Abstract
Spring pollinosis has become a part of life for many people throughout the world. A wide range of knowledge about the allergenic potential of individual pollen allergen types is documented well, but the starting point of the pollen allergens expression regulation in plants itself is still not fully answered. Expression analysis of pollen allergens does not yet have any specific protocols or methods developed, despite a very good sequence background available in public bioinformatics databases. However, research in this area of interest has a great application potential for breeding and biotechnology of allergenic plants that may benefit from the knowledge of the expression of allergen coding genes in individual varieties or genotypes. Here, a brief review of up-to-date knowledge about the coding sequences of central and eastern European spring pollen allergens is introduced together with real-time based analysis of the expression of two of the main pollen allergens–PR protein type and profilin type of birch and hazelnut.
1. North, Central and Eastern Europe Spring Pollinosis Sources
Bronchial asthma, allergic rhinitis and allergic conjunctivitis—these are some of symptoms of pollinosis. Seasonal pollen allergy is the most common allergic disease with the significantly increasing tendency. Nowadays, it affects up to one fifth of the population and it can occur at any time during life, but most often between the 15th and 25th year. Whether the allergy appears during life, is an interaction among heredity factors, intestinal microflora, nutrition and the length of time breast-fed [1].
The most important cause of the IgE-mediated allergy is pollen and its proteins from trees and grasses [2]. When thinking of trees, not only is pollen the source of allergens, but also fruits and seeds. In this review, the main objective is a summarization of the current knowledge and already developed methods for the analysis of the expression of pollen allergens per se, directly in pollen. Analyses such as these are important, especially when considering that every pollinosis season differs when compared to previous ones. As will be seen further in the text, not only the amount of pollen in the atmosphere, which is strongly dependent on the weather conditions, but also the place of the growth decides the level of allergen expression, i.e., releasing of allergenic molecules. Variations exist in allergen content in pollen grains and, moreover, allergens are also released via submicroscopic particles [3,4]. This is why quantifying different aspects of allergen characteristics has become realized [5].
The most allergenic tree pollen is produced in spring in Europe by alder, hazel, yew, elm, willow, ash, birch, plane and poplar. The main period of pollen release with peak periods depends on the region. Alder and hazel are the first to flower (December–April) followed by willow and elm (February–April), then ash and birch (March–May) and finally poplar (March–April). All of these trees collectively are the most important source of European spring pollinosis (Table 1) and the main allergens of the highly allergenic trees are well defined at the protein level and accessible in different allergen databases, such as WHO/IUIS (World Health Organization/International Union of Immunological Societes) Allergen Nomenclature (www.allergen.org); Allergome (www.allergome.org); SDAP (Structural Database of Allergenic Proteins) (www.fermi.utmb.edu); AllFam (www.meduniwien.ac.at) or Allergen Online (www.allergenonline.org). Almost all known tree pollen allergens are low-molecular weight intracellular proteins or glycoproteins and are released after touching an aquatic surface such as human mucous membranes. Many comprehensive descriptions of the structural characteristics of individual functions of pollen allergenic proteins are given in the literature [6,7], which is why only a very brief description of the main allergenic proteins is given here in Table 1. They exist in different trees as proteins with significant sequence homology and cross-reactive epitopes [8].

Table 1.
Allergenic proteins characteristics of main European spring pollinosis.
2. Spring Tree Pollinosis—Actual Allergen Nucleic Acids Sequences and Function Knowledge
Pollen from different species of monocotyledonic or and dicotyledonic plants is regarded to be the most frequent and active allergen source. Pollen allergens are low molecular weight proteins or glycoproteins and are located intracellular in the pollen grains. Their physiological activation starts after pollen hydration on mucosal surfaces or by liberation from small allergen containing respirable particles [14].
Currently, it is still in most cases impossible to purify and describe all of the major and minor allergens of natural allergen sources that are active in aqueous buffers. Thanks to the advance in the field of molecular genetics, this problem can be overcome by using cDNAs for recombinant production of allergens from their sources [8].
