An Incomplete European Barcode Library Has a Strong Impact on the Identification Success of Lepidoptera from Greece
Abstract
:1. Introduction
2. Materials and Methods
3. Results
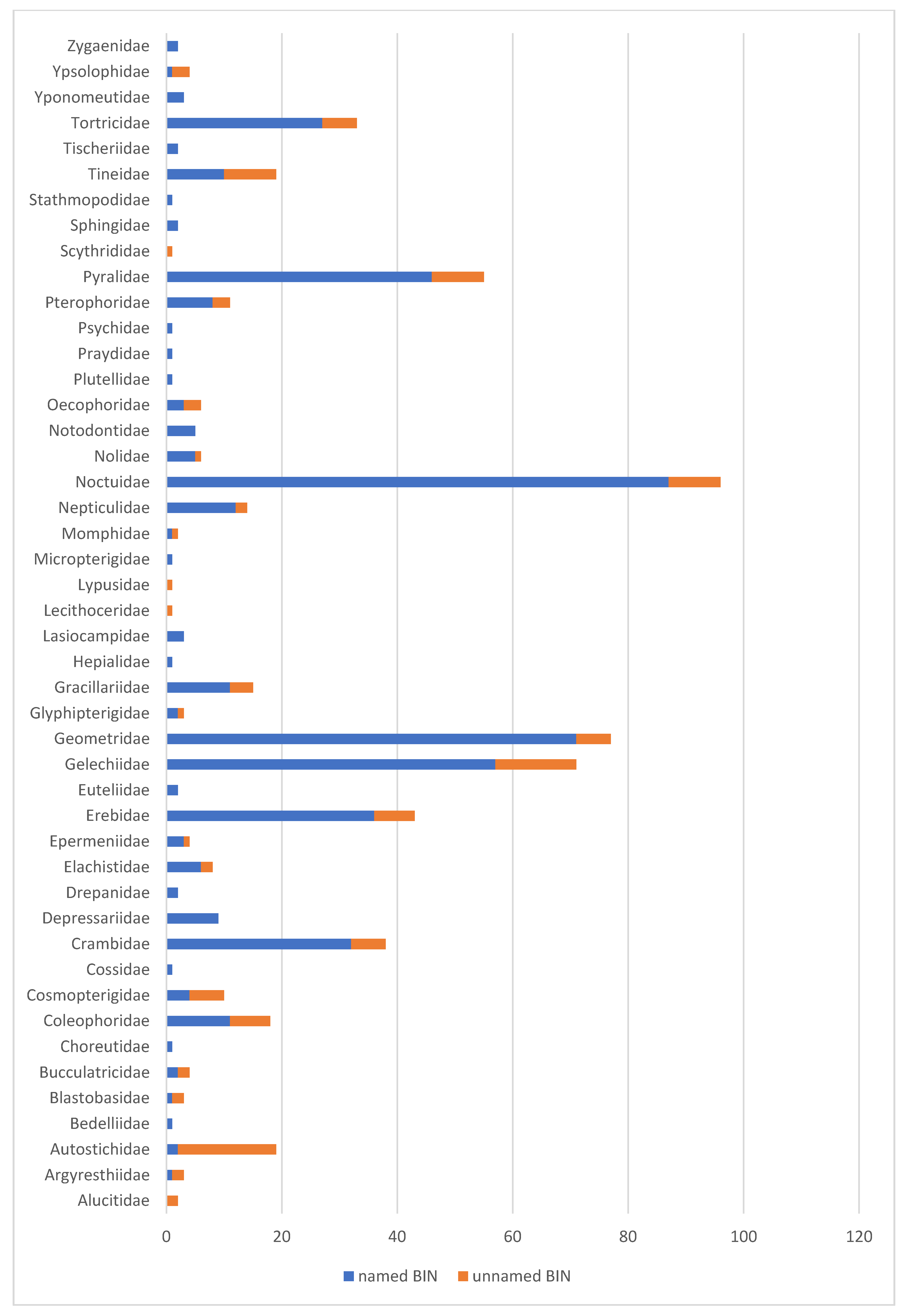
3.1. General Overview
3.2. Unidentified BINs—Incomplete Barcode Library
3.3. Potential Cryptic Diversity
3.4. BINs Attributed to Linnaean Names
3.5. New Faunistic Records
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Roslin, T.; Somervuo, P.; Pentinsaari, M.; Hebert, P.D.N.; Agda, J.; Ahlroth, P.; Anttonen, P.; Aspi, J.; Blagoev, G.; Blanco, S.; et al. A molecular-based identification resource for the arthropods of Finland. Mol. Ecol. Res. 2021, 22, 803–822. [Google Scholar] [CrossRef] [PubMed]
- Hausmann, A.; Haszprunar, G.; Segerer, A.H.; Speidel, W.; Behounek, G.; Hebert, P.D.N. Now DNA barcoded: The butterflies and larger moths of Germany (Lepidoptera: Rhopalocera, Macroheterocera). Spixiana 2011, 34, 47–58. [Google Scholar]
- Huemer, P.; Mutanen, M.; Sefc, K.M.; Hebert, P.D.N. Testing DNA Barcode Performance in 1000 Species of European Lepidoptera: Large Geographic Distances Have Small Genetic Impacts. PLoS ONE 2014, 9, e115774. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huemer, P.; Wiesmair, B. DNA-Barcoding der Tagfalter (Lepidoptera, Papilionoidea) Österreichs—unbekannte genetische Vielfalt im Zentrum Europas. Wiss. Jahrb. Tirol. Landesmuseen 2017, 10, 8–33. [Google Scholar]
- Huemer, P.; Wieser, C.; Stark, W.; Hebert, P.D.N.; Wiesmair, B. DNA barcode library of megadiverse Austrian Noctuoidea (Lepidoptera)—a nearly perfect match of Linnean taxonomy. Biodivers. Data J. 2019, 7, e37734. [Google Scholar] [CrossRef] [PubMed]
- Huemer, P.; Hebert, P.D.N. DNA Barcode Bibliothek der Schmetterlinge Südtirols und Tirols (Italien, Österreich)—Impetus für integrative Artdifferenzierung im 21. Jahrhundert. Gredleriana 2016, 16, 141–164. [Google Scholar]
