EventDTW: An Improved Dynamic Time Warping Algorithm for Aligning Biomedical Signals of Nonuniform Sampling Frequencies
Abstract
1. Introduction
1.1. DTW for Different Sampling Frequencies
1.2. Existing DTW Improvement Methods
1.3. Existing DTW Evaluation Metrics
2. Materials and Methods
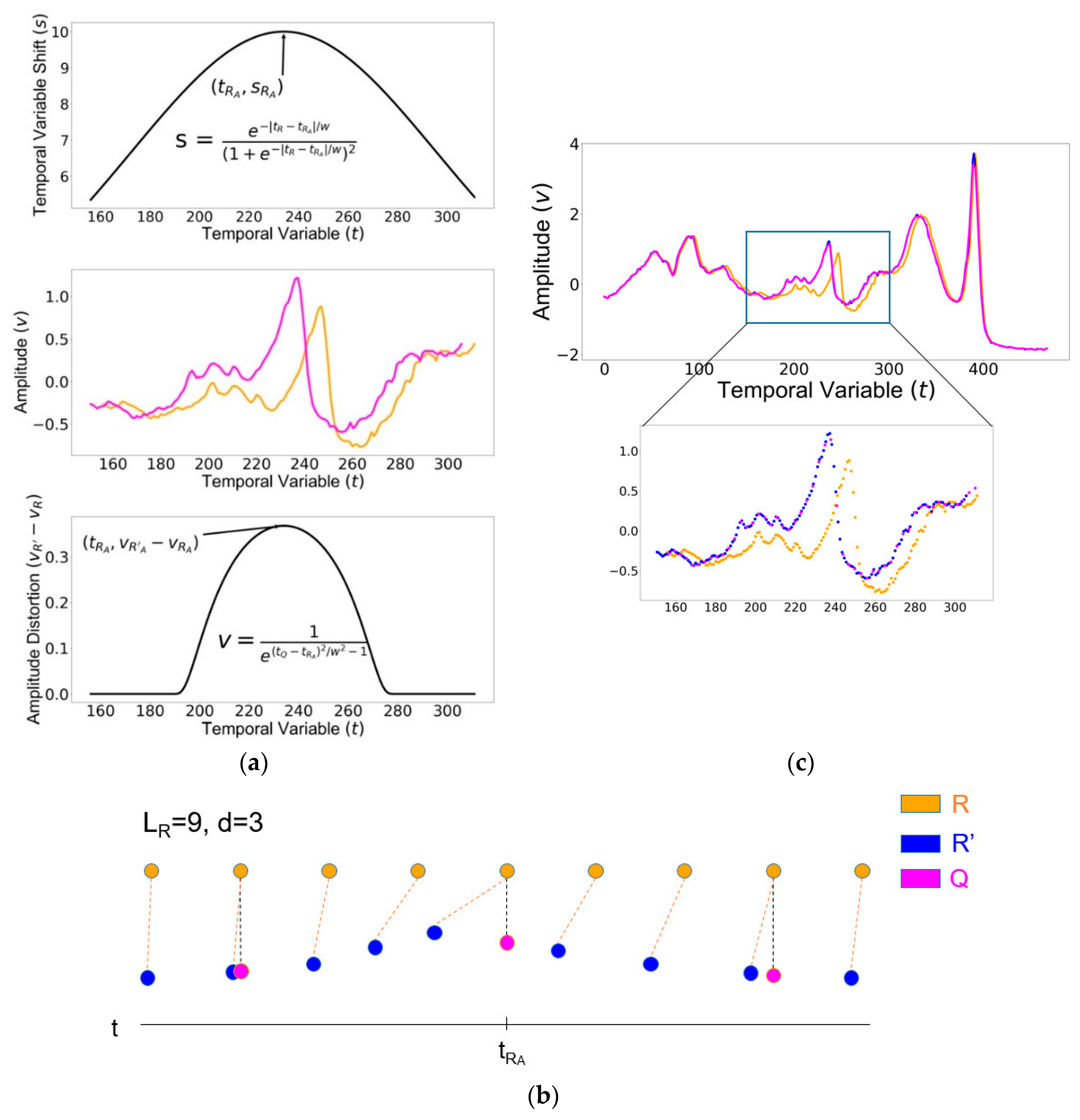
2.1. Companion Signal Generation
2.1.1. Generating
2.1.2. Generating
2.2. Calculation of the Optimal Alignment
