Ultrasensitive Electrochemical Detection of Clostridium perfringens DNA Based Morphology-Dependent DNA Adsorption Properties of CeO2 Nanorods in Dairy Products
Abstract
1. Introduction
2. Experimental
2.1. Materials and Reagents
- ssDNA: GCT CCT TTG GTT GAA TGA TG
- tDNA: CAT CAT TCA ACC AAA GGA GC
- one base-mismatched DNA: CAG CAT TCA ACC AAA GGA GC
- three base-mismatched DNA: CAG CAT TCA ACT AAC GGA GC
- non-complementary DNA: GGC GAG CGT TAT CCG GAT TT
2.2. Synthesis of Nanoceria
2.3. Apparatus and Characterization
2.4. Preparation of the CeO2/CHIT/GCE
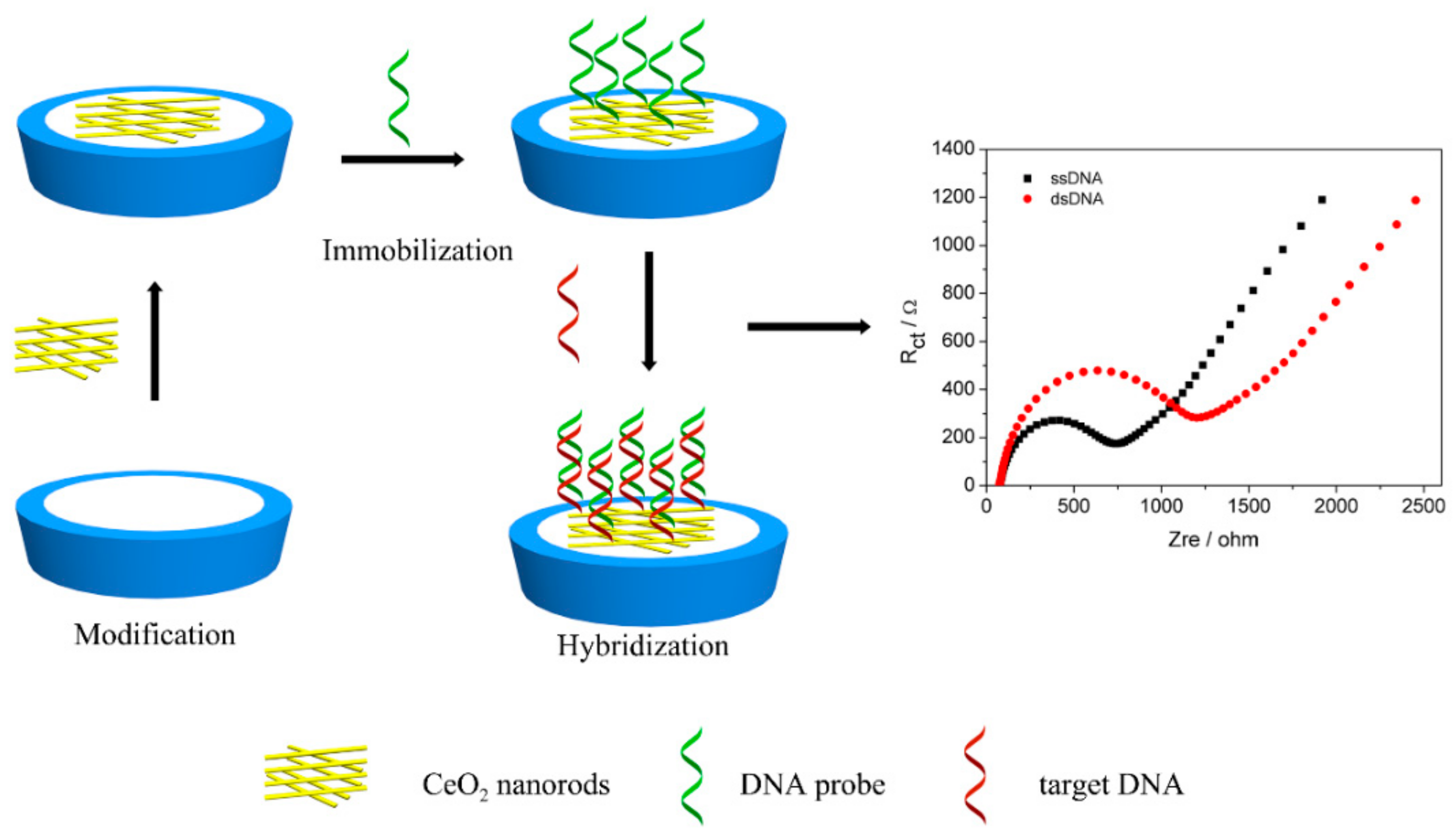
2.5. DNA Probe Immobilization and Hybridization
2.6. Electrochemical Measurements
2.7. Real Sample Assay
3. Results and Discussion
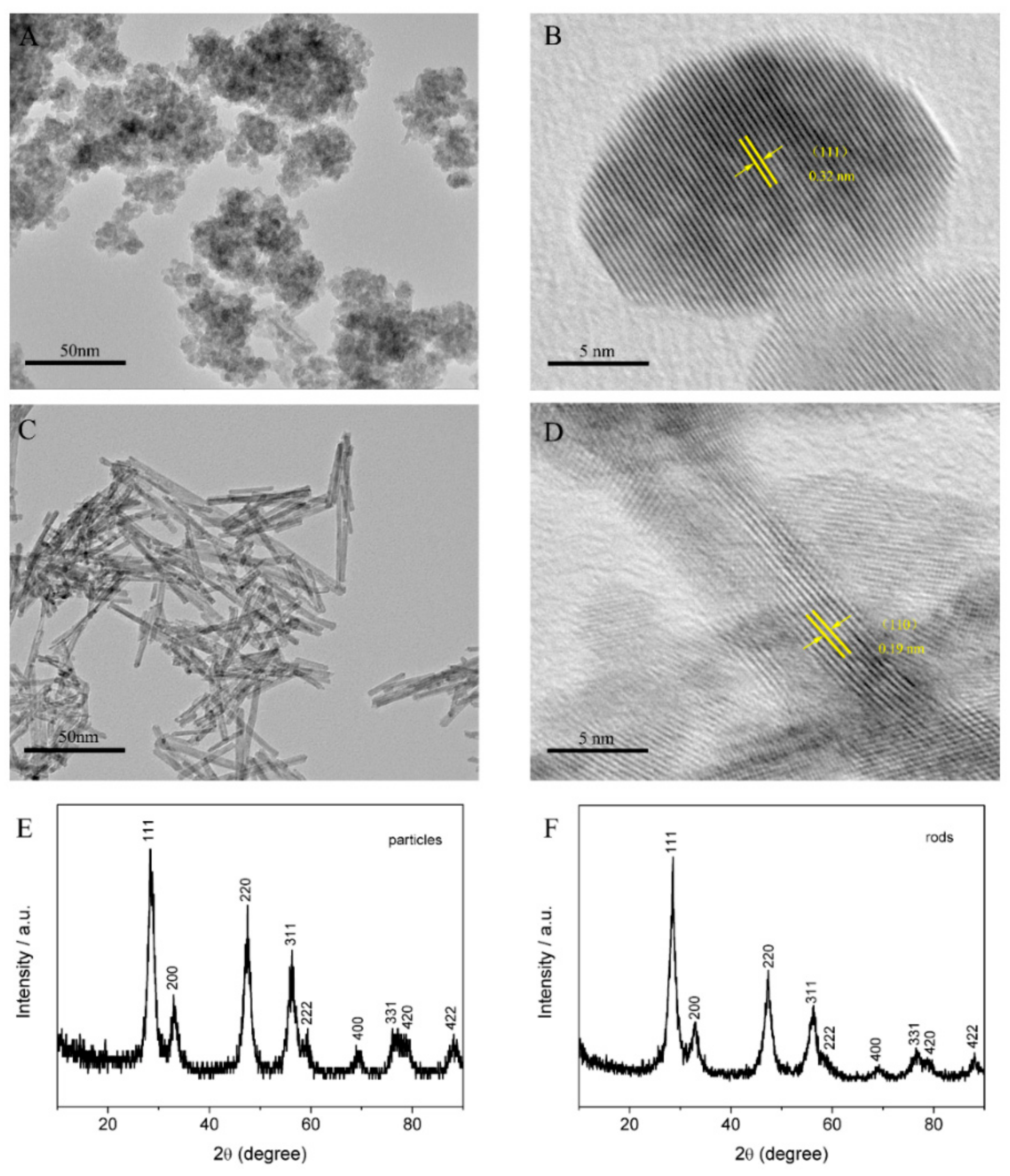
3.1. Morphological and Structural Characterization
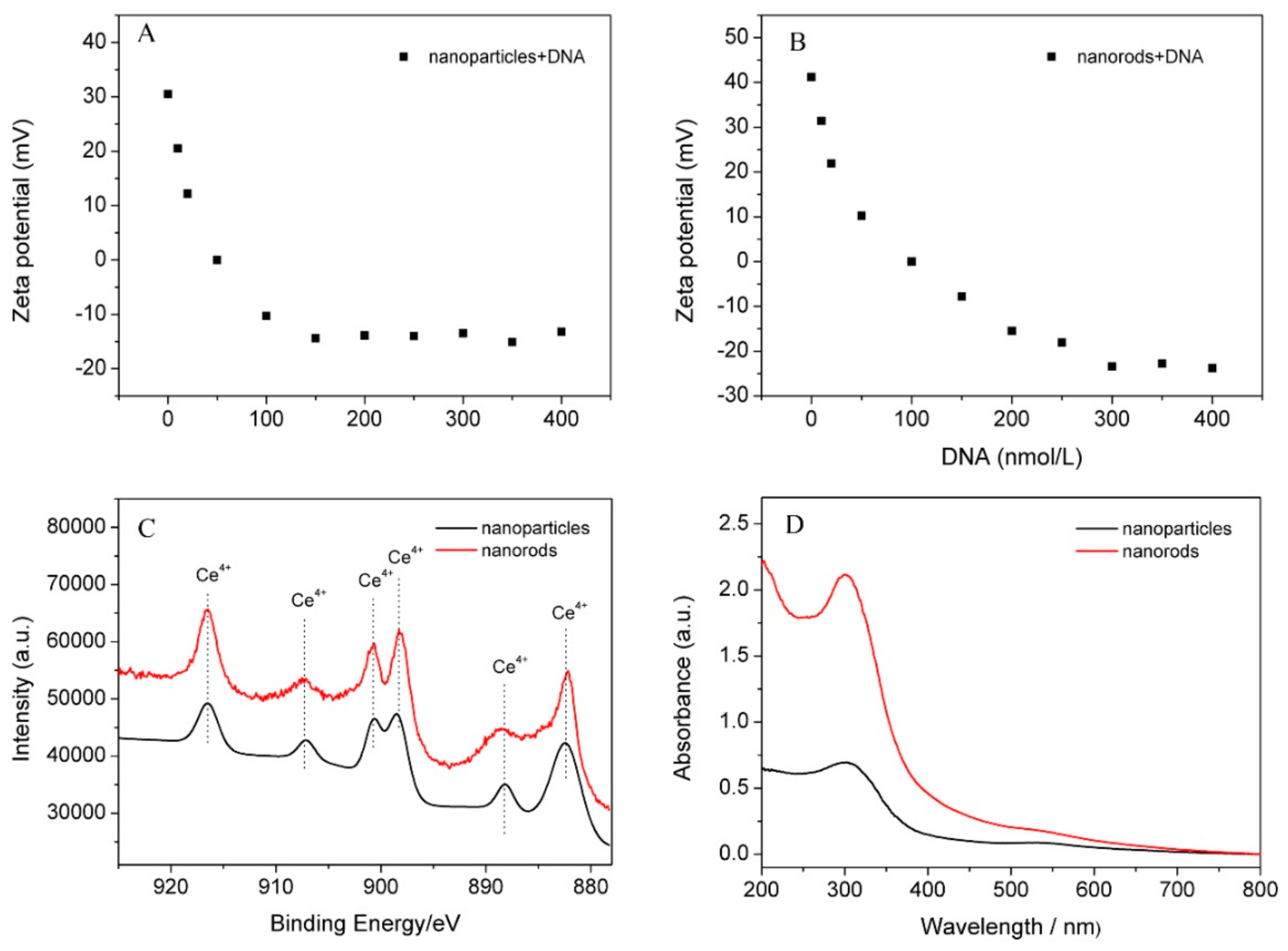
3.2. The Surface State of the Synthesized Nanoceria
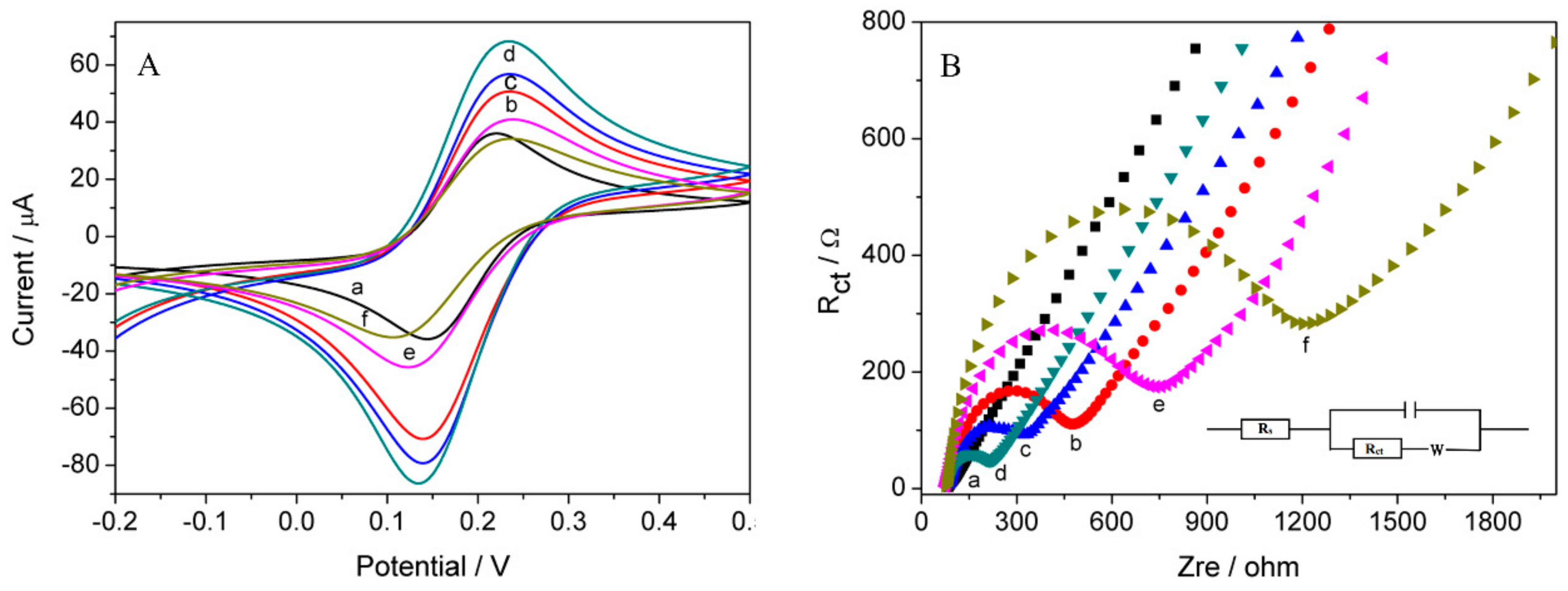
3.3. Electrochemical Characterization of Biosensor
3.4. Electrochemical Studies of DNA Immobilization
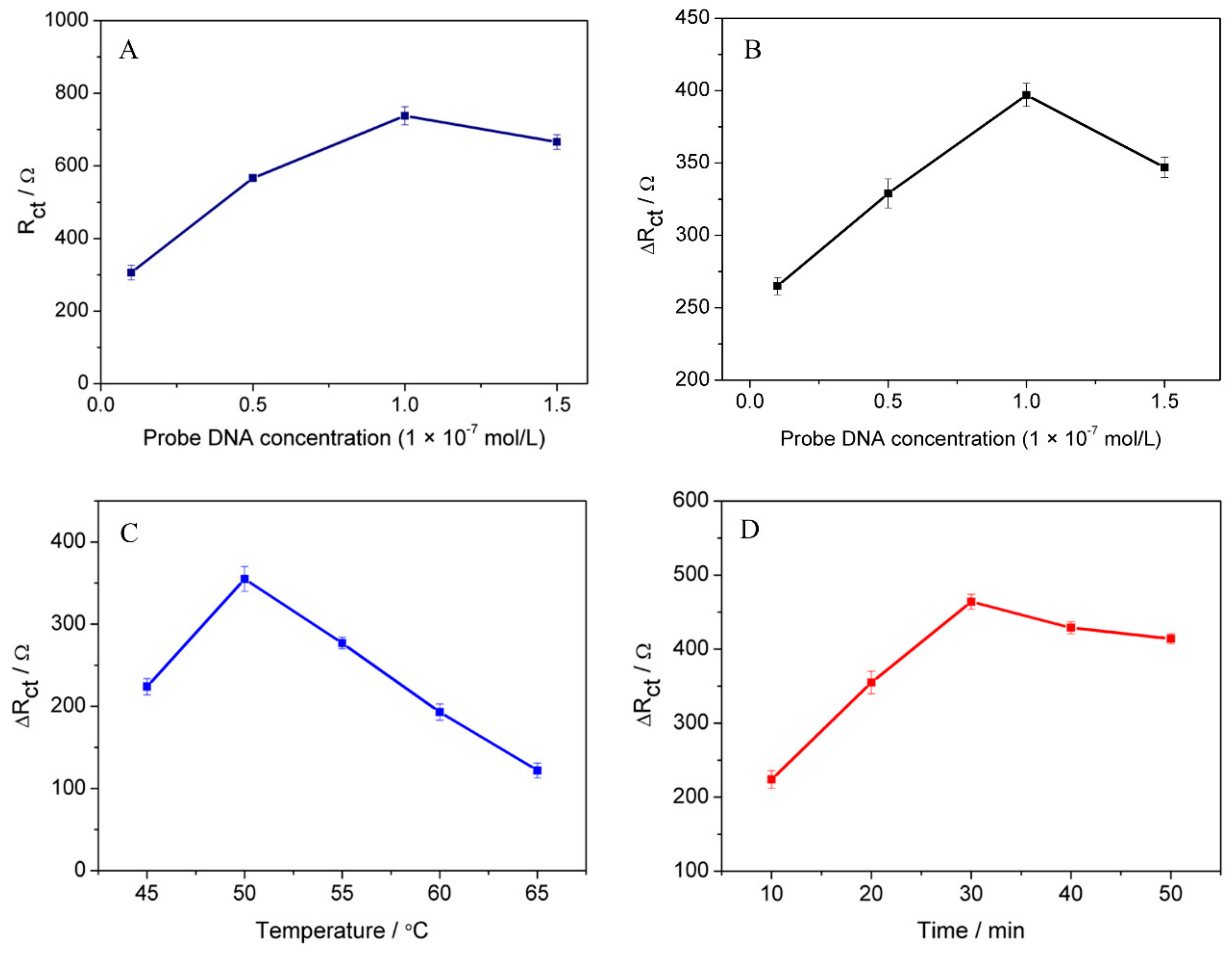
3.5. Optimization of DNA Hybridization
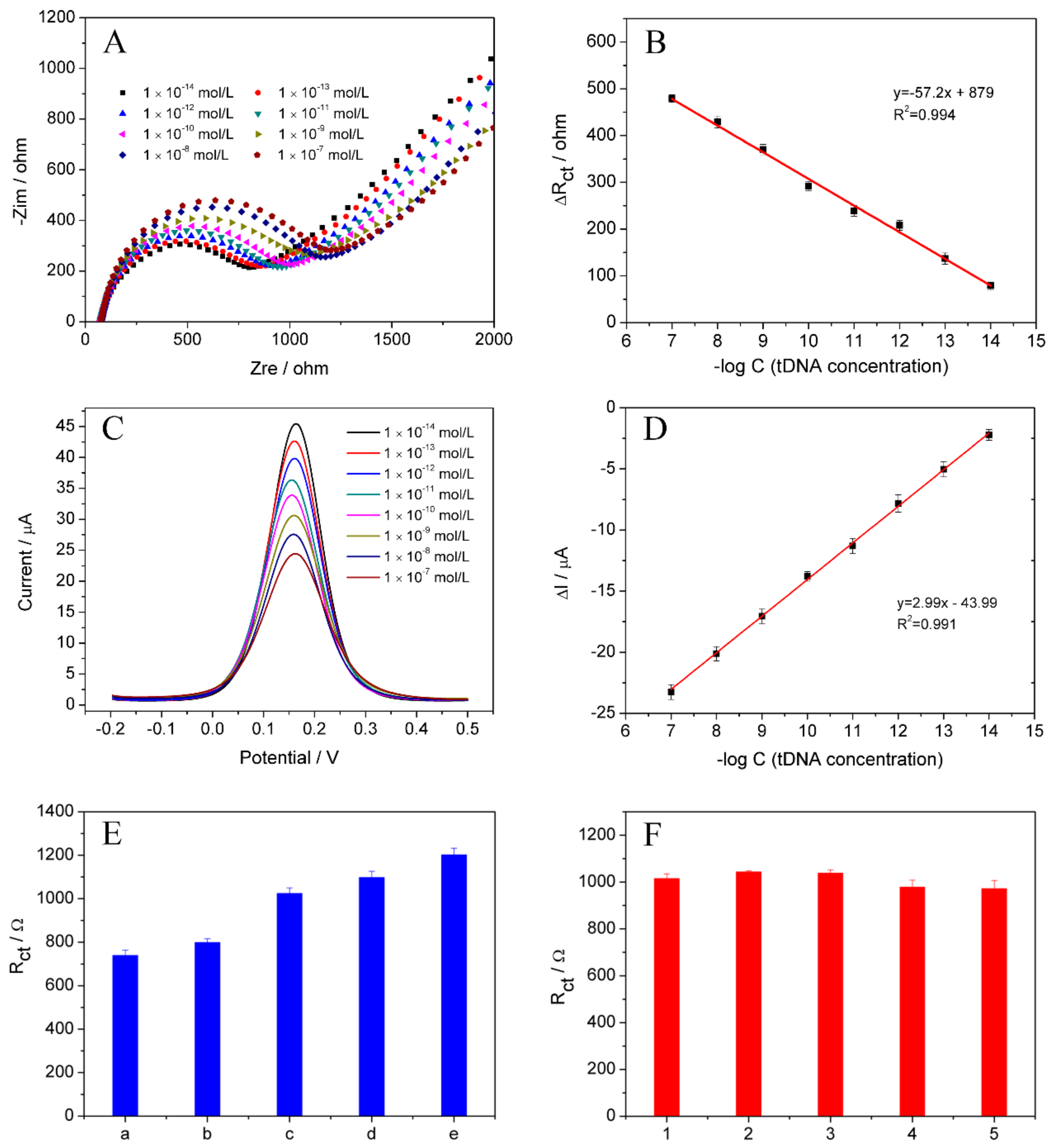
3.6. Quantitative Analysis of C. perfringens Sequence
3.7. The Selectivity of DNA Biosensor
3.8. The Producibility, Stability, and Regeneration Ability of DNA Biosensor
3.9. Detection of tDNA in Dairy Products
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Law, J.W.F.; Ab Mutalib, N.S.; Chan, K.G.; Lee, L.H. Rapid methods for the detection of foodborne bacterial pathogens: Principles, applications, advantages and limitations. Front. Microbiol. 2015, 5, 770–788. [Google Scholar] [CrossRef] [PubMed]
- Petit, L.; Gibert, M.; Popoff, M.R. Clostridium perfringens: Toxinotype and genotype. Trends Microbiol. 1999, 7, 104–110. [Google Scholar] [CrossRef]
- Brynestad, S.; Granum, P.E. Clostridium perfringens and foodborne infections. Int. J. Food Microbiol. 2002, 74, 195–202. [Google Scholar] [CrossRef]
