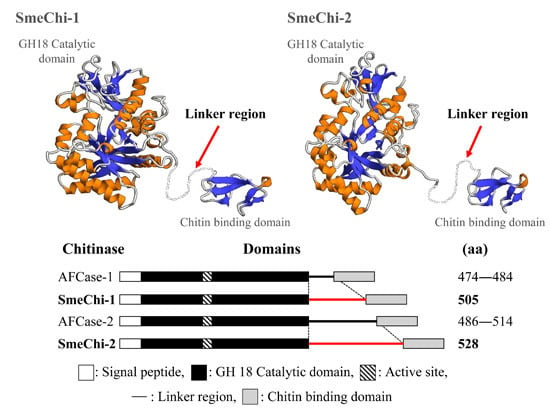
Stomach Chitinase from Japanese Sardine Sardinops melanostictus: Purification, Characterization, and Molecular Cloning of Chitinase Isozymes with a Long Linker
Abstract
:1. Introduction
2. Results and Discussion
2.1. Purification of SmeChiA and SmeChiB
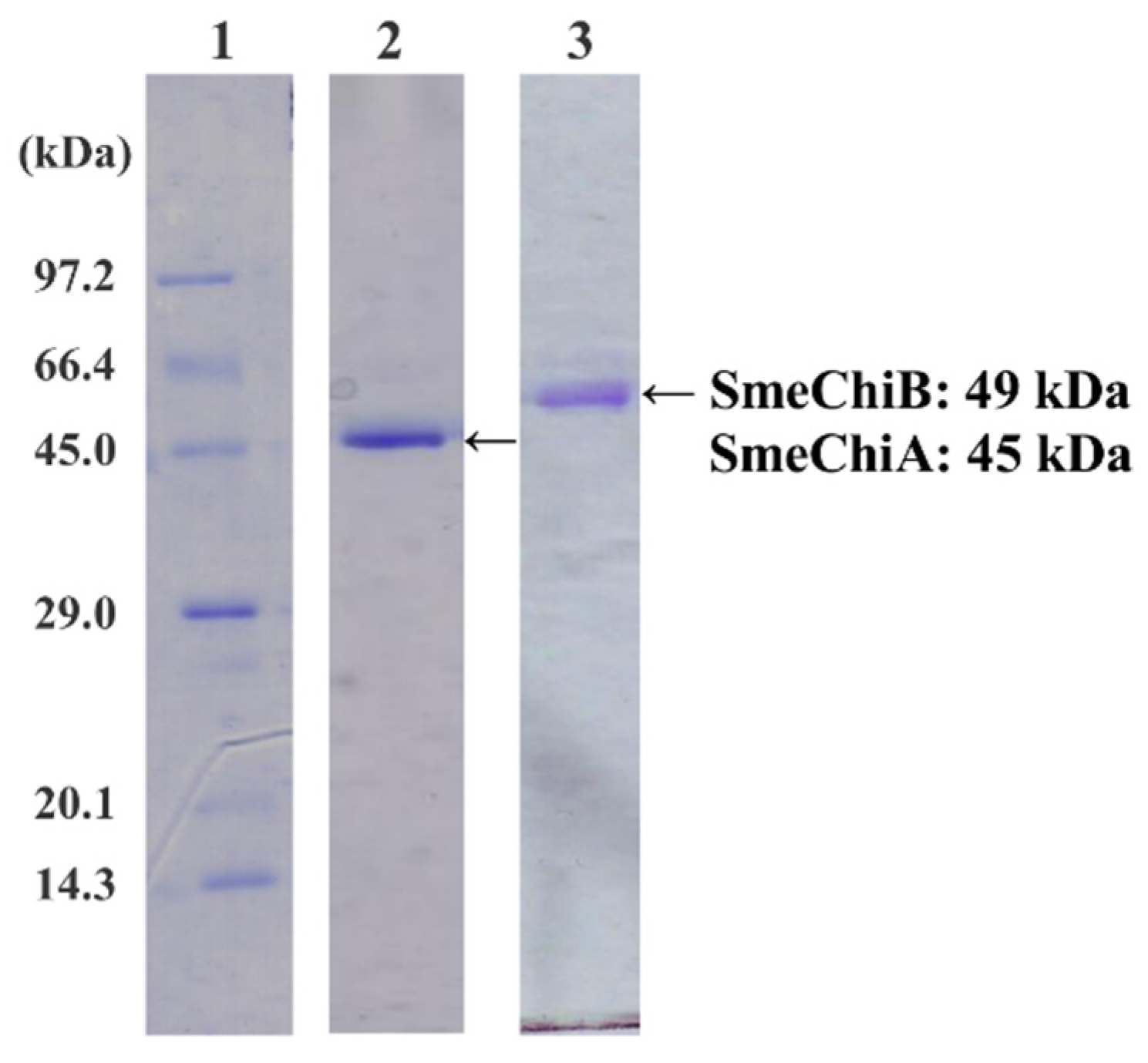
2.2. Effect of pH on SmeChiA and SmeChiB Activities

2.3. Substrate Specificities of SmeChiA and SmeChiB toward pNp-(GlcNAc)n (n = 1–4) and (GlcNAc)5
| Substrate | Specific Activity (U/mg) | |
|---|---|---|
| SmeChiA | SmeChiB | |
| p-nitrophenyl-(GlcNAc) (G-P) | ND | ND |
| p-nitrophenyl-(GlcNAc)2 (G-G-P) | 0.660 | 0.219 |
| p-nitrophenyl-(GlcNAc)3 (G-G-G-P) | 0.081 | 0.877 |
| p-nitrophenyl-(GlcNAc)4 (G-G-G-G-P) | 0.004 | 0.089 |
| Fish | AFCase-1 | AFCase-2 | References | ||
|---|---|---|---|---|---|
| Chitinase | Cleavage Patterns | Chitinase | Cleavage Patterns | ||
| Sardinops melanostictus | SmeChiA (45 kDa) | 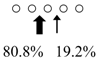 | SmeChiB (49 kDa) | 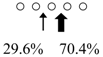 | This study |
| Sebastiscus marmoratus | SmChiA (46 kDa) | 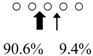 | SmChiC (56 kDa) | 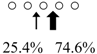 | [17] |
| SmChiB (52 kDa) | 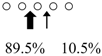 | ||||
| Parapristipoma trilineatum | PtChiA (50 kDa) | 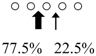 | PtChiB (60 kDa) | 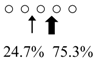 | [16] |
| Pennahia argentata | PaChiA (42 kDa) | 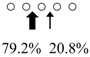 | PaChiB (56 kDa) | 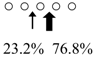 | [14,15] |
2.4. Substrate Specificities of SmeChiA toward Insoluble Substrates
| Substrate | Specific Activity (U/mg) |
|---|---|
| Crab shell α-chitin | 0.922 |
| Shrimp shell α-chitin | 0.303 |
| Squid pen β-chitin | 1.45 |
| Chitin nanofiber | 33.4 |
