Advances in Biosensors Technology for Detection and Characterization of Extracellular Vesicles
Abstract
:1. Introduction
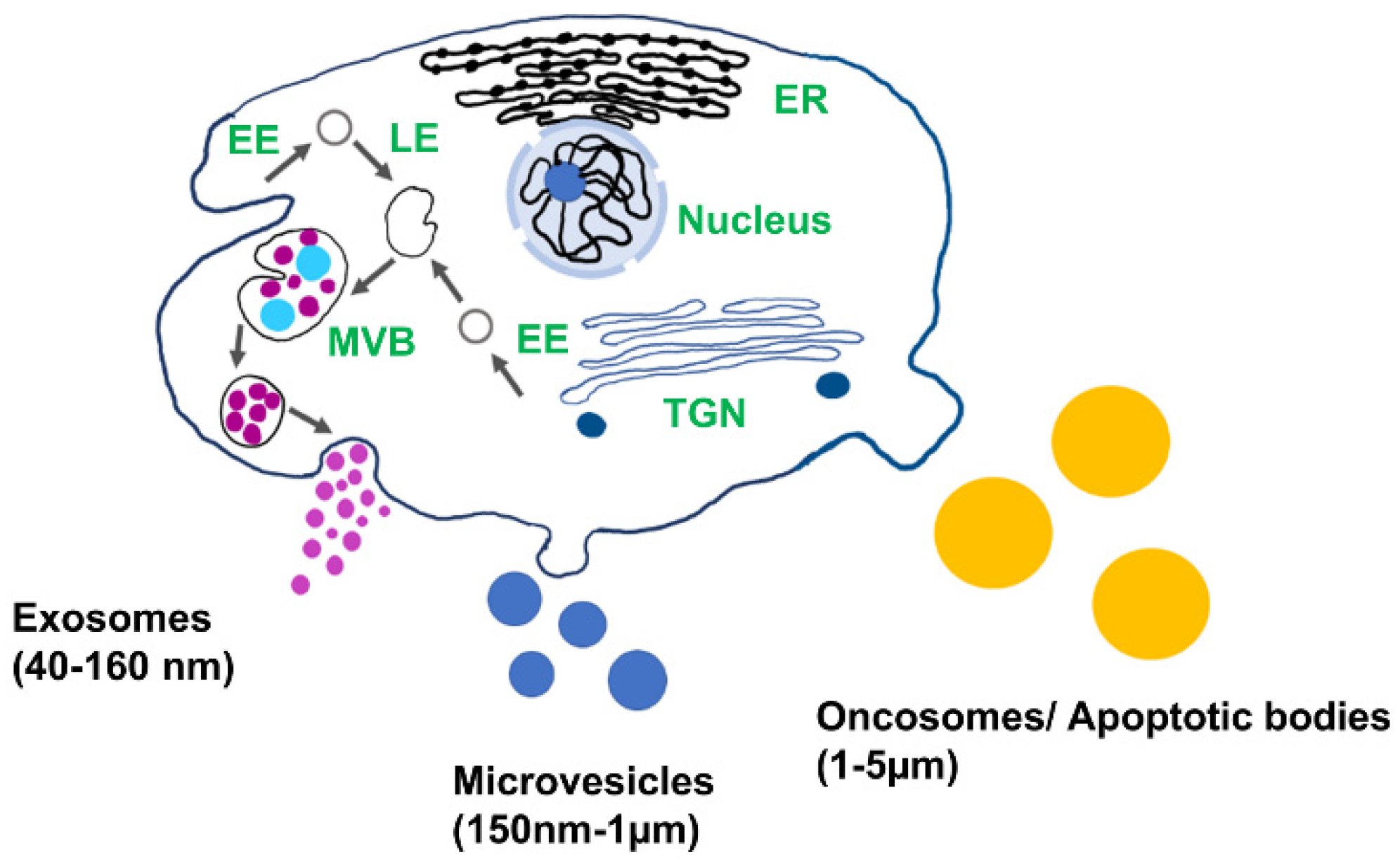
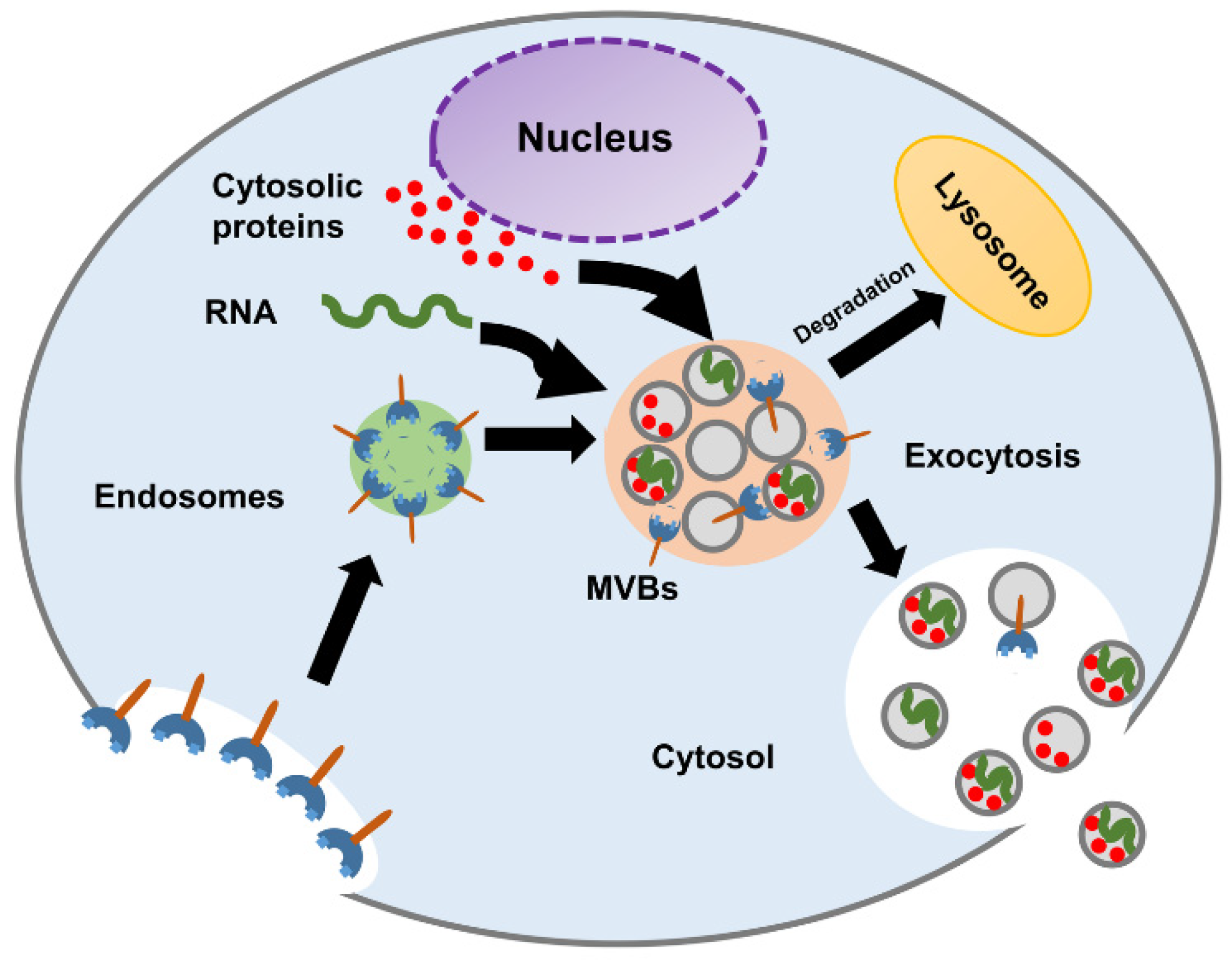
2. Exosome’s Biogenesis, Molecular Cargo, and Function
- MVBs fuse with lysosomes and degrade their content.
