Design of Artificial Riboswitches as Biosensors
Abstract
1. Introduction
2. Components
2.1. RNA Aptamers
2.2. Read-Out Mechanisms
2.3. Composition of Functional Switches
3. Theory
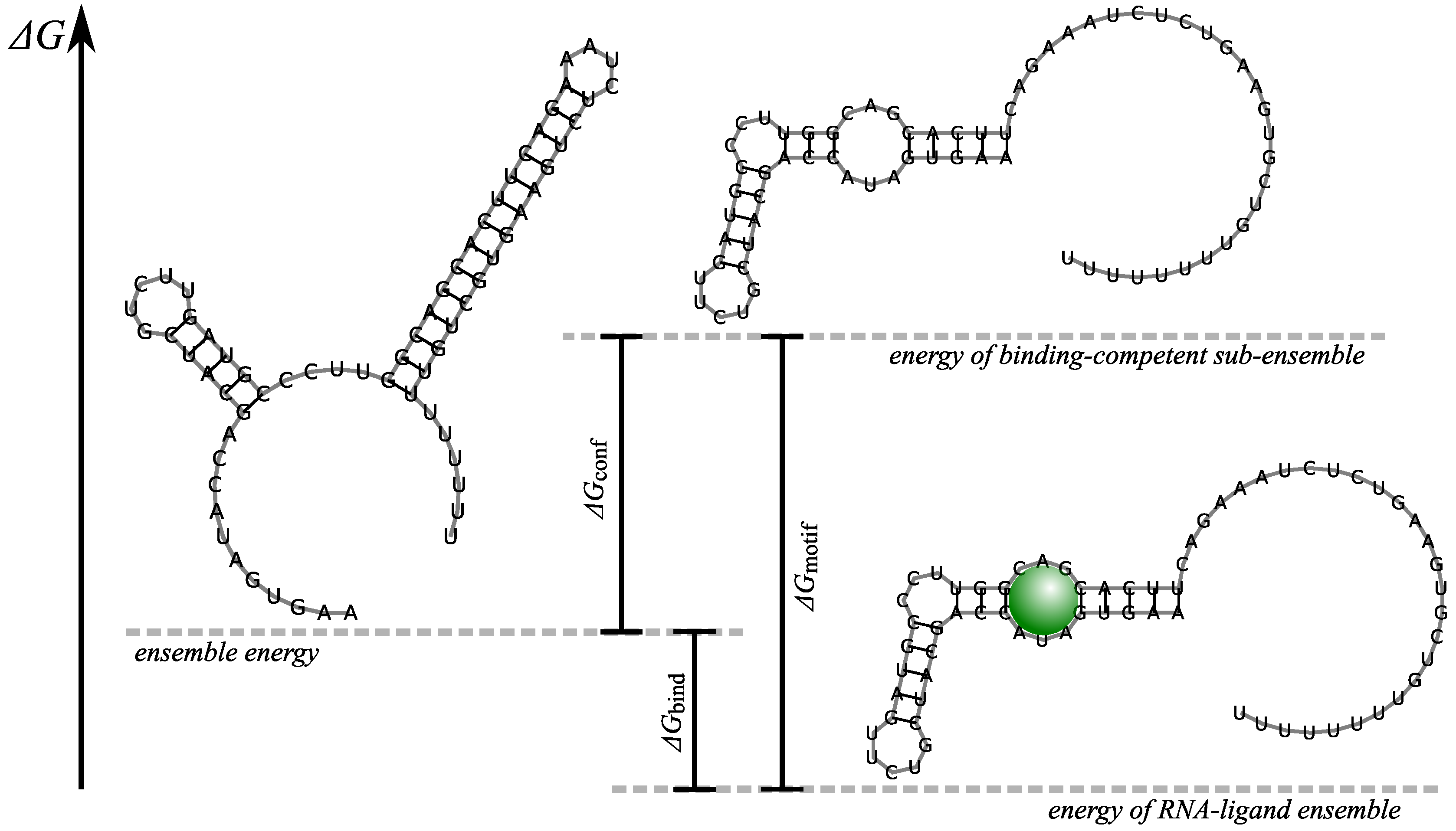
3.1. Thermodynamics of RNA Folding
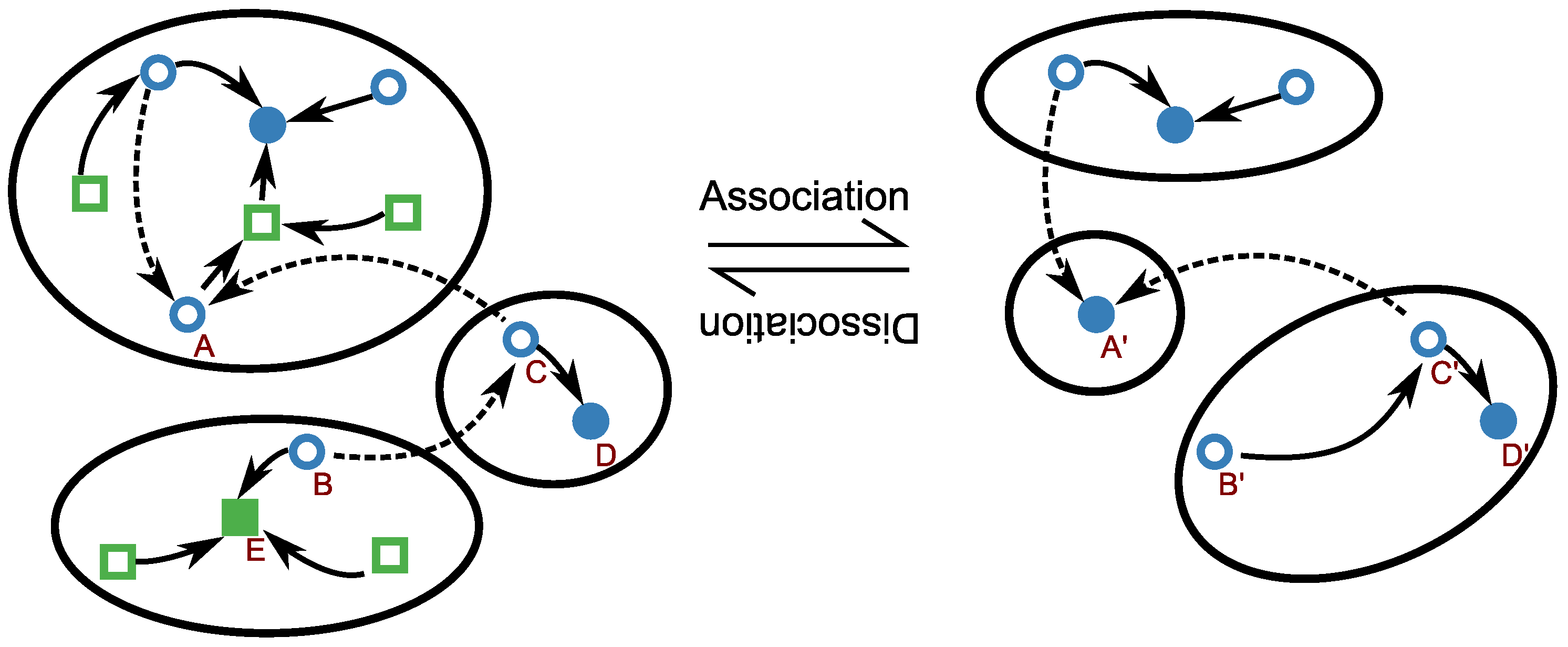
3.2. Kinetic of RNA Folding
3.3. Thermodynamic and Kinetic Design Principles
3.4. Fold Changes for Activators and Repressors
4. Practical Designs
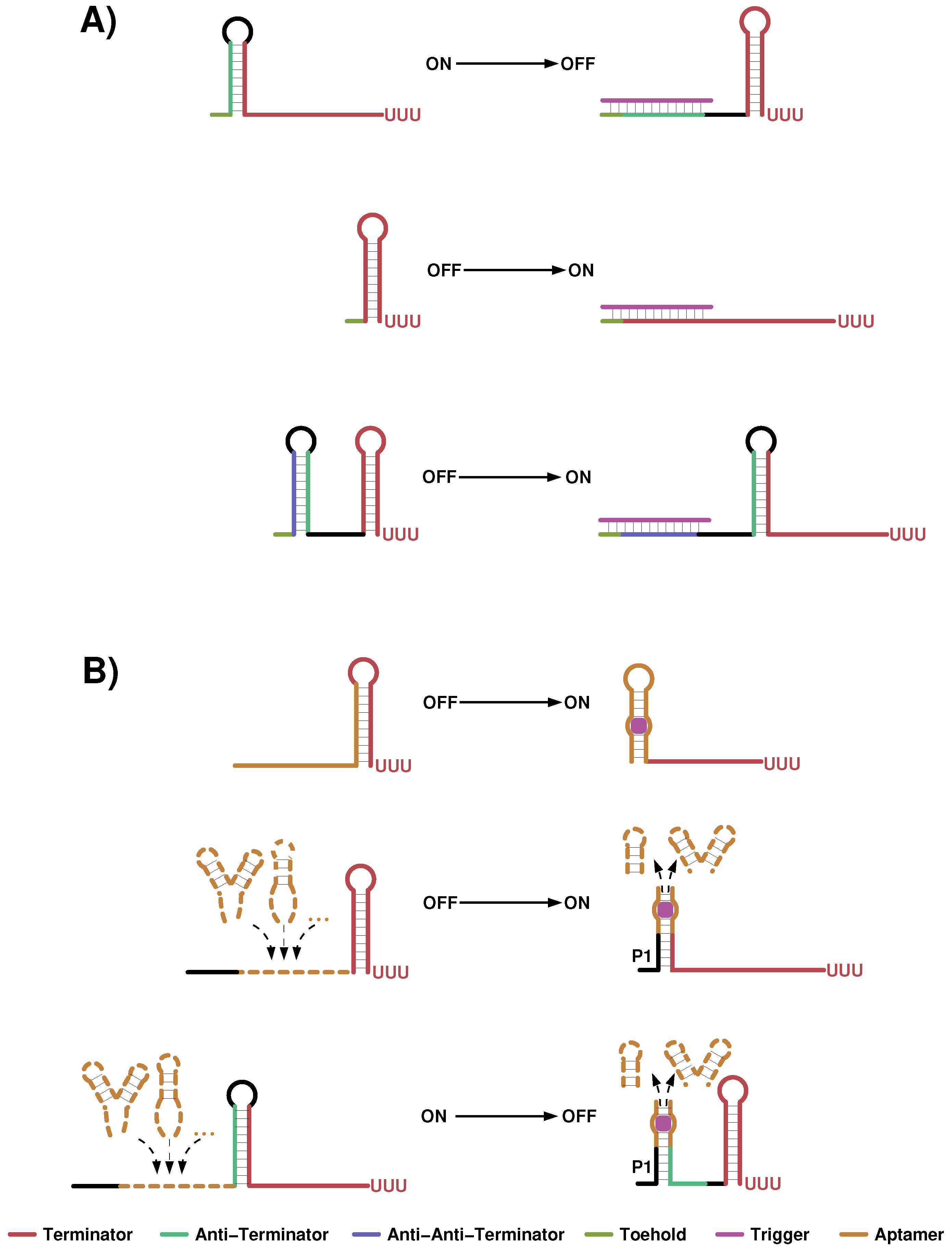
4.1. Small Transcription Regulating RNAs
4.2. Transcription Regulating Riboswitch Design
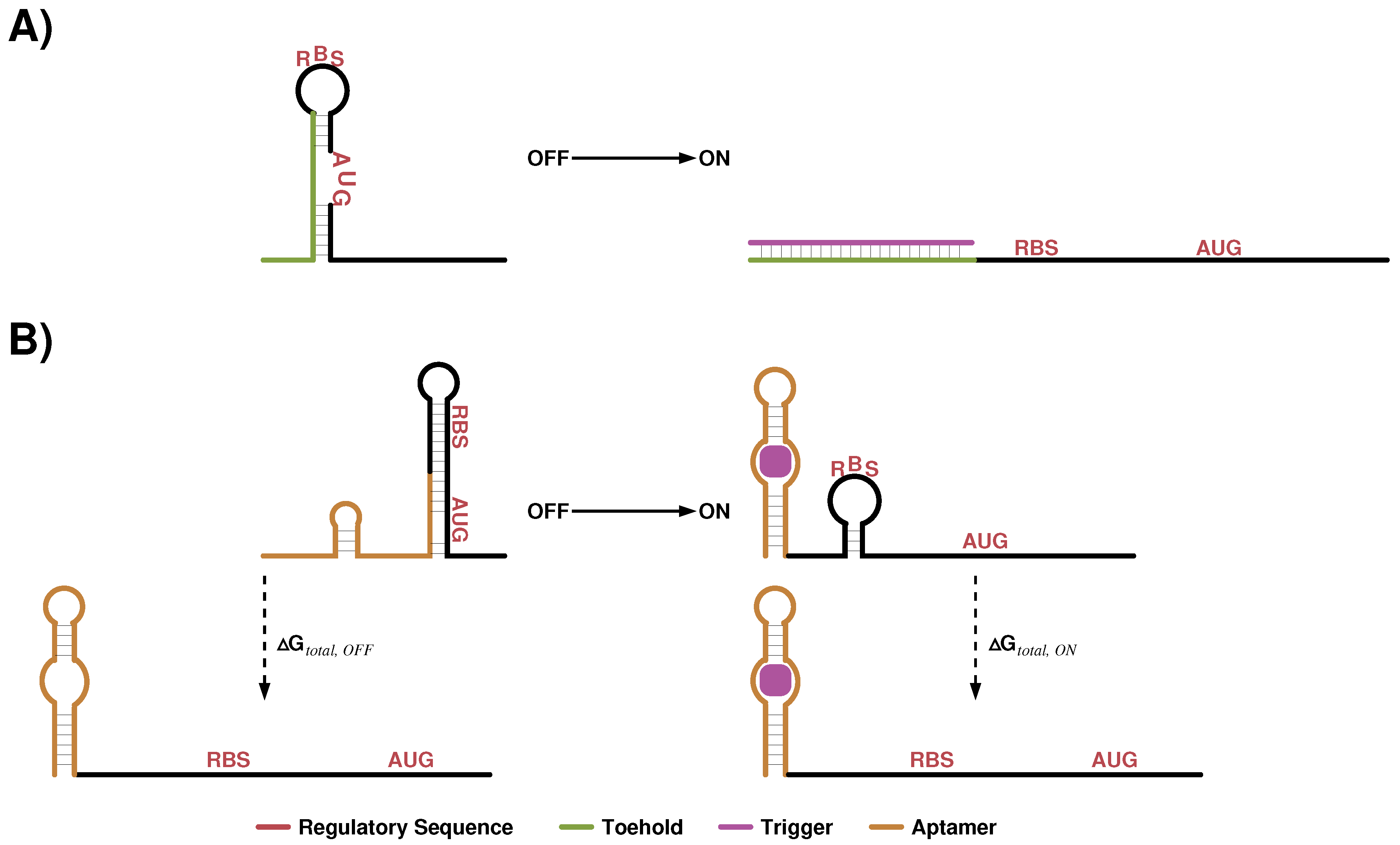
4.3. Small Translation Activation RNAs
4.4. Translation Regulating Riboswitch Design
5. Summary
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| RBS | Ribosome Binding Site |
| mRNA | messenger RNA |
| sRNA | small RNA |
| UTR | untranslated region |
| nt | nucleotide |
| GFP | Green Fluorescent Protein |
| SELEX | Systematic Evolution of Ligands by EXponential enrichment |
| SD | Shine Dalgarno sequence |
| DFHBI | 3,5-difluoro-4-hydroxybenzylidene imidazolinone |
| TO1 | thiazole orange |
| HHR | hammerhead ribozyme |
| TX-TL | transcription-translation |
| ITC | isothermal titration calorimetry |
References
- Cech, T.R.; Steitz, J.A. The Noncoding RNA Revolution—Trashing Old Rules to Forge New Ones. Cell 2014, 157, 77–94. [Google Scholar] [CrossRef] [PubMed]
- Edwards, A.L.; Batey, R.T. Riboswitches: A Common RNA Regulatory Element. Nat. Educ. 2010, 3, 9. [Google Scholar]
- Narberhaus, F. Translational control of bacterial heat shock and virulence genes by temperature-sensing mRNAs. RNA Biol. 2010, 7, 84–89. [Google Scholar] [CrossRef] [PubMed]
- Nechooshtan, G.; Elgrably-Weiss, M.; Sheaffer, A.; Westhof, E.; Altuvia, S. A pH-responsive riboregulator. Genes Dev. 2009, 23, 2650–2662. [Google Scholar] [CrossRef] [PubMed]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Carothers, J.M.; Goler, J.A.; Keasling, J.D. Chemical synthesis using synthetic biology. Curr. Opin. Biotechnol. 2009, 20, 498–503. [Google Scholar] [CrossRef] [PubMed]
- Machtel, P.; Bąkowska-Żywicka, K.; Żywicki, M. Emerging applications of riboswitches—From antibacterial targets to molecular tools. J. Appl. Genet. 2016, 57, 531–541. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.C.; Bloom, R.J.; Smolke, C.D. Engineering biological systems with synthetic RNA molecules. Mol. Cell 2011, 43, 915–926. [Google Scholar] [CrossRef] [PubMed]
- Slomovic, S.; Pardee, K.; Collins, J.J. Synthetic biology devices for in vitro and in vivo diagnostics. Proc. Natl. Acad. Sci. USA 2015, 112, 14429–14435. [Google Scholar] [CrossRef] [PubMed]
- Suess, B.; Fink, B.; Berens, C.; Stentz, R.; Hillen, W. A theophylline responsive riboswitch based on helix slipping controls gene expression in vivo. Nucleic Acids Res. 2004, 32, 1610–1614. [Google Scholar] [CrossRef] [PubMed]
- Berens, C.; Suess, B. Riboswitch engineering—Making the all-important second and third steps. Curr. Opin. Biotechnol. 2015, 31, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Chappell, J.; Watters, K.E.; Takahashi, M.K.; Lucks, J.B. A renaissance in RNA synthetic biology: new mechanisms, applications and tools for the future. Curr. Opin. Chem. Biol. 2015, 28, 47–56. [Google Scholar] [CrossRef] [PubMed]
- McKeague, M.; Wong, R.S.; Smolke, C.D. Opportunities in the design and application of RNA for gene expression control. Nucleic Acids Res. 2016, 44, 2987–2999. [Google Scholar] [CrossRef] [PubMed]
