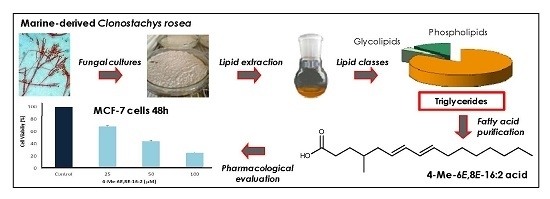
The Marine-Derived Fungus Clonostachys rosea, Source of a Rare Conjugated 4-Me-6E,8E-hexadecadienoic Acid Reducing Viability of MCF-7 Breast Cancer Cells and Gene Expression of Lipogenic Enzymes
Abstract
:1. Introduction
2. Results and Discussion
2.1. Lipid and Fatty Acid Composition of C. rosea—OSMAC Approach
| Culture Medium | Lipid Content (%, dw) | Fatty Acids (Mean ± SD) (% Total FA) | ||||
|---|---|---|---|---|---|---|
| 16:0 | 4-Me-6,8-16:2 | 18:2 and 18:1 | 18:0 | Others FA * | ||
| PDA | 12.9 ± 0.3 | 16.1 ± 0.9 | 8.0 ± 3.0 | 55.0 ± 4.0 | 10.0 ± 2.0 | 12.0 ± 7.0 |
| MES | 29.7 ± 0.4 | 20.0 ± 5.0 | 9.0 ± 2.0 | 52.5 ± 13.5 | 8.0 ± 3.0 | 10.0 ± 5.0 |
| CYA | 8.7 ± 0.8 | 15.7 ± 0.5 | 14.0 ± 1.0 | 60.0 ± 17.0 | 6.0 ± 0.1 | 4.0 ± 3.0 |
| YES | 30.7 ± 0.4 | 14.0 ± 2.0 | 11.0 ± 1.0 | 64.0 ± 6.0 | 6.0 ± 1.0 | 4.0 ± 2.0 |
| DCA | 14.3 ± 0.4 | 16.0 ± 1.0 | 23.0 ± 0.8 | 44.6 ± 4.7 | 5.0 ± 1.0 | 10.0 ± 6.0 |
| MEA | 14.9 ± 0.2 | 17.5 ± 0.4 | 8.0 ± 2.0 | 57.0 ± 2.0 | 9.0 ± 0.5 | 9.0 ± 6.0 |
| Lipid production | Main FA Composition (% total FA) | |||
|---|---|---|---|---|
| Lipid Classes | % of Total Lipids | Saturated FA * | Unsaturated FA * | 4-Me-6,8-16:2 |
| Triglycerides | 84 ± 7 | 28 ± 2 | 58 ± 7 | 20 ± 4 |
| Glycolipids | 4 ± 2 | 46 ± 3 | 44 ± 2 | 5 ± 1 |
| Phospholipids | 12 ± 6 | 29 ± 3 | 64 ± 6 | 3 ± 1 |
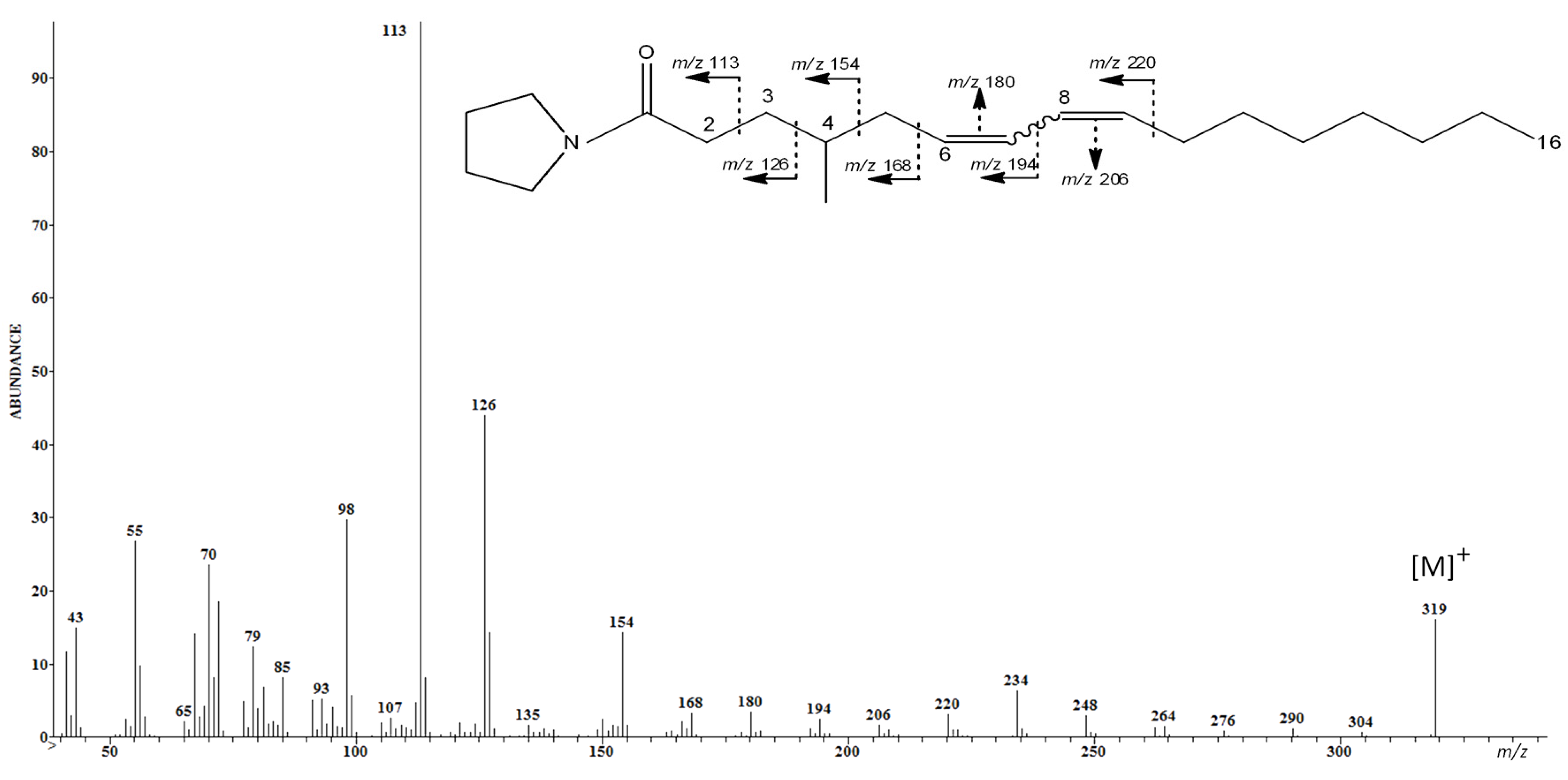
2.2. Characterization of 4-Me-6-8-Hexadecadienoic Acid Structure (Isomer C)
| Positions | δC | δH (Integral, Mult., J = Hz) | |
|---|---|---|---|
| 1 | 174.4 | - | - |
| 2 | 31.9 | 1.33 | (1H, m) |
| 2.33 | (1H, m) | ||
| 3 | 31.8 | 1.33 | (1H, m) |
| 2.33 | (1H, m) | ||
| 4 | 32.9 | 1.48 | (1H, m) |
| 5 | 39.8 | 1.99 | (1H, dt, 7.2, 6.8) |
| 2.05 | (1H, dt, 7.2, 6.8) | ||
| 6 | 129.8 | 5.54 | (1H, m) |
| 7 | 130.1 | 5.99 | (1H, m) |
| 8 | 132.03 | 5.99 | (1H, m) |
| 9 | 132.9 | 5.54 | (1H, m) |
| 10 | 32.6 | 2.05 | (2H, m) |
| 11 | 29.2 | 1.44 | (2H, m) |
| 12 | 31.4 | 1.33 | (1H, m) |
| 1.70 | (1H, m) | ||