Obtaining the allergen-encoded cDNAs, three different methods are applied. The first is based on immunoscreening, the second is based on amino acids determination, and the third usesthe sequence similarity at the protein and nucleic acid level. The workflow for the first one is: mRNA isolation from allergenic pollen → cDNA synthesis → construction of expressed cDNA library → IgE immunoscreening → sequencing. Today, using this method together with robotic-based screening provides cost effective and rapid identification of allergens, even from complex allergenic sources [15]. The workflow for the second is: Allergen protein purification → determination of amino acid composition and N-terminal amino acid sequence → oligonucleotides design → PCR cloning or screening of cDNA libraries → sequencing. The workflow for the third is: Known cDNA sequences and amino acids sequences comparison → degenerate primers design → the first-strand cDNA amplification and amplification of 5′ cDNA ends → cloning and sequencing of cDNA. Comparing them to the immunoscreening method, the PCR based technology is much more effective and the growing amount of data stored in public databases of allergens facilitate allergen cloning using this approach [16]. Described methods was used for the isolation of cDNA of following spring pollen allergens: Bet v1, Bet v2, Bet v3, Bet v4, Aln g 4, Cor a 1 and Aln g 1 [16,17,18,19,20]. When considering north, central and eastern Europe, the most allergenic tree pollen is produced by the Order of Fagales. It comprises the three main allergenic families—Betulaceae, Corylaceae and Fagaceae. Pollen allergens that are known for their genera are listed in Table 2.

Table 2.
Accession codes of nucleotide sequences for spring pollinosis allergens.
3. Birch Pollen Allergens—Actual DNA/RNA Sequence Data
Bet v1: A major birch allergen originated in Betulla verrucosa is actually the most known and described as one with in total three subfamilies that were identified by their sequences similarity comparison [22]. All three subfamilies are abundant in vascular plants. The first, pathogenesis-related protein family PR-10, is expressed in plants following the signals as pathogen attack or abiotic stress. They are most concentrated in reproductive tissues—pollen, seeds and fruits [23]. This birch allergen protein was identified to posses a very similar structure with the human lipocalin 2 [24]. Both were invented to have the specific structures that allowed them to bind iron and only in the situation when no iron is binding on birch Bet v1 protein it becomes an allergen by the manner of affecting of Th2 cells of the human immune system [24]. Park et al. [25] reported the ribonuclease activity of pathogenesis-related proteins with the function in antiviral pathway. The other subfamily of Bet v1 allergens was described as a group of major latex proteins and ripening-related proteins in the latex of opium poppy [26,27]. The last of the Bet v1 subfamilies comprises proteins containing members with S-norcoclaurin synthase activity and are involved in alkaloid biosynthesis [28].
In total, nine Bet v1 isoforms are known as to be naturally occurring, a, b, c, d, e, f, g, j and i. When regarding the induction of type I allergic symptoms, they differ among themselves by the ability to react with the IgE from patients and it is reported [29] that the IgE binding activity, when compared in vitro and in vivo, is characterized directly by the six amino acid residues at different positions of the Bet v1 molecule. Betula verrucosa/pendula major pollen allergen is well defined on the nucleotide level and, in total, 47 isoallergens sequences are available in the NCBI database for mRNA with the sequence identities. Here, the conserved parts were obtained first by megablast algorithm and, subsequently, the nucleotide differences were found among them as summarized in Figure 1. The fylogenetic similarity of the sequences of Bet v1 isoallergens with the gene coding for the major birch pollen allergen Betv1 (X15877.1) is illustrated in the dendrogram constructed by NJ procedure (Figure 2).

Figure 1.
Numbers of nucleotide differences among Bet v1 isoforms for the conserved part based on the NCBI data. * No sequence homology found; all the isosoforms are compared to: X15877.1 and the sequences are coded as follow: 1, AF124839.1; 2, AF124838.1; 3, AF124837.1; 4, AJ002110.1; 5, AJ002109.1; 6, AJ002108.1; 7, AJ002107.1; 8, AJ002106.1; 9, X77200.1; 10, X77272.1; 11, X77274.1; 12, X77273.1; 13, X77271.1; 14, X77270.1; 15, X77268.1; 16, X77267.1; 17, X77266.1; 18, X77265.1; 19, X77269.1; 20, X77599.1; 21, X77600.1; 22, X77601.1; 23, Y12560.1; 24, AJ006915.1; 25, AJ006914.1; 26, AJ006913.1; 27, AJ006912.1; 28, AJ006911.1; 29, AJ006910.1; 30, AJ006909.1; 31, AJ006908.1; 32, AJ006907.1; 33, AJ006905.1; 34, AJ006904.1; 35, AJ006903.1; 36, AJ006906.1; 37, Z80106.1; 38, Z80105.1; 39, Z80103.1; 40, Z80102.1; 41, Z80101.1; 42, Z80100.1; 43, Z80099.1; 44, Z80098.1; 45, Z80104.1; 46, X82028.1; and 47, X81972.1.
