A Sensitive and Label-Free Pb(II) Fluorescence Sensor Based on a DNAzyme Controlled G-Quadruplex/Thioflavin T Conformation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
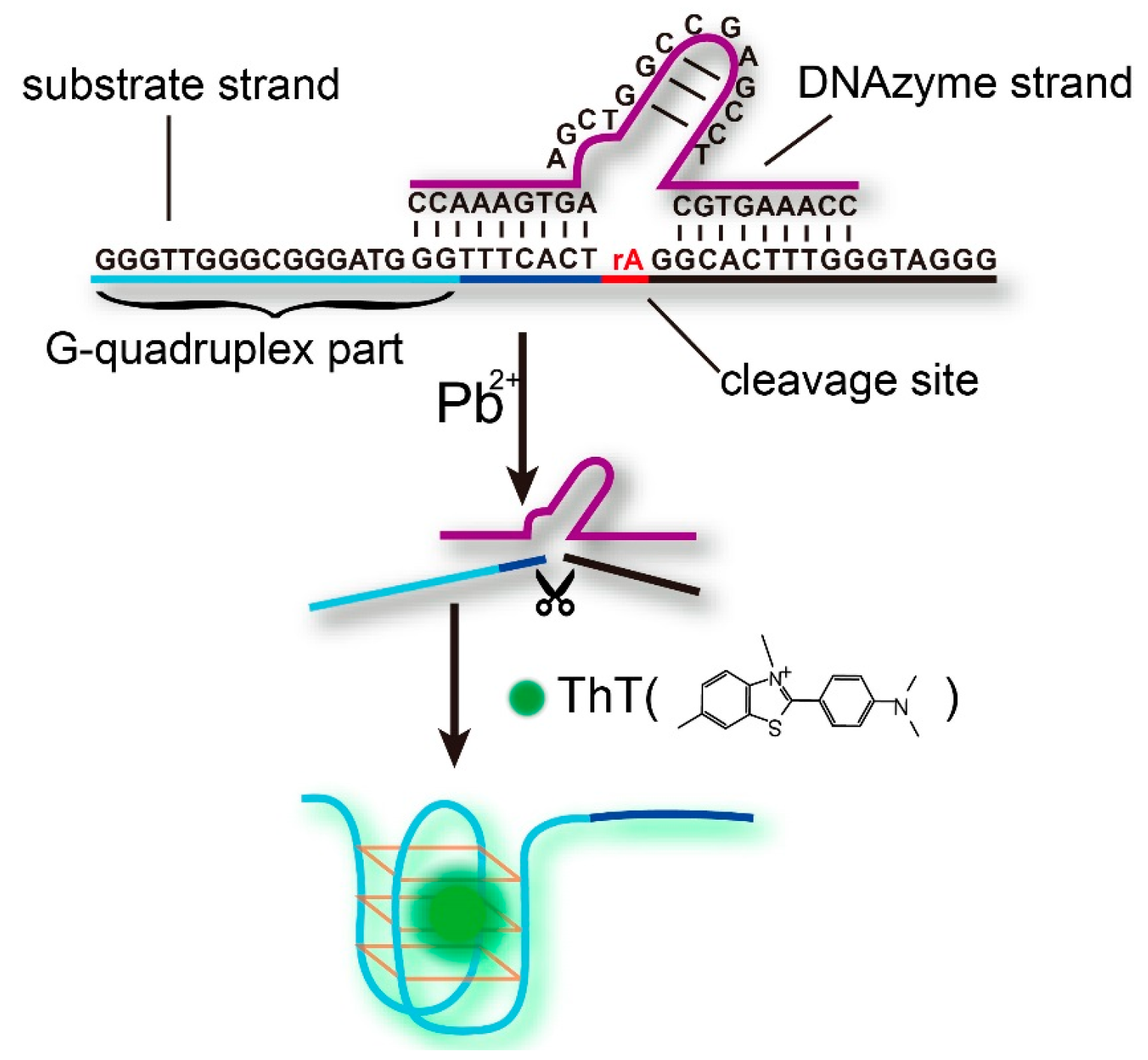
- Pb-DNAzyme:5′-CCAAAGTGCTCCGAGCCGGTCGAAGTGAAACC-3′
- Pb-Sub:5′-GGGTTGGGCGGGATGGGTTTCACTrAGGCACTTTGGGTAGGG-3′, (rA represents an adenosine ribonucleotide).
