2.1. RISC-V Processors
Several processors were considered for this work, namely microcontrollers with
ARM and
Xtensa (e.g., ESP32) based cores. However, due to energy constrains derived from the project’s application domain, an FPGA was targeted as the preferred hardware platform. The RISC-V Instruction Set Architecture (ISA) was first developed in 2010, at UC Berkeley, and has been gaining popularity in both academia and industry [
7]. The supported ISA is fundamental to a CPU, since, together with the compiler, it links the hardware and the software by mapping the high level software constructs into low level instructions that the CPU can and will execute [
8]. Compared with other architectures, the RISC-V provides a number of advantages that makes it especially attractive for development [
9]:
The open-source nature allows any interested entity access to the source Intellectual Property (IP) of the cores without any licensing issue;
Even though the development of the ISA is open-source, its major features are already well defined and stable, which also attracts software development;
Additional functionalities are available through a set of extensions, which are also well defined after stabilization;
Due to its modular nature, the ISA is suitable for both high performance and low power integrated circuit applications. Additionally, specialized application processors that feature dedicated accelerators are also supported by the ISA.
This rising support is mainly due to it’s open-source characteristic and the advantages listed above, which enables developers to create their own cores following the RISC-V ISA guidelines. However, the other core attributes must be defined by the developer. Therefore, a wide range of different memory interfaces (examples:
AXI and
Wishbone) and configurations is used to add peripherals to the core, access memories or control execution. One processor can use separate AXI interfaces to access instruction or data memories, while another core (i.e., processor) uses a single Wishbone bus to access both [
9]. This heterogeneity complicates the choice of the right processor for a specific use-case, among all the options core landscape. Due to the complexity of the implementations, many cores, although supported by variety of toolchains, will not work out-of-the-box in the specific user target platform. In the scope of this work we focused in open-source, FPGA-optimized implementations. Other requirements were introduce in order to narrow down the number of cores explored, such as:
As some singularities of ASIC-targeted designs, for instance the memory generator usage, might make them unusable for an FPGA implementation, these kind of designs were discouraged for this work;
Cores must be able to support the recent versions of the ISA in order to be targeted by the recent versions of the software toolchain.
Following these criteria, some notable cores have been summarized in
Table 1. This table displays some open-source RISC-V processors and some of the most important features when selecting a processor for a given application.
In [
9], the authors performed a thorough evaluation of the performance and efficiency of some of the cores listed in
Table 1. Their selection of cores for a close evaluation consists of:
Piccolo and Flute,
Orca,
PicoRV32,
SweRV,
Taiga and
VexRiscv, also, as a “gold standard”, they used a 5 and a 8 stage proprietary cores. As these cores have configurations that differ from one another, the authors were careful to make the same configurations for all cores, allowing a fair comparison between them (e.g., cache was disabled for all that support this feature). To evaluate them in real hardware, the authors also used the so called TaPaSCo [
11], a system-on-chip generator for FPGAs, to easily generate bitstreams for a variety of platforms without having to deal with low level integration details. The hardware used consisted in an array of Xilinx made FPGA platforms: (1) AU250; (2) PYNQ; (3) VC709; (4) ZCU102. The chosen evaluation metrics were
Single Core Performance and, since the target hardware are FPGAs,
Resource Utilization.
To evaluate the performance, there are many widely used benchmarking options, most notably, there is the CoreMark and the DMIPS. For the latter, the results were quite surprising. The two proprietary cores (the authors did not mentioned their names nor their authors), which in theory had a high level of optimization, actually performed quite poorly when compared with some open source cores. With this benchmark, we can also observe that some cores have less Instructions Per Clock (IPC) but achieve reasonable results with higher working frequencies, such as the PicoRV32. The results were similar in both benchmarks; the most performing cores were the Taiga, VexRiscv and the Orca.
In terms of resource usage the results were also surprising. The cores that actually had a better performance were the ones that also consumed the least FPGA resources. Meaning that those cores, especially Taiga, Orca and VexRiscv are highly optimized in terms of performance and resource usage.
