Improving Temporal Stability and Accuracy for Endoscopic Video Tissue Classification Using Recurrent Neural Networks
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics and Information Governance
2.2. Datasets and Clinical Taxonomy
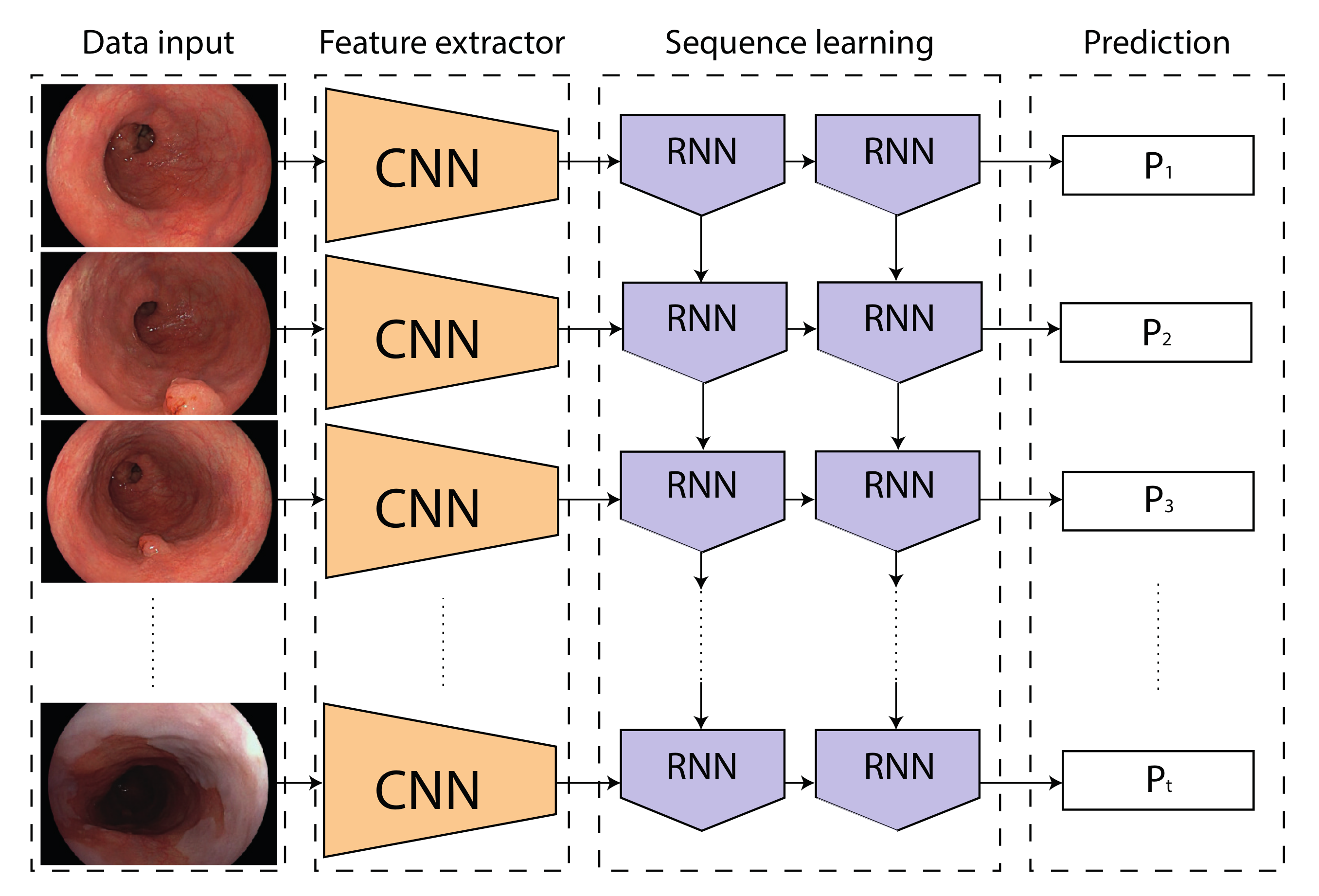
2.3. Network Architectures and Training Protocol
3. Experiments
3.1. Metrics
3.2. Statistical Analysis
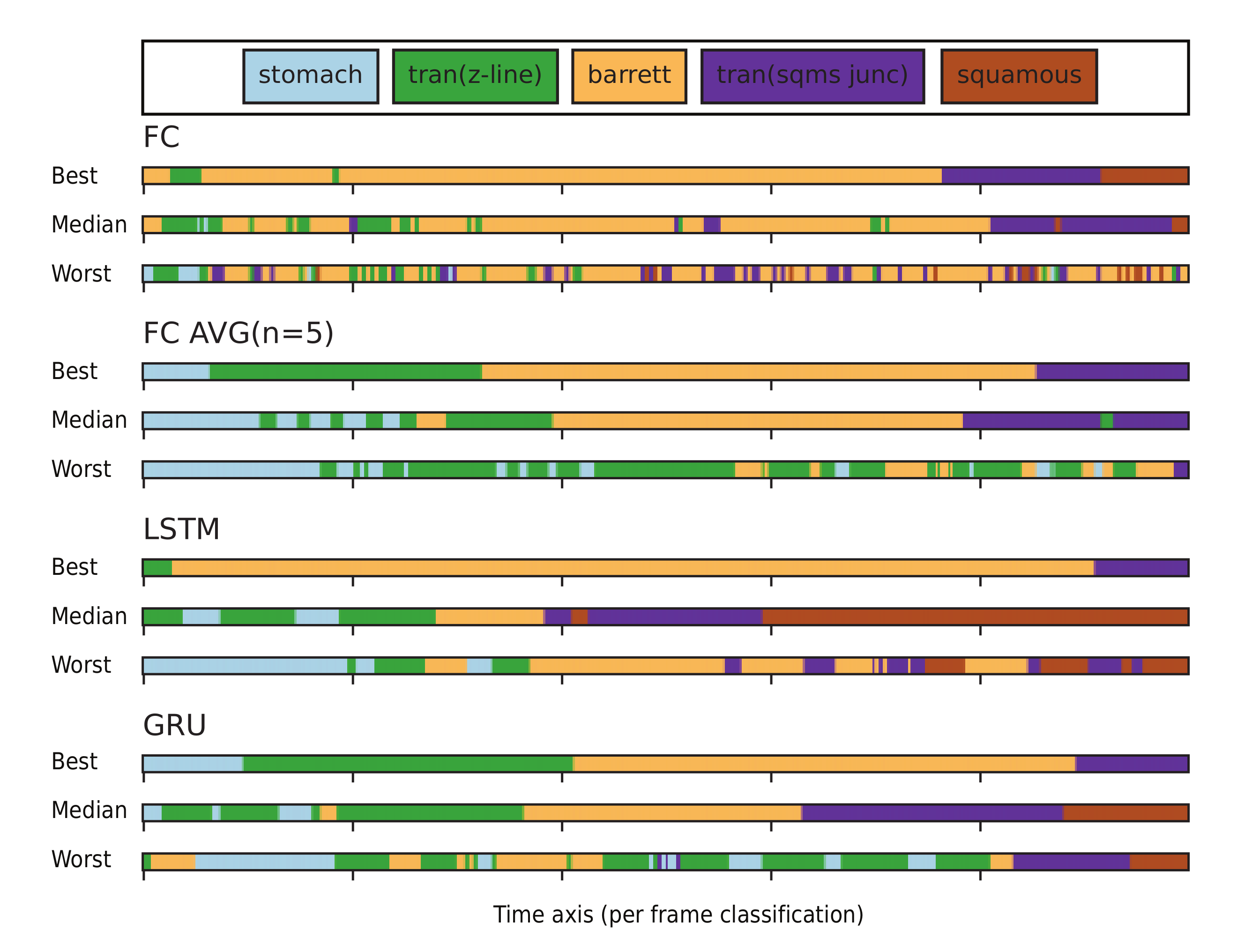
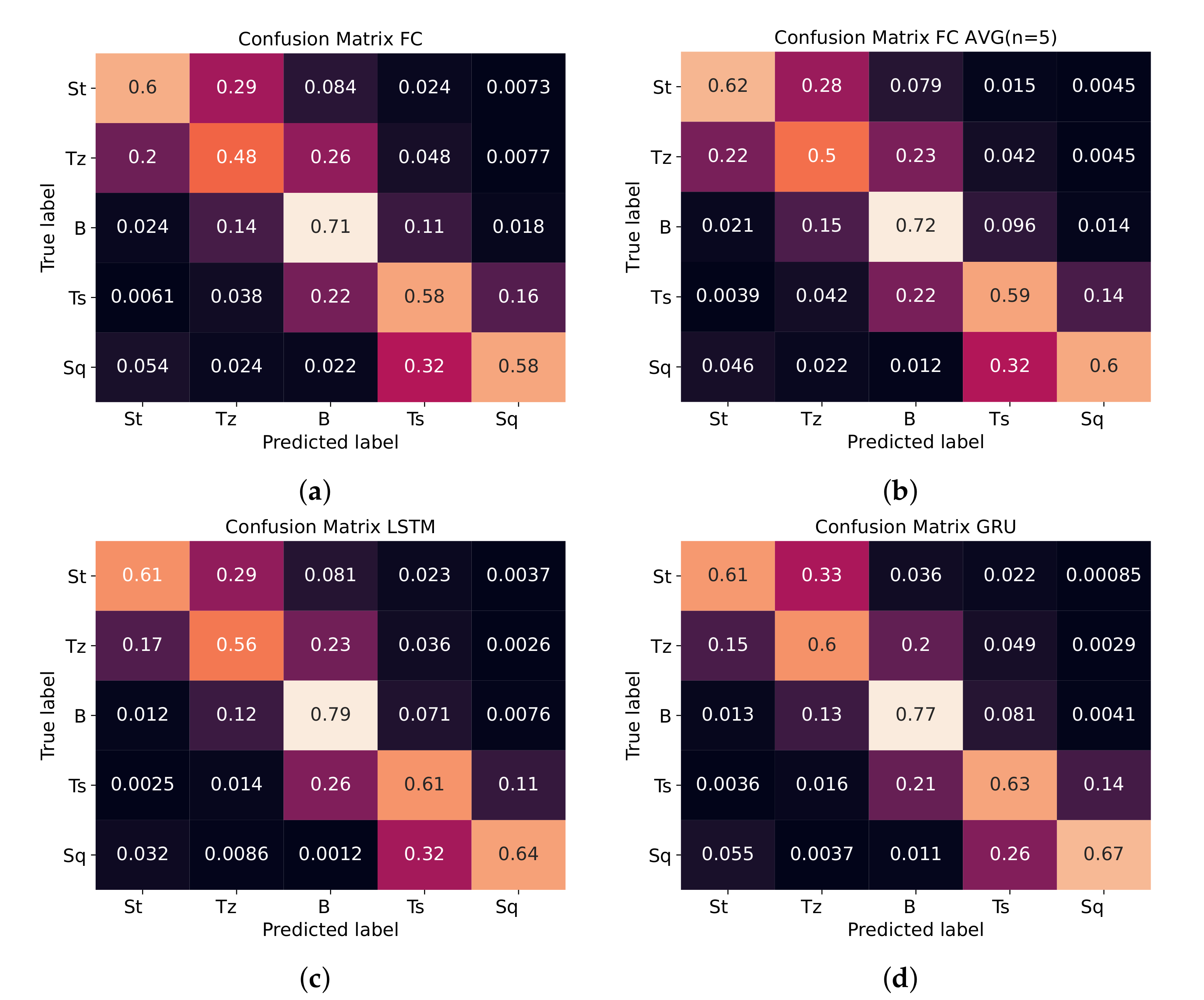
4. Results
5. Discussion and Conclusions
Author Contributions
Funding
Conflicts of Interest
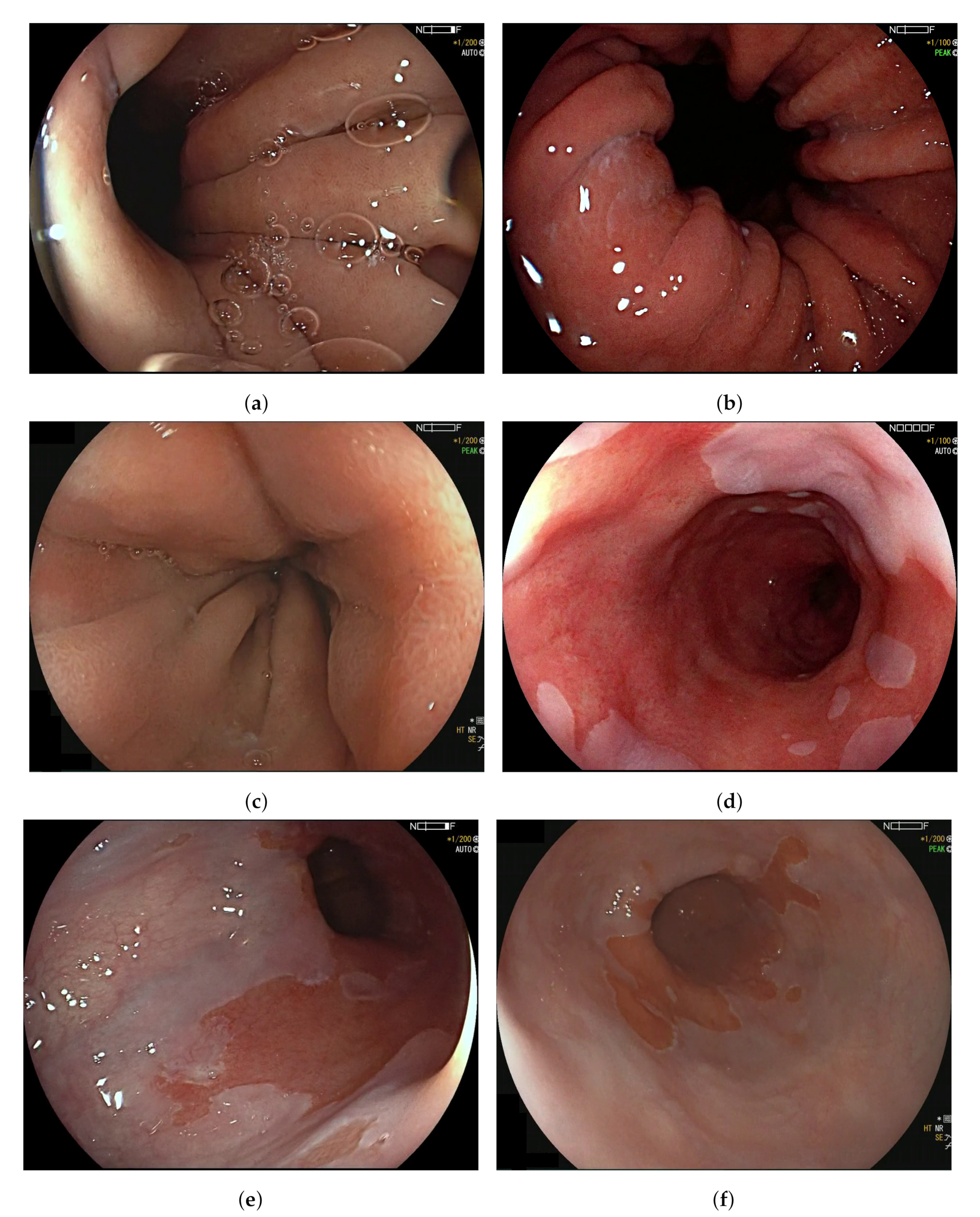
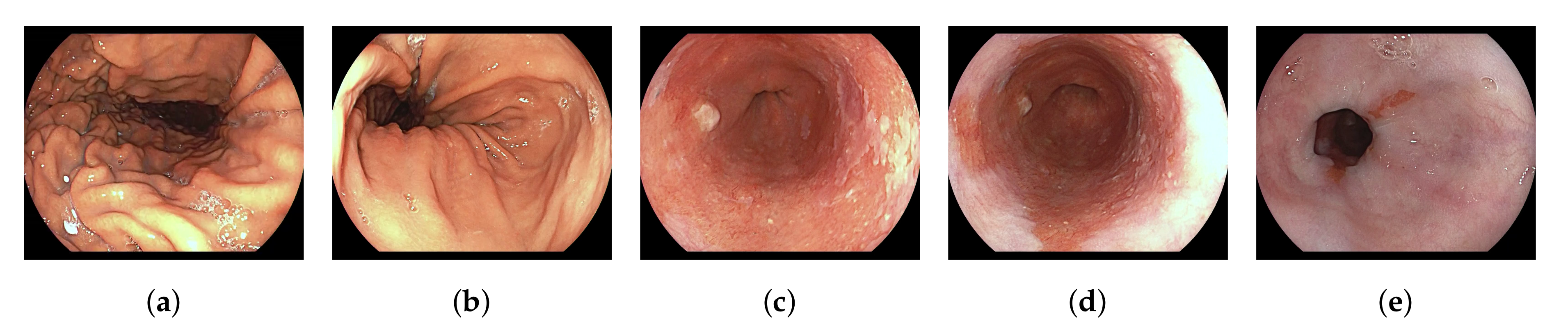
Appendix A. Ambiguous Organ Classes
References
- Groof, J.D.; van der Sommen, F.; van der Putten, J.; Struyvenberg, M.R.; Zinger, S.; Curvers, W.L.; Pech, O.; Meining, A.; Neuhaus, H.; Bisschops, R.; et al. The Argos project: The development of a computer-aided detection system to improve detection of Barrett’s neoplasia on white light endoscopy. United Eur. Gastroenterol. J. 2019, 7, 538–547. [Google Scholar] [CrossRef] [PubMed]
- Guck, J.; Schinkinger, S.; Lincoln, B.; Wottawah, F.; Ebert, S.; Romeyke, M.; Lenz, D.; Erickson, H.M.; Ananthakrishnan, R.; Mitchell, D.; et al. Optical deformability as an inherent cell marker for testing malignant transformation and metastatic competence. Biophys. J. 2005, 88, 3689–3698. [Google Scholar] [CrossRef] [PubMed]
