Real-Time Tau Protein Detection by Sandwich-Based Piezoelectric Biosensing: Exploring Tubulin as a Mass Enhancer
Abstract
1. Introduction
2. Materials and Methods
2.1. Reagents and Buffers
2.2. Piezoelectric Apparatus
2.3. Sensor Chip Functionalization and Detection Format
2.4. Detection of Tau Protein
3. Results and Discussion
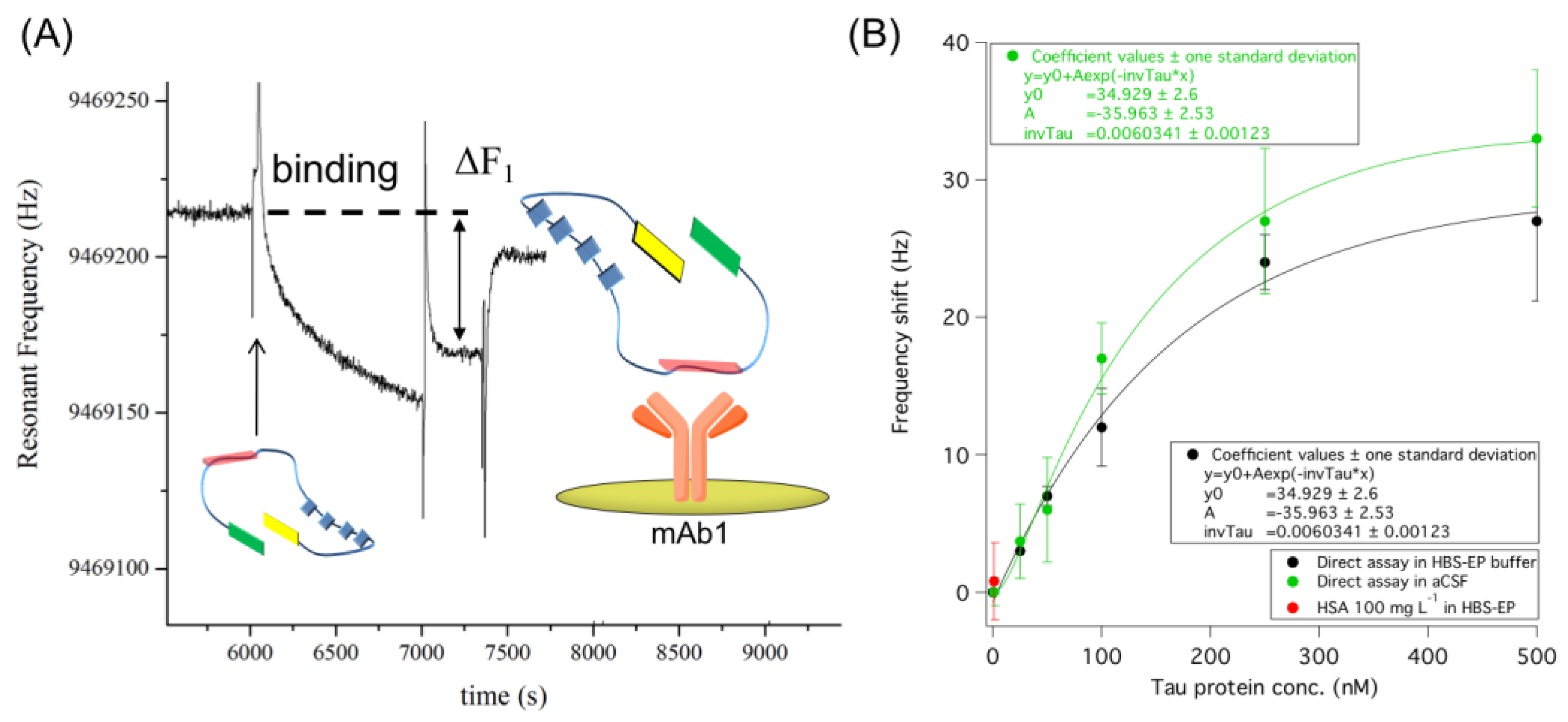
3.1. Direct Immuno-Based Detection of Tau in Buffer and aCSF
3.2. Improving Sensor Performances by Affinity-Based Mass Enhancement
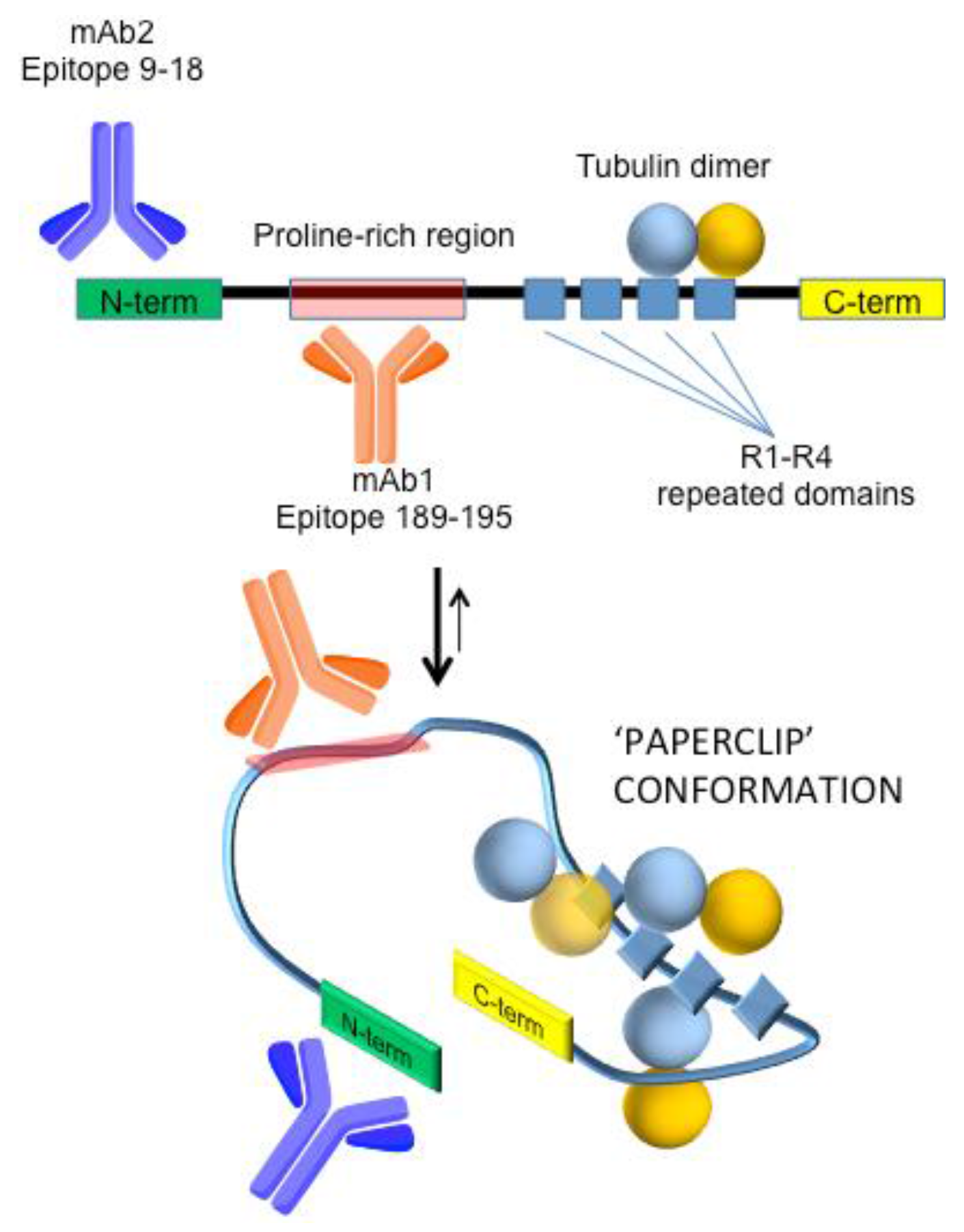
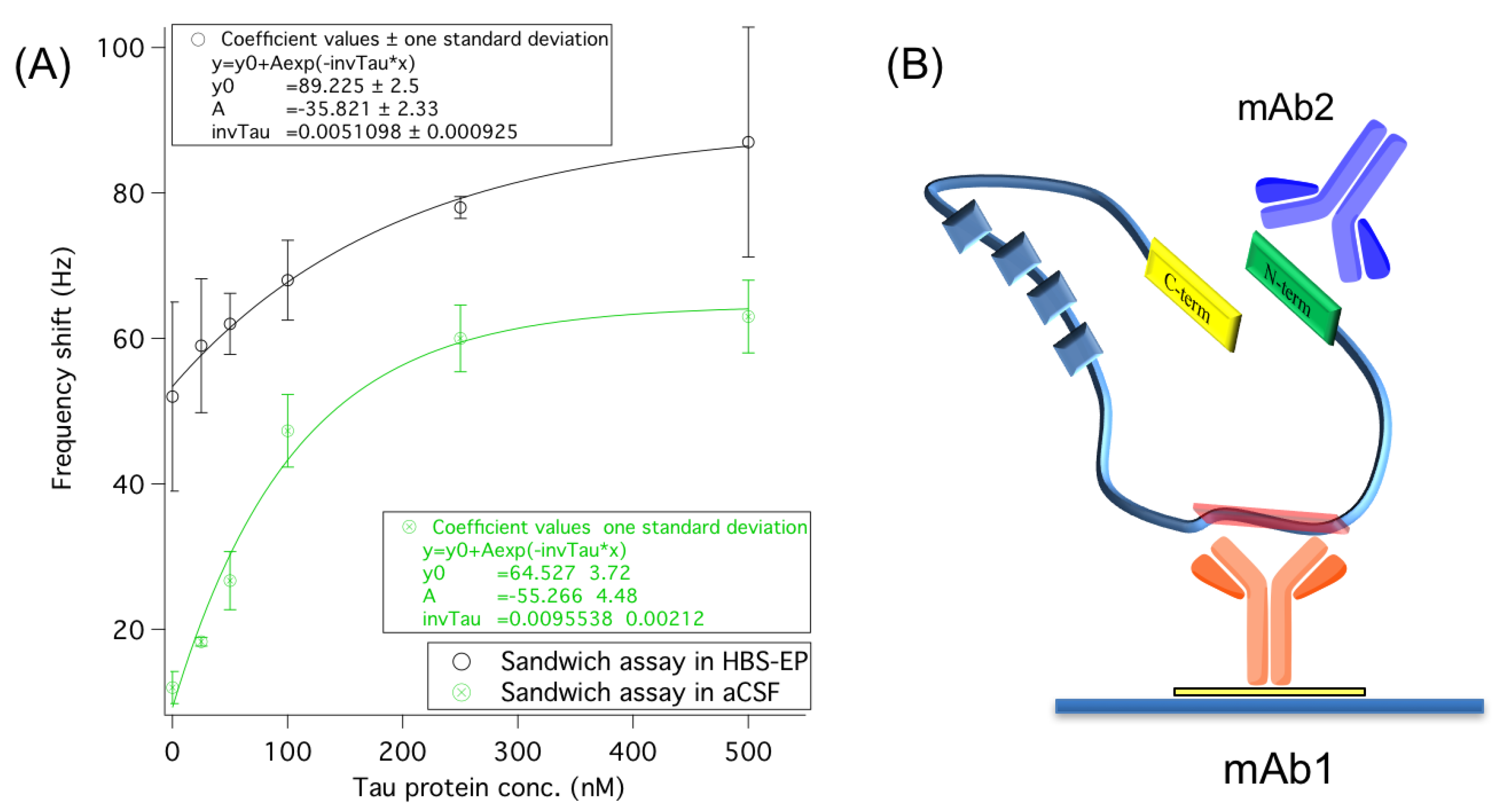
3.2.1. Secondary Monoclonal Antibody
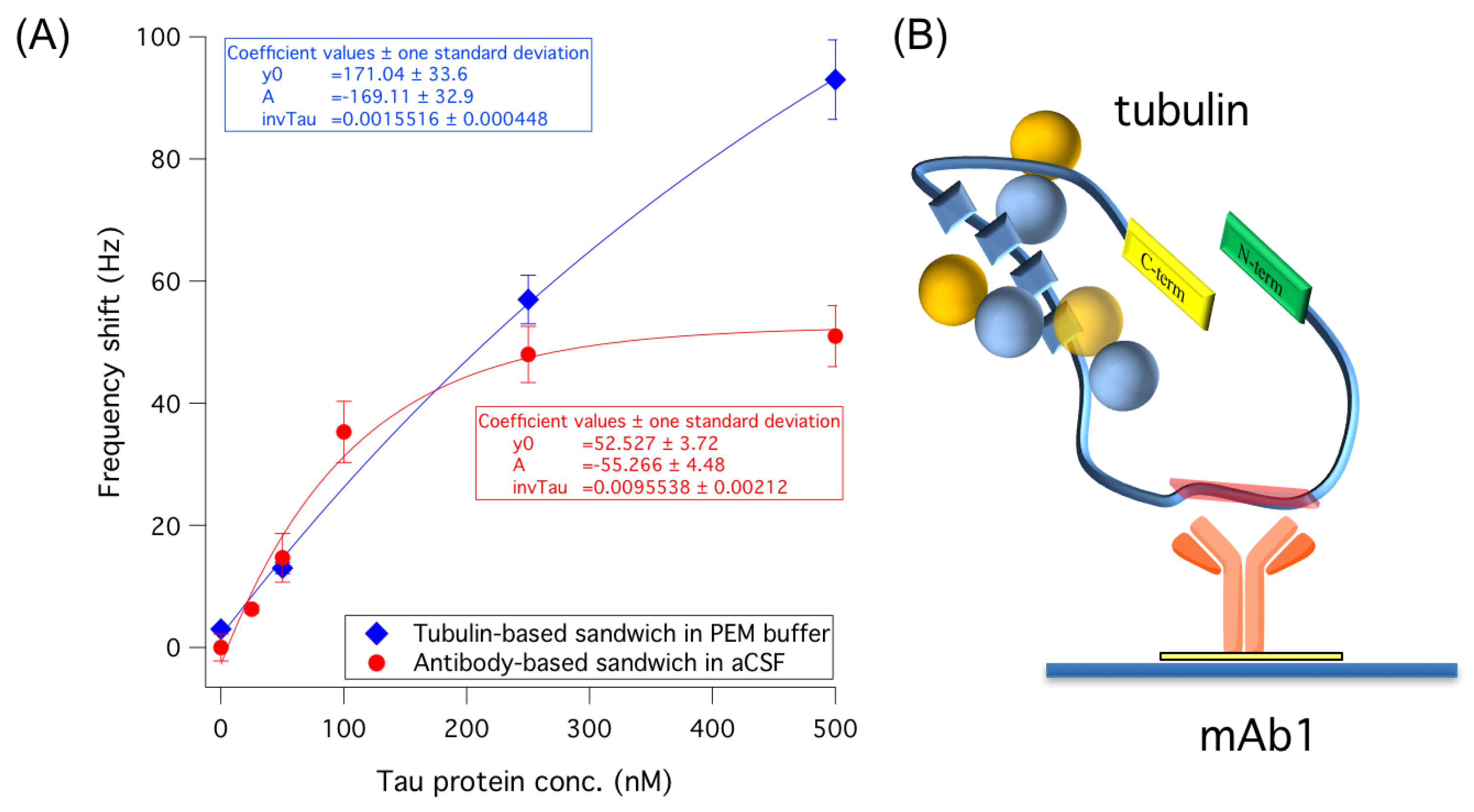
3.2.2. Use of Tubulin as an Alternative to Antibodies in a Sandwich-Like Assay
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- The World Alzheimer Report 2016. Available online: www.alz.co.uk/research/world-report-2016 (accessed on 14 December 2017).
- Lewczuk, P.; Lelental, N.; Lachmann, I.; Holzer, M.; Flach, K.; Brandner, S.; Engelborghs, S.; Teunissen, C.E.; Zetterberg, H.; Molinuevo, J.L.; et al. Non-phosphorylated tau as a potential biomarker of Alzheimer’s disease: Analytical and diagnostic characterization. J. Alzheimer’s Dis. 2017, 55, 159–170. [Google Scholar] [CrossRef] [PubMed]
- Kulic, L.; Unschuld, P.G. Recent advances in cerebrospinal fluid biomarkers for the detection of preclinical Alzheimer’s disease. Curr. Opin. Neurol. 2016, 29, 749–755. [Google Scholar] [CrossRef] [PubMed]
