Automated Detection of Presymptomatic Conditions in Spinocerebellar Ataxia Type 2 Using Monte Carlo Dropout and Deep Neural Network Techniques with Electrooculogram Signals
Abstract
1. Introduction
2. Uncertainty Quantification in Deep Learning
- The given task is naturally vague or
- The models are not appropriate to describe the data.
- Aleatoric uncertainty or data uncertainty.
- Epistemic uncertainty or knowledge uncertainty.
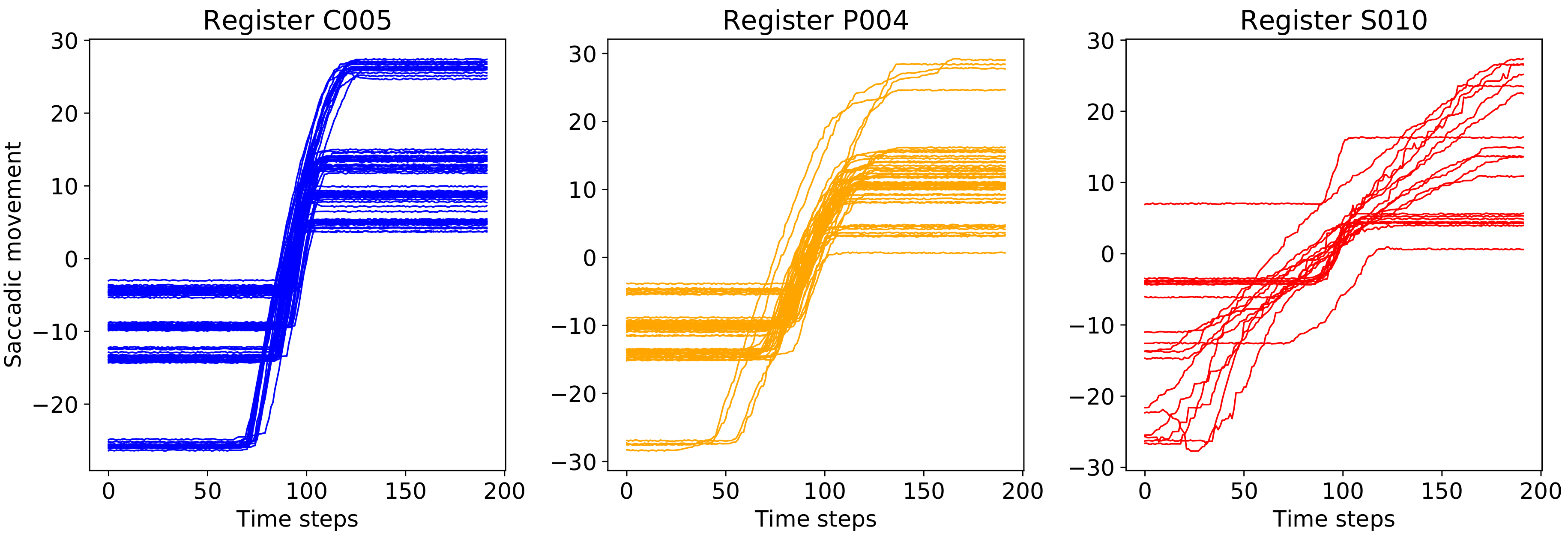
3. Materials
4. Methods
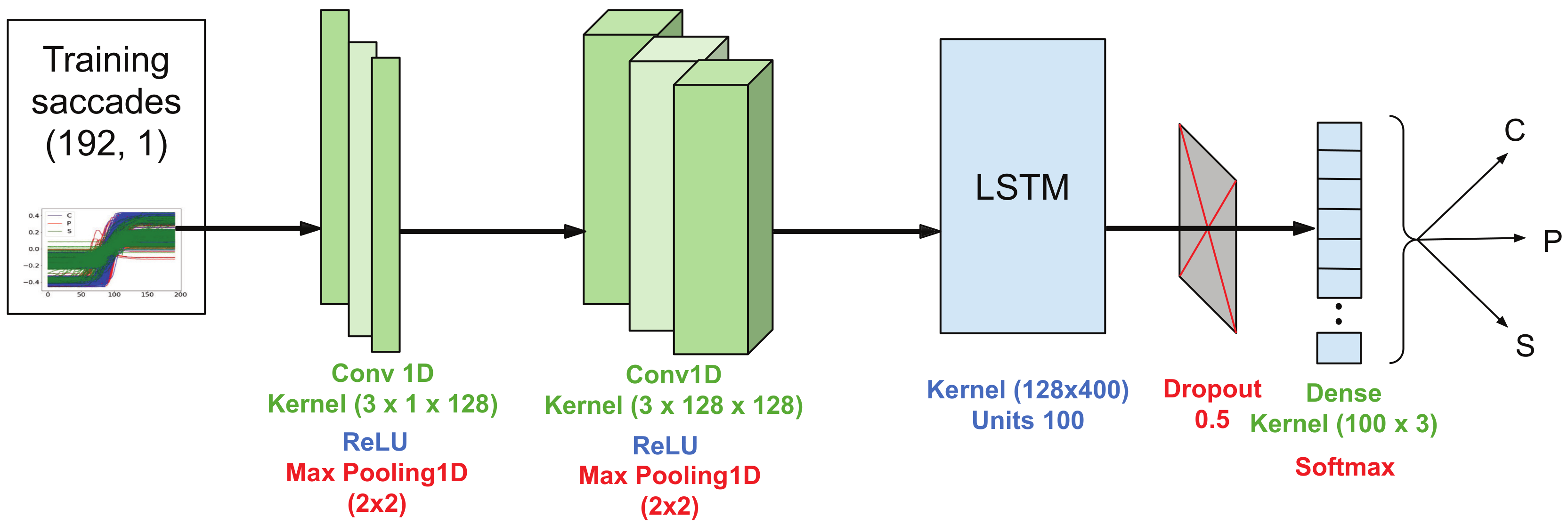
4.1. Deep Modeling with Monte Carlo Dropout for EOG Saccades
| Algorithm 1: Calculation of class probabilities, labeling of individual saccades and computation of mean accuracy over the N runs of the MCD. This procedure is applied to both validation and test registers with constituent saccades |
 |
| Algorithm 2: MCD ensemble (MCDE) calculation of class probabilities and labeling for individual saccades and computation of accuracy. This procedure is applied to both validation and test registers with constituent saccades |
 |
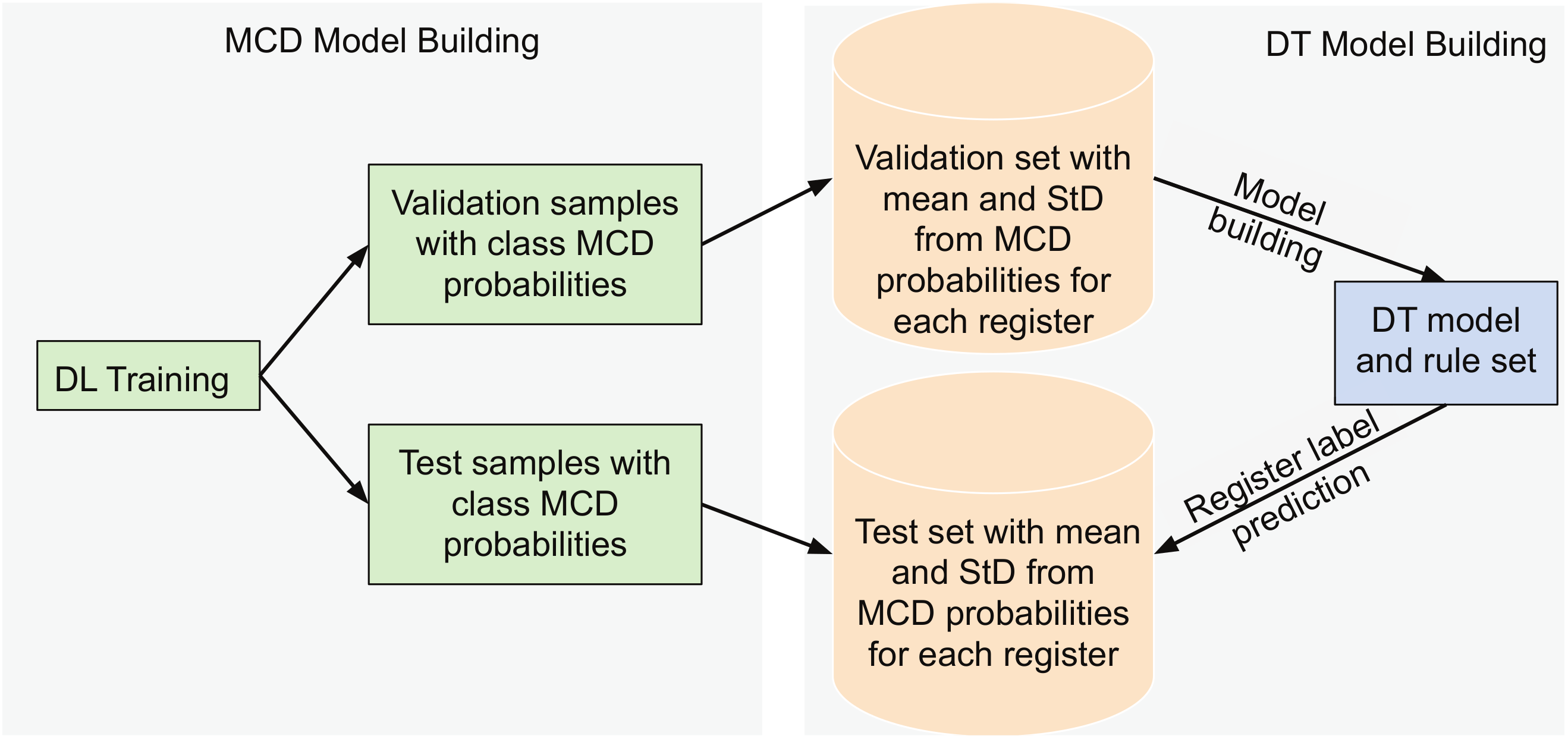
4.2. Feature Extraction and Classification of Registers via Decision Trees
| Algorithm 3: DT model construction on the validation registers and prediction on the test registers |
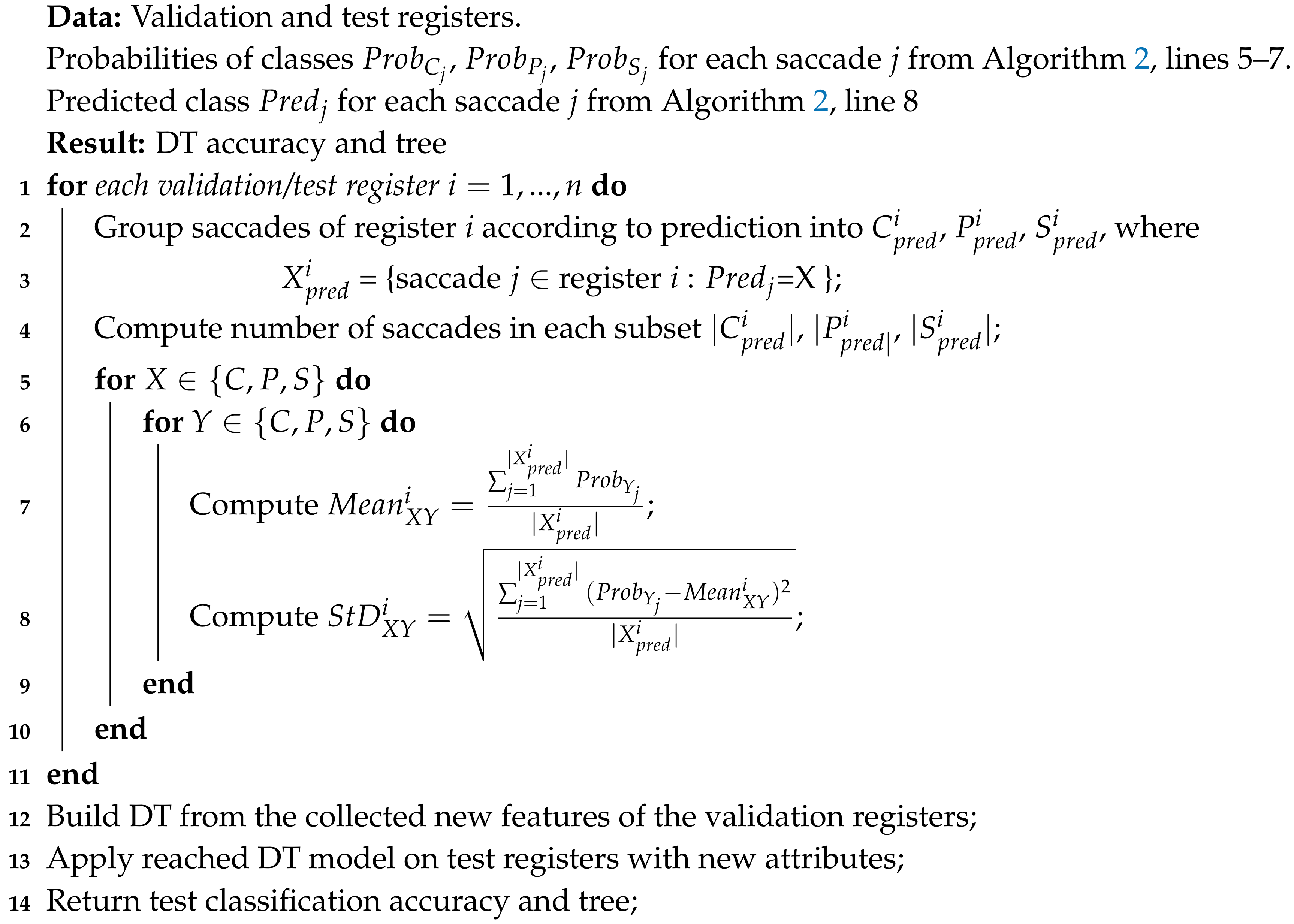 |
5. Experimental Results
5.1. Experimental Setup
5.2. Results and Visualization
5.3. Discussion
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| EOG | Electrooculogram |
| SCA2 | Spinocerebellar ataxia type 2 |
| DL | Deep learning |
| DNN | Deep neural networks |
| UQ | Uncertainty Quantification |
| MCD | Monte Carlo dropout |
| MCDE | Monte Carlo dropout ensemble |
| CNN | Convolutional neural network |
| LSTM | Long short-term memory |
| DT | Decision tree |
References
