Electrochemical Immuno- and Aptamer-Based Assays for Bacteria: Pros and Cons over Traditional Detection Schemes
Abstract
1. Introduction
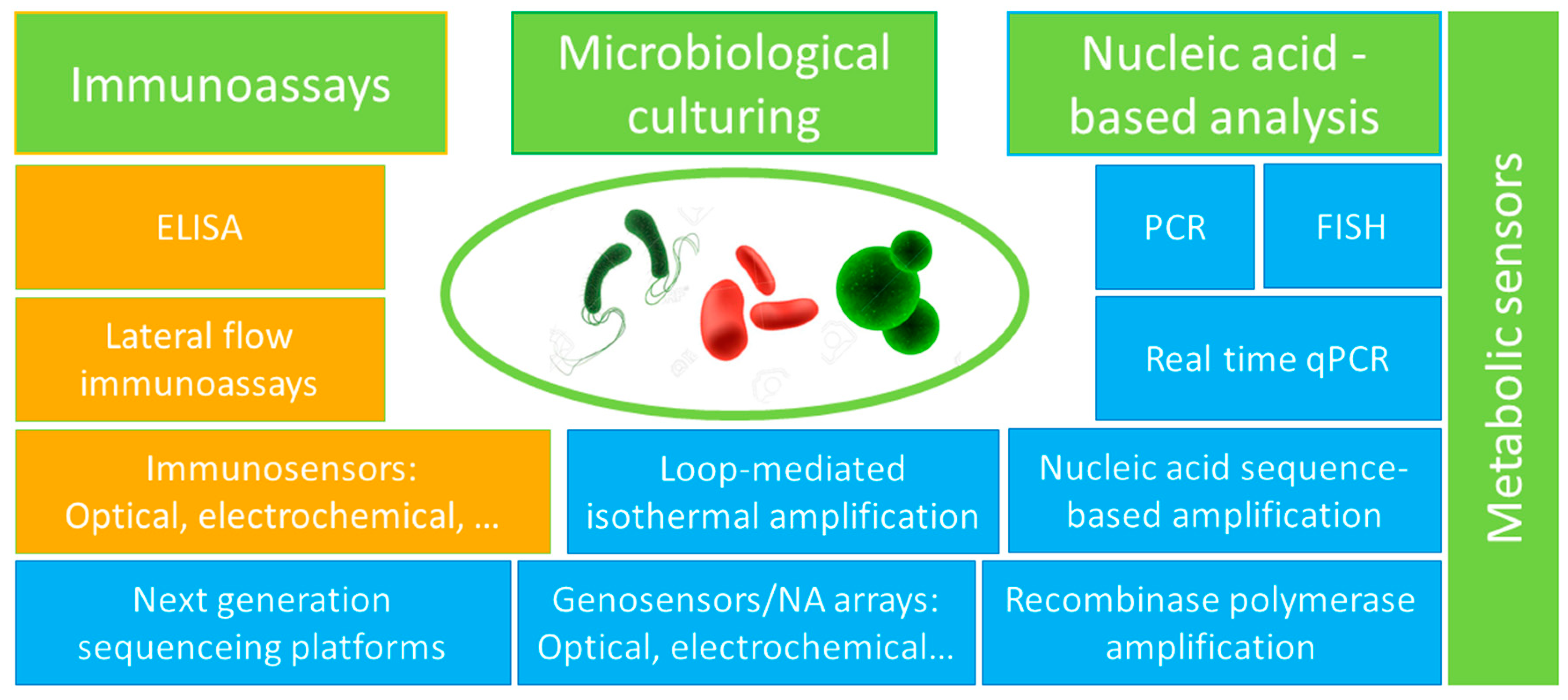
2. Methodological Approaches for Bacterial Detection and Quantification
- -
- NA sequence-based amplification (NASBA, amplifies and detects bacterial messenger RNA, more sensitive and fast (less than 90 min) than PCR, no interference from dead cells’ DNA, however, too expensive for environmental applications and suffers from errors in amplification and quantification following the amplification step) [23];
- -
- Loop-mediated isothermal amplification (LAMP, less expensive than PCR, more sensitive and faster (1 h) DNA amplification at 60–65 °C, less sensitive to inhibitors) [24];
- -
- Recombinase polymerase amplification (RPA, fast (<20 min) amplification of DNA/RNA at 37–42 °C; can be integrated with other, portable detection devices, however, it faces primers design difficulties and requires post-amplification purification digestion) [25].
3. Immunoassays and Aptamer-Based Assays with Optical Detection
4. Electrochemical Immunoassays
4.1. Electrochemical ELISA
4.2. Electrochemical Immunoassays (Not Enzyme-Linked)
4.2.1. Antibodies and Aptamers Based Assays
4.2.2. Antimicrobial Peptides (AMP) Based Assays
4.2.3. Bacteriophages Based Assays
4.3. Whole Cell Imprinted Polymer Sensors as Alternative to E-Immunoassays
4.4. Electro-Optical Immunoassays
4.5. Electrochemical Immunoanalysis of Whole Cells with A Nanopore Technology
5. Future Perspectives
Funding
Conflicts of Interest
References
- Velusamy, V.; Arshak, K.; Korostynska, O.; Oliwa, K.; Adley, C. An overview of foodborne pathogen detection: In the perspective of biosensors. Biotechnol. Adv. 2010, 28, 232–254. [Google Scholar] [CrossRef]
- WHO. Available online: https://www.who.int/water_sanitation_health/publications/jmp-report-2019/en/; https://www.who.int/water_sanitation_health/diseases-risks/en/ (accessed on 25 September 2020).
- Law, J.W.-F.; Ab Mutalib, N.-S.; Chan, K.-G.; Lee, L.-H. Rapid methods for the detection of foodborne bacterial pathogens: Principles, applications, advantages and limitations. Front. Microbiol. 2015, 5, 770. [Google Scholar] [CrossRef]
- Dye, C. After 2015: Infectious diseases in a new era of health and development. Philos. Trans. R. Soc. B Biol. Sci. 2014, 369, 20130426. [Google Scholar] [CrossRef] [PubMed]
- Koch, G.H.; Brongers, M.P.; Thompson, N.G.; Virmani, Y.P.; Payer, J.H. Cost of corrosion in the United States. In Handbook of Environmental Degradation of Materials; William Andrew Inc.: New York, NY, USA, 2005; pp. 3–24. [Google Scholar]
- Jansen, H.J.; Breeveld, F.J.; Stijnis, C.; Grobusch, M.P. Biological warfare, bioterrorism, and biocrime. Clin. Microbiol. Infect. 2014, 20, 488–496. [Google Scholar] [CrossRef] [PubMed]
- WHO. Guidelines for drinking-water quality. In Surveillance and Control of Community Supplies; World Health Organization: Geneva, Switzerland, 1997; Volume 3. [Google Scholar]
- U.S. Department of Health and Human Services; Public Health Service; Food and Drug Administration. Grade “A” Pasteurized Milk Ordinance; Public Health Service, Food and Drug Administration: Arden, NC, USA, 2017.