| Colloidal chitin | 0.922 |
2.5. Molecular Cloning of the Two Chitinase cDNAs

2.6. Tissue Expressions of SmeChi-1 and SmeChi-2
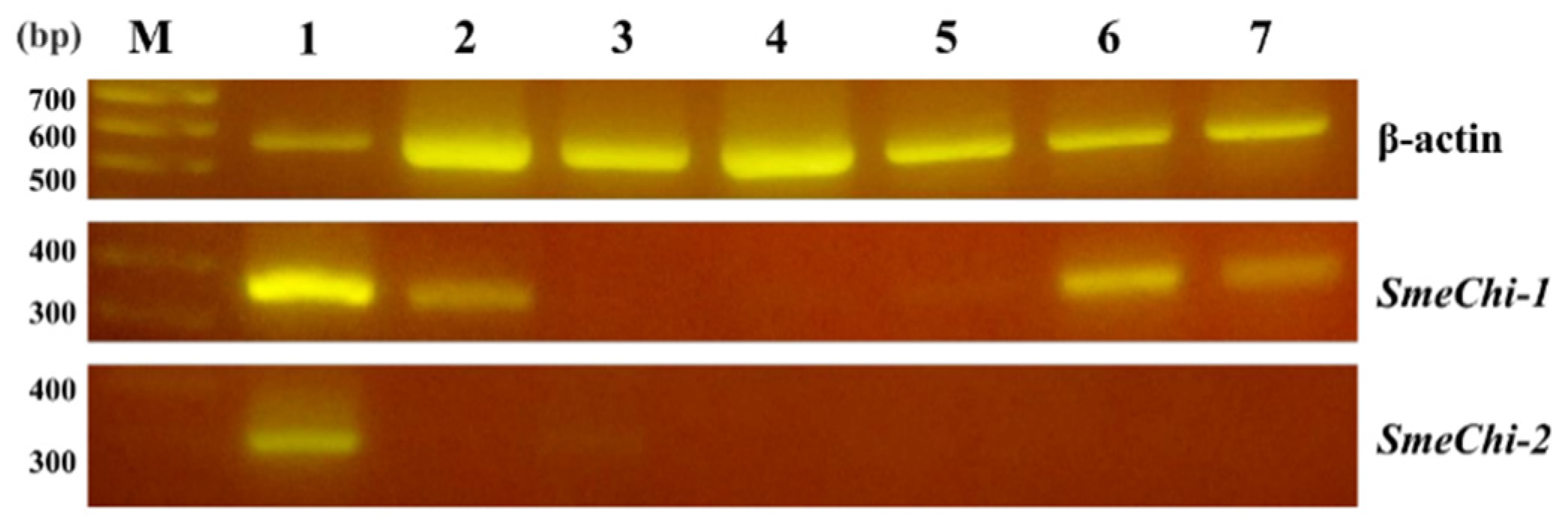
2.7. Phylogenetic Analysis of SmeChi-1 and SmeChi-2
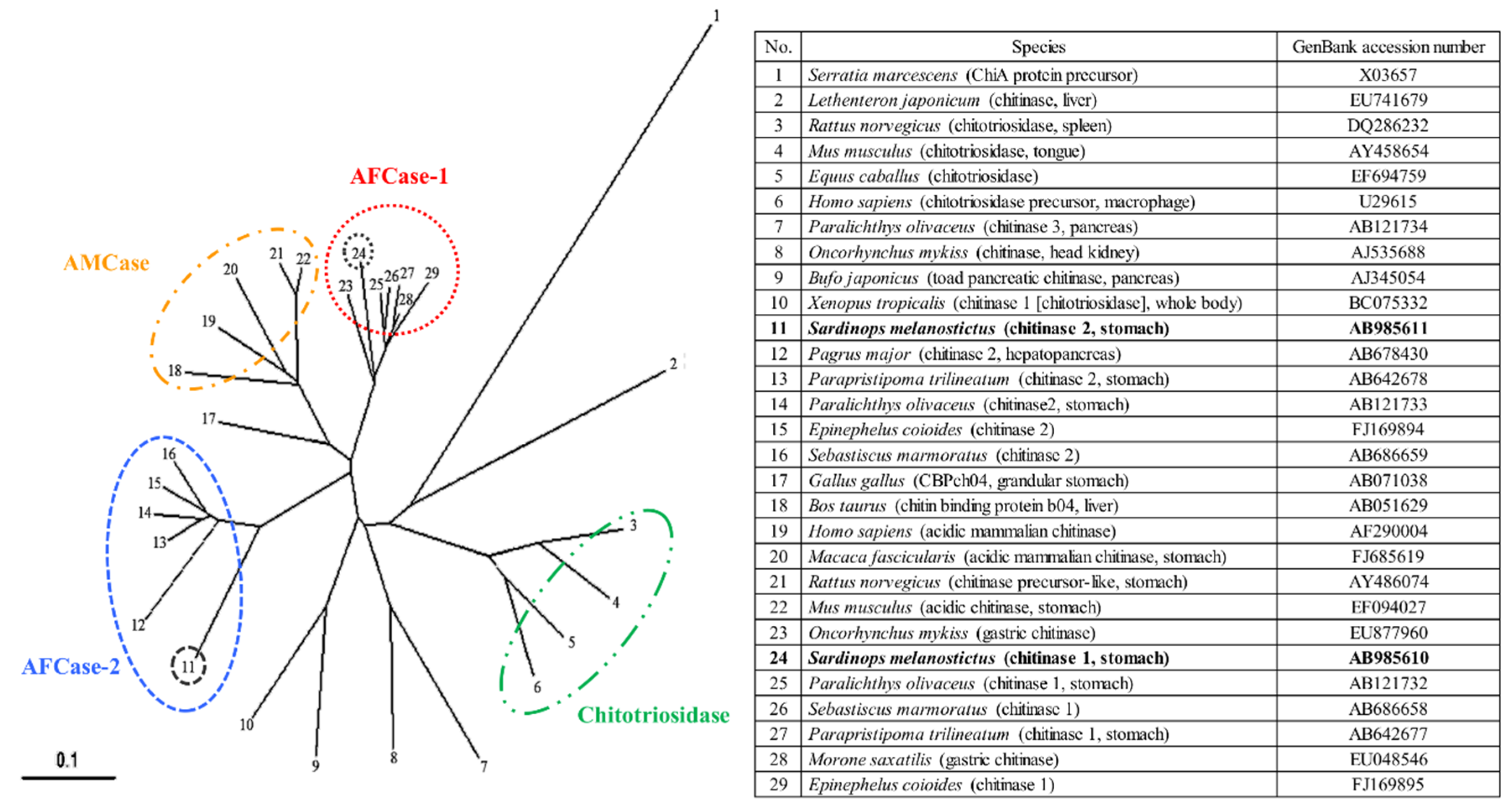
2.8. Prediction of the 3D Structural Models of SmeChi-1 and SmeChi-2
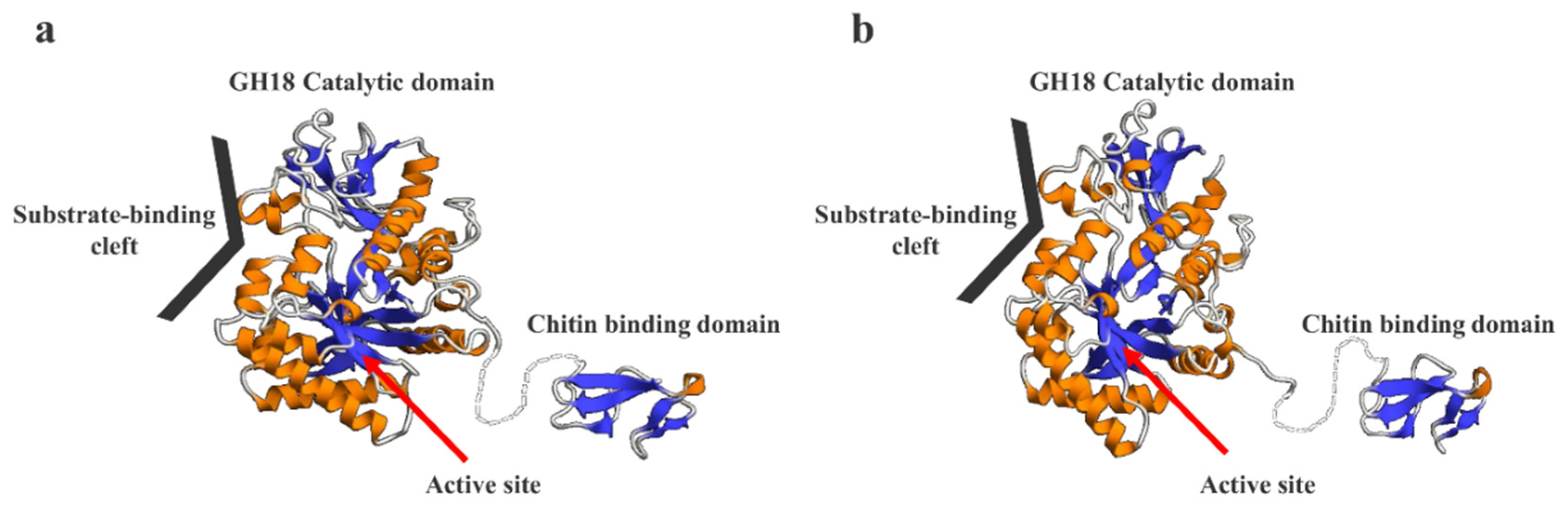
3. Experimental Section
3.1. Chemicals
3.2. Purification of SmeChiA and SmeChiB
3.3. Chitinase Activity Assay
3.4. Effect of pH on Chitinase Activity
3.5. Protein Measurement
3.6. Gel Electrophoresis
3.7. N-Terminal Amino Acid Sequence Analysis
3.8. Cloning of S. melanostictus cDNA
3.9. Nucleotide Sequence Analysis
3.10. Phylogenetic Analysis of SmeChi-1 and SmeChi-2
3.11. Prediction of the 3D Structure Models of SmeChi-1 and SmeChi-2
3.12. Tissue-Specific Gene Expressions of SmeChi-1 and SmeChi-2
4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| (GlcNAc)n | N-acetylchitooligosaccharides |
| GlcNAc | N-acetyl-d-glucosamine |
| pNp-(GlcNAc)n | p-nitrophenyl N-acetylchitooligosaccharides |
| HPLC | high-performance liquid chromatography |
| RACE | rapid amplification of cDNA ends |
| SDS-PAGE | sodium dodecyl sulfate polyacrylamide electrophoresis |
References
- Gooday, G.W. Aggressive and defensive roles for chitinases. In Chitin and Chitinases; Jolles, P., Muzzarelli, R.A.A., Eds.; Birkhäuser Verlag: Basel, Switzerland, 1999; pp. 157–169. [Google Scholar]
- Muzzarellia, R.A.A.; Boudrant, J.; Meyerc, D.; Mannod, N.; DeMarchisd, M.; Paoletti, M.G. Current views on fungal chitin/chitosan, human chitinases, food preservation, glucans, pectins and inulin: A tribute to Henri Braconnot, precursor of the carbohydrate polymers science, on the chitin bicentennial. Carbohydr. Polym. 2012, 87, 995–1012. [Google Scholar] [CrossRef]