- MVBs fuse with the plasma membrane and release their content into the extracellular space.
3. Conventional Methods for Isolation and Detection of Exosomes
4. Sensors for Detection and Molecular Characterization of Exosomes
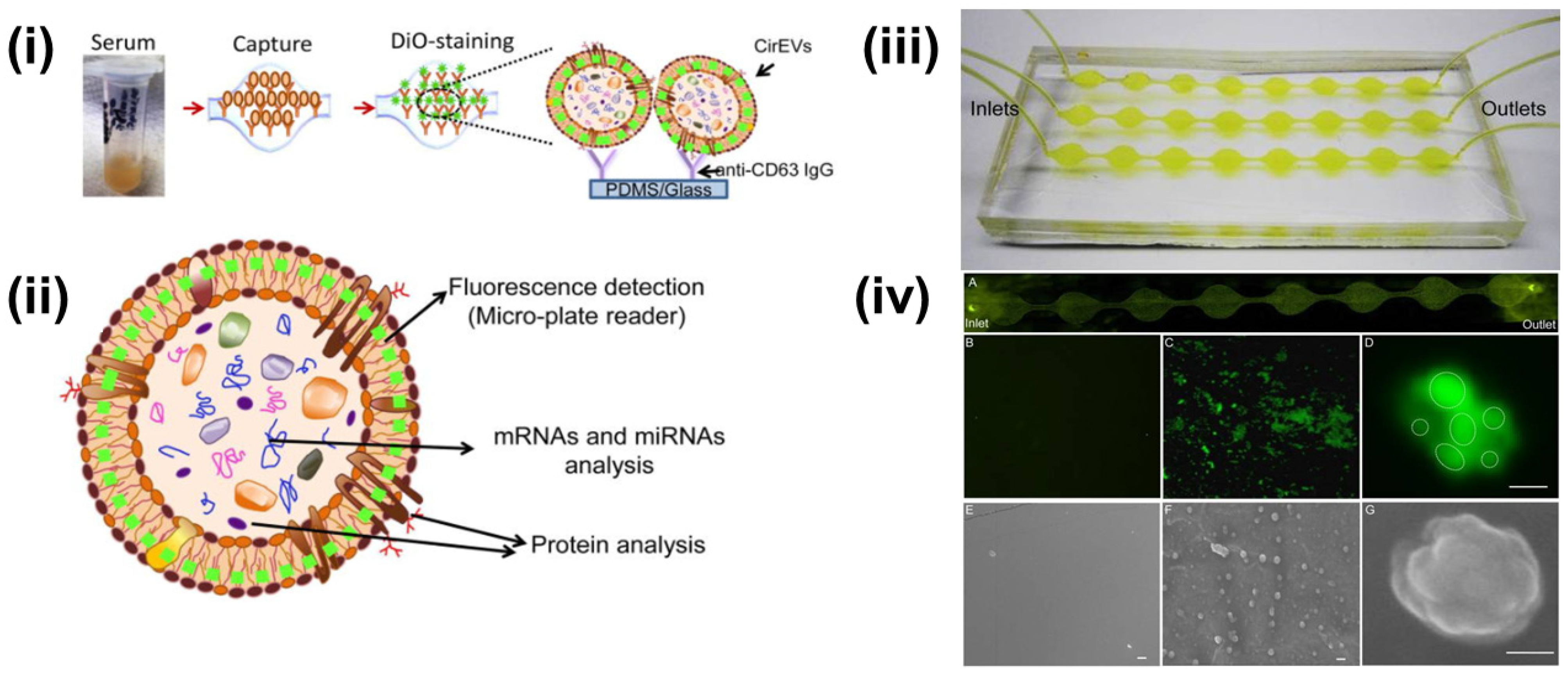
4.1. Fluorescence-Based Sensors
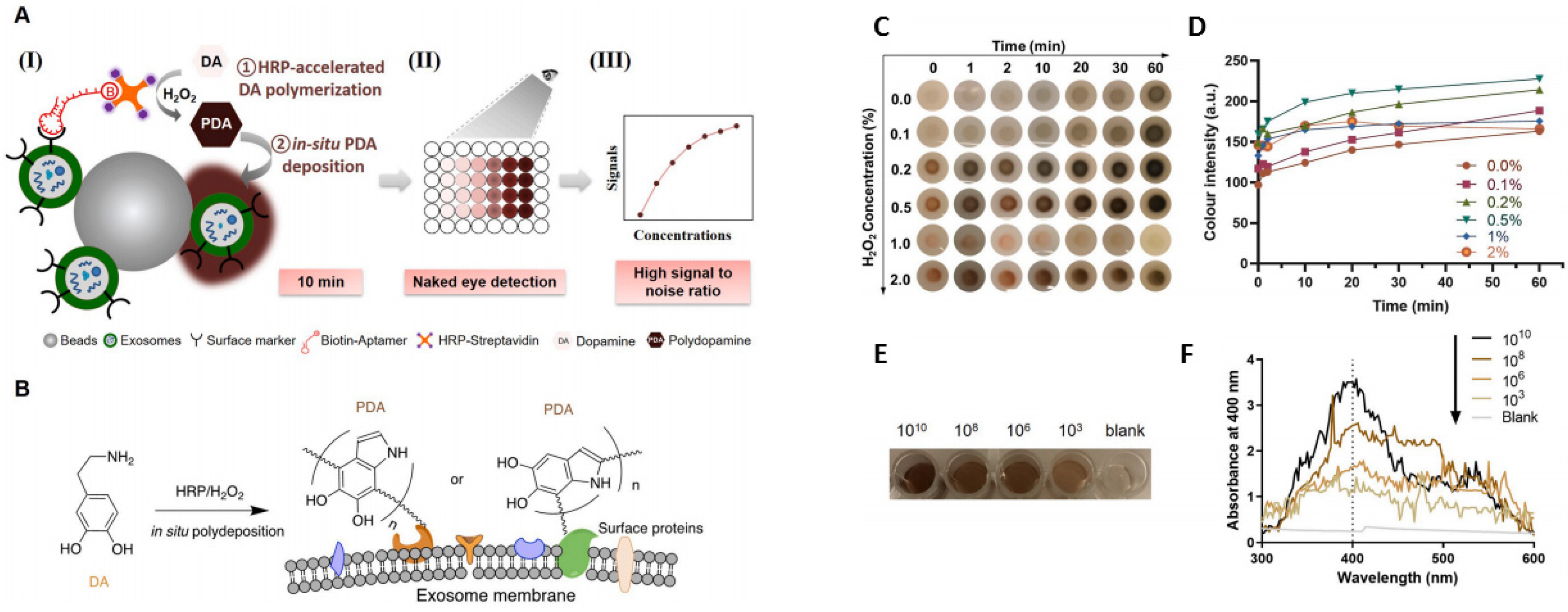
4.2. Colorimetric Sensors
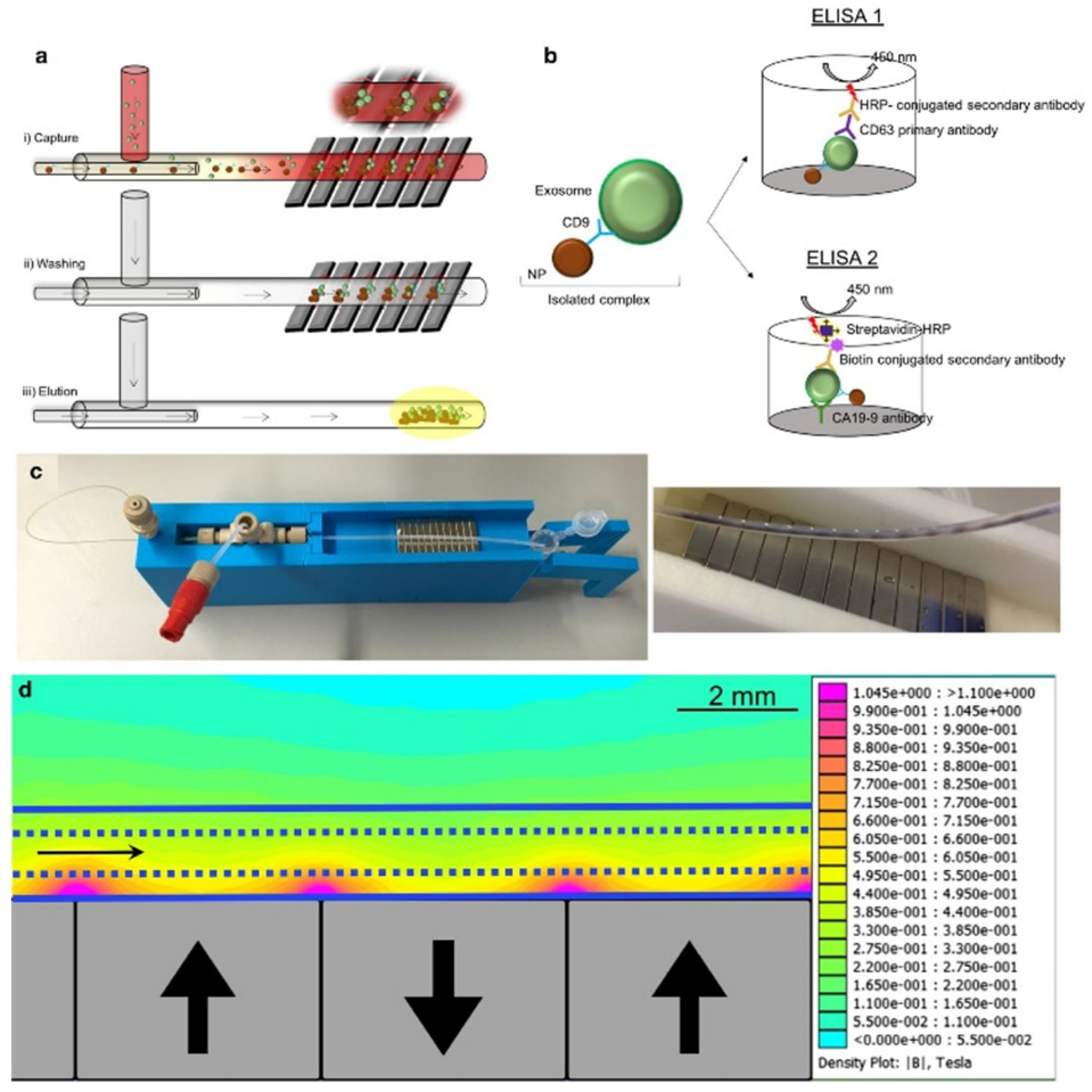
4.3. Magnetic Sensors
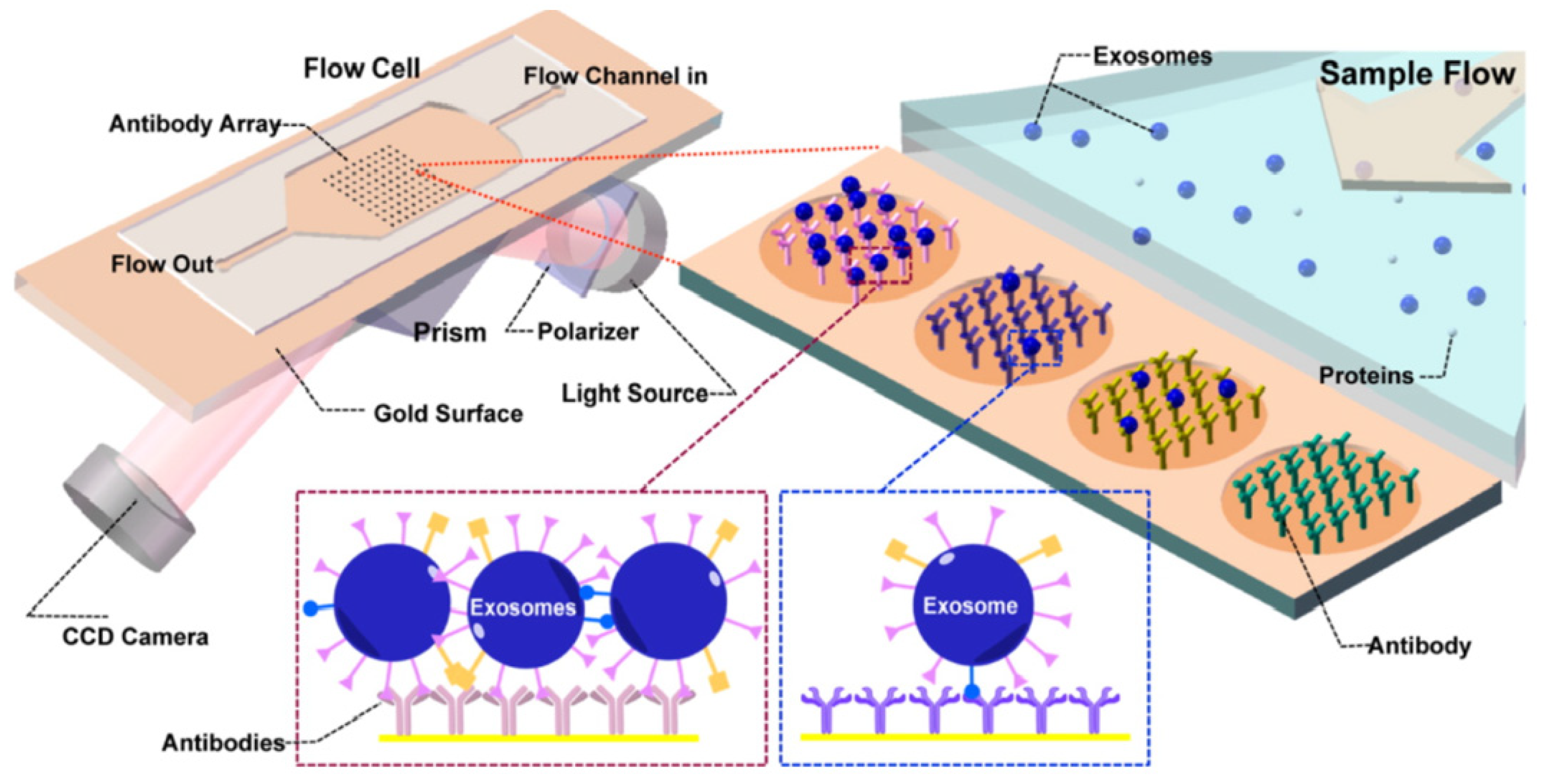
4.4. Surface Plasmon Resonance (SPR) Sensors
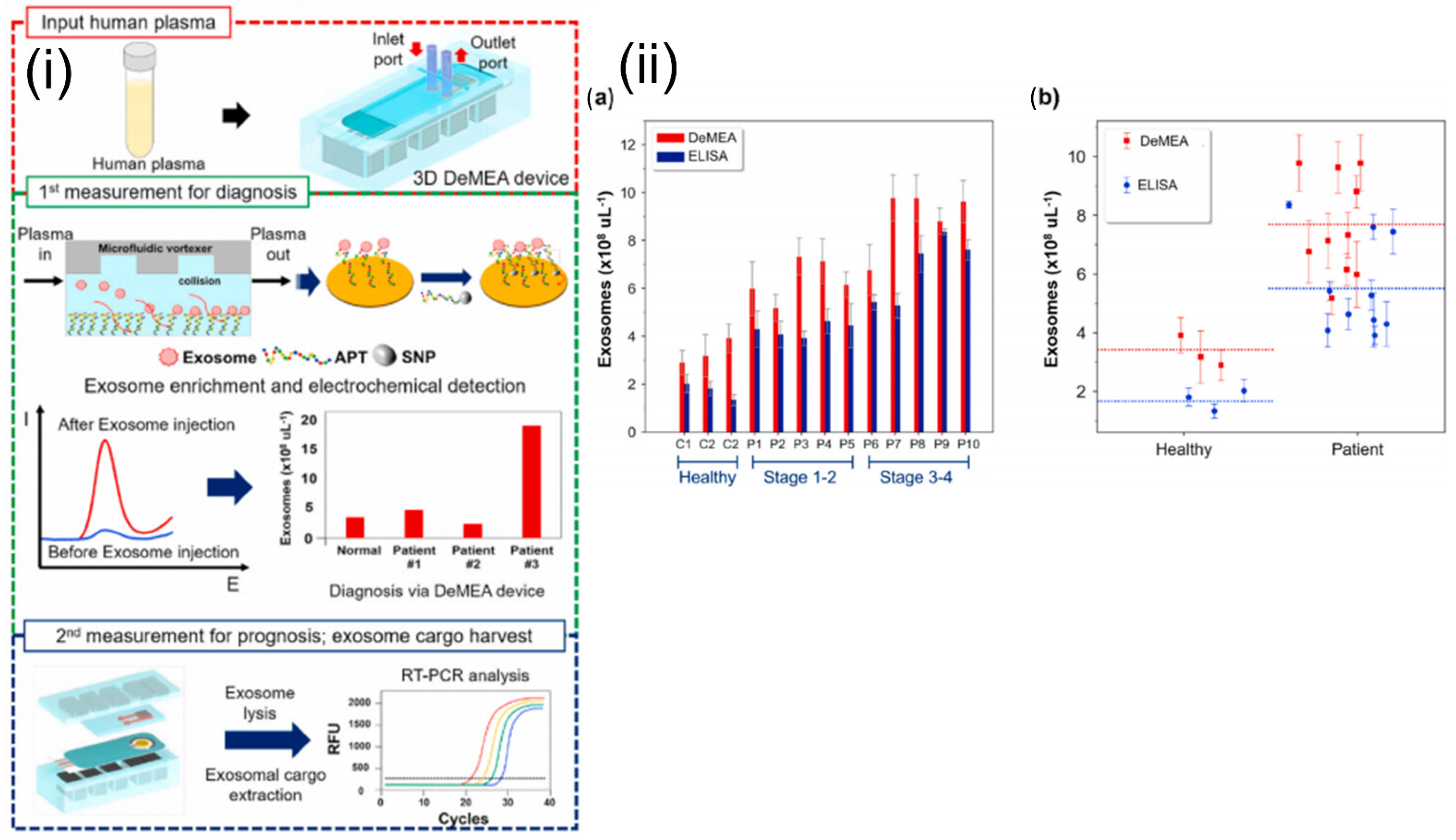
4.5. Electrochemical Sensors
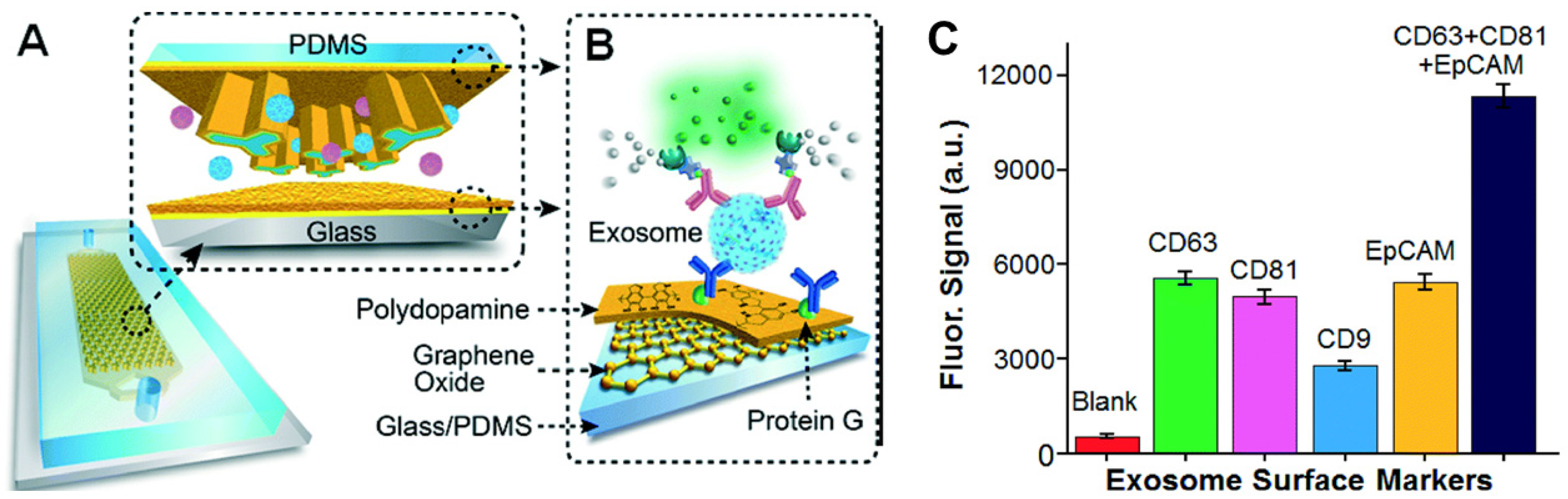
4.6. Immunoaffinity Sensors
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Colombo, M.; Moita, C.; van Niel, G.; Kowal, J.; Vigneron, J.; Benaroch, P.; Manel, N.; Moita, L.F.; Théry, C.; Raposo, G. Analysis of ESCRT functions in exosome biogenesis, composition and secretion highlights the heterogeneity of extracellular vesicles. J. Cell Sci. 2013, 126, 5553–5565. [Google Scholar] [CrossRef] [Green Version]
- Katsuda, T.; Kosaka, N.; Ochiya, T. The roles of extracellular vesicles in cancer biology: Toward the development of novel cancer biomarkers. Proteomics 2014, 14, 412–425. [Google Scholar] [CrossRef] [PubMed]
- EL Andaloussi, S.; Mäger, I.; Breakefield, X.O.; Wood, M.J.A. Extracellular vesicles: Biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 2013, 12, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Margolis, L.; Sadovsky, Y. The biology of extracellular vesicles: The known unknowns. PLoS Biol. 2019, 17, e3000363. [Google Scholar] [CrossRef] [PubMed]
- Szabo, G.; Momen-Heravi, F. Extracellular Vesicles and Exosomes. In The Liver; Wiley: Hoboken, NJ, USA, 2020; pp. 1022–1027. [Google Scholar]
- LeBleu, V.S.; Kalluri, R. Exosomes as a Multicomponent Biomarker Platform in Cancer. Trends Cancer 2020, 6, 767–774. [Google Scholar] [CrossRef]
- Pang, H.; Luo, S.; Xiao, Y.; Xia, Y.; Li, X.; Huang, G.; Xie, Z.; Zhou, Z. Emerging Roles of Exosomes in T1DM. Front. Immunol. 2020, 11, 3099. [Google Scholar] [CrossRef]
- Hejrati, A.; Hasani, B.; Esmaili, M.; Bashash, D.; Tavakolinia, N.; Zafari, P. Role of exosome in autoimmunity, with a particular emphasis on rheumatoid arthritis. Int. J. Rheum. Dis. 2021, 24, 159–169. [Google Scholar] [CrossRef] [PubMed]
- Cañas, J.A.; Sastre, B.; Rodrigo-Muñoz, J.M.; del Pozo, V. Exosomes: A new approach to asthma pathology. Clin. Chim. Acta 2019, 495, 139–147. [Google Scholar] [CrossRef]
- Wang, L.; Zhang, L. Circulating Exosomal miRNA as Diagnostic Biomarkers of Neurodegenerative Diseases. Front. Mol. Neurosci. 2020, 13, 53. [Google Scholar] [CrossRef] [Green Version]
- Kalluri, R.; LeBleu, V.S. The biology, function, and biomedical applications of exosomes. Science 2020, 367. [Google Scholar] [CrossRef]
- Lin, J.; Li, J.; Huang, B.; Liu, J.; Chen, X.; Chen, X.-M.; Xu, Y.-M.; Huang, L.-F.; Wang, X.-Z. Exosomes: Novel Biomarkers for Clinical Diagnosis. Sci. World J. 2015, 2015, 1–8. [Google Scholar] [CrossRef]
- Cooper, J.M.; Wiklander, P.B.O.; Nordin, J.Z.; Al-Shawi, R.; Wood, M.J.; Vithlani, M.; Schapira, A.H.V.; Simons, J.P.; El-Andaloussi, S.; Alvarez-Erviti, L. Systemic exosomal siRNA delivery reduced alpha-synuclein aggregates in brains of transgenic mice. Mov. Disord. 2014, 29, 1476–1485. [Google Scholar] [CrossRef] [Green Version]
- van Niel, G.; Charrin, S.; Simoes, S.; Romao, M.; Rochin, L.; Saftig, P.; Marks, M.S.; Rubinstein, E.; Raposo, G. The Tetraspanin CD63 Regulates ESCRT-Independent and -Dependent Endosomal Sorting during Melanogenesis. Dev. Cell 2011, 21, 708–721. [Google Scholar] [CrossRef] [Green Version]