2.3. Design of Error Rate Metric
2.4. Design of the Singularity Score
2.5. EventDTW
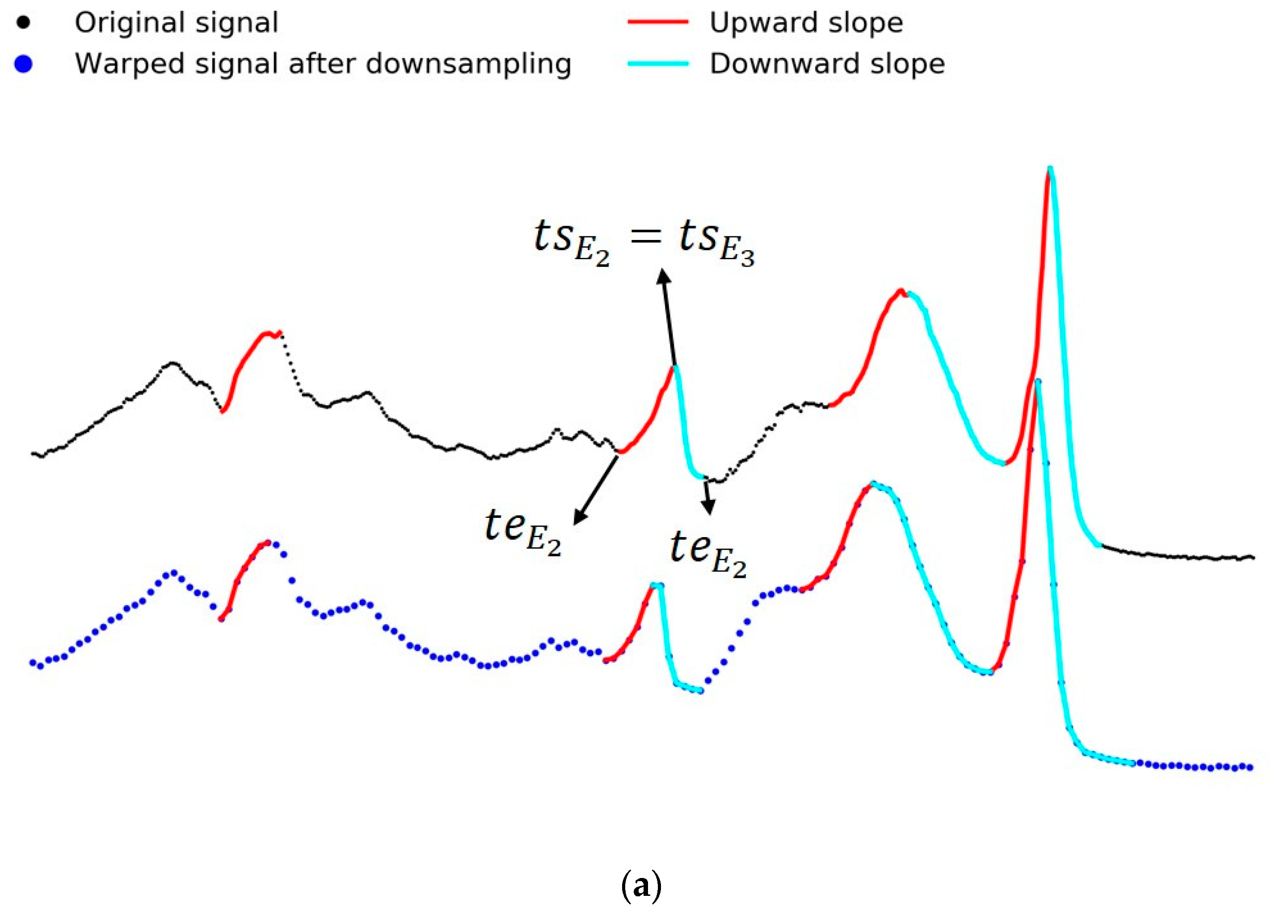
2.5.1. Event Detection and Selection
- Select an entire upslope with monotonically increasing observations in the signal. An upslope is defined such that it is a contiguous subset of observations starting with the lowest point and continuously increasing until there is a decrease or stagnation from one point to another.