- Etzel, M.; Mörl, M. Synthetic Riboswitches: From Plug and Pray toward Plug and Play. Biochemistry 2017, 56, 1181–1198. [Google Scholar] [CrossRef] [PubMed]
- Nahvi, A.; Sudarsan, N.; Ebert, M.S.; Zou, X.; Brown, K.L.; Breaker, R.R. Genetic control by a metabolite binding mRNA. Chem. Biol. 2002, 9, 1043–1049. [Google Scholar] [CrossRef]
- Mironov, A.S.; Gusarov, I.; Rafikov, R.; Lopez, L.E.; Shatalin, K.; Kreneva, R.A.; Perumov, D.A.; Nudler, E. Sensing small molecules by nascent RNA: A mechanism to control transcription in bacteria. Cell 2002, 111, 747–756. [Google Scholar] [CrossRef]
- Jenison, R.D.; Gill, S.C.; Pardi, A.; Polisky, B. High-resolution molecular discrimination by RNA. Science 1994, 263, 1425–1429. [Google Scholar] [CrossRef] [PubMed]
- Jucker, F.M.; Phillips, R.M.; McCallum, S.A.; Pardi, A. Role of a heterogeneous free state in the formation of a specific RNA-theophylline complex. Biochemistry 2003, 42, 2560–2567. [Google Scholar] [CrossRef] [PubMed]
- Harvey, I.; Garneau, P.; Pelletier, J. Inhibition of translation by RNA-small molecule interactions. RNA 2002, 8, 452–463. [Google Scholar] [CrossRef] [PubMed]
- Desai, S.K.; Gallivan, J.P. Genetic screens and selections for small molecules based on a synthetic riboswitch that activates protein translation. J. Am. Chem. Soc. 2004, 126, 13247–13254. [Google Scholar] [CrossRef] [PubMed]
- Fowler, C.C.; Brown, E.D.; Li, Y. A FACS-based approach to engineering artificial riboswitches. Chembiochem 2008, 9, 1906–1911. [Google Scholar] [CrossRef] [PubMed]
- Wachsmuth, M.; Findeiß, S.; Weissheimer, N.; Stadler, P.F.; Mörl, M. de novo design of a synthetic riboswitch that regulates transcription termination. Nucleic Acids Res. 2013, 41, 2541–2551. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.W.; Zhao, L.; Pardi, A.; Xia, T. Ultrafast Dynamics Show That the Theophylline and 3-Methylxanthine Aptamers Employ a Conformational Capture Mechanism for Binding Their Ligands. Biochemistry 2010, 49, 2943–2951. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, G.R.; Jenison, R.D.; Wick, C.L.; Simorre, J.P.; Pardi, A. Interlocking structural motifs mediate molecular discrimination by a theophylline-binding RNA. Nat. Struct. Biol. 1997, 4, 644–649. [Google Scholar] [CrossRef] [PubMed]
- Wunnicke, D.; Strohbach, D.; Weigand, J.E.; Appel, B.; Feresin, E.; Suess, B.; Müller, S.; Steinhoff, H.J. Ligand-induced conformational capture of a synthetic tetracycline riboswitch revealed by pulse EPR. RNA 2011, 17, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Berens, C.; Thain, A.; Schroeder, R. A tetracycline-binding RNA aptamer. Bioorg. Med. Chem. 2001, 9, 2549–2556. [Google Scholar] [CrossRef]
- Hanson, S.; Bauer, G.; Fink, B.; Suess, B. Molecular analysis of a synthetic tetracycline-binding riboswitch. RNA 2005, 11, 503–511. [Google Scholar] [CrossRef] [PubMed]
- Suess, B.; Hanson, S.; Berens, C.; Fink, B.; Schroeder, R.; Hillen, W. Conditional gene expression by controlling translation with tetracycline-binding aptamers. Nucleic Acids Res. 2003, 31, 1853–1858. [Google Scholar] [CrossRef] [PubMed]
- Weigand, J.E.; Suess, B. Tetracycline aptamer-controlled regulation of pre-mRNA splicing in yeast. Nucleic Acids Res. 2007, 35, 4179–4185. [Google Scholar] [CrossRef] [PubMed]
- Xiao, H.; Edwards, T.E.; Ferré-D’Amaré, A.R. Structural basis for specific, high-affinity tetracycline binding by an in vitro evolved aptamer and artificial riboswitch. Chem. Biol. 2008, 15, 1125–1137. [Google Scholar] [CrossRef] [PubMed]
