Abstract
Biodiversity hotspots often suffer from a lack of taxonomic knowledge, particularly those in tropical regions. However, accurate taxonomic knowledge is needed to support sustainable management of biodiversity, especially when it is harvested for human sustenance. Sundaland, the biodiversity hotspot encompassing the islands of Java, Sumatra, Borneo, and Peninsular Malaysia, is one of those. With more than 900 species, its freshwater ichthyofauna includes a large number of medium- to large-size species, which are targeted by inland fisheries. Stock assessment requires accurate taxonomy; however, several species groups targeted by inland fisheries are still poorly known. One of those cases is the cyprinid genus Barbonymus. For this study, we assembled a consolidated DNA barcode reference library for Barbonymus spp. of Sundaland, consisting of mined sequences from BOLD, as well as newly generated sequences for hitherto under-sampled islands such as Borneo. A total of 173 sequences were analyzed using several DNA-based species delimitation methods. We unambiguously detected a total of 6 Molecular Operational Taxonomic Units (MOTUs) and were able to resolve several conflicting assignments to the species level. Furthermore, we clarified the identity of MOTUs occurring in Java.
1. Introduction
Sundaland, comprising the islands of Java, Bali, Sumatra, Borneo, and peninsular Malaysia, constitutes one of the world’s largest biodiversity hotspots [1,2]. With circa 900 freshwater fish species, half of which are endemic, the ichthyofauna of this biogeographical region is particularly rich, with a density of 0.8 species per km2, a value twice as large as that observed in Brazil and the Democratic Republic of Congo [3]. This large diversity is critically threatened, mostly due to the alarming rate of deforestation over the past few decades [4,5,6], in conjunction with pollution [7] and watershed fragmentation through the development of dams for irrigation and hydroelectric power [8]. Furthermore, the diversity of freshwater fishes in Sundaland is still not sufficiently understood [9], hampering conservation efforts.
Freshwater fishes constitute a major source of animal protein in Southeast Asia, where inland fisheries rank among the world’s most productive, with Indonesia repeatedly at the top [10,11,12,13]. Sundaland hosts a substantial amount of medium to large-size species, all above 30 cm in maximum standard length [3], which are targeted by inland fisheries [11]. Common targets include genera such as snakeheads (Channa spp.), catfishes (Ompok spp., Hemibagrus spp.), and various cyprinid genera including Barbonymus spp. and Leptobarbus spp. [11,14]. Although common, the taxonomy of these genera is poorly understood. Species boundaries are unclear, and the diversity for a number of genera has likely been underestimated [15,16,17].
A good example for this is the cyprinid genus Barbonymus, for which species numbers range from 5 in Fishbase [18] to 10 in Eschmeyer’s Catalog of Fishes [19], the latter following the checklist of Southeast Asian freshwater fishes by Kottelat [17] (Table 1). All Barbonymus species occur in Sundaland, except B. altus (Günther, 1868), which was described from the Chao Phraya River in Thailand (Text S1 in Supplementary Material). Of the nine species occurring in Sundaland, three, B. sunieri (Weber and de Beaufort, 1916), B. strigatus (Boulenger, 1894), and B. platysoma (Bleeker, 1855), have been described based on a single specimen, and none of them have ever been observed since their original description [20]. Among the six remaining species, B. balleroides (Valenciennes, in Cuvier and Valenciennes, 1842), B. gonionotus (Bleeker, 1849) and B. schwanefeldii (Bleeker, 1864) are widespread in watersheds flowing to the Java Sea, where they are frequently targeted by inland fisheries. Barbonymus collingwoodii (Günther, 1868) and B. mahakkamensis (Ahl, 1922) are endemic to North and East Borneo, respectively, and B. belinka (Bleeker, 1860) is an endemic species of the west coast of Sumatra.

Table 1.
List of available nominal species of Barbonymus including species names in original descriptions, authors, species names in Eschmeyer’s Catalog of Fishes and Fishbase, and current status following [17].
DNA barcoding, the use of cytochrome oxidase I (COI) as a species tag for automated identification, opened new perspectives for the characterization of Sundaland’s ichthyofauna by helping to clarify taxonomic confusion within several groups [15,21,22], by identifying discrepancies in historical species records [9] and by detecting a substantial amount of morphologically similar, yet highly divergent lineages (i.e., cryptic diversity) within numerous species [9,15,21,22,23,24,25,26,27,28]. Several molecular studies that aimed at characterizing patterns of genetic diversity in Barbonymus led to conflicting species identities associated with sequences submitted to international repositories [9,16,29,30,31,32,33,34,35,36].
As part of an ongoing project that seeks to build a DNA barcode reference library for the ichthyofauna of Sundaland [9,22], we generated new barcode records for Barbonymus species, which, together with previously published sequences, cover the diversity of the genus in the region. The objective of the present study is to re-examine Barbonymus species boundaries and their distribution ranges using DNA-based species delimitation methods. By including DNA barcode records of specimens collected near type localities, we are also reappraising published Barbonymus sequences.
2. Materials and Methods
2.1. Sampling and Collection Management
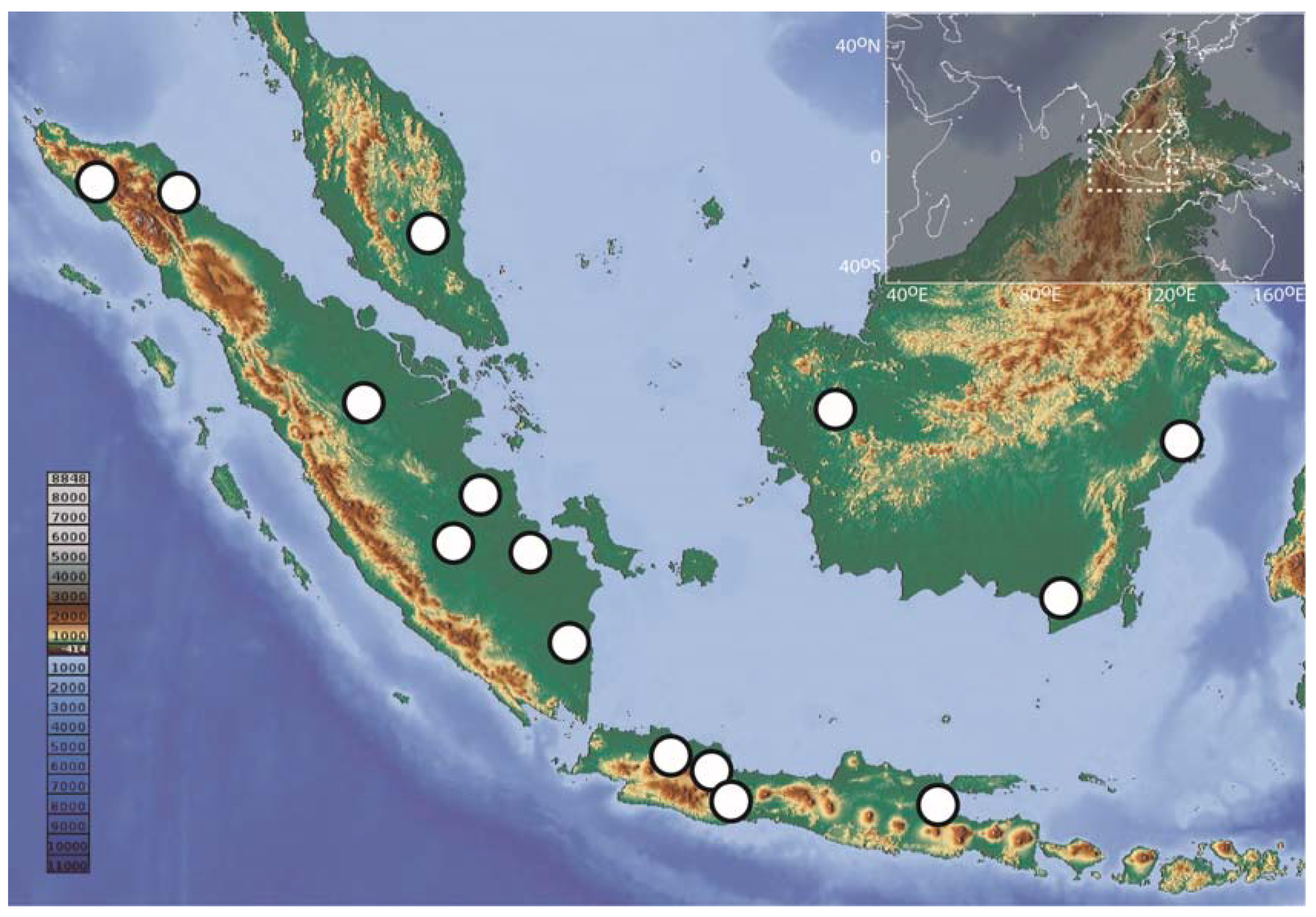
Specimens were captured using various methods including electrofishing, seine nets, cast nets and gill nets, as well as by visiting fish markets in Sundaland (Figure 1; DS-BARBONYM, dx.doi.org/10.5883/DS-BARBONYM, accessed on 15 January 2021). Specimens were photographed and individually labeled, and voucher specimens were preserved in a 5% formalin solution. A fin clip or a muscle biopsy was taken for each specimen and fixed in a 96% ethanol solution for further genetic analyses. Both tissue and voucher specimens were deposited in the national collections at the Museum Zoologicum Bogoriense (MZB) in the Research Centre for Biology (RCB) of the Indonesian Institute of Sciences (LIPI).
