Karstic Landscapes Are Foci of Species Diversity in the World’s Third-Largest Vertebrate Genus Cyrtodactylus Gray, 1827 (Reptilia: Squamata; Gekkonidae)
Abstract
1. Introduction
2. Materials and Methods
2.1. Habitat Preferences and Ecotypes
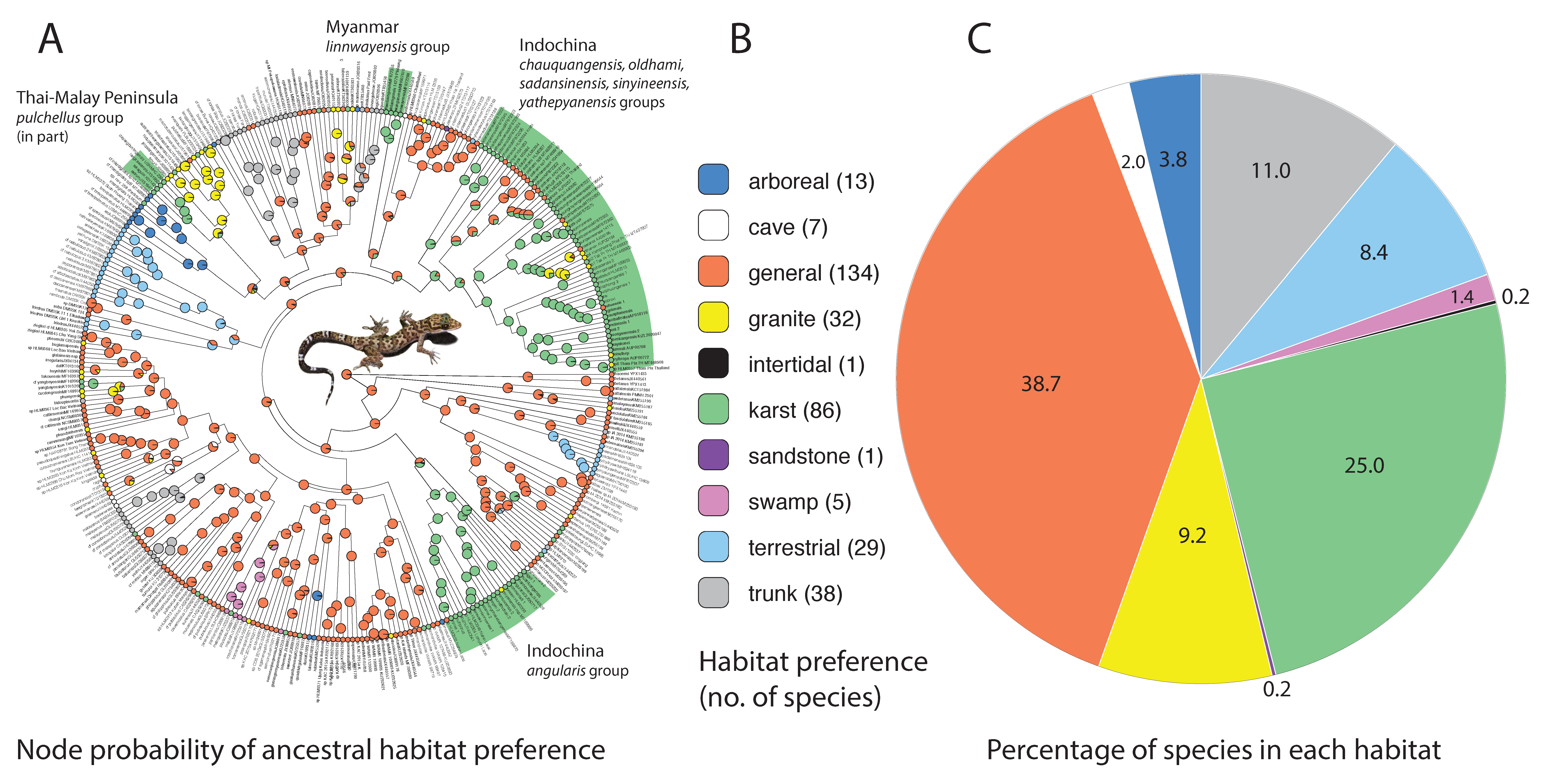
- General (Figure 4A). Species that use the majority of the microhabitats in their immediate surroundings in whatever environment they inhabit. The microhabitats may include rocks of all types (when present), logs, tree trunks (with or without holes and crevices), and all vegetative structures of various dimensions, the ground, and human-made structures in many cases. No particular microhabitat is notably preferred over any other although some species may be most often observed in low vegetation.