- Shipovskov, S.; Saunders, A.M.; Nielsen, J.S.; Hansen, M.H.; Gothelf, K.V.; Ferapontova, E.E. Electrochemical sandwich assay for attomole analysis of DNA and RNA from beer spoilage bacteria Lactobacillus brevis. Biosens. Bioelectron. 2012, 37, 99–106. [Google Scholar] [CrossRef] [PubMed]
- McEntire, J.; Acheson, D.; Siemens, A.; Eilert, S.; Robach, M. The Public Health Value of Reducing Salmonella Levels in Raw Meat and Poultry. Food Prot. Trends 2014, 34, 386–392. [Google Scholar]
- Lee, A.; Mirrett, S.; Reller, L.B.; Weinstein, M.P. Detection of bloodstream Infections in adults: How many blood cultures are needed? J. Clin. Microbiol. 2007, 45, 3546–3548. [Google Scholar] [CrossRef]
- Proctor, L.M.; Creasy, H.H.; Fettweis, J.M.; Lloyd-Price, J.; Mahurkar, A.; Zhou, W.; Buck, G.A.; Snyder, M.P.; Strauss, J.F.; Weinstock, G.M.; et al. The Integrative Human Microbiome Project. Nature 2019, 569, 641–648. [Google Scholar]
- Carding, S.; Verbeke, K.; Vipond, D.T.; Corfe, B.M.; Owen, L.J. Dysbiosis of the gut microbiota in disease. Microb. Ecol. Health Dis. 2015, 26, 26191. [Google Scholar] [CrossRef]
- WHO. New Report Calls for Urgent Action to Avert Antimicrobial Resistance Crisis; WHO: Geneva, Switzerland, 2019. [Google Scholar]
- Nolan, T.; Hands, R.E.; Bustin, S.A. Quantification of mRNA using real time RT-PCR. Nat. Protoc. 2006, 1, 1559–1582. [Google Scholar] [CrossRef]
- Wang, Y.; Salazar, J.K. Culture-independent rapid detection methods for bacterial pathogens and toxins in food matrices. Compr. Rev. Food Sci. Food Saf. 2016, 15, 183–205. [Google Scholar] [CrossRef]
- Costea, P.I.; Zeller, G.; Sunagawa, S.; Pelletier, E.; Alberti, A.; Levenez, F.; Tramontano, M.; Driessen, M.; Hercog, R.; Jung, F.-E.; et al. Towards standards for human fecal sample processing in metagenomic studies. Nat. Biotechnol. 2017, 35, 1069. [Google Scholar] [CrossRef]
- Ganz, K.; Gill, A. Inhibition of polymerase chain reaction for the detection of Escherichia coli O157:H7 and Salmonella enterica on walnut kernels. Food Microbiol. 2013, 35, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Lin, C.-W.; Wang, J. Advances in rapid detection methods for foodborne pathogens. J. Microbiol. Biotechnol. 2014, 24, 297–312. [Google Scholar] [CrossRef] [PubMed]
- Margot, H.; Stephan, R.; Guarino, S.; Jagadeesan, B.; Chilton, D.; O’Mahony, E.; Iversen, C. Inclusivity, exclusivity and limit of detection of commercially available real-time PCR assays for the detection of Salmonella. Int. J. Food Microbiol. 2013, 165, 221–226. [Google Scholar] [CrossRef] [PubMed]
- Nocker, A.; Cheung, C.-Y.; Camper, A.K. Comparison of propidium monoazide with ethidium monoazide for differentiation of live vs. dead bacteria by selective removal of DNA from dead cells. J. Microbiol. Meth. 2006, 67, 310–320. [Google Scholar] [CrossRef]
- Li, J.; Macdonald, J. Advances in isothermal amplification: Novel strategies inspired by biological processes. Biosens. Bioelectron. 2015, 64, 196–211. [Google Scholar] [CrossRef]
- Hønsvall, B.K.; Robertson, L.J. From research lab to standard environmental analysis tool: Will NASBA make the leap? Water Res. 2017, 109, 389–397. [Google Scholar] [CrossRef]
- Li, Y.; Fan, P.; Zhou, S.; Zhang, L. Loop-mediated isothermal amplification (LAMP): A novel rapid detection platform for pathogens. Microb. Pathog. 2017, 107, 54–61. [Google Scholar] [CrossRef]
- Lobato, I.M.; O’Sullivan, C.K. Recombinase polymerase amplification: Basics, applications and recent advances. TrAC Tr. Anal. Chem. 2018, 98, 19–35. [Google Scholar] [CrossRef]
- Ferapontova, E. Basic concepts and recent advances in electrochemical analysis of nucleic acids. Curr. Opin. Electrochem. 2017, 5, 218–225. [Google Scholar] [CrossRef]
- Simoska, O.; Stevenson, K.J. Electrochemical sensors for rapid diagnosis of pathogens in real time. Analyst 2019, 144, 6461–6478. [Google Scholar] [CrossRef] [PubMed]
- Severgnini, M.; Cremonesi, P.; Consolandi, C.; De Bellis, G.; Castiglioni, B. Advances in DNA microarray technology for the detection of foodborne pathogens. Food Bioproc. Technol. 2011, 4, 936–953. [Google Scholar] [CrossRef]
- McLoughlin, K.S. Microarrays for pathogen detection and analysis. Brief. Funct. Gen. 2011, 10, 342–353. [Google Scholar] [CrossRef] [PubMed]
- Deurenberg, R.H.; Bathoorn, E.; Chlebowicz, M.A.; Couto, N.; Ferdous, M.; García-Cobos, S.; Kooistra-Smid, A.M.D.; Raangs, E.C.; Rosema, S.; Veloo, A.C.M.; et al. Application of next generation sequencing in clinical microbiology and infection prevention. J. Biotechnol. 2017, 243, 16–24. [Google Scholar] [CrossRef] [PubMed]
- Vincent, A.T.; Derome, N.; Boyle, B.; Culley, A.I.; Charette, S.J. Next-generation sequencing (NGS) in the microbiological world: How to make the most of your money. J. Microbiol. Meth. 2017, 138, 60–71. [Google Scholar] [CrossRef]
- Czilwik, G.; Messinger, T.; Strohmeier, O.; Wadle, S.; von Stetten, F.; Paust, N.; Roth, G.; Zengerle, R.; Saarinen, P.; Niittymäki, J.; et al. Rapid and fully automated bacterial pathogen detection on a centrifugal-microfluidic LabDisk using highly sensitive nested PCR with integrated sample preparation. Lab Chip 2015, 15, 3749–3759. [Google Scholar] [CrossRef]
- Yan, H.; Zhu, Y.; Zhang, Y.; Wang, L.; Chen, J.; Lu, Y.; Xu, Y.; Xing, W. Multiplex detection of bacteria on an integrated centrifugal disk using bead-beating lysis and loop-mediated amplification. Sci. Rep. 2017, 7, 1460. [Google Scholar] [CrossRef]
- Waage, A.S.; Vardund, T.; Lund, V.; Kapperud, G. Detection of Small Numbers of Campylobacter jejuni and Campylobacter coli Cells in Environmental Water, Sewage, and Food Samples by a Seminested PCR Assay. Appl. Environ. Microbiol. 1999, 65, 1636–1643. [Google Scholar] [CrossRef]
- Ruiz-Rueda, O.; Soler, M.; Calvó, L.; García-Gil, J.L. Multiplex Real-time PCR for the Simultaneous Detection of Salmonella spp. and Listeria monocytogenes in Food Samples. Food Anal. Methods 2010, 4, 131–138. [Google Scholar] [CrossRef]
- Wolffs, P.F.; Glencross, K.; Thibaudeau, R.; Griffiths, M.W. Direct quantitation and detection of salmonellae in biological samples without enrichment, using two-step filtration and real-time PCR. Appl. Environ. Microbiol. 2006, 72, 3896–3900. [Google Scholar] [CrossRef] [PubMed]
- Vinayaka, A.C.; Ngo, T.A.; Kant, K.; Engelsmann, P.; Dave, V.P.; Shahbazi, M.-A.; Wolff, A.; Bang, D.D. Rapid detection of Salmonella enterica in food samples by a novel approach with combination of sample concentration and direct PCR. Biosens. Bioelectron. 2019, 129, 224–230. [Google Scholar] [CrossRef] [PubMed]
- Xiao, L.; Zhang, Z.; Sun, X.; Pan, Y.; Zhao, Y. Development of a quantitative real-time PCR assay for viable Salmonella spp. without enrichment. Food Control 2015, 57, 185–189. [Google Scholar] [CrossRef]
- Ma, K.; Deng, Y.; Bai, Y.; Xu, D.; Chen, E.; Wu, H.; Li, B.; Gao, L. Rapid and simultaneous detection of Salmonella, Shigella, and Staphylococcus aureus in fresh pork using a multiplex real-time PCR assay based on immunomagnetic separation. Food Control 2014, 42, 87–93. [Google Scholar] [CrossRef]
- Kim, G.; Moon, J.-H.; Moh, C.-Y.; Lim, J.-g. A microfluidic nano-biosensor for the detection of pathogenic Salmonella. Biosens. Bioelectron. 2015, 67, 243–247. [Google Scholar] [CrossRef]
- Li, J.; Zhai, L.; Bie, X.; Lu, Z.; Kong, X.; Yu, Q.; Lv, F.; Zhang, C.; Zhao, H. A novel visual loop-mediated isothermal amplification assay targeting gene62181533 for the detection of Salmonella spp. in foods. Food Control 2016, 60, 230–236. [Google Scholar] [CrossRef]
- Yamazaki, W.; Ishibashi, M.; Kawahara, R.; Inoue, K. Development of a loop-mediated isothermal amplification assay for sensitive and rapid detection of Vibrio parahaemolyticus. BMC Microbiol. 2008, 8, 163. [Google Scholar] [CrossRef]
- Shao, Y.; Zhu, S.; Jin, C.; Chen, F. Development of multiplex loop-mediated isothermal amplification-RFLP (mLAMP-RFLP) to detect Salmonella spp. and Shigella spp. in milk. Int. J. Food Microbiol. 2011, 148, 75–79. [Google Scholar] [CrossRef]