- Becerra-García, R.A.; García-Bermúdez, R.; Joya, G.; Fernández-Higuera, A.; Velázquez-Rodríguez, C.; Velázquez-Mariño, M.; Cuevas-Beltrán, F.; García-Lagos, F.; Rodráguez-Labrada, R. Data mining process for identification of non-spontaneous saccadic movements in clinical electrooculography. Neurocomputing 2017, 250, 28–36. [Google Scholar] [CrossRef]
- Stoean, C.; Stoean, R.; Becerra-García, R.A.; García-Bermúdez, R.; Atencia, M.; García-Lagos, F.; Velázquez-Pérez, L.; Joya, G. Unsupervised Learning as a Complement to Convolutional Neural Network Classification in the Analysis of Saccadic Eye Movement in Spino-Cerebellar Ataxia Type 2. In Advances in Computational Intelligence; Rojas, I., Joya, G., Catala, A., Eds.; Springer International Publishing: Cham, Switzerland, 2019; pp. 26–37. [Google Scholar]
- Stoean, R.; Stoean, C.; Becerra-García, R.A.; García-Bermúdez, R.; Atencia, M.; García-Lagos, F.; Velázquez-Pérez, L.; Joya, G. A Hybrid Unsupervised—Deep Learning Tandem for Electrooculography Time Series Analysis. PLoS ONE 2020, in press. [Google Scholar]
- Gal, Y.; Ghahramani, Z. Dropout as a Bayesian Approximation: Representing Model Uncertainty in Deep Learning. In Proceedings of the 33rd International Conference on International Conference on Machine Learning (ICML’16), New York, NY, USA, 20–22 June 2016; Volume 48, pp. 1050–1059. [Google Scholar]
- Hastie, T.; Tibshirani, R.; Friedman, J.H. The Elements of Statistical Learning: Data Mining, Inference, and Prediction, 2nd ed.; Springer Series in Statistics; Springer: New York, NY, USA, 2009; p. 745. [Google Scholar]
- Abdar, M.; Yen, N.Y.; Hung, J.C.S. Improving the diagnosis of liver disease using multilayer perceptron neural network and boosted decision trees. J. Med Biol. Eng. 2018, 38, 953–965. [Google Scholar] [CrossRef]