| 13 | 29.4 | 1.33 | (2H, m) |
| 14 | 29.1 | 1.33 | (2H, m) |
| 15 | 22.6 | 1.33 | (2H, m) |
| 16 | 14.1 | 0.86 | (3H, t, 6.8) |
| 17 | 19.1 | 0.88 | (3H, d, 6.4) |
| –OCH3 | 51.5 | 3.66 | (3H, s) |

2.3. 4-Me-6E,8E-Hexadecadienoic Acid Reduces Viability of Human MCF-7 Breast Cancer Cells
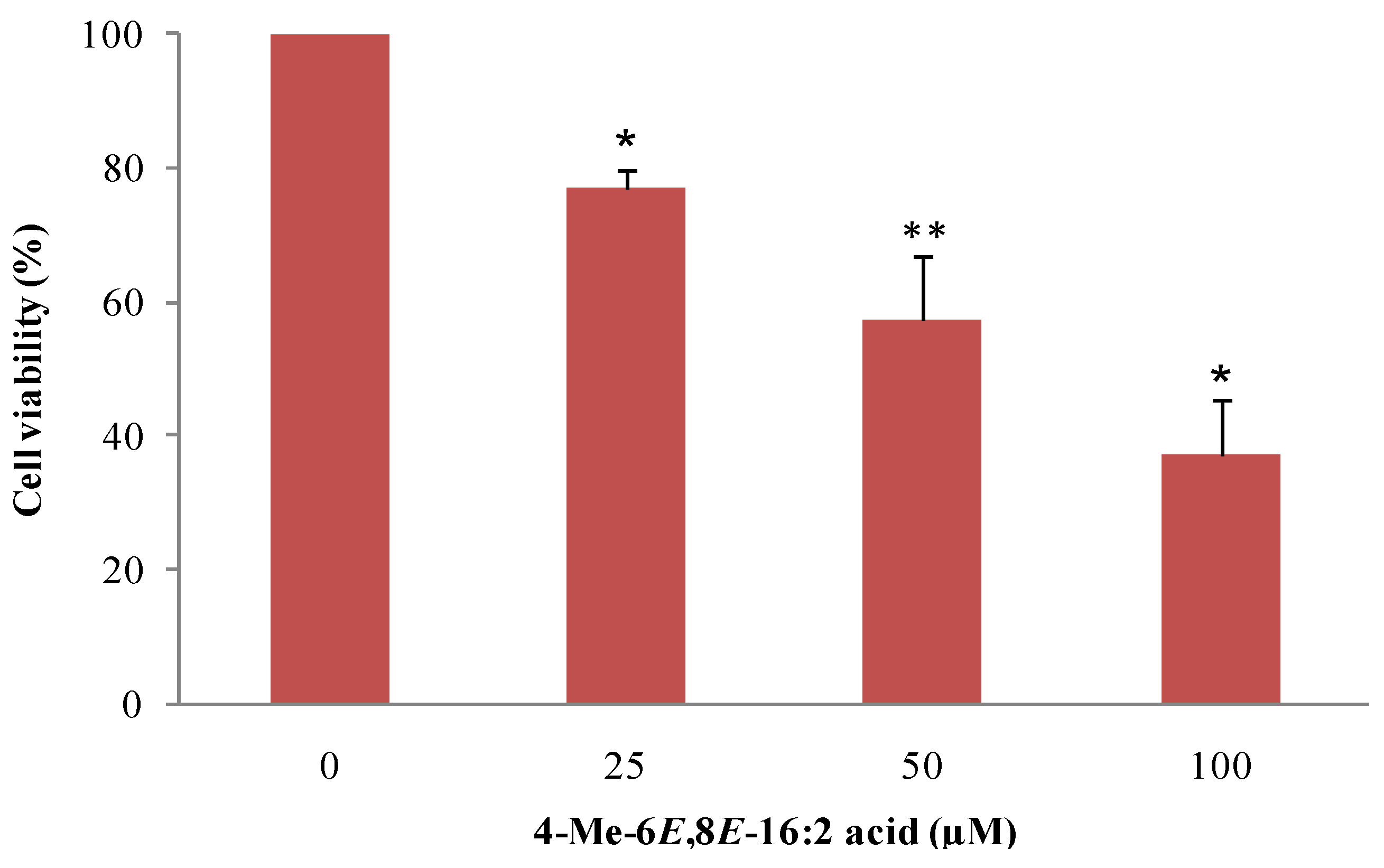
2.4. 4-Me-6E,8E-Hexadecadienoic Acid Reduces Gene Expression of Lipogenic Enzymes
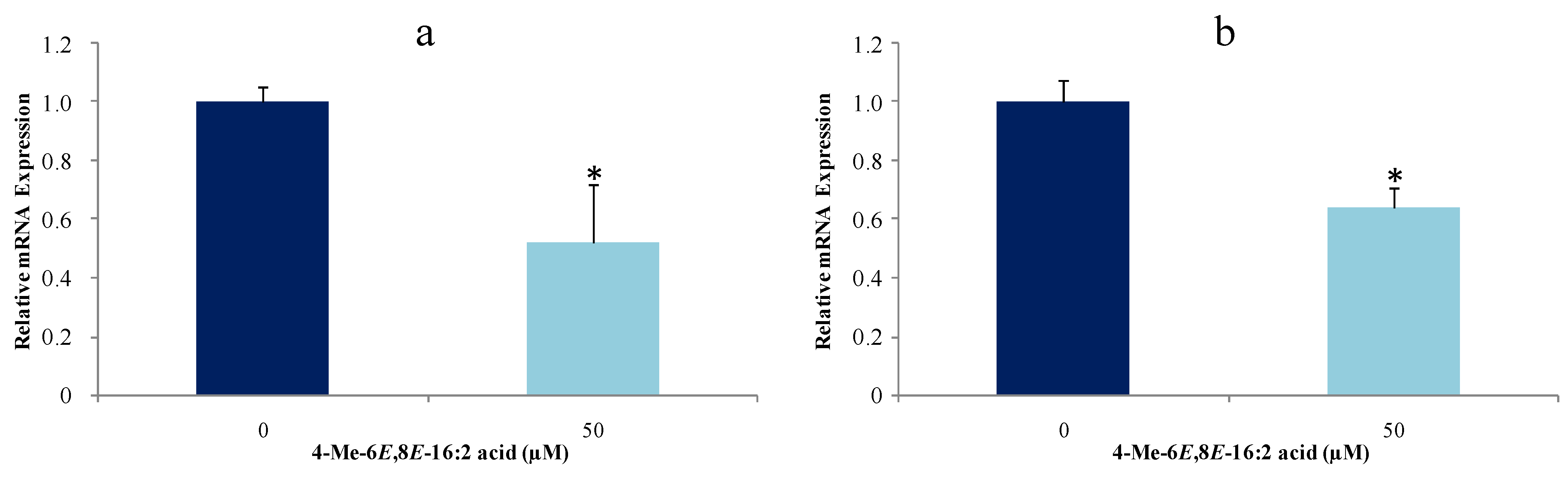
3. Experimental Section
3.1. Clonostachys rosea Strain MMS1090
3.2. Culture Media for OSMAC Approach
3.3. Lipid Extraction and Separation of Lipid Classes
3.4. Preparation of Fatty Acid Methyl Esters and N-Acyl Pyrrolidides
3.5. Gas Chromatography-Mass Spectrometry Analysis
3.6. Production, Purification and Structural Elucidation of 4-Me-6,8-16:2 Fatty Acid
3.7. Cell Viability Assay
3.8. Gene Expression of Lipogenic Enzymes
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| CFA | Conjugated fatty acid |
| FA | Fatty acid |
| FAME | Fatty acid methyl ester |
| GC-MS | Gas chromatography coupled to mass spectrometry |
| NAP | N-acyl pyrrolidide |
| OSMAC | One strain many compounds |
| PUFA | Polyunsaturated fatty acid |
| TL | Total lipids |
| TG | Triglycerides |
| ACC | Acetyl CoA carboxylase |
| FAS | Fatty acid synthase |
References
- Li, Q.; Wang, M.Y. Use food industry waste to produce microbial oil. Sci. Technol. Food Ind. 1997, 6, 65–69. [Google Scholar]
- Li, Q.; Du, W.; Liu, D. Perspectives of microbial oils for biodiesel production. Appl. Microbiol. Biotechnol. 2008, 80, 749–756. [Google Scholar] [CrossRef] [PubMed]
- Hassan, M.; Blanc, P.J.; Granger, L.M.; Pareilleux, A.; Goma, G. Influence of nitrogen and iron limitations on lipid production by Cryptococcus curvatus grown in batch and fed-batch culture. Process Biochem. 1996, 31, 355–361. [Google Scholar] [CrossRef]
- Stahl, P.D.; Klug, M.J. Characterization and differentiation of filamentous fungi based on fatty acid composition. Appl. Environ. Microbiol. 1996, 62, 4136–4146. [Google Scholar] [PubMed]