2.2. FPGA-Based CNN Accelerator
This section presents methodologies from previous works that aim to accelerate the inference of CNNs and recent approaches to ECG identification. The first section focuses on conventional CNNs and next section (
Section 2.3) specifically targets BCNNs and ECG identification.
FPGA-based CNN accelerators aim to improve inference performance by parallelizing the CNN forward propagation. There are 6 main ways to parallelize a CNN’s inference [
12]: intra convolution parallelism, inter convolution parallelism, inter feature map parallelism, intra feature map parallelism, inter layer parallelism and batch parallelism. This section will review previous works based on the implementation of 2D CNNs.
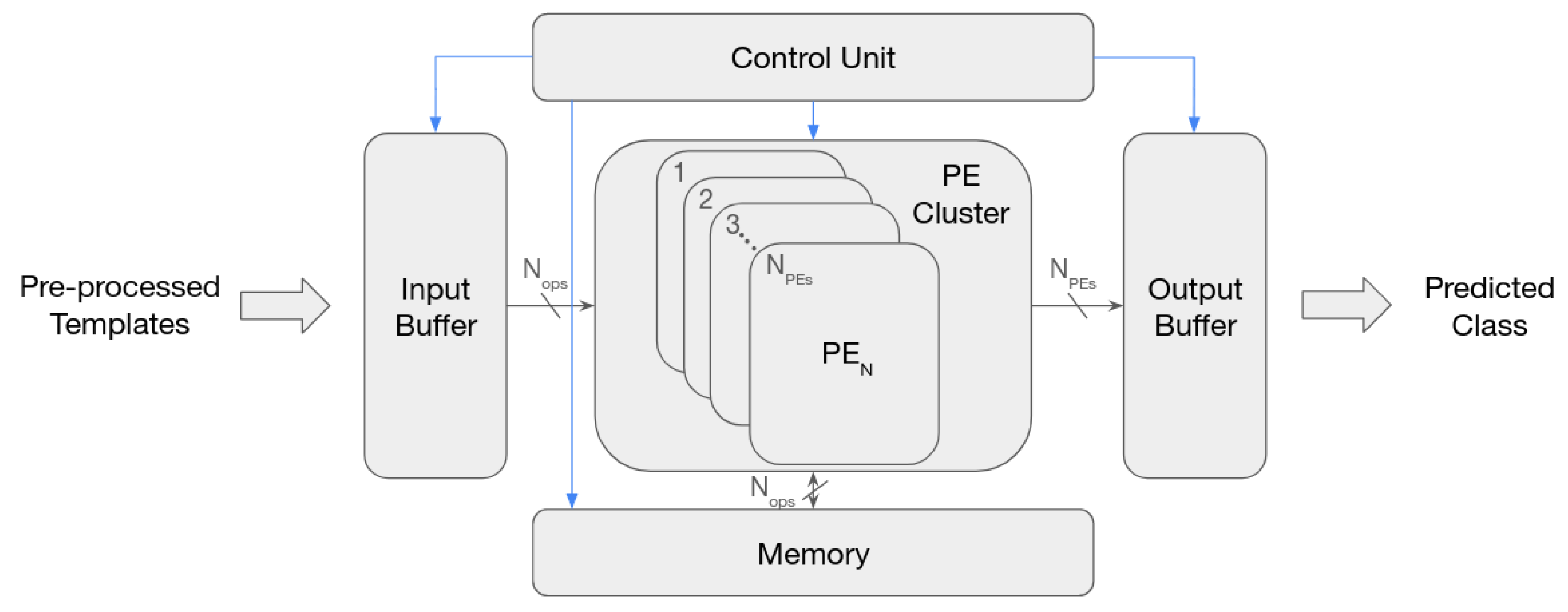
Some previous FPGA-based CNN accelerators [
13,
14,
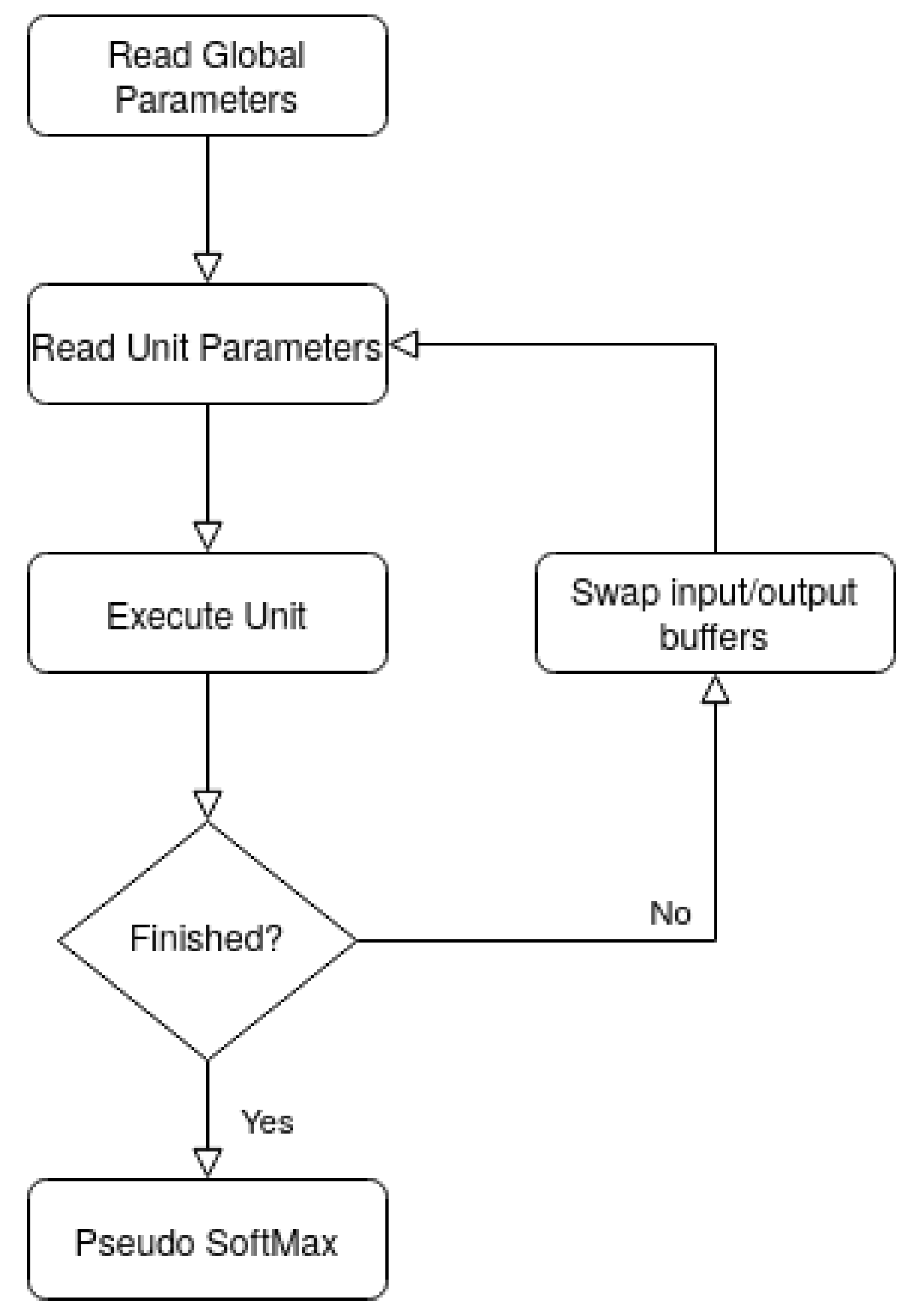
15] follow a standard architecture that entails: an external memory, used to hold the CNN’s parameters and input/output feature maps; an input and output buffer to cache the input and output feature maps; a Processing Element (PE) that processes the operations required to compute the outputs using the cached inputs and the weights that are maintained in a PE’s internal buffer; and a controller which controls the overall execution.
