Building a Plant DNA Barcode Reference Library for a Diverse Tropical Flora: An Example from Queensland, Australia
Abstract
:1. Introduction
2. Methods
3. Results and Discussion
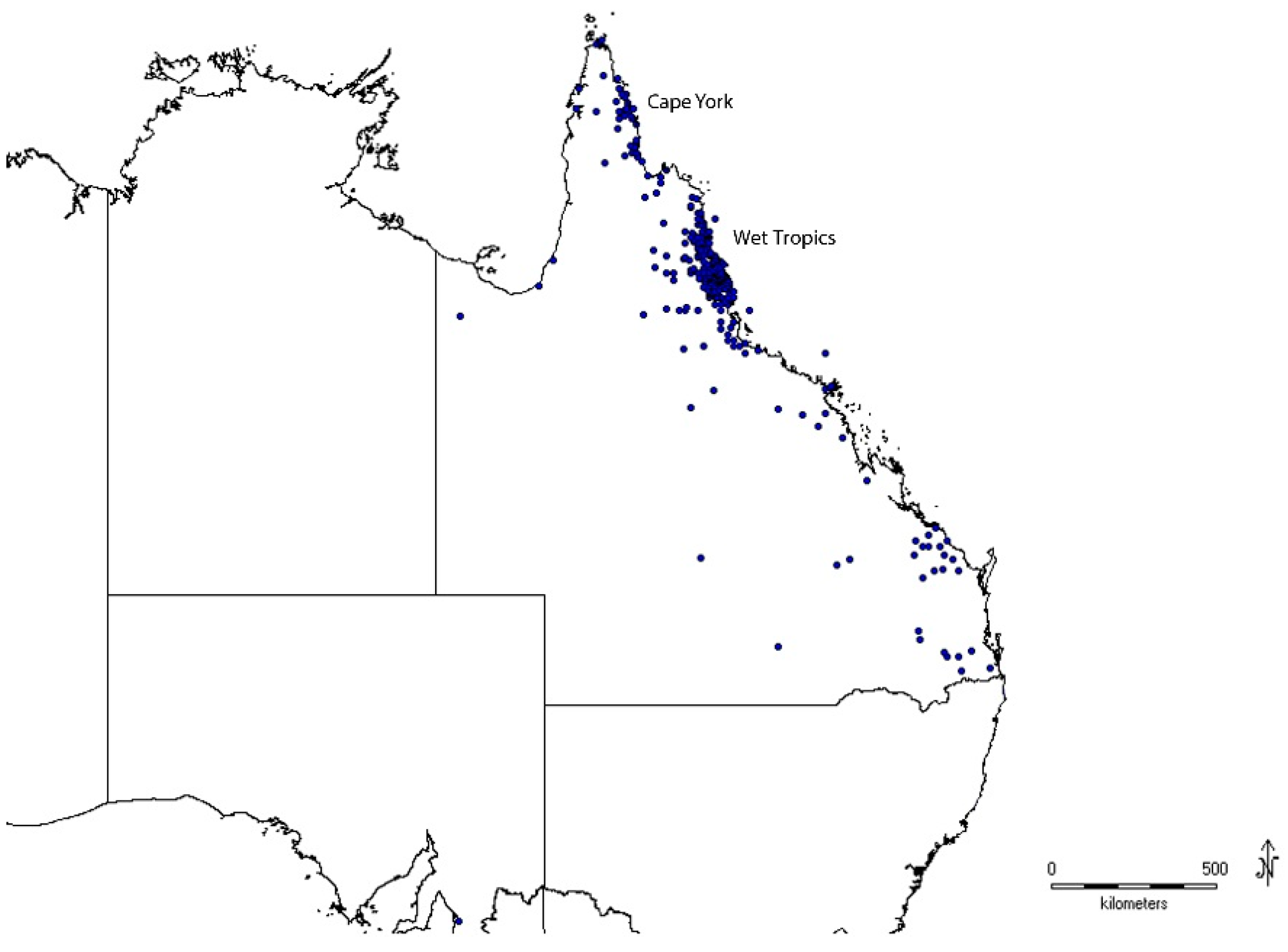
3.1. Taxonomic Coverage
3.2. Variation within Species across Geographic Breaks
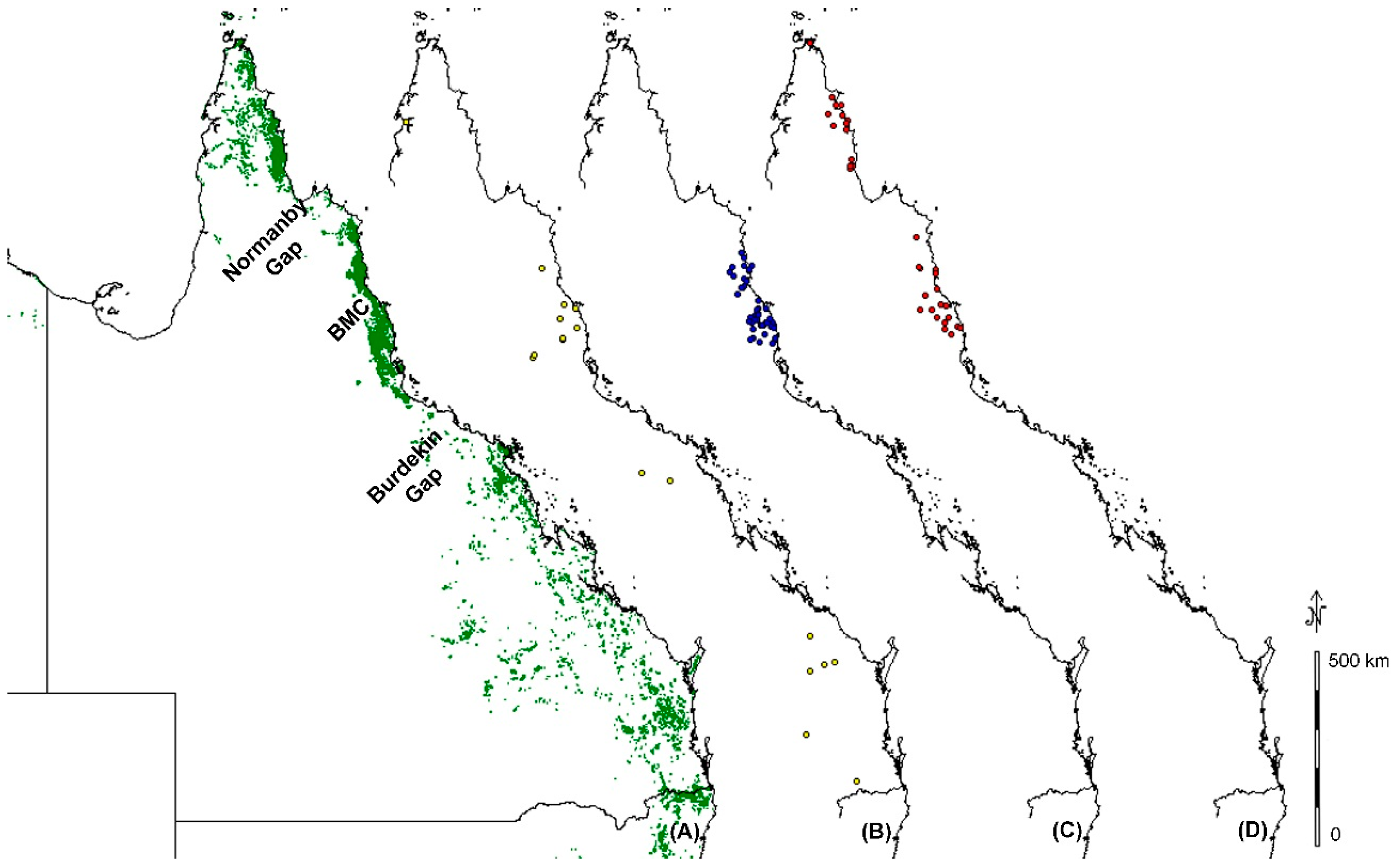
3.3. Burdekin Gap
3.4. Black Mountain Corridor
3.5. Normanby Gap
4. Conclusions
4.1. DNA Barcodes as a Tool for Studying Biogeography
4.2. Data Resources
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Corlett, R.T. Flora and reproductive phenology of the rain-forest at Bukit-Timah, Singapore. J. Trop. Ecol. 1990, 6, 55–63. [Google Scholar] [CrossRef]
- Costion, C.M.; Liston, J.; Kitalong, A.H.; Iida, A.; Lowe, A.J. Using the ancient past for establishing current threat in poorly inventoried regions. Biol. Conserv. 2012, 147, 153–162. [Google Scholar] [CrossRef]
- Pearson, D.L.; Hamilton, A.L.; Erwin, T.L. Recovery Plan for the Endangered Taxonomy Profession. Bioscience 2011, 61, 58–63. [Google Scholar] [CrossRef]
- Tancoigne, E.; Dubois, A. Taxonomy: No Decline, but inertia. Cladistics 2013, 29, 567–570. [Google Scholar] [CrossRef]