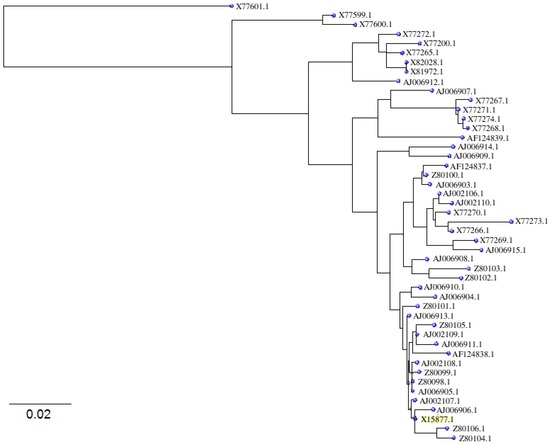
Figure 2.
Sequences fylogenesis of Bet v1 isoallergens. The isoforms correspond with the number codes in Figure 1.
4. Minor Central and Eastern Pollen Allergens—Actual DNA/RNA Sequence Data
Closely-related and cross-reactive allergens with the Bet v1 were found in the pollen of some other trees from the order Fagales such as Corylus avellana, Alnus glutinosa or Castanea sativa (Table 2). For this allergen, the IgE cross-reactivity among Bet v1 and homologous allergens from fruits or vegetables that belong to the Rosaceae, Apiaceae or Fabaceae is often described [22].
Fra e 1 is amain pollen allergen of ash that is widely distributed in both central and northern Europe. The clinical relevance of the ash pollen allergen is quite difficult to value because its pollination overlaps with that of Betulaceae [30]. Currently, three main isoforms of Fra e 1 are stored in the sequence database with the following accession numbers: AY652744.1, AY377127.1 and AF526295.1. All of them are for mRNA. The sequence identity between AY652744.1 and AF526295.1 is 99%. In total, four differences exist in their sequences: in AF526295.1 G is replaced by C in the 135th position, C is replaced by G in the 271st position, G is replaced by A in the 300th position and C is replaced by T in the 380th position. The sequence identity between AY652744.1 and AY377127.1 is 89% with the differences listed in the Table 3.

Table 3.
Nucleotide differences between ash Fra e 1 allergen mRNA isoforms.
Aln g 1, the major allergen in pollen of alder, is also a member of the Bet v1 family. Only a sequence of mRNA actually exists in the public databases with the accession number S50892. Valenta et al. [31] has performed the alignments among the selected major allergens— Bet v1, Aln g 1 and Cor a 1. In hybridization, similar binding pattern of Bet v1 cDNA probe was detected to pollen RNAs from all of the analyzed species. This confirms the high homology of the mRNAs coding for the analyzed allergens. Based on these observations, an extract with only one major allergen from those analyzed should be sufficient for diagnostic and therapeutic purposes, as the authors have concluded [31].
5. Hazelnut Pollen Allergens—Actual DNA/RNA Sequence Data
Avery different bioinformatics situation exist for Corylus avellana major allergen Cor a 1. Currently, 1649 profilin sequences for different plant species are available at NCBI GenBank database and about half of them were isolated from pollen. Jimenez-Lopez et al. [7] reported in their study that no data regarding interspecific comparisons and cultivars sequence variability is available for pollen profilins in the literature and the presence of polymorphism is described in a low number of sequences in different plant species.
For hazel pollen profilin, in total eight isoallergens for mRNA and two sequences for DNA on the nucleotide level sequences are available in the NCBI database. Performing the sequence alignment, there can be recognized three groups that posses no identities among the groups but only within them (Table 4). The approach based on finding of the most conserved sequences cannot be applied for all of them but only for an individual group. The above-mentioned fully mirror the actual molecular knowledge situation about the profilins. Profilins are a multigene family that were summarized by Jimenez-Lopez et al [7] as having the functions in actin dynamics, beside their allergenic potential, which was described for olive, birch, maize, hazel and timothy-grass.