- Select the key slopes. We selected the slopes with the highest 20% elevation in based on empirical trials. Elevation is defined as the product of the number of observations in a slope and the amplitude difference between the maximum point and the minimum point of the slope.
- Every upslope in should match an upslope in based on proximity in timestamps. If a slope in is not matched, it is excluded.
- Set the peak of the upslope as an event and propagate the information along the slope as per below.
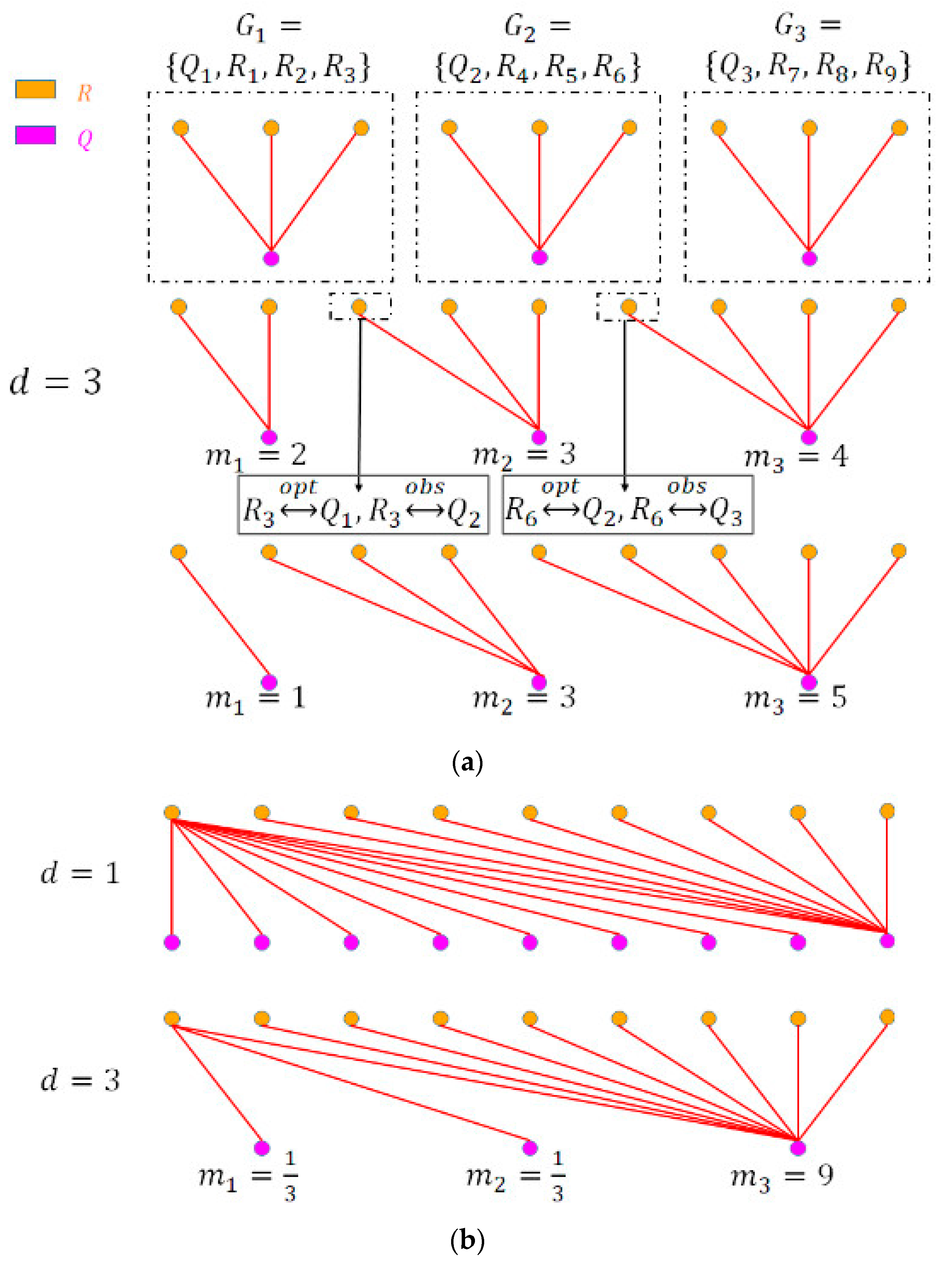
2.5.2. Event Information Propagation
3. Results
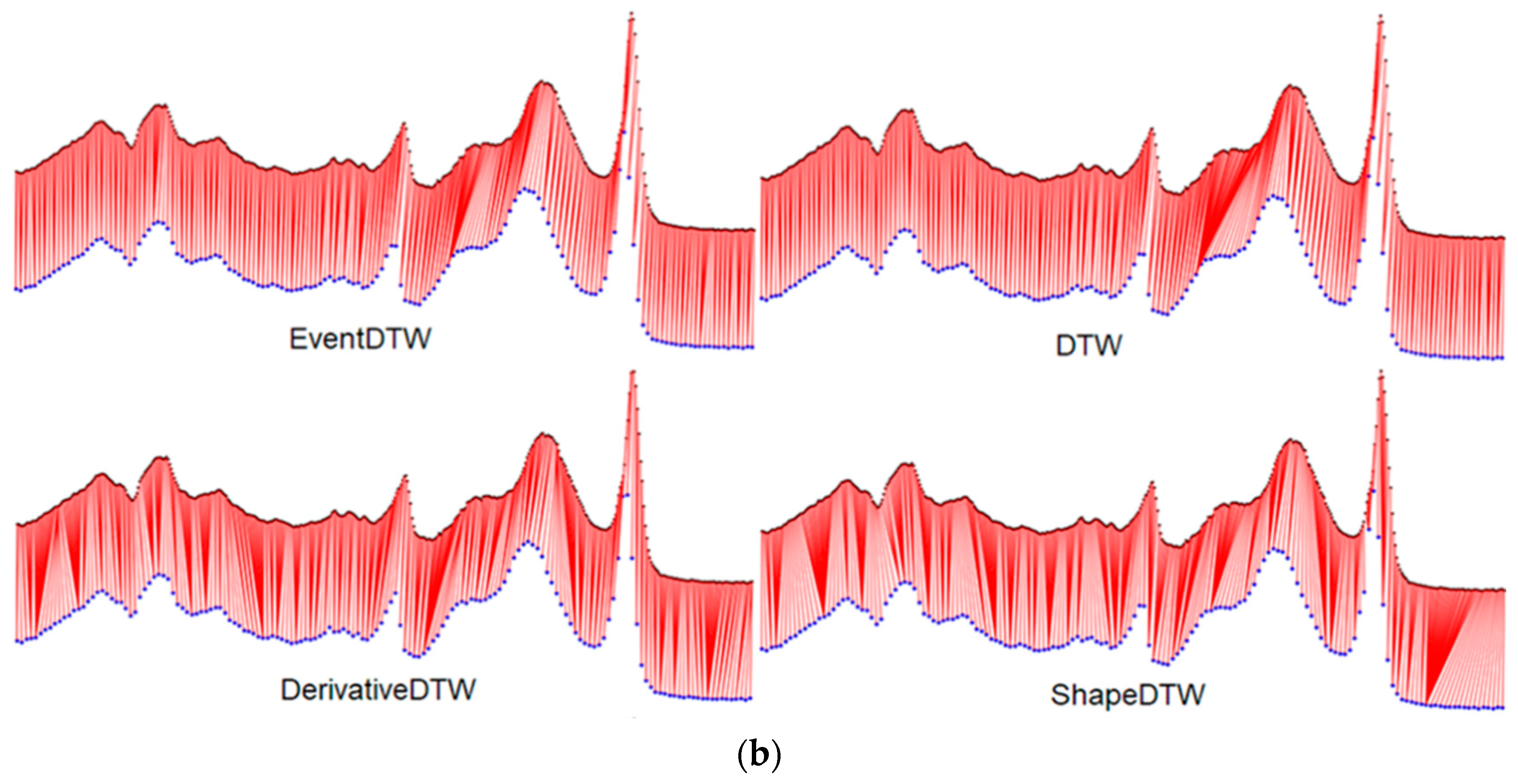
3.1. Results of Alignments on the University of California, Riverside (UCR) Time Series Classification Archive
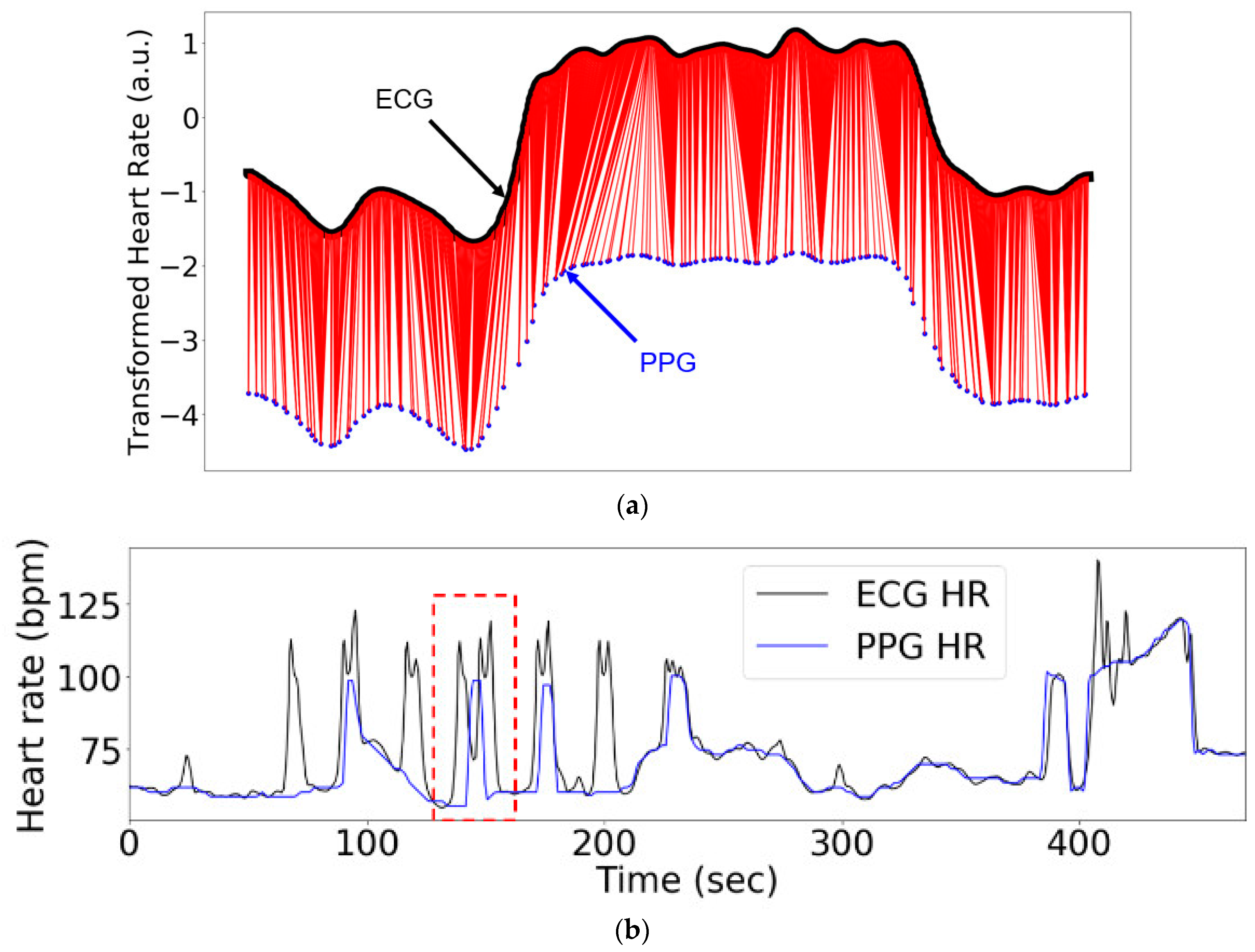
3.2. Results of Alignments on ECG/PPG Heart Rate Datasets