- Huemer, P.; Karsholt, O. Commented checklist of European Gelechiidae (Lepidoptera). ZooKeys 2020, 921, 65–140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huemer, P.; Karsholt, O.; Aarvik, L.; Berggren, K.; Bidzilya, O.; Junnilainen, J.; Landry, J.-F.; Mutanen, M.; Nupponen, K.; Segerer, A.; et al. DNA barcode library for European Gelechiidae (Lepidoptera) suggests greatly underestimated species diversity. ZooKeys 2020, 921, 141–157. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Vaamonde, C.; Kirichenko, N.; Cama, A.; Doorenweerd, C.; Godfray, H.C.J.; Guiguet, A.; Gomboc, S.; Huemer, P.; Landry, J.-F.; Laštůvka, A.; et al. Evaluating DNA Barcoding for Species Identification and Discovery in European Gracillariid Moths. Front. Ecol. Evol. 2021, 9, 626752. [Google Scholar] [CrossRef]
- Hausmann, A.; Godfray, H.C.J.; Huemer, P.; Mutanen, M.; Rougerie, R.; van Nieukerken, E.J.; Ratnasingham, S.; Hebert, P.D.N. Genetic Patterns in European Geometrid Moths Revealed by the Barcode Index Number (BIN) System. PLoS ONE 2013, 8, e84518. [Google Scholar] [CrossRef]
- Dinca, V.; Dapporto, L.; Somervuo, P.; Vodă, R.; Cuvelier, S.; Gascoigne-Pees, M.; Huemer, P.; Mutanen, M.; Hebert, P.D.N.; Vila, R. High resolution DNA barcode library for European butterflies reveals continental patterns of mitochondrial genetic diversity. Comm. Biol. 2021, 4, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Mutanen, M.; Kivelä, S.M.; Vos, R.A.; Doorenweerd, C.; Ratnasingham, S.; Hausmann, A.; Huemer, P.; Dincă, V.; van Nieukerken, E.J.; Lopez-Vaamonde, C.; et al. Species-Level Para- and Polyphyly in DNA Barcode Gene Trees: Strong Operational Bias in European Lepidoptera. Syst. Biol. 2016, 65, 1024–1040. [Google Scholar] [CrossRef] [PubMed]
- Ratnasingham, S.; Hebert, P.D.N. BOLD: The Barcode of Life Data System. Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lepiforum: Website zur Bestimmung von Schmetterlingen (Lepidoptera) und Ihren Präimaginalstadien. Available online: http://www.lepiforum.eu/bh/downloads/2021-iii-20/Lepiforums-Europaliste_Schmetterlinge_Version_8.3_Stand_2021_03_20 (accessed on 28 December 2021).
