Gut Microbiota and Iron: The Crucial Actors in Health and Disease
Abstract
1. Introduction
2. Mammalian Gut Microbiome in Health
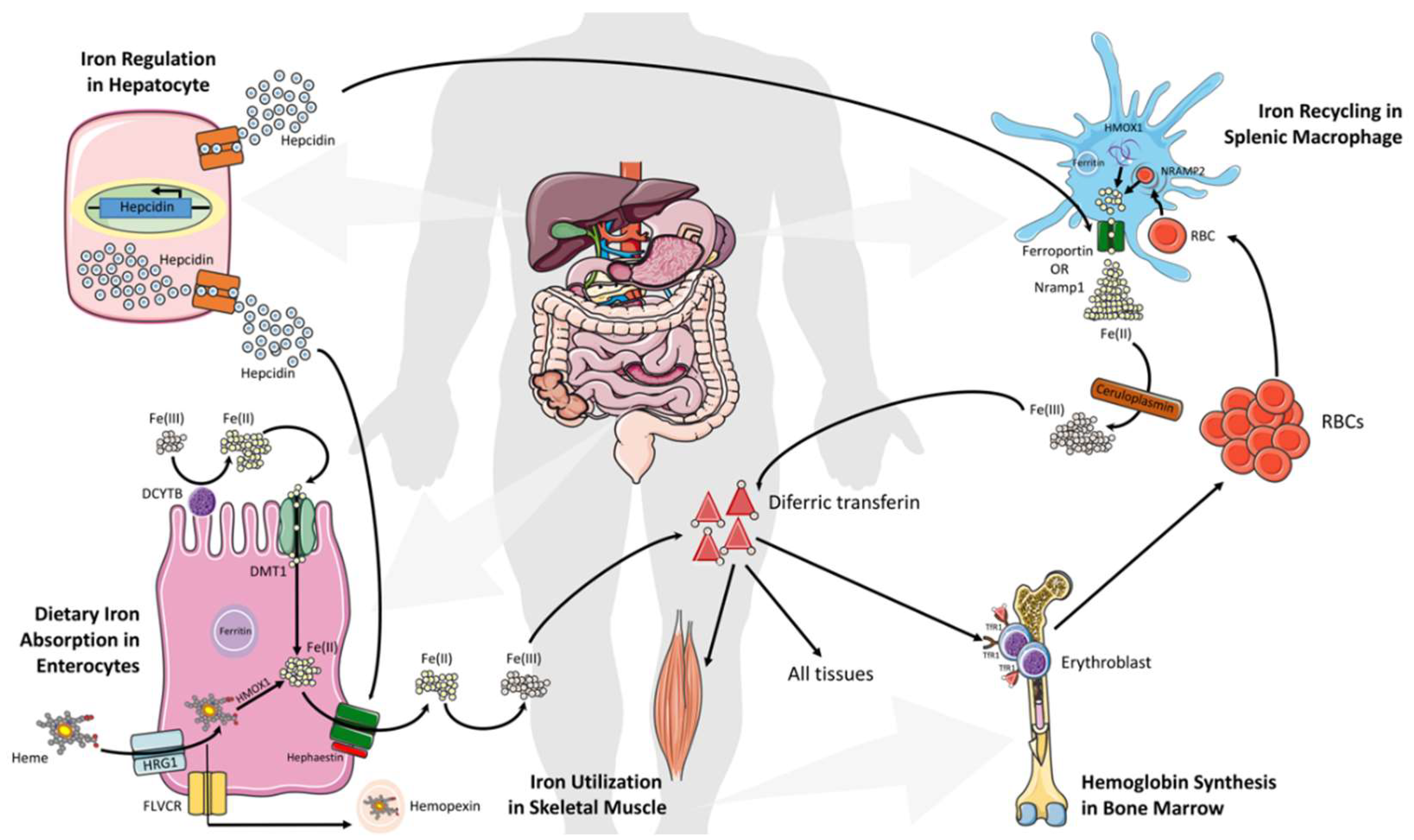
3. Systemic Iron Metabolism and Homeostasis
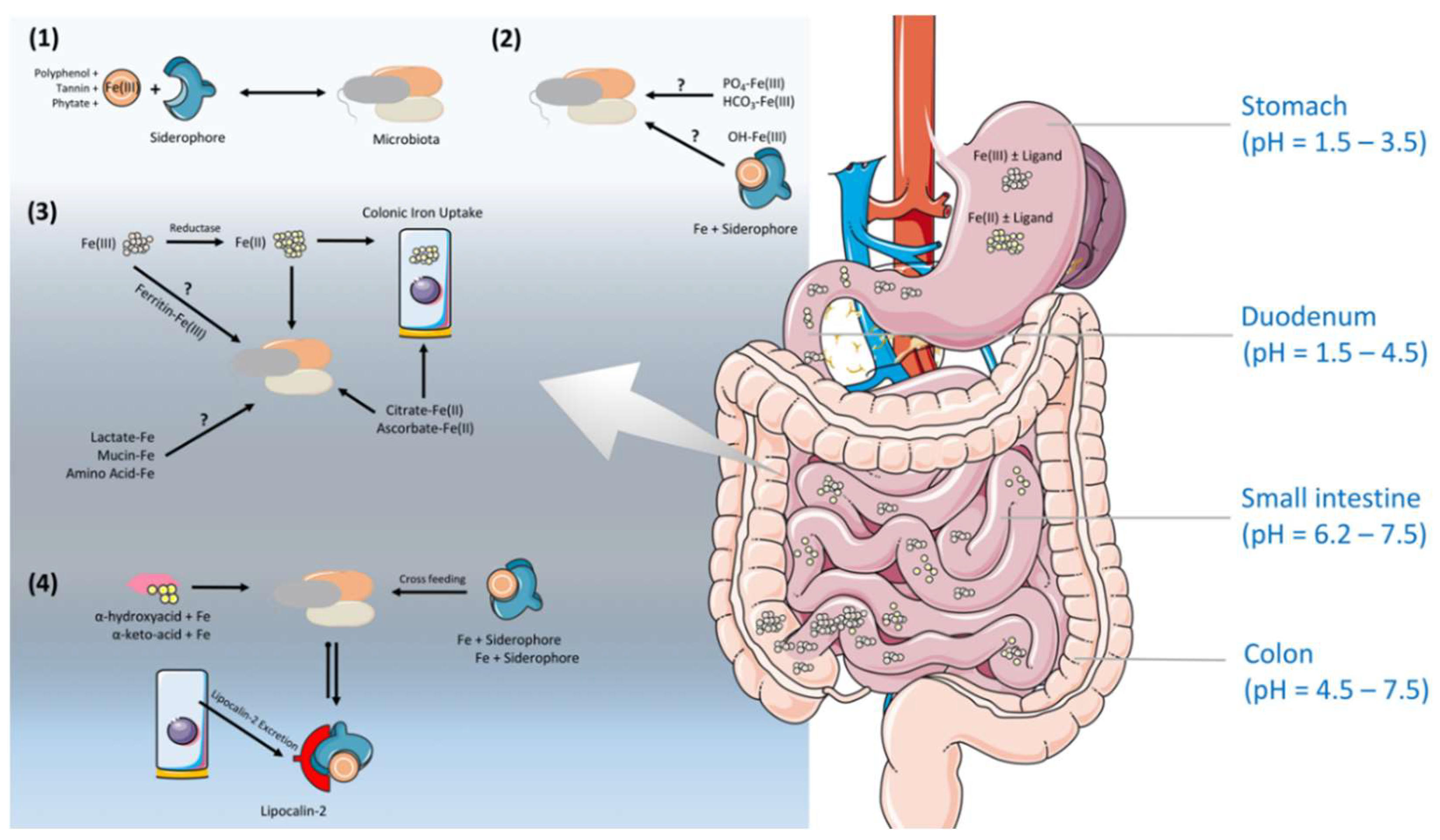
4. Iron Regulation Along the Gastrointestinal Tract (GIT) Under the Shade of the Gut Microbiota
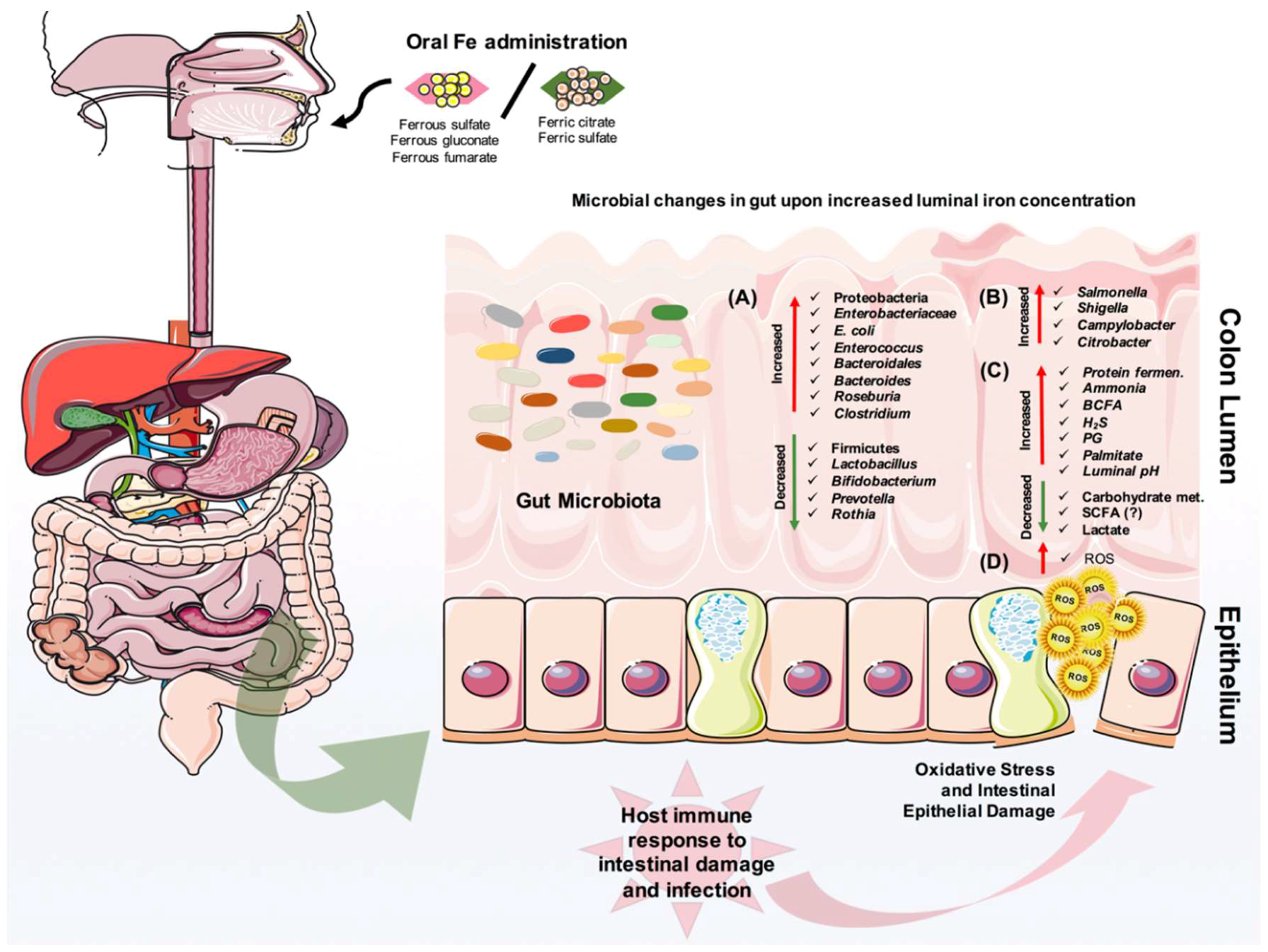
5. The Effect of Iron on Gut Microbiota and Pathogens
6. Iron and Inflammatory Bowel Disease (IBD)
7. Iron and Colorectal Cancer
8. Concluding Remarks
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ACD | Anaemia of chronic disease |
| BCFA | Branched chain fatty acids |
| BMPs | Bone morphogenetic proteins |
| CD | Crohn’s disease |
| DCYTB | Duodenal Cytochrome B |
| DMT1 | Divalent Metal Transporter 1 |
| GF | Germ-free |
| GI | Gastrointestinal |
| H2O2 | Hydrogen peroxide |
| HEPH | Hephaestin |
| HMOX1 | Heme Oxygenase 1 |
| IBD | Inflammatory bowel disease |
| IBS | Irritable bowel syndrome |
| IDA | Iron deficiency anemia |
| Nramp1 | Natural Resistance-Associated |
| PG | Phosphatidylglycerol |
| RBC | Red blood cell |
| ROS | Reactive oxygen species |
| SCFA | Short-chain fatty acids |
| SLC40A1 | Solute Carrier Family 40 Member 1 |
| SLC46A1 | Solute Carrier Family 46 Member 1 |
| SNP | Single nucleotide polymorphisms |
| SOD | Superoxide dismutase |
| SPF | Specific pathogen-free |
| UC | Ulcerative colitis |
References
- Wessling-Resnick, M. Iron homeostasis and the inflammatory response. Annu. Rev. Nutr. 2010, 30, 105–122. [Google Scholar] [CrossRef] [PubMed]
- Nairz, M.; Schroll, A.; Sonnweber, T.; Weiss, G. The struggle for iron—A metal at the host-pathogen interface. Cell Microbiol. 2010, 12, 1691–1702. [Google Scholar] [CrossRef] [PubMed]
- Markel, T.A.; Crisostomo, P.R.; Wang, M.; Herring, C.M.; Meldrum, K.K.; Lillemoe, K.D.; Meldrum, D.R. The struggle for iron: Gastrointestinal microbes modulate the host immune response during infection. J. Leukoc. Biol. 2007, 81, 393–400. [Google Scholar] [CrossRef] [PubMed]
- Sheftel, A.D.; Mason, A.B.; Ponka, P. The long history of iron in the Universe and in health and disease. Biochim. Biophys. Acta 2012, 1820, 161–187. [Google Scholar] [CrossRef] [PubMed]
- Ganz, T. Iron and infection. Int. J. Hematol. 2018, 107, 7–15. [Google Scholar] [CrossRef] [PubMed]
- Stein, J.; Hartmann, F.; Dignass, A.U. Diagnosis and management of iron deficiency anemia in patients with IBD. Nat. Rev. Gastroenterol. Hepatol. 2010, 7, 599–610. [Google Scholar] [CrossRef] [PubMed]
- Goodnough, L.T.; Nemeth, E.; Ganz, T. Detection, evaluation, and management of iron-restricted erythropoiesis. Blood 2010, 116, 4754–4761. [Google Scholar] [CrossRef] [PubMed]
- Cherayil, B.J. Iron and immunity: Immunological consequences of iron deficiency and overload. Arch. Immunol. Ther. Exp. 2010, 58, 407–415. [Google Scholar] [CrossRef] [PubMed]
- Hooper, L.V.; Wong, M.H.; Thelin, A.; Hansson, L.; Falk, P.G.; Gordon, J.I. Molecular analysis of commensal host-microbial relationships in the intestine. Science 2001, 291, 881–884. [Google Scholar] [CrossRef] [PubMed]
- Hooper, L.V.; Midtvedt, T.; Gordon, J.I. How host-microbial interactions shape the nutrient environment of the mammalian intestine. Annu. Rev. Nutr. 2002, 22, 283–307. [Google Scholar] [CrossRef] [PubMed]
- Hooper, L.V.; Macpherson, A.J. Immune adaptations that maintain homeostasis with the intestinal microbiota. Nat. Rev. Immunol. 2010, 10, 159–169. [Google Scholar] [CrossRef] [PubMed]
- Stecher, B.; Macpherson, A.J.; Hapfelmeier, S.; Kremer, M.; Stallmach, T.; Hardt, W.D. Comparison of Salmonella enterica serovar Typhimurium colitis in germfree mice and mice pretreated with streptomycin. Infect. Immun. 2005, 73, 3228–3241. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, B.; Portugal, S.; Tran, T.M.; Gozzelino, R.; Ramos, S.; Gomes, J.; Regalado, A.; Cowan, P.J.; d’Apice, A.J.; Chong, A.S.; et al. Gut microbiota elicits a protective immune response against malaria transmission. Cell 2014, 159, 1277–1289. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, B.; Schibli, S.; Macpherson, A.J.; Sokollik, C. D-lactic Acidosis: Successful Suppression of D-lactate-Producing Lactobacillus by Probiotics. Pediatrics 2018. [Google Scholar] [CrossRef] [PubMed]
- Soares, M.P.; Yilmaz, B. Microbiota Control of Malaria Transmission. Trends Parasitol. 2016, 32, 120–130. [Google Scholar] [CrossRef] [PubMed]
- Uchimura, Y.; Fuhrer, T.; Li, H.; Lawson, M.A.; Zimmermann, M.; Yilmaz, B.; Zindel, J.; Ronchi, F.; Sorribas, M.; Hapfelmeier, S.; et al. Antibodies Set Boundaries Limiting Microbial Metabolite Penetration and the Resultant Mammalian Host Response. Immunity 2018, 49, 545–559. [Google Scholar] [CrossRef] [PubMed]
