Abstract
The ectomycorrhizal basidiomycetes genus Thelephora has been understudied in subtropical ecosystems. Many species of Thelephora are important edible and medicinal fungi, with substantial economic value. Two new Thelephora species, T. grandinioides and T. wuliangshanensis spp. nov. are proposed here based on a combination of morphological features and molecular evidence. Thelephora grandinioides is characterized by laterally stipitate basidiocarps with a grandinoid hymenial surface, a monomitic hyphal system with clamped generative hyphae, and the presence of tubular and septated cystidia and subglobose to globose basidiospores measuring as 5.3–7.4 × 4–6.5 µm. Thelephora wuliangshanensis is characterized by infundibuliform basidiocarps, radially black striate on the pileus, a smooth, umber to coffee hymenial surface, a monomitic hyphal system with thick-walled generative hyphae, and basidiospores that turn greenish grey to buff in 5% KOH. Phylogenetic analyses of rDNA internal transcribed spacer region (ITS) and nuclear large subunit region (nrLSU) showed that the two new Thelephora are phylogenetically distinct: T. grandinioides is sister to T. aurantiotincta and T. sikkimensis, while T. wuliangshanensis is sister to a clade comprising T. austrosinensis and T. aurantiotincta with high support as well.
1. Introduction
Thelephora Ehrh. ex Willd., the genus type of Thelephoraceae Chevall. is one of the most important taxa in basidiomycetes [1,2,3,4,5]. They are widely distributed worldwide, especially in the northern temperate and tropical regions [1,2,3,4,5]. Thelephora is a fairly well-studied ectomycorrhizal basidiomycetes genus with basidiocarps of various shape; the entire genus forms ectomycorrhizal relationships with diverse plants and significant contribution to plant health and ecosystem stability [4,6,7,8,9,10,11,12]. As mycorrhiza-formers, Thelephora play a very important role in pioneer microhabitats of coniferous forests [13,14]. Acting as white rot fungi, they also can decompose dead wood [14,15].
Some species of Thelephora are economically important edible and medicinal mushrooms. Thelephora ganbajun M. Zang is one of the most popular edible fungi in China and some East Asian countries [14,15,16,17,18,19,20,21]. Thelephora is typified by T. terrestris Ehrh. ex Willd. [22], and the genus is characterized by its diverse forms of basidiomycetes as stereoid, clavarioid, cantharelloid, spathulate, pleuropodally pileate to resupinate; hymenophore smooth to slightly wrinkled and often cyanescent in KOH; pileus surface glabrous to strigose, even or faintly ribbed or papillose; hymenium continuous, usually on inferior side, sometimes amphigenous in some species; hyphal system monomitic with clamped generative hyphae; basidia 4-spored; basidiospores subhyaline to brownish, ornamented, typically muricate, verruculose or echinulate, even or slightly rough-walled in a few species, and inamyloid [1,3,12,22]. As of 2008, fifty species of Thelephora have been accepted [23], and some new species have been reported in recent years [4,5,14,24,25,26,27]. Index Fungorum [28] shows 871 specific and infraspecific names in Thelephora. However, to date, 62 species of Thelephora have been accepted [3,4,5,14,23,24,25,26,27].
Thelephora share similar characteristics with Tomentella Pers. ex Pat. especially in the form, size, and type of spore ornamentations [3,29,30]. Based on phylogenetic analyses using rDNA internal transcribed spacer region (ITS) sequences showed that the species of Thelephora mixed with Tomentella, revealing that both genera are closely related, but it is well-known that the phylogenetic analyses of ITS loci are insufficient to resolve phylogenetic relationships among closely related taxa [4,5,31,32]. Based on ITS and nrLSU analyses, Vizzini et al. [25] showed that Thelephora and Tomentella species do not separate to two monophyletic groups but they are intermixed and form a well-supported monophyletic clade (Thelephora/Tomentella clade). Back to traditional method, the most important characteristic for distinguishing Thelephora and Tomentella is the form of the basidiocarps (resupinate in Tomentella; erect, with varied forms, to partially resupinate in Thelephora) [24,29,33]. Das et al. [26] proposed that other features such as the hymenophore surface needed to be observed to determine whether it could act as a more informative characteristic than the highly variable stipitate/resupinate configuration of basidiocarps. Phylogenetic analyses of combined ITS and nrLSU dataset in Basidiomycota revealed that Thelephora is sister to Tomentella nested in Thelephoraceae while the limits between both genera are not yet clear [5,26,34]. While ITS and nrLSU sequences alone cannot resolve phylogenetic relationships in this complex group of species [5,34]. Vizzini et al. [25] mentioned that in the future Thelephora and Tomentella will be considered as one genus merging Tomentella into Thelephora.
With this work we intend to identify two Thelephora species found in southern China based on morphology and phylogeny, and provide full descriptions, color photographs, a detailed comparison of two new species with closely related taxa and a phylogenetic tree to show the placement of two new species.
2. Materials and Methods
2.1. Specimens Collection and Herbarium Specimen Preparation
Four samples of Thelephora were collected in Yunnan (Figure 1A–C) viz. CLZhao 3406 (Holotype) from the Wuliangshan National Nature Reserve, Huangcaoling, Jingdong County, Puer, at latitude 24°18′ N and longitude 101°05′ E, at 2113 m above sea level, 1 October 2017; CLZhao 3408 from the Xieqipo Forest Park, Zhenyuan County, Puer, at latitude 24°18′ N and longitude 101°05′ E, at 1350 m above sea level, 1 October 2017; CLZhao 4107 (Holotype) from the Wuliangshan National Nature Reserve, Huangcaoling, Jingdong County, Puer, at latitude 24°23′ N and longitude 100°45′ E, at 2313 m above sea level, 5 October 2017; CLZhao 21020 from the Wuliangshan National Nature Reserve, Huangcaoling, Jingdong County, Puer, at latitude 23°57′ N and longitude 100°57′ E, 8 October 2020. The fruiting bodies were observed growing on the ground of pine-broadleaved mixed forest. Photographs of the fruiting bodies were taken in the field, macromorphological characteristics were recorded and then the fruiting bodies were collected. The collected fruiting bodies were dried in an electric food dehydrator at 40 °C, then sealed and stored in an envelope bag. They were then transported to mycology laboratory of Southwest Forestry University, Kunming where microscopic morphology and phylogeny were studied.