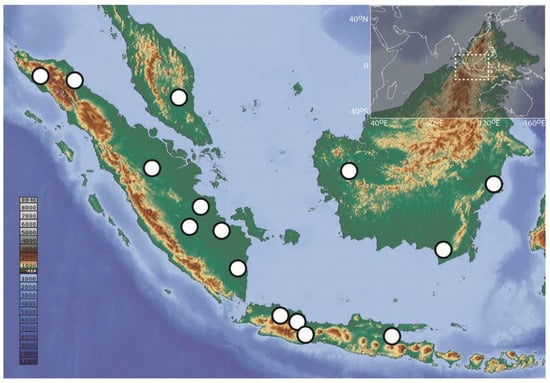
Figure 1.
Collecting sites in Sundaland for the 173 DNA barcode records of Barbonymus analyzed in this study.
2.2. Sequencing and International Repositories
Genomic DNA was extracted from the muscle tissue samples using a Qiagen DNeasy 96 tissue extraction kit following manufacturer’s specifications. A 651 bp segment from the 5′ region of the cytochrome oxidase I gene (COI) was amplified using the primer cocktail C_FishF1t1/C_FishR1t1 [37]. Polymerase Chain Reaction (PCR) amplifications were done on a Veriti 96-well Fast thermocycler (ABI—Applied Biosystems) with a final volume of 10.0 μL containing 5.0 μL buffer 2X, 3.3 μL ultrapure water, 1.0 μL each primer (10 μM), 0.2 μL enzyme Phire Hot Start II DNA polymerase (5U), and 0.5 μL of DNA template (~50 ng). The following thermocycler regime was used: initial denaturation at 98 °C for 5 min followed by 30 cycles denaturation at 98 °C for 5 s, annealing at 56 °C for 20 s, and extension at 72 °C for 30 s, followed by a final extension step at 72 °C for 5 min. PCR products were purified with ExoSap-IT (USB Corporation, Cleveland, OH, USA) and sequenced in both directions. Sequencing reactions were performed at the Centre for Biodiversity Genomics, University of Guelph, Canada, using the BigDye Terminator v3.1 Cycle Sequencing Ready Reaction kit following standard protocols described in [38]. Sequencing was performed on an ABI 3730xl capillary sequencer (Applied Biosystems). Sequences and collateral information were deposited on BOLD [39] and are available as a public data set (dx.doi.org/10.5883/DS-BARBONYM, accessed on 15 January 2021, Table S1).
2.3. Genetic Species Delimitation and Phylogenetic Inferences
Several methods for species delineation based on DNA sequences have been proposed [40,41,42,43]. Each of these have different properties, particularly when dealing with singletons (i.e., lineages represented by a single sequence) or heterogeneous speciation rates among lineages [44]. A combination of different approaches is increasingly used to overcome potential pitfalls arising from uneven sampling [22,45,46,47,48]. We used six different sequence-based methods of species delimitation to identify the Molecular Operational Taxonomic Unit (MOTU): (1) Refined Single Linkage (RESL) as implemented in BOLD and used to generate Barcode Index Numbers (BIN) [42], (2) Automatic Barcode Gap Discovery (ABGD) [41], (3) Poisson Tree Process (PTP) in its single (sPTP) and multiple rates version (mPTP) as implemented in the stand-alone software mptp_0.2.3 [43,49], and (4) General Mixed Yule-Coalescent (GMYC) in its single (sGMYC) and multiple threshold version (mGMYC) as implemented in the R package Splits 1.0–19 [50].
Both the mPTP algorithm and the GYMC use phylogenetic trees as input files. We reconstructed a maximum likelihood (ML) tree for the former using RAxML [51] based on a GTR + I + Γ substitution model. For the GYMC algorithm, we calculated an ultrametric, fully resolved tree using the Bayesian approach implemented in BEAST 2.6.2 [52]. Sequences were collapsed into haplotypes prior to reconstructing the ultrametric tree using RAxML, and Bayesian reconstruction was based on a strict-clock prior of 1.2% per million year [53]. Two Markov chains of 20 million each were run independently using Yule pure birth and GTR + I + Γ substitution models. Trees were sampled every 5000 states, after an initial burn-in period of 5 million. Both runs were combined with trees resampled every 20,000 states using LogCombiner 2.6.2, and the maximum credibility tree was constructed using TreeAnnotator 2.6.2 [52].
A final COI gene tree was reconstructed using the SpeciesTreeUCLN algorithm of the StarBEAST2 package [54]. This approach implements a mixed-model including a coalescent component within species and a diversification component between species that allows accounting for variations of substitution rates within and between species [55]. SpeciesTreeUCLN jointly reconstructs gene trees and species trees and therefore requires the designation of species, which were determined using the consensus of our species delimitation analyses. The SpeciesTreeUCLN analysis was performed with the same parameters as mentioned above.
Kimura 2-parameter (K2P) [56] pairwise genetic distances were calculated using the R package Ape 5.4 [57]. Maximum intraspecific and nearest neighbor genetic distances were calculated from the pairwise K2P distance matrix using the R package Spider 1.5 [58]. Haplotype extraction and haplotype network reconstruction were performed for the most widespread species using the R package pegas 1.0 [59].