- Lösel, D.M. Fungal lipids. In Microbial Lipids; Ratledge, C., Wilkinson, S.G., Eds.; Academic Press: London, UK, 1988; Volume 1, pp. 699–806. [Google Scholar]
- Brennan, P.J.; Griffin, P.F.S.; Lösel, D.M.; Tyrrell, D. The lipids of fungi. Prog. Chem. Fats Other Lipids 1975, 14, 49–89. [Google Scholar] [CrossRef]
- Pohl, C.H.; Botha, A.; Kock, J.L.F.; Coetzee, D.J.; Botes, P.J. The production of γ-linolenic acid by selected members of the Dikaryomycota grown on different carbon sources. Antonie Leeuwenhoek 1997, 72, 191–199. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, N.; Dubois, N.; Wielgosz-Collin, G.; du Pont, T.R.; Bergé, J.P.; Pouchus, Y.F.; Barnathan, G. Lipid content and fatty acid composition of a marine-derived Trichoderma longibrachiatum strain cultured by agar surface and submerged fermentations. Process Biochem. 2007, 42, 676–680. [Google Scholar] [CrossRef]
- Oleinikova, G.K.; Zhuravleva, O.I.; Yurchenko, A.N.; Sobolevskaya, M.P.; Kirichuk, N.N.; Afiyatullov, S.S. Non-polar compounds and free fatty acids from several marine isolates of fungi of the genus Aspergillus. Chem. Nat. Compd. 2013, 48, 1065–1066. [Google Scholar] [CrossRef]
- Oleinikova, G.K.; Smetanina, O.F.; Khudyakova, Y.V.; Kirichuk, N.N.; Afiyatullov, S.S. Non-polar compounds and free fatty acids from marine isolates of mycelial fungi. Chem. Nat. Compd. 2013, 49, 499–500. [Google Scholar] [CrossRef]
- Devi, P.; Shridhar, M.P.D.; DeSouza, L.; Naik, C.G. Cellular fatty acid composition of marine-derived fungi. Indian J. Geo-Mar. Sci. 2006, 35, 359–363. [Google Scholar]
- Schroers, H.J.; Samuels, G.J.; Seifert, K.A.; Gams, W. Classification of the mycoparasite Gliocladium roseum in Clonostachys as C. rosea, its relationship to Bionectria ochroleuca, and notes on other Gliocladium-like fungi. Mycologia 1999, 91, 365. [Google Scholar] [CrossRef]
- Stadler, M.; Schulz, B. High energy biofuel from endophytic fungi? Trends Plant Sci. 2009, 14, 353–355. [Google Scholar] [CrossRef] [PubMed]
