Predicting the Cochlear Dead Regions Using a Machine Learning-Based Approach with Oversampling Techniques
Abstract
:1. Introduction
2. Materials and Methods
2.1. Subjects
2.2. TEN (HL) Test
2.3. Model Development
2.4. Statistical Analysis and Model Evaluation Methods
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Moore, B.C.; Huss, M.; Vickers, D.A.; Glasberg, B.R.; Alcantara, J.I. A test for the diagnosis of dead regions in the cochlea. Br. J. Audiol. 2000, 34, 205–224. [Google Scholar] [CrossRef] [PubMed]
- Preminger, J.E.; Carpenter, R.; Ziegler, C.H. A clinical perspective on cochlear dead regions: Intelligibility of speech and subjective hearing aid benefit. J. Am. Acad. Audiol. 2005, 16, 600–613. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huss, M.; Moore, B.C. Dead regions and pitch perception. J. Acoust. Soc. Am. 2005, 117, 3841–3852. [Google Scholar] [CrossRef] [PubMed]
- Pepler, A.; Munro, K.J.; Lewis, K.; Kluk, K. Prevalence of Cochlear Dead Regions in New Referrals and Existing Adult Hearing Aid Users. Ear Hear. 2014, 35, e99–e109. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.-S.; Park, H.; Hong, S.H.; Chung, W.-H.; Cho, Y.-S.; Moon, I.J. Predicting cochlear dead regions in patients with hearing loss through a machine learning-based approach: A preliminary study. PLoS ONE 2019, 14, e0217790. [Google Scholar] [CrossRef] [PubMed]
- Moore, B.C.; Glasberg, B.R.; Stone, M.A. New version of the TEN test with calibrations in dB HL. Ear Hear. 2004, 25, 478–487. [Google Scholar] [CrossRef] [PubMed]
- Carhart, R.; Jerger, J.F. Preferred method for clinical determination of pure-tone thresholds. J. Speech Hear. Disord. 1959, 24, 330–345. [Google Scholar] [CrossRef]
- Ahadi, M.; Milani, M.; Malayeri, S. Prevalence of cochlear dead regions in moderate to severe sensorineural hearing impaired children. Int. J. Pediatr. Otorhinolaryngol. 2015, 79, 1362–1365. [Google Scholar] [CrossRef] [PubMed]
- Strobl, C.; Malley, J.; Tutz, G. An introduction to recursive partitioning: Rationale, application, and characteristics of classification and regression trees, bagging, and random forests. Psychol. Methods 2009, 14, 323–348. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, W.Y.; Chen, C.H.; Tseng, Y.J.; Tsai, Y.T.; Chang, C.Y.; Wang, H.Y.; Chen, C.K. Predicting post-stroke activities of daily living through a machine learning-based approach on initiating rehabilitation. Int. J. Med. Inform. 2018, 111, 159–164. [Google Scholar] [CrossRef] [PubMed]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. SMOTE: Synthetic minority over-sampling technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Kuhn, M. The caret package. J. Stat. Softw. 2009, 28. [Google Scholar]
- Hothorn, T.; Hornik, K.; Zeileis, A. Unbiased recursive partitioning: A conditional inference framework. J. Comput. Graph. Stat. 2006, 15, 651–674. [Google Scholar] [CrossRef] [Green Version]
- Torgo, L.; Torgo, M.L. Package ‘Dmwr’. Comprehensive R Archive Network. 2013. Available online: http://www2.uaem.mx/r-mirror/web/packages/DMwR/DMwR.pdf (accessed on 25 October 2021).