In works such as [
13,
14], a PE performs concurrent Multiply and Accumulate (MAC) operations to solve a single 2D convolution. This is an example of intra convolution parallelism.
The authors of [
13] take advantage of inter convolution parallelism. Their accelerator implements a PE that is capable of processing multiple input feature maps, by summing the result of multiple concurrent 2D convolutions with the objective of computing a single 3D convolution necessary to produce an element of a output feature map.
By having an array of PEs, where each PE is designed to compute a single output feature map at the time, the approaches found in [
13,
15,
16] are able to concurrently process multiple output feature maps, which is a form of inter feature map parallelism.
In [
17], the authors implemented all layers in a pipelined structure that enables the execution of all layers concurrently, while requiring a substantial amount of FPGA resources. This is an example of inter layer parallelism. In the same work, the authors propose a
divide and conquer strategy in the computation of fully connected layers that, if executed all at once, require a substantial amount of memory to hold all the operands. They divide the operation into multiple simple sub-convolutions, whose results can be accumulated to get the final result; this is an example of batch parallelism.
In [
18], the authors note that in modern CNN architecures, such as [
19], in deeper convolutional layers, the number of input/output channels surpass the actual input/output feature map size. In such CNNs, the authors argue that intra feature map parallelism is preferred over inter feature map parallelism.
2.3. BCNN-Based Optimization
Quantization of the network’s parameters is a popular practice to save memory usage and increase computational performance. The parameters are typically represented by 32-bit floating-point values, however, the hardware required to process floating-point data is much more complex and slower, comparatively to what is needed to handle integer data [
20]. An approach commonly found in the literature is to quantize floating point values to 8- or 16-bit integer values, but a more extreme quantization can be performed. Binary Neural Network (BNN)s, first proposed by Courbariaux et al. in [
20], introduced the concept of constraining the activations and the parameters of a Neural Network (NN) to either +1 or −1 [
20], allowing a 1-bit representation. This new quantization paradigm helps minimize the memory footprint.
FPGA implementations of BCNN follow a similar architecture as conventional CNNs. The main difference lies with the datapath structure of each PE, where the multipliers and adder trees seen in conventional CNN accelerators are replaced with logic that implements the
XNOR Dot Product (XNP) operation (described in [
20]). In [
21], the authors propose a pipelined PE datapath consisting of four stages:
XNOR,
popcount, accumulation and Batch Normalization (BN) + binarization. Works such as [
16,
18,
21,
22] employ intra and inter convolution parallelism, in addition to inter feature map parallelism.
The novel ECG-ID-BNet [
6], a BCNN that implements ECG classification, was used as the ECG identification method and it is detailed in
Table 2, where unit refers to a sequential stack of a convolutional/fully connected layer, a max pooling layer (if present) and a batch normalization layer. The BCNN was evaluated on real-world data from the Physionet Computing in Cardiology Challenge 2017 dataset [
23] (containing 8528 ECG recordings lasting from 9 to just over 60 seconds, acquired at the hand palms using the AliveCore (
https://www.alivecor.com/, accessed on: 1 October 2021) device and from our own dataset collected containing ECG data collected at the thighs using an experimental device integrated in a toilet seat cover [
4] (further described in [
6]).
From the AliveCore dataset we randomly selected ECG recordings classified as normal and with 60 s duration for 50 different subjects, while our dataset contains ECG recordings lasting approximately 180 seconds for 10 different subjects. The best results with ECG-ID-BNet were obtained using 4 convolutional and 1 fully-connected units, and it managed to achieve a
and
accuracy on the AliveCore dataset and our dataset, respectively. Comparatively with the state-of-the art [
24] these results exhibit similar accuracy, hence demonstrating that the BCNN approach does not degrade the quality of the recognition, with the advantage of improved computational performance.