- Trunk (Figure 4B). These are species generally found on the trunks and large branches of large trees at varying heights and often take refuge in cracks, crevices, or holes in the trunks. They may occasionally occur on large granite rocks but only if the rocks are near the trees. These species are generally the largest and most robust species in the genus [22,23,24]. None have been reported to have prehensile tails although some species may coil the tail horizontally similar to that seen in arboreal species.
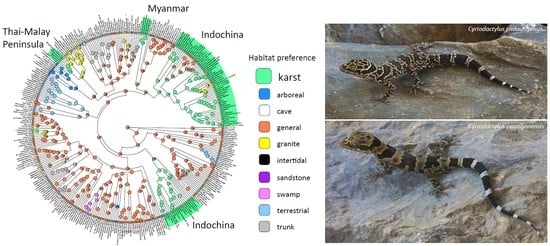
- Karst (Figure 4C). These are generally more gracile species that are restricted to habitats where limestone rock (karst) is present. Individuals use this substrate (including cliff faces, small rocks, and boulders) as well as adjacent vegetation. If caves are present, they will enter only into the twilight zone and usually no deeper than 50 m from the entrance [14]. Despite what has been written about many karst-associated species being cave species or cave adapted (e.g., [25]), none truly are and most are more commonly found on the outside of caves (see below). These species do not occur in habitats lacking karstic substrates.
- Cave (Figure 4D). These are species that occur exclusively in the cave-like environments formed by large granite boulders. Open spaces between the boulders can be quite extensive and contain areas where very little light penetrates. These species rarely occur on the out-facing (i.e., the forest-side) surfaces of the boulders and for the most part, are restricted to the spaces between the boulders at varying depths below the surface of the ground in extremely low levels of illumination. These are truly cave-adapted species with notably thin, gracile bodies, long limbs, flat heads, large eyes, and faded color patterns [13,26,27].
- Terrestrial (Figure 4E). These are species that generally occur only on the ground and may take refuge beneath natural and human-made surface objects. They may occasionally be found on the tops of small rocks (when present) or on the bases of small trees and shrubs but never higher than 1 m above the ground. These species are relatively small and notably squat, with short fat tails, thick heads, and short digits [28,29].
- Arboreal (Figure 4F). These are cryptically colored species [30,31] generally restricted to small branches, leaves, trunks of varying sizes, and shrubs. Some may take refuge beneath exfoliating bark often as high or higher than three meters above the ground. These species are rarely observed on the ground or lower than 1.5 m above the ground. In such instances, it is usually during windy and/or rainy nights (perhaps forced down from higher up; [32]; authors pers. obs.) or during egg laying. All species have a prehensile tail used as a climbing aid [31,32,33] that is often carried in a coiled, elevated position.
- Granite (Figure 4H). These are generally more robust, strongly tuberculated species found in forested habitats bearing large granite boulders (not just small, scattered, granite rocks or rocks of other types). Vegetation is often used, especially by hatchlings and juveniles, but individuals occur more commonly on the granite boulders in all planes of orientation. These species do not occur in forested areas lacking granite boulders.
- Intertidal (Figure 4I). This category contains a single species that occurs exclusively in the rocky intertidal zones of small islands in the Seribuat Archipelago off the southeastern coast of Peninsular Malaysia and avoids nearby forested regions even if they lack other species of Cyrtodactylus [19,36].
- Sandstone (Figure 4J). This category was not included in Grismer et al. [6]. It contains a single species endemic to a forested sandstone massif isolated in the lowlands of northwestern Cambodia [11]. This species is known to forage only on the surface or within crevices of sandstone rocks and was not observed on the nearby vegetation [37]. This species is similar in body shape to closely related granite-associated species (Grismer unpublished).
