Genetic Diversity in 19th Century Barley (Hordeum vulgare) Reflects Differing Agricultural Practices and Seed Trade in Jämtland, Sweden
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Material
2.2. Genetic Analysis
2.3. Data Analysis
3. Results
3.1. Genotyping of Historical and Extant Accessions
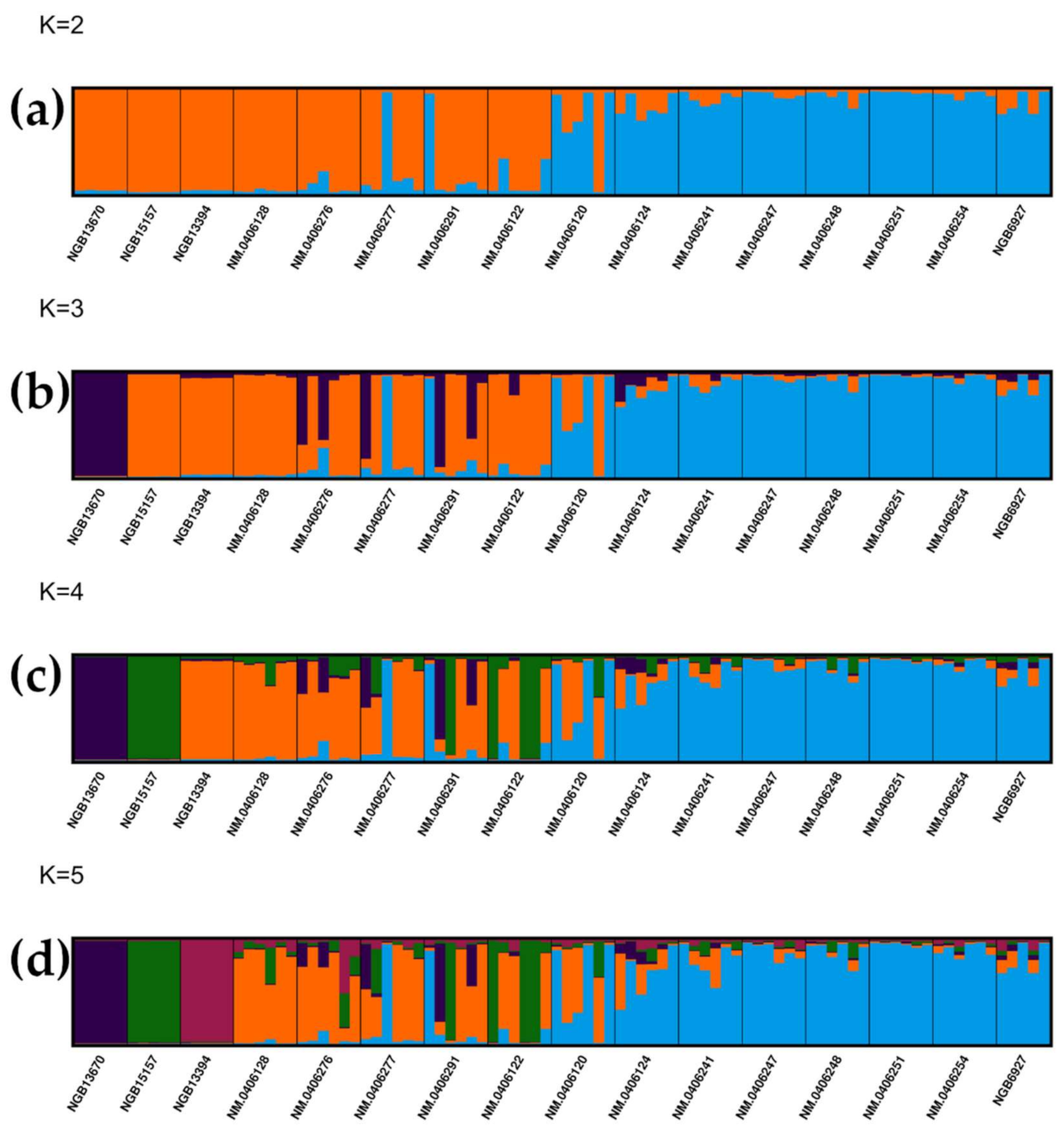
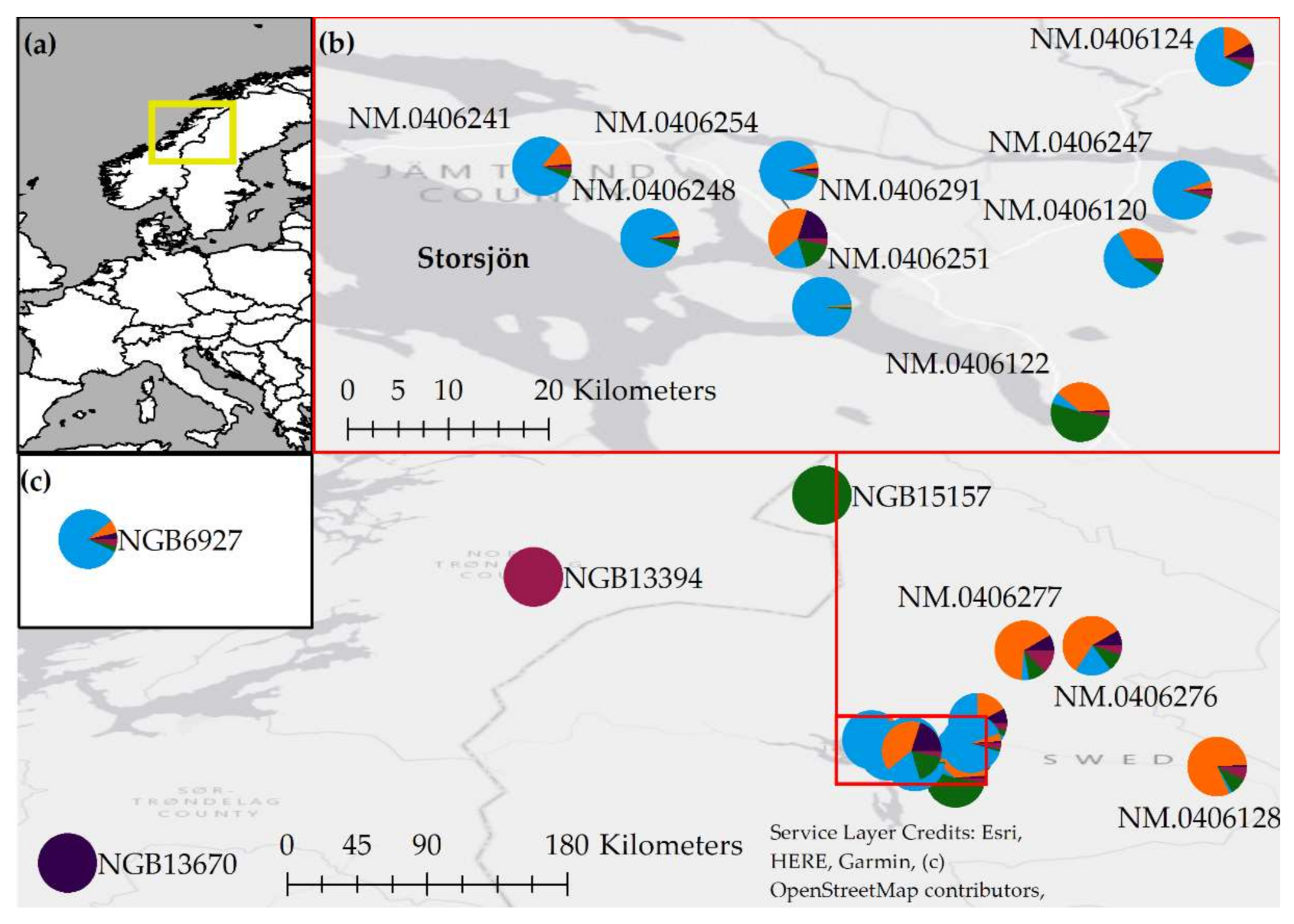
3.2. Analyses of Population Structure Suggest a Central and a Peripheral Genetic Cluster in Jämtland
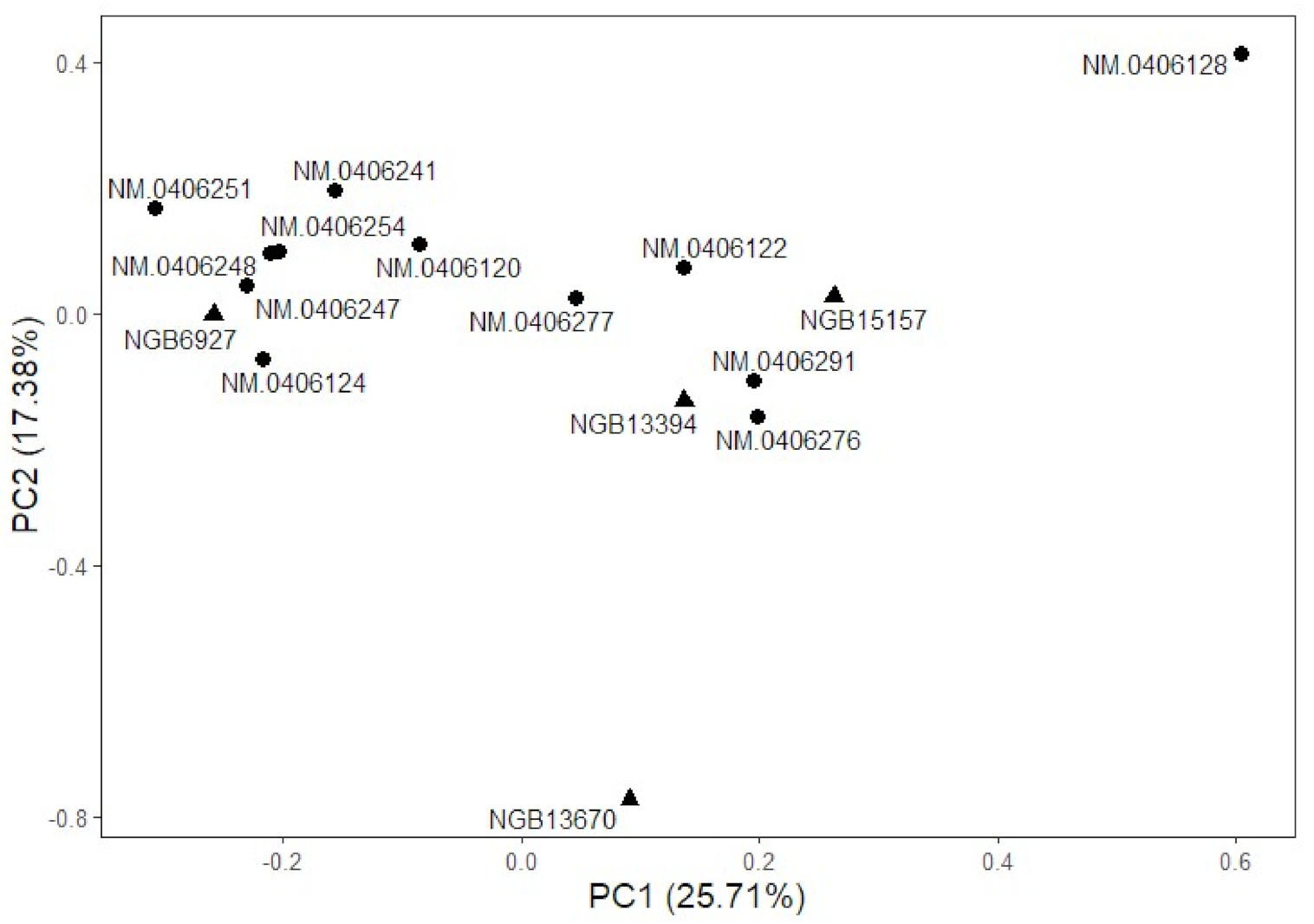
3.3. Differentiation of Accessions Is Partly Explained by Isolation by Distance
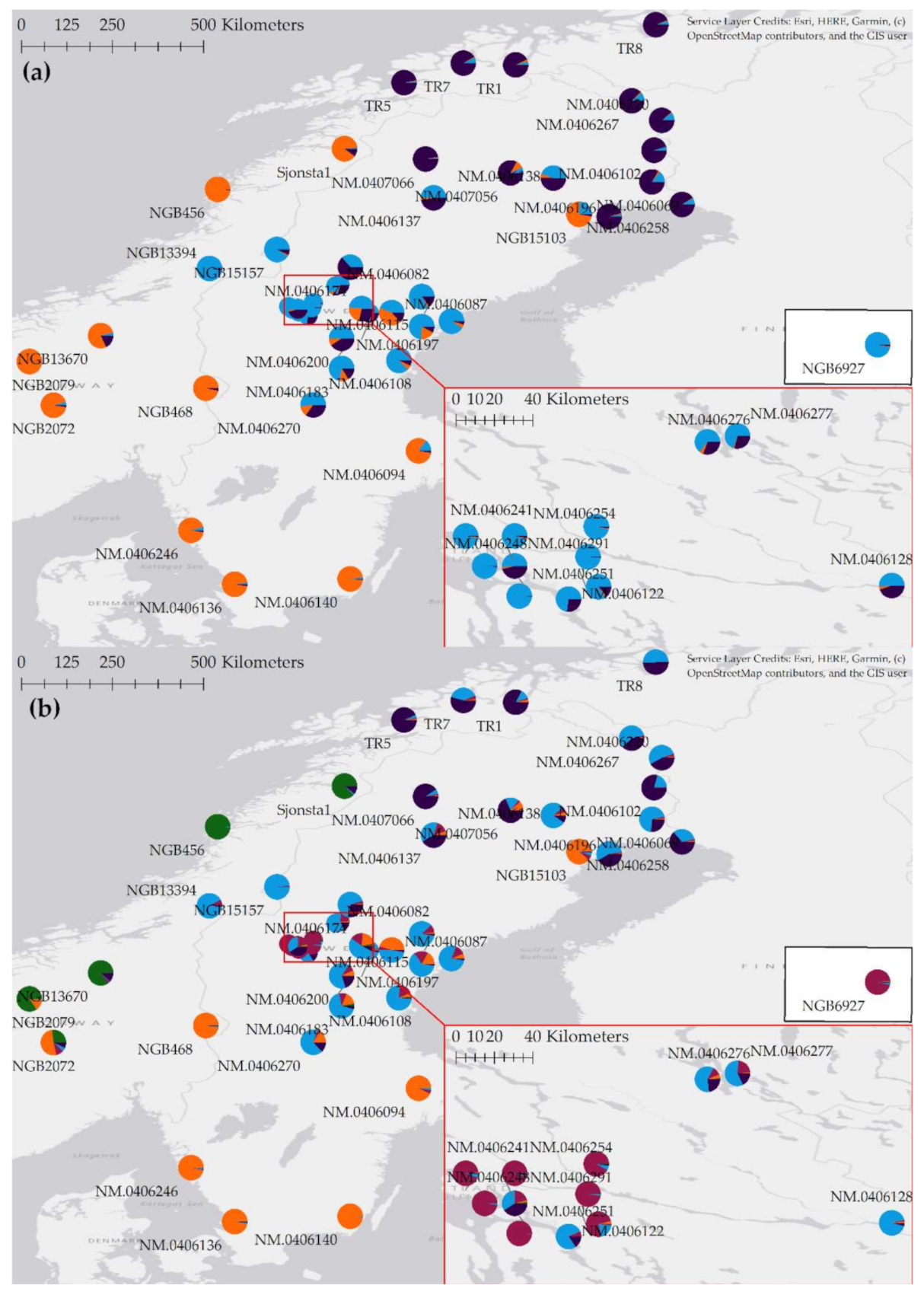
3.4. Barley from Jämtland Both Belong to and Differ from the Genetic Cluster of Barley in Central Scandinavia
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zeven, A.C. Landraces: A review of definitions and classifications. Euphytica 1998, 104, 127–139. [Google Scholar] [CrossRef]
- Leino, M.W. “Primitive” Landraces: Swedish Farmers’ Perception on Seed Traits before the Era of Crop Improvement, in Seedways. The Circulation, Control and Care of Plants in a Warmer World; Karlsson, A.R.B., Ed.; Vitterhets Historie och Antikvitets Akademien: Stockholm, Sweden, 2021. [Google Scholar]