- Conde-Vancells, J.; Rodriguez-Suarez, E.; Embade, N.; Gil, D.; Matthiesen, R.; Valle, M.; Elortza, F.; Lu, S.C.; Mato, J.M.; Falcon-Perez, J.M. Characterization and Comprehensive Proteome Profiling of Exosomes Secreted by Hepatocytes. J. Proteome Res. 2008, 7, 5157–5166. [Google Scholar] [CrossRef] [Green Version]
- Subra, C.; Grand, D.; Laulagnier, K.; Stella, A.; Lambeau, G.; Paillasse, M.; De Medina, P.; Monsarrat, B.; Perret, B.; Silvente-Poirot, S.; et al. Exosomes account for vesicle-mediated transcellular transport of activatable phospholipases and prostaglandins. J. Lipid Res. 2010, 51, 2105–2120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Witas, R.; Chaput, D.; Khan, H.; Stevens, S.M.; Kang, D. Isolation and Proteomic Analysis of Microvesicles and Exosomes from HT22 Cells and Primary Neurons. In Neuroproteomics; Humana Press: New York, NY, USA, 2017; pp. 255–267. [Google Scholar]
- Liang, B.; Peng, P.; Chen, S.; Li, L.; Zhang, M.; Cao, D.; Yang, J.; Li, H.; Gui, T.; Li, X.; et al. Characterization and proteomic analysis of ovarian cancer-derived exosomes. J. Proteom. 2013, 80, 171–182. [Google Scholar] [CrossRef]
- Théry, C.; Zitvogel, L.; Amigorena, S. Exosomes: Composition, biogenesis and function. Nat. Rev. Immunol. 2002, 2, 569–579. [Google Scholar] [CrossRef]
- Farooqi, A.A.; Desai, N.N.; Qureshi, M.Z.; Librelotto, D.R.N.; Gasparri, M.L.; Bishayee, A.; Nabavi, S.M.; Curti, V.; Daglia, M. Exosome biogenesis, bioactivities and functions as new delivery systems of natural compounds. Biotechnol. Adv. 2018, 36, 328–334. [Google Scholar] [CrossRef]
- Lin, S.; Yu, Z.; Chen, D.; Wang, Z.; Miao, J.; Li, Q.; Zhang, D.; Song, J.; Cui, D. Progress in Microfluidics-Based Exosome Separation and Detection Technologies for Diagnostic Applications. Small 2020, 16, 1903916. [Google Scholar] [CrossRef] [PubMed]
- Doyle, L.; Wang, M. Overview of Extracellular Vesicles, Their Origin, Composition, Purpose, and Methods for Exosome Isolation and Analysis. Cells 2019, 8, 727. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morita, E.; Sandrin, V.; Chung, H.-Y.; Morham, S.G.; Gygi, S.P.; Rodesch, C.K.; Sundquist, W.I. Human ESCRT and ALIX proteins interact with proteins of the midbody and function in cytokinesis. EMBO J. 2007, 26, 4215–4227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Géminard, C.; de Gassart, A.; Blanc, L.; Vidal, M. Degradation of AP2 During Reticulocyte Maturation Enhances Binding of Hsc70 and Alix to a Common Site on TfR for Sorting into Exosomes. Traffic 2004, 5, 181–193. [Google Scholar] [CrossRef] [PubMed]
- Witwer, K.W.; Buzás, E.I.; Bemis, L.T.; Bora, A.; Lässer, C.; Lötvall, J.; Nolte-‘t Hoen, E.N.; Piper, M.G.; Sivaraman, S.; Skog, J.; et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell. Vesicles 2013, 2, 20360. [Google Scholar] [CrossRef]
- Crescitelli, R.; Lässer, C.; Szabó, T.G.; Kittel, A.; Eldh, M.; Dianzani, I.; Buzás, E.I.; Lötvall, J. Distinct RNA profiles in subpopulations of extracellular vesicles: Apoptotic bodies, microvesicles and exosomes. J. Extracell. Vesicles 2013, 2, 20677. [Google Scholar] [CrossRef] [PubMed]
- Bullock, M.; Silva, A.; Kanlikilicer-Unaldi, P.; Filant, J.; Rashed, M.; Sood, A.; Lopez-Berestein, G.; Calin, G. Exosomal Non-Coding RNAs: Diagnostic, Prognostic and Therapeutic Applications in Cancer. Non-Coding RNA 2015, 1, 53–68. [Google Scholar] [CrossRef] [PubMed]
- Nahand, J.S.; Mahjoubin-Tehran, M.; Moghoofei, M.; Pourhanifeh, M.H.; Mirzaei, H.R.; Asemi, Z.; Khatami, A.; Bokharaei-Salim, F.; Mirzaei, H.; Hamblin, M.R. Exosomal miRNAs: Novel players in viral infection. Epigenomics 2020, 12, 353–370. [Google Scholar] [CrossRef] [PubMed]
- Mao, L.; Li, X.; Gong, S.; Yuan, H.; Jiang, Y.; Huang, W.; Sun, X.; Dang, X. Serum exosomes contain ECRG4 mRNA that suppresses tumor growth via inhibition of genes involved in inflammation, cell proliferation, and angiogenesis. Cancer Gene Ther. 2018, 25, 248–259. [Google Scholar] [CrossRef]