- Rahman, M.M.; Davis, D.N. Addressing the class imbalance problem in medical datasets. Int. J. Mach. Learn. Comput. 2013, 3, 224. [Google Scholar] [CrossRef]
- Oh, S.-J.; Lee, I.-W.; Wang, S.-G.; Kong, S.-K.; Kim, H.-K.; Goh, E.-K. Extratympanic Observation of Middle and Inner Ear Structures in Rodents Using Optical Coherence Tomography. Clin. Exp. Otorhinolaryngol. 2020, 13, 106–112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aazh, H.; Moore, B.C. Dead regions in the cochlea at 4 kHz in elderly adults: Relation to absolute threshold, steepness of audiogram, and pure-tone average. J. Am. Acad. Audiol. 2007, 18, 97–106. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, H.Y.; Seo, Y.M.; Kim, K.A.; Kang, Y.S.; Cho, C.S. Clinical Application of the Threshold Equalizing Noise Test in Patients with Hearing Loss of Various Etiologies: A Preliminary Study. J. Audiol. Otol. 2015, 19, 20–25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cox, R.M.; Alexander, G.C.; Johnson, J.; Rivera, I. Cochlear dead regions in typical hearing aid candidates: Prevalence and implications for use of high-frequency speech cues. Ear Hear. 2011, 32, 339–348. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moore, B.C. Prevalence of dead regions in subjects with sensorineural hearing loss. Ear Hear. 2007, 28, 231–241. [Google Scholar] [CrossRef] [PubMed]
- Halpin, C.; Rauch, S.D. Clinical implications of a damaged cochlea: Pure tone thresholds vs information-carrying capacity. Otolaryngol. Head Neck Surg. 2009, 140, 473–476. [Google Scholar] [CrossRef] [PubMed]
