Native Population Structure beyond Hatchery Introgression in the Endemic Sicilian Trout
Abstract
1. Introduction
2. Methods
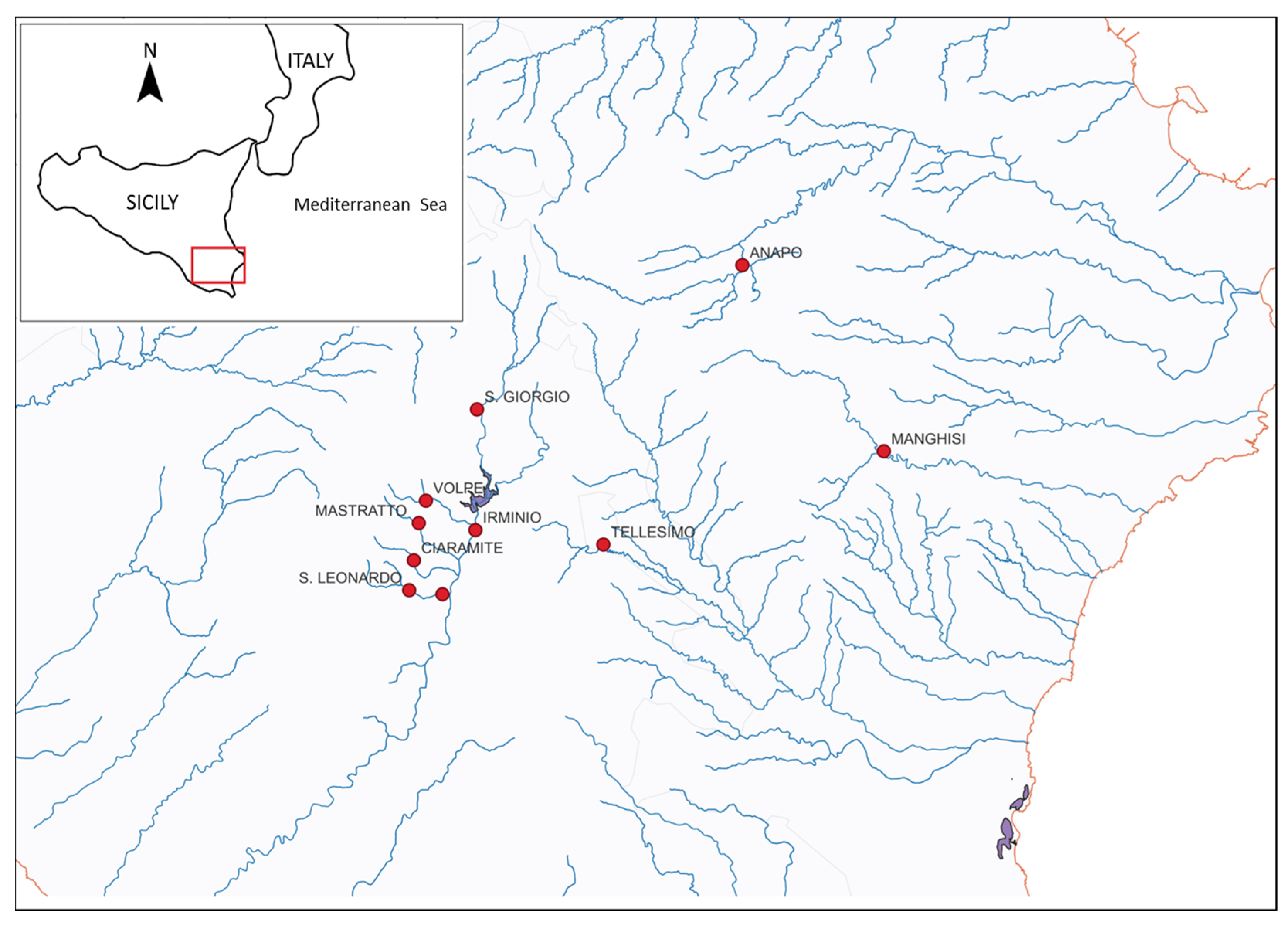
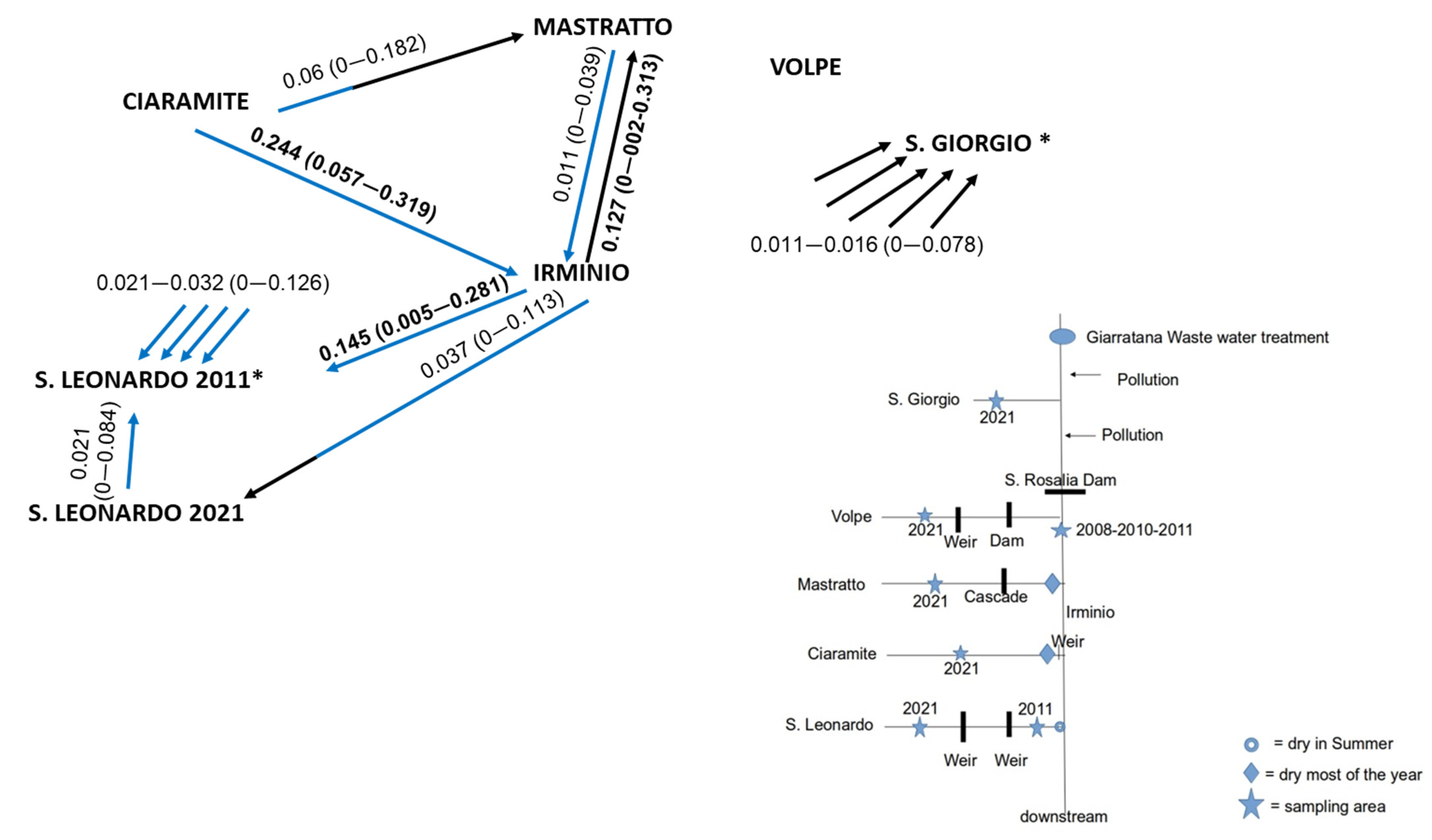
2.1. Studied Area and Sampling
2.2. Microsatellites and LDH-C Genotyping
2.3. Data Analysis from Genotyping
2.4. Mitochondrial DNA Sequencing
3. Results
3.1. Hatchery Introgression and Gene Diversity within Populations
3.2. Temporal Genetic Differentiation
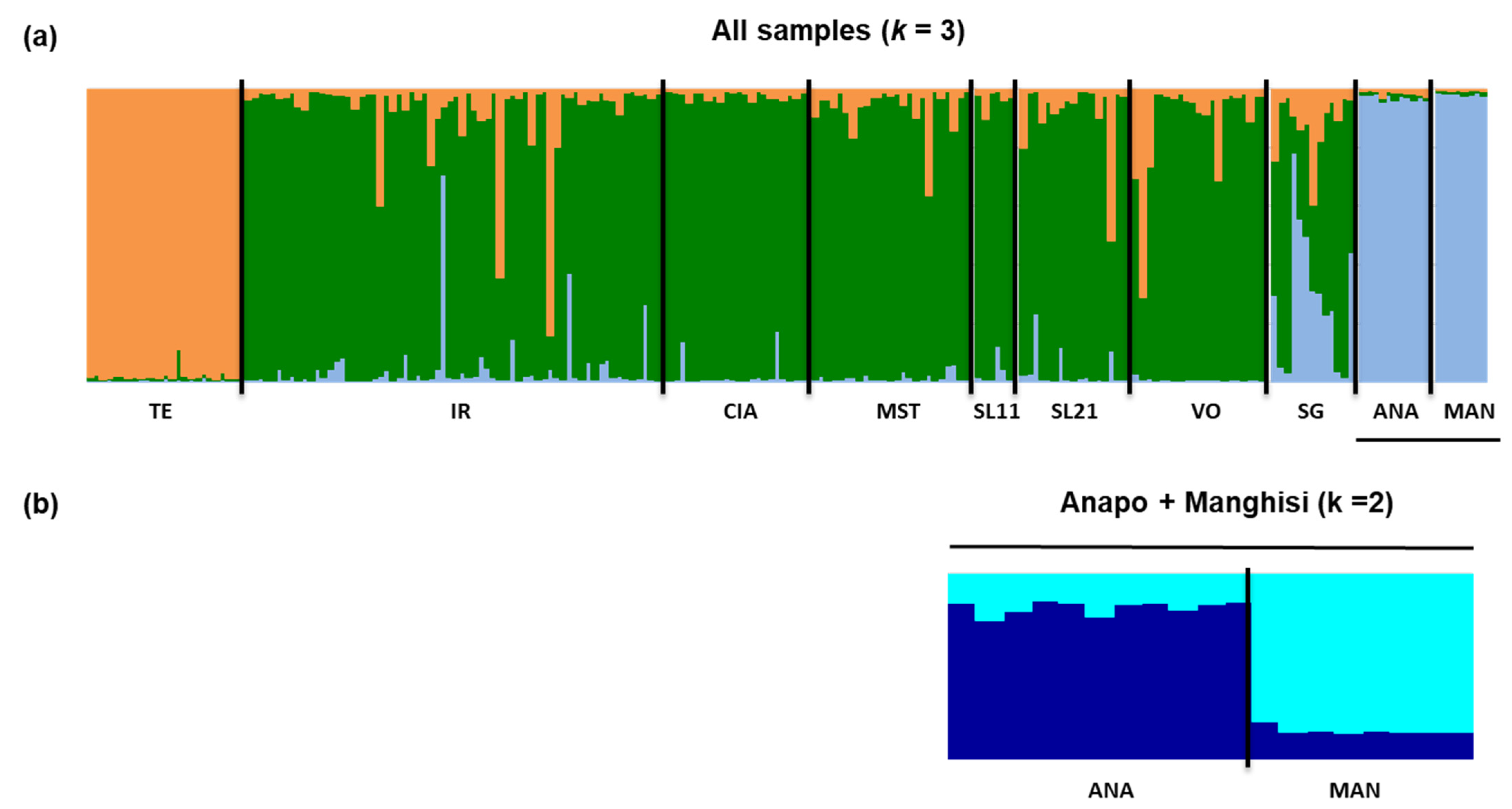
3.3. Genetic Differentiation and Population Structure
3.4. mtDNA-CR Sequencing
4. Discussion
4.1. Hatchery Introgression and Disruption of Gene Diversity
4.2. Population Structure beyond Hatchery Introgression
4.3. Conclusions and Conservation Management Strategies
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Filipe, A.F.; Markovic, D.; Pletterbauer, F.; Tisseuil, C.; De Wever, A.; Schmutz, S.; Bonada, N.; Freyhof, J. Forecasting Fish Distribution along Stream Networks: Brown Trout (Salmo trutta) in Europe. Divers. Distrib. 2013, 19, 1059–1071. [Google Scholar] [CrossRef]
- Mostafavi, H.; Kambouzia, J. Impact of Climate Change on the Distribution of Brown Trout, Salmo trutta Linnaeus, 1758 (Teleostei: Salmonidae) Using Ensemble Modelling Approach in Iran. Iran. J. Ichthyol. 2019, 6, 73–81. [Google Scholar] [CrossRef]
- Zava, B.; Beller, P.; Chiari, P.; Nardi, P.A.; Violani, C.; Bernini, F. Faunistic and taxonomic notes on Salmo trutta macrostigma (Dum.) from Sicily. Atti IV Convegno Naz. A.I.I.A.D. Distrib. Fauna Ittica Ital. Riva Gard 1991, 413–422. [Google Scholar]
- Zava, B.; Beller, P.; Chiari, P.; Nardi, P.A.; Violani, C.; Bernini, F. Salmo cettii Rafinesque-Schmaltz, 1810, an early name for the Sicilian trout. Atti Soc. Ital. Sci. Nat. Mus. Civ. Stor. Nat. Milano 1996, 137, 65–66. [Google Scholar]
- Kottelat, M.; Freyhof, J. Handbook of European Freshwater Fishes; Publications Kottelat: Cornol, Switzerland, 2007. [Google Scholar]
- Duchi, A. Flank Spot Number and Its Significance for Systematics, Taxonomy and Conservation of the near-Threatened Mediterranean Trout Salmo Cettii: Evidence from a Genetically Pure Population. J. Fish Biol. 2018, 92, 254–260. [Google Scholar] [CrossRef]
- Schöffmann, J.; Sušnik, S.; Snoj, A. Phylogenetic Origin of Salmo trutta L 1758 from Sicily, Based on Mitochondrial and Nuclear DNA Analyses. Hydrobiologia 2007, 575, 51–55. [Google Scholar] [CrossRef]
- Fruciano, C.; Pappalardo, A.M.; Tigano, C.; Ferrito, V. Phylogeographical Relationships of Sicilian Brown Trout and the Effects of Genetic Introgression on Morphospace Occupation. Biol. J. Linn. Soc. 2014, 112, 387–398. [Google Scholar] [CrossRef]