- Zhang, Y.; Cai, F.; Liu, J.; Chang, H.; Liu, L.; Yang, A.; Liu, X. Transfer RNA-derived fragments as potential exosome tRNA-derived fragment biomarkers for osteoporosis. Int. J. Rheum. Dis. 2018, 21, 1659–1669. [Google Scholar] [CrossRef]
- Möhrmann, L.; Huang, H.J.; Hong, D.S.; Tsimberidou, A.M.; Fu, S.; Piha-Paul, S.A.; Subbiah, V.; Karp, D.D.; Naing, A.; Krug, A.; et al. Liquid Biopsies Using Plasma Exosomal Nucleic Acids and Plasma Cell-Free DNA Compared with Clinical Outcomes of Patients with Advanced Cancers. Clin. Cancer Res. 2018, 24, 181–188. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tellez-Gabriel, M.; Knutsen, E.; Perander, M. Current Status of Circulating Tumor Cells, Circulating Tumor DNA, and Exosomes in Breast Cancer Liquid Biopsies. Int. J. Mol. Sci. 2020, 21, 9457. [Google Scholar] [CrossRef]
- Record, M.; Silvente-Poirot, S.; Poirot, M.; Wakelam, M.O. Extracellular vesicles: Lipids as key components of their biogenesis and functions. J. Lipid Res. 2018, 59, 1316–1324. [Google Scholar] [CrossRef] [Green Version]
- Skotland, T.; Hessvik, N.P.; Sandvig, K.; Llorente, A. Exosomal lipid composition and the role of ether lipids and phosphoinositides in exosome biology. J. Lipid Res. 2019, 60, 9–18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iliescu, F.; Vrtačnik, D.; Neuzil, P.; Iliescu, C. Microfluidic Technology for Clinical Applications of Exosomes. Micromachines 2019, 10, 392. [Google Scholar] [CrossRef] [Green Version]
- Bennit, H.R.F.; Gonda, A.; McMullen, J.R.W.; Kabagwira, J.; Wall, N.R. Peripheral Blood Cell Interactions of Cancer-Derived Exosomes Affect Immune Function. Cancer Microenviron. 2019, 12, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Kong, F.-L.; Wang, X.-P.; Li, Y.-N.; Wang, H.-X. The role of exosomes derived from cerebrospinal fluid of spinal cord injury in neuron proliferation in vitro. Artif. Cells Nanomed. Biotechnol. 2018, 46, 200–205. [Google Scholar] [CrossRef] [Green Version]
- Hou, R.; Li, Y.; Sui, Z.; Yuan, H.; Yang, K.; Liang, Z.; Zhang, L.; Zhang, Y. Advances in exosome isolation methods and their applications in proteomic analysis of biological samples. Anal. Bioanal. Chem. 2019, 411, 5351–5361. [Google Scholar] [CrossRef]
- Klingeborn, M.; Skiba, N.P.; Stamer, W.D.; Rickman, C.B. Isolation of Retinal Exosome Biomarkers from Blood by Targeted Immunocapture. In Retinal Degenerative Diseases; Springer: Berlin, Germany, 2019; pp. 21–25. [Google Scholar]
- Sharma, P.; Ludwig, S.; Muller, L.; Hong, C.S.; Kirkwood, J.M.; Ferrone, S.; Whiteside, T.L. Immunoaffinity-based isolation of melanoma cell-derived exosomes from plasma of patients with melanoma. J. Extracell. Vesicles 2018, 7, 1435138. [Google Scholar] [CrossRef] [PubMed]
- Cui, B.; Liu, C.; Yao, S. One-Step RT-PCR for Detection of Micrornas in Exosomes Using Droplet Microfluidics. In Proceedings of the 2020 IEEE 33rd International Conference on Micro Electro Mechanical Systems (MEMS), Vancouver, BC, Canada, 18–20 January 2020; pp. 142–146. [Google Scholar]
- Nwokwu, C.D.; Bari, S.M.I.; Hutson, K.H.; Brausell, C.; Nestorova, G.G. ExoPRIME: Solid-phase immunoisolation and OMICS analysis of surface-marker-specific exosomal subpopulations. Talanta 2022, 236, 122870. [Google Scholar] [CrossRef]
- Guerreiro, E.M.; Vestad, B.; Steffensen, L.A.; Aass, H.C.D.; Saeed, M.; Øvstebø, R.; Costea, D.E.; Galtung, H.K.; Søland, T.M. Efficient extracellular vesicle isolation by combining cell media modifications, ultrafiltration, and size-exclusion chromatography. PLoS ONE 2018, 13, e0204276. [Google Scholar] [CrossRef] [Green Version]
- Mitchell, M.D. Exosome enrichment by ultracentrifugation and size exclusion chromatography. Front. Biosci. 2018, 23, 4621. [Google Scholar] [CrossRef] [Green Version]