- Abdar, M.; Makarenkov, V. CWV-BANN-SVM ensemble learning classifier for an accurate diagnosis of breast cancer. Measurement 2019, 146, 557–570. [Google Scholar] [CrossRef]
- Wang, X.; Zhao, Y.; Pourpanah, F. Recent Advances in Deep Learning. Int. J. Mach. Learn. Cyber. 2020, 11, 747–750. [Google Scholar] [CrossRef]
- Basiri, M.E.; Abdar, M.; Cifci, M.A.; Nemati, S.; Acharya, U.R. A novel method for sentiment classification of drug reviews using fusion of deep and machine learning techniques. Knowl. Based Syst. 2020, 105949. [Google Scholar] [CrossRef]
- Sherkatghanad, Z.; Akhondzadeh, M.; Salari, S.; Zomorodi-Moghadam, M.; Abdar, M.; Acharya, U.R.; Khosrowabadi, R.; Salari, V. Automated detection of autism spectrum disorder using a convolutional neural network. Front. Neurosci. 2019, 13, 1325. [Google Scholar] [CrossRef] [PubMed]
- Shen, D.; Wu, G.; Suk, H.I. Deep learning in medical image analysis. Annu. Rev. Biomed. Eng. 2017, 19, 221–248. [Google Scholar] [CrossRef]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; Van Der Laak, J.A.; Van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- Stoean, R. Analysis on the potential of an EA-surrogate modelling tandem for deep learning parametrization: An example for cancer classification from medical images. Neural Comput. Appl. 2020, 32, 313–322. [Google Scholar] [CrossRef]
- Mittal, S.; Stoean, C.; Kajdacsy-Balla, A.; Bhargava, R. Digital Assessment of Stained Breast Tissue Images for Comprehensive Tumor and Microenvironment Analysis. Front. Bioeng. Biotechnol. 2019, 7, 246. [Google Scholar] [CrossRef] [PubMed]
- García-Garaluz, E.; Atencia, M.; Joya, G.; García-Lagos, F.; Sandoval, F. Hopfield networks for identification of delay differential equations with an application to dengue fever epidemics in Cuba. Neurocomputing 2011, 74, 2691–2697. [Google Scholar] [CrossRef]