- De Waard, J.R.; Ivanova, N.V.; Hajibabaei, M.; Hebert, P.D.N. Assembling DNA barcodes: Analytical protocols. In Methods in Molecular Biology: Environmental Genomics; Martin, C.C., Ed.; Humana Press Inc.: Totowa, NJ, USA, 2008; pp. 275–293. [Google Scholar]
- Ortiz, A.S.; Rubio, R.M.; Guerrero, J.J.; Garre, M.J.; Serrano, J.; Hebert, P.D.N.; Hausmann, A. Close congruence between Barcode Index Numbers (Bins) and species boundaries in the Erebidae (Lepidoptera: Noctuoidea) of the Iberian Peninsula. Biodiv. Data J. 2017, 5, e19840. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ratnasingham, S.; Hebert, P.D.N. A DNA-based registry for all animal species: The Barcode Index Number (BIN) System. PLoS ONE 2013, 8, e66213. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gozmány, L. The Lepidoptera of Greece and Cyprus. Fauna Graecia IX; Hellenic Zoological Society: Athens, Greece, 2012; Volume 1. [Google Scholar]
- Karsholt, O.; van Nieukerken, E.J. Lepidoptera, Moths. Fauna Europaea 2013, Version 2021.12. Available online: https://fauna-eu.org (accessed on 1 December 2021).
- Gaytán, Á.; Bergsten, J.; Canelo, T.; Pérez-Izquierdo, C.; Santoro, M.; Bonal, R. DNA Barcoding and geographical scale effect: The problems of undersampling genetic diversity hotspots. Ecol. Evol. 2020, 10, 10754–10772. [Google Scholar] [CrossRef] [PubMed]
| Taxon | Family |
|---|---|
| Coleophora helgada (Anikin, 2005) | Coleophoridae |
| Coleophora gardesanella Toll, 1953 | Coleophoridae |
| Eteobalea siciliae (Riedl, 1966) | Cosmopterigidae |
| Agriphila brioniellus (Zerny, 1914) | Crambidae |
| Friedlanderia cicatricella (Hübner, 1824) | Crambidae |
| Spoladea recurvalis (Fabricius, 1775) | Crambidae |
| Elachista argentella (Clerck, 1759) | Elachistidae |
| Elachista atricomella Stainton, 1849 | Elachistidae |
| Elachista biatomella (Stainton, 1848) | Elachistidae |
| Elachista chrysodesmella Zeller, 1850 | Elachistidae |
| Elachista obliquella Stainton, 1854 | Elachistidae |
| Anarsia leberonella Réal, 1994 | Gelechiidae |
| Aproaerema sangiella (Stainton, 1863) | Gelechiidae |
| Aristotelia subdecurtella (Stainton, 1859) | Gelechiidae |
| Ivanauskiella occitanica (Nel & Varenne, 2013) | Gelechiidae |
| Mesophleps oxycedrella (Millière, 1871) | Gelechiidae |
| Scrobipalpa superstes Povolný, 1977 | Gelechiidae |
| Thiotricha subocellea (Stephens, 1834) | Gelechiidae |
| Eupithecia ultimaria Boisduval, 1840 | Geometridae |
| Stigmella obliquella (Heinemann, 1862) | Nepticulidae |
| Bryophila felina (Eversmann, 1852) | Noctuidae |
| Batia inexpectella Jäckh, 1972 | Oecophoridae |
| Acrobasis bithynella Zeller, 1848 | Pyralidae |
| Acrobasis fallouella (Ragonot, 1871) | Pyralidae |
| Assara concicolella (Constant, 1884) | Pyralidae |
| Ceutolopha isidis (Zeller, 1867) | Pyralidae |
| Phycita torrenti Agenjo, 1962 | Pyralidae |
| Tischeria decidua Wocke, 1876 | Tischeriidae |
| Clepsis burgasiensis (Rebel, 1916) | Tortricidae |
| Cydia rymarczyki Varenne & Nel, 2013 | Tortricidae |
| Lobesia bicinctana (Duponchel, 1844) | Tortricidae |
| Neocochylis dubitana (Hübner, 1799) | Tortricidae |
| Pammene herrichiana (Heinemann, 1854) | Tortricidae |
| Ypsolopha alpella (Denis & Schiffermüller, 1775) | Ypsolophidae |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huemer, P.; Mutanen, M. An Incomplete European Barcode Library Has a Strong Impact on the Identification Success of Lepidoptera from Greece. Diversity 2022, 14, 118. https://doi.org/10.3390/d14020118
Huemer P, Mutanen M. An Incomplete European Barcode Library Has a Strong Impact on the Identification Success of Lepidoptera from Greece. Diversity. 2022; 14(2):118. https://doi.org/10.3390/d14020118
Chicago/Turabian StyleHuemer, Peter, and Marko Mutanen. 2022. "An Incomplete European Barcode Library Has a Strong Impact on the Identification Success of Lepidoptera from Greece" Diversity 14, no. 2: 118. https://doi.org/10.3390/d14020118
APA StyleHuemer, P., & Mutanen, M. (2022). An Incomplete European Barcode Library Has a Strong Impact on the Identification Success of Lepidoptera from Greece. Diversity, 14(2), 118. https://doi.org/10.3390/d14020118