2.2. Mitochondrial DNA
2.3. Phylogenetic Analyses
2.4. Ancestral State Reconstruction
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tolentino, P.J.; Navidad, J.R.L.; Angeles, M.D.; Fernandez, D.A.P.; Villanueva, E.L.C.; Obeña, R.D.R.; Buot, J.I.E. Review: Biodiversity of forests over limestone in Southeast Asia with emphasis on the Philippines. Biodiversitas J. Biol. Divers. 2020, 21, 1597–1613. [Google Scholar] [CrossRef]
- Sodhi, N.S.; Koh, L.P.; Brook, B.W.; Ng, P.K.L. Southeast Asian biodiversity: An impending disaster. Trends Ecol. Evol. 2004, 19, 654–660. [Google Scholar] [CrossRef] [PubMed]
- Sodhi, N.S.; Posa, M.R.C.; Lee, T.M.; Bickford, D.; Koh, L.P.; Brook, B.W. The state and conservation of Southeast Asian biodiversity. Biodivers. Conserv. 2010, 19, 317–328. [Google Scholar] [CrossRef]
- Clements, R.; Ng, P.K.; Lu, X.X.; Ambu, S.; Schilthuizen, M.; Bradshaw, C.J. Using biogeographical patterns of endemic land snails to improve conservation planning for limestone karsts. Biol. Conserv. 2008, 141, 2751–2764. [Google Scholar] [CrossRef]
- Clements, R.; Sodhi, N.S.; Schilthuizen, M.; Ng, P.K.L. Limestone karst of Southeast Asia: Imperiled arks of biodiversity. Bioscience 2006, 56, 733–742. [Google Scholar] [CrossRef]
- Grismer, L.L.; Wood, P.L., Jr.; Le, M.D.; Quah, E.S.H.; Grismer, J.L. Evolution of habitat preference in 243 species of Bent--toed geckos (Genus Cyrtodactylus Gray, 1827) with a discussion of karst habitat conservation. Ecol. Evol. 2020, 10, 13717–13730. [Google Scholar] [CrossRef]
- Barjadze, S.; Murvanidze, M.; Arabuli, T.; van Mumladze, L.; Pkhakadze, V.; Djanashvili, R.; Salakaia, M. Annotated List of Inver-tebrates of the Georgian Karst Caves; Georgian Academic Book: Tbilisi, Georgia, 2015; pp. 1–120. [Google Scholar]
- Chen, W.H.; Zhang, Y.M.; Li, Z.Y.; Nguyen, Q.H.; Nguyen, T.H.; Shui, Y. Hemiboea crystallina, a new species of Gesneriaceae from karst regions of China and Vietnam. Phytotaxa 2018, 336, 23–56. [Google Scholar] [CrossRef]
- Chin, S.C. The limestone hill flora of Malaya: Part 1. Gard. Bull. Sing. 1977, 30, 165–220. [Google Scholar]
- Chung, W.-H.; Zhang, Y.-M.; Li, Z.-Y.; Nguyen, Q.-H.; Nguyen, T.-H.; Shui, Y.-M. Phylogenetic analyses of Begonia sect. Coelocentrum and allied limestone species of China shed light on the evolution of Sino-Vietnamese karst flora. Bot. Stud. 2014, 55, 1–35. [Google Scholar] [CrossRef]
- Grismer, L.L.; Wood, P.L.; Poyarkov, N.A.; Le, M.D.; Kraus, F.; Agarwal, I.; Oliver, P.M.; Nguyen, S.N.; Nguyen, T.Q.; Karuna-rathna, S.; et al. Phylogenetic partitioning of the third-largest vertebrate genus in the world, Cyrtodactylus Gray, 1827 (Reptilia; Squamata; Gekkonidae) and its relevance to taxonomy and conservation. Vert. Zool. 2021, 71, 101–154. [Google Scholar] [CrossRef]
- Uetz, P.; Freed, P.; Hošek, J. The Reptile Database. Available online: http://www.reptile-database.org (accessed on 14 February 2021).