- Van der Putten, J.; de Groof, J.; van der Sommen, F.; Struyvenberg, M.; Zinger, S.; Curvers, W.; Schoon, E.; Bergman, J.; de With, P.H. Informative frame classification of endoscopic videos using convolutional neural networks and hidden Markov models. In Proceedings of the 2019 IEEE International Conference on Image Processing (ICIP), Taipei, Taiwan, 22–25 September 2019; pp. 380–384. [Google Scholar]
- Wang, S.; Cong, Y.; Cao, J.; Yang, Y.; Tang, Y.; Zhao, H.; Yu, H. Scalable gastroscopic video summarization via similar-inhibition dictionary selection. Artif. Intell. Med. 2016, 66, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Giordano, D.; Murabito, F.; Palazzo, S.; Pino, C.; Spampinato, C. An AI-based Framework for Supporting Large Scale Automated Analysis of Video Capsule Endoscopy. In Proceedings of the 2019 IEEE EMBS International Conference on Biomedical & Health Informatics (BHI), Chicago, IL, USA, 19–22 May 2019; pp. 1–4. [Google Scholar]
- Rezvy, S.; Zebin, T.; Braden, B.; Pang, W.; Taylor, S.; Gao, X. Transfer learning for Endoscopy disease detection and segmentation with mask-RCNN benchmark architecture. In Proceedings of the 2020 IEEE 17th International Symposium on Biomedical Imaging, Oxford, UK, 3 April 2020; p. 17. [Google Scholar]