- Wang, S.L.; Liang, T.W.; Yen, Y.H. Bioconversion of chitincontaining wastes for the production of enzymes and bioactive materials. Carbohydr. Polym. 2011, 84, 732–742. [Google Scholar] [CrossRef]
- Wang, Z.; Zheng, L.H.; Yang, S.L.; Niu, R.L.; Chu, E.; Lin, X.K. N-acetylchitooligosaccharide is a potent angiogenic inhibitor both in vivo and in vitro. Biochem. Biophy. Res. Commun. 2007, 357, 26–31. [Google Scholar] [CrossRef] [PubMed]
- Matsumiya, M. Enzymatic production of N-acetyl-d-glucosamine using crude enzyme from the liver of squids. Food Sci. Technol. Res. 2004, 10, 296–299. [Google Scholar] [CrossRef]
- Chen, J.K.; Shen, C.R.; Liu, C.L. N-Acetylglucosamine: Production and applications. Mar. Drugs 2010, 8, 2493–2516. [Google Scholar] [CrossRef] [PubMed]
- Henrissat, B.; Bairoch, A. New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem. J. 1993, 293, 781–788. [Google Scholar] [CrossRef] [PubMed]
- Terwisscha van Scheltinga, A.C.; Kalk, K.H.; Beintema, J.J.; Dijkstra, B.W. Crystal structures of hevamine, a plant defence protein with chitinase and lysozyme activity, and its complex with an inhibitor. Structure 1994, 2, 1181–1189. [Google Scholar] [CrossRef]
- Van Aalten, D.M.F.; Synstad, B.; Brurberg, M.B.; Hough, E.; Riise, B.W.; Eijsink, V.G.H.; Wierenga, R.K. Structure of a two-domain chitotriosidase from Serratia marcescens at 1.9-A resolution. Proc. Natl. Acad. Sci. USA 2000, 97, 5842–5847. [Google Scholar] [CrossRef] [PubMed]
- Hart, P.J.; Pfluger, H.D.; Monzingo, A.F.; Hollis, T.; Ropbertus, J.D. The refined crystal structure of an endochitinase from Hordeum vulgare L. seeds at 1.8 A resolution. J. Mol. Biol. 1995, 248, 402–413. [Google Scholar] [CrossRef]