- Wise, M.G.; Siragusa, G.R. Quantitative Detection of Clostridium perfringens in the Broiler Fowl Gastrointestinal Tract by Real-Time PCR. Appl. Environ. Microbiol. 2005, 71, 3911–3916. [Google Scholar] [CrossRef] [PubMed]
- Sharma, H.; Mutharasan, R. Review of biosensors for foodborne pathogens and toxins. Sens. Actuators B 2013, 183, 535–549. [Google Scholar] [CrossRef]
- Yang, Z.Y.; Shim, W.B.; Kim, K.Y.; Chung, D.H. Rapid detection of enterotoxigenic Clostridium perfringens in meat samples using immunomagnetic separation polymerase chain reaction (IMS-PCR). J. Agric. Food Chem. 2010, 58, 7135–7140. [Google Scholar] [CrossRef] [PubMed]
- Albini, S.; Brodard, I.; Jaussi, A.; Wollschlaeger, N.; Frey, J.; Miserez, R.; Abril, C. Real-time multiplex PCR assays for reliable detection of Clostridium perfringens toxin genes in animal isolates. Vet. Microbiol. 2008, 127, 179–185. [Google Scholar] [CrossRef] [PubMed]
- McCourt, M.T.; Finlay, D.A.; Laird, C.; Smyth, J.A.; Bell, C.; Ball, H.J. Sandwich ELISA detection of Clostridium perfringens cells and alpha-toxin from field cases of necrotic enteritis of poultry. Vet. Microbiol. 2005, 106, 259–264. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.N.; Liu, F.; Liu, C.; Liu, L.L.; Li, Y.; Pu, X.Y. Induction of an electrochemiluminescence sensor for DNA detection of Clostridium perfringens based on rolling circle amplification. Anal. Methods 2014, 6, 1558–1562. [Google Scholar] [CrossRef]
- Kang, X.B.; Chen, Q.S.; Liang, X.Y.; Pang, G.C. Fabrication of Bacillus cereus electrochemical immunosensor based on double-layer gold nanoparticles and chitosan. Sens. Actuators B 2013, 177, 1010–1016. [Google Scholar] [CrossRef]
- Huang, H.Y.; Bai, W.Q.; Dong, C.X.; Guo, R.; Liu, Z.H. An ultrasensitive electrochemical DNA biosensor based on graphene/Au nanorod/polythionine for human papillomavirus DNA detection. Biosens. Bioelectron. 2015, 68, 442–446. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.X.; Nan, F.X.; Zhao, J.L.; Yang, T.; Ge, T.; Jiao, K. A label-free ultrasensitive electrochemical DNA sensor based on thin-layer MoS2 nanosheets with high electrochemical activity. Biosens. Bioelectron. 