Investigation of Machine Learning Approaches for Traumatic Brain Injury Classification via EEG Assessment in Mice
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal Data
2.2. Fluid Percussion Injury (FPI) and EEG/Electromyography (EMG) Sleep-Wake Recordings
2.3. EEG and Sleep/Wake Scoring
2.4. Analysis Implementation
2.4.1. Rule-Based Machine Learning Algorithms
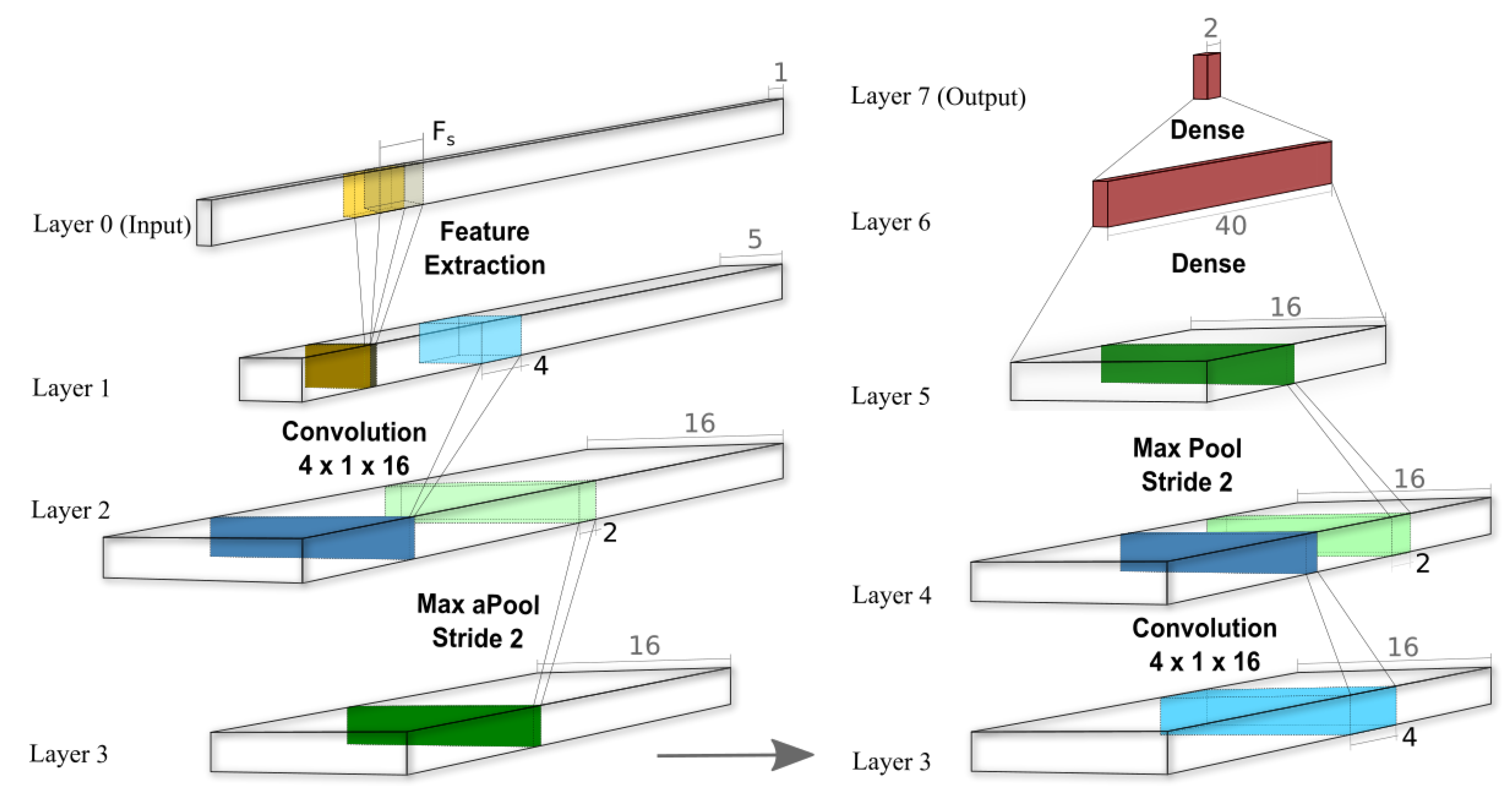
2.4.2. Convolutional Neural Network (CNN)
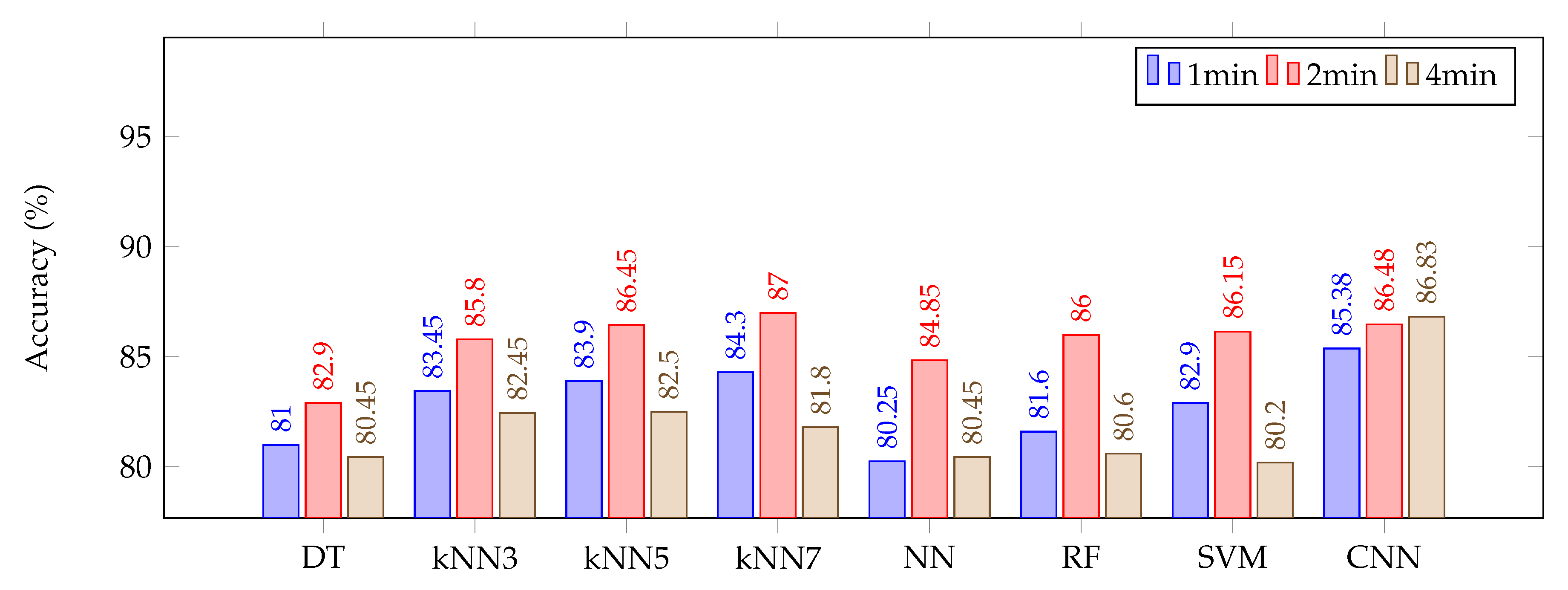
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Dewan, M.C.; Rattani, A.; Gupta, S.; Baticulon, R.E.; Hung, Y.C.; Punchak, M.; Agrawal, A.; Adeleye, A.O.; Shrime, M.G.; Rubiano, A.M.; et al. Estimating the global incidence of traumatic brain injury. J. Neurosurg. 2018, 1, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Sandsmark, D.K.; Elliott, J.E.; Lim, M.M. Sleep-wake disturbances after traumatic brain injury: Synthesis of human and animal studies. Sleep 2017, 40. [Google Scholar] [CrossRef] [PubMed]
- Ruff, R. Two decades of advances in understanding of mild traumatic brain injury. J. Head Trauma Rehabil. 2005, 20, 5–18. [Google Scholar] [CrossRef] [PubMed]
- Jamaludin, Z.; Mokhtar, M.N.A. Intelligent Manufacturing and Mechatronics: Proceedings of the 2nd Symposium on Intelligent Manufacturing and Mechatronics-SympoSIMM 2019, 8 July 2019, Melaka, Malaysia; Springer: Melaka, Malaysia, 2019. [Google Scholar]
- Penttonen, M.; Buzsáki, G. Natural logarithmic relationship between brain oscillators. Thalamus Relat. Syst. 2003, 2, 145–152. [Google Scholar] [CrossRef]
- Lim, M.M.; Elkind, J.; Xiong, G.; Galante, R.; Zhu, J.; Zhang, L.; Lian, J.; Rodin, J.; Kuzma, N.N.; Pack, A.I.; et al. Dietary therapy mitigates persistent wake deficits caused by mild traumatic brain injury. Sci. Transl. Med. 2013, 5, 215ra173. [Google Scholar] [CrossRef]