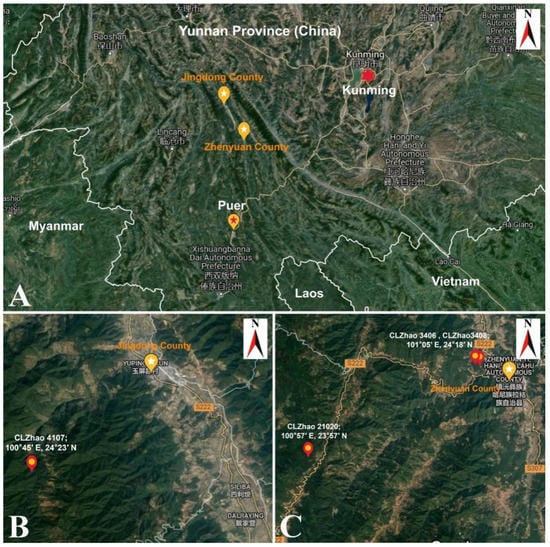
Figure 1.
The localities of new Thelephora species. (A) Jingdong county and Zhenyuan county in Yunnan province. (B) The locality of Thelephora wuliangshanensis strain (Holotype: CLZhao 4107) in Jingdong county. (C) The locality of T. wuliangshanensis (CLZhao 21020) and T. grandinioides (Holotype: CLZhao 3406; CLZhao 3408) in Zhenyuan county. Source: Map data ©2021 Google.
2.2. Morphology
The specimens studied are deposited at the herbarium of Southwest Forestry University (SWFC), Kunming, Yunnan Province, China. Macromorphological descriptions were based on field notes and photos captured in the field and lab. Color terminology followed Petersen [35]. Micromorphological data were obtained from the dried specimens, and were observed under a light microscope following Dai [36]. The following abbreviations were used: KOH = 5% potassium hydroxide water solution, CB = Cotton Blue, CB– = acyanophilous, IKI = Melzer’s reagent, IKI– = both inamyloid and indextrinoid, L = mean spore length (arithmetic average for all spores), W = mean spore width (arithmetic average for all spores), Q = variation in the L/W ratios between the specimens studied, n = a/b (number of spores (a) measured from given number (b) of specimens).
2.3. Molecular Phylogeny
A conventional cetyl trimethylammonium bromide (CTAB) plant genome rapid extraction kit (DN14, Aidlab Biotechnologies Co., Ltd., Beijing, China) was used to obtain genomic DNA from dried specimens, according to the manufacturer’s instructions [37]. Amplification reactions were performed in a 30 μL reaction volume composed of 15 μL 2 × FastTaq Premix (a mixture of FastTaq TM DNA Polymerase, buffer, dNTP Mixture, and stabilizer) (Beijing Qingke Biological Technology Co., Ltd., Beijing, China), 1 μL of each of the reverse and forward primers (Beijing Kinco Biotechnology Co., Ltd., Kunming Branch, China), 8.5 μL doble distilled water (ddH2O), and 1–1.2 μL DNA. ITS region was amplified with primer pair ITS5 and ITS4 [38]. The nrLSU was amplified with primer pairs LR0R and LR7 [39]. PCR procedure for ITS followed: initial denaturation at 95 °C for 3 min, followed by 35 cycles at 94 °C for 40 s, 58 °C for 45 s and 72 °C for 1 min, and a final extension of 72 °C for 10 min. For the nrLSU regions, PCR amplification conditions were used as follows: initial denaturation of 1 min at 94 °C, followed by 35 cycles of denaturation at 94 °C for 30 s, 1 min of annealing at 48 °C, 90 s extension at 72 °C, and a final extension of 10 min at 72 °C. PCR products were purified and sequenced at Kunming Tsingke Biological Technology Limited Company, Kunming, Yunnan Province, China. All newly generated sequences were deposited in NCBI GenBank/UNITE (Table 1). Sequences were aligned in MAFFT 7 (https://mafft.cbrc.jp/alignment/server/, accessed on 3 December 2021) using G-INS-i strategy for ITS combined dataset, and manually adjusted in BioEdit [40]. Aligned dataset was deposited in TreeBase (submission ID 28432). Odontia fibrosa (Berk. and M.A. Curtis) Kõljalg and O. ferruginea Pers. were selected as outgroup for phylogenetic analyses of combined dataset [25,26].

Table 1.
Names, vouchers, location, and corresponding GenBank/UNITE accession numbers of taxa used in this study. The newly generated sequences are shown in black bold and T indicates the type.
Maximum parsimony (MP) analysis was applied to the ITS and nrLSU dataset followed Zhao and Wu [37]. Tree construction procedure was performed in PAUP* version 4.0b10 [48]. All characters were equally weighted and gaps were treated as missing data. Trees were inferred using the heuristic search option with TBR branch swapping and 1000 random sequence additions. Max-trees were set to 5000, branches of zero length were collapsed and all parsimonious trees were saved. Clade robustness was assessed using bootstrap analysis with 1000 replicates [49]. Descriptive tree statistics: tree length (TL), consistency index (CI), retention index (RI), rescaled consistency index (RC), and homoplasy index (HI) were calculated for each Maximum Parsimonious Tree generated. Datamatrix was also analyzed using Maximum Likelihood (ML) approach with RAxML-HPC2 through the Cipres Science Gateway (www.phylo.org, accessed on 3 December 2021) [50]. Branch support for ML analysis was determined by 1000 bootstrap replicates.
MrModeltest 2.3 [51] was used to determine the best-fit evolution model for the data set for Bayesian inference (BI). BI was calculated with MrBayes 3.1.2 [52]. Four Markov chains were run for 2 runs from random starting trees for 160 thousand generations for ITS. The first one-fourth of all generations was discarded as burn-in. The majority rule consensus tree of all remaining trees was calculated. Branches were considered as significantly supported if they received maximum likelihood bootstrap value >60%, maximum parsimony bootstrap value >50%, or Bayesian posterior probabilities >0.90.