- Scarano, S.; Lisi, S.; Ravelet, C.; Peyrin, E.; Minunni, M. Detecting Alzheimer’s disease biomarkers: From antibodies to new bio-mimetic receptors and their application to established and emerging bioanalytical platforms—A critical review. Anal. Chim. Acta 2016, 94, 21–37. [Google Scholar] [CrossRef]
- Blennow, K.; Hampel, H.; Weiner, M.; Zetterberg, H. Cerebrospinal fluid and plasma biomarkers in Alzheimer disease. Nat. Rev. Neurol. 2010, 6, 131–144. [Google Scholar] [CrossRef] [PubMed]
- Shaw, L.M.; Korecka, M.; Clark, C.M.; Lee, V.M.-Y.; Trojanowski, J.Q. Biomarkers of neurodegeneration for diagnosis and monitoring therapeutics. Nat. Rev. Drug Discov. 2007, 6, 295–303. [Google Scholar] [CrossRef] [PubMed]
- Hernández, F.; Avila, J. Tauopathies. Cell Mol. Life Sci. 2007, 64, 2219–2233. [Google Scholar] [CrossRef] [PubMed]
- Ganesh, H.V.S.; Chow, A.M.; Kerman, K. Recent advances in biosensors for neurodegenerative disease detection. Trends Anal. Chem. 2016, 79, 363–370. [Google Scholar] [CrossRef]
- Kaushik, A.; Jayant, R.D.; Tiwari, S.; Vashist, A.; Nair, M. Nano-biosensors to detect beta-amyloid for Alzheimer’s disease management. Biosens. Bioelectron. 2016, 80, 273–287. [Google Scholar] [CrossRef] [PubMed]
- Saijo, E.; Ghetti, B.; Zanusso, G.; Oblak, A.; Furman, J.L.; Diamond, M.I.; Kraus, A.; Caughey, B. Ultrasensitive and selective detection of 3-repeat tau seeding activity in Pick disease brain and cerebrospinal fluid. Acta Neuropathol. 2017, 133, 751–765. [Google Scholar] [CrossRef] [PubMed]