- Min, J.; Baeumner, A.J. Highly sensitive and specific detection of viable Escherichia coli in drinking water. Anal. Biochem. 2002, 303, 186–193. [Google Scholar] [CrossRef]
- O’Grady, J.; Lacey, K.; Glynn, B.; Smith, T.J.; Barry, T.; Maher, M. tmRNA—A novel high-copy-number RNA diagnostic target--its application for Staphylococcus aureus detection using real-time NASBA. FEMS Microbiol. Lett. 2009, 301, 218–223. [Google Scholar] [CrossRef]
- Song, Y.; Li, W.; Duan, Y.; Li, Z.; Deng, L. Nicking enzyme-assisted biosensor for Salmonella enteritidis detection based on fluorescence resonance energy transfer. Biosens. Bioelectron. 2014, 55, 400–404. [Google Scholar] [CrossRef] [PubMed]
- Guo, D.; Liu, B.; Liu, F.; Cao, B.; Chen, M.; Hao, X.; Feng, L.; Wang, L. Development of a DNA Microarray for Molecular Identification of All 46 Salmonella O Serogroups. Appl. Environ. Microbiol. 2013, 79, 3392. [Google Scholar] [CrossRef] [PubMed]
- Huang, A.; Qiu, Z.; Jin, M.; Shen, Z.; Chen, Z.; Wang, X.; Li, J.-W. High-throughput detection of food-borne pathogenic bacteria using oligonucleotide microarray with quantum dots as fluorescent labels. Int. J. Food Microbiol. 2014, 185, 27–32. [Google Scholar] [CrossRef] [PubMed]
- Tortajada-Genaro, L.A.; Rodrigo, A.; Hevia, E.; Mena, S.; Niñoles, R.; Maquieira, Á. Microarray on digital versatile disc for identification and genotyping of Salmonella and Campylobacter in meat products. Anal. Bioanal. Chem. 2015, 407, 7285–7294. [Google Scholar] [CrossRef] [PubMed]
- Varshney, M.; Yang, L.; Su, X.-L.; Li, Y. Magnetic Nanoparticle-Antibody Conjugates for the Separation of Escherichia coli O157:H7 in Ground Beef. J. Food Prot. 2005, 68, 1804–1811. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Wu, S.; Duan, N.; Ma, X.; Xia, Y.; Chen, J.; Ding, Z.; Wang, Z. A sensitive gold nanoparticle-based colorimetric aptasensor for Staphylococcus aureus. Talanta 2014, 127, 163–168. [Google Scholar] [CrossRef]
- Zhong, Z.; Gao, X.; Gao, R.; Jia, L. Selective capture and sensitive fluorometric determination of Pseudomonas aeruginosa by using aptamer modified magnetic nanoparticles. Microchim. Acta 2018, 185, 377. [Google Scholar] [CrossRef]
- Chaivisuthangkura, P.; Pengsuk, C.; Longyant, S.; Sithigorngul, P. Evaluation of monoclonal antibody based immunochromatographic strip test for direct detection of Vibrio cholerae O1 contamination in seafood samples. J. Microbiol. Methods 2013, 95, 304–311. [Google Scholar] [CrossRef]
- Shen, Z.; Hou, N.; Jin, M.; Qiu, Z.; Wang, J.; Zhang, B.; Wang, X.; Wang, J.; Zhou, D.; Li, J. A novel enzyme-linked immunosorbent assay for detection of Escherichia coli O157:H7 using immunomagnetic and beacon gold nanoparticles. Gut Pathog. 2014, 6, 14. [Google Scholar] [CrossRef]
- Wu, W.; Li, J.; Pan, D.; Li, J.; Song, S.; Rong, M.; Li, Z.; Gao, J.; Lu, J. Gold nanoparticle-based enzyme-linked antibody-aptamer sandwich assay for detection of Salmonella Typhimurium. ACS Appl. Mater Interfaces 2014, 6, 16974–16981. [Google Scholar] [CrossRef]
- Jin, B.; Wang, S.; Lin, M.; Jin, Y.; Zhang, S.; Cui, X.; Gong, Y.; Li, A.; Xu, F.; Lu, T.J. Upconversion nanoparticles based FRET aptasensor for rapid and ultrasenstive bacteria detection. Biosens. Bioelectron. 2017, 90, 525–533. [Google Scholar] [CrossRef] [PubMed]
- Shin, W.R.; Sekhon, S.S.; Rhee, S.K.; Ko, J.H.; Ahn, J.Y.; Min, J.; Kim, Y.H. Aptamer-Based Paper Strip Sensor for Detecting Vibrio fischeri. ACS Comb. Sci. 2018, 20, 261–268. [Google Scholar] [CrossRef] [PubMed]
- Mouffouk, F.; da Costa, A.M.; Martins, J.; Zourob, M.; Abu-Salah, K.M.; Alrokayan, S.A. Development of a highly sensitive bacteria detection assay using fluorescent pH-responsive polymeric micelles. Biosens. Bioelectron. 2011, 26, 3517–3523. [Google Scholar] [CrossRef] [PubMed]
- Cho, I.H.; Mauer, L.; Irudayaraj, J. In-situ fluorescent immunomagnetic multiplex detection of foodborne pathogens in very low numbers. Biosens. Bioelectron. 2014, 57, 143–148. [Google Scholar] [CrossRef] [PubMed]
- Chunglok, W.; Wuragil, D.K.; Oaew, S.; Somasundrum, M.; Surareungchai, W. Immunoassay based on carbon nanotubes-enhanced ELISA for Salmonella enterica serovar Typhimurium. Biosens. Bioelectron. 2011, 26, 3584–3589. [Google Scholar] [CrossRef]
- Cho, I.H.; Irudayaraj, J. In-situ immuno-gold nanoparticle network ELISA biosensors for pathogen detection. Int. J. Food Microbiol. 2013, 164, 70–75. [Google Scholar] [CrossRef]
- Kumar, S.; Balakrishna, K.; Batra, H.V. Enrichment-ELISA for Detection of Salmonella typhi From Food and Water Samples. Biomed. Environ. Sci. 2008, 21, 137–143. [Google Scholar] [CrossRef]
- Ohk, S.H.; Bhunia, A.K. Multiplex fiber optic biosensor for detection of Listeria monocytogenes, Escherichia coli O157:H7 and Salmonella enterica from ready-to-eat meat samples. Food Microbiol. 2013, 33, 166–171. [Google Scholar] [CrossRef]
- Bruno, J.G. Application of DNA aptamers and quantum dots to lateral flow test strips for detection of foodborne pathogens with improved sensitivity versus colloidal gold. Pathogens 2014, 3, 341–355. [Google Scholar] [CrossRef]
- Jin, B.; Yang, Y.; He, R.; Park, Y.I.; Lee, A.; Bai, D.; Li, F.; Lu, T.J.; Xu, F.; Lin, M. Lateral flow aptamer assay integrated smartphone-based portable device for simultaneous detection of multiple targets using upconversion nanoparticles. Sens. Actuators B Chem. 2018, 276, 48–56. [Google Scholar] [CrossRef]
- Fronczek, C.F.; You, D.J.; Yoon, J.Y. Single-pipetting microfluidic assay device for rapid detection of Salmonella from poultry package. Biosens. Bioelectron. 2013, 40, 342–349. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Knoll, W.; Dostalek, J. Bacterial pathogen surface plasmon resonance biosensor advanced by long range surface plasmons and magnetic nanoparticle assays. Anal. Chem. 2012, 84, 8345–8350. [Google Scholar] [CrossRef] [PubMed]
- Park, T.S.; Yoon, J.Y. Smartphone Detection of Escherichia coli From Field Water Samples on Paper Microfluidics. IEEE Sens. J. 2015, 15, 1902–1907. [Google Scholar] [CrossRef]
- Sharma, H.; Mutharasan, R. Rapid and sensitive immunodetection of Listeria monocytogenes in milk using a novel piezoelectric cantilever sensor. Biosens. Bioelectron. 2013, 45, 158–162. [Google Scholar] [CrossRef]
- Ozalp, V.C.; Bayramoglu, G.; Erdem, Z.; Arica, M.Y. Pathogen detection in complex samples by quartz crystal microbalance sensor coupled to aptamer functionalized core-shell type magnetic separation. Anal. Chim. Acta 2015, 853, 533–540. [Google Scholar] [CrossRef]
- Rahman, M.R.T.; Lou, Z.; Wang, H.; Ai, L. Aptamer immobilized magnetoelastic sensor for the determination of Staphylococcus aureus. Anal. Lett. 2015, 48, 2414–2422. [Google Scholar] [CrossRef]
- Salam, F.; Uludag, Y.; Tothill, I.E. Real-time and sensitive detection of Salmonella Typhimurium using an automated quartz crystal microbalance (QCM) instrument with nanoparticles amplification. Talanta 2013, 115, 761–767. [Google Scholar] [CrossRef]
- Campbell, G.A.; Mutharasan, R. A Method of measuring Escherichia coli O157:H7 at 1 Cell/mL in 1 liter sample using antibody functionalized piezoelectric-excited millimeter-sized cantilever sensor. Environ. Sci. Technol. 2007, 41, 1668–1674. [Google Scholar] [CrossRef]
- Wan, Y.; Lin, Z.; Zhang, D.; Wang, Y.; Hou, B. Impedimetric immunosensor doped with reduced graphene sheets fabricated by controllable electrodeposition for the non-labelled detection of bacteria. Biosens. Bioelectron. 2011, 26, 1959–1964. [Google Scholar] [CrossRef]
- Wan, Y.; Zhang, D.; Wang, Y.; Hou, B. A 3D-impedimetric immunosensor based on foam Ni for detection of sulfate-reducing bacteria. Electrochem. Commun. 2010, 12, 288–291. [Google Scholar] [CrossRef]
- La Belle, J.T.; Shah, M.; Reed, J.; Nandakumar, V.; Alford, T.L.; Wilson, J.W.; Nickerson, C.A.; Joshi, L. Label-Free and Ultra-Low Level Detection of Salmonella enterica Serovar Typhimurium Using Electrochemical Impedance Spectroscopy. Electroanalysis 2009, 21, 2267–2271. [Google Scholar] [CrossRef]
- Esteban-Fernández de Ávila, B.; Pedrero, M.; Campuzano, S.; Escamilla-Gómez, V.; Pingarrón, J.M. Sensitive and rapid amperometric magnetoimmunosensor for the determination of Staphylococcus aureus. Anal. Bioanal. Chem. 2012, 403, 917–925. [Google Scholar] [CrossRef] [PubMed]