- Smith, K.; McCoy, K.D.; Macpherson, A.J. Use of axenic animals in studying the adaptation of mammals to their commensal intestinal microbiota. Semin. Immunol. 2007, 19, 59–69. [Google Scholar] [CrossRef] [PubMed]
- Cahenzli, J.; Balmer, M.L.; McCoy, K.D. Microbial-immune cross-talk and regulation of the immune system. Immunology 2013, 138, 12–22. [Google Scholar] [CrossRef] [PubMed]
- Ganal, S.C.; Sanos, S.L.; Kallfass, C.; Oberle, K.; Johner, C.; Kirschning, C.; Lienenklaus, S.; Weiss, S.; Staeheli, P.; Aichele, P.; et al. Priming of Natural Killer Cells by Nonmucosal Mononuclear Phagocytes Requires Instructive Signals from Commensal Microbiota. Immunity 2012, 37, 171–186. [Google Scholar] [CrossRef] [PubMed]
- Macpherson, A.J.; Yilmaz, B.; Limenitakis, J.P.; Ganal-Vonarburg, S.C. IgA Function in Relation to the Intestinal Microbiota. Annu. Rev. Immunol. 2018, 36, 359–381. [Google Scholar] [CrossRef] [PubMed]
- Holmes, E.; Li, J.V.; Athanasiou, T.; Ashrafian, H.; Nicholson, J.K. Understanding the role of gut microbiome-host metabolic signal disruption in health and disease. Trends Microbiol. 2011, 19, 349–359. [Google Scholar] [CrossRef] [PubMed]
- Clayton, T.A.; Lindon, J.C.; Cloarec, O.; Antti, H.; Charuel, C.; Hanton, G.; Provost, J.P.; Le Net, J.L.; Baker, D.; Walley, R.J.; et al. Pharmaco-metabonomic phenotyping and personalized drug treatment. Nature 2006, 440, 1073–1077. [Google Scholar] [CrossRef] [PubMed]
- Grundmann, O. The Gut Microbiome and Pre-systemic Metabolism: Current State and Evolving Research. J. Drug Metab. Toxicol. 2010, 1, 1–7. [Google Scholar] [CrossRef]
- Nieuwdorp, M.; Gilijamse, P.W.; Pai, N.; Kaplan, L.M. Role of the microbiome in energy regulation and metabolism. Gastroenterology 2014, 146, 1525–1533. [Google Scholar] [CrossRef] [PubMed]
- De Aguiar Vallim, T.Q.; Tarling, E.J.; Edwards, P.A. Pleiotropic roles of bile acids in metabolism. Cell Metab. 2013, 17, 657–669. [Google Scholar] [CrossRef] [PubMed]
- Sekirov, I.; Russell, S.L.; Antunes, L.C.M.; Finlay, B.B. Gut Microbiota in Health and Disease. Physiol. Rev. 2010, 90, 859–904. [Google Scholar] [CrossRef] [PubMed]
- Peterson, D.A.; Frank, D.N.; Pace, N.R.; Gordon, J.I. Metagenomic approaches for defining the pathogenesis of inflammatory bowel diseases. Cell Host Microbe 2008, 3, 417–427. [Google Scholar] [CrossRef] [PubMed]
- Maloy, K.J.; Powrie, F. Intestinal homeostasis and its breakdown in inflammatory bowel disease. Nature 2011, 474, 298–306. [Google Scholar] [CrossRef] [PubMed]
- Mazmanian, S.K.; Round, J.L.; Kasper, D.L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 2008, 453, 620–625. [Google Scholar] [CrossRef] [PubMed]
- Kamada, N.; Chen, G.Y.; Inohara, N.; Nunez, G. Control of pathogens and pathobionts by the gut microbiota. Nat. Immun. 2013, 14, 685–690. [Google Scholar] [CrossRef] [PubMed]
- Distrutti, E.; Monaldi, L.; Ricci, P.; Fiorucci, S. Gut microbiota role in irritable bowel syndrome: New therapeutic strategies. World J. Gastroenterol. 2016, 22, 2219–2241. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.Y.; Tsai, C.N.; Lee, Y.S.; Lin, C.Y.; Huang, K.Y.; Chao, H.C.; Lai, M.W.; Chiu, C.H. Intestinal microbiome in children with severe and complicated acute viral gastroenteritis. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Duvallet, C.; Gibbons, S.M.; Gurry, T.; Irizarry, R.A.; Alm, E.J. Meta-analysis of gut microbiome studies identifies disease-specific and shared responses. Nat. Commun. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Org, E.; Parks, B.W.; Joo, J.W.J.; Emert, B.; Schwartzman, W.; Kang, E.Y.; Mehrabian, M.; Pan, C.; Knight, R.; Gunsalus, R.; et al. Genetic and environmental control of host-gut microbiota interactions. Genome Res. 2015, 25, 1558–1569. [Google Scholar] [CrossRef] [PubMed]
- Rothschild, D.; Weissbrod, O.; Barkan, E.; Kurilshikov, A.; Korem, T.; Zeevi, D.; Costea, P.I.; Godneva, A.; Kalka, I.N.; Bar, N.; et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 2018, 555, 210. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.Z.; van Sommeren, S.; Huang, H.; Ng, S.C.; Alberts, R.; Takahashi, A.; Ripke, S.; Lee, J.C.; Jostins, L.; Shah, T.; et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 2015, 47, 979–986. [Google Scholar] [CrossRef] [PubMed]
- De Lange, K.M.; Moutsianas, L.; Lee, J.C.; Lamb, C.A.; Luo, Y.; Kennedy, N.A.; Jostins, L.; Rice, D.L.; Gutierrez-Achury, J.; Ji, S.G.; et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 2017, 49, 256–261. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, B.; Spalinger, M.R.; Biedermann, L.; Franc, Y.; Fournier, N.; Rossel, J.B.; Juillerat, P.; Rogler, G.; Macpherson, A.J.; Scharl, M. The presence of genetic risk variants within PTPN2 and PTPN22 is associated with intestinal microbiota alterations in Swiss IBD cohort patients. PLoS ONE 2018, 13, e0199664. [Google Scholar] [CrossRef] [PubMed]