- Chan, H.-N.; Xu, D.; Ho, S.-L.; Shing Wong, M.; Li, H.-W. Ultra-sensitive detection of protein biomarkers for diagnosis of Alzheimer’s disease. Chem. Sci. 2017, 8, 4012–4018. [Google Scholar] [CrossRef]
- Jeganathan, S.; von Bergen, M.; Brutlach, H.; Steinhoff, H.J.; Mandelkow, E. Global hairpin folding of tau in solution. Biochemistry 2006, 45, 2283–2293. [Google Scholar] [CrossRef] [PubMed]
- Demeritte, T.; Priya, B.; Nellore, V.; Kanchanapally, R.; Sinha, S.S.; Pramanik, A.; Chavva, S.R.; Ray, P.C. Hybrid Graphene Oxide Based Plasmonic-Magnetic Multifunctional Nanoplatform for Selective Separation and Label-Free Identification of Alzheimer’s Disease Biomarkers. ACS Appl. Mater. Interfaces 2015, 7, 13693–13700. [Google Scholar] [CrossRef] [PubMed]
- Lisi, S.; Scarano, S.; Fedeli, S.; Cicchi, S.; Pascale, E.; Ravelet, C.; Peyrin, E.; Minunni, M. Toward sensitive immuno-based detection of tau protein by surface plasmon resonance coupled to carbon nanostructures as signal amplifiers. Biosens. Bioelectron. 2017, 93, 289–292. [Google Scholar] [CrossRef] [PubMed]
- Okuno, H.; Mori, K.; Okada, T.; Yokoyama, Y.; Suzuki, H. Development of aggregation inhibitors for amyloid-β peptides and their evaluation by quartz-crystal microbalance. Chem. Biol. Drug Des. 2007, 69, 356–361. [Google Scholar] [CrossRef] [PubMed]