- Grismer, L.L.; Grismer, J.L. A re-evaluation of the phylogenetic relationships of the Cyrtodactylus condorensis group (Squamata; Gekkonidae) and a suggested protocol for the characterization of rock-dwelling ecomorphology in Cyrtodactylus. Zootaxa 2017, 4300, 486–504. [Google Scholar] [CrossRef]
- Grismer, L.L.; Wood, P.L.; Thura, M.K.; Zin, T.; Quah, E.S.H.; Murdoch, M.L.; Grismer, M.S.; Lin, A.; Kyaw, H.; Lwin, N. Twelve new species of Cyrtodactylus Gray (Squamata: Gekkonidae) from isolated limestone habitats in east-central and southern Myanmar demonstrate high localized diversity and unprecedented microendemism. Zool. J. Linn. Soc. 2017, 182, 862–959. [Google Scholar] [CrossRef]
- Hikida, T. Bornean Gekkonid Lizards of the Genus Cyrtodactylus (Lacertilia: Gekkonidae). Jpn. J. Herpetol. 1990, 13, 91–107. [Google Scholar] [CrossRef][Green Version]
- Johnson, C.B.; Evan Quah, S.H.; Anuar, S.; Muin, M.A.; Wood, J.P.L.; Grismer, J.L.; Greer, L.F.; Onn, C.K.; Ahmad, N.; Bauer, A.M.; et al. Phylogeography, geographic variation, and taxonomy of the Bent-toed Gecko Cyrtodactylus quadrivirgatus Taylor, 1962 from Peninsular Malaysia with the description of a new swamp dwelling species. Zootaxa 2012, 3406, 39–58. [Google Scholar] [CrossRef]
- Kraus, F. Taxonomic partitioning of Cyrtodactylus louisiadensis (Lacertilia: Gekkonidae) from Papua New Guinea. Zootaxa 2008, 1883, 1–27. [Google Scholar] [CrossRef]
- Welton, L.J.; Siler, C.D.; Diesmos, A.C.; Brown, R.M. Phylogeny-based species delimitation of southern Philippines bent-toed geckos and a new species of Cyrtodactylus (Squamata: Gekkonidae) from western Mindanao and the Sulu Archipelago. Zootaxa 2010, 2390, 49–68. [Google Scholar] [CrossRef]
- Grismer, L.L. Lizards of Peninsular Malaysia, Singapore and Their Adjacent Archipelagos; Edition Chimaira: Frankfürt am Main, Germany, 2011; pp. 1–728. [Google Scholar]
- Grismer, L.L.; Wood, P.L., Jr.; Thura, M.K.; Quah, E.S.; Murdoch, M.L.; Grismer, M.S.; Herr, M.W.; Lin, A.; Kyaw, H. Three more new species of Cyrtodactylus (Squamata: Gekkonidae) from the Salween Basin of eastern Myanmar underscore the urgent need for the conservation of karst habitats. J. Nat. Hist. 2018, 52, 1243–1294. [Google Scholar] [CrossRef]
- Grismer, L.L.; Wood, P.L., Jr.; Quah, E.S.H.; Grismer, M.S.; Thura, M.K.; Oaks, J.R.; Lin, A. Two new species of Cyrtodactylus Gray, 1827 (Squamata: Gekkonidae) from a karstic archipelago in the Salween Basin of southern Myanmar (Burma). Zootaxa 2020, 4718, 151–183. [Google Scholar] [CrossRef]
- Nielsen, S.V.; Oliver, P.M. Morphological and genetic evidence for a new karst specialist lizard from New Guinea (Cyrtodactylus: Gekkonidae). R. Soc. Open Sci. 2017, 4, 170781. [Google Scholar] [CrossRef]
- Oliver, P.; Krey, K.; Mumpuni; Richards, S. A new species of bent-toed gecko (Cyrtodactylus, Gekkonidae) from the North Papuan Mountains. Zootaxa 2011, 2930, 22–32. [Google Scholar] [CrossRef]
- Oliver, P.M.; Richards, S.J.; Mumpuni, M.; Rösler, H. The Knight and the King: Two new species of giant bent-toed gecko (Cyrtodactylus, Gekkonidae, Squamata) from northern New Guinea, with comments on endemism in the North Papuan Mountains. ZooKeys 2016, 562, 105–130. [Google Scholar] [CrossRef] [PubMed]
- Ellis, M.; Pauwels, O.S.G. The bent-toed geckos (Cyrtodactylus) of the caves and karst of Thailand. Cave Karst Sci. 2012, 39, 16–22. [Google Scholar]