2.2. Apparatus
2.3. DNAzyme-Based Fluorescence Sensor for Pb2+
3. Results
3.1. The Principle of Pb2+ Detection
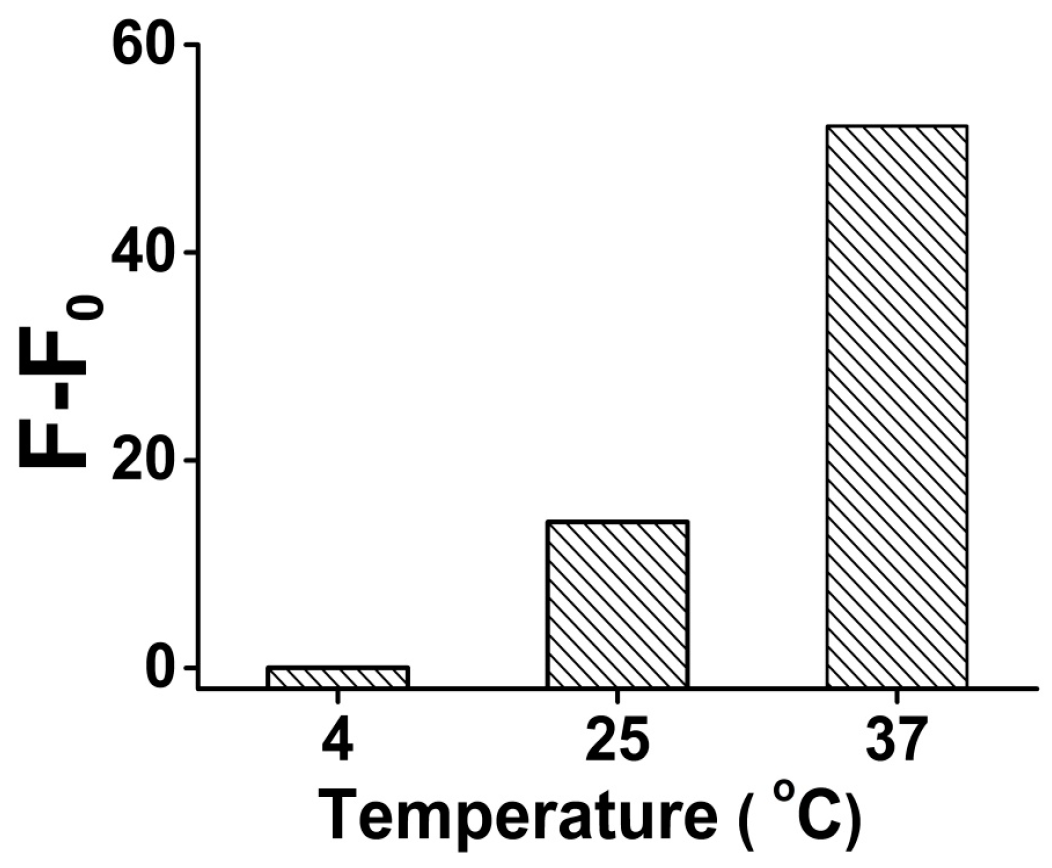
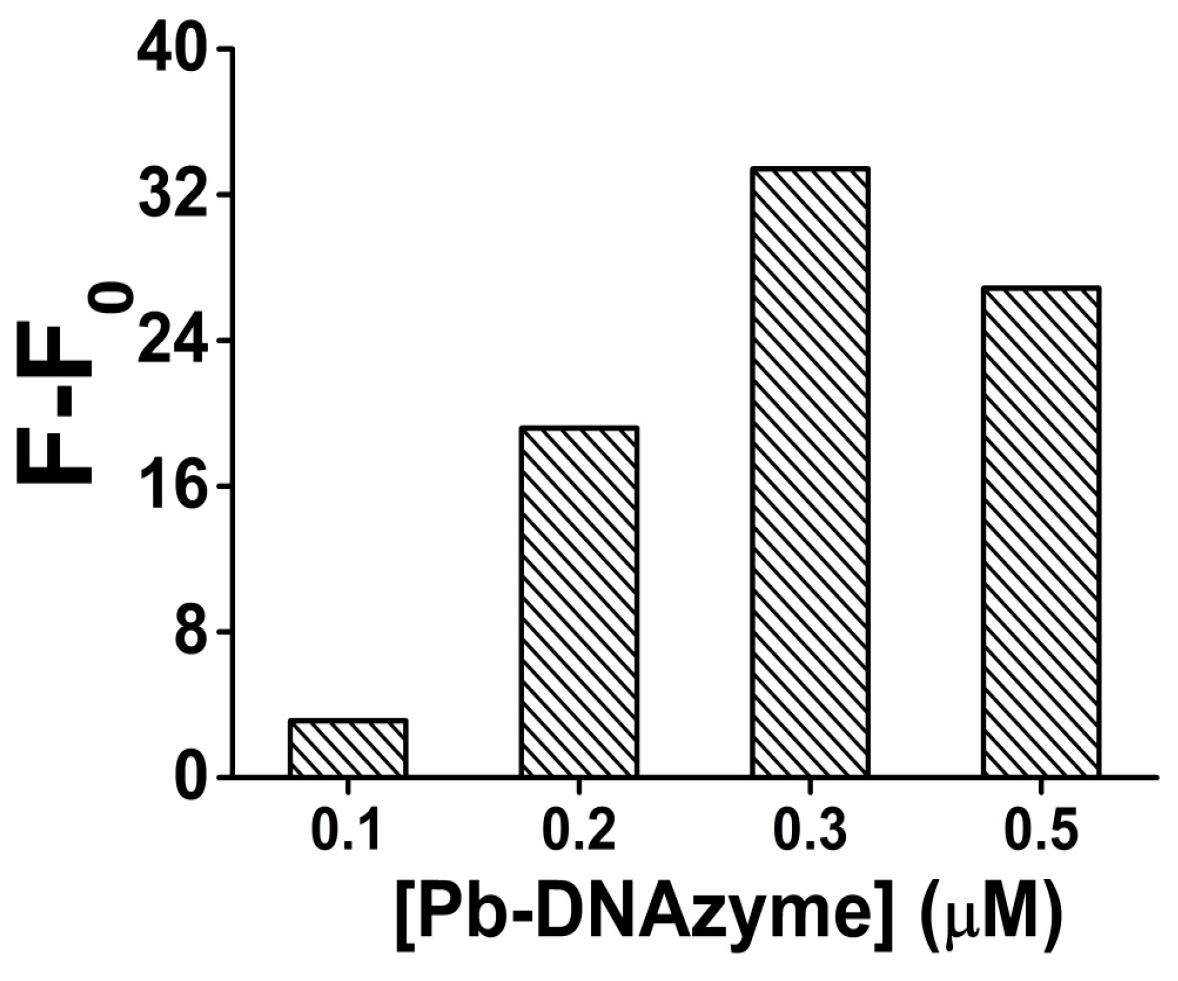
3.2. Optimization of the Detection Conditions
3.3. Quantification of Pb2+
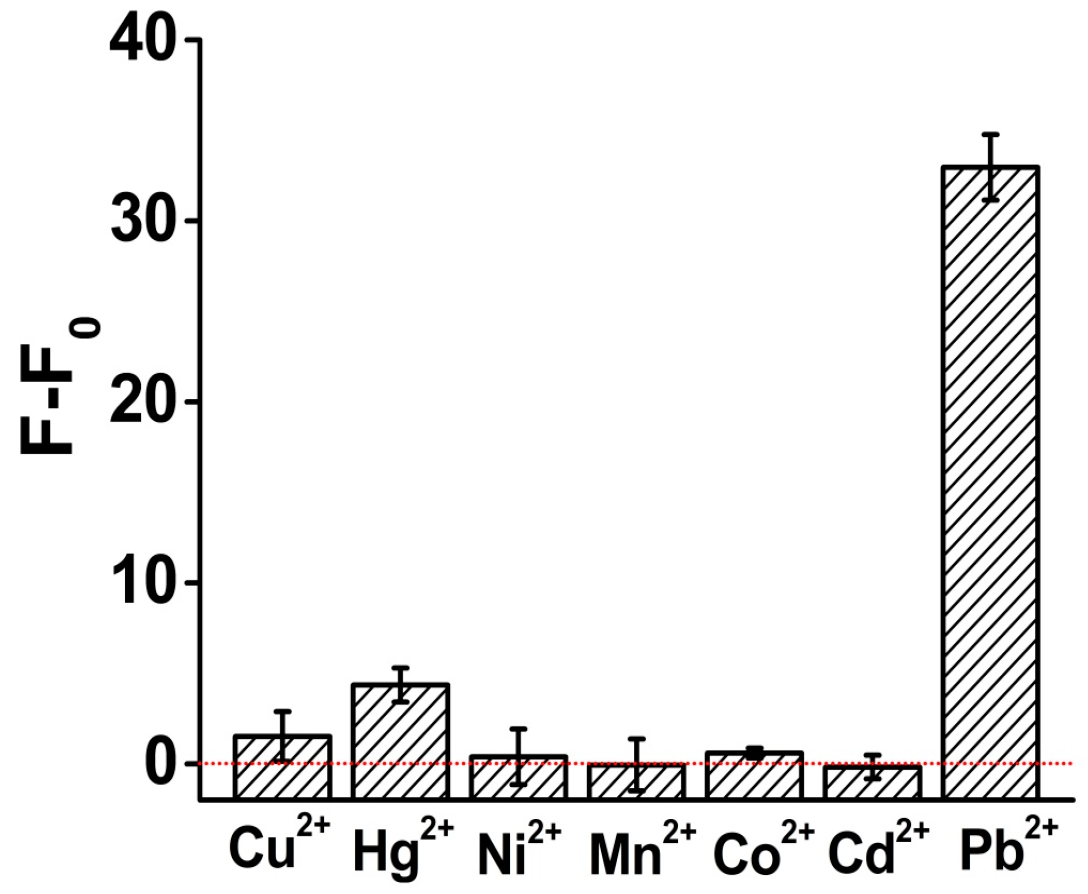
3.4. Specificity of Pb2+ Analysis
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Raj, S.; Shankaran, D.R. Curcumin based biocompatible nanofibers for lead ion detection. Sens. Actuators B Chem. 2016, 226, 318–325. [Google Scholar] [CrossRef]
- Shang, Y.; Zhang, Y.; Li, P.; Lai, J.; Kong, X.-Y.; Liu, W.; Xiao, K.; Xie, G.; Tian, Y.; Wen, L. DNAzyme tunable lead (II) gating based on ion-track etched conical nanochannels. Chem. Commun. 2015, 51, 5979–5981. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Lu, Y. A colorimetric lead biosensor using DNAzyme-directed assembly of gold nanoparticles. J. Am. Chem. Soc. 2003, 125, 6642–6643. [Google Scholar] [CrossRef] [PubMed]
- Kwon, J.Y.; Jang, Y.J.; Lee, Y.J.; Kim, K.M.; Seo, M.S.; Nam, W.; Yoon, J. A highly selective fluorescent chemosensor for Pb2+. J. Am. Chem. Soc. 2005, 127, 10107–10111. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.Q.; Peng, C.; Li, D.; Zhuo, L.; He, S.J.; Wang, L.H.; Huang, Q.; Xu, Q.H.; Fan, C.H. Metal ion-modulated graphene-DNAzyme interactions: Design of a nanoprobe for fluorescent detection of lead(II) ions with high sensitivity, selectivity and tunable dynamic range. Chem. Commun. 2011, 47, 6278–6280. [Google Scholar] [CrossRef] [PubMed]
- Behbahani, M.; Hassanlou, P.G.; Amini, M.M.; Omidi, F.; Esrafili, A.; Farzadkia, M.; Bagheri, A. Application of solvent-assisted dispersive solid phase extraction as a new, fast, simple and reliable preconcentration and trace detection of lead and cadmium ions in fruit and water samples. Food Chem. 2015, 187, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, V.M.P.; Barbosa, A.F.; Bettini, J.; Luccas, P.O.; Figueiredo, E.C. Direct extraction of lead (II) from untreated human blood serum using restricted access carbon nanotubes and its determination by atomic absorption spectrometry. Talanta 2016, 147, 478–484. [Google Scholar] [CrossRef] [PubMed]