- Contreras-Naranjo, J.C.; Wu, H.-J.; Ugaz, V.M. Microfluidics for exosome isolation and analysis: Enabling liquid biopsy for personalized medicine. Lab Chip 2017, 17, 3558–3577. [Google Scholar] [CrossRef] [PubMed]
- Greening, D.W.; Xu, R.; Ji, H.; Tauro, B.J.; Simpson, R.J. A Protocol for Exosome Isolation and Characterization: Evaluation of Ultracentrifugation, Density-Gradient Separation, and Immunoaffinity Capture Methods. In Proteomic Profiling; Humana Press: New York, NY, USA, 2015; pp. 179–209. [Google Scholar]
- Lobb, R.; Möller, A. Size Exclusion Chromatography: A Simple and Reliable Method for Exosome Purification. In Extracellular Vesicles; Humana Press: New York, NY, USA, 2017; pp. 105–110. [Google Scholar]
- Navajas, R.; Corrales, F.J.; Paradela, A. Serum Exosome Isolation by Size-Exclusion Chromatography for the Discovery and Validation of Preeclampsia-Associated Biomarkers. In Proteomics for Biomarker Discovery; Humana Press: New York, NY, USA, 2019; pp. 39–50. [Google Scholar]
- Soares Martins, T.; Catita, J.; Martins Rosa, I.; Odete, A.B.; da Cruz e Silva, O.; Henriques, A.G. Exosome isolation from distinct biofluids using precipitation and column-based approaches. PLoS ONE 2018, 13, e0198820. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xiao, D.; Ohlendorf, J.; Chen, Y.; Taylor, D.D.; Rai, S.N.; Waigel, S.; Zacharias, W.; Hao, H.; McMasters, K.M. Identifying mRNA, MicroRNA and Protein Profiles of Melanoma Exosomes. PLoS ONE 2012, 7, e46874. [Google Scholar] [CrossRef] [Green Version]
- Alvarez, M.L.; Khosroheidari, M.; Kanchi Ravi, R.; DiStefano, J.K. Comparison of protein, microRNA, and mRNA yields using different methods of urinary exosome isolation for the discovery of kidney disease biomarkers. Kidney Int. 2012, 82, 1024–1032. [Google Scholar] [CrossRef] [Green Version]
- He, L.; Zhu, D.; Wang, J.; Wu, X. A highly efficient method for isolating urinary exosomes. Int. J. Mol. Med. 2018, 43, 83–90. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Theodoraki, M.; Hong, C.; Donnenberg, V.S.; Donnenberg, A.D.; Whiteside, T.L. Evaluation of Exosome Proteins by on-Bead Flow Cytometry. Cytom. Part A 2021, 99, 372–381. [Google Scholar] [CrossRef]
- van der Vlist, E.J.; Nolte-’t Hoen, E.N.M.; Stoorvogel, W.; Arkesteijn, G.J.A.; Wauben, M.H.M. Fluorescent labeling of nano-sized vesicles released by cells and subsequent quantitative and qualitative analysis by high-resolution flow cytometry. Nat. Protoc. 2012, 7, 1311–1326. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Pertierra, E.; Oliveira-Rodríguez, M.; Matos, M.; Gutiérrez, G.; Moyano, A.; Salvador, M.; Rivas, M.; Blanco-López, M.C. Extracellular Vesicles: Current Analytical Techniques for Detection and Quantification. Biomolecules 2020, 10, 824. [Google Scholar] [CrossRef]
- Wang, C.; Ding, Q.; Plant, P.; Basheer, M.; Yang, C.; Tawedrous, E.; Krizova, A.; Boulos, C.; Farag, M.; Cheng, Y.; et al. Droplet digital PCR improves urinary exosomal miRNA detection compared to real-time PCR. Clin. Biochem. 2019, 67, 54–59. [Google Scholar] [CrossRef]
- Cho, S.M.; Shin, S.; Kim, Y.; Song, W.; Hong, S.G.; Jeong, S.H.; Kang, M.S.; Lee, K.A. A novel approach for tuberculosis diagnosis using exosomal DNA and droplet digital PCR. Clin. Microbiol. Infect. 2020, 26, e1–e942. [Google Scholar] [CrossRef]
- Anderson, W.; Lane, R.; Korbie, D.; Trau, M. Observations of Tunable Resistive Pulse Sensing for Exosome Analysis: Improving System Sensitivity and Stability. Langmuir 2015, 31, 6577–6587. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, M.; Shimizu, K.; Rahman, M.; Ishikawa, H.; Takase, H.; Ugawa, S.; Okada, A.; Inoshima, Y. Efficient method for isolation of exosomes from raw bovine milk. Drug Dev. Ind. Pharm. 2019, 45, 359–364. [Google Scholar] [CrossRef] [PubMed]