- Modarres, M.H.; Kuzma, N.N.; Kretzmer, T.; Pack, A.I.; Lim, M.M. EEG slow waves in traumatic brain injury: Convergent findings in mouse and man. Neurobiol. Sleep Circadian Rhythm. 2017, 2, 59–70. [Google Scholar] [CrossRef]
- Arbour, C.; Khoury, S.; Lavigne, G.J.; Gagnon, K.; Poirier, G.; Montplaisir, J.Y.; Carrier, J.; Gosselin, N. Are NREM sleep characteristics associated to subjective sleep complaints after mild traumatic brain injury? Sleep Med. 2015, 16, 534–539. [Google Scholar] [CrossRef]
- Gosselin, N.; Lassonde, M.; Petit, D.; Leclerc, S.; Mongrain, V.; Collie, A.; Montplaisir, J. Sleep following sport-related concussions. Sleep Med. 2009, 10, 35–46. [Google Scholar] [CrossRef]
- Orff, H.J.; Ayalon, L.; Drummond, S.P. Traumatic brain injury and sleep disturbance: A review of current research. J. Head Trauma Rehabil. 2009, 24, 155–165. [Google Scholar] [CrossRef]
- Rapp, P.E.; Keyser, D.O.; Albano, A.; Hernandez, R.; Gibson, D.B.; Zambon, R.A.; Hairston, W.D.; Hughes, J.D.; Krystal, A.; Nichols, A.S. Traumatic brain injury detection using electrophysiological methods. Front. Human Neurosci. 2015, 9, 11. [Google Scholar] [CrossRef]
- Cao, C.; Tutwiler, R.L.; Slobounov, S. Automatic classification of athletes with residual functional deficits following concussion by means of EEG signal using support vector machine. IEEE Trans. Neural Syst. Rehabil. Eng. 2008, 16, 327–335. [Google Scholar] [PubMed]
- Dixon, C.E.; Lyeth, B.G.; Povlishock, J.T.; Findling, R.L.; Hamm, R.J.; Marmarou, A.; Young, H.F.; Hayes, R.L. A fluid percussion model of experimental brain injury in the rat. J. Neurosurg. 1987, 67, 110–119. [Google Scholar] [CrossRef] [PubMed]