- Strobel, G.A.; Knighton, B.; Kluck, K.; Ren, Y.; Livinghouse, T.; Griffin, M.; Spakowicz, D.; Sears, J. The production of myco-diesel hydrocarbons and their derivatives by the endophytic fungus Gliocladium roseum (NRRL 50072). Microbiology 2008, 154, 3319–3328. [Google Scholar] [CrossRef] [PubMed]
- Bode, H.B.; Bethe, B.; Höfs, R.; Zeeck, A. Big effects from small changes: Possible ways to explore nature’s chemical diversity. ChemBioChem 2002, 3, 619–627. [Google Scholar] [CrossRef]
- El Roz, A.; Bard, J.M.; Huvelin, J.M.; Nazih, H. The anti-proliferative and pro-apoptotic effects of the trans9,trans11 conjugated linoleic acid isomer on MCF-7 breast cancer cells are associated with LXR activation. Prostaglandins Leukot. Essent. Fatty Acids 2013, 88, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, A.; Wu, J.; Barbour, S.; Fang, X. Lysophosphatidic acid activates lipogenic pathways and de novo lipid synthesis in ovarian cancer cells. J. Biol. Chem. 2012, 287, 24990–25000. [Google Scholar] [CrossRef] [PubMed]
- Ratledge, C. Regulation of lipid accumulation in oleaginous micro-organisms. Biochem. Soc. Trans. 2002, 30, 1047–1050. [Google Scholar] [CrossRef] [PubMed]
- Economou, C.N.; Aggelis, G.; Pavlou, S.; Vayenas, D.V. Single cell oil production from rice hulls hydrolysate. Bioresour. Technol. 2011, 102, 9737–9742. [Google Scholar] [CrossRef] [PubMed]
- Ratledge, C. Fatty acid biosynthesis in microorganisms being used for single cell oil production. Biochimie 2004, 86, 807–815. [Google Scholar] [CrossRef] [PubMed]
- Weete, J.D. Fungal lipids. In Lipid Biochemistry of Fungi and Other Organisms; Springer: New York, NY, USA, 1980; pp. 9–48. [Google Scholar]
- Goodridge, A.G. Dietary regulation of gene expression: Enzymes involved in carbohydrate and lipid metabolism. Annu. Rev. Nutr. 1987, 7, 157–185. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, J.; Kishino, S.; Ando, A.; Sugimoto, S.; Mihara, K.; Shimizu, S. Production of conjugated fatty acids by lactic acid bacteria. J. Biosci. Bioeng. 2005, 100, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Kishino, S.; Ogawa, J.; Yokozeki, K.; Shimizu, S. Microbial production of conjugated fatty acids. Lipid Technol. 2009, 21, 177–181. [Google Scholar] [CrossRef]
