Colony Fingerprint-Based Discrimination of Staphylococcus species with Machine Learning Approaches
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains
2.2. Lens-Less Imaging System
2.3. Imaging Processing
2.4. Discrimination Analysis
3. Results
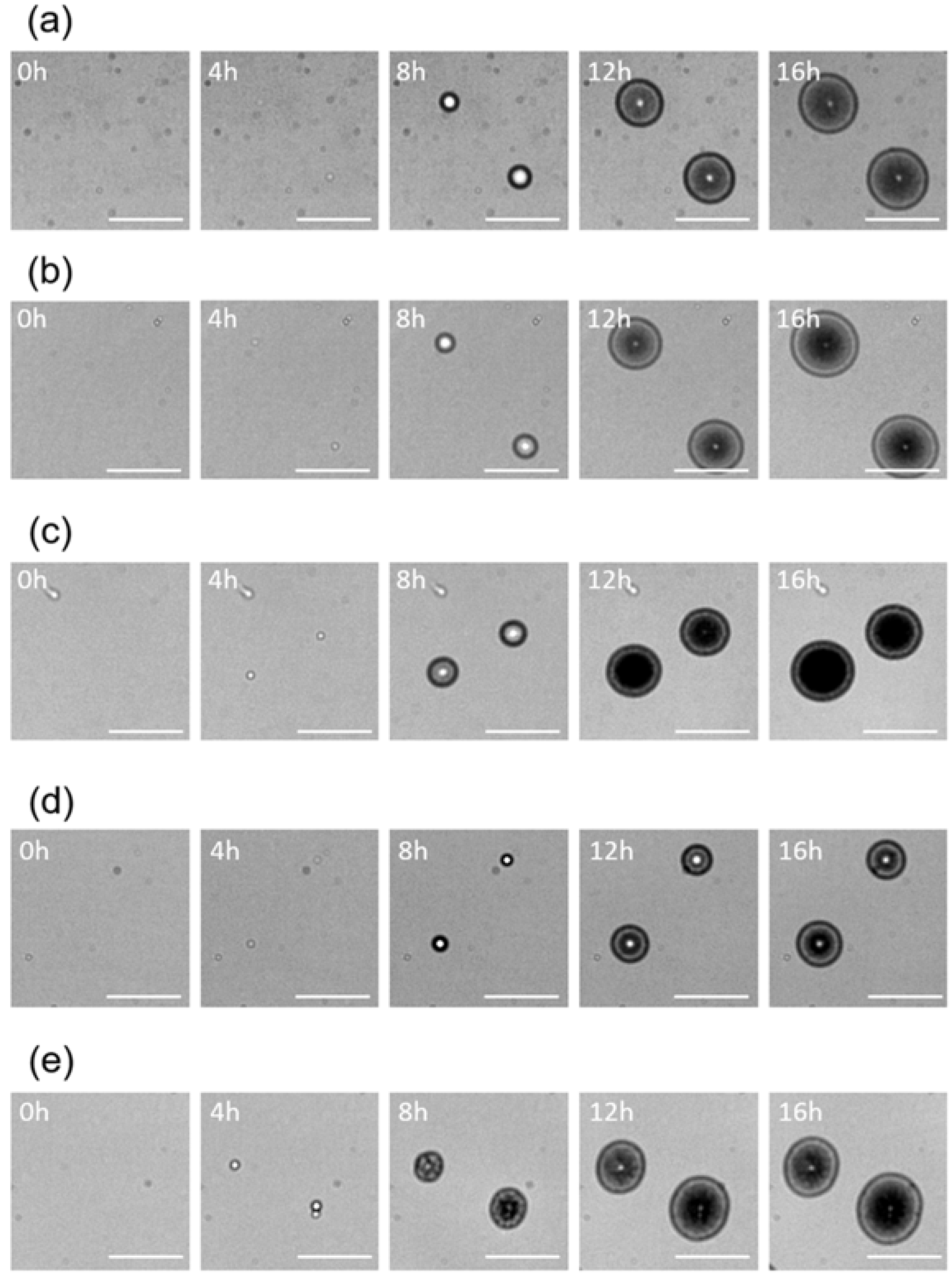
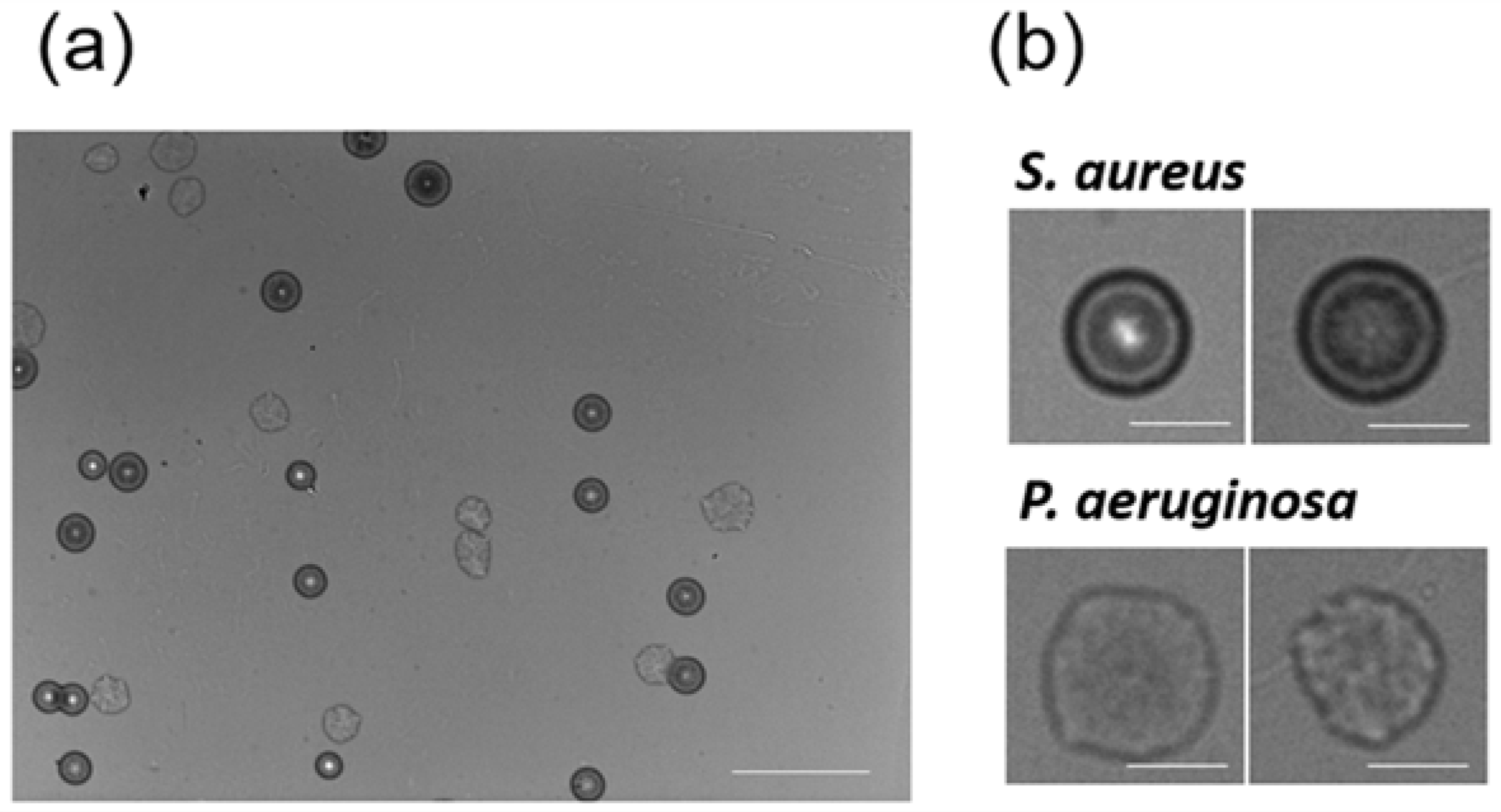
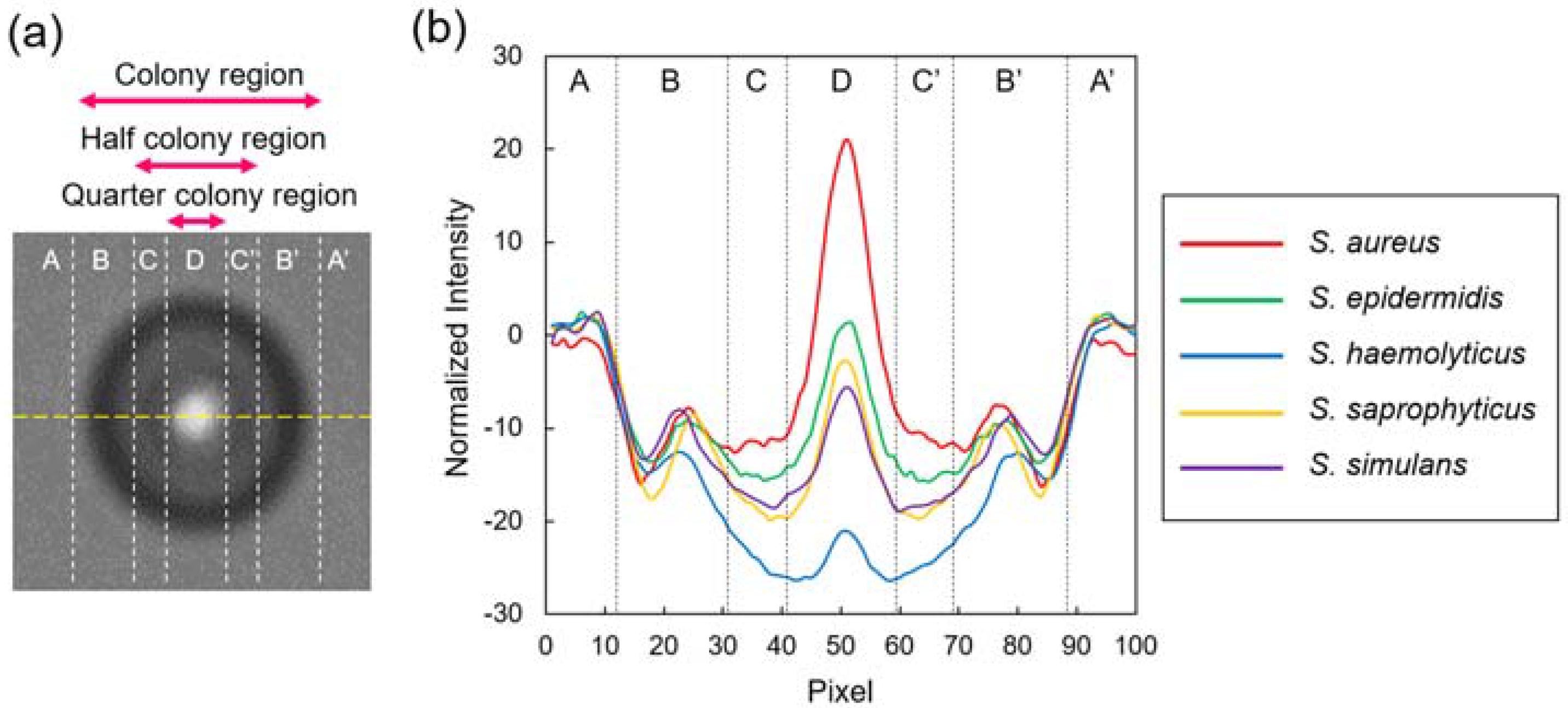
3.1. Colony Fingerprints of Staphylococcus spp.
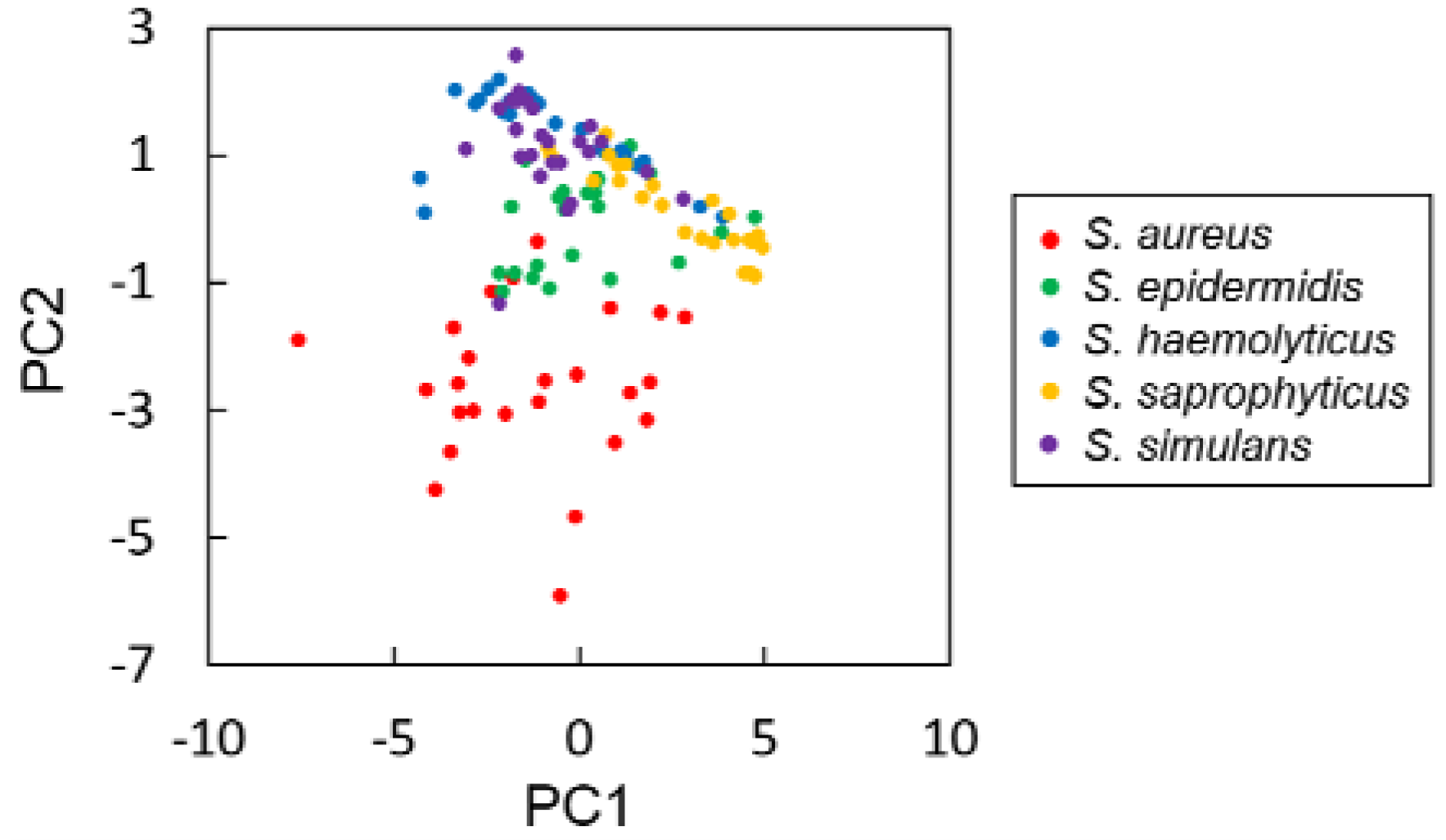
3.2. Discriminative Parameters for Colony Fingerprinting of Staphylococcus spp.
3.3. Comparison of Machine Learning Approaches for Discrimination
3.4. Colony Fingerprints of S. aureus in the Presence of Another Bacterium
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Schleifer, K.-H.; Bell, J.A. Staphylococcaceae; Springer: New York, NY, USA, 2009; Volume 3, p. 392. [Google Scholar]
- Hennekinne, J.A.; De Buyser, M.L.; Dragacci, S. Staphylococcus aureus and its food poisoning toxins: Characterization and outbreak investigation. FEMS Microbiol. Rev. 2012, 36, 815–836. [Google Scholar] [CrossRef] [PubMed]
- Asperger, H.; Zangerl, P. Staphylococcus aureus; Academic Press: San Diego, CA, USA, 2003. [Google Scholar]