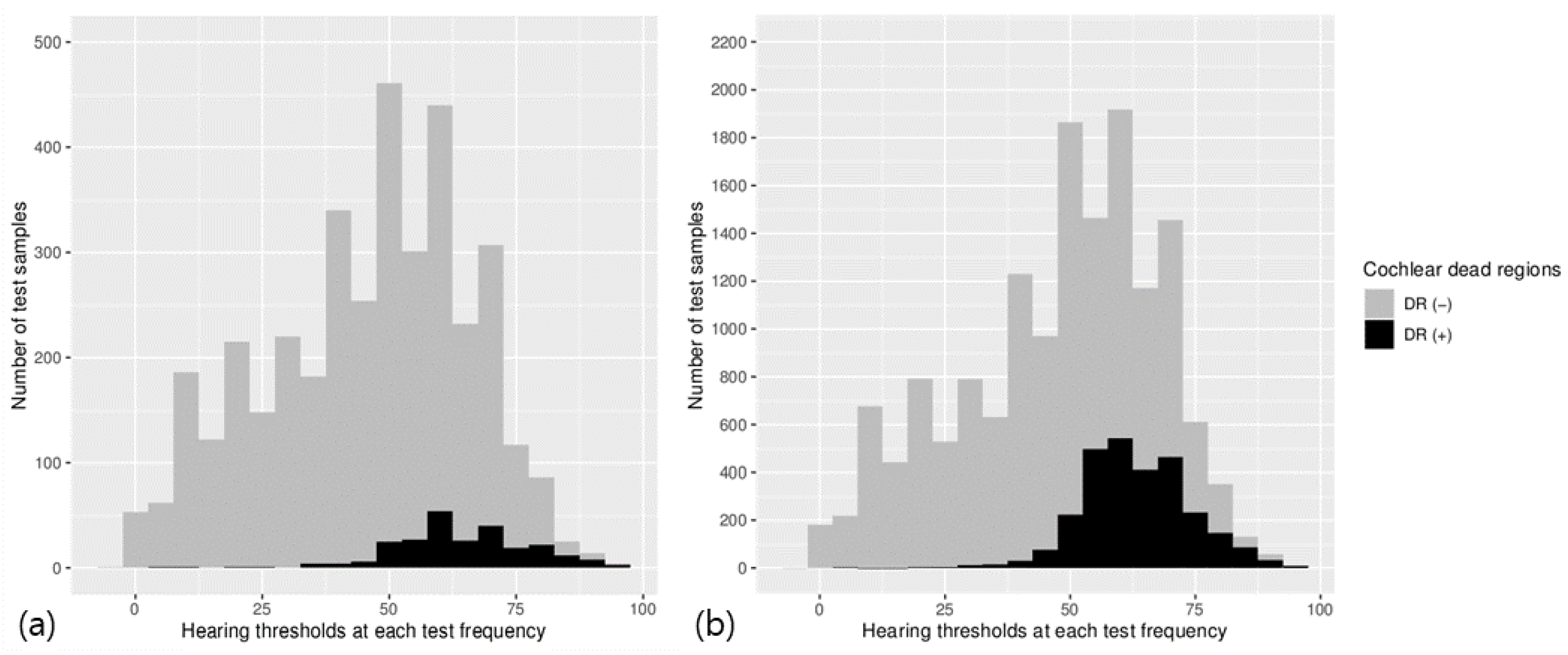
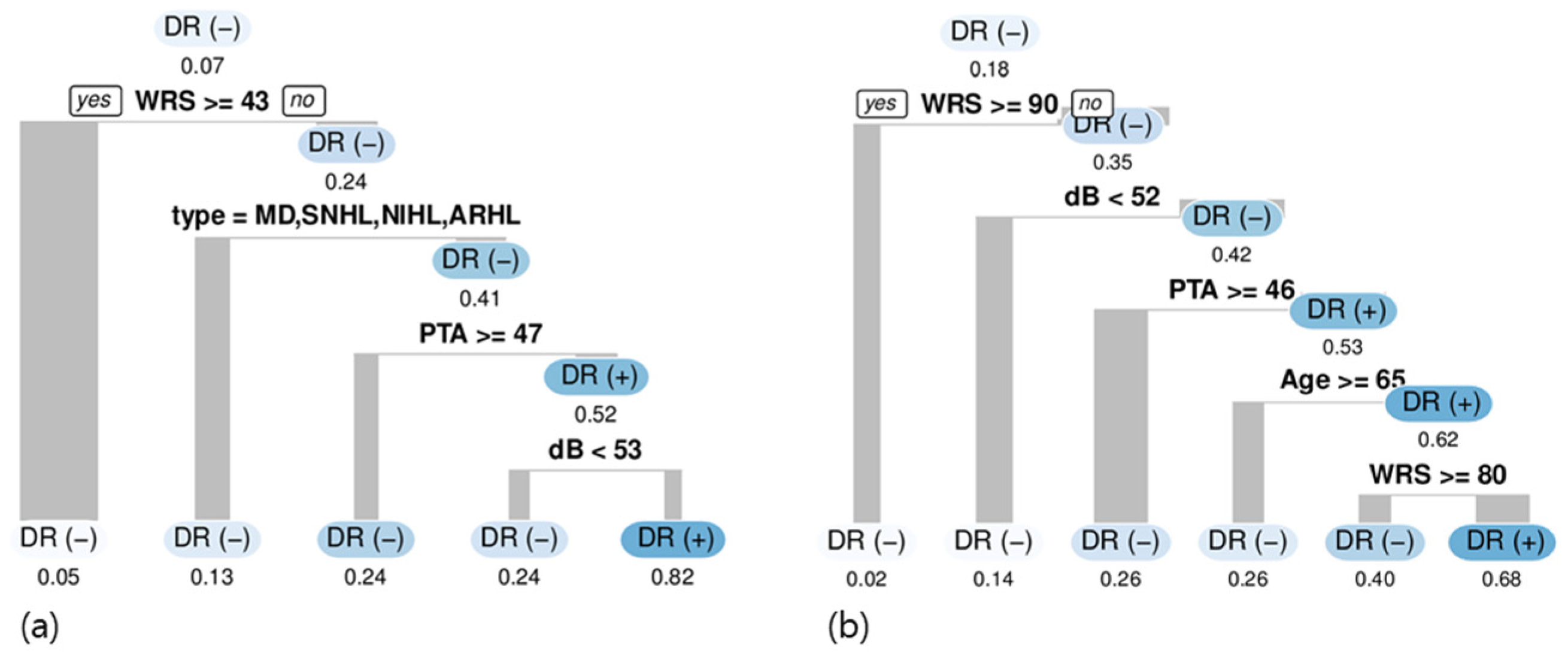
| Original Data (555 Ears) | Oversampled Data (15,494 Samples) | |
|---|---|---|
| Side | ||
| Right | 285 (51.35%) | 7857 (50.71%) |
| Left | 270 (48.65%) | 7637 (49.29%) |
| PTA (dB) | 44.8 ± 16.0 | 33.4 ± 13.1 |
| WRS (%) | 82.1 ± 23.9 | 78.9 ± 23.8 |
| Types of diseases | ||
| SNHL with unknown etiology | 114 (20.54%) | 3513 (22.67%) |
| SSNHL | 99 (17.84%) | 2649 (17.10%) |
| VS | 39 (7.03%) | 1339 (8.64%) |
| MD | 65 (11.71%) | 1832 (11.82%) |
| NIHL | 70 (12.61%) | 1882 (12.15%) |
| ARHL | 168 (30.27%) | 4279 (27.62%) |
| Original Data | Oversampled Data | |||||
|---|---|---|---|---|---|---|
| Odds Ratio | 95% Confidence Interval | p-Value | Odds Ratio | 95% Confidence Interval | p-Value | |
| Age | 0.99 | 0.98–1.01 | 0.36 | 0.99 | 0.99–1.00 | <0.001 * |
| Sex (reference: Female) Male | 0.42 | 0.29–0.61 | <0.001 * | 0.52 | 0.48–0.61 | <0.001 * |
| PTA (dB) | 0.94 | 0.92–0.96 | <0.001 * | 0.95 | 0.94–0.96 | <0.001 * |
| WRS (reference: ≥40) <40 | 3.77 | <0.001 * | 1.90 | 1.67–2.17 | <0.001 * | |
| Pure tone threshold of each frequency (dB) | 1.11 | 1.09–1.13 | <0.001 * | 1.11 | 1.10–1.12 | <0.001 * |
| Types of diseases | ||||||
| (reference: SNHL) | ||||||
| SSNHL | 1.45 | 0.88–2.41 | 0.15 | 1.56 | 1.31–1.85 | <0.001 * |
| VS | 2.40 | 1.36–4.23 | 0.002 * | 2.67 | 2.19–3.24 | <0.001 * |
| MD | 0.36 | 0.18–0.73 | 0.004 * | 0.51 | 0.41–0.63 | <0.001 * |
| NIHL | 0.46 | 0.18–1.15 | 0.10 | 1.05 | 0.82–1.34 | 0.70 |