- Monzingo, A.F.; Marcotte, E.M.; Hart, P.J.; Robertus, J.D. Chitinases, chitosanases, and lysozymes can be divided into procaryotic and eucaryotic families sharing a conserved core. Nat. Struct. Biol. 1996, 3, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Matsumiya, M.; Mochizuki, A. Purification and characterization of chitinase from the stomach of common mackerel Scomber japonicas. Bull. Coll. Agric. Vet. Med. Nihon Univ. 1995, 52, 131–136. [Google Scholar]
- Matsumiya, M.; Arakane, Y.; Haga, A.; Muthukrishnan, S.; Kramer, K.J. Substrate specificity of chitinases from two species of fish, greenling, Hexagrammos otakii, and common mackerel, Scomber japonicus, and the insect, tobacco hornworm, Manduca sexta. Biosci. Biotechnol. Biochem. 2006, 70, 971–979. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Miyauchi, K.; Mochizuki, A.; Matsumiya, M. Purification and characterization of chitinase from the stomach of silver croaker Pennahia argentatus. Protein Expr. Purif. 2009, 65, 214–222. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Miyauchi, K.; Matsumiya, M. Purification and characterization of a 56 kDa chitinase isozyme (PaChiB) from the stomach of silver croaker Pennahia argentatus. Biosci. Biotechnol. Biochem. 2012, 76, 971–979. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Kondo, Y.; Matsumiya, M. Purification, characterization, and molecular cloning of chitinases from the stomach of the threeline grunt Parapristipoma trilineatum. Process. Biochem. 2013, 48, 1324–1334. [Google Scholar] [CrossRef]
- Ikeda, M.; Shirase, D.; Sato, T.; Ueda, M.; Hirabayashi, S.; Matsumiya, M. Primary structure and enzymatic properties of chitinase isozymes purified from the stomach of the marbled rockfish Sebastiscus marmoratus. J. Chitin Chitosan Sci. 2014, 2, 106–116. [Google Scholar] [CrossRef]
- Matsumiya, M.; Miyauchi, K.; Mochizuki, A. Characterization of 38 kDa and 42 kDa chitinase isozymes from the liver of Japanese squid Todarodes pacificus. Fish. Sci. 2002, 68, 603–609. [Google Scholar] [CrossRef]