- Villa, T.C.; Maxted, N.; Scholten, M.; Ford-Lloyd, B. Defining and identifying crop landraces. Plant Genet. Resour. 2005, 3, 373–384. [Google Scholar] [CrossRef] [Green Version]
- Harlan, J.R. Our vanishing genetic resources. Science 1975, 188, 618–621. [Google Scholar] [CrossRef]
- Berg, T. Landraces and folk varieties: A conceptual reappraisal of terminology. Euphytica 2009, 166, 423–430. [Google Scholar] [CrossRef]
- Forsberg, N.E.; Russell, J.; Macaulay, M.; Leino, M.W.; Hagenblad, J. Farmers without borders—Genetic structuring in century old barley (Hordeum vulgare). Heredity 2015, 114, 195–206. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pusadee, T.; Wongtamee, A.; Rerkasem, B.; Olsen, K.M.; Jamjod, S. Farmers drive genetic diversity of Thai Purple rice (Oryza sativa L.) landraces. Econ. Bot. 2019, 73, 76–85. [Google Scholar] [CrossRef]
- De Luca, D.; Cennamo, P.; Del Guacchio, E.; Di Novella, R.; Caputo, P. Conservation and genetic characterisation of common bean landraces from Cilento region (southern Italy): High differentiation in spite of low genetic diversity. Genetica 2018, 146, 29–44. [Google Scholar] [CrossRef]
- Hagenblad, J.; Morales, J. An Evolutionary Approach to the History of Barley (Hordeum vulgare) Cultivation in the Canary Islands. Afr. Archaeol. Rev. 2020, 37, 579–595. [Google Scholar] [CrossRef] [PubMed]