Table 4.
Profilin sequence of hazelnut stored in the NCBI database.
Profilin is a class of plant pan-allergen, as the profilins from very different plant species are responsible for similar allergic reactions in sensitized patients. Moreover, 20% of all pollen allergic patients react to plant profilins and many of the pollen expressed proteins that bind on IgE in allergic patients have been found to be profilins [32]. Profilins are known to be expressed by up to ten different genes in plants [33]. Today, two main groups are recognized depending on their manner of expression. The first group is expressed constitutively and in all plant tissues and the second one is expressed only in the reproductive tissues [34]. The distribution of profilin in pollen depends on the stage of development and the specific isoforms expressed [35].
6. Spring Tree Pollinosis—Expression Analysis of Bet v1, Bet v2, Cor a 1 and Cor a 2 in Pollen Grains from DifferentIn Situ Conditions
Real-time PCR based expression analysis of food allergens per se is well established for almost all food allergens [36,37]. For pollen allergens, the situation regarding the expression of allergen genes in the pollen grains is very different. The application of real-time PCR in the analysis of the expression level of pollen allergens from differentin situ conditions is only in the experimental phase and no normalized methods actually exist. Real-time PCR was reported as efficient in pollen monitoring by [38]. The authors have used nuclear ribosomal RNA genes as target for the quantization to improve conservation within species. The whole genome gene expression analysis was performed for olive pollen samples by [39], but the expression of specific pollen allergens directly in pollen was reported to be rare [40,41].
Real-time PCR has the potential to be used in routine comparative analysis of allergen expressionin situ and the subsequent health and land management can be applied, such as in the case of the results of different levels of birch Bet v1 expression. In the study of [40], different localities of Ukraine were chosen for the comparative study. Bet v1 expression in the pollen from different part of cities was from 0.77 to 2x higher as the pollen collected in the forest. Bet v1 expression in the pollen from village 0.55x higher when comparing to the forest sample. These findings have started the discussion about the presence of birch trees in the specific part of Kiyev (personal communication with Katerina Garkava—Institute of Ecological Safety, NAU in Kyiv). This is why setting up the validated methods will have very practical outcomes. In the case of the expression levels of CorA allergen in the Corylus avellana, the situation was reported to be similar. The level of CorA expression was within the range of 0.532 to 1.206x higher for the samples from urbanized area when compared to the sample from the village [41]. Very different data were obtained for the expression of hazelnut pollen profilin. The same samples as in the analysis of CorA allergen expression were used, but the results have shown a high variation in the abundance of profilin allergen transcripts. Expression levels were in the range of 2.957 up to the 52.936 higher when comparing samples from urbanized area to the sample from the village. In the downtown area, the profilin expression reached on average 21 times higher values and, in the sample from cement plant area, the expression is reported to be 52 times higher [41]. The interaction between air pollution and allergenic vegetation was reported previously [42] to be a consequence of the coverage of air pollution by micro-particulates. The results of the transcriptomic analysis [40,41] show that the effect of the higher allergenic potential of pollen grains in urbanized area has synergic background and the expression of allergens is higher there per se.
Real-time PCR is reported by many authors as a very efficient tool in many specific applications such as gene expression [43], identification and quantification of different pathogens [44] or authentication of food [45]. For the purposes of detection and quantification of airborne allergenic pollen taxa, real-time PCR was also applied. Longhi et al. [38] have reported the technique as a rapid, accurate, and automated for pollen grain quantitation and control.
Differences among levels of expression are well documented for many allergens and for different stages of the growth [46,47]. Wani et al. [48] reported that different expression levels of allergen were obtained for variable climatic conditions.
Today, different approaches can be applied for the development of real-time PCR protocol [49,50]. Regarding the pros of data mining, the first step of the transcriptomic analysis (Figure 3) in the case that mRNA sequences of allergens are known and stored in public databases is the sequences alignment. This is aimed to identify their conserved and variable parts. The most appropriate regions for primer designation can be mined by BLAST analysis [51] against the individual sequences. Using this approach, verification steps of the product specificity checking must be a part of the analysis work-flow. Here, HRM analysis or restriction cleavage can be chosen beside the melting analysis of the amplified target.