- Sanz, N. Phylogeographic History of Brown Trout: A Review. In Brown Trout: Biology, Ecology and Management; John Wiley & Sons Ltd.: Hoboken, NJ, USA, 2018; pp. 17–63. [Google Scholar] [CrossRef]
- Snoj, A.; Marić, S.; Bajec, S.S.; Berrebi, P.; Janjani, S.; Schöffmann, J. Phylogeographic Structure and Demographic Patterns of Brown Trout in North-West Africa. Mol. Phylogenet. Evol. 2011, 61, 203–211. [Google Scholar] [CrossRef]
- Snoj, A.; Bravničar, J.; Marić, S.; Sušnik Bajec, S.; Benaissa, H.; Schöffmann, J. Nuclear DNA Reveals Multiple Waves of Colonisation, Reticulate Evolution and a Large Impact of Stocking on Trout in North-West Africa. Hydrobiologia 2021, 848, 3389–3405. [Google Scholar] [CrossRef]
- Hashemzadeh Segherloo, I.; Freyhof, J.; Berrebi, P.; Ferchaud, A.L.; Geiger, M.; Laroche, J.; Levin, B.A.; Normandeau, E.; Bernatchez, L. A Genomic Perspective on an Old Question: Salmo Trouts or Salmo trutta (Teleostei: Salmonidae)? Mol. Phylogenet. Evol. 2021, 162, 107204. [Google Scholar] [CrossRef]
- Berrebi, P.; Caputo Barucchi, V.; Splendiani, A.; Muracciole, S.; Sabatini, A.; Palmas, F.; Tougard, C.; Arculeo, M.; Marić, S. Brown Trout (Salmo trutta L.) High Genetic Diversity around the Tyrrhenian Sea as Revealed by Nuclear and Mitochondrial Markers. Hydrobiologia 2019, 826, 209–231. [Google Scholar] [CrossRef]
- Lobón-Cerviá, J.; Esteve, M.; Berrebi, P.; Duchi, A.; Lorenzoni, M.; Young, K.A. Trout and Char of Central and Southern Europe and Northern Africa. In Trout and Char of the World; Kershner, J., Williams, J., Gresswell, R., Lobón-Cerviá, J., Eds.; American Fisheries Society: Atlantic City, NJ, USA, 2019; pp. 379–410. [Google Scholar] [CrossRef]
- Freyhof, J.; Kottelat, M. Salmo cettii. IUCN Red List of Threatened Species Version 3.1. 2008. Available online: http://www.iucnredlist.org/details/135528/0 (accessed on 1 June 2022).
- Freyhof, J.; Bergner, L.; Ford, M. Threatened Fishes of the Mediterranean Basin Biodiversity Hotspot: Distribution, Extinction risk and impact of hydropower. EuroNatur RiverWatch 2020, I-viii, 1–348. [Google Scholar] [CrossRef]
- Rondinini, C.; Battistoni, A.; Peronace, V.; Teofili, C. Lista Rossa Dei Vertebrati Italiani. WWF Ital. Sett. Divers. Biol. 2013, 56, 56. [Google Scholar]
- Duchi, A. Preliminary data on the trouts of Torrent Tellesimo (Ragusa, Sicilia). Atti Soc. Ital. Sci. Nat. Mus. Civ. Stor. Nat. 1988, 129, 167–172. (In Italian) [Google Scholar]
- Duchi, A. Fecundity, Egg and Alevin Size in the River Irminio Population of the Threatened Salmo Cettii Rafinesque-Schmaltz, 1810 (Sicily, Italy). J. Appl. Ichthyol. 2011, 27, 868–872. [Google Scholar] [CrossRef]