- Ngo, V.T. Two new cave-dwelling species of Cyrtodactylus Gray (Squamata: Gekkonidae) from Southwestern Vietnam. Zootaxa 2008, 1909, 37–51. [Google Scholar] [CrossRef]
- Nguyen, S.N.; Orlov, N.L.; Darevsky, I.S. Descriptions of two new species of the genus Cyrtodactylus Gray, 1827 (Squamata: Sauria: Gekkonidae) from southern Vietnam. J. Herpetol. 2006, 13, 215–226. [Google Scholar] [CrossRef]
- Agarwal, I. Two new species of ground-dwelling Cyrtodactylus (Geckoella) from the Mysore Plateau, south India. Zootaxa 2016, 4193, 228–244. [Google Scholar] [CrossRef] [PubMed]
- Grismer, L.L.; Wood, P.L., Jr.; Thura, M.K.; Win, N.M.; Quah, E.S.H. Two more new species of the Cyrtodactylus peguensis group (Squamata: Gekkonidae) from the fringes of the Ayeyarwady Basin, Myanmar. Zootaxa 2019, 4577, 274–294. [Google Scholar] [CrossRef] [PubMed]
- Grismer, L.L.; Shahrul, A.; Quah, E.; Muin, M.A.; Chan, K.O.; Grismer, J.L.; Ahmad, N. A new spiny, prehensile-tailed species of Cyrtodactylus (Squamata: Gekkonidae) from Peninsular Malaysia with a preliminary hypothesis of relationships based on morphology. Zootaxa 2010, 2625, 40–52. [Google Scholar] [CrossRef]
- Harvey, M.B.; O’connell, K.A.; Wostl, E.; Riyanto, A.; Kurniawan, N.; Smith, E.N.; Grismer, L.L. Redescription of Cyrtodactylus lateralis (Werner) (Squamata: Gekkonidae) and phylogeny of the prehensile-tailed Cyrtodactylus. Zootaxa 2016, 4107, 517–540. [Google Scholar] [CrossRef] [PubMed]
- Dring, J.C.M. Amphibians and reptiles from northern Trengganu, Malaysia, with descriptions of two new geckos: Cnemaspis and Cyrtodactylus. Bull. British Mus. (Nat. Hist.) 1979, 34, 181–241. [Google Scholar]
- Grismer, L.L. On the distribution and identification of Cyrtodactylus brevipalmatus Smith, 1923, and Cyrtodactylus elok, Dring, 1979. Raffles Bull. Zool. 2008, 56, 177–179. [Google Scholar] [CrossRef]
- Grismer, L.L.; Wood, P.L., Jr.; Lim, K.K.P. Cyrtodactylus majulah, a new species of bent-toed gecko (Reptilia: Squamata: Gek-konidae) from Singapore and the Riau Archipelago. Raffles Bull. Zool. 2012, 60, 487–499. [Google Scholar]
- Riyanto, A.; Grismer, L.L.; Wood, P.L., Jr. Cyrtodactylus rosichonariefi sp. nov. (Squamata: Gekkonidae), a new swamp-dwelling bent-toed gecko from Bunguran Island (Great Natuna), Indonesia. Zootaxa 2015, 3964, 114–124. [Google Scholar] [CrossRef] [PubMed]
- Youmans, T.M.; Grismer, L.L. A new species of Cyrtodactylus (Reptilia: Squamata: Gekkonidae) from the Seribuat Archipelago, West Malaysia. Herpetol. Nat. Hist. 2006, 10, 61–70. [Google Scholar]
- Geissler, P.; Hartmann, T.; Ihlow, F.; Neang, T.; Seng, R.; Wagner, P.; Böhme, W. Herpetofauna of the Phnom Kulen National Park, northern Cambodia—An annotated checklist. Cambodian J. Nat. Hist. 2019, 1, 40–63. [Google Scholar]
- Wood, P.L., Jr.; Heinicke, M.P.; Jackman, T.R.; Bauer, A.M. Phylogeny of bent-toed geckos (Cyrtodactylus) reveals a west to east pattern of diversification. Mol. Phylo. Evol. 2012, 65, 992–1003. [Google Scholar] [CrossRef]
- Hillis, D.M.; Moritz, C.; Mable, B.K.; Graur, D. Molecular Systematics, 2nd ed.; Sinauer Associates, Inc.: Sunderland, MA, USA, 1996; pp. 1–456. [Google Scholar] [CrossRef]
- Katoh, M.; Kuma, M. MAFTT: A novel method for rapid sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [PubMed]
- Maddison, W.P.; Maddison, D.R. Mesquite: A Modular System for Evolutionary Analysis. Version 3.04. 2015. Available online: http://mesquiteproject.org (accessed on 23 December 2020).