- Faith, J.J.; Guruge, J.L.; Charbonneau, M.; Subramanian, S.; Seedorf, H.; Goodman, A.L.; Clemente, J.C.; Knight, R.; Heath, A.C.; Leibel, R.L.; et al. The long-term stability of the human gut microbiota. Science 2013, 341, 1237439. [Google Scholar] [CrossRef] [PubMed]
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Schloissnig, S.; Arumugam, M.; Sunagawa, S.; Mitreva, M.; Tap, J.; Zhu, A.; Waller, A.; Mende, D.R.; Kultima, J.R.; Martin, J.; et al. Genomic variation landscape of the human gut microbiome. Nature 2013, 493, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Pantopoulos, K. Regulation of cellular iron metabolism. Biochem. J. 2011, 434, 365–381. [Google Scholar] [CrossRef] [PubMed]
- Dev, S.; Babitt, J.L. Overview of iron metabolism in health and disease. Hemodial. Int. 2017, 21 (Suppl. 1), S6–S20. [Google Scholar] [CrossRef]
- McKie, A.T.; Barrow, D.; Latunde-Dada, G.O.; Rolfs, A.; Sager, G.; Mudaly, E.; Mudaly, M.; Richardson, C.; Barlow, D.; Bomford, A.; et al. An iron-regulated ferric reductase associated with the absorption of dietary iron. Science 2001, 291, 1755–1759. [Google Scholar] [CrossRef] [PubMed]
- Nunez, G.; Sakamoto, K.; Soares, M.P. Innate Nutritional Immunity. J. Immunol. 2018, 201, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Hentze, M.W.; Muckenthaler, M.U.; Galy, B.; Camaschella, C. Two to tango: Regulation of Mammalian iron metabolism. Cell 2010, 142, 24–38. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Bandyopadhyay, U. Free heme toxicity and its detoxification systems in human. Toxicol. Lett. 2005, 157, 175–188. [Google Scholar] [CrossRef] [PubMed]
- Chua, A.C.; Graham, R.M.; Trinder, D.; Olynyk, J.K. The regulation of cellular iron metabolism. Crit. Rev. Clin. Lab. Sci. 2007, 44, 413–459. [Google Scholar] [CrossRef] [PubMed]
- Gunshin, H.; Mackenzie, B.; Berger, U.V.; Gunshin, Y.; Romero, M.F.; Boron, W.F.; Nussberger, S.; Gollan, J.L.; Hediger, M.A. Cloning and characterization of a mammalian proton-coupled metal-ion transporter. Nature 1997, 388, 482–488. [Google Scholar] [CrossRef] [PubMed]
- Qiu, A.; Jansen, M.; Sakaris, A.; Min, S.H.; Chattopadhyay, S.; Tsai, E.; Sandoval, C.; Zhao, R.; Akabas, M.H.; Goldman, I.D. Identification of an intestinal folate transporter and the molecular basis for hereditary folate malabsorption. Cell 2006, 127, 917–928. [Google Scholar] [CrossRef] [PubMed]
- Wallander, M.L.; Leibold, E.A.; Eisenstein, R.S. Molecular control of vertebrate iron homeostasis by iron regulatory proteins. Biochim. Biophys. Acta 2006, 1763, 668–689. [Google Scholar] [CrossRef] [PubMed]
- Ganz, T. Erythropoietic regulators of iron metabolism. Free Radic. Biol. Med. 2018. [Google Scholar] [CrossRef] [PubMed]
- Cherayil, B.J.; Ellenbogen, S.; Shanmugam, N.N. Iron and intestinal immunity. Curr. Opin. Gastroenterol. 2011, 27, 523–528. [Google Scholar] [CrossRef] [PubMed]
- Nemeth, E.; Tuttle, M.S.; Powelson, J.; Vaughn, M.B.; Donovan, A.; Ward, D.M.; Ganz, T.; Kaplan, J. Hepcidin regulates cellular iron efflux by binding to ferroportin and inducing its internalization. Science 2004, 306, 2090–2093. [Google Scholar] [CrossRef] [PubMed]
- Deugnier, Y.; Turlin, B. Pathology of hepatic iron overload. Semin. Liver Dis. 2011, 31, 260–271. [Google Scholar] [CrossRef] [PubMed]
- Deugnier, Y.; Turlin, B. Pathology of hepatic iron overload. World J. Gastroenterol. 2007, 13, 4755–4760. [Google Scholar] [CrossRef] [PubMed]
- Ovesen, L.; Bendtsen, F.; Tage-Jensen, U.; Pedersen, N.T.; Gram, B.R.; Rune, S.J. Intraluminal pH in the stomach, duodenum, and proximal jejunum in normal subjects and patients with exocrine pancreatic insufficiency. Gastroenterology 1986, 90, 958–962. [Google Scholar] [CrossRef]
- Jacobs, A.; Miles, P.M. Intraluminal transport of iron from stomach to small-intestinal mucosa. Br. Med. J. 1969, 4, 778–781. [Google Scholar] [CrossRef] [PubMed]
- Hedrich, S.; Schlomann, M.; Johnson, D.B. The iron-oxidizing proteobacteria. Microbiology 2011, 157, 1551–1564. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, K.; Bjarnason, I.; Laftah, A.H.; Latunde-Dada, G.O.; Simpson, R.J.; McKie, A.T. Expression of iron absorption genes in mouse large intestine. Scand. J. Gastroenterol. 2005, 40, 169–177. [Google Scholar] [CrossRef] [PubMed]
- Cowart, R.E. Reduction of iron by extracellular iron reductases: Implications for microbial iron acquisition. Arch. Biochem. Biophys. 2002, 400, 273–281. [Google Scholar] [CrossRef]