- Duchi, A.; Occhipinti, G. Produzione sperimentale di novellame di trota macrostigma, Salmo (trutta) macrostigma Dum, in provincia di Ragusa (Sicilia). Atti V Conv. AIIAD-Montecchio Magg. (VI) 1994, 379–386. (In Italian) [Google Scholar]
- Bobbio, L.; Cannas, R.; Cau, A.; Deiana, A.M.; Duchi, A.; Gandolfi, G.; Tagliavini, J. Variabilità mitocondriale in specie di salmonidi, con particolare riferimento alle forme “macrostigma”. Atti Del VI Convegno Dell’ A.I.I.A.D. Varese Ligure 1996, 42–49. [Google Scholar]
- Sanz, N.; Araguas, R.M.; Fernández, R.; Vera, M.; García-Marín, J.L. Efficiency of Markers and Methods for Detecting Hybrids and Introgression in Stocked Populations. Conserv. Genet. 2009, 10, 225–236. [Google Scholar] [CrossRef]
- McMeel, O.M.; Hoey, E.M.; Ferguson, A. Partial Nucleotide Sequences, and Routine Typing by Polymerase Chain Reaction-Restriction Fragment Length Polymorphism, of the Brown Trout (Salmo trutta) Lactate Dehydrogenase, LDH-C1*90 And*100 Alleles. Mol. Ecol. 2001, 10, 29–34. [Google Scholar] [CrossRef]
- Van Oosterhout, C.; Hutchinson, W.F.; Wills, D.P.M.; Shipley, P. Micro-Checker: Software for Identifying and Correcting Genotyping Errors in Microsatellite Data. Mol. Ecol. Notes 2004, 4, 535–538. [Google Scholar] [CrossRef]
- Goudet, J. FSTAT (Version 1.2): A Computer Program to Calculate F-Statistics. J. Hered. 1995, 86, 485–486. [Google Scholar] [CrossRef]
- Kalinowski, S.T. HP-RARE 1.0: A computer program for performing rarefaction on measures of allelic richness. Mol. Ecol. Notes 2005, 5, 187–189. [Google Scholar] [CrossRef]
- Raymond, M.; Rousset, F. GENEPOP (version 1.2): Population genetics software for exact tests and ecumenicism. J. Hered. 1995, 86, 248–249. [Google Scholar] [CrossRef]
- Rice, W.R. Analyzing Tables of Statistical Tests. Evolution 1989, 43, 223–225. [Google Scholar] [CrossRef]
- Cornuet, J.M.; Luikart, G. Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 1996, 144, 2001–2014. [Google Scholar] [CrossRef]
- Dieringer, D.; Schlötterer, C. Microsatellite analyser (MSA): A platform independent analysis tool for large microsatellite data sets. Mol. Ecol. Notes 2003, 3, 167–169. [Google Scholar] [CrossRef]
- Wilson, G.A.; Rannala, B. Bayesian inference of recent migration rates using multilocus genotypes. Genetics 2003, 163, 1177–1191. [Google Scholar] [CrossRef]