- Kurnia, D.; Akiyama, K.; Hayashi, H. 10-Phenyl-[11]-cytochalasans from indonesian mushroom Microporellus subsessilis. Phytochemistry 2007, 68, 697–702. [Google Scholar] [CrossRef] [PubMed]
- Nagao, K.; Yanagita, T. Conjugated fatty acids in food and their health benefits. J. Biosci. Bioeng. 2005, 100, 152–157. [Google Scholar] [CrossRef] [PubMed]
- Martin, J.C.; Valeille, K. Conjugated linoleic acids: All the same or to everyone its own function? Reprod. Nutr. Dev. 2002, 42, 525–536. [Google Scholar] [CrossRef] [PubMed]
- Yuan, G.F.; Chen, X.E.; Li, D. Conjugated linolenic acids and their bioactivities: A review. Food Funct. 2014, 5, 1360–1368. [Google Scholar] [CrossRef] [PubMed]
- Stachowska, E. Conjugated dienes of linoleic acid and tumorigenesis. Ann. Acad. Med. Stetin. 2007, 54, 122–125. [Google Scholar]
- Ip, C.; Singh, M.; Thompson, H.J.; Scimeca, J.A. Conjugated linoleic acid suppresses mammary carcinogenesis and proliferative activity of the mammary gland in the rat. Cancer Res. 1994, 54, 1212–1215. [Google Scholar] [PubMed]
- Kabara, J.J.; Swieczkowski, D.M.; Conley, A.J.; Truant, J.P. Fatty acids and derivatives as antimicrobial agents. Antimicrob. Agents Chemother. 1972, 2, 23–28. [Google Scholar] [CrossRef] [PubMed]
- Ip, C.; Dong, Y.; Ip, M.M.; Banni, S.; Carta, G.; Angioni, E.; Murru, E.; Spada, S.; Melis, M.P.; Saebo, A. Conjugated linoleic acid isomers and mammary cancer prevention. Nutr. Cancer 2002, 43, 52–58. [Google Scholar] [CrossRef] [PubMed]
- Moutsioulis, A.A.; Rule, D.C.; Murrieta, C.M.; Bauman, D.E.; Lock, A.L.; Barbano, D.M.; Carey, G.B. Human breast milk enrichment in conjugated linoleic acid after consumption of a conjugated linoleic acid-rich food product: A pilot study. Nutr. Res. 2008, 28, 437–442. [Google Scholar] [CrossRef] [PubMed]