- Jones, H.; Lister, D.L.; Bower, M.A.; Leigh, F.J.; Smith, L.M.; Jones, M.K. Approaches and constraints of using existing landrace and extant plant material to understand agricultural spread in prehistory. Plant Genet. Resour. 2008, 6, 98–112. [Google Scholar] [CrossRef]
- Van Heerwaarden, J.; Doebley, J.; Briggs, W.H.; Glaubitz, J.C.; Goodman, M.M.; Gonzalez, J.D.; Ross-Ibarra, J. Genetic signals of origin, spread, and introgression in a large sample of maize landraces. Proc. Natl. Acad. Sci. USA 2011, 108, 1088–1092. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chebotar, S.; Roder, M.S.; Korzun, V.I.; Borner, A. Genetic integrity of ex situ genebank collections. Cell. Mol. Biol. Lett. 2002, 7, 437–444. [Google Scholar]
- Parzies, H.; Spoor, W.; Ennos, R. Genetic diversity of barley landrace accessions (Hordeum vulgare ssp. vulgare) conserved for different lengths of time in ex situ gene banks. Heredity 2000, 84, 476–486. [Google Scholar] [CrossRef]
- Hagenblad, J.; Zie, J.; Leino, M.W. Exploring the population genetics of genebank and historical landrace varieties. Genet. Resour. Crop Evol. 2012, 59, 1185–1199. [Google Scholar] [CrossRef]