- Falush, D.; Stephens, M.; Pritchard, J.K. Inference of population structure using multilocus genotype data: Dominant markers and null alleles. Mol. Ecol. Notes 2007, 7, 574–578. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Earl, D.A.; VonHoldt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Jombart, T. Adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020; ISBN 3-900051-07-0. Available online: https://www.R-project.org/ (accessed on 20 June 2022).
- Jombart, T.; Devillard, S.; Balloux, F. Discriminant Analysis of Principal Components: A New Method for the Analysis of Genetically Structured Populations. BMC Genet. 2010, 11, 94. [Google Scholar] [CrossRef]
- Excoffier, L.; Laval, G.; Schneider, S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol. Bioinform. 2005, 1, 47–50. [Google Scholar] [CrossRef]
- Cortey, M.; García-Marín, J.L. Evidence for Phylogeographically Informative Sequence Variation in the Mitochondrial Control Region of Atlantic Brown Trout. J. Fish Biol. 2002, 60, 1058–1063. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Librado, T.; Rozas, J. DnaSP v5: A software for comprehensive analyses of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef]
- Suárez, J.; Bautista, J.M.; Almodóvar, A.; Machordom, A. Evolution of the Mitochondrial Control Region in Palaearctic Brown Trout (Salmo trutta) Populations: The Biogeographical Role of the Iberian Peninsula. Heredity 2001, 87, 198–206. [Google Scholar] [CrossRef]
- Sommani, E.; Sommani, E. Osservazioni sulla sistematica ed ecologia delle trote nell’Italia meridionale. Boll. Pesca Idrobiol. 1950, 5, 170–187. [Google Scholar]
- Duchi, A. Extant Because Important or Important Because Extant? On the Scientific Importance and Conservation of a Genetically Pure Sicilian Population of the Threatened Salmo cettii Rafinesque, 1810. Cybium 2020, 44, 41–44. [Google Scholar] [CrossRef]
- Sanz, N.; Cortey, M.; Pla, C.; García-Marín, J.L. Hatchery Introgression Blurs Ancient Hybridization between Brown Trout (Salmo trutta) Lineages as Indicated by Complementary Allozymes and MtDNA Markers. Biol. Conserv. 2006, 130, 278–289. [Google Scholar] [CrossRef]
- Splendiani, A.; Ruggeri, P.; Giovannotti, M.; Caputo Barucchi, V. Role of Environmental Factors in the Spread of Domestic Trout in Mediterranean Streams. Freshw. Biol. 2013, 58, 2089–2101. [Google Scholar] [CrossRef]
- Hansen, M.M.; Ruzzante, D.E.; Nielsen, E.E.; Mensberg, K.L.D. Microsatellite and Mitochondrial DNA Polymorphism Reveals Life-History Dependent Interbreeding between Hatchery and Wild Brown Trout (Salmo trutta L.). Mol. Ecol. 2000, 9, 583–594. [Google Scholar] [CrossRef] [PubMed]
- Vera, M.; Garcia-Marin, J.L.; Martinez, P.; Araguas, R.M.; Bouza, C. Identification and Conservation of Remnant Genetic Resources of Brown Trout in Relict Populations from Western Mediterranean Streams. Hydrobiologia 2013, 707, 29–45. [Google Scholar] [CrossRef]
