S.G.; Hwang, Y.O.; Kim, S.J. A cold-adapted epoxide hydrolase from a strict marine bacterium, Sphingophyxis alaskensis. J. Microbiol. Biotechnol. 2008, 18, 1445–1452. [Google Scholar] [PubMed]
- Alterio, V.; Aurilia, V.; Romanelli, A.; Parracino, A.; Saviano, M.; D’Auria, S.; De Simone, G. Crystal structure of an S-formylglutathione hydrolase from Pseudoalteromonas haloplanktis TAC125. Biopolymers 2010, 93, 669–677. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.W.; Yoo, W.; Park, S.-H.; Le, L.T.H.L.; Jeong, C.-S.; Ryu, B.H.; Shin, S.C.; Kim, H.-W.; Park, H.; Kim, K.K.; et al. Structural and functional characterization of a novel cold-active S-formylglutathione hydrolase (SfSFGH) homolog from Shewanella frigidimarina, a psychrophilic bacterium. Microb. Cell Fact. 2019, 18, 140. [Google Scholar] [CrossRef] [PubMed]
- Ramya, L.N.; Pulicherla, K.K. Molecular insights into cold active polygalacturonase enzyme for its potential application in food processing. J. Food Sci. Technol. 2015, 52, 5484–5496. [Google Scholar] [CrossRef] [PubMed]
- Elleuche, S.; Qoura, F.M.; Lorenz, U.; Rehn, T.; Brück, T.; Antranikian, G. Cloning, expression and characterization of the recombinant cold-active type-I pullulanase from Shewanella arctica. J. Mol. Catal. B Enzym. 2015, 116, 70–77. [Google Scholar] [CrossRef]
- Turkiewicz, M.; Pazgier, M.; Donachie, S.; Kalinowska, H. Invertase and a-glucosidase production by the endemic Antarctic marine yeast Leucosporidium antarcticum. Pol. Polar Res. 2005, 26, 125–136. [Google Scholar]
- Violot, S.; Aghajari, N.; Czjzek, M.; Feller, G.; Sonan, G.K.; Gouet, P.; Gerday, C.; Haser, R.; Receveur-Brechot, V. Structure of a full length psychrophilic cellulase from Pseudoalteromonas haloplanktis revealed by X-ray diffraction and small angle X-ray scattering. J. Mol. Biol. 2005, 348, 1211–1224. [Google Scholar] [CrossRef]
- Lonhienne, T.; Zoidakis, J.; Vorgias, C.E.; Feller, G.; Gerday, C.; Bouriotis, V. Modular structure, local flexibility and cold-activity of a novel chitobiase from a psychrophilic Antarctic bacterium. J. Mol. Biol. 2001, 310, 291–297. [Google Scholar] [CrossRef]
- Rina, M.; Pozidis, C.; Mavromatis, K.; Tzanodaskalaki, M.; Kokkinidis, M.; Bouriotis, V. Alkaline phosphatase from the Antarctic strain TAB5. Properties and psychrophilic adaptations. Eur. J. Biochem. 2000, 267, 1230–1238. [Google Scholar] [CrossRef] [PubMed]
- Koutsioulis, D.; Wang, E.; Tzanodaskalaki, M.; Nikiforaki, D.; Deli, A.; Feller, G.; Heikinheimo, P.; Bouriotis, V. Directed evolution on the cold adapted properties of TAB5 alkaline phosphatase. Protein Eng. Des. Sel. 2008, 21, 319–327. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://www.neb.com/products/m0289-antarctic-phosphatase#Product%20Information (accessed on 15 July 2019).
- Tsuruta, H.; Mikami, B.; Higashi, T.; Aizono, Y. Crystal structure of cold-active alkaline phosphatase from the psychrophile Shewanella sp. Biosci. Biotechnol. Biochem. 2010, 74, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://www.takarabio.com/products/cloning/modifying-enzymes/nucleases/ cryonase-cold-active-nuclease (accessed on 15 July 2019).
- Available online: https://arcticzymes.com/technology/hl-exol/ (accessed on 15 July 2019).