- Zomorodi-Moghadam, M.; Abdar, M.; Davarzani, Z.; Zhou, X.; Pławiak, P.; Acharya, U.R. Hybrid particle swarm optimization for rule discovery in the diagnosis of coronary artery disease. Expert Syst. 2019, e12485. [Google Scholar] [CrossRef]
- Pourpanah, F.; Zhang, B.; Ma, R.; Hao, Q. Non-intrusive human motion recognition using distributed sparse sensors and the genetic algorithm based neural network. In Proceedings of the 2018 IEEE SENSORS, New Delhi, India, 28–31 October 2018; pp. 1–4. [Google Scholar]
- Pourpanah, F.; Shi, Y.; Lim, C.P.; Hao, Q.; Tan, C.J. Feature selection based on brain storm optimization for data classification. Appl. Soft Comput. 2019, 80, 761–775. [Google Scholar] [CrossRef]
- Plawiak, P.; Tadeusiewicz, R. Approximation of phenol concentration using novel hybrid computational intelligence methods. Int. J. Appl. Math. Comput. Sci. 2014, 24, 165–181. [Google Scholar] [CrossRef]
- Pławiak, P. An estimation of the state of consumption of a positive displacement pump based on dynamic pressure or vibrations using neural networks. Neurocomputing 2014, 144, 471–483. [Google Scholar] [CrossRef]
- Alam, S.; Kwon, K.C.; Alam, A.; Abbass, M.Y.; Imtiaz, S.M.; Kim, N. Trajectory-Based Air-Writing Recognition Using Deep Neural Network and Depth Sensor. Sensors 2020, 20, 376. [Google Scholar]
- Rengasamy, D.; Jafari, M.; Rothwell, B.; Chen, X.; Figueredo, G.P. Deep Learning with Dynamically Weighted Loss Function for Sensor-Based Prognostics and Health Management. Sensors 2020, 20, 723. [Google Scholar] [CrossRef]
- Yildirim, O.; Baloglu, U.B.; Tan, R.S.; Ciaccio, E.J.; Acharya, U.R. A new approach for arrhythmia classification using deep coded features and LSTM networks. Comput. Methods Programs Biomed. 2019, 176, 121–133. [Google Scholar] [CrossRef]
- Plawiak, P.; Acharya, U.R. Novel deep genetic ensemble of classifiers for arrhythmia detection using ECG signals. Neural Comput. Appl. 2019, 1–25. [Google Scholar]