- Kotarek, J.A.; Moss, M.A. Impact of phospholipid bilayer saturation on amyloid-beta protein aggregation intermediate growth: A quartz crystal microbalance analysis. Anal. Biochem. 2010, 399, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Johnson, K.C.; Moss, M.A. Quartz crystal microbalance analysis of growth kinetics for aggregation intermediates of the amyloid-beta protein. Anal. Biochem. 2008, 378, 15–24. [Google Scholar] [CrossRef]
- Wang, S.X.; Acha, D.; Shah, A.J.; Hills, F.; Roitt, I.; Demosthenous, A.; Bayford, R.H. Detection of the tau protein in human serum by a sensitive four-electrode electrochemical biosensor. Biosens. Bioelectron. 2017, 92, 482–488. [Google Scholar] [CrossRef] [PubMed]
- Esteves-Villanueva, J.O.; Trzeciakiewicz, H.; Martic, S. A protein-based electrochemical biosensor for detection of tau protein, a neurodegenerative disease biomarker. Analyst 2014, 139, 2823–2831. [Google Scholar] [CrossRef] [PubMed]
- Jahnke, H.G.; Rothermel, A.; Sternberger, I.; Mack, T.G.A.; Kurz, R.G.; Pänke, O.; Striggow, F.; Robitzki, A.A. An impedimetric microelectrode-based array sensor for label-free detection of tau hyperphosphorylation in human cells. Lab Chip 2009, 9, 1422–1428. [Google Scholar] [CrossRef] [PubMed]
- Kristofikova, Z.; Ripova, D.; Hegnerová, K.; Sirova, J.; Homola, J. Protein τ-Mediated Effects on Rat Hippocampal Choline Transporters CHT1 and τ-Amyloid β Interactions. Neurochem. Res. 2013, 38, 1949–1959. [Google Scholar] [CrossRef] [PubMed]