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Myers, C.; Rabiner, L.; Rosenberg, A. Performance tradeoffs in dynamic time warping algorithms for isolated word recognition. IEEE Trans. Acoust. Speech Signal Process. 1980, 28, 623–635. [Google Scholar] [CrossRef]
- Hachaj, T.; Piekarczyk, M. Evaluation of Pattern Recognition Methods for Head Gesture-Based Interface of a Virtual Reality Helmet Equipped with a Single IMU Sensor. Sensors 2019, 19, 5408. [Google Scholar] [CrossRef] [PubMed]
- Gineke, T.H.; Marcel, A.; Reinders, J.T.; Hendriks, E.A. Multi-dimensional dynamic time warping for gesture recognition. In Proceedings of the Thirteenth annual conference of the Advanced School for Computing and Imaging, Heijen, The Netherlands, 13–15 June 2007; Volume 300. [Google Scholar]
- Izakian, H.; Witold, P.; Iqbal, J. Fuzzy clustering of time series data using dynamic time warping distance. Eng. Appl. Artif. Intell. 2015, 39, 235–244. [Google Scholar] [CrossRef]
- Viceconti, M.; Clapworthy, G.; Testi, D.; Taddei, F.; McFarlane, N. Multimodal fusion of biomedical data at different temporal and dimensional scales. Comput. Methods Programs Biomed. 2011, 102, 227–237. [Google Scholar] [CrossRef]
- Burns, A.; Greene, B.R.; McGrath, M.J.; O’Shea, T.J.; Kuris, B.; Ayer, S.M.; Stroiescu, F.; Cionca, V. SHIMMER™—A wireless sensor platform for noninvasive biomedical research. IEEE Sens. J. 2010, 10, 1527–1534. [Google Scholar] [CrossRef]
- Sakoe, H.; Chiba, S. Dynamic programming algorithm optimization for spoken word recognition. IEEE Trans. Acoust. Speech Signal Proc. 1978, 26, 43–49. [Google Scholar] [CrossRef]
- Keogh Eamonn, J.; Michael Pazzani, J. Derivative dynamic time warping. In Proceedings of the 2001 SIAM International Conference on Data Mining. Society for Industrial and Applied Mathematics, Chicago, IL, USA, 5–7 April 2001. [Google Scholar]
- Chang, C.-Y.; Huang, D.-A.; Sui, Y.; Fei, L.; Niebles, J.C. D3TW: Discriminative Differentiable Dynamic Time Warping for Weakly Supervised Action Alignment and Segmentation. In Proceedings of the 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Long Beach, CA, USA, 16–20 June 2019; pp. 3541–3550. [Google Scholar]
- Cifuentes, J.; Pham, M.T.; Moreau, R.; Prieto, F.; Boulanger, P. Surgical gesture classification using Dynamic Time Warping and affine velocity. In Proceedings of the 2017 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Seogwipo, Korea, 11–15 July 2017; pp. 2275–2278. [Google Scholar]
- Zhao, J.; Laurent, I. shapeDTW: Shape Dynamic Time Warping. arXiv 2016, arXiv:1606.01601. [Google Scholar] [CrossRef]
- Berndt, D.; Clifford, J. Using dynamic time warping to find patterns in time series. AAAI-94 Workshop on Knowledge Discovery in Databases (KDD-94), Seattle, 31 July–1 August 1994. Available online: https://www.aaai.org/Library/Workshops/ws94-03.php (accessed on 6 May 2020).