- Cohen, J.P.; Luck, M.; Honari, S. Distribution matching losses can hallucinate features in medical image translation. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Cham, Switzerland, 2018; pp. 529–536. [Google Scholar]
- Tanno, R.; Worrall, D.; Kaden, E.; Ghosh, A.; Grussu, F.; Bizzi, A.; Sotiropoulos, S.N.; Criminisi, A.; Alexander, D.C. Uncertainty Quantification in Deep Learning for Safer Neuroimage Enhancement. arXiv 2019, arXiv:1907.13418. [Google Scholar]
- Malinin, A. Uncertainty Estimation in Deep Learning with Application to Spoken Language Assessment. Ph.D. Thesis, University of Cambridge, Cambridge, UK, 2019. [Google Scholar]
- Labach, A.; Salehinejad, H.; Valaee, S. Survey of dropout methods for deep neural networks. arXiv 2019, arXiv:1904.13310. [Google Scholar]
- Nguyen, C.; Do, T.T.; Carneiro, G. Uncertainty in model-agnostic meta-learning using variational inference. In Proceedings of the IEEE Winter Conference on Applications of Computer Vision, Snowmass Village, CO, USA, 1–5 March 2020; pp. 3090–3100. [Google Scholar]
- Wen, Y.; Tran, D.; Ba, J. BatchEnsemble: An Alternative Approach to Efficient Ensemble and Lifelong Learning. arXiv 2020, arXiv:2002.06715. [Google Scholar]
- Jungo, A.; Meier, R.; Ermis, E.; Herrmann, E.; Reyes, M. Uncertainty-driven sanity check: Application to postoperative brain tumor cavity segmentation. arXiv 2018, arXiv:1806.03106. [Google Scholar]
- Jungo, A.; McKinley, R.; Meier, R.; Knecht, U.; Vera, L.; Pérez-Beteta, J.; Molina-García, D.; Pérez-García, V.M.; Wiest, R.; Reyes, M. Towards uncertainty-assisted brain tumor segmentation and survival prediction. In International MICCAI Brainlesion Workshop; Springer: Cham, Switzerland, 2017; pp. 474–485. [Google Scholar]