- Costion, C.; Ford, A.; Cross, H.; Crayn, D.; Harrington, M.; Lowe, A. Plant DNA Barcodes Can Accurately Estimate Species Richness in Poorly Known Floras. PLoS ONE 2011, 6, e26841. [Google Scholar] [CrossRef] [PubMed]
- Kesanakurti, P.R.; Fazekas, A.J.; Burgess, K.S.; Percy, D.M.; Newmaster, S.G.; Graham, S.W.; Barrett, S.C.H.; Hajibabaei, M.; Husband, B.C. Spatial patterns of plant diversity below-ground as revealed by DNA barcoding. Mol. Ecol. 2011, 20, 1289–1302. [Google Scholar] [CrossRef] [PubMed]
- Mahadani, P.; Gosh, S.K. DNA Barcoding: A Tool for Species Identification from Herbal Juices. DNA Barcodes 2013, 1, 35–38. [Google Scholar] [CrossRef]
- Sin, W.M.; Tam, Y.K.; Tsui, S.; Ng, C.; Mok, C.; Ha, W. An integrated and validated DNA-based protocol developed to fight against commercial frauds: A Case of Fraudulent Substitutions for Deer Products. DNA Barcodes 2013, 1, 27–34. [Google Scholar] [CrossRef]
- Vesterinen, E.J.; Lilley, T.; Laine, V.N.; Wahlberg, N. Next Generation Sequencing of Fecal DNA Reveals the Dietary Diversity of the Widespread Insectivorous Predator Daubenton’s Bat (Myotis daubentonii) in Southwestern Finland. PLoS ONE 2013, 8, e82168. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Robledo, C.; Erickson, D.L.; Staines, C.L.; Erwin, T.L.; Kress, W.J. Tropical Plant-Herbivore Networks: Reconstructing Species Interactions Using DNA Barcodes. PLoS ONE 2013. [Google Scholar] [CrossRef] [PubMed]
- Ratnasingham, S.; Hebert, P.D.N. BOLD: The Barcode of Life Data System (www.barcodinglife.org). Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Census of the Queensland Flora. Available online: http://www.qld.gov.au/environment/plants-animals/plants/herbarium/flora-census/index.html (accessed on 15 August 2015).
- Crisp, M.D.; Laffan, S.; Linder, H.P.; Monro, A. Endemism in the Australian flora. J. Biogeogr. 2001, 28, 183–198. [Google Scholar] [CrossRef]
- Wet Tropics Management Authority. State of the Wet Tropics Report 2013–2014. Wet Tropics Management Authority, Cairns, QLD, 2014. Available online: http://www.wettropics.gov.au/site/user-assets/docs/b5-wtma-report-2013-2014.pdf (accessed on 24 February 2016).
- CCDB Protocols, DNA Extraction. Available online: http://ccdb.ca/docs/CCDB_DNA_Extraction-Plants.pdf (accessed on 15 August 2015).