2015, 64, 386–391. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.L.; Li, X.; Yan, S.L.; Wang, M.M.; Liu, P.; Dong, Y.L.; Zhang, C.C. Single-walled carbon nanotubes-carboxyl functionalized graphene oxide based electrochemical DNA biosensor for thermolabilehemolysin gene detection. Anal. Methods 2015, 7, 5303–5310. [Google Scholar] [CrossRef]
- Oretskaya, T.S.; Romanova, E.A.; Andreev, S.Y.; Antsypovich, S.I.; Tóth, C.; Gajdos, V.; Hianik, T. A novel method of introducing hydrophobic moieties into oligonucleotides for covalent and non-covalent immobilization on electrode surfaces. Bioelectrochemistry 2002, 56, 47–51. [Google Scholar] [CrossRef]
- Jarczewska, M.; Górski, Ł.; Malinowska, E. Electrochemical aptamer-based biosensors as potential tools for clinical diagnostics. Anal. Methods 2016, 8, 3861–3877. [Google Scholar] [CrossRef]
- Yang, F.; Jiang, X.Y.; Zhong, X.; Wei, S.P.; Yuan, R. Highly sensitive electrochemiluminescence detection of mucin1 based on V2O5 nanospheres as peroxidase mimetics to catalyze H2O2 for signal amplification. Sens. Actuators B 2018, 265, 126–133. [Google Scholar] [CrossRef]
- Hajdukiewicz, J.; Boland, S.; Kavanagh, P.; Leech, D. An enzyme-amplified amperometric DNA hybridisation assay using DNA immobilised in a carboxymethylated dextran film anchored to a graphite surface. Biosens. Bioelectron. 2010, 25, 1037–1042. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.W. Adsorption of DNA onto gold nanoparticles and graphene oxide: Surface science and applications. Phys. Chem. Chem. Phys. 2012, 14, 10485–10496. [Google Scholar] [CrossRef] [PubMed]
- He, S.J.; Song, B.; Li, D.; Zhu, C.F.; Qi, W.P.; Wen, Y.Q.; Wang, L.H.; Song, S.P.; Fang, H.P.; Fan, C.H. A Graphene Nanoprobe for Rapid, Sensitive, and Multicolor Fluorescent DNA Analysis. Adv. Funct. Mater. 2010, 20, 453–459. [Google Scholar] [CrossRef]