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Trifinopoulos, J.; Nguyen, L.-T.; von Haeseler, A.; Minh, B.Q. W-IQ-TREE: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016, 44, W232–W235. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.; von Haeseler, A.; Jermiin, L.S. Model Finder: Fast model selection for accurate phylogenetic estimates. Nat. Meth. 2017, 14, 587. [Google Scholar] [CrossRef]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef] [PubMed]
- Minh, B.Q.; Nguyen, M.A.T.; von Haeseler, A. Ultrafast approximation for phylogenetic bootstrap. Mol. Biol. Evol. 2013, 30, 1188–1195. [Google Scholar] [CrossRef] [PubMed]
- Bouckaert, R.; Heled, J.; Kühnert, D.; Vaughan, T.; Wu, C.-H.; Xie, D.; Suchard, M.A.; Rambaut, A.; Drummond, A.J. BEAST 2: A Software Platform for Bayesian Evolutionary Analysis. PLoS Comput. Biol. 2014, 10, e1003537. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the Gateway Computing Environments Workshop (GCE), New Orleans, LA, USA, 14 November 2010; pp. 1–8. [Google Scholar]
- Rambaut, A.; Suchard, M.A.; Xie, D.; Drummond, A.J. Tracer v1. 2014. Available online: https://bioweb.pasteur.fr/packages/pack@Tracer@v1.6 (accessed on 23 March 2021).
- Rambaut, A.; Drummond, A.J. TreeAnnotator v1. 7.0. 2013. Available online: https://bioweb.pasteur.fr/packages/pack@Tracer@v1.6 (accessed on 23 March 2021).
- Huelsenbeck, J.P.; Ronquist, F.; Nielsen, R.; Bollback, J.P. Bayesian inference of phylogeny and its impact on evolutionary biol- ogy. Science 2001, 294, 2310–2314. [Google Scholar] [CrossRef]
- Wilcox, T.P.; Zwickl, D.J.; Heath, T.A.; Hillis, D.M. Phylogenetic relationships of the Dwarf Boas and a comparison of Bayesian and bootstrap measures of phylogenetic support. Mol. Phylo. Evol. 2002, 25, 361–371. [Google Scholar] [CrossRef]
- Revell, L.J. Phytools: An R package for phylogenetic comparative biology (and other things). Meth. Ecol. Evol. 2012, 3, 217–223. [Google Scholar] [CrossRef]
- Paradis, E.; Schilep, K. Ape 5.0: An environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 2018, 35, 526–528. [Google Scholar] [CrossRef]
- Che, J.; Jiang, K.; Yan, F.; Zhang, Y.-P. Amphibians and Reptiles of Tibet-Diversity and Evolution; Science Press: Beijing, China, 2020; pp. 1–792. (In Chinese) [Google Scholar]
- Do, T. Characteristics of karst ecosystems of Vietnam and their vulnerability to human impact. Acta Geol. Sin. 2002, 75, 325–329. [Google Scholar] [CrossRef]
- Schilthuizen, M.; Liew, T.-S.; Bin Elahan, B.; Lackman-Ancrenaz, I. Effects of karst forest degradation on pulmonate and proso-branch land snail communities in Sabah, Malaysian Borneo. Conserv. Biol. 2005, 19, 949–954. [Google Scholar] [CrossRef]
- Sterling, E.J.; Hurley, M.M.; Le, M.D. Vietnam: A Natural History; Yale University Press: New Haven, CT, USA, 2006; pp. 1–456. [Google Scholar]