- Wang, Y.; Hou, Y.; Nie, P.; Wang, Y.; Ren, X.; Wei, Q.; Wang, Q. A Novel Cold-Adapted and Salt-Tolerant RNase R from Antarctic Sea-Ice Bacterium Psychrobacter sp. ANT206. Molecules 2019, 24, 2229. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://international.neb.com/products/m0372-antarctic-thermolabile-udg#Product%20 Information (accessed on 15 July 2019).
- Tsigos, I.; Velonia, K.; Smonou, I.; Bouriotis, V. Purification and characterization of an alcohol dehydrogenase from the Antarctic psychrophile Moraxella sp. TAE123. Eur. J. Biochem. 1998, 254, 356–362. [Google Scholar] [CrossRef]
- Galkin, A.; Kulakova, L.; Ashida, H.; Sawa, Y.; Esaki, N. Cold-adapted alanine dehydrogenases from two antarctic bacterial strains: Gene cloning, protein characterization, and comparison with mesophilic and thermophilic counterparts. Appl. Environ. Microbiol. 1999, 65, 4014–4020. [Google Scholar]
- Wang, Y.; Hou, Y.; Wang, Y.; Zheng, L.; Xu, X.; Pan, K.; Li, R.; Wang, Q. A Novel Cold-Adapted Leucine Dehydrogenase from Antarctic Sea-Ice Bacterium Pseudoalteromonas sp. ANT178. Mar. Drugs 2018, 16, 359. [Google Scholar] [CrossRef] [PubMed]
- Oikawa, T.; Yamamoto, N.; Shimoke, K.; Uesato, S.; Ikeuchi, T.; Fujioka, T. Purification, characterization, and overexpression of psychrophilic and thermolabile malate dehydrogenase of a novel antarctic psychrotolerant, Flavobacterium frigidimaris KUC-1. Biosci. Biotechnol. Biochem. 2005, 69, 2146–2154. [Google Scholar] [CrossRef]
- Fedoy, A.E.; Yang, N.; Martinez, A.; Leiros, H.K.; Steen, I.H. Structural and functional properties of isocitrate dehydrogenase from the psychrophilic bacterium Desulfotalea psychrophila reveal a cold-active enzyme with an unusual high thermal stability. J. Mol. Biol. 2007, 372, 130–149. [Google Scholar] [CrossRef]
- Yoneda, K.; Sakuraba, H.; Muraoka, I.; Oikawa, T.; Ohshima, T. Crystal structure of UDP-galactose 4-epimerase-like L-threonine dehydrogenase belonging to the intermediate short-chain dehydrogenase-reductase superfamily. FEBS J. 2010, 277, 5124–5132. [Google Scholar] [CrossRef]
- Merlino, A.; Russo Krauss, I.; Castellano, I.; De Vendittis, E.; Rossi, B.; Conte, M.; Vergara, A.; Sica, F. Structure and flexibility in cold-adapted iron superoxide dismutases: The case of the enzyme isolated from Pseudoalteromonas haloplanktis. J. Struct. Biol. 2010, 172, 343–352. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Z.; Jiang, Y.H.; Miao, J.L.; Wang, Q.F.; Zhang, B.T.; Li, G.Y. Purification and characterization of a cold-active iron superoxide dismutase from a Psychrophilic Bacterium, Marinomonas sp. NJ522. Biotechnol. Lett. 2006, 28, 85–88. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.F.; Wang, Y.F.; Hou, Y.H.; Shi, Y.L.; Han, H.; Miao, M.; Wu, Y.Y.; Liu, Y.P.; Yue, X.N.; Li, Y.J. Cloning, expression and biochemical characterization of recombinant superoxide dismutase from Antarctic psychrophilic bacterium Pseudoalteromonas sp. ANT506. J. Basic Microbiol. 2016, 56, 753–761. [Google Scholar] [CrossRef] [PubMed]
- Na, J.; Im, H.; Lee, K. Expression and Purification of Recombinant Superoxide Dismutase (PaSOD) from Psychromonas arctica in Escherichia coli. Bull. Korean Chem. Soc. 2011, 32, 2405–2409. [Google Scholar] [CrossRef][Green Version]