- Dou, D.; Ren, X.; Han, M.; Xu, X.; Ge, X.; Gu, Y.; Wang, X. Cancer-Associated Fibroblasts-Derived Exosomes Suppress Immune Cell Function in Breast Cancer via the miR-92/PD-L1 Pathway. Front. Immunol. 2020, 11, 2026. [Google Scholar] [CrossRef]
- Liu, C.; Guo, J.; Tian, F.; Yang, N.; Yan, F.; Ding, Y.; Wei, J.; Hu, G.; Nie, G.; Sun, J. Field-Free Isolation of Exosomes from Extracellular Vesicles by Microfluidic Viscoelastic Flows. ACS Nano 2017, 11, 6968–6976. [Google Scholar] [CrossRef] [Green Version]
- Gaillard, M.; Thuaire, A.; Nonglaton, G.; Agache, V.; Roupioz, Y.; Raillon, C. Biosensing extracellular vesicles: Contribution of biomolecules in affinity-based methods for detection and isolation. Analyst 2020, 145, 1997–2013. [Google Scholar] [CrossRef] [PubMed]
- Gurunathan, S.; Kang, M.-H.; Jeyaraj, M.; Qasim, M.; Kim, J.-H. Review of the Isolation, Characterization, Biological Function, and Multifarious Therapeutic Approaches of Exosomes. Cells 2019, 8, 307. [Google Scholar] [CrossRef] [Green Version]
- Wang, W.; Wu, C.; Jin, H. Exosomes in chronic inflammatory skin diseases and skin tumors. Exp. Dermatol. 2019, 28, 213–218. [Google Scholar] [CrossRef]
- Wang, J.; Ma, P.; Kim, D.H.; Liu, B.-F.; Demirci, U. Towards microfluidic-based exosome isolation and detection for tumor therapy. Nano Today 2021, 37, 101066. [Google Scholar] [CrossRef]
- Le, M.-C.N.; Fan, H.Z. Exosome isolation using nanostructures and microfluidic devices. Biomed. Mater. 2021, 16. [Google Scholar] [CrossRef]
- Lin, B.; Lei, Y.; Wang, J.; Zhu, L.; Wu, Y.; Zhang, H.; Wu, L.; Zhang, P.; Yang, C. Microfluidic-Based Exosome Analysis for Liquid Biopsy. Small Methods 2021, 5, 2001131. [Google Scholar] [CrossRef]
- Zhang, H.; Freitas, D.; Kim, H.S.; Fabijanic, K.; Li, Z.; Chen, H.; Mark, M.T.; Molina, H.; Martin, A.B.; Bojmar, L.; et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat. Cell Biol. 2018, 20, 332–343. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Vermesh, O.; Mani, V.; Ge, T.J.; Madsen, S.J.; Sabour, A.; Hsu, E.-C.; Gowrishankar, G.; Kanada, M.; Jokerst, J.V.; et al. The Exosome Total Isolation Chip. ACS Nano 2017, 11, 10712–10723. [Google Scholar] [CrossRef]
- Woo, H.-K.; Sunkara, V.; Park, J.; Kim, T.-H.; Han, J.-R.; Kim, C.-J.; Choi, H.-I.; Kim, Y.-K.; Cho, Y.-K. Exodisc for Rapid, Size-Selective, and Efficient Isolation and Analysis of Nanoscale Extracellular Vesicles from Biological Samples. ACS Nano 2017, 11, 1360–1370. [Google Scholar] [CrossRef]
- Yasui, T.; Yanagida, T.; Ito, S.; Konakade, Y.; Takeshita, D.; Naganawa, T.; Nagashima, K.; Shimada, T.; Kaji, N.; Nakamura, Y.; et al. Unveiling massive numbers of cancer-related urinary-microRNA candidates via nanowires. Sci. Adv. 2017, 3, e1701133. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Z.; Yang, Y.; Yamaguchi, H.; Hung, M.-C.; Kameoka, J. Isolation of cancer-derived extracellular vesicle subpopulations by a size-selective microfluidic platform. Biomicrofluidics 2020, 14, 034113. [Google Scholar] [CrossRef] [PubMed]
- Kanwar, S.S.; Dunlay, C.J.; Simeone, D.M.; Nagrath, S. Microfluidic device (ExoChip) for on-chip isolation, quantification and characterization of circulating exosomes. Lab Chip 2014, 14, 1891–1900. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Pan, W.; Liu, C.; Guo, J.; Shen, J.; Feng, J.; Luo, T.; Situ, B.; Zhang, Y.; An, T.; et al. Homogenous Magneto-Fluorescent Nanosensor for Tumor-Derived Exosome Isolation and Analysis. ACS Sens. 2020, 5, 2052–2060. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Zhang, L.; Cui, M.; Su, Y.; He, Y. Rapid and Accurate Detection of Lymph Node Metastases Enabled through Fluorescent Silicon Nanoparticles-Based Exosome Probes. Anal. Chem. 2021, 93, 10122–10131. [Google Scholar] [CrossRef] [PubMed]