- Leino, M.W.; Hagenblad, J.; Edqvist, J.; Strese, E.M. DNA preservation and utility of a historic seed collection. Seed Sci. Res. 2009, 19, 125–135. [Google Scholar] [CrossRef]
- Leino, M.W.; Hagenblad, J. Nineteenth century seeds reveal the population genetics of landrace barley (Hordeum vulgare). Mol. Biol. Evol. 2010, 27, 964–973. [Google Scholar] [CrossRef] [Green Version]
- Leino, M.W. Frösamlingar på museer–ny teknik gör värdelösa föremål värdefulla igen. Nord. Museol. 2010, 96. [Google Scholar] [CrossRef]
- Selçuk, A.; Forsberg, N.; Hagenblad, J.; Leino, M.W. Molecular genotyping of historical barley landraces reveals novel candidate regions for local adaption. Crop Sci. 2015, 55, 2766–2776. [Google Scholar]
- Forsberg, N.E.; Leino, M.W.; Hagenblad, J. Population structure in landrace barley (Hordeum vulgare L.) during the late 19th century crop failures in Fennoscandia. Heredity 2019, 123, 733–745. [Google Scholar] [CrossRef] [PubMed]
- Badr, A.; Rabey, H.E.; Effgen, S.; Ibrahim, H.H.; Pozzi, C.; Rohde, W.; Salamini, F. On the origin and domestication history of barley (Hordeum vulgare). Mol. Biol. Evol. 2000, 17, 499–510. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- FAO (Food and Agricultural Organization of the United Nations). FAOSTAT. 2020. Available online: http://www.fao.org/faostat (accessed on 23 June 2020).