- Kan, G.; Wen, H.; Wang, X.; Zhou, T.; Shi, C. Cloning and characterization of iron-superoxide dismutase in Antarctic yeast strain Rhodotorula mucilaginosa AN5. J. Basic Microbiol. 2017, 57, 680–690. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Sun, M.; Liu, W.; Zhang, B. Purification and characterization of a psychrophilic catalase from Antarctic Bacillus. Can. J. Microbiol. 2008, 54, 823–828. [Google Scholar] [CrossRef] [PubMed]
- Sarmiento, F.; Peralta, R.; Blamey, J.M. Cold and Hot Extremozymes: Industrial Relevance and Current Trends. Front. Bioeng. Biotechnol. 2015, 3, 148. [Google Scholar] [CrossRef] [PubMed]
- Ji, M.; Barnwell, C.V.; Grunden, A.M. Characterization of recombinant glutathione reductase from the psychrophilic Antarctic bacterium Colwellia psychrerythraea. Extremophiles 2015, 19, 863–874. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Han, H.; Cui, B.; Hou, Y.; Wang, Y.; Wang, Q. A glutathione peroxidase from Antarctic psychrotrophic bacterium Pseudoalteromonas sp. ANT506: Cloning and heterologous expression of the gene and characterization of recombinant enzyme. Bioengineered 2017, 8, 742–749. [Google Scholar] [CrossRef]
- Cotugno, R.; Rosaria Ruocco, M.; Marco, S.; Falasca, P.; Evangelista, G.; Raimo, G.; Chambery, A.; Di Maro, A.; Masullo, M.; De Vendittis, E. Differential cold-adaptation among protein components of the thioredoxin system in the psychrophilic eubacterium Pseudoalteromonas haloplanktis TAC 125. Mol. BioSyst. 2009, 5, 519–528. [Google Scholar] [CrossRef]
- Wang, Q.; Hou, Y.; Shi, Y.; Han, X.; Chen, Q.; Hu, Z.; Liu, Y.; Li, Y. Cloning, expression, purification, and characterization of glutaredoxin from Antarctic sea-ice bacterium Pseudoalteromonas sp. AN178. BioMed Res. Int. 2014, 2014, 246871. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Hou, Y.; Wang, Y.; Lu, Z.; Song, C.; Xu, Y.; Wei, N.; Wang, Q. Cloning, expression and enzymatic characteristics of a 2-Cys peroxiredoxin from Antarctic sea-ice bacterium Psychrobacter sp. ANT206. Int. J. Biol. Macromol. 2019, 129, 1047–1055. [Google Scholar] [CrossRef] [PubMed]
- Birolo, L.; Tutino, M.L.; Fontanella, B.; Gerday, C.; Mainolfi, K.; Pascarella, S.; Sannia, G.; Vinci, F.; Marino, G. Aspartate aminotransferase from the Antarctic bacterium Pseudoalteromonas haloplanktis TAC 125. Cloning, expression, properties, and molecular modelling. Eur. J. Biochem. 2000, 267, 2790–2802. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Wang, Q.; Hou, Y.; Hong, Y.; Han, X.; Yi, J.; Qu, J.; Lu, Y. Molecular cloning, expression and enzymatic characterization of glutathione S-transferase from Antarctic sea-ice bacteria Pseudoalteromonas sp. ANT506. Microbiol. Res. 2014, 169, 179–184. [Google Scholar] [CrossRef] [PubMed]
- Angelaccio, S.; Florio, R.; Consalvi, V.; Festa, G.; Pascarella, S. Serine hydroxymethyltransferase from the cold adapted microorganism Psychromonas ingrahamii: A low temperature active enzyme with broad substrate specificity. Int. J. Mol. Sci. 2012, 13, 1314–1326. [Google Scholar] [CrossRef] [PubMed]
- Albino, A.; Marco, S.; Di Maro, A.; Chambery, A.; Masullo, M.; De Vendittis, E. Characterization of a cold-adapted glutathione synthetase from the psychrophile Pseudoalteromonas haloplanktis. Mol. BioSyst. 2012, 8, 2405–2414. [Google Scholar] [CrossRef]