- Vaidyanathan, R.; Naghibosadat, M.; Rauf, S.; Korbie, D.; Carrascosa, L.G.; Shiddiky, M.J.A.; Trau, M. Detecting Exosomes Specifically: A Multiplexed Device Based on Alternating Current Electrohydrodynamic Induced Nanoshearing. Anal. Chem. 2014, 86, 11125–11132. [Google Scholar] [CrossRef] [Green Version]
- Xu, L.; Chopdat, R.; Li, D.; Al-Jamal, K.T. Development of a simple, sensitive and selective colorimetric aptasensor for the detection of cancer-derived exosomes. Biosens. Bioelectron. 2020, 169, 112576. [Google Scholar] [CrossRef]
- Zhang, Y.; Jiao, J.; Wei, Y.; Wang, D.; Yang, C.; Xu, Z. Plasmonic Colorimetric Biosensor for Sensitive Exosome Detection via Enzyme-Induced Etching of Gold Nanobipyramid@MnO 2 Nanosheet Nanostructures. Anal. Chem. 2020, 92, 15244–15252. [Google Scholar] [CrossRef]
- Issadore, D.; Min, C.; Liong, M.; Chung, J.; Weissleder, R.; Lee, H. Miniature magnetic resonance system for point-of-care diagnostics. Lab Chip 2011, 11, 2282. [Google Scholar] [CrossRef]
- Shao, H.; Chung, J.; Balaj, L.; Charest, A.; Bigner, D.D.; Carter, B.S.; Hochberg, F.H.; Breakefield, X.O.; Weissleder, R.; Lee, H. Protein typing of circulating microvesicles allows real-time monitoring of glioblastoma therapy. Nat. Med. 2012, 18, 1835–1840. [Google Scholar] [CrossRef]
- Sancho-Albero, M.; Sebastián, V.; Sesé, J.; Pazo-Cid, R.; Mendoza, G.; Arruebo, M.; Martín-Duque, P.; Santamaría, J. Isolation of exosomes from whole blood by a new microfluidic device: Proof of concept application in the diagnosis and monitoring of pancreatic cancer. J. Nanobiotechnol. 2020, 18, 150. [Google Scholar] [CrossRef]
- de Almeida, M.P.; Pereira, E.; Baptista, P.; Gomes, I.; Figueiredo, S.; Soares, L.; Franco, R. Gold Nanoparticles as (Bio) Chemical Sensors. Comp. Anal. Chem. 2014, 66, 529–567. [Google Scholar]
- Zeng, S.; Baillargeat, D.; Ho, H.-P.; Yong, K.-T. Nanomaterials enhanced surface plasmon resonance for biological and chemical sensing applications. Chem. Soc. Rev. 2014, 43, 3426. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Wang, K.; Cui, J.; Liu, H.; Bu, X.; Ma, H.; Wang, W.; Gong, H.; Lausted, C.; Hood, L.; et al. Label-Free Quantitative Detection of Tumor-Derived Exosomes through Surface Plasmon Resonance Imaging. Anal. Chem. 2014, 86, 8857–8864. [Google Scholar] [CrossRef] [Green Version]
- Chen, W.; Li, J.; Wei, X.; Fan, Y.; Qian, H.; Li, S.; Xiang, Y.; Ding, S. Surface plasmon resonance biosensor using hydrogel-AuNP supramolecular spheres for determination of prostate cancer-derived exosomes. Microchim. Acta 2020, 187, 590. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Shoaie, N.; Jahanpeyma, F.; Zhao, J.; Azimzadeh, M.; Al−Jamal, K.T. Optical, electrochemical and electrical (nano)biosensors for detection of exosomes: A comprehensive overview. Biosens. Bioelectron. 2020, 161, 112222. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Rahimian, A.; Son, K.; Shin, D.-S.; Patel, T.; Revzin, A. Development of an aptasensor for electrochemical detection of exosomes. Methods 2016, 97, 88–93. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kashefi-Kheyrabadi, L.; Kim, J.; Chakravarty, S.; Park, S.; Gwak, H.; Kim, S.-I.; Mohammadniaei, M.; Lee, M.-H.; Hyun, K.-A.; Jung, H.-I. Detachable microfluidic device implemented with electrochemical aptasensor (DeMEA) for sequential analysis of cancerous exosomes. Biosens. Bioelectron. 2020, 169, 112622. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Skog, J.; Hsu, C.-H.; Lessard, R.T.; Balaj, L.; Wurdinger, T.; Carter, B.S.; Breakefield, X.O.; Toner, M.; Irimia, D. Microfluidic isolation and transcriptome analysis of serum microvesicles. Lab Chip 2010, 10, 505–511. [Google Scholar] [CrossRef] [Green Version]