- FAO (Food and Agricultural Organization of the United Nations). Crop Prospects and Food Situation—Quarterly Global Report No. 1. Rome, 2020. Available online: http://www.fao.org/giews/reports/crop-prospects/en/ (accessed on 26 June 2020).
- Akar, T.; Avci, M.; Dusunceli, F. Barley: Post Harvest Operations; Food and Agriculture Organization (FAO) of the United Nations, The Central Research Institute for Field Crops: Ankara, Turkey, 2004; p. 64. [Google Scholar]
- Newton, A.C.; Flavell, A.J.; George, T.S.; Leat, P.; Mullholland, B.; Ramsay, L.; Revoredo-Giha, C.; Russell, J.; Steffenson, B.J.; Swanston, J.S.; et al. Crops that feed the world 4. Barley: A resilient crop? Strengths and weaknesses in the context of food security. Food Secur. 2011, 3, 141–178. [Google Scholar] [CrossRef]
- Pasam, R.K.; Sharma, R.; Walther, A.; Özkan, H.; Graner, A.; Kilian, B. Genetic diversity and population structure in a legacy collection of spring barley landraces adapted to a wide range of climates. PLoS ONE 2014, 9, e116164. [Google Scholar] [CrossRef] [Green Version]
- Poets, A.M.; Fang, Z.; Clegg, M.T.; Morrell, P.L. Barley landraces are characterized by geographically heterogeneous genomic origins. Genome Biol. 2015, 16, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Florence, A.; Ennos, R.A.; Hoad, S.P.; Hoebe, P.N. Variation in light interception traits in European spring barley landraces. Field Crops Res. 2019, 241, 107549. [Google Scholar] [CrossRef]
- Kumar, A.; Verma, R.P.; Singh, A.; Sharma, H.K.; Devi, G. Barley landraces: Ecological heritage for edaphic stress adaptations and sustainable production. Environ. Sustain. Indic. 2020, 6, 100035. [Google Scholar] [CrossRef]
- Sørensen, L.; Karg, S. The expansion of agrarian societies towards the north–new evidence for agriculture during the Mesolithic/Neolithic transition in Southern Scandinavia. J. Archaeol. Sci. 2014, 51, 98–114. [Google Scholar] [CrossRef]
- Vorren, K.-D. Farm development at the Arctic cereal limit in northern Norway—Continuity and discontinuities. Veg. Hist. Archaeobotany 2005, 14, 161–170. [Google Scholar] [CrossRef]
- Leino, M.W. Spannmål: Svenska Lantsorter; Nordiska Museets Förlag: Stockholm, Sweden, 2017. [Google Scholar]
- Hellström, P. Norrlands Jordbruk; Almqvist & Wiksell: Uppsala, Sweden, 1917. [Google Scholar]
- Brink, S. Marknader och forbönder. In Perspektiv på Härjedalen; Sporrong, U., Ed.; Svenska vyer: Sveg, Sweden, 1995; pp. 57–65. [Google Scholar]
- Welinder, S. The northern margin of cereal cultivation in Sweden during the Middle Ages. Fornvännen 2019, 114, 36–42. [Google Scholar]
- Wallin, J.-E.; Oskarsson, B. Odlingslandskapets Framväxt i Storsjöbygden: En Miljöhistorisk Studie i Järnåldersbygd, Baserad på Pollenanalyser och GIS-Baserade Kartanalyser; Jämtlands Läns Museum: Östersund, Sweden, 2002. [Google Scholar]
- Antonson, H. Jämtlands Äldre Agrarlandskap: Försök Till en Regional Indelning Baserad på Lantmäteriakter för Tiden 1650–1899; Kulturmiljöenheten, Länsstyr. i Jämtlands län: Östersund, Sweden, 1993. [Google Scholar]
- Statistics Sweden, Agricultural Statistics. 2020. Available online: http://www.statistikdatabasen.scb.se/pxweb/en/ssd/ (accessed on 13 July 2020).