- Romanowski, K.; Zaborin, A.; Fernandez, H.; Poroyko, V.; Valuckaite, V.; Gerdes, S.; Liu, D.C.; Zaborina, O.Y.; Alverdy, J.C. Prevention of siderophore- mediated gut-derived sepsis due to P. aeruginosa can be achieved without iron provision by maintaining local phosphate abundance: Role of pH. BMC Microbiol. 2011, 11, 212. [Google Scholar] [CrossRef] [PubMed]
- Salovaara, S.; Sandberg, A.S.; Andlid, T. Combined impact of pH and organic acids on iron uptake by Caco-2 cells. J. Agric. Food Chem. 2003, 51, 7820–7824. [Google Scholar] [CrossRef] [PubMed]
- Lund, E.K.; Wharf, S.G.; Fairweather-Tait, S.J.; Johnson, I.T. Increases in the concentrations of available iron in response to dietary iron supplementation are associated with changes in crypt cell proliferation in rat large intestine. J. Nutr. 1998, 128, 175–179. [Google Scholar] [CrossRef] [PubMed]
- Ohkawara, Y.; Bamba, M.; Nakai, I.; Kinka, S.; Masuda, M. The absorption of iron from the human large intestine. Gastroenterology 1963, 44, 611–614. [Google Scholar] [PubMed]
- Johnston, K.L.; Johnson, D.M.; Marks, J.; Srai, S.K.; Debnam, E.S.; Sharp, P.A. Non-haem iron transport in the rat proximal colon. Eur. J. Clin. Investig. 2006, 36, 35–40. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.; Yeoh, B.S.; Saha, P.; Tian, Y.; Singh, V.; Patterson, A.D.; Vijay-Kumar, M. Modulation of urinary siderophores by the diet, gut microbiota and inflammation in mice. J. Nutr. Biochem. 2017, 41, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Kruger, J.; Taylor, J.R.; Du, X.; De Moura, F.F.; Lonnerdal, B.; Oelofse, A. Effect of phytate reduction of sorghum, through genetic modification, on iron and zinc availability as assessed by an in vitro dialysability bioaccessibility assay, Caco-2 cell uptake assay, and suckling rat pup absorption model. Food Chem. 2013, 141, 1019–1025. [Google Scholar] [CrossRef] [PubMed]
- Sotelo, A.; Gonzalez-Osnaya, L.; Sanchez-Chinchillas, A.; Trejo, A. Role of oxate, phytate, tannins and cooking on iron bioavailability from foods commonly consumed in Mexico. Int. J. Food Sci. Nutr. 2010, 61, 29–39. [Google Scholar] [CrossRef] [PubMed]
- Hurrell, R.; Egli, I. Iron bioavailability and dietary reference values. Am. J. Clin. Nutr. 2010, 91, 1461S–1467S. [Google Scholar] [CrossRef] [PubMed]
- Van Dokkum, W. Significance of iron bioavailability for iron recommendations. Biol. Trace Elem. Res. 1992, 35, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Hallberg, L.; Brune, M.; Rossander, L. The role of vitamin C in iron absorption. Int. J. Vitam. Nutr. Res. Suppl. 1989, 30, 103–108. [Google Scholar] [PubMed]
- Salovaara, S.; Sandberg, A.S.; Andlid, T. Organic acids influence iron uptake in the human epithelial cell line Caco-2. J. Agric. Food Chem. 2002, 50, 6233–6238. [Google Scholar] [CrossRef] [PubMed]
- Blachier, F.; Vaugelade, P.; Robert, V.; Kibangou, B.; Canonne-Hergaux, F.; Delpal, S.; Bureau, F.; Blottiere, H.; Bougle, D. Comparative capacities of the pig colon and duodenum for luminal iron absorption. Can. J. Physiol. Pharmacol. 2007, 85, 185–192. [Google Scholar] [CrossRef] [PubMed]
- Tako, E.; Glahn, R.P.; Welch, R.M.; Lei, X.; Yasuda, K.; Miller, D.D. Dietary inulin affects the expression of intestinal enterocyte iron transporters, receptors and storage protein and alters the microbiota in the pig intestine. Br. J. Nutr. 2008, 99, 472–480. [Google Scholar] [CrossRef] [PubMed]
- Deschemin, J.C.; Noordine, M.L.; Remot, A.; Willemetz, A.; Afif, C.; Canonne-Hergaux, F.; Langella, P.; Karim, Z.; Vaulont, S.; Thomas, M.; et al. The microbiota shifts the iron sensing of intestinal cells. FASEB J. 2016, 30, 252–261. [Google Scholar] [CrossRef] [PubMed]
- Reddy, B.S.; Pleasants, J.R.; Wostmann, B.S. Effect of intestinal microflora on iron and zinc metabolism, and on activities of metalloenzymes in rats. J. Nutr. 1972, 102, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Forrester, R.H.; Conrad, M.E., Jr.; Crosby, W.H. Measurement of total body iron in animals using whole-body liquid scintillation detectors. Proc. Soc. Exp. Biol. Med. 1962, 111, 115–119. [Google Scholar] [CrossRef] [PubMed]
- Stern, P.; Kosak, R.; Misirlija, A. The problem of iron resorption. Experientia 1954, 10, 227. [Google Scholar] [CrossRef] [PubMed]
- Cremonesi, P.; Acebron, A.; Raja, K.B.; Simpson, R.J. Iron absorption: Biochemical and molecular insights into the importance of iron species for intestinal uptake. Pharmacol. Toxicol. 2002, 91, 97–102. [Google Scholar] [CrossRef] [PubMed]
- Raymond, K.N.; Dertz, E.A.; Kim, S.S. Enterobactin: An archetype for microbial iron transport. Proc. Natl. Acad. Sci. USA 2003, 100, 3584–3588. [Google Scholar] [CrossRef] [PubMed]