- Valík, L.; Sirotná, Z.; Liptáková, D. Growth characterisation of Staphylococcus aureus in milk: A quantitative approach. Czech J. Food Sci. 2009, 27, 443–453. [Google Scholar]
- Medveďová, A.; Valík, Ľ. Staphylococcus aureus: Characterisation and quantitative growth description in milk and artisanal raw milk cheese production. In Structure and Function of Food Engineering; InTech: London, UK, 2012. [Google Scholar]
- Chapman, G.H. The significance of sodium chloride in studies of Staphylococci. J. Bacteriol. 1945, 50, 201–203. [Google Scholar] [PubMed]
- Sperber, W.H.; Tatini, S.R. Interpretation of the tube coagulase test for identification of Staphylococcus aureus. Appl. Microbiol. 1975, 29, 502–505. [Google Scholar] [PubMed]
- Welker, M.; Moore, E.R. Applications of whole-cell matrix-assisted laser-desorption/ionization time-of-flight mass spectrometry in systematic microbiology. Syst. Appl. Microbiol. 2011, 34, 2–11. [Google Scholar] [CrossRef] [PubMed]
- Sandrin, T.R.; Goldstein, J.E.; Schumaker, S. MALDI TOF MS profiling of bacteria at the strain level: A review. Mass Spectrom. Rev. 2013, 32, 188–217. [Google Scholar] [CrossRef] [PubMed]
- Stöckel, S.; Kirchhoff, J.; Neugebauer, U.; Rösch, P.; Popp, J. The application of Raman spectroscopy for the detection and identification of microorganisms. J. Raman Spectrosc. 2016, 47, 89–109. [Google Scholar] [CrossRef]