- Dribe, M.; Olsson, M.; Svensson, P. Famines in the Nordic Countries, AD 536–1875; Lund University, Department of Economic History: Lund, Sweden, 2015. [Google Scholar]
- Osvald, H. Åkerns Nyttoväxter; Sv. Litteratur: Stockholm, Sweden, 1959. [Google Scholar]
- Anonymous. Allmänna Konst- och Industriutställningen i Stockholm: Officiel Katalog Öfver Industriafdelningen; Central-Tryckeriet: Stockholm, Sweden, 1897. [Google Scholar]
- Sveriges Hembygdsförbund. Sveriges Hembygdsförbund. 2020. Available online: https://www.hembygd.se/ (accessed on 17 April 2020).
- He, C.; Holme, J.; Anthony, J. SNP Genotyping: The KASP Assay, in Crop Breeding; Springer: New York, NY, USA, 2014; pp. 75–86. [Google Scholar]
- Moragues, M.; Comadran, J.; Waugh, R.; Milne, I.; Flavell, A.J.; Russell, J.R. Effects of ascertainment bias and marker number on estimations of barley diversity from high-throughput SNP genotype data. Theor. Appl. Genet. 2010, 120, 1525–1534. [Google Scholar] [CrossRef] [PubMed]
- Hagenblad, J.; Morales, J.; Leino, M.W.; Rodríguez-Rodríguez, A.C. Farmer fidelity in the Canary Islands revealed by ancient DNA from prehistoric seeds. J. Archaeol. Sci. 2017, 78, 78–87. [Google Scholar] [CrossRef]
- Jones, H.; Leigh, F.J.; Mackay, I.; Bower, M.A.; Smith, L.M.; Charles, M.P.; Jones, G.; Jones, M.K.; Brown, T.A.; Powell, W. Population-based resequencing reveals that the flowering time adaptation of cultivated barley originated east of the Fertile Crescent. Mol. Biol. Evol. 2008, 25, 2211–2219. [Google Scholar] [CrossRef] [Green Version]
- Ramsay, L.; Comadran, J.; Druka, A.; Marshall, D.F.; Thomas, W.T.; Macaulay, M.; MacKenzie, K.; Simpson, C.; Fuller, J.; Bonar, N.; et al. INTERMEDIUM-C, a modifier of lateral spikelet fertility in barley, is an ortholog of the maize domestication gene TEOSINTE BRANCHED 1. Nat. Genet. 2011, 43, 169–172. [Google Scholar] [CrossRef]
- Comadran, J.; Kilian, B.; Russell, J.; Ramsay, L.; Stein, N.; Ganal, M.; Shaw, P.; Bayer, M.; Thomas, W.; Marshall, D.; et al. Natural variation in a homolog of Antirrhinum CENTRORADIALIS contributed to spring growth habit and environmental adaptation in cultivated barley. Nat. Genet. 2012, 44, 1388. [Google Scholar] [CrossRef]
- Yan, L.; Fu, D.; Li, C.; Blechl, A.; Tranquilli, G.; Bonafede, M.A.; Sanchez, A.L.; Valarik, M.; Yasuda, S.H.; Dubcovsky, J. The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc. Natl. Acad. Sci. USA 2006, 103, 19581–19586. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xia, Y.; Li, R.; Bai, G.; Siddique, K.H.; Varshney, R.K.; Baum, M.; Yan, G.; Guo, P. Genetic variations of HvP5CS1 and their association with drought tolerance related traits in barley (Hordeum vulgare L.). Sci. Rep. 2017, 7, 1–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Komatsuda, T.; Pourkheirandish, M.; He, C.; Azhaguvel, P.; Kanamori, H.; Perovic, D.; Stein, N.; Graner, A.; Wicker, T.; Tagiri, A.; et al. Six-rowed barley originated from a mutation in a homeodomain-leucine zipper I-class homeobox gene. Proc. Natl. Acad. Sci. USA 2007, 104, 1424–1429. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. 2019. Available online: https://www.R-project.org/ (accessed on 10 May 2020).
- RStudio Team. RStudio: Integrated Development for R. 2018. Available online: http://www.rstudio.com/ (accessed on 10 May 2020).
- Wickham, H.; Averick, M.; Bryan, J.; Chang, W.; McGowan, L.D.; François, R.; Grolemund, G.; Hayes, A.; Henry, L.; Hester, J.; et al. Welcome to the Tidyverse. J. Open Source Softw. 2019, 4, 1686. [Google Scholar] [CrossRef]
- Jombart, T. adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef] [Green Version]
- Goudet, J.; Jombart, T. Hierfstat: Estimation and Tests of Hierarchical F-statistics. R Package Version 0.04-22 2015, 10. Available online: https://cran.r-project.org/web/packages/hierfstat/index.html (accessed on 27 June 2020).
- Tang, Y.; Horikoshi, M.; Li, W. ggfortify: Unified interface to visualize statistical results of popular R packages. R J. 2016, 8, 478–489. [Google Scholar] [CrossRef] [Green Version]
- Schauberger, P.; Walker, A. Openxlsx: Read, Write and Edit XLSX Files (Version 4.1.5). 2020. Available online: https://CRAN.R-project.org/package=openxlsx (accessed on 23 July 2020).