3. Results
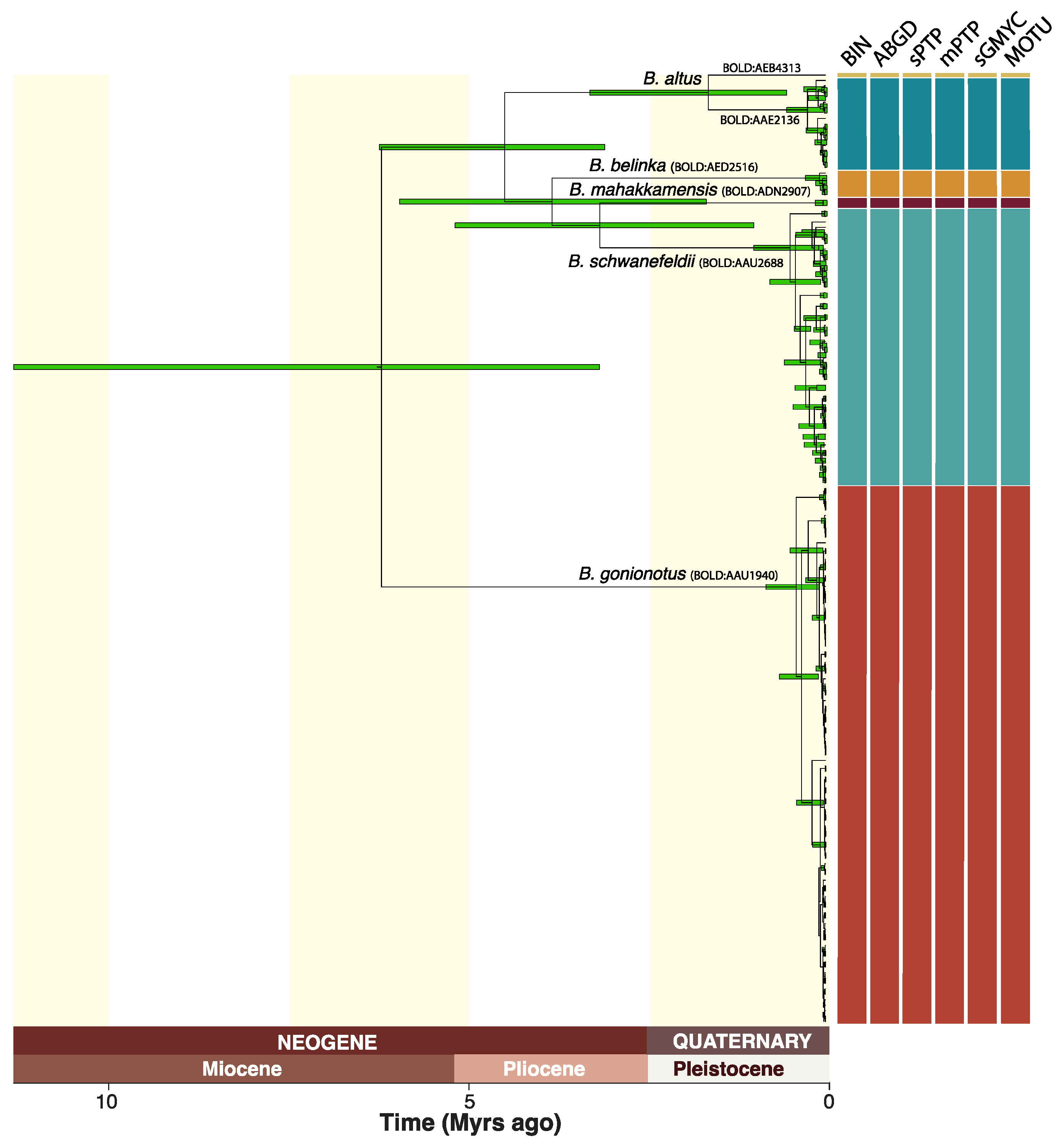
The total of 173 DNA barcodes used for this study comprised 154 sequences mined from BOLD and 19 sequences generated for Barbonymus specimens originating from Sumatra and Borneo. The newly generated sequences represent the first DNA barcode records of Barbonymus for Borneo. All the sequences were above 500 bp in length and no stop codons were detected, suggesting that the sequences collected represent functional coding regions. DNA-based species delimitation methods resulted in congruent delimitation schemes with 6 MOTUs for mPTP, sPTP, ABGD, RESL, and sGMYC (Figure 2; Table S1). However, mGMYC resulted in the delimitation of 31 highly incongruent MOTUs. Therefore, the mGMYC partitioning scheme was discarded. The final consensus scheme consisted of six MOTUs (Figure 2; Table 2) showing a distinct barcoding gap, which is defined as the lack of overlap between maximum intraspecific and minimum interspecific genetic distance. Maximum intraspecific distances ranged from 0 (BOLD:ADN2907, BOLD:AED2516) to 0.018 (BOLD:AAD1940) (Table 1). Minimum interspecific distances ranged from 0.026 for the two Barbonymus MOTUs (BOLD:AAE2136 and BOLD:AEB4313) to 0.08 for BOLD:AAD1940 (Table 1).
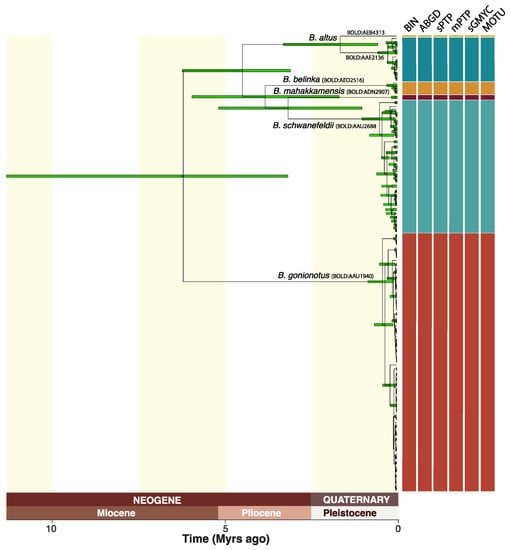
Figure 2.
Mitochondrial gene tree for the 173 DNA barcodes of Barbonymus inferred with SpeciesTreeUCLN, including 95% HPD interval for node age estimates, genetic species delimitation results for five methods (mGMYC discarded) and their 50% consensus, BOLD Barcode Index Numbers (BIN) for each MOTU, and revised species names.

Table 2.
Summary of genetic distances and MOTUs including species names, number of individuals analyzed, BOLD Barcode Index Number (BIN), maximum intraspecific and minimum interspecific K2P genetic distances.