- Bae, E.; Bai, N.; Aroonnual, A.; Bhunia, A.K.; Hirleman, E.D. Label-free identification of bacterial microcolonies via elastic scattering. Biotechnol. Bioeng. 2011, 108, 637–644. [Google Scholar] [CrossRef] [PubMed]
- Banada, P.P.; Guo, S.; Bayraktar, B.; Bae, E.; Rajwa, B.; Robinson, J.P.; Hirleman, E.D.; Bhunia, A.K. Optical forward-scattering for detection of Listeria monocytogenes and other Listeria species. Biosens. Bioelectron. 2007, 22, 1664–1671. [Google Scholar] [CrossRef] [PubMed]
- Banada, P.P.; Huff, K.; Bae, E.; Rajwa, B.; Aroonnual, A.; Bayraktar, B.; Adil, A.; Robinson, J.P.; Hirleman, E.D.; Bhunia, A.K. Label-free detection of multiple bacterial pathogens using light-scattering sensor. Biosens. Bioelectron. 2009, 24, 1685–1692. [Google Scholar] [CrossRef] [PubMed]
- Huff, K.; Aroonnual, A.; Littlejohn, A.E.; Rajwa, B.; Bae, E.; Banada, P.P.; Patsekin, V.; Hirleman, E.D.; Robinson, J.P.; Richards, G.P.; et al. Light-scattering sensor for real-time identification of Vibrio parahaemolyticus, Vibrio vulnificus and Vibrio cholerae colonies on solid agar plate. Microb. Biotechnol. 2012, 5, 607–620. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Kim, H.; Singh, A.K.; Aroonnual, A.; Bae, E.; Rajwa, B.; Fratamico, P.M.; Bhunia, A.K. Light scattering sensor for direct identification of colonies of Escherichia coli serogroups O26, O45, O103, O111, O121, O145 and O157. PLoS ONE 2014, 9, e105272. [Google Scholar] [CrossRef] [PubMed]
- Maeda, Y.; Dobashi, H.; Sugiyama, Y.; Saeki, T.; Lim, T.K.; Harada, M.; Matsunaga, T.; Yoshino, T.; Tanaka, T. Colony fingerprint for discrimination of microbial species based on lensless imaging of microcolonies. PLoS ONE 2017, 12, e0174723. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T.; Saeki, T.; Sunaga, Y.; Matsunaga, T. High-content analysis of single cells directly assembled on CMOS sensor based on color imaging. Biosens. Bioelectron. 2010, 26, 1460–1465. [Google Scholar] [CrossRef] [PubMed]
- Ozcan, A.; Demirci, U. Ultra wide-field lens-free monitoring of cells on-chip. Lab Chip 2008, 8, 98–106. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.H.; Lee, J.E. Real-time bacterial microcolony counting using on-chip microscopy. Sci. Rep. 2016, 6, 21473. [Google Scholar] [CrossRef] [PubMed]