- Tan, F.; Leung, P.H.M.; Liu, Z.-b.; Zhang, Y.; Xiao, L.; Ye, W.; Zhang, X.; Yi, L.; Yang, M. A PDMS microfluidic impedance immunosensor for E. coli O157:H7 and Staphylococcus aureus detection via antibody-immobilized nanoporous membrane. Sens. Actuators B Chem. 2011, 159, 328–335. [Google Scholar] [CrossRef]
- Chemburu, S.; Wilkins, E.; Abdel-Hamid, I. Detection of pathogenic bacteria in food samples using highly-dispersed carbon particles. Biosens. Bioelectron. 2005, 21, 491–499. [Google Scholar] [CrossRef] [PubMed]
- Das, R.; Dhiman, A.; Kapil, A.; Bansal, V.; Sharma, T.K. Aptamer-mediated colorimetric and electrochemical detection of Pseudomonas aeruginosa utilizing peroxidase-mimic activity of gold NanoZyme. Anal. Bioanal. Chem. 2019, 411, 1229–1238. [Google Scholar] [CrossRef]
- de la Rica, R.; Baldi, A.; Fernández-Sánchez, C.; Matsui, H. Selective Detection of Live Pathogens via Surface-Confined Electric Field Perturbation on Interdigitated Silicon Transducers. Anal. Chem. 2009, 81, 3830–3835. [Google Scholar] [CrossRef]
- Joung, C.-K.; Kim, H.-N.; Im, H.-C.; Kim, H.-Y.; Oh, M.-H.; Kim, Y.-R. Ultra-sensitive detection of pathogenic microorganism using surface-engineered impedimetric immunosensor. Sens. Actuators B Chem. 2012, 161, 824–831. [Google Scholar] [CrossRef]
- Wang, Y.; Ping, J.; Ye, Z.; Wu, J.; Ying, Y. Impedimetric immunosensor based on gold nanoparticles modified graphene paper for label-free detection of Escherichia coli O157:H7. Biosens. Bioelectron. 2013, 49, 492–498. [Google Scholar] [CrossRef]
- Maalouf, R.; Fournier-Wirth, C.; Coste, J.; Chebib, H.; Saïkali, Y.; Vittori, O.; Errachid, A.; Cloarec, J.-P.; Martelet, C.; Jaffrezic-Renault, N. Label-free detection of bacteria by electrochemical impedance spectroscopy: comparison to surface plasmon resonance. Anal. Chem. 2007, 79, 4879–4886. [Google Scholar] [CrossRef]
- Neufeld, T.; Schwartz-Mittelmann, A.; Biran, D.; Ron, E.Z.; Rishpon, J. Combined phage typing and amperometric detection of released enzymatic activity for the specific identification and quantification of bacteria. Anal. Chem. 2003, 75, 580–585. [Google Scholar] [CrossRef]
- dos Santos, M.B.; Azevedo, S.; Agusil, J.P.; Prieto-Simón, B.; Sporer, C.; Torrents, E.; Juárez, A.; Teixeira, V.; Samitier, J. Label-free ITO-based immunosensor for the detection of very low concentrations of pathogenic bacteria. Bioelectrochemistry 2015, 101, 146–152. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Kulkarni, A.; Doepke, A.; Halsall, H.B.; Iyer, S.; Heineman, W.R. Carbohydrate-based label-free detection of Escherichia coli ORN 178 using electrochemical impedance spectroscopy. Anal. Chem. 2012, 84, 241–246. [Google Scholar] [CrossRef] [PubMed]
- Sedki, M.; Chen, X.; Chen, C.; Ge, X.; Mulchandani, A. Non-lytic M13 phage-based highly sensitive impedimetric cytosensor for detection of coliforms. Biosens. Bioelectron. 2020, 148, 111794. [Google Scholar] [CrossRef] [PubMed]
- Yao, L.; Wang, L.; Huang, F.; Cai, G.; Xi, X.; Lin, J. A microfluidic impedance biosensor based on immunomagnetic separation and urease catalysis for continuous-flow detection of E. coli O157:H7. Sens. Actuators B Chem. 2018, 259, 1013–1021. [Google Scholar] [CrossRef]
- Pankratov, D.; Bendixen, M.; Shipovskov, S.; Gosewinkel, U.; Ferapontova, E. Cellulase-linked immunomagnetic microbial assay on electrodes: Specific and sensitive detection of a single bacterial cell Anal. Chem. 2020, 92, 12451–12459. [Google Scholar] [CrossRef]
- Chan, K.Y.; Ye, W.W.; Zhang, Y.; Xiao, L.D.; Leung, P.H.; Li, Y.; Yang, M. Ultrasensitive detection of E. coli O157:H7 with biofunctional magnetic bead concentration via nanoporous membrane based electrochemical immunosensor. Biosens. Bioelectron. 2013, 41, 532–537. [Google Scholar] [CrossRef]
- Hua, R.; Hao, N.; Lu, J.; Qian, J.; Liu, Q.; Li, H.; Wang, K. A sensitive Potentiometric resolved ratiometric photoelectrochemical aptasensor for Escherichia coli detection fabricated with non-metallic nanomaterials. Biosens. Bioelectron. 2018, 106, 57–63. [Google Scholar] [CrossRef]
- Joung, C.K.; Kim, H.N.; Lim, M.C.; Jeon, T.J.; Kim, H.Y.; Kim, Y.R. A nanoporous membrane-based impedimetric immunosensor for label-free detection of pathogenic bacteria in whole milk. Biosens. Bioelectron. 2013, 44, 210–305. [Google Scholar] [CrossRef]
- Burrs, S.L.; Bhargava, M.; Sidhu, R.; Kiernan-Lewis, J.; Gomes, C.; Claussen, J.C.; McLamore, E.S. A paper based graphene-nanocauliflower hybrid composite for point of care biosensing. Biosens. Bioelectron. 2016, 85, 479–487. [Google Scholar] [CrossRef]
- Laczka, O.; Maesa, J.M.; Godino, N.; del Campo, J.; Fougt-Hansen, M.; Kutter, J.P.; Snakenborg, D.; Munoz-Pascual, F.X.; Baldrich, E. Improved bacteria detection by coupling magneto-immunocapture and amperometry at flow-channel microband electrodes. Biosens. Bioelectron. 2011, 26, 3633–3640. [Google Scholar] [CrossRef]
- Li, Y.; Fang, L.; Cheng, P.; Deng, J.; Jiang, L.; Huang, H.; Zheng, J. An electrochemical immunosensor for sensitive detection of Escherichia coli O157:H7 using C60 based biocompatible platform and enzyme functionalized Pt nanochains tracing tag. Biosens. Bioelectron. 2013, 49, 485–491. [Google Scholar] [CrossRef] [PubMed]
- dos Santos, M.B.; Agusil, J.P.; Prieto-Simon, B.; Sporer, C.; Teixeira, V.; Samitier, J. Highly sensitive detection of pathogen Escherichia coli O157:H7 by electrochemical impedance spectroscopy. Biosens. Bioelectron. 2013, 45, 174–180. [Google Scholar] [CrossRef]
- Altintas, Z.; Akgun, M.; Kokturk, G.; Uludag, Y. A fully automated microfluidic-based electrochemical sensor for real-time bacteria detection. Biosens. Bioelectron. 2018, 100, 541–548. [Google Scholar] [CrossRef] [PubMed]
- Tian, F.; Lyu, J.; Shi, J.; Tan, F.; Yang, M. A polymeric microfluidic device integrated with nanoporous alumina membranes for simultaneous detection of multiple foodborne pathogens. Sens. Actuators B Chem. 2016, 225, 312–318. [Google Scholar] [CrossRef]
- Zhang, F.; Keasling, J. Biosensors and their applications in microbial metabolic engineering. Trends Micorbiol. 2011, 19, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-P.; Lei, Q.-Y. Metabolite sensing and signaling in cell metabolism. Signal Transduct. Target. Ther. 2018, 3, 30. [Google Scholar] [CrossRef]
- Macario, A.J.L.; de MacArio, E.C. Monoclonal Antibodies against Bacteria; Academic Press: Orlando, FL, USA, 1986. [Google Scholar]
- Wang, L.; Wang, R.; Wei, H.; Li, Y. Selection of aptamers against pathogenic bacteria and their diagnostics application. World J. Microbiol. Biotechnol. 2018, 34, 149. [Google Scholar] [CrossRef]
- Banada, P.P.; Bhunia, A.K. Antibodies and Immunoassays for Detection of Bacterial Pathogens. In Principles of Bacterial Detection: Biosensors, Recognition Receptors and Microsystems; Zourob, M., Elwary, S., Turner, A., Eds.; Springer: New York, NY, USA, 2008; pp. 567–602. [Google Scholar]
- Li, L.; Li, Q.; Liao, Z.; Sun, Y.; Cheng, Q.; Song, Y.; Song, E.; Tan, W. Magnetism-resolved separation and fluorescence quantification for near-simultaneous detection of multiple pathogens. Anal. Chem. 2018, 90, 9621–9628. [Google Scholar] [CrossRef]
- Yu, M.; Wang, H.; Fu, F.; Li, L.; Li, J.; Li, G.; Song, Y.; Swihart, M.T.; Song, E. Dual-recognition Förster Resonance Energy Transfer based platform for one-step sensitive detection of pathogenic bacteria using fluorescent vancomycin–gold nanoclusters and aptamer–gold nanoparticles. Anal. Chem. 2017, 89, 4085–4090. [Google Scholar] [CrossRef]
- Hayman, R.B. Fiber Optic Biosensors for Bacterial Detection. In Principles of Bacterial Detection: Biosensors, Recognition Receptors and Microsystems; Zourob, M., Elwary, S., Turner, A., Eds.; Springer: New York, NY, USA, 2008; pp. 125–137. [Google Scholar]
- Eltzov, E.; Marks, R.S. Miniaturized flow stacked immunoassay for detecting Escherichia coli in a single step. Anal. Chem. 2016, 88, 6441–6449. [Google Scholar] [CrossRef]