- Ma, R.; Van Mol, W.; Adams, F. Determination of cadmium, copper and lead in environmental samples. An evaluation of flow injection on-line sorbent extraction for flame atomic absorption spectrometry. Anal. Chim. Acta 1994, 285, 33–43. [Google Scholar] [CrossRef]
- Jackson, S.E.; Pearson, N.J.; Griffin, W.L.; Belousova, E.A. The application of laser ablation-inductively coupled plasma-mass spectrometry to in situ U-Pb zircon geochronology. Chem. Geol. 2004, 211, 47–69. [Google Scholar] [CrossRef]
- Liang, Q.; Jing, H.; Gregoire, D.C. Determination of trace elements in granites by inductively coupled plasma mass spectrometry. Talanta 2000, 51, 507–513. [Google Scholar] [CrossRef]
- Elfering, H.; Andersson, J.T.; Poll, K.G. Determination of organic lead in soils and waters by hydride generation inductively coupled plasma atomic emission spectrometry. Analyst 1998, 123, 669–674. [Google Scholar] [CrossRef]
- Ochsenkühn-Petropoulou, M.; Ochsenkühn, K.M. Comparison of inductively coupled plasma-atomic emission spectrometry, anodic stripping voltammetry and instrumental neutron-activation analysis for the determination of heavy metals in airborne particulate matter. Fresenius’ J. Anal. Chem. 2001, 369, 629–632. [Google Scholar] [CrossRef]
- Liu, J.; Cao, Z.; Lu, Y. Functional nucleic acid sensors. Chem. Rev. 2009, 109, 1948–1998. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Lu, Y. A DNAzyme catalytic beacon sensor for paramagnetic Cu2+ ions in aqueous solution with high sensitivity and selectivity. J. Am. Chem. Soc. 2007, 129, 9838–9839. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Brown, A.K.; Meng, X.; Cropek, D.M.; Istok, J.D.; Watson, D.B.; Lu, Y. A catalytic beacon sensor for uranium with parts-per-trillion sensitivity and millionfold selectivity. Proc. Natl. Acad. Sci. USA 2007, 104, 2056–2061. [Google Scholar] [CrossRef] [PubMed]