- Georlette, D.; Jonsson, Z.O.; Van Petegem, F.; Chessa, J.; Van Beeumen, J.; Hubscher, U.; Gerday, C. A DNA ligase from the psychrophile Pseudoalteromonas haloplanktis gives insights into the adaptation of proteins to low temperatures. Eur. J. Biochem. 2000, 267, 3502–3512. [Google Scholar] [CrossRef] [PubMed]
- De Luca, V.; Vullo, D.; Del Prete, S.; Carginale, V.; Osman, S.M.; AlOthman, Z.; Supuran, C.T.; Capasso, C. Cloning, characterization and anion inhibition studies of a gamma-carbonic anhydrase from the Antarctic bacterium Colwellia psychrerythraea. Bioorg. Med. Chem. 2016, 24, 835–840. [Google Scholar] [CrossRef] [PubMed]
- De Luca, V.; Vullo, D.; Del Prete, S.; Carginale, V.; Scozzafava, A.; Osman, S.M.; AlOthman, Z.; Supuran, C.T.; Capasso, C. Cloning, characterization and anion inhibition studies of a new gamma-carbonic anhydrase from the Antarctic bacterium Pseudoalteromonas haloplanktis. Bioorg. Med. Chem. 2015, 23, 4405–4409. [Google Scholar] [CrossRef]
- Angeli, A.; Del Prete, S.; Osman, S.M.; AlOthman, Z.; Donald, W.A.; Capasso, C.; Supuran, C.T. Activation Studies of the gamma-Carbonic Anhydrases from the Antarctic Marine Bacteria Pseudoalteromonas haloplanktis and Colwellia psychrerythraea with Amino Acids and Amines. Mar. Drugs 2019, 17, 238. [Google Scholar] [CrossRef]
- Truong, L.V.; Tuyen, H.; Helmke, E.; Binh, L.T.; Schweder, T. Cloning of two pectate lyase genes from the marine Antarctic bacterium Pseudoalteromonas haloplanktis strain ANT/505 and characterization of the enzymes. Extremophiles 2001, 5, 35–44. [Google Scholar] [CrossRef] [PubMed]
- Do, H.; Kim, S.J.; Lee, C.W.; Kim, H.W.; Park, H.H.; Kim, H.M.; Park, H.; Park, H.; Lee, J.H. Crystal structure of UbiX, an aromatic acid decarboxylase from the psychrophilic bacterium Colwellia psychrerythraea that undergoes FMN-induced conformational changes. Sci. Rep. 2015, 5, 8196. [Google Scholar] [CrossRef]
- Do, H.; Lee, C.W.; Han, S.J.; Lee, S.G.; Kim, H.J.; Park, H.; Lee, J.H. Purification, crystallization and preliminary X-ray crystallographic studies of FMN-bound and FMN-free forms of aromatic acid decarboxylase (CpsUbiX) from the psychrophilic bacterium Colwellia psychrerythraea 34H. Acta Crystallogr. Sect. F Struct. Biol. Commun. 2014, 70, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Do, H.; Yun, J.S.; Lee, C.W.; Choi, Y.J.; Kim, H.Y.; Kim, Y.J.; Park, H.; Chang, J.H.; Lee, J.H. Crystal Structure and Comparative Sequence Analysis of GmhA from Colwellia psychrerythraea Strain 34H Provides Insight into Functional Similarity with DiaA. Mol. Cells 2015, 38, 1086–1095. [Google Scholar] [CrossRef] [PubMed]
- See Too, W.C.; Few, L.L. Cloning of triose phosphate isomerase gene from an antarctic psychrophilic Pseudomonas sp. by degenerate and splinkerette PCR. World J. Microbiol. Biotechnol. 2010, 26, 1251–1259. [Google Scholar] [CrossRef] [PubMed]
- Rentier-Delrue, F.; Mande, S.C.; Moyens, S.; Terpstra, P.; Mainfroid, V.; Goraj, K.; Lion, M.; Hol, W.G.; Martial, J.A. Cloning and overexpression of the triosephosphate isomerase genes from psychrophilic and thermophilic bacteria. Structural comparison of the predicted protein sequences. J. Mol. Biol. 1993, 229, 85–93. [Google Scholar] [CrossRef] [PubMed]
- D’Amico, S.; Marx, J.C.; Gerday, C.; Feller, G. Activity-stability relationships in extremophilic enzymes. J. Biol. Chem. 2003, 278, 7891–7896. [Google Scholar] [CrossRef] [PubMed]