- Verma, J.; Saxena, S.; Babu, S.G. ELISA-Based Identification and Detection of Microbes. In Analyzing Microbes; Springer: Berlin/Heidelberg, Germany, 2013; pp. 169–186. [Google Scholar]
- Anton, G.; Wilson, R.; Yu, Z.H.; Prehn, C.; Zukunft, S.; Adamski, J.; Heier, M.; Meisinger, C.; Romisch-Margl, W.; Wang-Sattler, R.; et al. Pre-analytical sample quality: Metabolite ratios as an intrinsic marker for prolonged room temperature exposure of serum samples. PLoS ONE 2015, 10, e0121495. [Google Scholar] [CrossRef] [PubMed]
- Henning, D.; Nielsen, K. Peroxidase-labelled monoclonal antibodies for use in enzyme immunoassay. J. Immunoass. 1987, 8, 297–307. [Google Scholar] [CrossRef] [PubMed]
- Gosling, J.P. A decade of development in immunoassay methodology. Clin. Chem. 1990, 36, 1408–1427. [Google Scholar] [CrossRef] [PubMed]
- Porstmann, T.; Kiessig, S.T. Enzyme immunoassay techniques. An overview. J. lmmunol. Methods 1992, 150, 5–21. [Google Scholar] [CrossRef]
- Khatkhatay, M.I.; Desai, M. A comparison of performances of four enzymes used in ELISA with special reference to beta-lactamase. J. Immunoass. 1999, 20, 151–183. [Google Scholar] [CrossRef]
- Liu, C.; Skaldin, M.; Wu, C.; Lu, Y.; Zavialov, A.V. Application of ADA1 as a new marker enzyme in sandwich ELISA to study the effect of adenosine on activated monocytes. Sci. Rep. 2016, 6, 31370. [Google Scholar] [CrossRef]
- Qu, Z.; Xu, H.; Xu, P.; Chen, K.; Mu, R.; Fu, J.; Gu, H. Ultrasensitive ELISA using enzyme-loaded nanospherical brushes as labels. Anal. Chem. 2014, 86, 9367–9371. [Google Scholar] [CrossRef]
- Engvall, E. The ELISA, enzyme-linked immunosorbent assay. Clin. Chem. 2010, 56, 319–320. [Google Scholar] [CrossRef]
- Elder, B.L.; Boraker, D.K.; Fives-Taylor, P.M. Whole-bacterial cell enzyme-linked immunosorbent assay for Streptococcus sanguis fimbrial antigens. J. Clin. Microbiol. 1982, 16, 141–144. [Google Scholar] [CrossRef]
- Aydin, S. A short history, principles, and types of ELISA, and our laboratory experience with peptide/protein analyses using ELISA. Peptides 2015, 72, 4–15. [Google Scholar] [CrossRef]
- Mirhosseini, S.A.; Fooladi, A.A.I.; Amani, J.; Sedighian, H. Production of recombinant flagellin to develop ELISA-based detection of Salmonella Enteritidis. Braz. J. Microbiol. 2017, 48, 774–781. [Google Scholar] [CrossRef] [PubMed]
- Hans, R.; Yadav, P.K.; Sharma, P.K.; Boopathi, M.; Thavaselvam, D. Development and validation of immunoassay for whole cell detection of Brucella abortus and Brucella melitensis. Sci. Rep. 2020, 10, 8543. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; He, J.; Cao, X.; Huang, K.; Luo, Y.; Xu, W. Development of a double-antibody sandwich ELISA for rapid detection of Bacillus Cereus in food. Sci. Rep. 2016, 6, 16092. [Google Scholar] [CrossRef] [PubMed]
- Sharifi, S.; Vahed, S.Z.; Ahmadian, E.; Dizaj, S.M.; Eftekhari, A.; Khalilov, R.; Ahmadi, M.; Hamidi-Asl, E.; Labib, M. Detection of pathogenic bacteria via nanomaterials-modified aptasensors. Biosens. Bioelectron. 2020, 150, 111933. [Google Scholar] [CrossRef]
- Galikowska, E.; Kunikowska, D.; Tokarska-Pietrzak, E.; Dziadziuszko, H.; Los, J.M.; Golec, P.; Wegrzyn, G.; Los, M. Specific detection of Salmonella enterica and Escherichia coli strains by using ELISA with bacteriophages as recognition agents. Eur. J. Clin. Microbiol. Infect. Dis. 2011, 30, 1067–1073. [Google Scholar] [CrossRef]
- Eltzov, E.; Guttel, S.; Kei, A.L.Y.; Sinawang, P.D.; Ionescu, R.E.; Marks, R.S. Lateral flow immunoassays—From paper strip to smartphone technology. Electroanalysis 2015, 27, 2116–2130. [Google Scholar] [CrossRef]
- Zhao, Y.; Wang, H.; Zhang, P.; Sun, C.; Wang, X.; Wang, X.; Yang, R.; Wang, C.; Zhou, L. Rapid multiplex detection of 10 foodborne pathogens with an up-converting phosphor technology-based 10-channel lateral flow assay. Sci. Rep. 2016, 6, 21342. [Google Scholar] [CrossRef]
- Kim, H.J.; Kwon, C.; Lee, B.S.; Noh, H. One-step sensing of foodborne pathogenic bacteria using a 3D paper-based device. Analyst 2019, 144, 2248–2255. [Google Scholar] [CrossRef]
- Jung, B.Y.; Jung, S.C.; Kweon, C.H. Development of a rapid immunochromatographic strip for detection of Escherichia coli O157. J. Food Prot. 2005, 68, 2140–2143. [Google Scholar] [CrossRef]
- Xu, D.; Wu, X.; Li, B.; Li, P.; Ming, X.; Chen, T.; Wei, H.; Xu, F. Rapid detection of Campylobacter jejuni using fluorescent microspheres as label for immunochromatographic strip test. Food Sci. Biotechnol. 2013, 2, 585–591. [Google Scholar] [CrossRef]
- Niu, K.; Zheng, X.; Huang, C.; Xu, K.; Zhi, Y.; Shen, H.; Jia, N. A colloidal gold nanoparticle-based immunochromatographic test strip for rapid and convenient detection of Staphylococcus aureus. J. Nanosci. Nanotechnol. 2014, 14, 5151–5156. [Google Scholar] [CrossRef] [PubMed]
- Ferapontova, E.E. Electrochemical assays for microbial analysis: How far they are from solving microbiota and microbiome challenges. Curr. Opin. Electrochem. 2020, 19, 153–161. [Google Scholar] [CrossRef]
- Kuss, S.; Amin, H.M.A.; Compton, R.G. Electrochemical detection of pathogenic bacteria—Recent strategies, advances and challenges. Chem. Asian J. 2018, 13, 2758–2769. [Google Scholar] [CrossRef]
- Campuzano, S.; de Ávila, B.E.-F.; Yuste, J.; Pedrero, M.; García, J.L.; García, P.; García, E.; Pingarrón, J.M. Disposable amperometric magnetoimmunosensors for the specific detection of Streptococcus pneumoniae. Biosens. Bioelectron. 2010, 26, 1225–1230. [Google Scholar] [CrossRef]
- Liébana, S.; Lermo, A.; Campoy, S.; Cortés, M.P.; Alegret, S.; Pividori, M.I. Rapid detection of Salmonella in milk by electrochemical magneto-immunosensing. Biosens. Bioelectron. 2009, 25, 510–513. [Google Scholar] [CrossRef] [PubMed]
- Ruan, C.; Yang, L.; Li, Y. Immunobiosensor chips for detection of Escherichia coli O157:H7 using electrochemical impedance spectroscopy. Anal. Chem. 2002, 74, 4814–4820. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhou, D. Magnetic particle-based ultrasensitive biosensors for diagnostics. Expert Rev. Mol. Diagn. 2012, 12, 565–571. [Google Scholar] [CrossRef]
- Nemr, C.R.; Smith, S.J.; Liu, W.; Mepham, A.H.; Mohamadi, R.M.; Labib, M.; Kelley, S.O. Nanoparticle-mediated capture and electrochemical detection of methicillin-resistant Staphylococcus aureus. Anal. Chem. 2019, 91, 2847–2853. [Google Scholar] [CrossRef]
- Ferapontova, E.E.; Hansen, M.N.; Saunders, A.M.; Shipovskov, S.; Sutherland, D.S.; Gothelf, K.V. Electrochemical DNA sandwich assay with a lipase label for attomole detection of DNA. Chem. Commun. 2010, 46, 1836–1838. [Google Scholar] [CrossRef]
- Fapyane, D.; Ferapontova, E.E. Electrochemical assay for a total cellulase activity with improved sensitivity. Anal. Chem. 2017, 89, 3959–3965. [Google Scholar] [CrossRef]
- Fapyane, D.; Nielsen, J.S.; Ferapontova, E.E. Electrochemical enzyme-linked sandwich assay with a cellulase label for ultrasensitive analysis of synthetic DNA and cell-isolated RNA. ACS Sens. 2018, 3, 2104–2111. [Google Scholar] [CrossRef] [PubMed]
- Malecka, K.; Pankratov, D.; Ferapontova, E.E. Femtomolar electroanalysis of a breast cancer biomarker HER-2/neu protein in human serum by the cellulase-linked sandwich assay on magnetic beads. Anal. Chim. Acta 2019, 1077, 140–149. [Google Scholar] [CrossRef] [PubMed]
- Zhong, M.; Yang, L.; Yang, H.; Cheng, C.; Deng, W.; Tan, Y.; Xie, Q.; Yao, S. An electrochemical immunobiosensor for ultrasensitive detection of Escherichia coli O157:H7 using CdS quantum dots-encapsulated metal-organic frameworks as signal-amplifying tags. Biosens. Bioelectron. 2019, 126, 493–500. [Google Scholar] [CrossRef] [PubMed]
- Kuss, S.; Couto, R.A.S.; Evans, R.M.; Lavender, H.; Tang, C.C.; Compton, R.G. Versatile electrochemical sensing platform for bacteria. Anal. Chem. 2019, 91, 4317–4322. [Google Scholar] [CrossRef]
- Zhou, Y.; Marar, A.; Kner, P.; Ramasamy, R.P. Charge-directed immobilization of bacteriophage on nanostructured electrode for whole-cell electrochemical biosensors. Anal. Chem. 2017, 89, 5734–5741. [Google Scholar] [CrossRef]