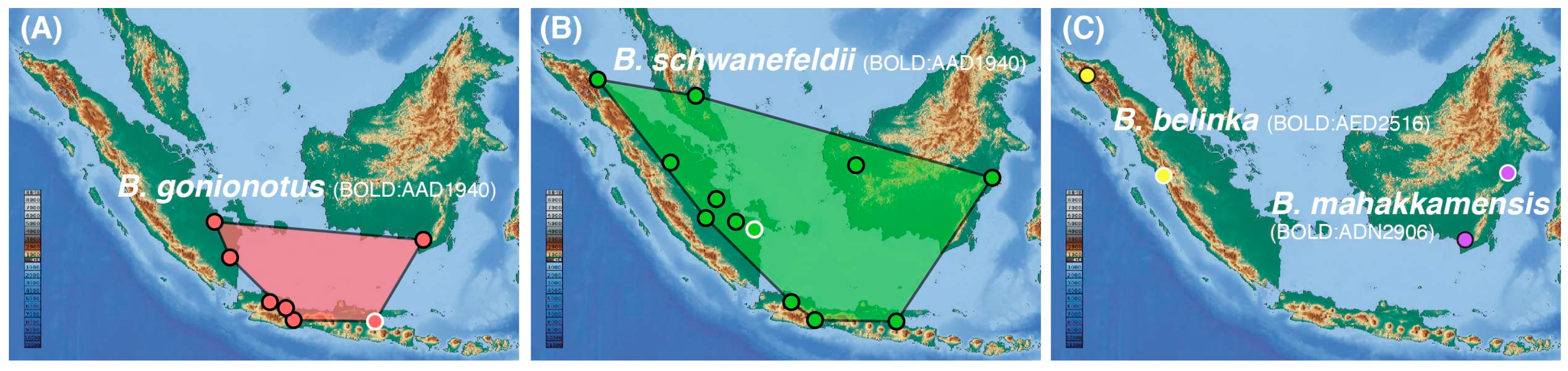
Conflicting species-level assignments were detected, particularly for previously published records from Java [9], where BIN BOLD:ADD1940 and BOLD:AAU0688, initially assigned to B. balleroides and B. gonionotus, match B. gonionotus and B. schwanefeldii, respectively (Table S1). These results extend the occurrence of B. schwanefeldii to Java Island and question the occurrence of B. balleroides in Java (Figure 3). Along the same line, the BIN BOLD:AED2516, initially assigned to B. gonionotus and highlighted as a potentially new taxon [16], likely corresponds to B. belinka, because B. collingwoodii is endemic to North Borneo, B. mahakkamensis corresponds to a distinct lineage (BOLD:ADN2907, Figure 2) restricted to East Borneo (Figure 3), and the occurrence of B. balleroides in Sumatra is uncertain (Text S1).
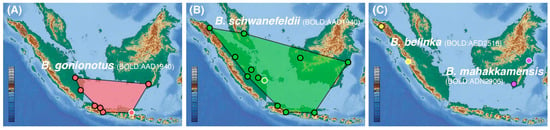
Figure 3.
Revised range distribution and type localities (white circle) for (A) B. gonionotus (only records in Sundaland with geocoordinates are shown, its range distribution expands northward to Thailand and India), (B) B. schwanefeldii (only records in Sundaland are shown), (C) B. belinka, and B. mahakkamensis. B. altus occurs in mainland Asia.
The Bayesian gene tree based on delimited MOTUs suggests close phylogenetic affinities between the Sundaland MOTUs corresponding to B. belinka, B. mahakkamensis, and B. schwanefeldii, with a Most Recent Common Ancestor (MRCA) dated at 3.8 Ma (Figure 2). This group is more closely related to MOTUs of B. altus from mainland Asia, with a MRCA dated at 4.5 Ma, than the MOTU assigned to B. gonionotus, which diverged from other Barbonymus MOTUs about 6 Ma.
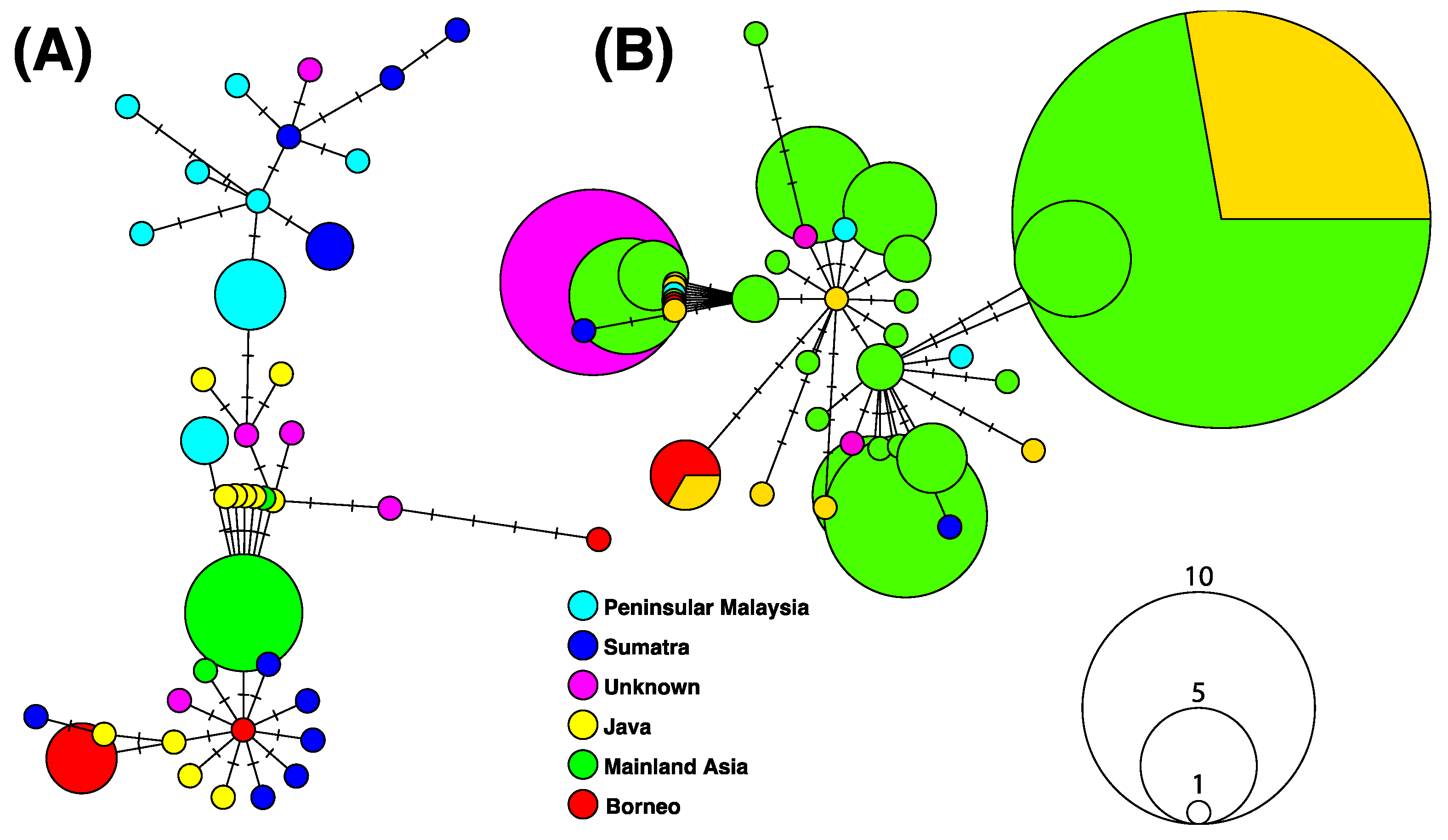
Intraspecific phylogeographic patterns were further explored for B. gonionotus and B. schwanefeldii using sequences with revised species assignment (Figure 4). A total of 40 haplotypes was observed for both B. gonionotus and B. schwanefeldii. Haplotype networks were markedly different for both species, with a star-like structure for B. schwanefeldii (Figure 4B) and a more scattered network for B. gonionotus (Figure 4A). Most islands of Sundaland host haplotypes scattered across networks for both species; however, mainland Asia is much more represented in the haplotype network of B. schwanefeldii (Figure 4B) than in B. gonionotus (Figure 4A).