- BOLD Specimen Data Submission Protocol. Available online: http://www.boldsystems.org/index.php/resources/handbookchapter=3_submissions.html§ion=data_submissions (accessed on 15 August 2015).
- CCDB Protocols, PCR Amplification. Available online: http://ccdb.ca/docs/CCDB_Amplification-Plants.pdf (accessed on 15 August 2015).
- CCDB Protocols, Sequencing. Available online: http://ccdb.ca/docs/CCDB_Sequencing.pdf (accessed on 15 August 2015).
- Australian Plant Name Index. Available online: https://www.anbg.gov.au/apni/ (accessed on 15 August 2015).
- Hilbert, D.W.; Graham, A.; Hopkins, M.S. Glacial and interglacial refugia within a long-term rainforest refugium: The Wet Tropics Bioregion of NE Queensland, Australia. Palaeogeography. Palaeoclimatol. Palaeoecol. 2007, 251, 104–118. [Google Scholar] [CrossRef]
- Chapple, D.G.; Hoskin, C.J.; Chapple, S.N.J.; Thompson, M.B. Phylogeographic divergence in the widespread delicate skink (Lampropholis delicata) corresponds to dry habitat barriers in eastern Australia. Bmc. Evol. Biol. 2011. [Google Scholar] [CrossRef] [PubMed]
- Moritz, C.; Hoskin, C.; Graham, C.H.; Hugall, A.; Moussalli, A. Historical biogeography, diversity and conservation of Australia’s tropical rainforest herpetofauna. Conserv. Biol. Ser. (Cambridge) 2005, 10, 243–264. [Google Scholar]
- Schneider, C.; Moritz, C. Rainforest refugia and evolution in Australia’s Wet Tropics. Proc. Natl. Acad. Sci. USA 1999, 266, 191–196. [Google Scholar]
- Hugall, A.; Moritz, C.; Moussalli, A.; Stanisic, J. Reconciling paleodistribution models and comparative phylogeography in the Wet Tropics rainforest land snail Gnarosophia bellendenkerensis (Brazier 1875). Proc. Natl. Acad. Sci. USA 2002, 99, 6112–6117. [Google Scholar] [CrossRef] [PubMed]
- Rossetto, M.; Crayn, D.; Ford, A.; Mellick, R.; Sommerville, K. The influence of environment and life-history traits on the distribution of genes and individuals: A Comparative Study of 11 Rainforest Trees. Mol. Ecol. 2009, 18, 1422–1438. [Google Scholar] [CrossRef] [PubMed]
- BOLD Systems. Public Data Portal—Record List. BATF Download URL: http://www.boldsystems.org/index.php/Public_BINSearch?searchtype=records (accessed on 24 February 2016).
- BOLD Systems. Data Portal. Available online: http://www.boldsystems.org (accessed on 24 February 2016).