- Jafari, H.; Amiri, M.; Abdi, E.; Navid, S.L.; Bouckaert, J.; Jijie, R.; Boukherroub, R.; Szunerits, S. Entrapment of uropathogenic E. coli cells into ultra-thin sol-gel matrices on gold thin films: A low cost alternative for impedimetric bacteria sensing. Biosens. Bioelectron. 2019, 124–125, 161–166. [Google Scholar] [CrossRef]
- Ranjbar, S.; Nejad, M.A.F.; Parolo, C.; Shahrokhian, S.; Merkoçi, A. Smart chip for visual detection of bacteria using the electrochromic properties of polyaniline. Anal. Chem. 2019, 91, 14960–14966. [Google Scholar] [CrossRef]
- Cheng, M.S.; Lau, S.H.; Chow, V.T.; Toh, C.-S. Membrane-based electrochemical nanobiosensor for Escherichia coli detection and analysis of cells viability. Environ. Sci. Technol. 2011, 45, 6453–6459. [Google Scholar] [CrossRef]
- Ahmed, A.; Rushworth, J.V.; Wright, J.D.; Millner, P.A. Novel impedimetric immunosensor for detection of pathogenic bacteria Streptococcus pyogenes in human saliva. Anal. Chem. 2013, 85, 12118–12125. [Google Scholar] [CrossRef]
- Chiriacò, M.S.; Parlangeli, I.; Sirsi, F.; Poltronieri, P.; Primiceri, E. Impedance sensing platform for detection of the food pathogen Listeria monocytogenes. Electronics 2018, 7, 347. [Google Scholar] [CrossRef]
- Labib, M.; Zamay, A.S.; Kolovskaya, O.S.; Reshetneva, I.T.; Zamay, G.S.; Kibbee, R.J.; Sattar, S.A.; Zamay, T.N.; Berezovski, M.V. Aptamer-based impedimetric sensor for bacterial typing. Anal. Chem. 2012, 84, 8114–8117. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Fu, Y.; Fang, W.; Li, Y. Electrochemical impedance immunosensor based on self-assembled monolayers for rapid detection of Escherichia coli O157:H7 with signal amplification using lectin. Sensors 2015, 15, 19212–19224. [Google Scholar] [CrossRef] [PubMed]
- Jasim, I.; Shen, Z.; Mlaji, Z.; Yuksek, N.S.; Abdullah, A.; Liu, J.; Dastider, S.G.; El-Dweik, M.; Zhang, S.; Almasri, M. An impedance biosensor for simultaneous detection of low concentration of Salmonella serogroups in poultry and fresh produce samples. Biosens. Bioelectron. 2019, 126, 292–300. [Google Scholar] [CrossRef] [PubMed]
- Brosel-Oliu, S.; Ferreira, R.; Uria, N.; Abramova, N.; Gargallo, R.; Muñoz-Pascual, F.-X.; Bratov, A. Novel impedimetric aptasensor for label-free detection of Escherichia coli O157:H7. Sens. Actuators B Chem. 2018, 255, 2988–2995. [Google Scholar] [CrossRef]
- Salimian, R.; Kékedy-Nagy, L.; Ferapontova, E.E. Specific picomolar detection of a breast cancer biomarker HER-2/neu protein in serum: Electrocatalytically amplified electroanalysis by the aptamer/PEG-modified electrode. Chem. Electro. Chem. 2017, 4, 872–879. [Google Scholar] [CrossRef]
- Jijie, R.; Kahlouche, K.; Barras, A.; Yamakawa, N.; Bouckaert, J.; Gharbi, T.; Szunerits, S.; Boukherroub, R. Reduced graphene oxide/polyethylenimine based immunosensor for the selective and sensitive electrochemical detection of uropathogenic Escherichia coli. Sens. Actuators B Chem. 2018, 260, 255–263. [Google Scholar] [CrossRef]
- Qiao, Z.; Fu, Y.; Lei, C.; Li, Y. Advances in antimicrobial peptides-based biosensing methods for detection of foodborne pathogens: A review. Food Control 2020, 112, 107116. [Google Scholar] [CrossRef]
- Mannoor, M.S.; Zhang, S.; Link, A.J.; McAlpine, M.C. Electrical detection of pathogenic bacteria via immobilized antimicrobial peptides. Proc. Natl. Acad. Sci. USA 2010, 107, 19207–19212. [Google Scholar] [CrossRef]
- Janczuk-Richter, M.; Marinović, I.; Niedziółka-Jönsson, J.; Szot-Karpińska, K. Recent applications of bacteriophage-based electrodes: A mini-review. Electrochem. Commun. 2019, 99, 11–15. [Google Scholar] [CrossRef]
- Moghtader, F.; Congur, G.; Zareie, H.M.; Erdem, A.; Piskin, E. Impedimetric detection of pathogenic bacteria with bacteriophages using gold nanorod deposited graphite electrodes. RSC Adv. 2016, 6, 97832–97839. [Google Scholar] [CrossRef]
- Richter, Ł.; Bielec, K.; Leśniewski, A.; Łoś, M.; Paczesny, J.; Hołyst, R. Dense layer of bacteriophages ordered in alternating electric field and immobilized by surface chemical modification as sensing element for bacteria detection. ACS Appl. Mater. Interfaces 2017, 9, 19622–19629. [Google Scholar] [CrossRef]
- Niyomdecha, S.; Limbut, W.; Numnuam, A.; Kanatharana, P.; Charlermroj, R.; Karoonuthaisiri, N.; Thavarungkul, P. Phage-based capacitive biosensor for Salmonella detection. Talanta 2018, 188, 658–664. [Google Scholar] [CrossRef]
- Golabi, M.; Kuralay, F.; Jager, E.W.H.; Beni, V.; Turner, A.P.F. Electrochemical bacterial detection using poly(3-aminophenylboronic acid)-based imprinted polymer. Biosens. Bioelectron. 2017, 93, 87–93. [Google Scholar] [CrossRef] [PubMed]
- MagaÑA, S.; Schlemmer, S.M.; Leskinen, S.D.; Kearns, E.A.; Lim, D.V. Automated Dead-End Ultrafiltration for Concentration and Recovery of Total Coliform Bacteria and Laboratory-Spiked Escherichia coli O157:H7 from 50-Liter Produce Washes To Enhance Detection by an Electrochemiluminescence Immunoassay. J. Food Prot. 2013, 76, 1152–1160. [Google Scholar] [CrossRef] [PubMed]
- Spehar-Délèze, A.-M.; Julich, S.; Gransee, R.; Tomaso, H.; Dulay, S.B.; O’Sullivan, C.K. Electrochemiluminescence (ECL) immunosensor for detection of Francisella tularensis on screen-printed gold electrode array. Anal. Bioanal. Chem. 2016, 408, 7147–7153. [Google Scholar] [CrossRef]
- Guo, Z.; Sha, Y.; Hu, Y.; Yu, Z.; Tao, Y.; Wu, Y.; Zeng, M.; Wang, S.; Li, X.; Zhou, J.; et al. Faraday cage-type electrochemiluminescence immunosensor for ultrasensitive detection of Vibrio vulnificus based on multi-functionalized graphene oxide. Anal. Bioanal. Chem. 2016, 408, 7203–7211. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Chen, X.; Zhang, L.; Gao, J.; Ma, Q. Electrochemiluminescence detection of Escherichia coli O157:H7 Based on a novel polydopamine surface imprinted polymer biosensor. ACS Appl. Mater. Interfaces 2017, 9, 5430–5436. [Google Scholar] [CrossRef] [PubMed]
- Hao, N.; Zhang, X.; Zhou, Z.; Hua, R.; Zhang, Y.; Liu, Q.; Qian, J.; Li, H.; Wang, K. AgBr nanoparticles/3D nitrogen-doped graphene hydrogel for fabricating all-solid-state luminol-electrochemiluminescence Escherichia coli aptasensors. Biosens. Bioelectron. 2017, 97, 377–383. [Google Scholar] [CrossRef]
- Yue, H.; He, Y.; Fan, E.; Wang, L.; Lu, S.; Fu, Z. Label-free electrochemiluminescent biosensor for rapid and sensitive detection of Pseudomonas aeruginosa using phage as highly specific recognition agent. Biosens. Bioelectron. 2017, 94, 429–432. [Google Scholar] [CrossRef]
- Kulasingam, V.; Jung, B.P.; Blasutig, I.M.; Baradaran, S.; Chan, M.K.; Aytekin, M.; Colantonio, D.A.; Adeli, K. Pediatric reference intervals for 28 chemistries and immunoassays on the Roche cobas® 6000 analyzer—A CALIPER pilot study. Clin. Biochem. 2010, 43, 1045–1050. [Google Scholar] [CrossRef]
- Reta, N.; Michelmore, A.; Saint, C.P.; Prieto-Simon, B.; Voelcker, N.H. Label-free bacterial toxin detection in water supplies using porous silicon nanochannel sensors. ACS Sens. 2019, 4, 1515–1523. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Zheng, Q.; Lin, J.; Yuk, H.-G.; Guo, L. Immuno- and nucleic acid-based current technologies for Salmonella detection in food. Eur. Food Res. Technol. 2020, 246, 373–395. [Google Scholar] [CrossRef]
- Ohlsson, P.; Evander, M.; Petersson, K.; Mellhammar, L.; Lehmusvuori, A.; Karhunen, U.; Soikkeli, M.; Seppä, T.; Tuunainen, E.; Spangar, A.; et al. Integrated Acoustic Separation, Enrichment, and Microchip Polymerase Chain Reaction Detection of Bacteria from Blood for Rapid Sepsis Diagnostics. Anal. Chem. 2016, 88, 9403–9411. [Google Scholar] [CrossRef] [PubMed]
- Mettakoonpitak, J.; Boehle, K.; Nantaphol, S.; Teengam, P.; Adkins, J.A.; Srisa-Art, M.; Henry, C.S. Electrochemistry on Paper-based Analytical Devices: A Review. Electroanalysis 2016, 28, 1420–1436. [Google Scholar] [CrossRef]
- Shen, L.-L.; Zhang, G.-R.; Etzold, B.J.M. Paper-Based Microfluidics for Electrochemical Applications. ChemElectroChem 2020, 7, 10–30. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://www.pointofcare.abbott/int/en/offerings/istat/istat-test-cartridges (accessed on 25 September 2020).