- Kerman, K.; Kim, D.K.; Hiep, H.M.; Tamiya, E. Detection of Alzheimer’s tau protein using localised surface plasmon resonance-based immunochip. Talanta 2008, 74, 1038–1042. [Google Scholar] [CrossRef]
- Esteves-Villanueva, J.O.; Martic-Milne, S. Electrochemical detection of anti-tau antibodies binding to tau protein and inhibition of GSK-3β-catalyzed phosphorylation. Anal. Biochem. 2016, 496, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Ackmann, M.; Wiech, H.; Mandelkow, E. Nonsaturable binding indicates clustering of tau on the microtubule surface in a paired helical filament-like conformation. J. Biol. Chem. 2000, 275, 30335–30343. [Google Scholar] [CrossRef] [PubMed]
- Drubin, D.; Kirschner, M. Tau protein function in living cells. J. Cell. Biol. 1986, 103, 2739–2746. [Google Scholar] [CrossRef] [PubMed]
- Devred, F.; Barbier, P.; Douillard, S.; Monasterio, O.; Andreu, J.M.; Peyrot, V. Tau induces ring and microtubule formation from alphabeta-tubulin dimers under nonassembly conditions. Biochemistry 2004, 43, 10520–10531. [Google Scholar] [CrossRef] [PubMed]
- Kadavath, H.; Hofele, R.V.; Biernat, J.; Kumar, S.; Tepper, K.; Urlaub, H.; Mandelkow, E.; Zweckstetter, M. Tau stabilizes microtubules by binding at the interface between tubulin heterodimers. Proc. Natl. Acad. Sci. USA 2015, 112, 7501–7506. [Google Scholar] [CrossRef] [PubMed]
- Folker, E.; Baker, B.; Goodson, H. Phylogenetic analysis of the formin homology 2 domain. Mol. Biol. Cell 2005, 16, 1–13. [Google Scholar] [CrossRef]