2.4. Pairwise Homoplasy Test
The Genealogical concordance phylogenetic species recognition analysis (GCPSR) is a tool used to check significant recombinant events. The data were analyzed using SplitsTree 4 with the pairwise homoplasy Φw, PHI test to determine the recombination level within closely related species [53,54,55]. One-locus dataset (ITS and nrLSU) with closely related species were used for the analyses. PHI results lower than 0.05 (Φw < 0.05) indicates a significant recombination is present in the dataset. The relationships between closely related taxa were visualized by constructing split graphs from the concatenated datasets, using the LogDet transformation and splits decomposition options.
3. Results
3.1. Molecular Phylogeny
ITS+nrLSU dataset (Figure 2) included 14 sequences representing 9 species, ITS dataset (Figure 3) included 42 sequences representing 23 species, and the nrLSU dataset (Figure 4) consisted of 27 sequences representing 18 species. The ITS+nrLSU dataset had an aligned length of 1887 characters, of which 1514 characters are constant, 123 are variable and parsimony-uninformative, and 250 are parsimony-informative. Maximum parsimony analysis yielded 1 equally parsimonious trees (TL = 616, CI = 0.7403, HI = 0.2597, RI = 0.7217, RC = 0.5343). Best model for the ITS dataset estimated and applied in the Bayesian analysis was GTR + I + G (lset nst = 6, rates = invgamma; prset statefreqpr = dirichlet (1, 1, 1, 1)). Bayesian analysis and ML analysis resulted in a similar topology to MP analysis with an average standard deviation of split frequencies = 0.006269. Estimated base frequencies; A = 0.249341, C = 0.218571, G = 0.275562, T = 0.256526; substitution rates AC = 1.282736, AG = 4.546435, AT = 0.637878, CG = 0.715981, CT = 11.556530, GT = 1.000000; proportion of invariable sites I = 0.502830; distribution shape parameter α = 0.545315. The ITS dataset had an aligned length of 727 characters, of which 355 characters are constant, 87 are variable and parsimony-uninformative, and 285 are parsimony-informative. Maximum parsimony analysis yielded 1 equally parsimonious trees (TL = 1026, CI = 0.5312, HI = 0.4688, RI = 0.7579, RC = 0.4026). Best model for the ITS dataset estimated and applied in the Bayesian analysis was GTR + I + G (lset nst = 6, rates = invgamma; prset statefreqpr = dirichlet (1, 1, 1, 1)). Bayesian analysis and ML analysis resulted in a similar topology to MP analysis with an average standard deviation of split frequencies = 0.009479. Estimated base frequencies; A = 0.219763, C = 0.258778, G = 0.243142, T = 0.278317; substitution rates AC = 1.165884, AG = 5.728342, AT = 0.893563, CG = 0.776446, CT = 8.106172, GT = 1.000000; proportion of invariable sites I = 0.125398; distribution shape parameter α = 0.548666. The nrLSU dataset had an aligned length of 1393 characters, of which 1203 characters are constant, 63 are variable and parsimony-uninformative, and 127 are parsimony-informative. Maximum parsimony analysis yielded 1 equally parsimonious trees (TL = 360, CI = 0.5611, HI = 0.4389, RI = 0.5741, RC = 0.3221). Best model for the nrLSU dataset estimated and applied in the Bayesian analysis was GTR + I + G (lset nst = 6, rates = invgamma; prset statefreqpr = dirichlet (1, 1, 1, 1)). Bayesian analysis and ML analysis resulted in a similar topology to MP analysis with an average standard deviation of split frequencies = 0.009402. Estimated base frequencies; A = 0.263240, C = 0.200643, G = 0.293575, T = 0.242542; substitution rates AC = 1.213148, AG = 7.899379 AT = 0.709533, CG = 0.649904, CT = 18.992697, GT = 1.000000; proportion of invariable sites I = 0.629441; distribution shape parameter α = 0.755239.
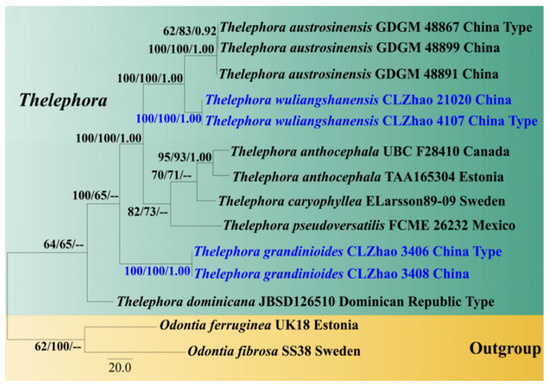

Figure 2.
Maximum parsimony strict consensus tree based on the ITS+nrLSU sequences. Bootstrap support values for maximum likelihood (ML) equal to or higher than 60%, Bayesian Probability (PP) equal to or higher than 0.90, and maximum parsimony (MP) bootstrap proportions equal to or higher than 50% are mentioned above the branches (MP/ML/PP). Strains of the newly described species are depicted in blue.
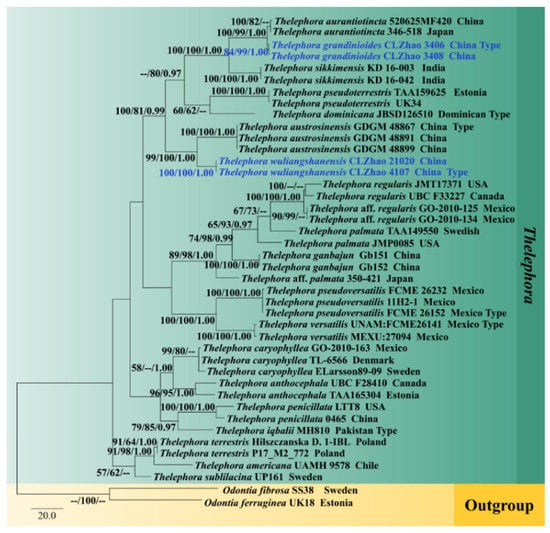
Figure 3.
Maximum parsimony consensus tree based on the ITS sequences. Bootstrap support values for maximum likelihood (ML) equal to or higher than 60%, Bayesian Probability (PP) equal to or higher than 0.90, and maximum parsimony (MP) bootstrap proportions equal to or higher than 50% are mentioned above the branches (MP/ML/PP). Strains of the newly described species are depicted in blue.