- Ali, H.; Sharif, M.; Yasmin, M.; Rehmani, M.H.; Riaz, F. A survey of feature extraction and fusion of deep learning for detection of abnormalities in video endoscopy of gastrointestinal-tract. Artif. Intell. Rev. 2019, 1–73. [Google Scholar] [CrossRef]
- Du, W.; Rao, N.; Liu, D.; Jiang, H.; Luo, C.; Li, Z.; Gan, T.; Zeng, B. Review on the Applications of Deep Learning in the Analysis of Gastrointestinal Endoscopy Images. IEEE Access 2019, 7, 142053–142069. [Google Scholar] [CrossRef]
- Van der Putten, J.; de Groof, J.; van der Sommen, F.; Struyvenberg, M.; Zinger, S.; Curvers, W.; Schoon, E.; Bergman, J.; de With, P.H.N. First steps into endoscopic video analysis for Barrett’s cancer detection: Challenges and opportunities. In Medical Imaging 2020: Computer-Aided Diagnosis; International Society for Optics and Photonics: San Diego, CA, USA, 2020; Volume 11314, p. 1131431. [Google Scholar]
- Yao, G.; Liu, X.; Lei, T. Action Recognition with 3D ConvNet-GRU Architecture. In Proceedings of the 3rd ACM International Conference on Robotics, Control and Automation, Chengdu, China, 11–13 August 2018; pp. 208–213. [Google Scholar]