- Kruskall, J.B.; Liberman, M. The symmetric time warping algorithm: From continuous to discrete. In Time Warps, String Edits and Macromolecules: The Theory and Practice of String Comparison; Addison-Wesley: Boston, MA, USA, 1983. [Google Scholar]
- Itakura, F. Minimum prediction residual principle applied to speech recognition. IEEE Trans. Acoust. Speech Signal Proc. 1975, 23, 52–72. [Google Scholar] [CrossRef]
- Abdullah, M.; Eamonn, J. Keogh: Extracting Optimal Performance from Dynamic Time Warping. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 2129–2130. [Google Scholar]
- Kirchhoff, H.; Lerch, A. Evaluation of features for audio-to-audio alignment. J. New Music Res. 2011, 40, 27–41. [Google Scholar] [CrossRef]
- Petitjean, F.; Forestier, G.; Webb, G.I.; Nicholson, A.E.; Chen, Y.; Keogh, E. Dynamic time warping averaging of time series allows faster and more accurate classification. In Proceedings of the 2014 IEEE international conference on data mining, New York, NY, USA, 24–28 February 2014. [Google Scholar]
- Rosa, M.; Fugmann, E.; Pinto, G.; Nunes, M. An anchored dynamic time-warping for alignment and comparison of swallowing acoustic signals. In Proceedings of the 2017 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Seogwipo, Korea, 11–15 July 2017; pp. 2749–2752. [Google Scholar]
- Douglass, A.C.S.; Harley, J.B. Dynamic Time Warping Temperature Compensation for Guided Wave Structural Health Monitoring. IEEE Trans. Ultrason. Ferroelectr. Freq. Control 2018, 65, 851–861. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Eamonn, K.; Hu, B.; Nurjahan, B.; Anthony, B.; Abdullah, M.; Gustavo, B. The UCR Time Series Classification Archive. Available online: www.cs.ucr.edu/~eamonn/time_series_data/ (accessed on 20 August 2019).
- Laub, P.J.; Taimre, T.; Pollett, P.K. Hawkes processes. arXiv 2015, arXiv:1507.02822. [Google Scholar]
- Rakthanmanon, T.; Campana, B.; Mueen, A.; Batista, G.; Westover, B.; Zhu, Q.; Zakaria, J.; Keogh, E. Searching and mining trillions of time series subsequencesunder dynamic time warping. In Proceedings of the 18th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Beijing, China, 12–16 August 2012; p. 262. [Google Scholar]
- Bent, B.; Goldstein, B.A.; Kibbe, W.A.; Dunn, J.P. Investigating sources of inaccuracy in wearable optical heart rate sensors. NPJ Digit. Med. 2020, 3, 1–9. [Google Scholar] [CrossRef] [PubMed]
| Algorithm | Percent of Best Algorithm by Singularity Score | Percent of Best Algorithm by Error Rate |
|---|---|---|
| eDTW | 42% | 37% |
| DTW | 32% | 49% |
| sDTW | 7% | 4% |
| dDTW | 19% | 11% |
| Algorithm | Percent of Best Algorithm by Singularity Score |
|---|---|
| eDTW | 76% |
| DTW | 0% |
| sDTW | 0% |
| dDTW | 24% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, Y.; Qi, Y.; Wang, W.K.; Bent, B.; Avram, R.; Olgin, J.; Dunn, J. EventDTW: An Improved Dynamic Time Warping Algorithm for Aligning Biomedical Signals of Nonuniform Sampling Frequencies. Sensors 2020, 20, 2700. https://doi.org/10.3390/s20092700
Jiang Y, Qi Y, Wang WK, Bent B, Avram R, Olgin J, Dunn J. EventDTW: An Improved Dynamic Time Warping Algorithm for Aligning Biomedical Signals of Nonuniform Sampling Frequencies. Sensors. 2020; 20(9):2700. https://doi.org/10.3390/s20092700
Chicago/Turabian StyleJiang, Yihang, Yuankai Qi, Will Ke Wang, Brinnae Bent, Robert Avram, Jeffrey Olgin, and Jessilyn Dunn. 2020. "EventDTW: An Improved Dynamic Time Warping Algorithm for Aligning Biomedical Signals of Nonuniform Sampling Frequencies" Sensors 20, no. 9: 2700. https://doi.org/10.3390/s20092700
APA StyleJiang, Y., Qi, Y., Wang, W. K., Bent, B., Avram, R., Olgin, J., & Dunn, J. (2020). EventDTW: An Improved Dynamic Time Warping Algorithm for Aligning Biomedical Signals of Nonuniform Sampling Frequencies. Sensors, 20(9), 2700. https://doi.org/10.3390/s20092700