- Forster, U.; Weigand, J.E.; Trojanowski, P.; Suess, B.; Wachtveitl, J. Conformational dynamics of the tetracycline-binding aptamer. Nucleic Acids Res. 2012, 40, 1807–1817. [Google Scholar] [CrossRef] [PubMed]
- Hanson, S.; Berthelot, K.; Fink, B.; McCarthy, J.E.G.; Suess, B. Tetracycline-aptamer-mediated translational regulation in yeast. Mol. Microbiol. 2003, 49, 1627–1637. [Google Scholar] [CrossRef] [PubMed]
- Müller, M.; Weigand, J.E.; Weichenrieder, O.; Suess, B. Thermodynamic characterization of an engineered tetracycline-binding riboswitch. Nucleic Acids Res. 2006, 34, 2607–2617. [Google Scholar] [CrossRef] [PubMed]
- Wittmann, A.; Suess, B. Engineered riboswitches: Expanding researcher’s toolbox with synthetic RNA regulators. FEBS Lett. 2012, 586, 2076–2083. [Google Scholar] [CrossRef] [PubMed]
- Groher, F.; Suess, B. Synthetic riboswitches—A tool comes of age. Biochim. Biophys. Acta (BBA)—Gene Regul. Mech. 2014, 1839, 964–973. [Google Scholar] [CrossRef] [PubMed]
- Weigand, J.E.; Sanchez, M.; Gunnesch, E.B.; Zeiher, S.; Schroeder, R.; Suess, B. Screening for engineered neomycin riboswitches that control translation initiation. RNA 2008, 14, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Weigand, J.E.; Schmidtke, S.R.; Will, T.J.; Duchardt-Ferner, E.; Hammann, C.; Wöhnert, J.; Suess, B. Mechanistic insights into an engineered riboswitch: A switching element which confers riboswitch activity. Nucleic Acids Res. 2011, 39, 3363–3372. [Google Scholar] [CrossRef] [PubMed]
- Duchardt-Ferner, E.; Gottstein-Schmidtke, S.R.; Weigand, J.E.; Ohlenschlager, O.; Wurm, J.P.; Hammann, C.; Suess, B.; Wöhnert, J. What a Difference an OH Makes: Conformational Dynamics as the Basis for the Ligand Specificity of the Neomycin-Sensing Riboswitch. Angew. Chem. 2016, 55, 1527–1530. [Google Scholar] [CrossRef] [PubMed]
- Dixon, N.; Duncan, J.N.; Geerlings, T.; Dunstan, M.S.; McCarthy, J.E.G.; Leys, D.; Micklefield, J. Reengineering orthogonally selective riboswitches. Proc. Natl. Acad. Sci. USA 2010, 107, 2830–2835. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.C.; Lowe, P.T.; Robinson, C.J.; Vincent, H.A.; Dixon, N.; Leigh, J.; Micklefield, J. Rational Re-engineering of a Transcriptional Silencing PreQ1Riboswitch. J. Am. Chem. Soc. 2015, 137, 9015–9021. [Google Scholar] [CrossRef] [PubMed]
- Soboleski, M.R.; Oaks, J.; Halford, W.P. Green fluorescent protein is a quantitative reporter of gene expression in individual eukaryotic cells. FASEB J. 2005, 19, 440–442. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.H. (Ed.) Experiments in Molecular Genetics; Cold Spring Habor Laboratory Press: Cold Spring Habor, NY, USA, 1972. [Google Scholar]
- Lynch, S.A.; Desai, S.K.; Sajja, H.K.; Gallivan, J.P. A high-throughput screen for synthetic riboswitches reveals mechanistic insights into their function. Chem. Biol. 2007, 14, 173–184. [Google Scholar] [CrossRef] [PubMed]
- Podolsky, T.; Fong, S.T.; Lee, B.T.O. Direct Selection of Tetracycline-Sensitive Escherichia coli Cells Using Nickel Salts. Plasmid 1996, 36, 112–115. [Google Scholar] [CrossRef] [PubMed]
- Stavropoulos, T.A.; Strathdee, C.A. Expression of the tetA(C) tetracycline efflux pump in Escherichia coli confers osmotic sensitivity. FEMS Microbiol. Lett. 2000, 190, 147–150. [Google Scholar] [CrossRef] [PubMed]
- Muranaka, N.; Abe, K.; Yokobayashi, Y. Mechanism-guided library design and dual genetic selection of synthetic OFF riboswitches. Chembiochem 2009, 10, 2375–2381. [Google Scholar] [CrossRef] [PubMed]