- Yue-Hei Ng, J.; Hausknecht, M.; Vijayanarasimhan, S.; Vinyals, O.; Monga, R.; Toderici, G. Beyond short snippets: Deep networks for video classification. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 4694–4702. [Google Scholar]
- Donahue, J.; Anne Hendricks, L.; Guadarrama, S.; Rohrbach, M.; Venugopalan, S.; Saenko, K.; Darrell, T. Long-term recurrent convolutional networks for visual recognition and description. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 2625–2634. [Google Scholar]
- Hashimoto, R.; Requa, J.; Tyler, D.; Ninh, A.; Tran, E.; Mai, D.; Lugo, M.; Chehade, N.E.H.; Chang, K.J.; Karnes, W.E.; et al. Artificial intelligence using convolutional neural networks for real-time detection of early esophageal neoplasia in Barrett’s esophagus (with video). Gastrointest. Endosc. 2020. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Zhou, F.; Bailey, A.; Braden, B.; East, J.; Lu, X.; Rittscher, J. A deep learning framework for quality assessment and restoration in video endoscopy. arXiv 2019, arXiv:1904.07073. [Google Scholar]
- Van der Putten, J.; Struyvenberg, M.; de Groof, J.; Curvers, W.; Schoon, E.; Baldaque-Silva, F.; Bergman, J.; van der Sommen, F.; de With, P.H.N. Endoscopy-Driven Pretraining for Classification of Dysplasia in Barrett’s Esophagus with Endoscopic Narrow-Band Imaging Zoom Videos. Appl. Sci. 2020, 10, 3407. [Google Scholar] [CrossRef]