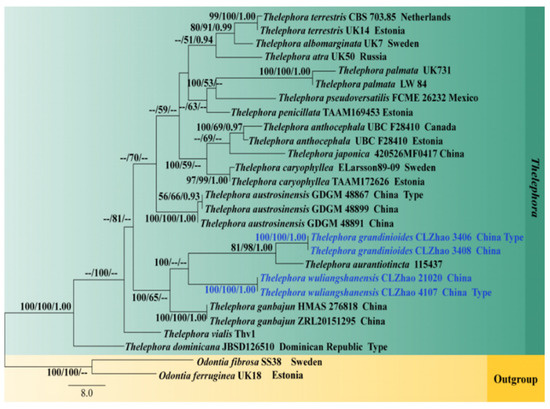
Figure 4.
Maximum parsimony strict consensus tree based on the nrLSU sequences. Bootstrap support values for maximum likelihood (ML) equal to or higher than 60%, Bayesian Probability (PP) equal to or higher than 0.90, and maximum parsimony (MP) bootstrap proportions equal to or higher than 50% are mentioned above the branches (MP/ML/PP). Strains of the newly described species are depicted in blue.
The phylogram inferred from ITS+nrLSU, ITS, and nrLSU sequences (Figure 2, Figure 3 and Figure 4) demonstrated that our specimens formed two isolated branches within Thelephora while T. grandinioides is sister to T. aurantiotincta Corner, and T. sikkimensis K. Das, Hembrom and Kuhar, and T. wuliangshanensis are sister to a clade comprising T. austrosinensis T.H. Li and T. Li and T. aurantiotincta.
Application of PHI test to the ITS and nrLSU tree-locus sequences revealed no recombination level within phylogenetically related species. No significant recombination events were observed between Thelephora grandinioides and T. wuliangshanensis and phylogenetically closely related species viz. T. austrosinensis, T. ganbajun, and T. sikkimensis (Figure 5 and Figure 6). The test results of ITS sequence dataset show Φw = 0.8271 (Φw > 0.05) no recombination is present in the two new species with T. aurantiotincta, T. austrosinensis, T. dominicana Angelini, Losi and Vizzini, T. ganbajun, T. pseudoterrestris Corner, T. sikkimensis and T. vialis (Figure 5). The test results of nrLSU sequence dataset show Φw = 0.9964 (Φw > 0.05) no recombination is present in the two new species with T. aurantiotincta, T. austrosinensis, T. dominicana, T. ganbajun, T. pseudoterrestris and T. vialis (Figure 6).
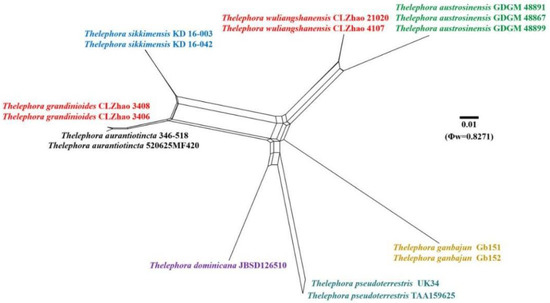
Figure 5.
Split graphs showing the results of PHI test for the ITS data of Thelephora grandinioides and T. wuliangshanensis and closely related taxa using LogDet transformation and splits decomposition. PHI test results Φw ≤ 0.05 indicate that there is significant recombination within the dataset. New taxa are in red while, closly related species to new species are in other colors.
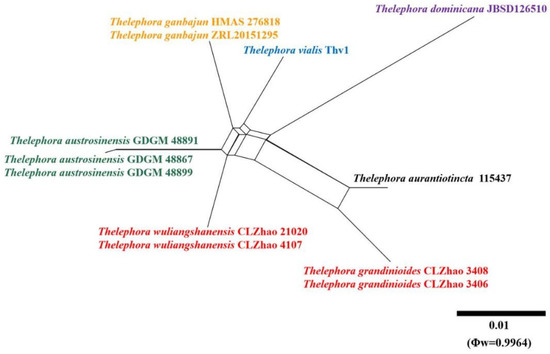
Figure 6.
Split graphs showing the results of PHI test for the nrLSU data of Thelephora grandinioides and T. wuliangshanensis and closely related taxa using LogDet transformation and splits decomposition. PHI test results Φw ≤ 0.05 indicate that there is significant recombination within the dataset. New taxa are in red while, closly related species to new species are in other colors.
3.2. Taxonomy
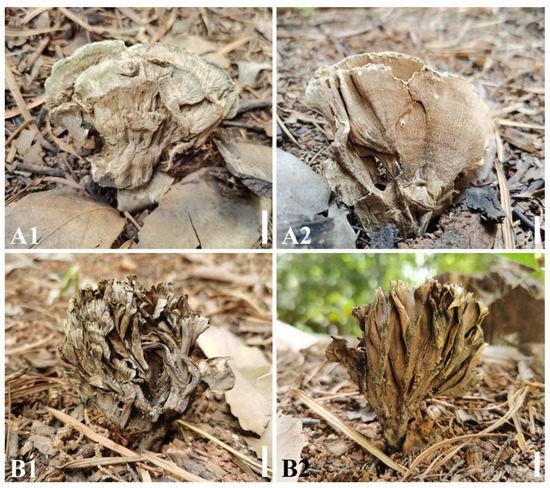

Figure 7.
Basidiocarps of Thelephora grandinioides (A: Holotype CLZhao 3406) and T. wuliangshanensis (B: Holotype CLZhao 4107). (A1) Basidiocarp surface of T. grandinioides; (A2) Hymenial surface of T. grandinioides; (B1) Basidiocarp surface of T. wuliangshanensis; (B2) Hymenial surface of T. wuliangshanensis. Bars: (A1–B2) = 1 cm.
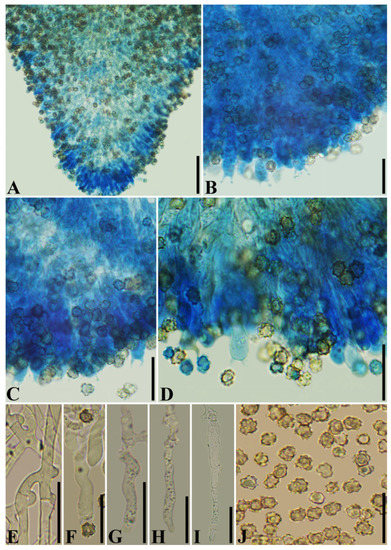
Figure 8.