- Muranaka, N.; Sharma, V.; Nomura, Y.; Yokobayashi, Y. An efficient platform for genetic selection and screening of gene switches in Escherichia coli. Nucleic Acids Res. 2009, 37, e39. [Google Scholar] [CrossRef] [PubMed]
- Topp, S.; Gallivan, J.P. Guiding bacteria with small molecules and RNA. J. Am. Chem. Soc. 2007, 129, 6807–6811. [Google Scholar] [CrossRef] [PubMed]
- Bren, A.; Eisenbach, M. How signals are heard during bacterial chemotaxis: Protein-protein interactions in sensory signal propagation. J. Bacteriol. 2000, 182, 6865–6873. [Google Scholar] [CrossRef] [PubMed]
- Topp, S.; Gallivan, J.P. Random walks to synthetic riboswitches-a high-throughput selection based on cell motility. Chembiochem 2008, 9, 210–213. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Deutscher, M.P. A uridine-rich sequence required for translation of prokaryotic mRNA. Proc. Natl. Acad. Sci. USA 1992, 89, 2605–2609. [Google Scholar] [CrossRef] [PubMed]
- Paige, J.S.; Wu, K.Y.; Jaffrey, S.R. RNA Mimics of Green Fluorescent Protein. Science 2011, 333, 637–642. [Google Scholar] [CrossRef] [PubMed]
- Litke, J.L.; You, M.; Jaffrey, S.R. Developing Fluorogenic Riboswitches for Imaging Metabolite Concentration Dynamics in Bacterial Cells. Meth. Enzymol. 2016, 572, 315–333. [Google Scholar] [PubMed]
- You, M.; Litke, J.L.; Jaffrey, S.R. Imaging metabolite dynamics in living cells using a Spinach-based riboswitch. Proc. Natl. Acad. Sci. USA 2015, 112, 2756–2765. [Google Scholar] [CrossRef] [PubMed]
- Ketterer, S.; Gladis, L.; Kozica, A.; Meier, M. Engineering and characterization of fluorogenic glycine riboswitches. Nucleic Acids Res. 2016, 44, 5983–5992. [Google Scholar] [CrossRef] [PubMed]
- Kellenberger, C.A.; Wilson, S.C.; Sales-Lee, J.; Hammond, M.C. RNA-Based Fluorescent Biosensors for Live Cell Imaging of Second Messengers Cyclic di-GMP and Cyclic AMP-GMP. J. Am. Chem. Soc. 2013, 135, 4906–4909. [Google Scholar] [CrossRef] [PubMed]
- Paige, J.S.; Nguyen-Duc, T.; Song, W.; Jaffrey, S.R. Fluorescence Imaging of Cellular Metabolites with RNA. Science 2012, 335, 1194. [Google Scholar] [CrossRef] [PubMed]
- Filonov, G.S.; Moon, J.D.; Svensen, N.; Jaffrey, S.R. Broccoli: Rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 2014, 136, 16299–16308. [Google Scholar] [CrossRef] [PubMed]
- Dolgosheina, E.V.; Jeng, S.C.Y.; Panchapakesan, S.S.S.; Cojocaru, R.; Chen, P.S.K.; Wilson, P.D.; Hawkins, N.; Wiggins, P.A.; Unrau, P.J. RNA mango aptamer-fluorophore: A bright, high-affinity complex for RNA labeling and tracking. ACS Chem. Biol. 2014, 9, 2412–2420. [Google Scholar] [CrossRef] [PubMed]
- Sunbul, M.; Jäschke, A. Contact-Mediated Quenching for RNA Imaging in Bacteria with a Fluorophore-Binding Aptamer. Angew. Chem. Int. Ed. 2013, 52, 13401–13404. [Google Scholar] [CrossRef] [PubMed]
- Arora, A.; Sunbul, M.; Jäschke, A. Dual-colour imaging of RNAs using quencher- and fluorophore-binding aptamers. Nucleic Acids Res. 2015, 43. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Fei, J.; Leslie, B.J.; Han, K.Y.; Kuhlman, T.E.; Ha, T. Tandem Spinach Array for mRNA Imaging in Living Bacterial Cells. Sci. Rep. 2015, 5, 17295. [Google Scholar] [CrossRef] [PubMed]
- Kaempfer, R. RNA sensors: Novel regulators of gene expression. EMBO Rep. 2003, 4, 1043–1047. [Google Scholar] [CrossRef] [PubMed]
- Gusarov, I.; Nudler, E. The Mechanism of Intrinsic Transcription Termination. Mol. Cell 1999, 3, 495–504. [Google Scholar] [CrossRef]
- Santangelo, T.J.; Artsimovitch, I. Termination and antitermination: RNA polymerase runs a stop sign. Nat. Rev. Microbiol. 2011, 9, 319–329. [Google Scholar] [CrossRef] [PubMed]