- Saeki, T.; Hosokawa, M.; Lim, T.K.; Harada, M.; Matsunaga, T.; Tanaka, T. Digital cell counting device integrated with a single-cell array. PLoS ONE 2014, 9, e89011. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T.; Sunaga, Y.; Hatakeyama, K.; Matsunaga, T. Single-cell detection using a thin film transistor photosensor with micro-partitions. Lab Chip 2010, 10, 3348–3354. [Google Scholar] [CrossRef] [PubMed]
- Yoshino, T.; Takai, K.; Negishi, R.; Saeki, T.; Kanbara, H.; Kikuhara, Y.; Matsunaga, T.; Tanaka, T. Rapid imaging and detection of circulating tumor cells using a wide-field fluorescence imaging system. Anal. Chim. Acta 2017, 969, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Saeki, T.; Sugamura, Y.; Hosokawa, M.; Yoshino, T.; Lim, T.K.; Harada, M.; Matsunaga, T.; Tanaka, T. Simple and rapid CD4 testing based on large-field imaging system composed of microcavity array and two-dimensional photosensor. Biosens. Bioelectron. 2015, 67, 350–355. [Google Scholar] [CrossRef] [PubMed]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Otsu, N. A threshold selection method from gray-level histograms. Automatica 1975, 11, 23–27. [Google Scholar] [CrossRef]
- R. Development Core Team. R: A Language and Environment for Statistical Computing. 2013. Available online: http://softlibre.unizar.es/manuales/aplicaciones/r/fullrefman.pdf (accessed on 19 August 2018).