| ARHL | 0.96 | 0.53–1.74 | 0.88 | 0.88 | 0.73–1.07 | 0.21 |
| Frequency | ||||||
| (reference: 1000 Hz) | ||||||
| 500 Hz | 1.36 | 0.74–2.53 | 0.32 | 0.66 | 0.55–0.86 | <0.001 * |
| 750 Hz | 1.12 | 0.60–2.07 | 0.73 | 0.73 | 0.60–0.90 | <0.001 * |
| 1500 Hz | 0.66 | 0.34–1.26 | 0.21 | 0.77 | 0.62–0.93 | 0.002 * |
| 2000 Hz | 0.82 | 0.44–1.53 | 0.53 | 0.73 | 0.66–0.97 | 0.009 * |
| 3000 Hz | 0.22 | 0.11–0.46 | <0.001 * | 0.19 | 0.17–0.27 | <0.001 * |
| 4000 Hz | 0.31 | 0.15–0.62 | <0.001 * | 0.13 | 0.12–0.20 | <0.001 * |
| Intercept | 0.01 | 0.00–0.02 | <0.001 * | 0.02 | 0.01–0.03 | <0.001 * |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chang, Y.-S.; Park, H.-S.; Moon, I.-J. Predicting the Cochlear Dead Regions Using a Machine Learning-Based Approach with Oversampling Techniques. Medicina 2021, 57, 1192. https://doi.org/10.3390/medicina57111192
Chang Y-S, Park H-S, Moon I-J. Predicting the Cochlear Dead Regions Using a Machine Learning-Based Approach with Oversampling Techniques. Medicina. 2021; 57(11):1192. https://doi.org/10.3390/medicina57111192
Chicago/Turabian StyleChang, Young-Soo, Hee-Sung Park, and Il-Joon Moon. 2021. "Predicting the Cochlear Dead Regions Using a Machine Learning-Based Approach with Oversampling Techniques" Medicina 57, no. 11: 1192. https://doi.org/10.3390/medicina57111192
APA StyleChang, Y.-S., Park, H.-S., & Moon, I.-J. (2021). Predicting the Cochlear Dead Regions Using a Machine Learning-Based Approach with Oversampling Techniques. Medicina, 57(11), 1192. https://doi.org/10.3390/medicina57111192