Microscopic structures of Thelephora grandinioides (holotype CLZhao 3406). (A) A section of hymenium; (B) A section of hymenium; (C) A section of hymenium; (D) A section of hymenium; (E) Generative hyphae with clamps; (F) Basidia; (G) Basidioles: (H) Tubular cystidia; (I) Septated cystidia; (J) Basidiospores. Bars: (A) = 50 µm, (B–I) = 20 µm.
MycoBank no.: MB840633.
Holotype—China, Yunnan Province, Puer, Zhenyuan County, Xieqipo Park, on the ground of pine-broadleaved mixed forest, 101°05′ E, 24°18′ N, 2113 m a.s.l., 1 October 2017, CLZhao 3406 (SWFC 00003406).
Etymology—grandinioides (Lat.): referring to the grandinoid hymenophore of the type specimens.
Basidiocarps—Annual, laterally stipitate, gregarious. Pilei medium-sized, coriaceous, infundibuliform, up to 9 cm long, 7 cm wide, 1.5 mm thick; fawn to isabelline when fresh, greyish brown on drying; proliferous from a central common base, rosulate, usually with several to many laterally confluent spathulate to flabelliform or valves, uplifted; the surface radially striate; margin thin, wavy. Hymenial surface grandinoid, olivaceous buff to clay-buff when fresh, clay-buff to slightly greyish brown on drying. Stipe cylindrical, up to 4 cm long, up to 1.5 cm in diameter. Context fleshy tough to leathery in fresh condition, corky to leathery in dried condition, up to 1 mm thick at the thickest portion of pileus, thinner at margin and thicker toward the base, pinkish buff to buff. Aculei, 6–8 per mm, 0.1–0.2 mm long, greyish brown. Odor mild when fresh, somewhat smelly when dried, or with the beef jerky flavor.
Hyphal system—Monomitic, generative hyphae with clamps, colorless, thick-walled, frequently branched, interwoven, 3–6.5 µm in diameter; IKI–, CB–; tissues turn to greenish grey to buff in KOH.
Hymenium—Cystidia of two types: (1) tubular cystidia, thick-walled, 35–60 × 5–7.5 µm; (2) septated cystidia, numerous, thick-walled, 40–75 × 6–8.5 µm; basidia cylindrical to clavate, slightly constricted in the middle to somewhat sinuous, with 4 sterigmata and a basal clamp, 27–62 × 5–7.5 µm, basidioles dominant, cylindrical, but slightly smaller than basidia.
Basidiospores—Subglobose to globose, nodulose to verrucose or ridged, echinulis 0.5–1 µm, fuscous vinaceous, thick-walled, with guttatae or not, IKI–, CB–, greenish grey to buff in 5% KOH, (5–)5.3–7.4(–7.8) × (3.8–)4–6.5(–7) µm (including ornamentations), L = 6.29 µm, W = 5.31 µm, Q = 1.18–1.21 (n = 60/2).
Additional specimens examined—China, Yunnan Province, Puer, Zhenyuan County, Xieqipo Forest Park, 101°05′ E, 24°18′ N, 1350 m a.s.l., on the ground of pine-broadleaved mixed forest, leg. C.L. Zhao, 1 October 2017, CLZhao 3408 (SWFC 00003408).
Notes—Thelephora grandinioides is phylogenetically closely related to T. aurantiotincta, T. dominicana, T. sikkimensis, while T. wuliangshanensis is sister to a clade comprising T. aurantiotincta and T. austrosinensis. The nucleotide differences of phylogenetically similar species to T. grandinioides are shown in Table 2. However, morphologically T. aurantiotincta differs from T. grandinioides by the larger basidiospores (6.5–9 × 5.5–6.5 μm vs. 5.3–7.4 × 4–6.5 μm) and shorter basidia (43–55 × 6.5–8 μm vs. 27–62 × 5–7.5 μm) [3] (Table 4). In addition, the results of BLAST queries in NCBI based on ITS and nrLSU separately are shown in Table 3.

Table 2.
The nucleotide differences of phylogenetically similar species to Thelephora grandinioides and T. sikkimensis.

Table 3.
The top ten species results of Blast search of Thelephora grandinioides and T. sikkimensis sequences.
Morphologically, T. grandinioides is similar to T. aurantiotincta, T. fuscella Ces. ex Lloyd, T. gelatinoidea Lloyd, T. griseozonata Cooke, T. intybacea Pers., T. japonica Yasuda and T. terrestris by having a grandinoid or odontoid hymenial surface. However, T. aurantiotincta differs from T. grandinioides by larger basidiospores (6.5–9 × 5.5–6.5 μm vs. 5.3–7.4 × 4–6.5 μm) and shorter basidia (43–55 × 6.5–8 μm vs. 27–62 × 5–7.5 μm) [3] (Table 4); T. fuscella differs in its shorter basidia (35–45 × 6–7 μm vs. 27–62 × 5–7.5 μm) with 2–4 sterigmata [3] (Table 4); T. gelatinoidea differs in having larger basidiospores (7–9.5 × 6–9 μm vs. 5.3–7.4 × 4–6.5 μm) and basidia (45–70 × 8–10 vs. 27–62 × 5–7.5 μm) [3] (Table 4); T. griseozonata separates from T. grandinioides by having larger basidiospores (8–12 × 5–8 μm vs. 5.3–7.4 × 4–6.5 μm) [3] (Table 4); T. intybacea differs from T. grandinioides by having larger basidiospores (8–12 × 6–9 μm vs. 5.3–7.4 × 4–6.5 μm) and basidia (45–90 × 9–12 vs. 27–62 × 5–7.5 μm) with 2–4 sterigmata [3] (Table 4); T. japonica differs from T. grandinioides by having larger basidiospores (7–10 ×6–8 μm vs. 5.3–7.4 × 4–6.5 μm) and smaller basidia (40–55 × 8–10 μm vs. 27–62 × 5–7.5 μm) [3] (Table 4); T. terrestris differs from T. grandinioides by having larger basidiospores (8–12 × 6–9 μm vs. 5.3–7.4 × 4–6.5 μm) and basidia (40–90 × 8–12 μm vs. 27–62 × 5–7.5 μm) [3] (Table 4).