- Sanz, N.; Fernández-Cebrián, R.; Casals, F.; Araguas, R.M.; García-Marín, J.L. Dispersal and Demography of Brown Trout, Salmo trutta, Inferred from Population and Family Structure in Unstable Mediterranean Streams. Hydrobiologia 2011, 671, 105–119. [Google Scholar] [CrossRef]
- Berrebi, P. Three Brown Trout Salmo trutta Lineages in Corsica Described through Allozyme Variation. J. Fish Biol. 2015, 86, 60–73. [Google Scholar] [CrossRef]
- Araguas, R.M.; Vera, M.; Aparicio, E.; Sanz, N.; Fernández-Cebrián, R.; Marchante, C.; García-Marín, J.L. Current Status of the Brown Trout (Salmo trutta) Populations within Eastern Pyrenees Genetic Refuges. Ecol. Freshw. Fish 2017, 26, 120–132. [Google Scholar] [CrossRef]
- Sanz, N.; Araguas, R.M.; Fernández-Cebrián, R.; Lobón-Cerviá, J. Factors modelling population structure in Brown trout Salmo trutta L.: Genetic monitoring of populations in Esva River (northwestern Spain). Hydrobiologia 2019, 837, 117–131. [Google Scholar] [CrossRef]
- García-Marín, J.L.; Utter, F.; Pla, C. Postglacial Colonization of Brown Trout in Europa Based on Distribuction of Allozyme Variants. Heredity 1999, 82, 46–56. [Google Scholar] [CrossRef]
- Pujolar, J.M.; Lucarda, A.N.; Simonato, M.; Patarnello, T. Restricted Gene Flow at the Micro- and Macro-Geographical Scale in Marble Trout Based on MtDNA and Microsatellite Polymorphism. Front. Zool. 2011, 8, 7. [Google Scholar] [CrossRef]
- Thaulow, J.; Borgstrøm, R.; Heun, M. Brown Trout Population Structure Highly Affected by Multiple Stocking and River Diversion in a High Mountain National Park. Conserv. Genet. 2013, 14, 145–158. [Google Scholar] [CrossRef]
- Klütsch, C.F.C.; Maduna, S.N.; Polikarpova, N.; Forfang, K.; Aspholm, P.E.; Nyman, T.; Eiken, H.G.; Amundsen, P.A.; Hagen, S.B. Genetic Changes Caused by Restocking and Hydroelectric Dams in Demographically Bottlenecked Brown Trout in a Transnational Subarctic Riverine System. Ecol. Evol. 2019, 9, 6068–6081. [Google Scholar] [CrossRef]
- Vera, M.; Martinez, P.; Bouza, C. Stocking Impact, Population Structure and Conservation of Wild Brown Trout Populations in Inner Galicia (NW Spain), an Unstable Hydrologic Region. Aquat. Conserv. Mar. Freshw. Ecosyst. 2018, 28, 435–443. [Google Scholar] [CrossRef]
- Sanz, N.; García-Marín, J.-L.; Pla, C. Divergence of Brown Trout (Salmo trutta) within Glacial Refugia. Can. J. Fish. Aquat. Sci. 2000, 57, 2201–2210. [Google Scholar] [CrossRef]