- Hollenstein, M.; Hipolito, C.; Lam, C.; Dietrich, D.; Perrin, D.M. A highly selective DNAzyme sensor for mercuric ions. Angew. Chem. Int. Ed. 2008, 47, 4346–4350. [Google Scholar] [CrossRef] [PubMed]
- Lan, T.; Furuya, K.; Lu, Y. A highly selective lead sensor based on a classic lead DNAzyme. Chem. Commun. 2010, 46, 3896–3898. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.Y.; Gu, W.; Zhang, C.L.; Shi, X.H.; Xian, Y.Z. In situ regulation nanoarchitecture of Au nanoparticles/reduced graphene oxide colloid for sensitive and selective SERS detection of lead ions. J. Colloid Interface Sci. 2016, 465, 279–285. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-B.; Wang, Z.; Xing, H.; Xiang, Y.; Lu, Y. Catalytic and Molecular Beacons for Amplified Detection of Metal Ions and Organic Molecules with High Sensitivity. Anal. Chem. 2010, 82, 5005–5011. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Han, S.H.; Chung, B.H. Improving Pb2+ detection using DNAzyme-based fluorescence sensors by pairing fluorescence donors with gold nanoparticles. Biosens. Bioelectron. 2011, 26, 2125–2129. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Rowe, A.A.; Plaxco, K.W. Electrochemical detection of parts-per-billion lead via an electrode-bound DNAzyme assembly. J. Am. Chem. Soc. 2007, 129, 262–263. [Google Scholar] [CrossRef] [PubMed]
- Dalavoy, T.S.; Wernette, D.P.; Gong, M.; Sweedler, J.V.; Lu, Y.; Flachsbart, B.R.; Shannon, M.A.; Bohn, P.W.; Cropek, D.M. Immobilization of DNAzyme catalytic beacons on PMMA for Pb 2+ detection. Lab Chip 2008, 8, 786–793. [Google Scholar] [CrossRef] [PubMed]