- Le, M.H.; Jimenez, C.; Chainet, E.; Stambouli, V. A Label-Free Impedimetric DNA Sensor Based on a Nanoporous SnO2 Film: Fabrication and Detection Performance. Sensors 2015, 15, 10686–10704. [Google Scholar] [CrossRef] [PubMed]
- Zuo, L.M.; Qu, Q.; Li, L.; Ran, X.; Gui, J.W.; Wang, Q.; Cui, X.H.; Jiang, C.L. Electrochemical DNA Biosensor Based on Magnetite/Multiwalled Carbon Nanotubes/Chitosan Nanocomposite for Bacillus Cereus Detection of Potential Marker for Gold Prospecting. Electroanalysis 2018, 30, 910–920. [Google Scholar] [CrossRef]
- Erdem, A. Nanomaterial-based electrochemical DNA sensing strategies. Talanta 2007, 74, 318–325. [Google Scholar] [CrossRef] [PubMed]
- Malhotra, B.D.; Kaushik, A. Metal oxide-chitosan based nanocomposite for cholesterol biosensor. Thin Solid Films 2009, 518, 614–620. [Google Scholar] [CrossRef]
- Qu, X.M.; You, L.X.; Tian, X.C.; Zhang, B.W.; Mahadevan, G.D.; Jiang, Y.X.; Sun, S.G. CeO2 nanorods with high energy surfaces as electrocatalytical supports for methanol electrooxidation. Microchim. Acta 2015, 182, 1078–1084. [Google Scholar] [CrossRef]
- Deori, K.; Gupta, D.; Saha, B.; Deka, S. Design of 3-Dimensionally Self-Assembled CeO2 Nanocube as a Breakthrough Catalyst for Efficient Alkylarene Oxidation in Water. ACS Catal. 2014, 4, 3169–3179. [Google Scholar] [CrossRef]
- Zhang, W.; Yang, T.; Zhuang, X.M.; Guo, Z.Y.; Jiao, K. An ionic liquid supported CeO2 nanoshuttles-carbon nanotubes composite as a platform for impedance DNA hybridization sensing. Biosens. Bioelectron. 2009, 24, 2417–2422. [Google Scholar] [CrossRef] [PubMed]
- Imagawa, H.; Sun, S.H. Controlled Synthesis of Monodisperse CeO2 Nanoplates Developed from Assembled Nanoparticles. J. Phys. Chem. C 2012, 116, 2761–2765. [Google Scholar] [CrossRef]
- Tan, H.; Wang, J.; Yu, S.; Zhou, K. Support Morphology-Dependent Catalytic Activity of Pd/CeO2 for Formaldehyde Oxidation. Environ. Sci. Technol. 2015, 49, 8675–8682. [Google Scholar] [CrossRef] [PubMed]
- Kang, D.J.; Yu, X.L.; Ge, M.F. Morphology-dependent properties and adsorption performance of CeO2 for fluoride removal. Chem. Eng. J. 2017, 330, 36–43. [Google Scholar] [CrossRef]