- Tutino, M.L.; Duilio, A.; Moretti, M.A.; Sannia, G.; Marino, G. A rolling-circle plasmid from Psychrobacter sp. TA144: Evidence for a novel rep subfamily. Biochem. Biophys. Res. Commun. 2000, 274, 488–495. [Google Scholar] [CrossRef] [PubMed]
- Kazuoka, T.; Takigawa, S.; Arakawa, N.; Hizukuri, Y.; Muraoka, I.; Oikawa, T.; Soda, K. Novel psychrophilic and thermolabile L-threonine dehydrogenase from psychrophilic Cytophaga sp. strain KUC-1. J. Bacteriol. 2003, 185, 4483–4489. [Google Scholar] [CrossRef]
- Tutino, M.L.; Birolo, L.; Fontanella, B.; Mainolfi, K.; Vinci, F.; Sannia, G.; Marino, G. Aspartate aminotransferase from Moraxella TAC125: An unusual psychrophilic enzyme. In Cold-Adapted Organisms; Springer: Berlin/Heidelberg, Germany, 1999. [Google Scholar]
- Adrio, J.L.; Demain, A.L. Microbial enzymes: Tools for biotechnological processes. Biomolecules 2014, 4, 117–139. [Google Scholar] [CrossRef]
- Speciale, G.; Thompson, A.J.; Davies, G.J.; Williams, S.J. Dissecting conformational contributions to glycosidase catalysis and inhibition. Curr. Opin. Struct. Biol. 2014, 28, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Lombard, V.; Golaconda Ramulu, H.; Drula, E.; Coutinho, P.M.; Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014, 42, D490–D495. [Google Scholar] [CrossRef] [PubMed]
- Dalmaso, G.Z.; Ferreira, D.; Vermelho, A.B. Marine extremophiles: A source of hydrolases for biotechnological applications. Mar. Drugs 2015, 13, 1925–1965. [Google Scholar] [CrossRef] [PubMed]
- Shukla, T.P.; Wierzbicki, L.E. Beta-galactosidase technology: A solution to the lactose problem. CRC Crit. Rev. Food Technol. 1975, 5, 325–356. [Google Scholar] [CrossRef]
- Karan, R.; Capes, M.D.; DasSarma, P.; DasSarma, S. Cloning, overexpression, purification, and characterization of a polyextremophilic beta-galactosidase from the Antarctic haloarchaeon Halorubrum lacusprofundi. BMC Biotechnol. 2013, 13, 3. [Google Scholar] [CrossRef]
- Xu, K.; Tang, X.; Gai, Y.; Mehmood, M.; Xiao, X.; Wang, F. Molecular characterization of cold-inducible beta-galactosidase from Arthrobacter sp. ON14 isolated from Antarctica. J. Microbiol. Biotechnol. 2011, 21, 236–242. [Google Scholar] [PubMed]
- Bialkowska, A.M.; Cieslinski, H.; Nowakowska, K.M.; Kur, J.; Turkiewicz, M. A new beta-galactosidase with a low temperature optimum isolated from the Antarctic Arthrobacter sp. 20B: Gene cloning, purification and characterization. Arch. Microbiol. 2009, 191, 825–835. [Google Scholar] [CrossRef]
- Makowski, K.; Bialkowska, A.; Szczesna-Antczak, M.; Kalinowska, H.; Kur, J.; Cieslinski, H.; Turkiewicz, M. Immobilized preparation of cold-adapted and halotolerant Antarctic beta-galactosidase as a highly stable catalyst in lactose hydrolysis. FEMS Microbiol. Ecol. 2007, 59, 535–542. [Google Scholar] [CrossRef]
- Van de Voorde, I. Evaluation of the cold-active Pseudoalteromonas haloplanktis β-galactosidase enzyme for lactose hydrolysis in whey permeate as primary step of d-tagatose production. Process. Biochem. 2014, 49, 2134–2140. [Google Scholar] [CrossRef]
- Asghar, S.; Lee, C.R.; Park, J.S.; Chi, W.J.; Kang, D.K.; Hong, S.K. Identification and biochemical characterization of a novel cold-adapted 1,3-alpha-3,6-anhydro-L-galactosidase, Ahg786, from Gayadomonas joobiniege G7. Appl. Microbiol. Biotechnol. 2018, 102, 8855–8866. [Google Scholar] [CrossRef]