- Frawley, E.R.; Fang, F.C. The ins and outs of bacterial iron metabolism. Mol. Microbiol. 2014, 93, 609–616. [Google Scholar] [CrossRef] [PubMed]
- Andrews, S.C.; Robinson, A.K.; Rodriguez-Quinones, F. Bacterial iron homeostasis. FEMS Microbiol. Rev. 2003, 27, 215–237. [Google Scholar] [CrossRef]
- Archibald, F. Lactobacillus plantarum, an organism not requiring iron. FEMS Microbiol. Lett. 1983, 19, 29–32. [Google Scholar] [CrossRef]
- Weinberg, E.D. The Lactobacillus anomaly: Total iron abstinence. Perspect. Biol. Med. 1997, 40, 578–583. [Google Scholar] [CrossRef] [PubMed]
- Aguirre, J.D.; Clark, H.M.; McIlvin, M.; Vazquez, C.; Palmere, S.L.; Grab, D.J.; Seshu, J.; Hart, P.J.; Saito, M.; Culotta, V.C. A manganese-rich environment supports superoxide dismutase activity in a Lyme disease pathogen, Borrelia burgdorferi. J. Biol. Chem. 2013, 288, 8468–8478. [Google Scholar] [CrossRef] [PubMed]
- Posey, J.E.; Gherardini, F.C. Lack of a role for iron in the Lyme disease pathogen. Science 2000, 288, 1651–1653. [Google Scholar] [CrossRef] [PubMed]
- Neilands, J.B. Siderophores: Structure and function of microbial iron transport compounds. J. Biol. Chem. 1995, 270, 26723–26726. [Google Scholar] [CrossRef] [PubMed]
- Hai, L.; Limenitakis, J.P.; Fuhrer, T.; Geuking, M.B.; Lawson, M.B.; Wyss, M.; Brugiroux, S.; Keller, I.; Macpherson, J.A.; Rupp, S.; et al. The outer mucus layer hosts a distinct intestinal microbial niche. Nat. Commun. 2015, 6, 8292. [Google Scholar]
- Wandersman, C.; Stojiljkovic, I. Bacterial heme sources: The role of heme, hemoprotein receptors and hemophores. Curr. Opin. Microbiol. 2000, 3, 215–220. [Google Scholar] [CrossRef]
- Otto, B.R.; Sparrius, M.; Verweij-van Vught, A.M.; MacLaren, D.M. Iron-regulated outer membrane protein of Bacteroides fragilis involved in heme uptake. Infect. Immun. 1990, 58, 3954–3958. [Google Scholar] [PubMed]
- Mevissen-Verhage, E.A.; Marcelis, J.H.; Harmsen-Van Amerongen, W.C.; de Vos, N.M.; Verhoef, J. Effect of iron on neonatal gut flora during the first three months of life. Eur. J. Clin. Microbiol. 1985, 4, 273–278. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, M.B.; Chassard, C.; Rohner, F.; N’Goran, E.K.; Nindjin, C.; Dostal, A.; Utzinger, J.; Ghattas, H.; Lacroix, C.; Hurrell, R.F. The effects of iron fortification on the gut microbiota in African children: A randomized controlled trial in Cote d’Ivoire. Am. J. Clin. Nutr. 2010, 92, 1406–1415. [Google Scholar] [CrossRef] [PubMed]
- Balamurugan, R.; Mary, R.R.; Chittaranjan, S.; Jancy, H.; Shobana Devi, R.; Ramakrishna, B.S. Low levels of faecal lactobacilli in women with iron-deficiency anaemia in south India. Br. J. Nutr. 2010, 104, 931–934. [Google Scholar] [CrossRef] [PubMed]
- Jaeggi, T.; Kortman, G.A.; Moretti, D.; Chassard, C.; Holding, P.; Dostal, A.; Boekhorst, J.; Timmerman, H.M.; Swinkels, D.W.; Tjalsma, H.; et al. Iron fortification adversely affects the gut microbiome, increases pathogen abundance and induces intestinal inflammation in Kenyan infants. Gut 2015, 64, 731–742. [Google Scholar] [CrossRef] [PubMed]
- Dostal, A.; Fehlbaum, S.; Chassard, C.; Zimmermann, M.B.; Lacroix, C. Low iron availability in continuous in vitro colonic fermentations induces strong dysbiosis of the child gut microbial consortium and a decrease in main metabolites. FEMS Microbiol. Ecol. 2013, 83, 161–175. [Google Scholar] [CrossRef] [PubMed]
- Tompkins, G.R.; O’Dell, N.L.; Bryson, I.T.; Pennington, C.B. The effects of dietary ferric iron and iron deprivation on the bacterial composition of the mouse intestine. Curr. Microbiol. 2001, 43, 38–42. [Google Scholar] [CrossRef] [PubMed]
- Dostal, A.; Chassard, C.; Hilty, F.M.; Zimmermann, M.B.; Jaeggi, T.; Rossi, S.; Lacroix, C. Iron depletion and repletion with ferrous sulfate or electrolytic iron modifies the composition and metabolic activity of the gut microbiota in rats. J. Nutr. 2012, 142, 271–277. [Google Scholar] [CrossRef] [PubMed]
- Ettreiki, C.; Gadonna-Widehem, P.; Mangin, I.; Coeffier, M.; Delayre-Orthez, C.; Anton, P.M. Juvenile ferric iron prevents microbiota dysbiosis and colitis in adult rodents. World J. Gastroenterol. 2012, 18, 2619–2629. [Google Scholar] [CrossRef] [PubMed]
- Benoni, G.; Cuzzolin, L.; Zambreri, D.; Donini, M.; Del Soldato, P.; Caramazza, I. Gastrointestinal effects of single and repeated doses of ferrous sulphate in rats. Pharmacol. Res. 1993, 27, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Buhnik-Rosenblau, K.; Moshe-Belizowski, S.; Danin-Poleg, Y.; Meyron-Holtz, E.G. Genetic modification of iron metabolism in mice affects the gut microbiota. Biometals 2012, 25, 883–892. [Google Scholar] [CrossRef] [PubMed]