- Bolat, G.; Abaci, S. Non-Enzymatic Electrochemical Sensing of Malathion Pesticide in Tomato and Apple Samples Based on Gold Nanoparticles-Chitosan-Ionic Liquid Hybrid Nanocomposite. Sensors 2018, 18, 773. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.Y.; Lauder, H.; Cruwys, H.; Falletta, P.; Beaudette, L.A. Development and application of an oligonucleotide microarray and real-time quantitative PCR for detection of wastewater bacterial pathogens. Sci. Total Environ. 2008, 398, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Mai, H.X.; Sun, L.D.; Zhang, Y.W.; Si, R.; Feng, W.; Zhang, H.P.; Liu, H.C.; Yan, C.H. Shape-selective synthesis and oxygen storage behavior of ceria nanopolyhedra, nanorods, and nanocubes. J. Phys. Chem. B 2005, 109, 24380–24385. [Google Scholar] [CrossRef] [PubMed]
- Ding, N.; Yan, N.; Ren, C.L.; Chen, X.G. Colorimetric determination of melamine in dairy products by Fe3O4 magnetic nanoparticles-H2O2-ABTS detection system. Anal. Chem. 2010, 82, 5897–5899. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Gong, M.; Cao, Y.D.; Wang, C.A. Facile synthesis of well-dispersed CeO2-CuOx composite hollow spheres with superior Catalytic Activity for CO oxidation. RSC Adv. 2015, 5, 95133–95139. [Google Scholar] [CrossRef]
- Pereira, R.; Rocha-Santos, T.A.; Antunes, F.E.; Rasteiro, M.G.; Ribeiro, R.; Gonçalves, F.; Soares, A.M.; Lopes, I. Screening evaluation of the ecotoxicity and genotoxicity of soils contaminated with organic and inorganic nanoparticles: The role of ageing. J. Hazard. Mater. 2011, 194, 345–354. [Google Scholar] [CrossRef] [PubMed]
- Pautler, R.; Kelly, E.Y.; Huang, P.J.; Cao, J.; Liu, B.W.; Liu, J.J. Attaching DNA to nanoceria: Regulating oxidase activity and fluorescence quenching. ACS Appl. Mater. Interfaces 2013, 5, 6820–6825. [Google Scholar] [CrossRef] [PubMed]
- Gao, W.; Wei, X.P.; Wang, X.J.; Cui, G.W.; Liu, Z.H.; Tang, B. A competitive coordination-based CeO2 nanowire-DNA nanosensor: Fast and selective detection of hydrogen peroxide in living cells and in vivo. Chem. Commun. 2016, 52, 3643–3646. [Google Scholar] [CrossRef] [PubMed]