- Bishara, W.; Sikora, U.; Mudanyali, O.; Su, T.W.; Yaglidere, O.; Luckhart, S.; Ozcan, A. Holographic pixel super-resolution in portable lensless on-chip microscopy using a fiber-optic array. Lab Chip 2011, 11, 1276–1279. [Google Scholar] [CrossRef] [PubMed]
- Vercruysse, D.; Dusa, A.; Stahl, R.; Vanmeerbeeck, G.; de Wijs, K.; Liu, C.; Prodanov, D.; Peumans, P.; Lagae, L. Three-part differential of unlabeled leukocytes with a compact lens-free imaging flow cytometer. Lab Chip 2015, 15, 1123–1132. [Google Scholar] [CrossRef] [PubMed]
- Suchwalko, A.; Buzalewicz, I.; Podbielska, H. Bacteria identification in an optical system with optimized diffraction pattern registration condition supported by enhanced statistical analysis. Opt. Express 2014, 22, 26312–26327. [Google Scholar] [CrossRef] [PubMed]
- Buzalewicz, I.; Lizewski, K.; Kujawinska, M.; Podbielska, H. Degeneration of Fraunhofer diffraction on bacterial colonies due to their light focusing properties examined in the digital holographic microscope system. Opt. Express 2013, 21, 26493–26505. [Google Scholar] [CrossRef] [PubMed]
- Su, P.T.; Liao, C.T.; Roan, J.R.; Wang, S.H.; Chiou, A.; Syu, W.J. Bacterial colony from two-dimensional division to three-dimensional development. PLoS ONE 2012, 7, e48098. [Google Scholar] [CrossRef] [PubMed]
- Rosenstein, R.; Götz, F. What distinguishes highly pathogenic staphylococci from medium-and non-pathogenic? In Between Pathogenicity and Commensalism; Springer: Berlin, Germany, 2012; pp. 33–89. [Google Scholar]
- Rodriguez, S.B.; Thornton, M.A.; Thornton, R.J. Discrimination of wine lactic acid bacteria by Raman spectroscopy. J. Ind. Microbiol. Biotechnol. 2017, 44, 1167–1175. [Google Scholar] [CrossRef] [PubMed]
- De Bruyne, K.; Slabbinck, B.; Waegeman, W.; Vauterin, P.; De Baets, B.; Vandamme, P. Bacterial species identification from MALDI-TOF mass spectra through data analysis and machine learning. Syst. Appl. Microbiol. 2011, 34, 20–29. [Google Scholar] [CrossRef] [PubMed]
- Hettick, J.M.; Kashon, M.L.; Slaven, J.E.; Ma, Y.; Simpson, J.P.; Siegel, P.D.; Mazurek, G.N.; Weissman, D.N. Discrimination of intact mycobacteria at the strain level: A combined MALDI-TOF MS and biostatistical analysis. Proteomics 2006, 6, 6416–6425. [Google Scholar] [CrossRef] [PubMed]


| Classifier a | Parameters b | Accuracy c | Species | Sensitivity d | Specificity e | PPV f |
|---|---|---|---|---|---|---|
| LDA | μmax, G, D, H, Ed | 74.4% | S. aureus | 80.0% | 99.0% | 95.2% |
| (5 parameters) | S. epidermidis | 72.0% | 89.0% | 62.1% | ||
| S. haemolyticus | 64.0% | 99.0% | 94.1% | |||
| S. saprophyticus | 80.0% | 95.0% | 80.0% | |||
| S. simulans | 76.0% | 86.0% | 57.6% | |||
| LDA | μmax, G, I, I1/2, I1/4, D, Dc, H, En, Ed, W, R, Z, S | 79.2% | S. aureus | 84.0% | 99.0% | 95.5% |
| S. epidermidis | 76.0% | 86.0% | 57.6% | |||