- Constante, M.; Fragoso, G.; Lupien-Meilleur, J.; Calve, A.; Santos, M.M. Iron Supplements Modulate Colon Microbiota Composition and Potentiate the Protective Effects of Probiotics in Dextran Sodium Sulfate-induced Colitis. Inflamm. Bowel Dis. 2017, 23, 753–766. [Google Scholar] [CrossRef] [PubMed]
- Yeung, C.K.; Glahn, R.P.; Welch, R.M.; Miller, D.D. Prebiotics and iron Bioavailability—Is there a connection? J. Food Sci. 2005, 70, R88–R92. [Google Scholar] [CrossRef]
- Weinborn, V.; Valenzuela, C.; Olivares, M.; Arredondo, M.; Weill, R.; Pizarro, F. Prebiotics increase heme iron bioavailability and do not affect non-heme iron bioavailability in humans. Food Funct. 2017, 8, 1994–1999. [Google Scholar] [CrossRef] [PubMed]
- Kortman, G.A.; Boleij, A.; Swinkels, D.W.; Tjalsma, H. Iron availability increases the pathogenic potential of Salmonella typhimurium and other enteric pathogens at the intestinal epithelial interface. PLoS ONE 2012, 7, e29968. [Google Scholar] [CrossRef] [PubMed]
- Bougle, D.; Vaghefi-Vaezzadeh, N.; Roland, N.; Bouvard, G.; Arhan, P.; Bureau, F.; Neuville, D.; Maubois, J.L. Influence of short-chain fatty acids on iron absorption by proximal colon. Scand. J. Gastroenterol. 2002, 37, 1008–1011. [Google Scholar] [CrossRef] [PubMed]
- Chlosta, S.; Fishman, D.S.; Harrington, L.; Johnson, E.E.; Knutson, M.D.; Wessling-Resnick, M.; Cherayil, B.J. The iron efflux protein ferroportin regulates the intracellular growth of Salmonella enterica. Infect. Immun. 2006, 74, 3065–3067. [Google Scholar] [CrossRef] [PubMed]
- Paradkar, P.N.; De Domenico, I.; Durchfort, N.; Zohn, I.; Kaplan, J.; Ward, D.M. Iron depletion limits intracellular bacterial growth in macrophages. Blood 2008, 112, 866–874. [Google Scholar] [CrossRef] [PubMed]
- Olakanmi, O.; Schlesinger, L.S.; Britigan, B.E. Hereditary hemochromatosis results in decreased iron acquisition and growth by Mycobacterium tuberculosis within human macrophages. J. Leukoc. Biol. 2007, 81, 195–204. [Google Scholar] [CrossRef] [PubMed]
- Pagani, A.; Nai, A.; Corna, G.; Bosurgi, L.; Rovere-Querini, P.; Camaschella, C.; Silvestri, L. Low hepcidin accounts for the proinflammatory status associated with iron deficiency. Blood 2011, 118, 736–746. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, K.K.; Chen, G.Y.; Schieber, A.M.P.; Redford, S.E.; Shokhirev, M.N.; Leblanc, M.; Lee, Y.M.; Ayres, J.S. Cooperative Metabolic Adaptations in the Host Can Favor Asymptomatic Infection and Select for Attenuated Virulence in an Enteric Pathogen. Cell 2018. [Google Scholar] [CrossRef] [PubMed]
- Tamboli, C.P.; Neut, C.; Desreumaux, P.; Colombel, J.F. Dysbiosis in inflammatory bowel disease. Gut 2004, 53, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Kaur, N.; Chen, C.C.; Luther, J.; Kao, J.Y. Intestinal dysbiosis in inflammatory bowel disease. Gut Microbes 2011, 2, 211–216. [Google Scholar] [CrossRef] [PubMed]
- Kamada, N.; Seo, S.U.; Chen, G.Y.; Nunez, G. Role of the gut microbiota in immunity and inflammatory disease. Nat. Rev. Immunol. 2013, 13, 321–335. [Google Scholar] [CrossRef] [PubMed]
- Xavier, R.J.; Podolsky, D.K. Unravelling the pathogenesis of inflammatory bowel disease. Nature 2007, 448, 427–434. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. The Global Prevalence of Anaemia in 2011; WHO Press: Geneva, Switzarland, 2015. [Google Scholar]
- Dignass, A.U.; Gasche, C.; Bettenworth, D.; Birgegard, G.; Danese, S.; Gisbert, J.P.; Gomollon, F.; Iqbal, T.; Katsanos, K.; Koutroubakis, I.; et al. European consensus on the diagnosis and management of iron deficiency and anaemia in inflammatory bowel diseases. J. Crohns Colitis 2015, 9, 211–222. [Google Scholar] [CrossRef] [PubMed]
- Cherayil, B.J. Cross-talk between iron homeostasis and intestinal inflammation. Gut Microbes 2010, 1, 65–69. [Google Scholar] [CrossRef] [PubMed]
- Gasche, C.; Lomer, M.C.; Cavill, I.; Weiss, G. Iron, anaemia, and inflammatory bowel diseases. Gut 2004, 53, 1190–1197. [Google Scholar] [CrossRef] [PubMed]
- Lomer, M.C.E.; Cook, W.B.; Jan-Mohamed, H.J.B.; Hutchinson, C.; Liu, D.Y.; Hider, R.C.; Powell, J.J. Iron requirements based upon iron absorption tests are poorly predicted by haematological indices in patients with inactive inflammatory bowel disease. Br. J. Nutr. 2012, 107, 1806–1811. [Google Scholar] [CrossRef] [PubMed]
- Carrier, J.C.; Aghdassi, E.; Jeejeebhoy, K.; Allard, J.P. Exacerbation of dextran sulfate sodium-induced colitis by dietary iron supplementation: Role of NF-kappa B. Int. J. Colorectal. Dis. 2006, 21, 381–387. [Google Scholar] [CrossRef] [PubMed]