- Chiuman, W.; Li, Y. Efficient signaling platforms built from a small catalytic DNA and doubly labeled fluorogenic substrates. Nucleic Acids Res. 2007, 35, 401–405. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Li, B.; Wang, E. “Fitting” Makes “Sensing” Simple: Label-Free Detection Strategies Based on Nucleic Acid Aptamers. Accounts Chem. Res. 2013, 46, 203–213. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Yan, J.; Fang, Z.; Zhang, C.; Bai, W.; Yan, M.; Zhu, C.; Gao, C.; Chen, A. Highly sensitive and selective optical detection of lead(II) using a label-free fluorescent aptasensor. RSC Adv. 2016, 6, 90300–90304. [Google Scholar] [CrossRef]
- Li, T.; Wang, E.; Dong, S. Lead(II)-induced allosteric G-quadruplex DNAzyme as a colorimetric and chemiluminescence sensor for highly sensitive and selective Pb2+ detection. Anal. Chem. 2010, 82, 1515–1520. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Liu, K.; Lin, Y.; Chang, H. Fluorescence detection of Lead(II) ions through their induced catalytic activity of DNAzymes. Anal. Chem. 2011, 83, 225–230. [Google Scholar] [CrossRef] [PubMed]
- Fu, T.; Ren, S.L.; Gong, L.; Meng, H.M.; Cui, L.; Kong, R.M.; Zhang, X.B.; Tan, W.H. A label-free DNAzyme fluorescence biosensor for amplified detection of Pb2+-based on cleavage-induced G-quadruplex formation. Talanta 2016, 147, 302–306. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Lin, J.; Zhang, X.; Cai, S.; Wu, D.; Li, C.; Yang, S.; Zhang, J. Label-free fluorescent biosensor based on the target recycling and Thioflavin T-induced quadruplex formation for short DNA species of c-erbB-2 detection. Anal. Chim. Acta 2014, 817, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Mohanty, J.; Barooah, N.; Dhamodharan, V.; Harikrishna, S.; Pradeepkumar, P.I.; Bhasikuttan, A.C. Thioflavin T as an Efficient Inducer and Selective Fluorescent Sensor for the Human Telomeric G-Quadruplex DNA. J. Am. Chem. Soc. 2013, 135, 367–376. [Google Scholar] [CrossRef] [PubMed]
- Gabelica, V.R.; Maeda, R.; Fujimoto, T.; Yaku, H.; Murashima, T.; Sugimoto, N.; Miyoshi, D. Multiple and cooperative binding of fluorescence light-up probe thioflavin t with human telomere DNA G-Quadruplex. Biochemistry 2013, 52, 5620–5628. [Google Scholar] [CrossRef] [PubMed]