- Lubrano di Scandalea, M.; Perone, C.S.; Boudreau, M.; Cohen-Adad, J. Deep active learning for axon-myelin segmentation on histology data. arXiv 2019, arXiv:1907.05143. [Google Scholar]
- Guo, F.; Ng, M.; Goubran, M.; Petersen, S.E.; Piechnik, S.K.; Neubauer, S.; Wright, G. Improving Cardiac MRI Convolutional Neural Network Segmentation on Small Training Datasets and Dataset Shift: A Continuous Kernel Cut Approach. Med. Image Anal. 2020, 6, 101636. [Google Scholar] [CrossRef] [PubMed]
- Côté-Allard, U.; Fall, C.L.; Drouin, A.; Campeau-Lecours, A.; Gosselin, C.; Glette, K.; Laviolette, F.; Gosselin, B. Deep learning for electromyographic hand gesture signal classification using transfer learning. IEEE Trans. Neural Syst. Rehabil. Eng. 2019, 27, 760–771. [Google Scholar] [CrossRef]
- Cheng, X.; Henry, C.; Andriulli, F.P.; Person, C.; Wiart, J. A Surrogate Model Based on Artificial Neural Network for RF Radiation Modelling with High-Dimensional Data. Int. J. Environ. Res. Public Health 2020, 17, 2586. [Google Scholar] [CrossRef]
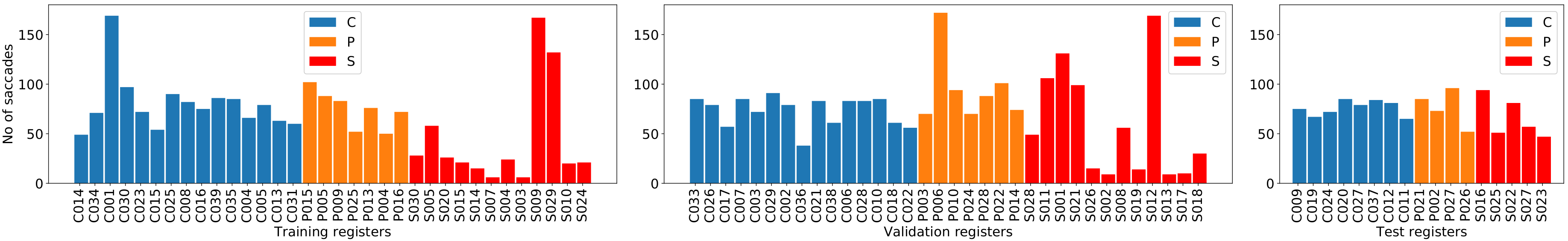
| Class | Average | StD | Min | Max |
|---|---|---|---|---|
| Control | 78.3 | 21.6 | 38 | 169 |
| Presymptomatic | 76.7 | 25 | 28 | 172 |
| Sick | 54 | 49 | 6 | 172 |
| Approach | Average (%) | Standard Deviation (%) | Minimum (%) | Maximum (%) |
|---|---|---|---|---|
| Saccade classification | ||||
| DL-MCD validation | 68.07 | 0.74 | 66.68 | 69.29 |
| DL-MCD Ensemble validation | 68.77 | 0.51 | 67.78 | 69.48 |
| CNN-LSTM validation | 70.38 | 0.28 | 69.7 | 70.7 |
| SVM validation | 60.06 | 0 | 60.06 | 60.06 |
| DL-MCD test | 70.28 | 1.15 | 68.57 | 72.19 |
| DL-MCD Ensemble test | 70.84 | 1.29 | 68.81 | 72.75 |
| CNN-LSTM test | 74.12 | 0.17 | 73.9 | 74.4 |
| SVM test | 63.18 | 0 | 63.18 | 63.18 |
| Register classification on the test set | ||||