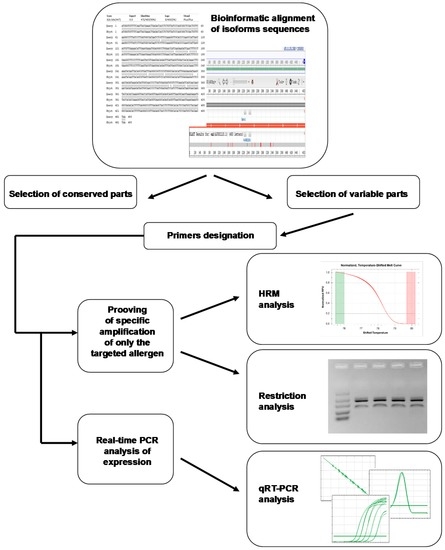
Figure 3.
Real-time PCR analysis work-flow of allergen expression based on data mining.
Another crucial step for the accurate quantification of analyzed transcripts is the identification of stable reference genes to normalize the target levels when using real-time PCR [52]. Those commonly used are housekeeping genes that are proven to be non-regulated. However, some studies reported high variation of tested housekeeping genes under different conditions and this is why no current universal reference gene exists [53], which is why the standard for using housekeeping genes is the approach where a set of reference genes (2–3 different genes) is validated [54].
Different housekeeping genes were reported as internal controls in real-time PCR analysis [55,56]. Cyclophylin, alpha-tubulin, and transcription factor CBF1 were tested as controls to normalize expression of Silver birch pollen Bet v1 allergen by [38]. Comparison of Corylus avellana pollen expression level using 18S rRNA as internal control was performed by [41]. HMG CoA reductase was validated for hazelnut pollen expression by [57].
7. Conclusions
The mechanism and regulation of allergens expression and the questions of regulatory processes is the research interest with a high impact for the future. All relevant findings will help to manage pollinosis that many people suffer with through the management of the biology of the allergens directly in the plants.
Acknowledgments
This work has been supported by the European Community under project No. 26220220180: Building Research Centre “AgroBioTech”.
Conflicts of Interest
The authors declare no conflict of interest.
List of Abbreviations
| BLAST | Basic Local Alignment Search Tool |
| HRM | High Resolution Melting |
| qRT-PCR | quantitative Real Time Polymerase Chain Reaction |
| NCBI | National Center for Biotechnology Information |
| PCR | Polymerase Chain Reaction |
References
- M’Rabet, L.; Vos, A.P.; Boehm, G.; Garssen, J. Breast-Feeding and Its Role in Early Development of the Immune System in Infants: Consequences for Health Later in Life. J. Nutr. 2008, 138, 1782S–1790S. [Google Scholar] [PubMed]
- Kay, A.B. Allergy and Allergic Diseases; Blackwell Publishing Ltd.: Oxford, UK, 2008. [Google Scholar]
- Buters, J.T.; Kasche, A.; Weichenmeier, I. Year-to-year variation in release of Bet v1 allergen from birch pollen: Evidence for geographical differences between West and South Germany. Int. Arch. Allergy Immunol. 2007, 145, 122–130. [Google Scholar] [CrossRef] [PubMed]
- Taylor, P.E.; Jacobson, K.W.; House, J.M.; Glovsky, M.M. Links between pollen, atopy and the asthma epidemic. Int. Arch. Allergy Immunol. 2007, 144, 162–170. [Google Scholar] [CrossRef] [PubMed]
- Rittenour, W.R.; Hamilton, R.G.; Beezhold, D.H.; Green, B.J. Immunologic, spectrophotometric and nucleic acid based methods for the detection and quantification of airborne pollen. J. Immunol. Methods 2012, 383, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Grote, M.; Vrtala, S.; Valenta, R. Monitoring of two allergens, Bet v1 and profillin, in dry and rehydrated birch pollen by immunogold electron microscopy and immunoblotting. J. Histochem. Cytochem. 1993, 41, 745–750. [Google Scholar] [CrossRef] [PubMed]
- Jimenez-Lopez, J.C.; Morales, S.; Castro, A.J. Characterization of Profilin Polymorphism in Pollen with a Focus on Multifunctionality. PLoS ONE 2012, 7, 2. [Google Scholar] [CrossRef] [PubMed]