- Óvári, L.; Calderon, S.K.; Lykhach, Y.; Libuda, J.; Erdőhelyi, A.; Papp, C.; Kiss, J.; Steinrück, H.-P. Near ambient pressure XPS investigation of the interaction of ethanol with Co/CeO2 (111). J. Catal. 2013, 307, 132–139. [Google Scholar] [CrossRef]
- Heckert, E.G.; Karakoti, A.S.; Seal, S.; Self, W.T. The role of cerium redox state in the SOD mimetic activity of nanoceria. Biomaterials 2008, 29, 2705–2709. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.; Wang, W.Z.; Zhang, L.; Zheng, Y.L.; Wang, Z. Insights into the Surface-Defect Dependence of Photoreactivity over CeO2 Nanocrystals with Well-Defined Crystal Facets. ACS Catal. 2015, 5, 4851–4858. [Google Scholar] [CrossRef]
- Cai, M.Z.; Zhu, L.; Ding, Y.P.; Wang, J.X.; Li, J.S.; Du, X.Y. Determination of sulfamethoxazole in foods based on CeO2/chitosan nanocomposite-modified electrodes. Mater. Sci. Eng. C-Mater. 2012, 32, 2623–2627. [Google Scholar] [CrossRef]
- Izadi, Z.; Sheikh-Zeinoddin, M.; Ensafi, A.A.; Soleimanian-Zad, S. Fabrication of an electrochemical DNA-based biosensor for Bacillus cereus detection in milk and infant formula. Biosens. Bioelectron. 2016, 80, 582–589. [Google Scholar] [CrossRef] [PubMed]
- Meenakshi, S.; Pandian, K.; Jayakumari, L.S.; Inbasekaran, S. Enhanced amperometric detection of metronidazole in drug formulations and urine samples based on chitosan protected tetrasulfonated copper phthalocyanine thin-film modified glassy carbon electrode. Mater. Sci. Eng. C-Mater. 2016, 59, 136–144. [Google Scholar] [CrossRef] [PubMed]
- Radhakrishnan, S.; Sumathi, C.; Dharuman, V.; Wilson, J. Polypyrrole nanotubes–polyaniline composite for DNA detection using methylene blue as intercalator. Anal. Methods 2013, 5, 1010–1015. [Google Scholar] [CrossRef]