- Choudhury, D.; Kamalesh Chaudhari, P.L.; John, R.; Dasgupta, A.K.; Pradeep, T.; Chakrabarti, G. Unprecedented inhibition of tubulin polymerization directed by gold nanoparticles inducing cell cycle arrest and apoptosis. Nanoscale 2013, 5, 4476–4489. [Google Scholar] [CrossRef] [PubMed]
- Hall, D.; Minton, A.P. Turbidity as a probe of tubulin polymerization kinetics: A theoretical and experimental re-examination. Anal. Biochem. 2005, 345, 198–213. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, D.; Scarano, S.; Lisi, S.; Palladino, P.; Minunni, M. Real-Time Tau Protein Detection by Sandwich-Based Piezoelectric Biosensing: Exploring Tubulin as a Mass Enhancer. Sensors 2018, 18, 946. https://doi.org/10.3390/s18040946
Li D, Scarano S, Lisi S, Palladino P, Minunni M. Real-Time Tau Protein Detection by Sandwich-Based Piezoelectric Biosensing: Exploring Tubulin as a Mass Enhancer. Sensors. 2018; 18(4):946. https://doi.org/10.3390/s18040946
Chicago/Turabian StyleLi, Dujuan, Simona Scarano, Samuele Lisi, Pasquale Palladino, and Maria Minunni. 2018. "Real-Time Tau Protein Detection by Sandwich-Based Piezoelectric Biosensing: Exploring Tubulin as a Mass Enhancer" Sensors 18, no. 4: 946. https://doi.org/10.3390/s18040946
APA StyleLi, D., Scarano, S., Lisi, S., Palladino, P., & Minunni, M. (2018). Real-Time Tau Protein Detection by Sandwich-Based Piezoelectric Biosensing: Exploring Tubulin as a Mass Enhancer. Sensors, 18(4), 946. https://doi.org/10.3390/s18040946