- Erichsen, K.; Milde, A.M.; Arslan, G.; Helgeland, L.; Gudbrandsen, O.A.; Ulvik, R.J.; Berge, R.K.; Hausken, T.; Berstad, A. Low-dose oral ferrous fumarate aggravated intestinal inflammation in rats with DSS-induced colitis. Inflamm. Bowel Dis. 2005, 11, 744–748. [Google Scholar] [CrossRef] [PubMed]
- Reifen, R.; Matas, Z.; Zeidel, L.; Berkovitch, Z.; Bujanover, Y. Iron supplementation may aggravate inflammatory status of colitis in a rat model. Digest. Dis. Sci. 2000, 45, 394–397. [Google Scholar] [CrossRef] [PubMed]
- Seril, D.N.; Liao, J.; Ho, K.L.K.; Warsi, A.; Yang, C.S.; Yang, G.Y. Dietary iron supplementation enhances DSS-induced colitis and associated colorectal carcinoma development in mice. Digest. Dis. Sci. 2002, 47, 1266–1278. [Google Scholar] [CrossRef] [PubMed]
- Werner, T.; Wagner, S.J.; Martinez, I.; Walter, J.; Chang, J.S.; Clavel, T.; Kisling, S.; Schuemann, K.; Haller, D. Depletion of luminal iron alters the gut microbiota and prevents Crohn’s disease-like ileitis. Gut 2011, 60, 325–333. [Google Scholar] [CrossRef] [PubMed]
- Gevers, D.; Kugathasan, S.; Denson, L.A.; Vazquez-Baeza, Y.; Van Treuren, W.; Ren, B.Y.; Schwager, E.; Knights, D.; Song, S.J.; Yassour, M.; et al. The Treatment-Naive Microbiome in New-Onset Crohn’s Disease. Cell Host Microbe 2014, 15, 382–392. [Google Scholar] [CrossRef] [PubMed]
- Juillerat, P.; Yilmaz, B.; Wiest, R.; Rogler, G.; Macpherson, A.J. The clinical determinants affect gut microbial profile of inflammatory bowel disease patients. J. Crohns Colitis 2018, 12, S14. [Google Scholar] [CrossRef]
- Morgan, X.C.; Tickle, T.L.; Sokol, H.; Gevers, D.; Devaney, K.L.; Ward, D.V.; Reyes, J.A.; Shah, S.A.; LeLeiko, N.; Snapper, S.B.; et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 2012, 13. [Google Scholar] [CrossRef] [PubMed]
- Norman, J.M.; Handley, S.A.; Baldridge, M.T.; Droit, L.; Liu, C.Y.; Keller, B.C.; Kambal, A.; Monaco, C.L.; Zhao, G.; Fleshner, P.; et al. Disease-Specific Alterations in the Enteric Virome in Inflammatory Bowel Disease. Cell 2015, 160, 447–460. [Google Scholar] [CrossRef] [PubMed]
- Schreiner, P.; Yilmaz, B.; Franc, Y.; Rossel, J.B.; Misselwitz, B.; Scharl, M.; Zeitz, J.; Frei, P.; Greuter, T.; Vavricka, S.; et al. Vegetarian and gluten-free diet in patients with IBD-associated with a different microbiota compared with omnivore IBD patients. J. Crohns Colitis 2018, 12, S549. [Google Scholar] [CrossRef]
- Sokol, H.; Leducq, V.; Aschard, H.; Pham, H.P.; Jegou, S.; Landman, C.; Cohen, D.; Liguori, G.; Bourrier, A.; Nion-Larmurier, I.; et al. Fungal microbiota dysbiosis in IBD. Gut 2017, 66, 1039–1048. [Google Scholar] [CrossRef] [PubMed]
- Willing, B.P.; Dicksved, J.; Halfvarson, J.; Andersson, A.F.; Lucio, M.; Zheng, Z.; Jarnerot, G.; Tysk, C.; Jansson, J.K.; Engstrand, L. A Pyrosequencing Study in Twins Shows That Gastrointestinal Microbial Profiles Vary With Inflammatory Bowel Disease Phenotypes. Gastroenterology 2010, 139, 1844–1854. [Google Scholar] [CrossRef] [PubMed]
- Zhu, A.; Kaneshiro, M.; Kaunitz, J.D. Evaluation and treatment of iron deficiency anemia: A gastroenterological perspective. Dig. Dis. Sci. 2010, 55, 548–559. [Google Scholar] [CrossRef] [PubMed]
- Kangaspunta, M.; Haapamaki, J.; Farkkila, M.; Arkkila, P. Inflammatory bowel disease and anemia: Intravenous iron treatment. Scand. J. Gastroenterol. 2018, 53, 430–434. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.; Clavel, T.; Smirnov, K.; Schmidt, A.; Lagkouvardos, I.; Walker, A.; Lucio, M.; Michalke, B.; Schmitt-Kopplin, P.; Fedorak, R.; et al. Oral versus intravenous iron replacement therapy distinctly alters the gut microbiota and metabolome in patients with IBD. Gut 2017, 66, 863–871. [Google Scholar] [CrossRef] [PubMed]
- Graf, E.; Eaton, J.W. Suppression of colonic cancer by dietary phytic acid. Nutr. Cancer 1993, 19, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Graf, E.; Eaton, J.W. Dietary suppression of colonic cancer. Fiber or phytate? Cancer 1985, 56, 717–718. [Google Scholar] [CrossRef]
- Nelson, R.L. Dietary iron and colorectal cancer risk. Free Radic. Biol. Med. 1992, 12, 161–168. [Google Scholar] [CrossRef]
- Ashmore, J.H.; Rogers, C.J.; Kelleher, S.L.; Lesko, S.M.; Hartman, T.J. Dietary Iron and Colorectal Cancer Risk: A Review of Human Population Studies. Crit. Rev. Food Sci. Nutr. 2016, 56, 1012–1020. [Google Scholar] [CrossRef] [PubMed]
- Babbs, C.F. Free radicals and the etiology of colon cancer. Free Radic. Biol. Med. 1990, 8, 191–200. [Google Scholar] [CrossRef]
- Nelson, R.L. Iron and colorectal cancer risk: Human studies. Nutr. Rev. 2001, 59, 140–148. [Google Scholar] [CrossRef] [PubMed]
- Nelson, R.L.; Davis, F.G.; Sutter, E.; Sobin, L.H.; Kikendall, J.W.; Bowen, P. Body iron stores and risk of colonic neoplasia. J. Natl. Cancer Inst. 1994, 86, 455–460. [Google Scholar] [CrossRef] [PubMed]