- De Hoon, M.J.L.; Makita, Y.; Nakai, K.; Miyano, S. Prediction of transcriptional terminators in Bacillus subtilis and related species. PLoS Comput. Biol. 2005, 1, e25. [Google Scholar] [CrossRef] [PubMed]
- Martin, F.H.; Tinoco, I. DNA-RNA hybrid duplexes containing oligo(dA:rU) sequences are exceptionally unstable and may facilitate termination of transcription. Nucleic Acids Res. 1980, 8, 2295–2300. [Google Scholar] [CrossRef] [PubMed]
- Wachsmuth, M.; Domin, G.; Lorenz, R.; Serfling, R.; Findeiß, S.; Stadler, P.F.; Mörl, M. Design criteria for synthetic riboswitches acting on transcription. RNA Biol. 2015, 12, 221–231. [Google Scholar] [CrossRef] [PubMed]
- Peters, J.M.; Vangeloff, A.D.; Landick, R. Bacterial Transcription Terminators: The RNA 3’-End Chronicles. J. Mol. Biol. 2011, 412, 793–813. [Google Scholar] [CrossRef] [PubMed]
- Dawid, A.; Cayrol, B.; Isambert, H. RNA synthetic biology inspired from bacteria: Construction of transcription attenuators under antisense regulation. Phys. Biol. 2009, 6, 25007. [Google Scholar] [CrossRef] [PubMed]
- Qi, L.; Lucks, J.B.; Liu, C.C.; Mutalik, V.K.; Arkin, A.P. Engineering naturally occurring trans-acting non-coding RNAs to sense molecular signals. Nucleic Acids Res. 2012, 40, 5775–5786. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, A.; Maeda, M. Aptazyme-based riboswitches as lable-free and detector-free seonsors for cofactors. Bioorg. Med. Chem. Lett. 2007, 17, 3156–3160. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, A.; Maeda, M. An artificial aptazyme-based riboswitch and its cascading system in E. coli. ChemBioChem 2008, 9, 206–209. [Google Scholar] [CrossRef] [PubMed]
- Nomura, Y.; Zhou, L.; Miu, A.; Yokobayashi, Y. Controlling mammalian gene expression by allosteric hepatitis virus ribozymes. ACS Synth. Biol. 2013, 2, 684–689. [Google Scholar] [CrossRef] [PubMed]
- Roth, A.; Weinberg, Z.; Chen, A.G.Y.; Kim, P.B.; Ames, T.D.; Breaker, R.R. A widespread self-cleaving ribozyme class is revealed by bioinformatics. Nat. Chem. Biol. 2014, 10, 56–60. [Google Scholar] [CrossRef] [PubMed]
- Felletti, M.; Wurmthaler, L.A.; Geiger, S.; Hartig, J.S. Twister ribozymes a highly versatile expression platforms for artificial riboswitches. Nat. Commun. 2016, 7, 12834. [Google Scholar] [CrossRef] [PubMed]
- Turner, D.H.; Mathews, D.H. NNDB: The nearest neighbor parameter database for predicting stability of nucleic acid secondary structure. Nucleic Acids Res. 2010, 38, 280–282. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M. On finding all suboptimal foldings of an RNA molecule. Science 1989, 244, 48–52. [Google Scholar] [CrossRef] [PubMed]
- Markham, N.R.; Zuker, M. UNAFold: Software for Nucleic Acid Folding and Hybridization. In Bioinformatics: Structure, Function and Applications; Keith, J.M., Ed.; Humana Press: Totowa, NJ, USA, 2008; pp. 3–31. [Google Scholar]
- Hofacker, I.L.; Fontana, W.; Stadler, P.F.; Bonhoeffer, L.S.; Tacker, M.; Schuster, P. Fast Folding and Comparison of RNA Secondary Structures. Monatsh. Chem. 1994, 125, 167–188. [Google Scholar] [CrossRef]
- Lorenz, R.; Bernhart, S.H.; Höner zu Siederdissen, C.; Tafer, H.; Flamm, C.; Stadler, P.F.; Hofacker, I.L. ViennaRNA Package 2.0. Alg. Mol. Biol. 2011, 6, 26. [Google Scholar] [CrossRef] [PubMed]
- Hajiaghayi, M.; Condon, A.; Hoos, H.H. Analysis of energy-based algorithms for RNA secondary structure prediction. BMC Bioinform. 2012, 13, 22. [Google Scholar] [CrossRef] [PubMed]