- Marth, K.; Germatuik, T.; Swoboda, I.; Valenta, R. Tree pollen allergens. In Allergens and Allergen Immunotherapy: Subcutaneous, Sublingual, and Oral, 5th ed.; Lockey, R.F., Ledford, D.K., Eds.; CRC Press: Boca Raton, FL, USA, 2014. [Google Scholar]
- Rodríguez, R.; Villalba, M.; Monsalve, R.I.; Batanero, E. The spectrum of olive pollen allergens. Int. Arch. Allergy Immunol. 2001, 125, 185–195. [Google Scholar] [CrossRef] [PubMed]
- Romano, P.G.N.; Horton, P.; Gray, J.E. The Arabidopsis Cyclophilin Gene Family. Plant Physiol. 2004, 134, 1268–1282. [Google Scholar] [CrossRef] [PubMed]
- Kumari, S.; Roy, S.; Singh, P.; Singla-Pareek, S.; Pareek, A. Cyclophilins: Proteins in search of function. Plant Signal. Behav. 2013, 8, e22734. [Google Scholar] [CrossRef] [PubMed]
- Luttkopf, D.; Muller, U.; Skov, P.S. Comparison of four variants of a major allergen in hazelnut (Corylusavellana) Cor a 1.04 with the major hazel pollen allergen Cor a 1.01. Mol. Immunol. 2002, 38, 515–525. [Google Scholar] [CrossRef]
- Pokoj, S.; Lauer, I.; Fotisch, K. Pichiapastoris is superior to E-coli for the production of recombinant allergenic non-specific lipid-transfer proteins. Protein Expr. Purif. 2010, 69, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Hayek, B.; Vangelista, L.; Pastore, A. Molecular and immunological characterization of a highly cross-reactive two EF-hand calcium-binding alder pollen allergens, Aln g 4: Structural basis for calcium-modulated IgE recognition. J. Immunol. 1998, 161, 7031–7039. [Google Scholar]
- Crameri, R. High throughput screening: A rapid way to recombinant allergens. Allergy 2001, 56, 30–34. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.S.; Shen, H.H.; Zheng, M.; Frewer, L.J.; Gilissen, W.J. Multidisciplinary Approaches to Allergies; Zhejiang University Press: Hangzhou, China, 2012. [Google Scholar]
- Breiteneder, H.; Pettenburger, K.; Bito, A. The gene coding for the major birch pollen allergen, Bet v 1, is highly homologous to a pea disease resistance response gene. EMBO J. 1989, 8, 1935–1938. [Google Scholar] [PubMed]
- Breiteneder, H.; Ferreira, F.; Hoffmann-Sommergruber, K. Four recombinant isoforms of Cora I, the major allergen of hazel pollen, show different IgE-binding properties. Eur. J. Biochem. 1993, 212, 355–362. [Google Scholar] [CrossRef] [PubMed]
- Seiberler, S.; Scheiner, O.; Kraft, D. Characterization of a birch pollen allergen, Bet v III, representing a novel class of Ca2+ binding proteins; specific expression in mature pollen and dependence of patients’ IgE binding on protein-bound Ca2+. EMBO J. 1994, 13, 3481–3486. [Google Scholar] [PubMed]
- Twardosz, A.; Hayek, B.; Seiberler, S. Molecular characterization, expression in Escherichia coli and epitope analysis of a two EF-hand calcium-binding birch pollen allergen, Bet v 4. Biochem. Biophys. Res. Commun. 1997, 239, 197–204. [Google Scholar] [CrossRef] [PubMed]
- Zuidmeer-Jongejan, L.; Fernández-Rivas, M.; Winter, M.G.T. Oil body-associated hazelnut allergens including oleosins are underrepresented in diagnostic extracts but associated with severe symptoms. Clin. Trans. Allergy 2014, 4. [Google Scholar] [CrossRef] [PubMed]
- Liscombe, D.K.; MacLeod, B.P.; Loukanina, N. Evidence for the monophyletic evolution of benzylisoquinoline alkaloid biosynthesis in angiosperms. Phytochemistry 2005, 66, 2501–2520. [Google Scholar] [CrossRef] [PubMed]
- Markovic-Housley, Z.; Degano, M.; Lamba, D. Crystal structure of a hypoallergenic isoform of the major birch pollen allergen Bet v 1 and its likely biological function as a plant steroid carrier. J. Mol. Biol. 2003, 325, 123–133. [Google Scholar] [CrossRef]