- Latinne, A.; Waengsothorn, S.; Herbreteau, V.; Michau, J.R.; Baillarguet, C. Thai limestone karsts: An impending biodiversity crisis. In Proceedings of the International Conference on Environment Supporting Food Energy Security: Crisis Opportunity, Bangkok, Thailand, 22–25 March 2001; pp. 176–187. [Google Scholar]
- Grismer, L.L.; Wood, P.L., Jr.; Anuar, S.; Davis, H.R.; Cobos, A.J.; Murdoch, M.L. A new species of karst forest Bent-toed Gecko (genus Cyrtodactylus Gray) not yet threatened by foreign cement companies and a summary of Peninsular Malaysia’s endemic karst forest herpetofauna and the need for its conservation. Zootaxa 2016, 4061, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Luo, Z.; Tang, S.; Jiang, Z.; Chen, J.; Fang, H.; Li, C. Conservation of terrestrial vertebrates in a global hotspot of karst in south-western China. Sci. Rep. 2016, 6, 25717. [Google Scholar] [CrossRef]
- von Oheimb, P.V.; von Oheimb, K.C.M.; Hirano, T.; Do, V.T.; Luong, H.V.; Ablett, J.; Pham, S.V.; Naggs, F. Competition matters: Determining the drivers of land snail community assembly among limestone karst areas in northern Vietnam. Ecol. Evol. 2017, 28, 4136–4149. [Google Scholar] [CrossRef] [PubMed]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grismer, L.; Wood, P.L.; Poyarkov, N.A.; Le, M.D.; Karunarathna, S.; Chomdej, S.; Suwannapoom, C.; Qi, S.; Liu, S.; Che, J.; et al. Karstic Landscapes Are Foci of Species Diversity in the World’s Third-Largest Vertebrate Genus Cyrtodactylus Gray, 1827 (Reptilia: Squamata; Gekkonidae). Diversity 2021, 13, 183. https://doi.org/10.3390/d13050183
Grismer L, Wood PL, Poyarkov NA, Le MD, Karunarathna S, Chomdej S, Suwannapoom C, Qi S, Liu S, Che J, et al. Karstic Landscapes Are Foci of Species Diversity in the World’s Third-Largest Vertebrate Genus Cyrtodactylus Gray, 1827 (Reptilia: Squamata; Gekkonidae). Diversity. 2021; 13(5):183. https://doi.org/10.3390/d13050183
Chicago/Turabian StyleGrismer, Lee, Perry L. Wood, Nikolay A. Poyarkov, Minh D. Le, Suranjan Karunarathna, Siriwadee Chomdej, Chatmongkon Suwannapoom, Shuo Qi, Shuo Liu, Jing Che, and et al. 2021. "Karstic Landscapes Are Foci of Species Diversity in the World’s Third-Largest Vertebrate Genus Cyrtodactylus Gray, 1827 (Reptilia: Squamata; Gekkonidae)" Diversity 13, no. 5: 183. https://doi.org/10.3390/d13050183
APA StyleGrismer, L., Wood, P. L., Poyarkov, N. A., Le, M. D., Karunarathna, S., Chomdej, S., Suwannapoom, C., Qi, S., Liu, S., Che, J., Quah, E. S. H., Kraus, F., Oliver, P. M., Riyanto, A., Pauwels, O. S. G., & Grismer, J. L. (2021). Karstic Landscapes Are Foci of Species Diversity in the World’s Third-Largest Vertebrate Genus Cyrtodactylus Gray, 1827 (Reptilia: Squamata; Gekkonidae). Diversity, 13(5), 183. https://doi.org/10.3390/d13050183