- Wongtangtintharn, S.; Oku, H.; Iwasaki, H.; Toda, T. Effect of branched-chain fatty acids on fatty acid biosynthesis of human breast cancer cells. J. Nutr. Sci. Vitaminol. (Tokyo) 2004, 50, 137–143. [Google Scholar] [CrossRef] [PubMed]
- Shultz, T.D.; Chew, B.P.; Seaman, W.R.; Luedecke, L.O. Inhibitory effect of conjugated dienoic derivatives of linoleic acid and β-carotene on the in vitro growth of human cancer cells. Cancer Lett. 1992, 63, 125–133. [Google Scholar] [CrossRef]
- Wang, L.S.; Huang, Y.W.; Liu, S.; Yan, P.; Lin, Y.C. Conjugated linoleic acid induces apoptosis through estrogen receptor alpha in human breast tissue. BMC Cancer 2008, 8, 208. [Google Scholar] [CrossRef] [PubMed]
- Petridou, A.; Mougios, V.; Sagredos, A. Supplementation with CLA: Isomer incorporation into serum lipids and effect on body fat of women. Lipids 2003, 38, 805–811. [Google Scholar] [CrossRef] [PubMed]
- Song, H.J.; Sneddon, A.A.; Heys, S.D.; Wahle, K.W.J. Regulation of fatty acid synthase (FAS) and apoptosis in estrogen-receptor positive and negative breast cancer cells by conjugated linoleic acids. Prostaglandins Leukot. Essent. Fatty Acids 2012, 87, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Wiggins, A.K.A.; Mason, J.K.; Thompson, L.U. Growth and gene expression differ over time in alpha-linolenic acid treated breast cancer cells. Exp. Cell Res. 2015, 333, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Swinnen, J.V.; Brusselmans, K.; Verhoeven, G. Increased lipogenesis in cancer cells: New players, novel targets. Curr. Opin. Clin. Nutr. Metab. Care 2006, 9, 358–365. [Google Scholar] [CrossRef] [PubMed]
- Menendez, J.A.; Lupu, R. Fatty acid synthase and the lipogenic phenotype in cancer pathogenesis. Nat. Rev. Cancer 2007, 7, 763–777. [Google Scholar] [CrossRef] [PubMed]
- Santos, C.R.; Schulze, A. Lipid metabolism in cancer. FEBS J. 2012, 279, 2610–2623. [Google Scholar] [CrossRef] [PubMed]
- Kuhajda, F.P. Fatty acid synthase and cancer: New application of an old pathway. Cancer Res. 2006, 66, 5977–5980. [Google Scholar] [CrossRef] [PubMed]