- Roohi, R.; Kuddus, M.; Saima, S. Cold-active detergent-stable extracellular α-amylase from Bacillus cereus GA6: Biochemical characteristics and its perspectives in laundry detergent formulation. J. Biochem. Technol. 2013, 4, 636–644. [Google Scholar]
- Qin, Y.; Huang, Z.; Liu, Z. A novel cold-active and salt-tolerant alpha-amylase from marine bacterium Zunongwangia profunda: Molecular cloning, heterologous expression and biochemical characterization. Extremophiles 2014, 18, 271–281. [Google Scholar] [CrossRef]
- Dornez, E.; Verjans, P.; Arnaut, F.; Delcour, J.A.; Courtin, C.M. Use of psychrophilic xylanases provides insight into the xylanase functionality in bread making. J. Agric. Food Chem. 2011, 59, 9553–9562. [Google Scholar] [CrossRef]
- Collins, T.; Meuwis, M.A.; Stals, I.; Claeyssens, M.; Feller, G.; Gerday, C. A novel family 8 xylanase, functional and physicochemical characterization. J. Biol. Chem. 2002, 277, 35133–35139. [Google Scholar] [CrossRef]
- Lopez-Otin, C.; Bond, J.S. Proteases: Multifunctional enzymes in life and disease. J. Biol. Chem. 2008, 283, 30433–30437. [Google Scholar] [CrossRef]
- Li, Q.; Yi, L.; Marek, P.; Iverson, B.L. Commercial proteases: Present and future. FEBS Lett. 2013, 587, 1155–1163. [Google Scholar] [CrossRef]
- Vojcic, L.; Pitzler, C.; Korfer, G.; Jakob, F.; Ronny, M.; Maurer, K.H.; Schwaneberg, U. Advances in protease engineering for laundry detergents. New Biotechnol. 2015, 32, 629–634. [Google Scholar] [CrossRef]
- Elleuche, S.; Schafers, C.; Blank, S.; Schroder, C.; Antranikian, G. Exploration of extremophiles for high temperature biotechnological processes. Curr. Opin. Microbiol. 2015, 25, 113–119. [Google Scholar] [CrossRef]
- Kuddus, M.; Ramteke, P.W. Recent developments in production and biotechnological applications of cold-active microbial proteases. Crit. Rev. Microbiol. 2012, 38, 330–338. [Google Scholar] [CrossRef]
- Tindbaek, N.; Svendsen, A.; Oestergaard, P.R.; Draborg, H. Engineering a substrate-specific cold-adapted subtilisin. Protein Eng. Des. Sel. 2004, 17, 149–156. [Google Scholar] [CrossRef]
- Bialkowska, A.M.; Krysiak, J.; Florczak, T.; Szulczewska, K.M.; Wanarska, M.; Turkiewicz, M. The psychrotrophic yeast Sporobolomyces roseus LOCK 1119 as a source of a highly active aspartic protease for the in vitro production of antioxidant peptides. Biotechnol. Appl. Biochem. 2018, 65, 726–738. [Google Scholar] [CrossRef]
- Kim, H.D.; Kim, S.M.; Choi, J.I. Purification, Characterization, and Cloning of a Cold-Adapted Protease from Antarctic Janthinobacterium lividum. J. Microbiol. Biotechnol. 2018, 28, 448–453. [Google Scholar] [CrossRef]
- Craik, C.S.; Page, M.J.; Madison, E.L. Proteases as therapeutics. Biochem. J. 2011, 435, 1–16. [Google Scholar] [CrossRef]
- Fornbacke, M.; Clarsund, M. Cold-adapted proteases as an emerging class of therapeutics. Infect. Dis. Ther. 2013, 2, 15–26. [Google Scholar] [CrossRef]
- Jaeger, K.E.; Eggert, T. Lipases for biotechnology. Curr. Opin. Biotechnol. 2002, 13, 390–397. [Google Scholar] [CrossRef]
- Joseph, B.; Ramteke, P.W.; Thomas, G. Cold active microbial lipases: Some hot issues and recent developments. Biotechnol. Adv. 2008, 26, 457–470. [Google Scholar] [CrossRef]
- Gotor-Fernández, V.; Brieva, R.; Gotor, V. Lipases: Useful biocatalysts for the preparation of pharmaceuticals. J. Mol. Catal. B Enzym. 2006, 40, 111–120. [Google Scholar] [CrossRef]