- Rousset, F. genepop’007: A Complete Re-Implementation of the Genepop Software for Windows and Linux. Mol. Ecol. Resour. 2008, 8, 103–106. [Google Scholar] [CrossRef]
- Slowikowski, K. ggrepel: Automatically Position Non-Overlapping Text Labels with’ggplot2’. R Package Version 0.8.2. 2020. Available online: https://cran.r-project.org/web/packages/ggrepel/index.html (accessed on 27 June 2020).
- Nei, M. Analysis of gene diversity in subdivided populations. Proc. Natl. Acad. Sci. USA 1973, 70, 3321–3323. [Google Scholar] [CrossRef] [Green Version]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef] [PubMed]
- Falush, D.; Stephens, M.; Pritchard, J.K. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 2003, 164, 1567–1587. [Google Scholar] [CrossRef] [PubMed]
- Falush, D.; Stephens, M.; Pritchard, J.K. Inference of population structure using multilocus genotype data: Dominant markers and null alleles. Mol. Ecol. Notes 2007, 7, 574–578. [Google Scholar] [CrossRef] [PubMed]
- Hubisz, M.J.; Falush, D.; Stephens, M.; Pritchard, J.K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour. 2009, 9, 1322–1332. [Google Scholar] [CrossRef] [Green Version]
- Kopelman, N.M.; Mayzel, J.; Jakobsson, M.; Rosenberg, N.A.; Mayrose, I. Clumpak: A program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 2015, 15, 1179–1191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- ESRI. ArcGIS Desktop. 2017. Available online: https://desktop.arcgis.com/en/arcmap/ (accessed on 10 May 2020).
- Lantmäteriet. Öppna Data. 2020. Available online: https://www.lantmateriet.se/sv/Kartor-och-geografisk-information/oppna-data/ (accessed on 6 August 2020).
- SMHI. Vegetationsperiod. 2020. Available online: https://www.smhi.se/kunskapsbanken/klimat/fenologi/vegetationsperiod-1.6270 (accessed on 6 August 2020).
- Jonsson, J. Historiska Vägar i Jämtlands Län; Jamtli Förlag: Östersund, Sweden, 2001. [Google Scholar]
- Salvesen, H. Jord i Jemtland: Bosetningshistoriske og Økonomiske Studier i Grenseland ca. 1200–1650; AB Wilénska Bokhandelns Förlag: Östersund, Sweden, 1979. [Google Scholar]
- Cross, R.; Wallace, A. Loss of genetic diversity from heterogeneous self-pollinating genebank accessions. Theor. Appl. Genet. 1994, 88, 885–890. [Google Scholar] [CrossRef]
- Lasky, J.R.; Upadhyaya, H.D.; Ramu, P.; Deshpande, S.; Hash, C.T.; Bonnette, J.; Juenger, T.E.; Hyma, K.; Acharya, C.; Mitchell, S.E.; et al. Genome-environment associations in sorghum landraces predict adaptive traits. Sci. Adv. 2015, 1, e1400218. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lister, D.L.; Thaw, S.; Bower, M.A.; Jones, H.; Charles, M.P.; Jones, G.; Smith, L.M.; Howe, C.J.; Brown, T.A.; Jones, M.K. Latitudinal variation in a photoperiod response gene in European barley: Insight into the dynamics of agricultural spread from ‘historic’specimens. J. Archaeol. Sci. 2009, 36, 1092–1098. [Google Scholar] [CrossRef]
| Accession | Farm/Village | Country | Collected | Lat | Long | Hs |
|---|---|---|---|---|---|---|
| NGB13670 | Oppdal | NOR | NA | 62.5943 | 9.6912 | 0.000 |
| NGB15157 | Jormvattnet | SWE | 1905 | 64.7189 | 14.0468 | 0.000 |
| NGB13394 | Snåsa | NOR | 1879 | 64.2465 | 12.3822 | 0.000 |
| NM.0406128 | Gevåg, Ragunda | SWE | 1896 | 63.1498 | 16.3310 | 0.074 |
| NM.0406276 | Tullingsås, Ström | SWE | 1896 | 63.8214 | 15.5062 | 0.074 |
| NM.0406277 | Risselås, Ström | SWE | 1896 | 63.8493 | 15.6085 | 0.065 |
| NM.0406291 | Rösta, Täng | SWE | 1896 | 63.2407 | 14.5651 | 0.086 |
| NM.0406122 | Backen, Brunflo | SWE | 1896 | 63.0855 | 14.8181 | 0.054 |
| NM.0406120 | Lungre, Kyrkås | SWE | 1896 | 63.2261 | 14.8659 | 0.067 |