- Currie, E.; Schulze, A.; Zechner, R.; Walther, T.C.; Farese, R.V., Jr. Cellular fatty acid metabolism and cancer. Cell Metab. 2013, 18, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Rajput, S.; Watabe, K.; Liao, D.F.; Cao, D. Acetyl-CoA carboxylase-a as a novel target for cancer therapy. Front. Biosci. Sch. Ed. 2010, 2, 515–526. [Google Scholar] [CrossRef]
- Gansler, T.S.; Hardman, W., III; Hunt, D.A.; Schaffel, S.; Hennigar, R.A. Increased expression of fatty acid synthase (OA-519) in ovarian neoplasms predicts shorter survival. Hum. Pathol. 1997, 28, 686–692. [Google Scholar] [CrossRef]
- Menendez, J.A.; Ropero, S.; Mehmi, I.; Atlas, E.; Colomer, R.; Lupu, R. Overexpression and hyperactivity of breast cancer-associated fatty acid synthase (oncogenic antigen-519) is insensitive to normal arachidonic fatty acid-induced suppression in lipogenic tissues but it is selectively inhibited by tumoricidal alpha-linolenic and gamma-linolenic fatty acids: A novel mechanism by which dietary fat can alter mammary tumorigenesis. Int. J. Oncol. 2004, 24, 1369–1383. [Google Scholar] [PubMed]
- Oku, H.; Wongtangtintharn, S.; Iwasaki, H.; Toda, T. Conjugated linoleic acid (CLA) inhibits fatty acid synthetase activity in vitro. Biosci. Biotechnol. Biochem. 2003, 67, 1584–1586. [Google Scholar] [CrossRef] [PubMed]
- Chujo, H.; Yamasaki, M.; Nou, S.; Koyanagi, N.; Tachibana, H.; Yamada, K. Effect of conjugated linoleic acid isomers on growth factor-induced proliferation of human breast cancer cells. Cancer Lett. 2003, 202, 81–87. [Google Scholar] [CrossRef]
- Bocca, C.; Bozzo, F.; Francica, S.; Colombatto, S.; Miglietta, A. Involvement of PPARγ and E-cadherin/β-catenin pathway in the antiproliferative effect of conjugated linoleic acid in MCF-7 cells. Int. J. Cancer 2007, 121, 248–256. [Google Scholar] [CrossRef] [PubMed]
- Thupari, J.N.; Pinn, M.L.; Kuhajda, F.P. Fatty acid synthase inhibition in human breast cancer cells leads to malonyl-CoA-induced inhibition of fatty acid oxidation and cytotoxicity. Biochem. Biophys. Res. Commun. 2001, 285, 217–223. [Google Scholar] [CrossRef] [PubMed]