- Griffiths, A.M.; Bright, D.; Stevens, J.R.; Hashemzadeh Segherloo, I.; Farahmand, H.; Abdoli, A.; Bernatchez, L.; Primmer, C.R.; Swatdipong, A.; Karami, M.; et al. Phylogenetic Status of Brown Trout Salmo trutta Populations in Five Rivers from the Southern Caspian Sea and Two Inland Lake Basins, Iran: A Morphogenetic Approach. J. Fish Biol. 2009, 81, 1136–1142. [Google Scholar] [CrossRef]
- Vera, M.; Sanz, N.; Hansen, M.M.; Almodóvar, A.; García-Marín, J.-L. Population and Family Structure of Brown Trout, Salmo trutta, in a Mediterranean Stream. Mar. Freshw. Res. 2010, 61, 672–681. [Google Scholar] [CrossRef]
- Fumagalli, L.; Snoj, A.; Jesenšek, D.; Balloux, F.; Jug, T.; Duron, O.; Brossier, F.; Crivelli, A.J.; Berrebi, P. Extreme Genetic Differentiation among the Remnant Populations of Marble Trout (Salmo marmoratus) in Slovenia. Mol. Ecol. 2002, 11, 2711–2716. [Google Scholar] [CrossRef]
- Ayllon, F.; Moran, P.; Garcia-Vazquez, E. Maintenance of a Small Anadromous Subpopulation of Brown Trout (Salmo trutta L.) by Straying. Freshw. Biol. 2006, 51, 351–358. [Google Scholar] [CrossRef]
- Lobón-Cerviá, J. Why, When and How Do Fish Populations Decline, Collapse and Recover? The Example of Brown Trout (Salmo trutta) in Rio Chaballos (Northwestern Spain). Freshw. Biol. 2009, 54, 1149–1162. [Google Scholar] [CrossRef]
- García-Marín, J.L.; Sanz, N.; Pla, C. Proportions of Native and Introduced Brown Trout in Adjacent Fished and Unfished Spanish Rivers. Conserv. Biol. 1998, 12, 313–319. [Google Scholar] [CrossRef]
- Hansen, M.M. Estimating the Long-Term Effects of Stocking Domesticated Trout into Wild Brown Trout (Salmo trutta) Populations: An Approach Using Microsatellite DNA Analysis of Historical and Contemporary Samples. Mol. Ecol. 2002, 11, 1003–1015. [Google Scholar] [CrossRef]
- Hansen, M.M.; Nielsen, E.E.; Ruzzante, D.E.; Bouza, C.; Mensberg, K.-L.D. Genetic monitoring of supportive breeding in brown trout (Salmo trutta L.), using microsatellite DNA markers. Can. J. Fish. Aquat. Sci. 2000, 57, 2130–2139. [Google Scholar] [CrossRef]
- Meraner, A.; Baric, S.; Pelster, B.; Dalla Via, J. Microsatellite DNA Data Point to Extensive but Incomplete Admixture in a Marble and Brown Trout Hybridisation Zone. Conserv. Genet. 2010, 11, 985–998. [Google Scholar] [CrossRef]
- Marić, S.; Sušnik Bajec, S.; Schöffmann, J.; Kostov, V.; Snoj, A. Phylogeography of Stream-Dwelling Trout in the Republic of Macedonia and a Molecular Genetic Basis for Revision of the Taxonomy Proposed by S. Karaman. Hydrobiologia 2016, 785, 249–260. [Google Scholar] [CrossRef]
- Wofford, J.E.B.; Gresswell, R.E.; Banks, M.A. Influence of Barriers to Movement on Within-Watershed Genetic Variation of Coastal Cutthroat Trout. Ecol. Appl. 2005, 15, 628–637. [Google Scholar] [CrossRef]