- Roth-Walter, F.; Gomez-Casado, C.; Pacios, L.F. Bet v 1 from birch pollen is a lipocalin-like protein acting as allergen only when devoid of iron by promoting Th2lymphocytes. J. Biol. Chem. 2014, 289, 17416–17421. [Google Scholar] [CrossRef] [PubMed]
- Park, C.J.; Kim, K.J.; Shin, R. Pathogenesis-related protein 10 isolated from hot pepper functions as a ribonuclease in an antiviral pathway. Plant J. 2004, 37, 186–198. [Google Scholar] [CrossRef] [PubMed]
- Osmark, P.; Boyle, B.; Brisson, N. Sequential and structural homology between intracellular pathogenesis-related proteins and a group of latex proteins. Plant Mol. Biol. 1998, 38, 1243–1246. [Google Scholar] [CrossRef] [PubMed]
- Vieths, S.; Scheurer, S.; Ballmer-Weber, B. Current understanding of cross-reactivity of food allergens and pollen. Ann. N. Y. Acad. Sci. 2002, 964, 47–68. [Google Scholar] [CrossRef] [PubMed]
- Samanani, N.; Liscombe, D.K.; Facchini, P.J. Molecular cloning and characterization of norcoclaurine synthase, an enzyme catalyzing the first committed step in benzylisoquinoline alkaloid biosynthesis. Plant J. 2004, 40, 302–313. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, F.; Ebner, C.; Kramer, B. Modulation of IgE reactivity of allergens by site-directed mutagenesis: Potential use of hypoallergenic variants for immunotherapy. FASEB J. 1998, 12, 231–242. [Google Scholar]
- Valenta, R.; Breiteneder, V.; Petternburger, K. Homology of the major birch-pollen allergen, Bet v I, with the major pollen allergens of alder, hazel, and hornbeam at the nucleic acid level as determined by cross-hybridization. J. Allergy Clin. Immun. 1991, 87, 677–682. [Google Scholar] [CrossRef]
- Mas, S.; Torres, M.; Garrodo-Arandia, M. Ash pollen immunoproteomics: Identification, Immunologic characterization, and sequencing of 6 new allergens. J. Allergy Clin. Immun. 2014, 133, 923–926. [Google Scholar] [CrossRef] [PubMed]
- Valenta, R.; Ferreira, F.; Grote, M. Identification of profilin as an actin-binding protein in higher plants. J. Biol. Chem. 1993, 268, 22777–22781. [Google Scholar] [PubMed]
- Ren, H.; Xiang, Y. The function of actin-binding proteins in pollen tube growth. Protoplasma 2007, 230, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Kandasamy, M.K.; McKinney, E.C.; Meagher, R.B. Plant profiling isovariants are distinctly regulated in vegetative and reproductive tissues. Cell Motil. Cytoskelet. 2002, 52, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Kovar, D.R.; Drøbak, B.K.; Staiger, C.J. Maize profilin isoforms are functionally distinct. Plant Cell 2000, 12, 583–598. [Google Scholar] [CrossRef] [PubMed]
- Lauer, I.; Alessandri, S.; Pokoj, S. Expression and characterization of three important panallergens from hazelnut. Mol. Nutr. Food Res. 2008, 52, 262–271. [Google Scholar] [CrossRef] [PubMed]
- Garino, C.; Locatelli, M.; Coïsson, J.D. Gene transcription analysis of hazelnut (Corylusavellana L.) allergens Cor a 1, Cora 8 and Cor a 11: A comparative study. Int. J. Food Sci. Technol. 2013, 48, 1208–1217. [Google Scholar] [CrossRef]
- Longhi, S.; Cristofori, A.; Gatto, P. Biomolecular identification of allergenic pollen: A new perspective for aerobiological monitoring? Ann. Allergy Asthma Immunol. 2009, 103, 508–514. [Google Scholar] [CrossRef]
- Aguerri, M.; Calzada, D.; Montaner, D. Differential gene-expression analysis defines a molecular pattern related to olive pollen allergy. J. Biol. Regul. Homeost. Agents 2013, 27, 337–350. [Google Scholar] [PubMed]
- Žiarovská, J.; Labajová, M.; Ražná, K. Changes in expression of BetV1 allergen of silver birch pollen in urbanized area of Ukraine. J. Environ. Sci. Health 2013, 48, 1479–1484. [Google Scholar] [CrossRef] [PubMed]