- Swinnen, J.V.; Vanderhoydonc, F.; Elgamal, A.A.; Eelen, M.; Vercaeren, I.; Joniau, S.; van Poppel, H.; Baert, L.; Goossens, K.; Heyns, W.; et al. Selective activation of the fatty acid synthesis pathway in human prostate cancer. Int. J. Cancer 2000, 88, 176–179. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dias, A.C.D.S.; Ruiz, N.; Couzinet-Mossion, A.; Bertrand, S.; Duflos, M.; Pouchus, Y.-F.; Barnathan, G.; Nazih, H.; Wielgosz-Collin, G. The Marine-Derived Fungus Clonostachys rosea, Source of a Rare Conjugated 4-Me-6E,8E-hexadecadienoic Acid Reducing Viability of MCF-7 Breast Cancer Cells and Gene Expression of Lipogenic Enzymes. Mar. Drugs 2015, 13, 4934-4948. https://doi.org/10.3390/md13084934
Dias ACDS, Ruiz N, Couzinet-Mossion A, Bertrand S, Duflos M, Pouchus Y-F, Barnathan G, Nazih H, Wielgosz-Collin G. The Marine-Derived Fungus Clonostachys rosea, Source of a Rare Conjugated 4-Me-6E,8E-hexadecadienoic Acid Reducing Viability of MCF-7 Breast Cancer Cells and Gene Expression of Lipogenic Enzymes. Marine Drugs. 2015; 13(8):4934-4948. https://doi.org/10.3390/md13084934
Chicago/Turabian StyleDias, Ana Camila Dos Santos, Nicolas Ruiz, Aurélie Couzinet-Mossion, Samuel Bertrand, Muriel Duflos, Yves-François Pouchus, Gilles Barnathan, Hassan Nazih, and Gaetane Wielgosz-Collin. 2015. "The Marine-Derived Fungus Clonostachys rosea, Source of a Rare Conjugated 4-Me-6E,8E-hexadecadienoic Acid Reducing Viability of MCF-7 Breast Cancer Cells and Gene Expression of Lipogenic Enzymes" Marine Drugs 13, no. 8: 4934-4948. https://doi.org/10.3390/md13084934
APA StyleDias, A. C. D. S., Ruiz, N., Couzinet-Mossion, A., Bertrand, S., Duflos, M., Pouchus, Y.-F., Barnathan, G., Nazih, H., & Wielgosz-Collin, G. (2015). The Marine-Derived Fungus Clonostachys rosea, Source of a Rare Conjugated 4-Me-6E,8E-hexadecadienoic Acid Reducing Viability of MCF-7 Breast Cancer Cells and Gene Expression of Lipogenic Enzymes. Marine Drugs, 13(8), 4934-4948. https://doi.org/10.3390/md13084934