| Order | Families | Genera | Species | Samples |
|---|---|---|---|---|
| Alismatales | 1 | 5 | 5 | 5 |
| Apiales | 4 | 10 | 13 | 26 |
| Aquifoliales | 2 | 4 | 4 | 5 |
| Arecales | 1 | 9 | 9 | 18 |
| Asparagales | 3 | 35 | 36 | 41 |
| Asterales | 3 | 6 | 6 | 9 |
| Austrobaileyales | 1 | 1 | 1 | 3 |
| Boraginales | 1 | 4 | 4 | 7 |
| Brassicales | 1 | 1 | 1 | 2 |
| Canellales | 1 | 1 | 1 | 3 |
| Caryophyllales | 3 | 4 | 4 | 5 |
| Celastrales | 1 | 8 | 9 | 22 |
| Commelinales | 2 | 4 | 4 | 5 |
| Cucurbitales | 2 | 5 | 6 | 10 |
| Dilleniales | 1 | 2 | 3 | 4 |
| Dioscoreales | 1 | 1 | 1 | 1 |
| Ericales | 6 | 18 | 53 | 112 |
| Escalloniales | 1 | 1 | 4 | 5 |
| Fabales | 2 | 28 | 38 | 66 |
| Gentianales | 4 | 44 | 69 | 134 |
| Huerteales | 1 | 1 | 1 | 3 |
| Lamiales | 6 | 23 | 25 | 45 |
| Laurales | 4 | 13 | 54 | 131 |
| Liliales | 3 | 4 | 4 | 6 |
| Magnoliales | 3 | 14 | 25 | 75 |
| Malpighiales | 15 | 44 | 68 | 129 |
| Malvales | 3 | 15 | 24 | 61 |
| Myrtales | 4 | 27 | 67 | 141 |
| Oxalidales | 3 | 11 | 20 | 38 |
| Pandanales | 1 | 1 | 1 | 1 |
| Paracryphiales | 1 | 2 | 2 | 3 |
| Piperales | 1 | 1 | 3 | 6 |
| Poales | 2 | 26 | 28 | 29 |
| Proteales | 1 | 18 | 20 | 40 |
| Ranunculales | 1 | 8 | 8 | 9 |
| Rosales | 6 | 21 | 36 | 65 |
| Santalales | 4 | 9 | 9 | 14 |
| Sapindales | 6 | 51 | 101 | 237 |
| Saxifragales | 1 | 2 | 2 | 4 |
| Solanales | 2 | 8 | 10 | 17 |
| Vitales | 1 | 5 | 6 | 8 |
| Zingiberales | 2 | 6 | 6 | 6 |
| Total | 113 | 496 | 857 | 1572 |
| Family | Species | Infra-Variation (bp) |
|---|---|---|
| Euphorbiaceae | Croton acronychioides F. Muell. | 2 |
| Malvaceae | Brachychiton australis (Schott & Endl.) A. Terracc. | 1 |
| Monimiaceae | Wilkiea macrophylla (A.Cunn.) A. DC. | 1 |
| Podocarpaceae | Podocarpus grayae de Laub. | 1 |
| Rubiaceae | Everistia vacciniifolia (F. Muell.) S. T. Reynolds & R. J. F. Hend. | 4 |
| Family | Species | Infra-Variation (bp) |
|---|---|---|
| Annonaceae | Goniothalamus australis Jessup | 1 |
| Apocynaceae | Tabernaemontana pandacaqui Lam. | 4 |
| Celastraceae | Hippocratea barbata F. Muell. | 2 |
| Eupomatiaceae | Eupomatia laurina R. Br. | 1 |
| Fabaceae | Archidendron ramiflorum (F. Muell.) Kosterm. | 1 |
| Fabaceae | Acacia celsa Tindale | 2 |
| Fabaceae | Castanospermum australe A. Cunn. ex Mudie | 8 |
| Fabaceae | Entada phaseoloides (L.) Merr. | 13 |
| Icacinaceae | Apodytes brachystylis F. Muell. | 2 |
| Lauraceae | Cryptocarya lividula B. Hyland | 1 |
| Lauraceae | Endiandra impressicosta C. K. Allen | 1 |
| Lauraceae | Endiandra leptodendron B. Hyland | 1 |
| Lauraceae | Cryptocarya bellendenkerana B. Hyland | 2 |
| Lauraceae | Cryptocarya densiflora Blume | 5 |
| Lauraceae | Endiandra insignis (F. M. Bailey) F. M. Bailey | 5 |
| Lauraceae | Endiandra longipedicellata C. T. White & W. D. Francis | 5 |
| Lauraceae | Beilschmiedia bancroftii (F. M. Bailey) C. T. White | 15 |
| Malvaceae | Argyrodendron peralatum (F. M. Bailey) Edlin ex J. H. Boas | 1 |
| Monimiaceae | Steganthera laxiflora (Benth.) Whiffin & Foreman | 4 |