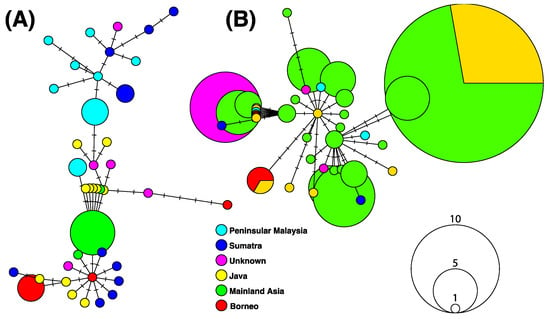
Figure 4.
Haplotype networks reconstructed based on 98 and 50 DNA barcodes available for B. gonionotus (A) and B. schwanefeldii (B), respectively. Numbers of changes are indicated by small segments on links. Circles represent haplotypes; the size of a circle is proportional to the haplotype frequency.
4. Discussion
The present study provides an update of the Barbonymus diversity in Sundaland through the aggregation of newly generated and recently published DNA barcode records, resulting in a reference library consisting of 173 sequences largely distributed across mainland Asia and Sundaland (the haplotype MK978151 was observed in five individuals in [16], resulting in a dataset in BOLD consisting of 169 sequences). DNA-based species delimitation methods largely agreed on the delineation of six MOTUs, except mGMYC with a much higher estimate, a fact that was already reported in other studies [22,27,46,48,60]. Aside from this noticeable exception, methods were concordant in their delimitations, and a barcoding gap was observed between all six MOTUs, with maximum K2P distances mostly <1% within MOTUs, thereby falling into previously observed ranges for Cypriniformes in Sundaland [9,16,21,22]. Along the same line, inferred ages of divergence among MOTUs are consistent with previous age estimates among Sundaland freshwater fishes, with most species originating during the last 5 Ma [21,22,26,28].
Discrepancies between BIN and species designation were observed within three MOTUs, corresponding to (1) BOLD:ADD1940 with sequences assigned to both B. gonionotus and B. balleroides, (2) BOLD:AAU0688 with sequences assigned to both B. gonionotus and B. schwanefeldii, and (3) BOLD:AED2516 with sequences assigned initially to B. gonionotus. Most discrepancies could be related to B. gonionotus, which appears scattered across these three MOTUs.
Most sequences of BOLD:ADD1940 from mainland Asia were attributed to B. gonionotus [29,32,33,36], while sequences from Sundaland were called B. balleroides [9,16]. Sequences from BOLD:AAU0688 from Java were assigned to B. gonionotus, while previously published and newly generated sequences from Sumatra and Borneo were assigned to B. schwanefeldii. These results suggest multiple misidentifications or large-scale introgressive hybridization between B. gonionotus and B. schwanefeldii, as the former is not reported to occur in Java [3,18,19,20]. While hybridization and introgression have been previously reported for Cypriniformes [61,62,63], such large-scale mitochondrial introgressions have never been reported for Sundaland fishes [22,23,25]. Along the same line, shared polymorphism through recent common ancestry might be responsible for these discrepancies; however, as inferred by this study, B. gonionotus is the most divergent species in Sundaland, and ancestral polymorphisms are usually detected across whole species distribution ranges [64], which is not the case here. The most likely hypothesis is that B. schwanefeldii occurs in Java, extending its range of distribution over all islands of Sundaland. Cases of translocation of B. schwanefeldii outside its distribution range have been reported, e.g., to Papua New Guinea [65], as a strategy to improve local fisheries by introducing species with fast growth rates. Numerous introductions have been reported in Java of both Sundaland species, non-native to Java, and exotic species [9]. This makes an introduction of B. schwanefeldii through translocations from either Sumatra or Borneo populations likely. However, the haplotype network of BOLD:AAU0688 indicates that Java specimens consist of mostly private haplotypes (Figure 4). If the B. schwanefeldii occurrence in Java resulted from recent introductions, most haplotypes would be shared with other populations in Sumatra and Borneo, which is not the case here. Furthermore, cases of MOTUs widely distributed in Sundaland were previously detected [15,28] and most BOLD:AAU0688 specimens from Java initially identified as B. gonionotus are juveniles and sub-adults, which makes misidentifications likely. These results suggest that Java populations previously assigned to B. gonionotus actually correspond to B. schwanefeldii (Figure 3).
The present study further questions the occurrence of B. balleroides in Java, which was comprehensively sampled recently [9,21], resulting in the discovery of two distinct Barbonymus MOTUs. The MOTU BOLD:AAD1940 was sampled at the type locality of B. gonionotus in Surabaya (Text S1, Figure 3), and all mainland Asia samples of this MOTU were previously assigned to B. gonionotus. This result suggests that Barbonymus populations previously assigned to B. balleroides [9,16,20] actually belong to B. gonionotus (Figure 4).
The MOTU BOLD:AED2516 likely corresponds to B. belinka. Barbonymus sunieri, B. strigatus, and B. platysoma have been described based on single specimens from either Java or North Borneo, none of which have been observed for decades. In addition, B. collingwoodii is endemic to North Borneo and B. mahakkamensis belongs to a distinct lineage (BOLD:ADN2907, Figure 2) restricted to East Borneo (Figure 3). The type locality of B. balleroides is unknown, but the holotype refers to the Indo-Australian region (Text S1). However, synonymies suggest ample distribution of B. balleroides in Java and Borneo, though its occurrence in Sumatra remains to be confirmed. Thus, B. belinka is the only name available for this MOTU in Sumatra, considering known distribution ranges of recently observed Barbonymus species.
A single case of unrecognized diversity is detected within B. altus, with two MOTUs detected including BOLD:AAE2136 and BOLD:AEB4313. The MOTU BOLD:AEB4313 corresponds to a singleton mined from GenBank and originating from a specimen sampled in the Mekong River (Vietnam), which is consistent with the identification as B. altus considering its type locality is the neighboring Chao Phraya River in Thailand. Raw electropherograms are not available for this sequence, which makes its quality assessment impossible; however, its placement within B. altus seems to confirm that this singleton does not result from poor sequence quality.
5. Conclusions
The present study confirms the utility of DNA barcoding for clarifying species identity and distribution ranges in cases of conflicting records. This is particularly evident in Java, where large conflicts between historical records and recent reappraisals based on DNA sequences were recently detected [9,21,66], suggesting large knowledge gaps. The lack of historical records for B. schwanefeldii in Java seems to indicate perpetuated misidentifications of Barbonymus populations. Individual translocations of B. schwanefeldii from Sumatra and Borneo to Java alone fail to account for its new occurrence. Our study suggests that both MOTUs reported from Java correspond to B. gonionotus and B. schwanefeldii and highlights the degree to which Barbonymus species are morphologically similar and difficult to distinguish based on meristic counts alone. However, DNA barcodes are clustered into MOTUs, which are unambiguously captured by most DNA-based species delimitation methods. Our revised DNA barcode reference library opens new perspectives for the management of the inland fisheries of Sundaland by enabling fast and reliable species level identification of Barbonymus spp. Considering the importance of Barbonymus spp. in local artisanal fisheries, and the difficulty of performing stock assessments at species level due to overlapping meristic counts among species, this library can be readily used as an alternative.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/d13070283/s1, Table S1: Results of the genetic species delimitation analyses. Text S1: Nomenclature of the ten nominal species of Barbonymus following Eschmeyer et al., 2018.