Table 4.
Comparison of Basidiospore, Basidia, Cystidia, Host, Substrate, Location and corresponding references of Thelelphora species. The characteristics of newly generated taxa are shown in black bold.
Thelephora grandinioides morphologically resembles T. ganbajun and T. ramarioides D.A. Reid due to the presence of cystidia. However, T. ganbajun differs from T. grandinioides by its larger basidiospores (7–12 × 6–8 μm vs. 5.3–7.4 × 4–6.5 μm), shorter basidia (25–35 × 9–12 μm vs. 27–62 × 5–7.5 μm), and larger cystidia (52–80 × 7–14 μm vs. 35–60 × 5–7.5 μm) [17] (Table 4); T. ramarioides differs from T. grandinioides by its larger basidiospores (7–12.5 × 5–8.5 μm vs. 5.3–7.4 × 4–6.5 μm) and smaller basidia (46–70 × 5–12 μm vs. 27–62 × 5–7.5 μm) [3] (Table 4).
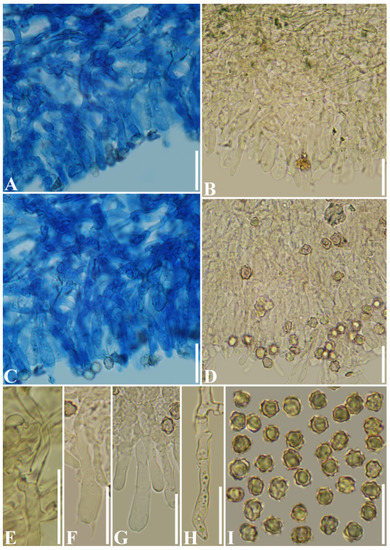
Figure 9.
Microscopic structures of Thelephora wuliangshanensis (Holotype CLZhao 4107). (A) A section of hymenium; (B) A section of hymenium; (C) A section of hymenium; (D) A section of hymenium; (E) Generative hyphae with clamps; (F) Basidia; (G) Basidioles: (H) Cystidia; (I) Basidiospores. Bars: (A–I) = 20 µm.
MycoBank no.: MB840634.
Holotype—China, Yunnan Province, Puer, Jingdong County, Huangcaoling, Wuliangshan National Nature Reserve, 100°45′ E, 24°23′ N, 2313 m a.s.l., on the ground of pine-broadleaved mixed forest, leg. C.L. Zhao, 5 October 2017, CLZhao 4107 (SWFC 00004107).
Etymology—wuliangshanensis (Lat.): referring to the provenance (Wuliangshan) of the type specimens.
Basidiocarps—Annual, laterally stipitate, gregarious. Pilei small to medium-sized, coriaceous, infundibuliform, up to 5.5 cm long, 4.5 cm wide, 1 mm thick; buff to salmon when fresh, pinkish buff to cinnamon-buff on drying; proliferous from a central common base, usually with several to many laterally confluent spathulate to flabelliform, uplifted; the surface radially black striate; margin thin, serrulate. Hymenial surface smooth, umber to coffee when fresh, coffee on drying. Stipe cylindrical, up to 2 cm long, up to 5 mm in diameter. Context fleshy tough in fresh condition, leathery in dried condition, up to 0.7 mm thick at the thickest portion of pileus, thinner at margin and thicker towards the base, pinkish buff. Odor mild when fresh, somewhat with the beef jerky flavor.
Hyphal system—Monomitic, generative hyphae with clamps, colorless, thick-walled, frequently branched, interwoven, 2.5–6 µm in diameter; IKI–, CB–; tissues turn to greenish grey in KOH.
Hymenium—Cystidia tubular, thick-walled, 28–55 × 3–7.5 µm; basidia barrel-shaped to slightly clavate, with 4 sterigmata and a basal clamp, 30–60 × 5–9.5 µm, basidioles dominant, clavate, but slightly smaller than basidia.
Basidiospores—Subglobose to globose, nodulose to verrucose, echinulis 0.5–1 µm, umber purple, thick-walled, with guttatae or not, IKI–, CB–, greenish grey to buff in 5% KOH, (5–)5.2–8.7(–9.3) × (3.7–)4.5–7.2(–7.6) µm (including ornamentations), L = 7 µm, W = 5.66 µm, Q = 1.23–1.25 (n = 60/2).
Additional specimens examined—China, Yunnan Province, Puer, Zhenyuan County, Huangcaoling, Wuliangshan National Nature Reserve, 100°57′ E, 23°57′ N, on the ground of pine-broadleaved mixed forest, 8 October 2020 CLZhao 21020 (SWFC 00021020).
Notes—Thelephora wuliangshanensis is sister to a clade comprising T. aurantiotincta and T. austrosinensis in phylogeny; and the nucleotide differences of phylogenetically similar species to T. sikkimensis are shown in Table 2. Thelephora sikkimensis differs from T. grandinioides by its shorter cystidia (30–50 μm vs. 35–60 μm) and hairy basidiocarp surface [26] (Table 4). Thelephora aurantiotincta separates from T. wuliangshanensis by smaller basidia (43–55 × 6.5–8 μm vs. 30–60 × 5–9.5 μm) [3] (Table 4); while T. austrosinensis differs from T. wuliangshanensis by its smaller basidiospores (5.7–6.3 × 5.0–5.4 μm vs. 5.2–8.7 × 4.5–7.2 μm) and basidia (15–24 × 5–7 μm vs. 30–60 × 5–9.5 μm) [27] (Table 4). In addition, the results of BLAST queries in NCBI based on ITS and nrLSU separately are shown in Table 3.
4. Discussion
In the present study, two new species, Thelephora grandinioides and T. wuliangshanensis are described based on phylogenetic analyses and morphological characteristics. In addition, the PHI test (Figure 5 and Figure 6) was carried out to confirm there is no recombination present in the two new species compared with closely related taxa.