- Raeymaekers, J.A.M.; Raeymaekers, D.; Koizumi, I.; Geldof, S.; Volckaert, F.A.M. Guidelines for Restoring Connectivity around Water Mills: A Population Genetic Approach to the Management of Riverine Fish. J. Appl. Ecol. 2009, 46, 562–571. [Google Scholar] [CrossRef]
- Birnie-Gauvin, K.; Larsen, M.H.; Nielsen, J.; Aarestrup, K. 30 Years of Data Reveal Dramatic Increase in Abundance of Brown Trout Following the Removal of a Small Hydrodam. J. Environ. Manag. 2017, 204, 467–471. [Google Scholar] [CrossRef]

| Sample (Code) | N | A (S.E.) | Ar (S.E.) | PA (freq > 0.1) | PAr (S.E.) | HS (S.E.) | q | HAT mtDNA* |
|---|---|---|---|---|---|---|---|---|
| Tellesimo (TE) | 24 | 2.125 (0.398) | 1.763 (0.257) | 0 | 0.12 (0.092) | 0.228 (0.077) | 0 | 0 (13) |
| Irminio 08 (IR08) | 10 | 4.5 (0.681) | 3.749 (0.523) | 0 | 0.23 (0.096) | 0.582 (0.096) | 0.350 | no data |
| Irminio 10 (IR10) | 33 | 6.125 (1.202) | 3.798 (0.524) | 1 (0) | 0.07 (0.031) | 0.575 (0.090) | 0.167 | 0.150 (20) |
| Irminio 11 (IR11) | 23 | 5.25 (0.921) | 3.788 (0.533) | 1 (0) | 0.1 (0.090) | 0.587 (0.094) | 0.196 | 0.176 (17) |
| Ciaramite (CIA) | 22 | 3.25 (0.526) | 2.832 (0.387) | 3 (0) | 0.02 (0.013) | 0.536 (0.082) | 0.409 | 0 (10) |
| Mastratto (MST) | 25 | 5.375 (0.905) | 3.611 (0.506) | 3 (0) | 0.27 (0.157) | 0.558 (0.091) | 0.333 | 0.25 (12) |
| San Leonardo 11 (SL11) | 6 | 3.25 (0.590) | 3.250 (0.590) | 0 | 0.02 (0.012) | 0.465 (0.105) | 0.167 | 0 (5) |
| San Leonardo 21 (SL21) | 17 | 4.25 (0.559) | 3.701 (0.375) | 1 (0) | 0.05 (0.044) | 0.507 (0.082) | 0.176 | 0 (7) |
| Volpe (VO) | 21 | 3.25 (0.559) | 2.666 (0.373) | 2 (0) | 0.13 (0.133) | 0.458 (0.085) | 0.265 | 0 (12) |
| San Giorgio (SG) | 13 | 3.25 (0.13) | 2.923 (0.260) | 1 (1) | 0.15 (0.121) | 0.512 (0.064) | 0.385 | 0.3 (10) |
| Irminio River | 170 | 4.80 (0.37) | 3.299 (0.393) | 10 (1) | 0.55 (0.185) | 0.531 (0.0116) | 0.272 | 0.129 (93) |
| Anapo (ANA) | 11 | 4.625 (0.498) | 3.839 (0.374) | 6 (4) | 0.73 (0.299) | 0.607 (0.061) | 0.227 | 0 (9) |
| Manghisi (MAN) | 8 | 4.625 (0.754) | 4.092 (0.613) | 3 (1) | 0.43 (0.229) | 0.567 (0.070) | 0.125 | 0 (7) |
| TE | IR208 | IR10 | IR11 | CIA | MST | SL11 | SL21 | VO | SG | ANA | MAN | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| TE | - | 0.395 | 0.398 | 0.412 | 0.418 | 0.388 | 0.428 | 0.402 | 0.388 | 0.432 | 0.566 | 0.691 |
| IR08 | 0.215 *** | - | 0.225 | 0.215 | 0.278 | 0.235 | 0.402 | 0.324 | 0.306 | 0.401 | 0.534 | 0.600 |
| IR10 | 0.169 *** | 0.005 ns | - | 0.146 | 0.251 | 0.203 | 0.337 | 0.244 | 0.281 | 0.366 | 0.492 | 0.548 |