| Strain | Technique | LOD a | Detection Range, CFU mL−1 | Interference Studies | Assay Time | Ref. |
|---|---|---|---|---|---|---|
| Campylobacter jejuni | PCR with pre-enrichment in suitable broths | 3 CFU/100 mL | - | Actinomyces pyogenes, Campylobacter coli, Enterobacter cloacae, Pseudomonas aeruginosa, Salmonella saintpaul, Yersinia enterocolitica | 48 h | [34] |
| Salmonella, Listeria monocytogene | qPCR with two fluorescently labelled primers | 5 CFU/25 mL | - | B. cerus, Campylobacter, E. aerogenes, E. cowanii, Cronobacter sakazakii, E. coli, E. faecalis, S. aureus, Shigella spp, Serratia liquefaciens, S. pneumoniae | <48 h | [35] |
| Salmonella | q-PCR with two-step pre-filtration on filter paper | 7.5 CFU/100 mL | - | - | 3 h | [36] |
| Salmonella typhimurium | direct PCR with immunomagnetic preconcentration | 2–3 | 6–6.4 × 104 | - | <3 h | [37] |
| Salmonella | PMA-qPCR | 36 (pure culture) and 100 (raw shrimp) | 36–3.6 × 108 (pure culture) and 100–1 × 108 (raw shrimp) | Vibrio parahaemolyticus, Listeria monocytogenes, E. coli O157:H7, S. aureus | 1–2 h | [38] |
| PMA was used to increase sensitivity | ||||||
| Salmonella sp. | Multiplex qPCR with immunomagnetic pre-concentration | 2 CFU/g | - | Listeria monocytogenes, E. coli, B. cerus, Streptomyces griseus, Pseudomonus aerug, Lactobacillus plantarum, E. faecalis, Streptococcus hemolyticus, Micrococcus luteus, P. aeruginosa, Clostridium sporogenes | <8 h | [39] |
| Shigella sp. | 6.8 CFU/g | |||||
| Staphylococcus aureus | 9.6 CFU/g | |||||
| Salmonella | Real-time RPA | 10 CFU/g (eggs) | - | Bacillus cereus, Campylobacter coli, E. coli O157:H7, L. casei, S. aureus, Pseudomonas aeruginosa, Vibrio vulnificus | 10 min | [40] |
| 100 CFU/g (chicken) | ||||||
| Salmonella | LAMP | 4.1 | - | Listeria monocytogenes, E. coli O157:H7, S. aureus, Yersinia enterocolitica, Proteus mirabilis, Shigella Flexner, Micrococcus luteus, Bacillus cereus, Enterobacter sakazakii, Pseudomonas fluorescens | 1 h | [41] |
| Vibrio parahaemolyticus | LAMP | 530 | - | Acinetobacter baumannii, Aeromonas hydrophila, Enterococcus faecalis, Haemophilus influenzae, Helicobacter pylori, Salmonella | 1 h | [42] |
| Salmonella and Shigella | Multiplexed LAMP Simultaneous detection of two bacterial species | 5 CFU/10 mL | - | S. aureus, E. coli, Bacillus cereus, Pseudomonas aeruginosa, Vibrio parahaemolyticus, Listeria monocytogenes | 20 h | [43] |
| E. coli | NASBA | 40 | - | Listeria monocytogenes, Shigella sonnei, Yersinia entero Colitica, Salmonella typhimurium | 40 min | [44] |
| Staphylococcus aureus | NASBA | 1–10 | - | Lactococcus lactis, Bacillus cereus, Listeria monocytogenes, Enterococcus faecalis, E. coli, Citrobacter freundii, Sal-monella, Streptococcus bovis, Klebsiella aerogenes | 3–4 h | [45] |
| Salmonella enteritidis. | FRET with CNP for signal enhancement | 150 | 100–3000 | Salmonella typhimurium, E. coli K88 | 2 h | [46] |
| Salmonella | DNA Micro-array | 2–8 CFU/g (tomato) | - | E. coli, Shigella, S. aureus, Pseudomonas aeruginosa, Citrobacter freundii, Vibrio cholera, Enterococcus fae-calis, Yersinia enterocolitica | <2 h | [47] |
| Salmonella | DNA Micro-array QD used in-place of fluorescent dyes | 10 | - | Vibrio parahaemolyticus, Vibrio fluvialis, Yersinia enterocolitica, Proteus sp.,S. aureus, Enterococcus faecalis, Campylobacter jejuni, β-hemolytic Streptococcus, Listeria monocytogenes | <2 h | [48] |
| Salmonella and Campylobacter | DNA Micro-array | 14–57 and 11–60 | - | Listeria monocytogenes, B. cereus, Cronobacter sakazakii, Citrobacter freundii, Klebsiella pneumonia, E. coli, Proteus vulgaris, Enterobacter aerogenes, Hafnia alvei, Serratia marcescens | 45 min | [49] |
| Strain/Analytical Scheme | Technique | LOD a, CFU mL−1 | Detection Range, CFU mL−1 | Interference Studies | Assay Time | Ref. |
|---|---|---|---|---|---|---|
| E. coli O157:H7Immunomagnetic assay, separation from complex matrix | Plate counting method | 16 | 1.6 × 101–7.2 × 107 | Salmonella enteritidis, Citrobacter freundii, Listeria monocytogenes | 15 min | [50] |
| Staphylococus aureus Aptaassay on microtiter plates | Colorimetric detection with AuNP as indicator | 9 | 10–106 | Vibrioparahemolyticus, Salmonella typhimurium, Streptococcus, E. coli, Enterobacter sakazakii, Listeria monocytogenes | 15 min | [51] |
| Pseudomonas aeruginosa Aptamer assay on MB | Fluorometric detection with magnetic separation | 1 | 10–108 | Listeria monocytogenes, S. aureus, Salmonella enterica, E. coli | 1.5 h | [52] |
| Vibrio cholerae O1 Sandwich immunoassay | Chromatographic | 5 × 105–106 | Shigella flexneri, Salmonella typhi, Pseudomonas aeruginosa, Proteus vulgaris, Klebsiella pneumonia, Enterobacter cloacae | 15 min | [53] | |
| E. coli O157:H7 Sandwich immunoassay | ELISA with HRP-TMB label and AuNP for signal amplification | 68 (PBS) | 6.8 × 102 (PBS) 6.8 × 103 (in food) | Salmonella senftenberg, Shigella sonnei, E. coli K12 | 3 h | [54] |
| Salmonella typhimurium Apta- and immunoassay on MB | Colorimetric; ELISA on MB with HRP/TMB, and AuNP for signal amplification | 1 × 103 | 1 × 103–1 × 108 | Salmonella typhi, Salmonella paratyphi, S. aureus, E. coli | 3 h | [55] |
| E. coli ATCC 8739 Apta-ssay on AuNP | FRET | 3 | 5–106 | E. coli DH5a, E. coli (ATCC 25922), Bacillus subtilis; S. aureus | - | [56] |
| Vibrio fischeri Sandwich aptaassay on paper | Colorimetric detection with AuNP | 40 | 40–4 × 105 | Vibrio parahemolyticus, E. coli, Bacillus subtilis, Shigella sonnei, S. aureus, Salmonella choleraesuis, Listeria monocytogenes | 10 min | [57] |
| E. coli O157:H7 Sandwich immunomagnetic assay | Fluorescence using pH sensitive fluorophore release detection labels | 15 | - | Streptococcus pneumoniae R6 | <3 h | [58] |
| E. coli O157:H7, Salmonella typhimurium, Listeria monocytogens Multiplex, Sandwich immunomagnetic assay | Fluorescence | <5 | - | No cross reactivity between target pathogens | 2 h | [59] |
| Salmonella enterica Sandwich and direct immunoassays | ELISA with CNT/HRP-TMB | 103 and 104 | - | - | 24 h (direct); 3 h (sandwich) | [60] |
| E. coli O157:H7, Salmonella typhimurium Sandwich immunomagnetic assay | ELISA with HRP/TMB and AuNP network for signal amplification | 3–15 | - | Listeria monocytogenes, Salmonella typhimurium, Salmonella enteritidis | 2 h | [61] |
| Salmonella enterica typhi Sandwich immunoassay with pre-enrichment in BPW | dot-ELISA, with Ab-HRP conjugate and 3,3 diaminobenzidine tetrahydrochloride | 104 before 102 after enrichment | - | - | 4 h, 10 h with enrichment | [62] |
| Listeria monocytogenes, E. coli O157:H7 and Salmonella enterica Sandwich immuno-fluorescence assay | Optical fiber; multiplexed simultaneous detection | 103 | - | Cross-reactivity tested with other target pathogens | <24 h | [63] |
| Escherichia coli Lateral flow aptaassay on QD | Colorimetric | 300–600 | - | Bacillus cereus, Enterococcus faecalis, Listeria monocytogenes, Salmonella enterica | 20 min | [64] |
| Salmonella Aptamer-based lateral flow assay | Colorimetric using up-conversion of NP for detection | 85 | 150–2000 | E. coli, S. aureus, Bacillus subtilis | 30 min | [65] |
| Salmonella typhimurium Immunoagglutination-based immunoassay | Optical Mie scattering of antigen-Ab clusters | 10 inconsistent with a 15 µL sample volume | 100–106 | - | 10 min (from 6 to 15 min) | [66] |
| E. coli O157:H7 Immunomagnetic pre-concentration | LRSP diffraction grated Au surface | 50 | 103 to 107 | E. coli K12 | 30 min | [67] |