Thelephora, a genus with diverse basidiocarp forms, are widely distributed worldwide [3,26,28]. Basidia form is an important characteristic of intraspecific identification in Thelephora, and most species of Thelephora have 4-spored basidia, there are 2-spored (T. erebia), 2–3-spored (T. atra and T. luzonensis), 2–4-spored (T. anthocephala, T. atrocitrina, T. brunneoviolacea, T. caryophyllea, T. cerberea, T. crassitexta, T. fragilis, T. fuscella, T. intybacea, T. japonica, T. magnifica, T. nigrescens, T. palmata, and T. terrestris) and sometimes there are 6–8-spored (T. ramarioides) [3,4,5,21] (Table 4).
Thelephora is closely related to Tomentella both in morphology and phylogeny [5,10,14,25,29,74]. Molecular phylogenetic analyses of previous studies showed that the taxa of Thelephora and Tomentella are non-monophyletic groups, and they are intermixed in molecular phylogeny [4,5,25,29,31,33]. Traditionally, the form of basidiocarps is the most important characteristic in distinguishing Thelephora and Tomentella, which are resupinate in Tomentella, but erect, with varied forms, to partially resupinate in Thelephora [3,14,24,25,29,34,74]. The variations in basidiocarp form may also complicate the characteristics of taxa [75,76], and the results of the morphological investigations and molecular phylogenetic analyses suggested that basidiocarp reduction happened several times independently across the evolution of thelephoroid fungi [14,25]. Taxa with reduced basidiocarps should be taken into account in the diagnoses of genera for which the initial descriptions did not cover a real spectrum of polymorphism and trends of morphological rationalization in connection with colonization of specific habitats [14,25,76]. According to molecular data, only one genus may be recognized, and Tomentella will be merged into Thelephora [25].
In the habitat and distribution, thelephoroid fungi have a circumglobal distribution, ranging from polar deserts [9] to tropical forests [3], but their peak diversity is observed within the boreal zone of the planet [14,33]. Most of the species in this group have ectomycorrhizal associations [77,78], but it is also capable of destroying wood debris as white rot producers [14,15]. The species of Thelephora are a widely distributed group found on six continents except Antarctica [3,5,14,17,26], mainly distributed across Europe in Austria, Bavaria, Belgium, Denmark, Estonia, France, Georgia, Germany, Italy, Netherlands, Norway, Poland, Russia, Slovenia, Spain, Sweden, UK and Ukraine [3,5,28,58,59,62,65,66,76]; additionally, the most-common substrata are hardwood and conifer [3,5]. It is also distributed in Asia (Borneo, China, Japan, India, Malaysia, Nepal, Pakistan, Philippines, Sri Lanka, and Singapore) [3,5,56,57,58,60,67,70,71], North America (Bahamas, Canada, Cuba, Dominican Republic, Haiti, Jamaica, Mexico, and USA) [3,5,14,25,56,57,58,62,63,69], South America (Uruguay) [3], Oceania (Australia and Papua New Guinea) [3], and Africa (Congo and Southern Africa) [3,5,56] seen in Table 4. Twenty one species of Thelephora have been reported from China (including our two new species), in which T. ganbajun and T. vialis Schwein. are the two most commonly reported taxa, and the former is one of the most popular edible fungi in Southwest China [5,17,18,19,20,60,61,67,68,69,70,71,72,73,74,75,76,77,78,79]. The diversity of Thelephora in China is still not well-known, especially in the subtropical and tropical regions and many recently described taxa of thelephoroid fungi are from these areas [5,67]. Thelephora grandinioides and T. wuliangshanensis are also from subtropics. According to our statistics, twenty-one Thelephora species have been recorded in China (Table 4), in which 7 are edible and 4 are medicinal (T. aurantiotincta, T. ganbajun, T. terrestris, and T. vialis) with anticancer properties, treat leukemia, boost immunity, and are an anti-allergic agent (Table 5). Fleshy to coriaceous basidiocarps, a mild odor, and beef jerky flavor are characteristics of T. grandinioides. Several Thelephora species are known as edible or medicinal mushrooms, while our new species are potential edibles thus, secondary metabolite analyses of the two new species should be carried out in the future.

Table 5.
Edible and medicinal species of Thelelphora.
Author Contributions
Conceptualization, C.-L.Z.; methodology, C.-L.Z. and X.-F.L.; software, C.-L.Z., X.-F.L. and S.T.; validation, C.-L.Z. and X.-F.L.; formal analysis, C.-L.Z. and X.-F.L.; investigation, C.-L.Z. and X.-F.L.; resources, C.-L.Z.; writing—original draft preparation, C.-L.Z. and X.-F.L.; writing—review and editing, C.-L.Z., S.C.K., S.T., J.-C.X., J.K. and X.-F.L.; visualization, C.-L.Z. and X.-F.L.; supervision, C.-L.Z.; project administration, C.-L.Z.; funding acquisition, C.-L.Z. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National NaturalScience Foundation of China (Project No. 32170004), Yunnan Fundamental Research Project, grant number 202001AS070043. S.T. thanks the International Postdoctoral Exchange Fellowship Program(number Y9180822S1), CAS President’s International Fellowship Initiative (PIFI)(number 2020PC0009), China Postdoctoral Science Foundation and the Yunnan HumanResources, and Social Security Department Foundation for funding herpostdoctoral research. S.C.K. thanks CASPresident’s International Fellowship Initiative (PIFI) young staff under the grantnumber: 2020FYC0002 for funding his postdoctoral research and the NationalScience Foundation of China (NSFC) for funding this research work under projectcode 31750110478. This work was partly supported by Chiang Mai University.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Publicly available datasets were analyzed in this study. This data can be found here: [https://www.ncbi.nlm.nih.gov/; https://www.mycobank.org/page/Simple%20names%20search; https://treebase.org/treebase-web/home.html, submission ID 28432; accessed on 3 December 2021].