| NM.0406124 | Österåsen, Häggenås | SWE | 1896 | 63.4035 | 14.9474 | 0.066 |
| NM.0406241 | Bjerte, Nälden | SWE | 1896 | 63.3056 | 14.3360 | 0.056 |
| NM.0406247 | Böle, Lit | SWE | 1896 | 63.2841 | 14.9096 | 0.059 |
| NM.0406248 | Silje, Rödön | SWE | 1896 | 63.2411 | 14.4327 | 0.046 |
| NM.0406251 | Kvarnsved, Frösön | SWE | 1896 | 63.1795 | 14.5867 | 0.049 |
| NM.0406254 | Landsom, Täng | SWE | 1896 | 63.2680 | 14.5571 | 0.057 |
| NGB6927 | Jämtland | SWE | NA | NA | NA | 0.044 |
| Name | Function | Reference |
|---|---|---|
| ppd-H1_ SNP48 | Photoperiod response, causative | [45] |
| Int-c_SNP124 | Spike morphology, modifier | [46] |
| HvCEN SNP9 | Spring or winter growth habit/earliness | [47] |
| HvFT1 i1_270 | Flowering time | [48] |
| HvFT1 prom_167 | Flowering time | [48] |
| HvP5CS1 SNP735 | Drought resistance | [49] |
| HvP5CS1 SNP846 | Drought resistance | [49] |
| Vrs1 E152 > F.S. | Spike morphology, causative | [50] |
| Vrs1 A40 > F.S. | Spike morphology, causative | [50] |
| Acc\Acc | NGB13670 | NGB15157 | NGB13394 | NM.0406128 | NM.0406276 | NM.0406277 | NM.0406291 | NM.0406122 | NM.0406120 | NM.0406124 | NM.0406241 | NM.0406247 | NM.0406248 | NM.0406251 | NM.0406254 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NGB15157 | 1.000 | ||||||||||||||
| NGB13394 | 1.000 | 1.000 | |||||||||||||
| NM.0406128 | 0.520 | 0.290 | 0.290 | ||||||||||||
| NM.0406276 | 0.437 | 0.344 | 0.344 | 0.099 | |||||||||||
| NM.0406277 | 0.508 | 0.442 | 0.392 | 0.124 | 0.088 | ||||||||||
| NM.0406291 | 0.349 | 0.341 | 0.280 | 0.113 | 0.074 | 0.076 | |||||||||
| NM.0406122 | 0.606 | 0.287 | 0.476 | 0.104 | 0.129 | 0.153 | 0.148 | ||||||||
| NM.0406120 | 0.539 | 0.430 | 0.396 | 0.160 | 0.146 | 0.118 | 0.086 | 0.167 | |||||||
| NM.0406124 | 0.481 | 0.516 | 0.423 | 0.252 | 0.225 | 0.179 | 0.141 | 0.284 | 0.141 | ||||||
| NM.0406241 | 0.591 | 0.540 | 0.501 | 0.252 | 0.218 | 0.192 | 0.184 | 0.246 | 0.101 | 0.150 | |||||
| NM.0406247 | 0.548 | 0.559 | 0.474 | 0.268 | 0.213 | 0.187 | 0.153 | 0.262 | 0.099 | 0.080 | 0.082 | ||||
| NM.0406248 | 0.635 | 0.597 | 0.573 | 0.297 | 0.221 | 0.234 | 0.185 | 0.282 | 0.099 | 0.173 | 0.084 | 0.079 | |||
| NM.0406251 | 0.662 | 0.639 | 0.612 | 0.326 | 0.278 | 0.237 | 0.218 | 0.328 | 0.157 | 0.119 | 0.157 | 0.072 | 0.134 | ||
| NM.0406254 | 0.564 | 0.544 | 0.494 | 0.272 | 0.214 | 0.184 | 0.151 | 0.266 | 0.097 | 0.084 | 0.065 | 0.031 | 0.078 | 0.077 | |
| NM.0406254 | 0.644 | 0.672 | 0.606 | 0.345 | 0.276 | 0.248 | 0.214 | 0.291 | 0.198 | 0.183 | 0.184 | 0.119 | 0.169 | 0.210 | 0.156 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Larsson, M.N.A.; Leino, M.W.; Hagenblad, J. Genetic Diversity in 19th Century Barley (Hordeum vulgare) Reflects Differing Agricultural Practices and Seed Trade in Jämtland, Sweden. Diversity 2021, 13, 315. https://doi.org/10.3390/d13070315
Larsson MNA, Leino MW, Hagenblad J. Genetic Diversity in 19th Century Barley (Hordeum vulgare) Reflects Differing Agricultural Practices and Seed Trade in Jämtland, Sweden. Diversity. 2021; 13(7):315. https://doi.org/10.3390/d13070315
Chicago/Turabian StyleLarsson, Martin N. A., Matti W. Leino, and Jenny Hagenblad. 2021. "Genetic Diversity in 19th Century Barley (Hordeum vulgare) Reflects Differing Agricultural Practices and Seed Trade in Jämtland, Sweden" Diversity 13, no. 7: 315. https://doi.org/10.3390/d13070315
APA StyleLarsson, M. N. A., Leino, M. W., & Hagenblad, J. (2021). Genetic Diversity in 19th Century Barley (Hordeum vulgare) Reflects Differing Agricultural Practices and Seed Trade in Jämtland, Sweden. Diversity, 13(7), 315. https://doi.org/10.3390/d13070315