Author Contributions
Conceptualization, H.D., A.S., T.S., D.W., D.S. and N.H.; methodology, A.S., H.D., D.S. and N.H.; software, A.S. and N.H.; validation, H.D., A.S., T.S., D.W., D.S. and N.H.; formal analysis, H.D., A.S. and N.H.; investigation, H.D., A.S., T.S., D.W., D.S. and N.H.; resources, H.D., A.S., T.S., S.S., U.N., D.S. and N.H.; data curation, H.D., D.S. and N.H.; writing—original draft preparation, H.D., A.S., D.S. and N.H.; writing—review and editing, T.S., D.W. and S.S.; visualization, H.D., A.S., T.S., D.W., D.S. and N.H.; supervision, D.W., D.S. and N.H.; project administration, N.H.; funding acquisition, D.S., D.W. and N.H. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the CFREF to the University of Guelph’s Food from Thought program, by the “Institut de Recherche pour le Développement” through annual funding and “Fonds d’Amorçage” (226F2ABIOS), by the French Embassy in Indonesia through the program “Science et Impact”, by Campus France through a Bio-Asia grant (BIOSHOT project) and by ANR (ANR-17-ASIE-0006).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data presented in this study are openly available in BOLD at [dx.doi.org/10.5883/DS-BARBONYM], accessed on 15 January 2021, reference number DS-BARBONYM.
Acknowledgments
We wish to thank Siti Nuramaliati Prijono, Witjaksono, Mohammad Irham, Marlina Adriyani, Ruliyana Susanti, Hari Sutrisno, Yuli Fitriana, Muhamad Syamsul Arifin Zein, late Sri Sulandari, and late Renny K. Hadiaty, at Research Centre for Biology (RCB-LIPI), as well as Joel Le Bail and Nicolas Gascoin at the French embassy in Jakarta for their continuous support. We are thankful to Daisy Wowor and Ujang Nurhaman at RCB-LIPI, and Sumanta and Bambang Dwisusilo at IRD Jakarta for their help during the field sampling. Finally, we acknowledge the reviewers who helped to improve this manuscript. A permit to collect fish was awarded to Nicolas Hubert (7/TKPIPA/FRP/SM/VII/2012, 68/EXT/SIP/FRP/SM/VIII/2013, 41/EXT/SIP/FRP/SM/VIII/2014, 361/SIP/FRP/E5/Dit.KI/IX/2015, 50/EXT/SIP/FRP/E5/Dit.KI/IX/2016, 45/EXT/SIP/FRP/E5/Dit.KI/VIII/2017, 392/SIP/FRP/E5/Dit.KI/XI/2018, and 200/E5/E5.4/SIP/2019). No experimentation was conducted on live specimens during this study. This publication has ISEM number 2021-135 SUD.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- Myers, N.; Mittermeier, R.A.; Mittermeier, C.G.; da Fonseca, G.A.B.; Kent, F. Biodiversity hotspots for conservation priorities. Nature 2000, 403, 853–858. [Google Scholar] [CrossRef]
- Mittermeier, R.A.; Turner, W.R.; Larsen, F.W.; Brooks, T.M.; Gascon, C. Global biodiversity conservation: The critical role of hotspots. In Biodiversity Hotspots; Springer: Berlin/Heidelberg, Germany, 2011; pp. 3–22. [Google Scholar]
- Hubert, N.; Wibowo, A.; Busson, F.; Caruso, D.; Sulandari, S.; Nafiqoh, N.; Pouyaud, L.; Rüber, L.; Avarre, J.C.; Herder, F.; et al. DNA barcoding Indonesian freshwater fishes: Challenges and prospects. DNA Barcodes 2015, 3, 144–169. [Google Scholar] [CrossRef] [Green Version]
- Imai, N.; Furukawa, T.; Tsujino, R.; Kitamura, S.; Yumoto, T. Factors affecting forest area change in Southeast Asia during 1980–2010. PLoS ONE 2018, 13, e0197391. [Google Scholar] [CrossRef] [Green Version]
- Laumonier, Y.; Uryu, Y.; Stüwe, M.; Budiman, A.; Setiabudi, B.; Hadian, O. Eco-floristic sectors and deforestation threats in Sumatra: Identifying new conservation area network priorities for ecosystem-based land use planning. Biodivers. Conserv. 2010, 19, 1153–1174. [Google Scholar] [CrossRef]
- Gaveau, D.L.A.; Sloan, S.; Molidena, E.; Yaen, H.; Sheil, D.; Abram, N.K.; Ancrenaz, M.; Nasi, R.; Quinones, M.; Wielaard, N. Four decades of forest persistence, clearance and logging on Borneo. PLoS ONE 2014, 9, e101654. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Garg, T.; Hamilton, S.E.; Hochard, J.P.; Kresch, E.P.; Talbot, J. (Not so) gently down the stream: River pollution and health in Indonesia. J. Environ. Econ. Manag. 2018, 92, 35–53. [Google Scholar] [CrossRef]
- Mulligan, M.; van Soesbergen, A.; Sáenz, L. GOODD, a global dataset of more than 38,000 georeferenced dams. Sci. Data 2020, 7, 1–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dahruddin, H.; Hutama, A.; Busson, F.; Sauri, S.; Hanner, R.; Keith, P.; Hadiaty, R.; Hubert, N. Revisiting the ichthyodiversity of Java and Bali through DNA barcodes: Taxonomic coverage, identification accuracy, cryptic diversity and identification of exotic species. Mol. Ecol. Resour. 2017, 17, 288–299. [Google Scholar] [CrossRef]
- FAO. Des Pêches et de L’aquaculture; FAO: Rome, Italy, 2018; ISBN 9789251306925. [Google Scholar]
- Koeshendrajana, S.; Cacho, O.J. Management Options for the Inland Fisheries Resource in South Sumatra, Indonesia: I Bioeconomic Model; Working Papers 12932; University of New England: Armidale, Australia, 2001. [Google Scholar]
- Coates, D. Inland capture fishery statistics of Southeast Asia: Current status and information needs. RAP Publ. 2002, 11, 114. [Google Scholar]
- Welcomme, R.L.; Baird, I.G.; Dudgeon, D.; Halls, A.; Lamberts, D.; Mustafa, M.G. Fisheries of the Rivers of Southeast Asia; Jon Wiley & Sons: Hoboken, NJ, USA, 2016; pp. 363–376. [Google Scholar]
- Muthmainnah, D.; Makmur, H.R.; Sawestri, S.; Supriyadi, F.; Fatah, K. The Features of Inland Fisheries in Southeast Asia: Inland Capture Fisheries and Its Status; Inland Fishery Resources Development and Management Department, Southeast: Samut Prakan, Thailand, 2019; ISBN 6024408161. [Google Scholar]
- Conte-Grand, C.; Britz, R.; Dahanukar, N.; Raghavan, R.; Pethiyagoda, R.; Tan, H.H.; Hadiaty, R.K.; Yaakob, N.S.; Rüber, L. Barcoding snakeheads (Teleostei, Channidae) revisited: Discovering greater species diversity and resolving perpetuated taxonomic confusions. PLoS ONE 2017, 12, e0184017. [Google Scholar] [CrossRef]
- Batubara, A.S.; Muchlisin, Z.A.; Efizon, D.; Elvyra, R.; Fadli, N.; RIZAL, S.; Siti-Azizah, M.N.; Wilkes, M. DNA barcoding (COI genetic marker) revealed hidden diversity of Cyprinid fish (Barbonymus spp.) from Aceh Waters, Indonesia. Biharean Biol. 2013, 15, 1. [Google Scholar]
- Kottelat, M. The fishes of the inland waters of Southeast Asia: A catalog and core bibliography of the fishes known to occur in freshwaters, mangroves and estuaries. Raffles Bull. Zool. 2013, (Suppl. 27), 1–663. [Google Scholar]
- Froese, R.; Pauly, D. Fishbase. Available online: http://www.fishbase.org (accessed on 15 January 2021).