- Nishino, R.; Suyama, A.; Ikeda, M.; Kakizaki, H.; Matsumiya, M. Purification and characterization of a liver chitinase from golden cuttlefish, Sepia esculenta. J. Chitin Chitosan Sci. 2014, 2, 238–243. [Google Scholar] [CrossRef]
- Ogino, T.; Tabata, H.; Ikeda, M.; Kakizaki, H.; Matsumiya, M. Purification of a chitinase from the posterior salivary gland of common octopus Octopus. vulgaris and its properties. J. Chitin Chitosan Sci. 2014, 2, 135–142. [Google Scholar] [CrossRef]
- Kurokawa, T.; Uji, S.; Suzuki, T. Molecular cloning of multiple chitinase genes in Japanese flounder, Paralichthys olivaceus. Comp. Biochem. Physiol. B 2004, 138, 255–264. [Google Scholar] [CrossRef] [PubMed]
- Kakizaki, H.; Ikeda, M.; Fukushima, H.; Matsumiya, M. Distribution of chitinolytic enzymes in the organs and cDNA cloning of chitinase isozymes from the stomach of two species of fish, chub mackerel (Scomber japonicus) and silver croaker (Pennahia argentata). Open J. Mar. Sci. 2015, 5, 398–411. [Google Scholar] [CrossRef]
- Suzuki, T.; Kakizaki, H.; Ikeda, M.; Matsumiya, M. Molecular cloning of a novel chitinase gene from blue shark (Prionace glauca; Chondrichthyes) stomach. J. Chitin Chitosan Sci. 2014, 2, 143–148. [Google Scholar] [CrossRef]
- Fujitani, N.; Hasegawa, H.; Kakizaki, H.; Ikeda, M.; Matsumiya, M. Molecular cloning of multiple chitinase genes in swimming crab Portunus trituberculatus. J. Chitin Chitosan Sci. 2014, 2, 149–156. [Google Scholar] [CrossRef]
- Huang, Q.S.; Yan, J.H.; Tang, J.Y.; Tao, Y.M.; Xie, X.L.; Wang, Y.; Wei, X.Q.; Yan, Q.H.; Chen, Q.X. Cloning and tissue expressions of seven chitinase family genes in Litopenaeus vannamei. Fish Shellfish Immunol. 2010, 29, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Proespraiwong, P.; Tassanakajon, A.; Rimphanitchayakit, V. Chitinases from the black tiger shrimp Penaeus monodon: Phylogenetics, expression and activities. Comp. Biochem. Physiol. B 2010, 156, 86–96. [Google Scholar] [CrossRef] [PubMed]