- Rivas, E. The four ingredients of single-sequence RNA secondary structure prediction. A unifying perspective. RNA Biol. 2013, 10, 1185–1196. [Google Scholar] [CrossRef] [PubMed]
- Andronescu, M.; Condon, A.; Turner, D.H.; Mathews, D.H. The determination of RNA folding nearest neighbor parameters. Methods Mol. Biol. 2014, 1097, 45–70. [Google Scholar] [PubMed]
- McCaskill, J.S. The Equilibrium Partition Function and Base Pair Binding Probabilities for RNA Secondary Structure. Biopolymers 1990, 29, 1105–1119. [Google Scholar] [CrossRef] [PubMed]
- Dimitrov, R.A.; Zuker, M. Prediction of hybridization and melting for double-stranded nucleic acids. Biophys. J. 2004, 87, 215–226. [Google Scholar] [CrossRef] [PubMed]
- Bernhart, S.H.; Tafer, H.; Mückstein, U.; Flamm, C.; Stadler, P.F.; Hofacker, I.L. Partition Function and Base Pairing Probabilities of RNA Heterodimers. Algorithms Mol. Biol. 2006, 1, 3. [Google Scholar] [CrossRef] [PubMed]
- Dirks, R.M.; Bois, J.S.; Schaeffer, J.M.; Winfree, E.; Pierce, N.A. Thermodynamic Analysis of Interacting Nucleic Acid Strands. SIAM Rev. 2007, 49, 65–88. [Google Scholar] [CrossRef]
- Mückstein, U.; Tafer, H.; Hackermüller, J.; Bernhard, S.B.; Stadler, P.F.; Hofacker, I.L. Thermodynamics of RNA-RNA Binding. Bioinformatics 2006, 22, 1177–1182. [Google Scholar] [CrossRef] [PubMed]
- Busch, A.; Richter, A.S.; Backofen, R. IntaRNA: Efficient prediction of bacterial sRNA targets incorporating target site accessibility and seed regions. Bioinformatics 2008, 24, 2849–2856. [Google Scholar] [CrossRef] [PubMed]
- Chitsaz, H.; Salari, R.; Sahinalp, S.C.; Backofen, R. A partition function algorithm for interacting nucleic acid strands. Bioinformatics 2009, 25, i365–i373. [Google Scholar] [CrossRef] [PubMed]
- Huang, F.W.D.; Qin, J.; Reidys, C.M.; Stadler, P.F. Partition Function and Base Pairing Probabilities for RNA-RNA Interaction Prediction. Bioinformatics 2009, 25, 2646–2654. [Google Scholar] [CrossRef] [PubMed]
- Xia, T.; SantaLucia, J., Jr.; Burkard, M.E.; Kierzek, R.; Schroeder, S.J.; Jiao, X.; Cox, C.; Turner, D.H. Thermodynamic parameters for an expanded nearest-neighbor model for formation of RNA duplexes with Watson-Crick base pairs. Biochemistry 1998, 37, 14719–14735. [Google Scholar] [CrossRef] [PubMed]
- Mückstein, U.; Tafer, H.; Bernhard, S.H.; Hernandez-Rosales, M.; Vogel, J.; Stadler, P.F.; Hofacker, I.L. Translational Control by RNA-RNA Interaction: Improved Computation of RNA-RNA Binding Thermodynamics. In BioInformatics Research and Development—BIRD 2008; Elloumi, M., Küng, J., Linial, M., Murphy, R.F., Schneider, K., Toma, C.T., Eds.; Springer: Berlin, Germany, 2008; Volume 13, pp. 114–127. [Google Scholar]
- Lorenz, R.; Hofacker, I.L.; Stadler, P.F. RNA Folding with Hard and Soft Constraints. Algorithms Mol. Biol. 2016, 11, 8. [Google Scholar] [CrossRef] [PubMed]
- Hackermüller, J.; Meisner, N.C.; Auer, M.; Jaritz, M.; Stadler, P.F. The Effect of RNA Secondary Structures on RNA-Ligand Binding and the Modifier RNA Mechanism: A Quantitative Model. Gene 2005, 345, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Flamm, C.; Fontana, W.; Hofacker, I.; Schuster, P. RNA folding kinetics at elementary step resolution. RNA 2000, 6, 325–338. [Google Scholar] [CrossRef] [PubMed]
- Gultyaev, A.P. The computer simulation of RNA folding involving pseudoknot formation. Nucleic Acids Res. 1991, 19, 2489–2494. [Google Scholar] [CrossRef] [PubMed]