- Bird, C.L.; Witte, J.S.; Swendseid, M.E.; Shikany, J.M.; Hunt, I.F.; Frankl, H.D.; Lee, E.R.; Longnecker, M.P.; Haile, R.W. Plasma ferritin, iron intake, and the risk of colorectal polyps. Am. J. Epidemiol. 1996, 144, 34–41. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.H.; Anderson, K.E.; Harnack, L.J.; Folsom, A.R.; Jacobs, D.R., Jr. Heme iron, zinc, alcohol consumption, and colon cancer: Iowa Women’s Health Study. J. Natl. Cancer Inst. 2004, 96, 403–407. [Google Scholar] [CrossRef] [PubMed]
- Larsson, S.C.; Adami, H.O.; Giovannucci, E.; Wolk, A. Re: Heme iron, zinc, alcohol consumption, and risk of colon cancer. J. Natl. Cancer Inst. 2005, 97, 232–233. [Google Scholar] [CrossRef] [PubMed]
- Kabat, G.C.; Miller, A.B.; Jain, M.; Rohan, T.E. A cohort study of dietary iron and heme iron intake and risk of colorectal cancer in women. Br. J. Cancer 2007, 97, 118–122. [Google Scholar] [CrossRef] [PubMed]
- Balder, H.F.; Vogel, J.; Jansen, M.C.; Weijenberg, M.P.; van den Brandt, P.A.; Westenbrink, S.; van der Meer, R.; Goldbohm, R.A. Heme and chlorophyll intake and risk of colorectal cancer in the Netherlands cohort study. Cancer Epidemiol. Biomarkers Prev. 2006, 15, 717–725. [Google Scholar] [CrossRef] [PubMed]
- Cross, A.J.; Ferrucci, L.M.; Risch, A.; Graubard, B.I.; Ward, M.H.; Park, Y.; Hollenbeck, A.R.; Schatzkin, A.; Sinha, R. A large prospective study of meat consumption and colorectal cancer risk: An investigation of potential mechanisms underlying this association. Cancer Res. 2010, 70, 2406–2414. [Google Scholar] [CrossRef] [PubMed]
- Nelson, R.L.; Persky, V.; Turyk, M. Determination of factors responsible for the declining incidence of colorectal cancer. Dis. Colon. Rectum 1999, 42, 741–752. [Google Scholar] [CrossRef] [PubMed]
- Sobhani, I.; Tap, J.; Roudot-Thoraval, F.; Roperch, J.P.; Letulle, S.; Langella, P.; Corthier, G.; Tran Van Nhieu, J.; Furet, J.P. Microbial dysbiosis in colorectal cancer (CRC) patients. PLoS ONE 2011, 6, e16393. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, J.K.; Holmes, E.; Kinross, J.; Burcelin, R.; Gibson, G.; Jia, W.; Pettersson, S. Host-gut microbiota metabolic interactions. Science 2012, 336, 1262–1267. [Google Scholar] [CrossRef] [PubMed]
- Huycke, M.M.; Gaskins, H.R. Commensal bacteria, redox stress, and colorectal cancer: Mechanisms and models. Exp. Biol. Med. 2004, 229, 586–597. [Google Scholar] [CrossRef]
- Gold, J.S.; Bayar, S.; Salem, R.R. Association of Streptococcus bovis bacteremia with colonic neoplasia and extracolonic malignancy. Arch. Surg. 2004, 139, 760–765. [Google Scholar] [CrossRef] [PubMed]
- McIntosh, G.H.; Royle, P.J.; Playne, M.J. A probiotic strain of L. acidophilus reduces DMH-induced large intestinal tumors in male Sprague-Dawley rats. Nutr. Cancer 1999, 35, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Kot, E.; Bezkorovainy, A. Binding of ferric iron to the cell walls and membranes of Bifidobacterium thermophilum: Effect of free radicals. J. Agric. Food Chem. 1999, 47, 4606–4610. [Google Scholar] [CrossRef] [PubMed]
- Vipperla, K.; O’Keefe, S.J. The microbiota and its metabolites in colonic mucosal health and cancer risk. Nutr. Clin. Pract. 2012, 27, 624–635. [Google Scholar] [CrossRef] [PubMed]
- Engle, S.J.; Ormsby, I.; Pawlowski, S.; Boivin, G.P.; Croft, J.; Balish, E.; Doetschman, T. Elimination of colon cancer in germ-free transforming growth factor beta 1-deficient mice. Cancer Res. 2002, 62, 6362–6366. [Google Scholar] [PubMed]
- Feng, Q.; Liang, S.; Jia, H.; Stadlmayr, A.; Tang, L.; Lan, Z.; Zhang, D.; Xia, H.; Xu, X.; Jie, Z.; et al. Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat. Commun. 2015, 6, 6528. [Google Scholar] [CrossRef] [PubMed]
- Girelli, D.; Ugolini, S.; Busti, F.; Marchi, G.; Castagna, A. Modern iron replacement therapy: Clinical and pathophysiological insights. Int. J. Hematol. 2018, 107, 16–30. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yilmaz, B.; Li, H. Gut Microbiota and Iron: The Crucial Actors in Health and Disease. Pharmaceuticals 2018, 11, 98. https://doi.org/10.3390/ph11040098
Yilmaz B, Li H. Gut Microbiota and Iron: The Crucial Actors in Health and Disease. Pharmaceuticals. 2018; 11(4):98. https://doi.org/10.3390/ph11040098
Chicago/Turabian StyleYilmaz, Bahtiyar, and Hai Li. 2018. "Gut Microbiota and Iron: The Crucial Actors in Health and Disease" Pharmaceuticals 11, no. 4: 98. https://doi.org/10.3390/ph11040098
APA StyleYilmaz, B., & Li, H. (2018). Gut Microbiota and Iron: The Crucial Actors in Health and Disease. Pharmaceuticals, 11(4), 98. https://doi.org/10.3390/ph11040098