| DL-MCDE-DT Accuracy | 81.18 | 7.16 | 72.12 | 94.12 |
| CNN-LSTM Accuracy | 76.47 | 0 | 76.47 | 76.47 |
| SVM Accuracy | 64.71 | 0 | 64.71 | 64.71 |
| DL-MCDE-DT Precision | 81.9 | 9.05 | 68.63 | 94.77 |
| DL-MCDE-DT Recall | 81.18 | 9.04 | 70.59 | 94.12 |
| DL-MCDE-DT F1-score | 80.93 | 9.16 | 69.36 | 93.87 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stoean, C.; Stoean, R.; Atencia, M.; Abdar, M.; Velázquez-Pérez, L.; Khosravi, A.; Nahavandi, S.; Acharya, U.R.; Joya, G. Automated Detection of Presymptomatic Conditions in Spinocerebellar Ataxia Type 2 Using Monte Carlo Dropout and Deep Neural Network Techniques with Electrooculogram Signals. Sensors 2020, 20, 3032. https://doi.org/10.3390/s20113032
Stoean C, Stoean R, Atencia M, Abdar M, Velázquez-Pérez L, Khosravi A, Nahavandi S, Acharya UR, Joya G. Automated Detection of Presymptomatic Conditions in Spinocerebellar Ataxia Type 2 Using Monte Carlo Dropout and Deep Neural Network Techniques with Electrooculogram Signals. Sensors. 2020; 20(11):3032. https://doi.org/10.3390/s20113032
Chicago/Turabian StyleStoean, Catalin, Ruxandra Stoean, Miguel Atencia, Moloud Abdar, Luis Velázquez-Pérez, Abbas Khosravi, Saeid Nahavandi, U. Rajendra Acharya, and Gonzalo Joya. 2020. "Automated Detection of Presymptomatic Conditions in Spinocerebellar Ataxia Type 2 Using Monte Carlo Dropout and Deep Neural Network Techniques with Electrooculogram Signals" Sensors 20, no. 11: 3032. https://doi.org/10.3390/s20113032
APA StyleStoean, C., Stoean, R., Atencia, M., Abdar, M., Velázquez-Pérez, L., Khosravi, A., Nahavandi, S., Acharya, U. R., & Joya, G. (2020). Automated Detection of Presymptomatic Conditions in Spinocerebellar Ataxia Type 2 Using Monte Carlo Dropout and Deep Neural Network Techniques with Electrooculogram Signals. Sensors, 20(11), 3032. https://doi.org/10.3390/s20113032