- Byrne, M.F.; Chapados, N.; Soudan, F.; Oertel, C.; Pérez, M.L.; Kelly, R.; Iqbal, N.; Chandelier, F.; Rex, D.K. Real-time differentiation of adenomatous and hyperplastic diminutive colorectal polyps during analysis of unaltered videos of standard colonoscopy using a deep learning model. Gut 2019, 68, 94–100. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Chen, H.; Dou, Q.; Qin, J.; Heng, P.A. Integrating online and offline three-dimensional deep learning for automated polyp detection in colonoscopy videos. IEEE J. Biomed. Health Inform. 2016, 21, 65–75. [Google Scholar] [CrossRef] [PubMed]
- Harada, S.; Hayashi, H.; Bise, R.; Tanaka, K.; Meng, Q.; Uchida, S. Endoscopic image clustering with temporal ordering information based on dynamic programming. In Proceedings of the 2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, 23–27 July 2019; pp. 3681–3684. [Google Scholar]
- Owais, M.; Arsalan, M.; Choi, J.; Mahmood, T.; Park, K.R. Artificial intelligence-based classification of multiple gastrointestinal diseases using endoscopy videos for clinical diagnosis. J. Clin. Med. 2019, 8, 986. [Google Scholar] [CrossRef] [PubMed]