| Moraceae | Streblus glaber (Merr.) Corner | 1 |
| Myristicaceae | Myristica globosa Warb. | 1 |
| Myrtaceae | Syzygium johnsonii (F. Muell.) B. Hyland | 1 |
| Myrtaceae | Syzygium kuranda (F. M. Bailey) B. Hyland | 1 |
| Myrtaceae | Leptospermum wooroonooran F. M. Bailey | 2 |
| Myrtaceae | Sphaerantia discolor Peter G. Wilson & B. Hyland | 4 |
| Myrtaceae | Rhodamnia sessiliflora Benth. | 9 |
| Phyllanthaceae | Glochidion sessiliflorum var. pedicellatum Airy Shaw | 7 |
| Rhamnaceae | Emmenosperma cunninghamii Benth. | 1 |
| Rubiaceae | Atractocarpus fitzalanii (F. Muell.) Puttock | 2 |
| Rubiaceae | Atractocarpus hirtus (F. Muell.) Puttock | 2 |
| Rutaceae | Melicope stipitata C. T. White & W. D. Francis | 2 |
| Sapindaceae | Harpullia rhyticarpa C. T. White & W. D. Francis | 2 |
| Sapindaceae | Diploglottis harpullioides S. T. Reynolds | 3 |
| Sapindaceae | Sarcotoechia cuneata Radlk. | 3 |
| Sapindaceae | Diploglottis smithii S. T. Reynolds | 5 |
| Sapindaceae | Rhysotoechia robertsonii (F. Muell.) Radlk. | 6 |
| Symplocaceae | Symplocos ampulliformis C. T. White | 5 |
| Family | Species | Infra-Variation (bp) |
|---|---|---|
| Annonaceae | Miliusa traceyi Jessup | 2 |
| Lauraceae | Endiandra longipedicellata C. T. White & W. D. Francis | 5 |
| Malvaceae | Argyrodendron polyandrum L. S. Sm. | 1 |
| Moraceae | Ficus variegata Blume | 2 |
| Myrtaceae | Gossia retusa N. Snow & Guymer | 4 |
| Myrtaceae | Rhodomyrtus macrocarpa Benth. | 4 |
| Myrtaceae | Syzygium fibrosum (F. M. Bailey) T. G. Hartley & L. M. Perry | 1 |
| Phyllanthaceae | Breynia cernua (Poir.) Mull. Arg. | 3 |
| Proteaceae | Helicia australasica F. Muell. | 3 |
| Rhamnaceae | Emmenosperma cunninghamii Benth. | 1 |
| Rubiaceae | Ixora timorensis Decne. | 3 |
| Rutaceae | Bosistoa brassii T. G. Hartley | 9 |
| Rutaceae | Medicosma sessiliflora (C. T. White) T. G. Hartley | 9 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Costion, C.M.; Lowe, A.J.; Rossetto, M.; Kooyman, R.M.; Breed, M.F.; Ford, A.; Crayn, D.M. Building a Plant DNA Barcode Reference Library for a Diverse Tropical Flora: An Example from Queensland, Australia. Diversity 2016, 8, 5. https://doi.org/10.3390/d8010005
Costion CM, Lowe AJ, Rossetto M, Kooyman RM, Breed MF, Ford A, Crayn DM. Building a Plant DNA Barcode Reference Library for a Diverse Tropical Flora: An Example from Queensland, Australia. Diversity. 2016; 8(1):5. https://doi.org/10.3390/d8010005
Chicago/Turabian StyleCostion, Craig M., Andrew J. Lowe, Maurizio Rossetto, Robert M. Kooyman, Martin F. Breed, Andrew Ford, and Darren M. Crayn. 2016. "Building a Plant DNA Barcode Reference Library for a Diverse Tropical Flora: An Example from Queensland, Australia" Diversity 8, no. 1: 5. https://doi.org/10.3390/d8010005
APA StyleCostion, C. M., Lowe, A. J., Rossetto, M., Kooyman, R. M., Breed, M. F., Ford, A., & Crayn, D. M. (2016). Building a Plant DNA Barcode Reference Library for a Diverse Tropical Flora: An Example from Queensland, Australia. Diversity, 8(1), 5. https://doi.org/10.3390/d8010005