- Kakizaki, H.; Hamaguchi, K.; Ikeda, M.; Matsumiya, M. Cloning of a novel chitinase cDNA from the stomach of the Coelacanth Latimeria chalumnae (sarcopterygii). J. Chitin Chitosan Sci. 2014, 2, 123–129. [Google Scholar] [CrossRef]
- Ifuku, S.; Nogi, M.; Abe, K.; Yoshioka, M.; Morimoto, M.; Saimoto, H.; Yano, H. Preparation of chitin nanofibers with a uniform width as α-chitin from crab shells. Biomacromolecules 2009, 10, 1584–1588. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.H.; Cai, G.P. Molecular cloning and characterization of rat chitotriosidase. DNA Seq. 2008, 19, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Ohno, M.; Togashi, Y.; Tsuda, K.; Okawa, K.; Kamaya, M.; Sakaguchi, M.; Sugahara, Y.; Oyama, F. Quantification of Chitinase mRNA Levels in Human and Mouse Tissues by Real-Time PCR: Species-Specific Expression of Acidic Mammalian Chitinase in Stomach Tissues. PLoS ONE 2013, 8, e67399. [Google Scholar]
- Miwa, S. Stomach. In Gyoruiseirigakunokiso; Aida, K., Ed.; Kouseisyakouseikaku: Tokyo, Japan, 2002; pp. 194–195. [Google Scholar]
- Matsumoto, T.; Nonaka, T.; Hashimoto, M.; Watanabe, T.; Mitsui, Y. Three-dimensional structure of the catalytic domain of chitinase Al from Bacillus circulars WL-12 at a very high resolution. Proc. Jpn. Acad. Ser. B 1999, 75, 269–274. [Google Scholar] [CrossRef]
- Watanabe, T; Ariga, Y.; Sato, U.; Toratani, T.; Hashimoto, M.; Nikaidou, N.; Kezuka, Y.; Nonaka, T.; Sugiyama, J. Aromatic residues within the substrate-binding cleft of Bacillus circulans chitinase A1 are essential for hydrolysis of crystalline chitin. Biochem. J. 2003, 376, 237–244. [Google Scholar]
- Shimahara, K.; Takiguchi, Y. Preparation of crustacean chitin. In Method in Enzymology; Wood, W.A., Kellogg, S., Eds.; Academic Press: New York, NY, USA, 1988; pp. 417–423. [Google Scholar]