- Fernández, A.; Shakhnovich, E.I. Activation-energy landscape for metastable RNA folding. Phys. Rev. A 1990, 42, 3657–3659. [Google Scholar] [CrossRef] [PubMed]
- Mironov, A.A.; Lebedev, V.F. A kinetic model of RNA folding. Biosystems 1993, 30, 49–56. [Google Scholar] [CrossRef]
- Tacker, M.; Fontana, W.; Stadler, P.F.; Schuster, P. Statistics of RNA Melting Kinetics. Eur. Biophys. J. 1994, 23, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Zhao, P.; Zhang, W.; Chen, S.J. Cotranscriptional folding kinetics of ribonucleic acid secondary structures. J. Chem. Phys. 2011, 135. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Voß, B. Analysing RNA-kinetics based on folding space abstraction. BMC Bioinform. 2014, 15, 60. [Google Scholar] [CrossRef] [PubMed]
- Wolfinger, M.T.; Svrcek-Seiler, W.A.; Flamm, C.; Hofacker, I.L.; Stadler, P.F. Exact Folding Dynamics of RNA Secondary Structures. J. Phys. A Math. Gen. 2004, 37, 4731–4741. [Google Scholar] [CrossRef]
- Wuchty, S.; Fontana, W.; Hofacker, I.L.; Schuster, P. Complete Suboptimal Folding of RNA and the Stability of Secondary Structures. Biopolymers 1999, 49, 145–165. [Google Scholar] [CrossRef]
- Mann, M.; Kucharík, M.; Flamm, C.; Wolfinger, M.T. Memory-efficient RNA energy landscape exploration. Bioinformatics 2014, 30, 2584–2591. [Google Scholar] [CrossRef] [PubMed]
- Sibani, P.; van der Pas, R.; Schön, J.C. The lid method for exhaustive exploration of metastable states of complex systems. Comput. Phys. Commun. 1999, 116, 17–27. [Google Scholar] [CrossRef]
- Klotz, T.; Kobe, S. “Valley Structures” in the phase space of a finite 3D Ising spin glass with ±I interactions. J. Phys. A Math. Gen. 1994, 27, L95–L100. [Google Scholar] [CrossRef]
- Doye, J.P.; Miller, M.A.; Welsh, D.J. Evolution of the potential energy surface with size for Lennard-Jones clusters. J. Chem. Phys. 1999, 111, 8417–8429. [Google Scholar] [CrossRef]
- Garstecki, P.; Hoang, T.; Cieplak, M. Energy Landscapes, supergraphs, and “folding funnels” in spin systems. Phys. Rev. E 1999, 60, 3219–3226. [Google Scholar] [CrossRef]
- Flamm, C.; Hofacker, I.L.; Stadler, P.F.; Wolfinger, M.T. Barrier Trees of Degenerate Landscapes. Z. Phys. Chem. 2002, 216, 155–173. [Google Scholar] [CrossRef]
- Kucharík, M.; Hofacker, I.L.; Stadler, P.F.; Qin, J. Basin Hopping Graph: A computational framework to characterize RNA folding landscapes. Bioinformatics 2014, 30, 2009–2017. [Google Scholar] [CrossRef] [PubMed]
- Kühnl, F.; Stadler, P.F.; Will, S. Tractable Kinetics of RNA—Ligand Interaction. In Bioinformatics Research and Applications: 12th International Symposium, ISBRA 2016; Bourgeois, A., Skums, P., Wan, X., Zelikovsky, A., Eds.; Springer: Berlin, Germany, 2016; Volume 9683, pp. 337–338. [Google Scholar]
- Kühnl, F.; Stadler, P.F.; Will, S. Tractable Analysis of RNA—Ligand Interaction Kinetics. BMC Bioinform. 2017, in press. [Google Scholar]
- Hofacker, I.L.; Flamm, C.; Heine, C.; Wolfinger, M.T.; Scheuermann, G.; Stadler, P.F. BarMap: RNA Folding on Dynamics Energy Landscapes. RNA 2010, 16, 1308–1316. [Google Scholar] [CrossRef] [PubMed]
- Taneda, A. Multi-objective genetic algorithm for pseudoknotted RNA sequence design. Front. Genet. 2012, 3, 36. [Google Scholar] [CrossRef] [PubMed]
- Hammer, S.; Tschiatschek, B.; Flamm, C.; Hofacker, I.L.; Findeiß, S. RNAblueprint: Flexible multiple target nucleic acid sequence design. Bioinformatics 2017, in press. [Google Scholar] [CrossRef] [PubMed]
- Flamm, C.; Hofacker, I.L.; Maurer-Stroh, S.; Stadler, P.F.; Zehl, M. Design of multistable RNA molecules. RNA 2001, 7, 254–265. [Google Scholar] [CrossRef] [PubMed]