- McIntosh, T.; Vink, R.; Noble, L.; Yamakami, I.; Fernyak, S.; Soares, H.; Faden, A. Traumatic brain injury in the rat: Characterization of a lateral fluid-percussion model. Neuroscience 1989, 28, 233–244. [Google Scholar] [CrossRef]
- Carbonell, W.S.; Maris, D.O.; McCall, T.; Grady, M.S. Adaptation of the fluid percussion injury model to the mouse. J. Neurosurg. 1998, 15, 217–229. [Google Scholar] [CrossRef]
- Safavian, S.R.; Landgrebe, D. A survey of decision tree classifier methodology. IEEE Trans. Syst. Man Cybern. 1991, 21, 660–674. [Google Scholar] [CrossRef]
- Abe, S. Support Vector Machines for Pattern Classification; Springer: London, UK, 2005; Volume 2. [Google Scholar]
- Peterson, L.E. K-nearest neighbor. Scholarpedia 2009, 4, 1883. [Google Scholar] [CrossRef]
- Cohen, M.X. Analyzing Neural Time Series Data: Theory and Practice; MIT Press: Cambridge, MA, USA, 2014. [Google Scholar]
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 2014, 15, 1929–1958. [Google Scholar]
- Perez, L.; Wang, J. The effectiveness of data augmentation in image classification using deep learning. arXiv 2017, arXiv:1712.04621. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 26 June–1 July 2016; pp. 770–778. [Google Scholar]
- Jafarlou, S.; Khorram, S.; Kothapally, V.; Hansen, J.H. Analyzing Large Receptive Field Convolutional Networks for Distant Speech Recognition. arXiv 2019, arXiv:1910.07047. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Lawhern, V.J.; Solon, A.J.; Waytowich, N.R.; Gordon, S.M.; Hung, C.P.; Lance, B.J. EEGNet: A compact convolutional neural network for EEG-based brain–computer interfaces. J. Neural Eng. 2018, 15, 056013. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Oh, S.L.; Hagiwara, Y.; Tan, J.H.; Adeli, H. Deep convolutional neural network for the automated detection and diagnosis of seizure using EEG signals. Comput. Biol. Med. 2018, 100, 270–278. [Google Scholar] [CrossRef] [PubMed]



| Subjects | 1 min | 2 min | 4 min |
|---|---|---|---|
| Sham102 | 736 | 352 | 168 |
| Sham103 | 637 | 275 | 101 |
| Sham104 | 922 | 427 | 186 |
| Sham107 | 684 | 316 | 146 |
| Sham108 | 780 | 364 | 164 |
| TBI102 | 901 | 429 | 201 |
| TBI103 | 271 | 81 | 16 |
| TBI104 | 207 | 61 | 12 |
| TBI106 | 458 | 181 | 59 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vishwanath, M.; Jafarlou, S.; Shin, I.; Lim, M.M.; Dutt, N.; Rahmani, A.M.; Cao, H. Investigation of Machine Learning Approaches for Traumatic Brain Injury Classification via EEG Assessment in Mice. Sensors 2020, 20, 2027. https://doi.org/10.3390/s20072027
Vishwanath M, Jafarlou S, Shin I, Lim MM, Dutt N, Rahmani AM, Cao H. Investigation of Machine Learning Approaches for Traumatic Brain Injury Classification via EEG Assessment in Mice. Sensors. 2020; 20(7):2027. https://doi.org/10.3390/s20072027
Chicago/Turabian StyleVishwanath, Manoj, Salar Jafarlou, Ikhwan Shin, Miranda M. Lim, Nikil Dutt, Amir M. Rahmani, and Hung Cao. 2020. "Investigation of Machine Learning Approaches for Traumatic Brain Injury Classification via EEG Assessment in Mice" Sensors 20, no. 7: 2027. https://doi.org/10.3390/s20072027
APA StyleVishwanath, M., Jafarlou, S., Shin, I., Lim, M. M., Dutt, N., Rahmani, A. M., & Cao, H. (2020). Investigation of Machine Learning Approaches for Traumatic Brain Injury Classification via EEG Assessment in Mice. Sensors, 20(7), 2027. https://doi.org/10.3390/s20072027