- Gotor-Fernández, V.; Busto, E.; Gotor, V. Candida antarctica Lipase B: An Ideal Biocatalyst for the Preparation of Nitrogenated Organic Compounds. Adv. Synth. Catal. 2006, 348, 797–812. [Google Scholar] [CrossRef]
- Maharana, A.K.; Singh, S.M. A cold and organic solvent tolerant lipase produced by Antarctic strain Rhodotorula sp. Y-23. J. Basic Microbiol. 2018, 58, 331–342. [Google Scholar] [CrossRef]
- Ramle, Z.; Rahim, R.A. Psychrophilic Lipase from Arctic Bacterium. Trop. Life Sci. Res. 2016, 27, 151–157. [Google Scholar] [CrossRef]
- Arifin, A.R.; Kim, S.J.; Yim, J.H.; Suwanto, A.; Kim, H.K. Isolation and biochemical characterization of Bacillus pumilus lipases from the Antarctic. J Microbiol. Biotechnol. 2013, 23, 661–667. [Google Scholar] [CrossRef]
- Florczak, T.; Daroch, M.; Wilkinson, M.C.; Bialkowska, A.; Bates, A.D.; Turkiewicz, M.; Iwanejko, L.A. Purification, characterisation and expression in Saccharomyces cerevisiae of LipG7 an enantioselective, cold-adapted lipase from the Antarctic filamentous fungus Geomyces sp. P7 with unusual thermostability characteristics. Enzyme Microb. Technol. 2013, 53, 18–24. [Google Scholar] [CrossRef]
- Kumar, V.; Sinha, A.K.; Makkar, H.P.; De Boeck, G.; Becker, K. Phytate and phytase in fish nutrition. J. Anim. Physiol. Anim. Nutr. 2012, 96, 335–364. [Google Scholar] [CrossRef]
- Rebello, S.; Jose, L.; Sindhu, R.; Aneesh, E.M. Molecular advancements in the development of thermostable phytases. Appl. Microbiol. Biotechnol. 2017, 101, 2677–2689. [Google Scholar] [CrossRef]
- Park, I.; Cho, J. The phytase from antarctic bacterial isolate, Pseudomonas sp. JPK1 as a potential tool for animal agriculture to reduce manure phosphorus excretion. Afr. J. Agric. Res. 2011, 6, 1398–1406. [Google Scholar]
- Awazu, N.; Shodai, T.; Takakura, H.; Kitagawa, M.; Mukai, H.; Kato, I. Microorganism-Derived Psychrophilic Endonuclease. U.S. Patent 8,034,597, 11 October 2011. [Google Scholar]
- Thallinger, B.; Prasetyo, E.N.; Nyanhongo, G.S.; Guebitz, G.M. Antimicrobial enzymes: An emerging strategy to fight microbes and microbial biofilms. Biotechnol. J. 2013, 8, 97–109. [Google Scholar] [CrossRef]
- See-Too, W.S.; Convey, P.; Pearce, D.A.; Chan, K.G. Characterization of a novel N-acylhomoserine lactonase, AidP, from Antarctic Planococcus sp. Microb. Cell Fact. 2018, 17, 179. [Google Scholar] [CrossRef]
- Medigue, C.; Krin, E.; Pascal, G.; Barbe, V.; Bernsel, A.; Bertin, P.N.; Cheung, F.; Cruveiller, S.; D’Amico, S.; Duilio, A.; et al. Coping with cold: The genome of the versatile marine Antarctica bacterium Pseudoalteromonas haloplanktis TAC125. Genome Res. 2005, 15, 1325–1335. [Google Scholar] [CrossRef]
- Methe, B.A.; Nelson, K.E.; Deming, J.W.; Momen, B.; Melamud, E.; Zhang, X.; Moult, J.; Madupu, R.; Nelson, W.C.; Dodson, R.J.; et al. The psychrophilic lifestyle as revealed by the genome sequence of Colwellia psychrerythraea 34H through genomic and proteomic analyses. Proc. Natl. Acad. Sci. USA 2005, 102, 10913–10918. [Google Scholar] [CrossRef]
- Marizcurrena, J.J.; Morel, M.A.; Brana, V.; Morales, D.; Martinez-Lopez, W.; Castro-Sowinski, S. Searching for novel photolyases in UVC-resistant Antarctic bacteria. Extremophiles 2017, 21, 409–418. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).