| (14 parameters) | S. haemolyticus | 76.0% | 99.0% | 95.0% | ||
| S. saprophyticus | 88.0% | 97.0% | 88.0% | |||
| S. simulans | 72.0% | 93.0% | 72.0% | |||
| k-NN | μmax, G, I, I1/2, I1/4, D, Dc, H, En, Ed, W, R, Z, S | 80.8% | S. aureus | 88.0% | 100.0% | 100.0% |
| S. epidermidis | 84.0% | 86.0% | 60.0% | |||
| (14 parameters) | S. haemolyticus | 76.0% | 97.0% | 86.4% | ||
| S. saprophyticus | 88.0% | 96.0% | 84.6% | |||
| S. simulans | 68.0% | 97.0% | 85.0% | |||
| NB | μmax, G, I, I1/2, I1/4, D, Dc, H, En, Ed, W, R, Z, S | 83.2% | S. aureus | 88.0% | 100.0% | 100.0% |
| S. epidermidis | 84.0% | 91.0% | 70.0% | |||
| (14 parameters) | S. haemolyticus | 76.0% | 97.0% | 86.4% | ||
| S. saprophyticus | 88.0% | 95.0% | 81.5% | |||
| S. simulans | 80.0% | 96.0% | 83.3% | |||
| ANN | μmax, G, I, I1/2, I1/4, D, Dc, H, En, Ed, W, R, Z, S | 99.2% | S. aureus | 100.0% | 100.0% | 100.0% |
| S. epidermidis | 100.0% | 100.0% | 100.0% | |||
| (14 parameters) | S. haemolyticus | 96.0% | 100.0% | 100.0% | ||
| S. saprophyticus | 100.0% | 99.0% | 96.2% | |||
| S. simulans | 100.0% | 100.0% | 100.0% | |||
| SVM | μmax, G, I, I1/2, I1/4, D, Dc, H, En, Ed, W, R, Z, S | 98.4% | S. aureus | 100.0% | 100.0% | 100.0% |
| S. epidermidis | 96.0% | 99.0% | 96.0% | |||
| (14 parameters) | S. haemolyticus | 100.0% | 100.0% | 100.0% | ||
| S. saprophyticus | 100.0% | 100.0% | 100.0% | |||
| S. simulans | 96.0% | 99.0% | 96.0% | |||
| RF | μmax, G, I, I1/2, I1/4, D, Dc, H, En, Ed, W, R, Z, S | 100.0% | S. aureus | 100.0% | 100.0% | 100.0% |
| S. epidermidis | 100.0% | 100.0% | 100.0% | |||
| (14 parameters) | S. haemolyticus | 100.0% | 100.0% | 100.0% | ||
| S. saprophyticus | 100.0% | 100.0% | 100.0% | |||
| S. simulans | 100.0% | 100.0% | 100.0% |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Maeda, Y.; Sugiyama, Y.; Kogiso, A.; Lim, T.-K.; Harada, M.; Yoshino, T.; Matsunaga, T.; Tanaka, T. Colony Fingerprint-Based Discrimination of Staphylococcus species with Machine Learning Approaches. Sensors 2018, 18, 2789. https://doi.org/10.3390/s18092789
Maeda Y, Sugiyama Y, Kogiso A, Lim T-K, Harada M, Yoshino T, Matsunaga T, Tanaka T. Colony Fingerprint-Based Discrimination of Staphylococcus species with Machine Learning Approaches. Sensors. 2018; 18(9):2789. https://doi.org/10.3390/s18092789
Chicago/Turabian StyleMaeda, Yoshiaki, Yui Sugiyama, Atsushi Kogiso, Tae-Kyu Lim, Manabu Harada, Tomoko Yoshino, Tadashi Matsunaga, and Tsuyoshi Tanaka. 2018. "Colony Fingerprint-Based Discrimination of Staphylococcus species with Machine Learning Approaches" Sensors 18, no. 9: 2789. https://doi.org/10.3390/s18092789
APA StyleMaeda, Y., Sugiyama, Y., Kogiso, A., Lim, T.-K., Harada, M., Yoshino, T., Matsunaga, T., & Tanaka, T. (2018). Colony Fingerprint-Based Discrimination of Staphylococcus species with Machine Learning Approaches. Sensors, 18(9), 2789. https://doi.org/10.3390/s18092789