- Zhang, P.; He, M.; Zeng, Y. Ultrasensitive microfluidic analysis of circulating exosomes using a nanostructured graphene oxide/polydopamine coating. Lab Chip 2016, 16, 3033–3042. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Tong, X.; Yang, L.; Yin, R.; Li, Y.; Zeng, D.; Wang, X.; Deng, K. A herringbone mixer based microfluidic device HBEXO-chip for purifying tumor-derived exosomes and establishing miRNA signature in pancreatic cancer. Sens. Actuators B Chem. 2021, 332, 129511. [Google Scholar] [CrossRef]
- Ishraq Bari, S.M.; Reis, L.G.; Holland, T.; Nestorova, G.G. Numerical analysis of optimal design parameters for a cell co-culture microfluidic platform with an integrated pressure-controlled valve. In Proceedings of the ASME International Mechanical Engineering Congress and Exposition, Online, 16–19 November 2020. [Google Scholar]
- Xu, H.; Liao, C.; Zuo, P.; Liu, Z.; Ye, B.-C. Magnetic-Based Microfluidic Device for On-Chip Isolation and Detection of Tumor-Derived Exosomes. Anal. Chem. 2018, 90, 13451–13458. [Google Scholar] [CrossRef] [PubMed]
| Isolation Technique | Isolation Principle | Appraisal Parameters | Advantages | Disadvantages | References | ||
|---|---|---|---|---|---|---|---|
| Time | Yield | Purity | |||||
| Ultracentrifugation | Density-based | * | * | ** | The gold standard | Time-consuming; low yield; moderate purity; | [38,46] |
| Immunoaffinity-capture | Affinity-based | ** | ** | *** | Highly specific and pure | Expensive; strict pH condition; constrained use | [40,41] |
| Size-exclusion chromatography | Vesicles Size | ** | ** | ** | Simple, rapid, moderate yield | Poor specificity; scaling problems | [44,47,48] |
| Polymer-based precipitation | Surface charge-based | * | *** | * | Simple and user-friendly, high yield | Expensive; low specificity; poor purity; scaling problems | [22,49] |
| Ultrafiltration | Molecular weight and size-based | ** | * | ** | Simple, no specific instrument | Clogging; low yield, low specificity, time-consuming | [50,51,52] |
| Sensor Type | Isolation Principle | Minimum Detection Limit | References |
|---|---|---|---|
| Fluorescence | Quantification of fluorescence signal | 6.56 × 107 EVs mL−1 | [74] |
| Colorimetric | Absorbance measurement of the colorimetric solution | 1.35 × 105 EVs mL−1 | [78] |
| Magnetic | Nuclear magnetic resonance | 4.39 × 103 EVs mL−1 | [93] |
| Surface Plasmon Resonance (SPR) | Resonant oscillation of the electrons stimulated by the incident light at the interface between a negative and a positive dielectric constant material | 105 EVs mL−1 | [85] |
| Electrochemical | Measurement of the electric current caused by the oxidation or reduction reactions | 106 EVs mL−1 | [87] |
| Immunoaffinity | Affinity-based isolation | 24 × 106 EVs mL−1 | [42] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bari, S.M.I.; Hossain, F.B.; Nestorova, G.G. Advances in Biosensors Technology for Detection and Characterization of Extracellular Vesicles. Sensors 2021, 21, 7645. https://doi.org/10.3390/s21227645
Bari SMI, Hossain FB, Nestorova GG. Advances in Biosensors Technology for Detection and Characterization of Extracellular Vesicles. Sensors. 2021; 21(22):7645. https://doi.org/10.3390/s21227645
Chicago/Turabian StyleBari, Saif Mohammad Ishraq, Faria Binte Hossain, and Gergana G. Nestorova. 2021. "Advances in Biosensors Technology for Detection and Characterization of Extracellular Vesicles" Sensors 21, no. 22: 7645. https://doi.org/10.3390/s21227645
APA StyleBari, S. M. I., Hossain, F. B., & Nestorova, G. G. (2021). Advances in Biosensors Technology for Detection and Characterization of Extracellular Vesicles. Sensors, 21(22), 7645. https://doi.org/10.3390/s21227645