- Ghatwary, N.; Zolgharni, M.; Janan, F.; Ye, X. Learning spatiotemporal features for esophageal abnormality detection from endoscopic videos. IEEE J. Biomed. Health Inform. 2020. [Google Scholar] [CrossRef]
- Graves, A.; Mohamed, A.R.; Hinton, G. Speech recognition with deep recurrent neural networks. In Proceedings of the 2013 IEEE international Conference on Acoustics, Speech and Signal Processing, Vancouver, BC, Canada, 26–31 May 2013; pp. 6645–6649. [Google Scholar]
- Wilcoxon, F. Individual comparisons by ranking methods. Biom. Bull. 1945, 1, 80–83. [Google Scholar] [CrossRef]
| Predicted Label | True Positive If Label Is: |
|---|---|
| Stomach (St) | St |
| Transition Z-line (Tz) | Tz, B |
| Barrett (B) | Tz, B, Ts |
| Transition squamous (Ts) | B, Ts |
| Squamous (Sq) | Sq |
| Label | N | Mean Accuracy (%) | |||
|---|---|---|---|---|---|
| FC | FC Avg (n = 5) | LSTM | GRU | ||
| Stomach (St) | 2593 | 59.6 | 62.2 | 60.7 | 61.3 |
| Tran. Z-line (Tz) | 2921 | 74.1 | 72.9 | 79.3 | 79.7 |
| Barrett (B) | 9444 | 95.7 | 96.1 | 98.0 | 98.3 |
| Tran. squamous (Ts) | 4215 | 79.5 | 81.3 | 87.0 | 83.9 |
| Squamous (Sq) | 755 | 58.3 | 59.9 | 63.8 | 67.4 |