- Ražná, K.; Bežo, M.; Nikolaieva, N. Variability of Corylus avellana, L. CorA and profilin pollen allergens expression. J. Environ. Sci. Health 2014, 49, 639–645. [Google Scholar] [CrossRef] [PubMed]
- D’Amato, G.; Liccardi, G.; D’Amato, M.; Holgatew, S. Environmental risk factors and allergic bronchial asthma. Clin. Exp. Allergy 2005, 35, 1113–1124. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, T.; Thomas, S. Gene expression and characterization of a stress-induced tyrosine decarboxylase from Arabidopsis thaliana. FEBS Lett. 2009, 583, 1895–1900. [Google Scholar] [CrossRef] [PubMed]
- Pochop, J.; Kačániová, M.; Hleba, L. Detection of Listeria monocytogenes in ready-to-eat food by step one real-time polymerase chain reaction. J. Environ. Sci. Health B 2012, 47, 212–216. [Google Scholar] [CrossRef] [PubMed]
- Farrokhi, R.; Joozani, R.J. Identification of pork genome in commercial meat extracts for Halal authentication by SYBR green I real-time PCR. Int. J. Food Sci. Technol. 2011, 46, 951–955. [Google Scholar] [CrossRef]
- Kang, I.H.; Srivastava, P.; Ozias-Akins, P.; Gallo, M. Temporal and Spatial Expression of the Major Allergens in Developing and Germinating Peanut Seed. Plant Physiol. 2007, 144, 836–845. [Google Scholar] [CrossRef] [PubMed]
- Platteau, C.; De Loose, M.; De Meulenaer, B.; Taverniers, I. Detection of allergenic ingredients using real-time PCR: A case study on hazelnut (Corylus avellena) and soy (Glycine max). J. Agric. Food Chem. 2011, 59, 10803–10814. [Google Scholar] [CrossRef] [PubMed]
- Wani, A.; Amin, Z. Airborne Pollen Allergy—Impact on Human Health; Lambert Academic Publishing: Saarbrücken, Germany, 2012. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−∆∆CT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Xie, L.; Chen, J. A novel procedure for absolute real-time quantification of gene expression patterns. Plant Methods 2012, 8, 9. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schaffer, A.A. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Phillips, M.A.; D’Auria, J.C.; Luck, K.; Gershenzon, J. Evaluation of Candidate Reference Genes for Real-Time Quantitative PCR of Plant Samples Using Purified cDNA as Template. Plant Mol. Biol. Rep. 2009, 27, 407–416. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, L.; Mauriat, M.; Guenin, S. The lack of a systematic validation of reference genes: A serious pitfall undervalued in reverse transcription-polymerase chain reaction (RT-PCR) analysis in plants. Plant Biotechnol. J. 2008, 6, 609–618. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.F.; Sun, S.S.M.; Yuan, D.Y. Validation of Candidate Reference Genes for the accurate normalization of Real-Time Quantitative RT-PCR Data in Rice during seed development. Plant Mol. Biol. Rep. 2010, 28, 49–57. [Google Scholar] [CrossRef]
- Nicot, N.; Hausman, J.F.; Hoffmann, L.; Evers, D. Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J. Exp. Bot. 2005, 56, 2907–2914. [Google Scholar] [CrossRef] [PubMed]
- Garg, R.; Sahoo, A.; Tyagi, A.K.; Jain, M. Validation of internal control genes for quantitative gene expression studies in chickpea (Cicer arietinum L.). Biochem. Biophys. Res. Commun. 2010, 396, 283–288. [Google Scholar] [CrossRef] [PubMed]
- Žiarovská, J.; Nikolajeva, N.; Garkava, K.; Brindza, J. Validation of HMG CoA reductase as internal control for hazelnut pollen allergens expression analysis. Austin J. Genet. Genom. Res. 2015, 2, 1–4. [Google Scholar]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).