- Eschmeyer, W.N.; Fricke, R.; van der Laan, R. Catalog of Fishes Electronic Version. Available online: http://www.calacademy.org/research/ichthyology/catalog/fishcatsearch.html (accessed on 15 January 2021).
- Kottelat, M.; Whitten, A.J.; Kartikasari, N.; Wirjoatmodjo, S. Freshwater Fishes of Western Indonesia and Sulawesi; Periplus Publishing Group: Hong Kong, China, 1993; ISBN 0945971605. [Google Scholar]
- Hubert, N.; Lumbantobing, D.; Sholihah, A.; Dahruddin, H.; Delrieu-Trottin, E.; Busson, F.; Sauri, S.; Hadiaty, R.; Keith, P. Revisiting species boundaries and distribution ranges of Nemacheilus spp. (Cypriniformes: Nemacheilidae) and Rasbora spp. (Cypriniformes: Cyprinidae) in Java, Bali and Lombok through DNA barcodes: Implications for conservation in a biodiversity hotspot. Conserv. Genet. 2019, 20, 517–529. [Google Scholar] [CrossRef]
- Sholihah, A.; Delrieu-Trottin, E.; Sukmono, T.; Dahruddin, H.; Risdawati, R.; Elvyra, R.; Wibowo, A.; Kustiati, K.; Busson, F.; Sauri, S.; et al. Disentangling the taxonomy of the subfamily Rasborinae (Cypriniformes, Danionidae) in Sundaland using DNA barcodes. Sci. Rep. 2020, 10, 2818. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, H.-C.; Abidin, M.Z.; Pulungan, C.P.; De Bruyn, M.; Mohd Nor, S.A. DNA barcoding reveals high cryptic diversity of freshwater halfbeak genus Hemirhamphodon from Sundaland. PLoS ONE 2016, 11, e0163596. [Google Scholar] [CrossRef] [PubMed]
- Beck, S.V.; Carvalho, G.R.; Barlow, A.; Ruber, L.; Hui Tan, H.; Nugroho, E.; Wowor, D.; Mohd Nor, S.A.; Herder, F.; Muchlisin, Z.A.; et al. Plio-Pleistocene phylogeography of the Southeast Asian Blue Panchax killifish, Aplocheilus panchax. PLoS ONE 2017, 12, e0179557. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nurul Farhana, S.; Muchlisin, Z.A.; Duong, T.Y.; Tanyaros, S.; Page, L.M.; Zhao, Y.; Adamson, E.A.S.; Khaironizam, M.Z.; de Bruyn, M.; Siti Azizah, M.N. Exploring hidden diversity in Southeast Asia’s Dermogenys spp. (Beloniformes: Zenarchopteridae) through DNA barcoding. Sci. Rep. 2018, 8, 10787. [Google Scholar] [CrossRef] [Green Version]
- Hutama, A.; Dahruddin, H.; Busson, F.; Sauri, S.; Keith, P.; Hadiaty, R.K.; Hanner, R.; Suryobroto, B.; Hubert, N. Identifying spatially concordant evolutionary significant units across multiple species through DNA barcodes: Application to the conservation genetics of the freshwater fishes of Java and Bali. Glob. Ecol. Conserv. 2017, 12, 170–187. [Google Scholar] [CrossRef]
- Delrieu-Trottin, E.; Durand, J.; Limmon, G.; Sukmono, T.; Sugeha, H.Y.; Chen, W.; Busson, F.; Borsa, P.; Dahruddin, H.; Sauri, S. Biodiversity inventory of the grey mullets (Actinopterygii: Mugilidae) of the Indo-Australian Archipelago through the iterative use of DNA-based species delimitation and specimen assignment methods. Evol. Appl. 2020, 13, 1451–1467. [Google Scholar] [CrossRef] [Green Version]
- Sholihah, A.; Delrieu-Trottin, E.; Condamine, F.L.; Wowor, D.; Rüber, L.; Pouyaud, L.; Agnèse, J.-F.; Hubert, N. Impact of Pleistocene Eustatic Fluctuations on Evolutionary Dynamics in Southeast Asian Biodiversity Hotspots. Syst. Biol. 2021. [Google Scholar] [CrossRef]
- Rahman, M.M.; Norén, M.; Mollah, A.R.; Kullander, S.O. Building a DNA barcode library for the freshwater fishes of Bangladesh. Sci. Rep. 2019, 9, 9382. [Google Scholar] [CrossRef] [PubMed]
- Aquino, L.M.G.; Tango, J.M.; Canoy, R.J.C.; Fontanilla, I.K.C.; Basiao, Z.U.; Ong, P.S.; Quilang, J.P. DNA barcoding of fishes of Laguna de Bay, Philippines. Mitochondrial DNA 2011, 22, 143–153. [Google Scholar] [CrossRef] [PubMed]
- Meganathan, P.; Austin, C.M.; Tam, S.M.; Chew, P.C.; Siow, R.; Rashid, Z.A.; Song, B.K. An Application of DNA Barcoding to the Malaysian Freshwater Fish Fauna: mtDNA COI Sequences Reveal Novel Haplotypes, Cryptic Species and Field-Based Misidentification; Monash University Malaysia: Selangor, Malaysia, 2015. [Google Scholar]
- Panprommin, D.; Soontornprasit, K.; Tuncharoen, S.; Iamchuen, N. Efficacy of DNA barcoding for the identification of larval fish species in the Upper and Middle Ing River, Thailand. Gene Rep. 2021, 23, 101057. [Google Scholar] [CrossRef]
- Barman, A.S.; Singh, M.; Singh, S.K.; Saha, H.; Singh, Y.J.; Laishram, M.; Pandey, P.K. DNA barcoding of freshwater fishes of Indo-Myanmar biodiversity hotspot. Sci. Rep. 2018, 8, 8579. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barman, A.S.; Singh, M.; Pandey, P.K. DNA barcoding and genetic diversity analyses of fishes of Kaladan River of Indo-Myanmar biodiversity hotspot. Mitochondrial DNA Part A 2018, 29, 367–378. [Google Scholar] [CrossRef]
- Esa, Y.B.; Siraj, S.S.; Daud, S.K.; Ryan, J.J.R.; Rahim, K.A.A.; Tan, S.G. Molecular systematics of mahseers (Cyprinidae) in Malaysia inferred from sequencing of a mitochondrial Cytochrome C Oxidase I (COI) gene. Pertanika J. Trop. Agric. Sci. 2008, 31, 263–269. [Google Scholar]
- Lakra, W.S.; Singh, M.; Goswami, M.; Gopalakrishnan, A.; Lal, K.K.; Mohindra, V.; Sarkar, U.K.; Punia, P.P.; Singh, K.V.; Bhatt, J.P. DNA barcoding Indian freshwater fishes. Mitochondrial DNA Part A 2016, 27, 4510–4517. [Google Scholar] [CrossRef]
- Ivanova, N.V.; Zemlak, T.S.; Hanner, R.H.; Hébert, P.D.N. Universal primers cocktails for fish DNA barcoding. Mol. Ecol. Notes 2007, 7, 544–548. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; deWaard, J.R.; Zakharov, E.; Prosser, S.W.J.; Sones, J.E.; McKeown, J.T.A.; Mantle, B.; La Salle, J. A DNA “barcode blitz”: Rapid digitization and sequencing of a natural history collection. PLoS ONE 2013, 8, e68535. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D.N. BOLD: The Barcode of Life Data System. Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef] [Green Version]
- Pons, J.; Barraclough, T.G.; Gomez-Zurita, J.; Cardoso, A.; Duran, D.P.; Hazell, S.; Kamoun, S.; Sumlin, W.D.; Vogler, A.P. Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Syst. Biol. 2006, 55, 595–606. [Google Scholar] [CrossRef] [Green Version]