| Overall | 19,931 | 82.2 | 83.0 | 85.9 | 85.6 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Boers, T.; van der Putten, J.; Struyvenberg, M.; Fockens, K.; Jukema, J.; Schoon, E.; van der Sommen, F.; Bergman, J.; de With, P. Improving Temporal Stability and Accuracy for Endoscopic Video Tissue Classification Using Recurrent Neural Networks. Sensors 2020, 20, 4133. https://doi.org/10.3390/s20154133
Boers T, van der Putten J, Struyvenberg M, Fockens K, Jukema J, Schoon E, van der Sommen F, Bergman J, de With P. Improving Temporal Stability and Accuracy for Endoscopic Video Tissue Classification Using Recurrent Neural Networks. Sensors. 2020; 20(15):4133. https://doi.org/10.3390/s20154133
Chicago/Turabian StyleBoers, Tim, Joost van der Putten, Maarten Struyvenberg, Kiki Fockens, Jelmer Jukema, Erik Schoon, Fons van der Sommen, Jacques Bergman, and Peter de With. 2020. "Improving Temporal Stability and Accuracy for Endoscopic Video Tissue Classification Using Recurrent Neural Networks" Sensors 20, no. 15: 4133. https://doi.org/10.3390/s20154133
APA StyleBoers, T., van der Putten, J., Struyvenberg, M., Fockens, K., Jukema, J., Schoon, E., van der Sommen, F., Bergman, J., & de With, P. (2020). Improving Temporal Stability and Accuracy for Endoscopic Video Tissue Classification Using Recurrent Neural Networks. Sensors, 20(15), 4133. https://doi.org/10.3390/s20154133