- Green, A.A.; Silver, P.A.; Collins, J.J.; Yin, P. Toehold Switches: De-Novo-Designed Regulators of Gene Expression. Cell 2014, 159, 925–939. [Google Scholar] [CrossRef] [PubMed]
- Espah-Borujeni, A.; Mishler, D.M.; Wang, J.; Huso, W.; Salis, H.M. Automated physics-based design of synthetic riboswitches from diverse RNA aptamers. Nucleic Acids Res. 2016, 44, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Chappell, J.; Takahashi, M.K.; Lucks, J.B. Creating small transcription activating RNAs. Nat. Chem. Biol. 2015, 11, 214–220. [Google Scholar] [CrossRef] [PubMed]
- Domin, G.; Findeiß, S.; Wachsmuth, M.; Will, S.; Stadler, P.F.; Mörl, M. Applicability of a computational design approach for synthetic riboswitches. Nucleic Acids Res. 2017, 45, 4108–4119. [Google Scholar] [CrossRef] [PubMed]
- Ceres, P.; Trausch, J.J.; Batey, R.T. Engineering modular ‘ON’ RNA switches using biological components. Nucleic Acids Res. 2013, 41, 10449–10461. [Google Scholar] [CrossRef] [PubMed]
- Ceres, P.; Garst, A.D.; Marcano-Velázquez, J.G.; Batey, R.T. Modularity of Select Riboswitch Expression Platforms Enables Facile Engineering of Novel Genetic Regulatory Devices. ACS Synth. Biol. 2013, 2, 463–472. [Google Scholar] [CrossRef] [PubMed]
- Meyer, S.; Chappell, J.; Sankar, S.; Chew, R.; Lucks, J.B. Improving fold activation of small transcription activating RNAs (STARs) with rational RNA engineering strategies. Biotechnol. Bioeng. 2016, 113, 216–225. [Google Scholar] [CrossRef] [PubMed]
- Xayaphoummine, A.; Viasnoff, V.; Harlepp, S.; Isambert, H. Encoding folding paths of RNA switches. Nucleic Acids Res. 2006, 35, 614–622. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.J.; Liu, P.; Nielsen, A.A.K.; Brophy, J.A.N.; Clancy, K.; Peterson, T.; Voigt, C.A. Characterization of 582 natural and synthetic terminators and quantification of their design constraints. Nat. Methods 2013, 10, 659–664. [Google Scholar] [CrossRef] [PubMed]
- Salis, H.M.; Mirsky, E.A.; Voigt, C.A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 2009, 27, 946–950. [Google Scholar] [CrossRef] [PubMed]
- Salis, H.M. The Ribosome Binding Site Calculator. In Methods in Enzymology; Elsevier BV: Amsterdam, The Netherlands, 2011; pp. 19–42. [Google Scholar]
- Borujeni, A.E.; Channarasappa, A.S.; Salis, H.M. Translation rate is controlled by coupled trade-offs between site accessibility, selective RNA unfolding and sliding at upstream standby sites. Nucleic Acids Res. 2013, 42, 2646–2659. [Google Scholar] [CrossRef] [PubMed]
- Mishler, D.M.; Gallivan, J.P. A family of synthetic riboswitches adopts a kinetic trapping mechanism. Nucleic Acids Res. 2014, 42, 6753–6761. [Google Scholar] [CrossRef] [PubMed]


© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Findeiß, S.; Etzel, M.; Will, S.; Mörl, M.; Stadler, P.F. Design of Artificial Riboswitches as Biosensors. Sensors 2017, 17, 1990. https://doi.org/10.3390/s17091990
Findeiß S, Etzel M, Will S, Mörl M, Stadler PF. Design of Artificial Riboswitches as Biosensors. Sensors. 2017; 17(9):1990. https://doi.org/10.3390/s17091990
Chicago/Turabian StyleFindeiß, Sven, Maja Etzel, Sebastian Will, Mario Mörl, and Peter F. Stadler. 2017. "Design of Artificial Riboswitches as Biosensors" Sensors 17, no. 9: 1990. https://doi.org/10.3390/s17091990
APA StyleFindeiß, S., Etzel, M., Will, S., Mörl, M., & Stadler, P. F. (2017). Design of Artificial Riboswitches as Biosensors. Sensors, 17(9), 1990. https://doi.org/10.3390/s17091990