- Puillandre, N.; Lambert, A.; Brouillet, S.; Achaz, G. ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Mol. Ecol. 2012, 21, 1864–1877. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D.N. A DNA-based registry for all animal species: The barcode index number (BIN) system. PLoS ONE 2013, 8, e66213. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kapli, P.; Zhang, J.; Kobert, K.; Pavlidis, P.; Stamatakis, A.; Flouri, T. Multi-rate Poisson Tree Processes for single-locus species delimitation under Maximum Likelihood and Markov Chain Monte Carlo. Bioinformatics 2017, 33, 1630–1638. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luo, A.; Ling, C.; Ho, S.Y.W.; Zhu, C.-D. Comparison of methods for molecular species delimitation across a range of speciation scenarios. Syst. Biol. 2018, 67, 830–846. [Google Scholar] [CrossRef] [Green Version]
- Kekkonen, M.; Mutanen, M.; Kaila, L.; Nieminen, M.; Hebert, P.D.N. Delineating Species with DNA Barcodes: A Case of Taxon Dependent Method Performance in Moths. PLoS ONE 2015, 10, e0122481. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kekkonen, M.; Hebert, P.D.N. DNA barcode-based delineation of putative species: Efficient start for taxonomic workflows. Mol. Ecol. Resour. 2014, 14, 706–715. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Hubert, N.; Huang, Y.; Wang, X.; Gan, X.; Peng, Z.; He, S. DNA barcoding the ichthyofauna of the Yangtze River: Insights from the molecular inventory of a mega-diverse temperate fauna. Mol. Ecol. Resour. 2019, 19, 1278–1291. [Google Scholar] [CrossRef]
- Limmon, G.; Delrieu-Trottin, E.; Patikawa, J.; Rijoly, F.; Dahruddin, H.; Busson, F.; Steinke, D.; Hubert, N. Assessing species diversity of Coral Triangle artisanal fisheries: A DNA barcode reference library for the shore fishes retailed at Ambon harbor (Indonesia). Ecol. Evol. 2020, 10, 3356–3366. [Google Scholar] [CrossRef]
- Zhang, J.; Kapli, P.; Pavlidis, P.; Stamatakis, A. A general species delimitation method with applications to phylogenetic placements. Bioinformatics 2013, 29, 2869–2876. [Google Scholar] [CrossRef] [Green Version]
- Fujisawa, T.; Barraclough, T.G. Delimiting species using single-locus data and the generalized mixed Yule coalescent approach: A revised method and evaluation on simulated data sets. Syst. Biol. 2013, 62, 707–724. [Google Scholar] [CrossRef] [Green Version]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Bouckaert, R.; Heled, J.; Kühnert, D.; Vaughan, T.; Wu, C.H.; Xie, D.; Suchard, M.A.; Rambaut, A.; Drummond, A.J. BEAST 2: A Software Platform for Bayesian Evolutionary Analysis. PLoS Comput. Biol. 2014, 10, e1003537. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bermingham, E.; McCafferty, S.; Martin, A.P. Fish biogeography and molecular clocks: Perspectives from the Panamanian isthmus. In Molecular Systematics of Fishes; Kocher, T.D., Stepien, C.A., Eds.; Academic Press: San Diego, CA, USA, 1997; pp. 113–128. [Google Scholar]
- Ogilvie, H.A.; Bouckaert, R.R.; Drummond, A.J. StarBEAST2 brings faster species tree inference and accurate estimates of substitution rates. Mol. Biol. Evol. 2017, 34, 2101–2114. [Google Scholar] [CrossRef]
- Ho, S.Y.W.; Larson, G. Molecular clocks: When timesare a-changin’. Trends Genet. 2006, 22, 79–83. [Google Scholar] [CrossRef]
- Kimura, M. A Simple Method for Estimating Evolutionary Rates of Base Substitutions through Comparative Studies of Nucleotide-Sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Paradis, E.; Schliep, K. ape 5.0: An environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 2019, 35, 526–528. [Google Scholar] [CrossRef]
- Brown, S.D.J.; Collins, R.A.; Boyer, S.; Lefort, C.; Malumbres-Olarte, J.; Vink, C.J.; Cruickshank, R.H. Spider: An R package for the analysis of species identity and evolution, with particular reference to DNA barcoding. Mol. Ecol. Resour. 2012, 12, 562–565. [Google Scholar] [CrossRef]
- Paradis, E. pegas: An {R} package for population genetics with an integrated--modular approach. Bioinformatics 2010, 26, 419–420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blair, C.; Bryson, J.R.W. Cryptic diversity and discordance in single-locus species delimitation methods within horned lizards (Phrynosomatidae: Phrynosoma). Mol. Ecol. Resour. 2017, 17, 1168–1182. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, Q.; Liu, S.; Yu, D.; Liu, H.; Danley, P.D. Mitochondrial capture and incomplete lineage sorting in the diversification of balitorine loaches (Cypriniformes, Balitoridae) revealed by mitochondrial and nuclear genes. Zool. Scr. 2012, 41, 233–247. [Google Scholar] [CrossRef]
- Hopkins, R.L.; Eisenhour, D.J. Hybridization of Lythrurus fasciolaris and Lythrurus umbratilis (Cypriniformes: Cyprinidae) in the Ohio River basin. Copeia 2008, 2008, 162–171. [Google Scholar] [CrossRef]
- Atsumi, K.; Nomoto, K.; Machida, Y.; Ichimura, M.; Koizumi, I. No reduction of hatching rates among F1 hybrids of naturally hybridizing three Far Eastern daces, genus Tribolodon (Cypriniformes, Cyprinidae). Ichthyol. Res. 2018, 65, 165–167. [Google Scholar] [CrossRef]
- Hubert, N.; Hanner, R. DNA barcoding, species delineation and taxonomy: A historical perspective. DNA Barcodes 2015, 3, 44–58. [Google Scholar] [CrossRef] [Green Version]
- Wibowo, A.; Atminarso, D.; Baumgartner, L.; Vasemagi, A. High prevalence of non-native fish species in a remote region of the Mamberamo River, Indonesia. Pac. Conserv. Biol. 2020, 26, 293–300. [Google Scholar] [CrossRef]
- Keith, P.; Lord, C.; Darhuddin, H.; Limmon, G.; Sukmono, T.; Hadiaty, R.; Hubert, N. Schismatogobius (Gobiidae) from Indonesia, with description of four new species. Cybium 2017, 41, 195–211. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).