- Shamsi, M.H.; Kraatz, H.B. The effects of oligonucleotide overhangs on the surface hybridization in DNA films: An impedance study. Analyst 2011, 136, 3107–3112. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.H.; Liao, X.L.; Ding, Y.T.; Gao, F.; Wang, Q.X. DNA biosensor based on a glassy carbon electrode modified with electropolymerized Eriochrome Black T. Microchim. Acta 2014, 181, 155–162. [Google Scholar] [CrossRef]
- Tezerjani, M.D.; Benvidi, A.; Rezaeinasab, M.; Jahanbani, S.; Moshtaghioun, S.M.; Youssefi, M.; Zarrini, K. An impedimeric biosensor based on a composite of graphene nanosheets and polyaniline as a suitable platform for prostate cancer sensing. Anal. Methods 2016, 8, 7507–7515. [Google Scholar] [CrossRef]
- Manzano, M.; Viezzi, S.; Mazerat, S.; Marks, R.S.; Vidic, J. Rapid and label-free electrochemical DNA biosensor for detecting hepatitis A virus. Biosens. Bioelectron. 2018, 100, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Schreiner, S.M.; Shudy, D.F.; Hatch, A.L.; Opdahl, A. Controlled and Efficient Hybridization Achieved with DNA Probes Immobilized Solely through Preferential DNA-Substrate Interactions. Anal. Chem. 2010, 82, 2803–2810. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.N.; Liu, F.; Zhang, L.Q.; Liu, L.L.; Liu, C.; Pu, X.Y. An electrochemical strategy with molecular beacon and hemin/G-quadruplex for the detection of Clostridium perfringens DNA on screen-printed electrodes. RSC Adv. 2014, 4, 57064–57070. [Google Scholar] [CrossRef]
- Wang, L.B.; Zheng, X.W.; Zhang, W.; Quan, X.Q.; Hu, Q.T.; Wu, W.J.; Zong, P.S.; Wu, M.Z. One-pot synthesis of carbon-decorated FePt nanoparticles and their application for label-free electrochemical impedance sensing of DNA hybridization. RSC Adv. 2013, 3, 9042–9046. [Google Scholar] [CrossRef]
- Zhang, K.Y.; Ma, H.Y.; Zhang, L.P.; Zhang, Y.Z. Fabrication of a Sensitive Impedance Biosensor of DNA Hybridization Based on Gold Nanoparticles Modified Gold Electrode. Electroanalysis 2008, 20, 2127–2133. [Google Scholar] [CrossRef]
| Electrode | Rs (Ω) | Rct (Ω) |
|---|---|---|
| GCE | 81.20 (1.095) | 979.9 (1.780) |
| CHIT/GCE | 76.06 (0.9024) | 373.8 (1.214) |