- Ohtakara, A. Chitinase and β-N-acetylhexosaminidase from Pycnoporus cinnabarinus. In Method in Enzymology; Wood, W.A., Kellogg, S.T., Eds.; Academic Press: New York, NY, USA, 1988; pp. 462–470. [Google Scholar]
- Koga, D.; Yoshioka, T.; Arakane, Y. HPLC analysis of anomeric formation and cleavage pattern by chitinolytic enzyme. Biosci. Biotechnol. Biochem. 1998, 62, 1643–1646. [Google Scholar] [CrossRef]
- Bradford, M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kawashima, S.; Ikehata, H.; Tada, C.; Ogino, T.; Kakizaki, H.; Ikeda, M.; Fukushima, H.; Matsumiya, M. Stomach Chitinase from Japanese Sardine Sardinops melanostictus: Purification, Characterization, and Molecular Cloning of Chitinase Isozymes with a Long Linker. Mar. Drugs 2016, 14, 22. https://doi.org/10.3390/md14010022
Kawashima S, Ikehata H, Tada C, Ogino T, Kakizaki H, Ikeda M, Fukushima H, Matsumiya M. Stomach Chitinase from Japanese Sardine Sardinops melanostictus: Purification, Characterization, and Molecular Cloning of Chitinase Isozymes with a Long Linker. Marine Drugs. 2016; 14(1):22. https://doi.org/10.3390/md14010022
Chicago/Turabian StyleKawashima, Satoshi, Hiroki Ikehata, Chihiro Tada, Tomohiro Ogino, Hiromi Kakizaki, Mana Ikeda, Hideto Fukushima, and Masahiro Matsumiya. 2016. "Stomach Chitinase from Japanese Sardine Sardinops melanostictus: Purification, Characterization, and Molecular Cloning of Chitinase Isozymes with a Long Linker" Marine Drugs 14, no. 1: 22. https://doi.org/10.3390/md14010022
APA StyleKawashima, S., Ikehata, H., Tada, C., Ogino, T., Kakizaki, H., Ikeda, M., Fukushima, H., & Matsumiya, M. (2016). Stomach Chitinase from Japanese Sardine Sardinops melanostictus: Purification, Characterization, and Molecular Cloning of Chitinase Isozymes with a Long Linker. Marine Drugs, 14(1), 22. https://doi.org/10.3390/md14010022