| IR11 | 0.186 *** | −0.012 ns | 0.004 ns | - | 0.273 | 0.209 | 0.340 | 0.240 | 0.270 | 0.386 | 0.519 | 0.586 |
| CIA | 0.293 *** | 0.042 * | 0.051 *** | 0.053 *** | - | 0.306 | 0.418 | 0.279 | 0.328 | 0.433 | 0.556 | 0.646 |
| MST | 0.184 *** | −0.001 ns | 0.013 * | 0.012 ns | 0.077 *** | - | 0.363 | 0.307 | 0.291 | 0.390 | 0.533 | 0.601 |
| SL11 | 0.361 *** | 0.134 ** | 0.079 *** | 0.095 *** | 0.178 *** | 0.117 *** | - | 0.332 | 0.353 | 0.404 | 0.590 | 0.614 |
| SL21 | 0.237 *** | 0.056 ** | 0.033 *** | 0.025 ** | 0.068 *** | 0.065 *** | 0.112 *** | - | 0.282 | 0.370 | 0.523 | 0.595 |
| VO | 0.258 *** | 0.063 ** | 0.068 *** | 0.066 *** | 0.124 *** | 0.074 *** | 0.142 *** | 0.097 *** | - | 0.367 | 0.575 | 0.669 |
| SG | 0.294 *** | 0.090 *** | 0.060 *** | 0.080 *** | 0.137 *** | 0.098 *** | 0.107 ** | 0.093 *** | 0.119 *** | - | 0.512 | 0.527 |
| ANA | 0.453 *** | 0.253 *** | 0.240 *** | 0.253 *** | 0.267 *** | 0.273 *** | 0.306 *** | 0.280 *** | 0.356 *** | 0.259 *** | - | 0.483 |
| MAN | 0.571 *** | 0.326 *** | 0.300 *** | 0.317 *** | 0.347* ** | 0.334 *** | 0.338 *** | 0.350 *** | 0.428 *** | 0.284 *** | 0.159 *** | - |
| Structure Tested (Samples Grouped) | Variance Component | % of Total | Fixation Index |
|---|---|---|---|
| Temporal (IR08 + IR10 + IR11) | |||
| Among groups | 0.324 | 15.41 | FCT = 0.154 *** |
| Within groups | 0.052 | 0.38 | FSC = 0.005 ns |
| k = 3, STRUCTURE (Irminio basin; ANA + MAN) | |||
| Among groups | 0.512 | 19.05 | FCT = 0.191 ** |
| Within groups | 0.159 | 5.93 | FSC = 0.073 *** |
| k = 4, STRUCTURE (Irminio basin) | |||
| Among groups | 0.556 | 20.52 | FCT = 0.205 ** |
| Within groups | 0.139 | 5.16 | FSC = 0.065 *** |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sanz, N.; Araguas, R.-M.; Giampiccolo, M.; Duchi, A. Native Population Structure beyond Hatchery Introgression in the Endemic Sicilian Trout. Diversity 2023, 15, 274. https://doi.org/10.3390/d15020274
Sanz N, Araguas R-M, Giampiccolo M, Duchi A. Native Population Structure beyond Hatchery Introgression in the Endemic Sicilian Trout. Diversity. 2023; 15(2):274. https://doi.org/10.3390/d15020274
Chicago/Turabian StyleSanz, Nuria, Rosa-Maria Araguas, Monica Giampiccolo, and Antonino Duchi. 2023. "Native Population Structure beyond Hatchery Introgression in the Endemic Sicilian Trout" Diversity 15, no. 2: 274. https://doi.org/10.3390/d15020274
APA StyleSanz, N., Araguas, R.-M., Giampiccolo, M., & Duchi, A. (2023). Native Population Structure beyond Hatchery Introgression in the Endemic Sicilian Trout. Diversity, 15(2), 274. https://doi.org/10.3390/d15020274