- Faverie, A.R.; Guédin, A.; Bedrat, A.; Yatsunyk, L.A.; Mergny, J.-L. Thioflavin T as a fluorescence light-up probe for G4 formation. Nucleic Acids Res. 2014. [Google Scholar] [CrossRef]


| DNA Probe | Linear Range | Detection Limit | Detection Methods | Detection Time | Reaction System | Reference |
|---|---|---|---|---|---|---|
| G-quadruplex | 1 ng/mL –1 mg/mL, R2 = 0.992 | 1 ng/mL (4.83 nM) | Fluorescence | 0.5 h | DsDNA’s conformational changes/PicoGreen | [25] |
| Allosteric G-quadruplex DNAzyme | 1 nM –316 nM, R2 = 0.980 | 1 nM | Chemiluminescence | 2 h | luminol-H2O2 | [26] |
| DNAzymes/AUR | 0 nM–1000 nM, R2 = 0.95 | 0.4 nM | Fluorescence | 3.5 h | G4/hemin/H2O2/AUR catalytic system | [27] |
| G-quadruplex DNAzyme | 5 nM–100 nM, R2 = 0.996 | 3 nM | Fluorescence | 3.5 h | catalytic beacons/G4/ZnPPIX | [28] |
| G-quadruplex DNAzyme | 10 nM–1000 nM, R2 = 0.997 | 0.06 nM | Fluorescence | 2.5 h | G4/ThT | This work |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wen, Y.; Wang, L.; Li, L.; Xu, L.; Liu, G. A Sensitive and Label-Free Pb(II) Fluorescence Sensor Based on a DNAzyme Controlled G-Quadruplex/Thioflavin T Conformation. Sensors 2016, 16, 2155. https://doi.org/10.3390/s16122155
Wen Y, Wang L, Li L, Xu L, Liu G. A Sensitive and Label-Free Pb(II) Fluorescence Sensor Based on a DNAzyme Controlled G-Quadruplex/Thioflavin T Conformation. Sensors. 2016; 16(12):2155. https://doi.org/10.3390/s16122155
Chicago/Turabian StyleWen, Yanli, Lele Wang, Lanying Li, Li Xu, and Gang Liu. 2016. "A Sensitive and Label-Free Pb(II) Fluorescence Sensor Based on a DNAzyme Controlled G-Quadruplex/Thioflavin T Conformation" Sensors 16, no. 12: 2155. https://doi.org/10.3390/s16122155
APA StyleWen, Y., Wang, L., Li, L., Xu, L., & Liu, G. (2016). A Sensitive and Label-Free Pb(II) Fluorescence Sensor Based on a DNAzyme Controlled G-Quadruplex/Thioflavin T Conformation. Sensors, 16(12), 2155. https://doi.org/10.3390/s16122155