| Escherichia coli Immunoassay in a paper-based microfluidic device | Optical Mie scattering of antigen-Ab clusters | 10 inconsistent with a 3.5 µL sample volume | 10 to 103 | - | 90 s | [68] |
| Salmonella typhimurium Sandwich immunoassay with magnetic pre-concentration | Fluorescence detection using QDNPs | 103 | 103–106 | E. coli | 30 min | [40] |
| RESONANCE-FREQUENCY-BASED IMMUNOASSAYS | ||||||
| Escherichia coli O157:H7 Immunoassay on Ab-modified glass b | Resonance frequency | 1 (in PBS) | - | - | 10 min | [69] |
| Salmonella enterica Aptamer-based assay on MB | Piezoelectric: QCM | 100 | 100–4 × 104 | E. coli | 40 min | [70] |
| S. aureus Aptamer-based assay | Magnetoelastic resonance frequency detection | 5 | 10–1 × 1011 | Listeria monocytogenes, E. coli, Enterobacter sakazakii, Streptococcus, Vibrio parahemolyticus | 25–26 min | [71] |
| Salmonella Sandwich immunoassay | Piezoelectric: QCM using AuNP labels for mass amplification | 10–20 | 10–105 | Klebsiella pneumonia, Enterobacteria spp, Pseudomona spp, S. aureus | 9 min | [72] |
| Listeria monocytogenes Sandwich immunoassay o | Resonance frequency detection on a sputtered gold/ lead-zirconate-titanate surface | 100 | 103–105 | - | 30 min | [73] |
| Strain/Analytical Scheme | Technique | LOD a, CFU mL−1 | Detection Range, CFU mL−1 | Interference Studies | Assay Time | Ref. |
|---|---|---|---|---|---|---|
| Sulphur reducing bacteria Immunoassay on chitosan doped rGS | EIS at 10 mV vs. Ag/AgCl with ferricyanide | 18 | 18–1.8 × 107 | Vibrio angillarum | - | [74] |
| Sulphur reducing bacteria Immunoassay on AuNP-modified Ni foam | EIS at 5 mV vs. Ag/AgC with ferricyanide | 21 | 2.1 × 101–2.1 × 107 | Vibrio anguillarum, E. coli | 2 h | [75] |
| Salmonella enterica Immunoassay on gold electrodes | EIS at 5 mV vs.Ag/AgCl with ferricyanide | 100 (10 CFU in 100 µL) | 100–10 × 104 | E. coli | 1.5 min (no data on incubation time) | [76] |
| S. aureus (protein A) Competitive magneto-immunoassay on TTF-AuSPE. | e-ELISA, HRP label; TTF mediator Amperometry at −0.15 V | 1(raw milk) | 1 to 107 | E. coli, Salmonella choleraesuis | 2 h | [77] |
| E. coli O157:H7, S. aureus Immunoassay on nano-porous alumina | EIS at 25 mV vs. Pt; no label | 100 | - | Both strains were used for specificity test | 2 h | [78] |
| E. coli, Listeria monocytogenes, Campylo-bacter jejuni Sandwich immunoassay with highly dispersed carbon particles | Electrochemical detection at 105 mV; HRP as a label, TMB as a substrate | 50, 10 and 50, respectively | 50–103, 10–1500, and 50–500 | - | 30 min | [79] |
| Pseudomonas aeruginosa Aptaassay on AuNP and AuNP/SPCE | Amperometry at 0.4 V with TMB | 60 | 60–60 × 107 | Vibrio cholera, Listeria monocytogens, S. aureus | 10 min (colorimetry) | [80] |
| E. coli IDE modified with anti-E. coli Ab | Impedance at 5 mV: no label, electric field perturbation | 300 | 102–104 | - | 1 h | [81] |
| E. coli O157:H7 Immunoassay at HA modified Au electrode | EIS with ferricyanide | 7 | 10–105 | S. aureus, Bacillus cereus, E. coli DH5a. | - | [82] |
| E. coli O157:H7 Immunoassay on AuNP modified rGO paper | EIS at 5 mV vs. Ag/AgCl with ferricyanide | 150 | 150–1.5 × 107 | S. aureus, Listeria monocytogenes, E. coli DH5a. | - | [83] |
| E. coli CIP 76.24 Immunoassay on polyclonal Ab/neuroavidin/SAM/Au | EIS with no indicator, at −0.6 V in aerated solutions | 10 | 10–105 and 103–107 for lysed cells | S. epidermis: Interference at ≥100 CFU mL−1 | 1 h incubation + detection | [84] |
| E. coli K12, MG1655 Phage typing & assaying activity of β-D-galactosidase in cell lysates, SPCE | Amperometry at 0.22 V, oxidation of enzymatically produced p-aminophenol | 1 CFU in 100 mL | 1–109 | Klebsiella pneumoniae | 6–8 h | [85] |
| E. coli O157:H7 Immunoassay on monoclonal Ab/ITO | EIS at 0.25 V with ferricyanide as a redox indicator | 10 | 10–106 | S. typhimurium, E. coli K12 | 0.8 h incubation + wash./detect. | [86] |
| E. coli ORN 178 Assay at carbohydrate modified SAM on Au | EIS at 5 mV vs. Ag/AgCl; with ferricyanide | 100 | 120 –2.5 × 103 | E. coli ORN 208 | <1 h | [87] |
| E. coli XL1-Blue; K12 Assay on non-lytic M13 phage/AuNP/GCE | EIS with ferricyanide redox indicator, at 0.15 V | 14 | 10–105 | Pseudomonas chlororaphis | 0.5 h incubation + wash./detect. | [88] |
| E. coli O157:H7 Sandwich immunoassay on MB at Au IDE, detected a response to urea hydrolysis by urease | EIS at 0 V, no indicator, label: urease/AuNP/aptamer; | 12 | 12–1.2 × 105 | S. typhimurium, Listeria monocytogenes | ca. 2 h | [89] |
| E. coli K12 and DH5α Sandwich immunoassay on MBs; on nitrocellulose modified Gr | Chronocoulometry at 0.3 V; no redox indicator; label: cellulase | 1 (PBS), 2 (milk) | 1–4 × 103 | E. agglomerans, S. aureus, Salmonella enteretidis, B. subtilis, P. putida | 3 h | [90] |
| E. coli O157:H7 Sandwich immunoassay on nanoporous alumina membrane | EIS at 25 mV/Pt; no label | 10 | 100–104 | - | - | [91] |
| E. coli Aptaassay on ITO modified with photoelectrochemical non-metallic NM | Potentiometric detection at 0.15 V (cathodic) and −0.4 V (anodic) (ratiometric detection) | 2.9 | 2.9–2.9 × 106 | - | 12 h | [92] |
| E. coli O157:H7 Immunoassay on nanoporous alumina membrane | EIS; no label | 10 (PBS) 83.7 (milk) | 10–105 | S. aureus, Bacillus cereus, E. coli DH5a. | - | [93] |
| E. coli O157:H7 Aptaassay on a paper modified with graphene nanoplatinum composite | EIS with ferricyanide indicator at 100 mV | 4 | 4–105 | - | 12 min | [94] |
| E. coli K12 Sandwich immunoassay | Amperometry at −0.35 V, HRP as a label; substrates: HQ/BQ andH2O2 | 55 (PBS) 100 (milk) | 102–108 | Pseudomonas putida | 1 h | [95] |
| E. coli O157:H7 Sandwich immunoassay with PtNCs coupled to GOD | Cyclic voltammetry from −0.15 V to 0.65 V | 15 | 32–3.2 × 106 | Salmonella typhi, Shigella dysenteriae Shigella flexneri | 30 min | [96] |
| E. coli O157:H7 Immunoassay on a SAM modified gold electrode | EIS with ferricyanide at 0 V vs. Ag/AgCl | 2 | 30–3 × 104 | Salmonella typhimurium | 45 min | [97] |
| E. coli Sandwich immunoassay on AuNP-structured electrode in an automated microfluidic chip | Ammperometry at –0.1 V, with an HRP label and TMB as a substrate | 50 | 50–106 | Shigella, Salmonella spp., Salmonella typhimurium, S. aureus | 30 min | [98] |
| E. coli O157:H7, S. aureus Nano-porous alumina membrane in a PEG-modified microfluiidc chip | Electrochemical impedance | 100 | 102–105 | E. coli O157:H7, S. aureus | <1 h | [99] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jamal, R.B.; Shipovskov, S.; Ferapontova, E.E. Electrochemical Immuno- and Aptamer-Based Assays for Bacteria: Pros and Cons over Traditional Detection Schemes. Sensors 2020, 20, 5561. https://doi.org/10.3390/s20195561
Jamal RB, Shipovskov S, Ferapontova EE. Electrochemical Immuno- and Aptamer-Based Assays for Bacteria: Pros and Cons over Traditional Detection Schemes. Sensors. 2020; 20(19):5561. https://doi.org/10.3390/s20195561
Chicago/Turabian StyleJamal, Rimsha Binte, Stepan Shipovskov, and Elena E. Ferapontova. 2020. "Electrochemical Immuno- and Aptamer-Based Assays for Bacteria: Pros and Cons over Traditional Detection Schemes" Sensors 20, no. 19: 5561. https://doi.org/10.3390/s20195561
APA StyleJamal, R. B., Shipovskov, S., & Ferapontova, E. E. (2020). Electrochemical Immuno- and Aptamer-Based Assays for Bacteria: Pros and Cons over Traditional Detection Schemes. Sensors, 20(19), 5561. https://doi.org/10.3390/s20195561