| CeO2/GCE | 80.15 (1.253) | 270.6 (2.627) |
| CeO2/CHIT/GCE | 74.31 (1.100) | 166.6 (1.704) |
| ssDNA/ CeO2/CHIT/GCE | 76.08 (0.6078) | 629.2 (0.8229) |
| dsDNA/ CeO2/CHIT/GCE | 76.01 (0.6954) | 1088 (0.8843) |
| Modified Material and Electrode | Detection Technique | Liner Range (mol/L) | Detection Limit (mol/L) | References |
|---|---|---|---|---|
| AuNPs/graphene/GCE | DPV | 10−12–10−6 | 10−12 | [50] |
| Au | ECL | 10−15–10−9 | 10−15 | [9] |
| CeO2/CHIT/GCE | EIS | 10−14–10−7 | 10−15 | this work |
| Samples | Added (pmol/L) | Found (pmol/L) | Recovery (%) | RSD (%) |
|---|---|---|---|---|
| pure milk | 10 | 9.77 | 97.7 | 1.66 |
| 20 | 19.9 | 99.5 | 2.74 | |
| 50 | 51.3 | 102.6 | 4.96 | |
| milk powder | 10 | 9.54 | 95.4 | 2.53 |
| 20 | 20.4 | 102 | 3.25 | |
| 50 | 48.9 | 97.8 | 3.45 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qian, X.; Qu, Q.; Li, L.; Ran, X.; Zuo, L.; Huang, R.; Wang, Q. Ultrasensitive Electrochemical Detection of Clostridium perfringens DNA Based Morphology-Dependent DNA Adsorption Properties of CeO2 Nanorods in Dairy Products. Sensors 2018, 18, 1878. https://doi.org/10.3390/s18061878
Qian X, Qu Q, Li L, Ran X, Zuo L, Huang R, Wang Q. Ultrasensitive Electrochemical Detection of Clostridium perfringens DNA Based Morphology-Dependent DNA Adsorption Properties of CeO2 Nanorods in Dairy Products. Sensors. 2018; 18(6):1878. https://doi.org/10.3390/s18061878
Chicago/Turabian StyleQian, Xingcan, Qing Qu, Lei Li, Xin Ran, Limei Zuo, Rui Huang, and Qiang Wang. 2018. "Ultrasensitive Electrochemical Detection of Clostridium perfringens DNA Based Morphology-Dependent DNA Adsorption Properties of CeO2 Nanorods in Dairy Products" Sensors 18, no. 6: 1878. https://doi.org/10.3390/s18061878
APA StyleQian, X., Qu, Q., Li, L., Ran, X., Zuo, L., Huang, R., & Wang, Q. (2018). Ultrasensitive Electrochemical Detection of Clostridium perfringens DNA Based Morphology-Dependent DNA Adsorption Properties of CeO2 Nanorods in Dairy Products. Sensors, 18(6), 1878. https://doi.org/10.3390/s18061878