Acknowledgments
We are grateful to Dong-Qiong Wang (Southwest Forestry University) for her help in Microstructure diagram and Yunnan Academy of Biodiversity, Southwest Forestry University for providing the facilities for molecular phylogenetic experiments.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Cunningham, G.H. The Thelephoraceae of Australia and New Zealand; New Zealand Department of Science and Industrial Research: Wellington, New Zealand, 1963; pp. 228–239. [Google Scholar] [CrossRef]
- Burt, E.A. The Thelephoraceae of North America I-XV; Hafner Publishing Company, Inc.: New York, NY, USA; London, UK, 1966; pp. 199–229. [Google Scholar] [CrossRef]
- Corner, E.J.H. A monograph of Thelephora (Basidiomycetes). Nova Hedwig. 1968, 27, 1–110. [Google Scholar]
- Ramírez-López, I.; Villegas-Ríos, M.; Salas-Lizana, R. Thelephora versatilis and Thelephora pseudoversatilis: Two new cryptic species with polymorphic basidiomes inhabiting tropical deciduous and sub-perennial forests of the Mexican Pacific coast. Mycologia 2015, 107, 346–358. [Google Scholar] [CrossRef]
- Li, T.; Li, T.H.; Song, B.; Hosen, M.I. Thelephora austrosinensis (Thelephoraceae), a new species close to T. ganbajun from southern China. Phytotaxa 2020, 471, 208–220. [Google Scholar] [CrossRef]
- Hilszczanska, D.; Sierota, Z. Persistence of ectomycorrhizas by Thelephora terrestris on outplanted Scots pine seedlings. Acta Mycol. 2006, 41, 313–318. [Google Scholar] [CrossRef]
- Iwanski, M.; Rudawski, M.; Leski, T. Mycorrhizal associations of nursery grown scots pine (Pinus sylvestris L.) seedlings in poland. Ann. For. Sci. 2006, 63, 715–723. [Google Scholar] [CrossRef]
- Flykt, E.; Timonen, S.; Pennanen, T. Variation of ectomycorrhizal colonisation in Norway spruce seedlings in Finnish forest nurseries. China Pharm. 2008, 42, 571–585. [Google Scholar] [CrossRef]
- Sha, T.; Xu, J.; Palanichamy, M.G.; Zhang, H.B.; Li, T.; Zhao, Z.W.; Zhang, Y.P. Genetic diversity of the endemic gourmet mushroom Thelephora ganbajun from south-western China. Microbiology 2008, 154, 3460–3468. [Google Scholar] [CrossRef]
- Yorou, S.N. Miscellaneous Contributions to the Anatomy and Molecular Phylogeny of Tropical African Resupinate Thelephorales. Ph.D. Thesis, University of Munich, Munich, Germany, 2008; p. 120. [Google Scholar]
- He, J.; Zhou, Z.; Yang, H.; Xu, J. Integrative management of commercialized wild mushroom: A case study of Thelephora ganbajun in Yunnan, southwest China. Environ. Manag. 2011, 48, 98–108. [Google Scholar] [CrossRef] [PubMed]
- Yangdo, R.; Kumar, S.; Sharma, Y.P. Three hitherto unreported macrofungi from cold arid region of Ladakh Province, Jammu and Kashmir, India. Kavaka 2015, 44, 42–44. [Google Scholar]
- Ezhov, O.N.; Zmitrovich, I.V. Lignotrophic basidiomycetes from pioneering microsites in boreal forests of the White Sea Region. Byulleten Moskovskogo Obshchestva Ispytateley Prirody. Otd. Biol. 2017, 122, 44–50. [Google Scholar]
- Zmitrovich, I.V.; Shchepin, O.N.; Malysheva, V.F.; Kalinovskaya, N.I.; Volobuev, S.V.; Myasnikov, A.G.; Ezhov, O.N.; Novozhilov, Y.K. Basidiome reduction in litter-inhabiting Thelephorales in boreal forest environments: Morphological and molecular evidence. Curr. Res. Environ. Appl. Mycol. 2018, 8, 360–371. [Google Scholar] [CrossRef]
- Bi, Z.S.; Zheng, G.Y.; Li, T.H. The Macrofungus Flora of China’s Guangdong Province; Chinese University Press: Hong Kong, 1993; p. 734. [Google Scholar]
- Teng, S.C. Notes on the Thelephoraceae and Hydnaceae from China. Sinensia 1934, 6, 9–13. [Google Scholar]
- Tai, F.L. Sylloge Fungorum Sinicorum; Science Press, Academia Sinica: Beijing, China, 1979; pp. 740–741. [Google Scholar]
- Zang, M. Some new and noteworthy higher fungi from eastern Himalayas. Acta Bot. Yunnanica 1987, 9, 81–88. [Google Scholar]
- Dai, Y.C.; Wei, Y.L.; Zhang, X.Q. An annotated checklist of non-poroid Aphyllophorales in China. Ann. Bot. Fenn. 2004, 41, 233–247. [Google Scholar] [CrossRef]
- Qin, W.M.; Dai, Y.C.; Wu, X.L. Thelephora caryophyllea-a corticioid fungus new to China. Guizhou Sci. 2009, 27, 19–21. [Google Scholar]
- Li, Y.; Li, T.H.; Yang, Z.L.; Bau, T.; Dai, Y.C. Atlas of Chinese Macrofungal Resources; Central China Farmers Publishing House: Zhengzhou, China, 2015; p. 1351. [Google Scholar]
- Willdenow, C.L. Florae Berolinensis Prodromus: Secundum Systema Linneanum ab Illustr. Viro ac eq. C.P. Thunbergio EMENDATUM Conscriptus; Impensis Wilhelmi Viewegii, Berolini: Berlin, Germany, 1787; p. 490. [Google Scholar]
- Kirk, P.M.; Cannon, P.; Stalpers, J. (Eds.) Dictionary of the Fungi, 10th ed.; CABI: Wallingford, UK, 2008; p. 686. [Google Scholar]
- Zecchin, G. Ⅱ genere Thelephora in Friuli-Ottavo contributo. Riv. Micol. 2008, 51, 117–125. [Google Scholar]
- Vizzini, A.; Angelini, C.; Losi, C.; Ercole, E. Thelephora dominicana (Basidiomycota, Thelephorales), a new species from the Dominican Republic, and preliminary notes on thelephoroid genera. Phytotaxa 2016, 265, 27–38. [Google Scholar] [CrossRef]
- Das, K.; Hembrom, M.E.; Ghosh, A.; Parihar, A.; Kuhar, F. Thelephora sikkimensis sp. nov. (Thelephoraceae) from the Eastern Himalayas (India). Nova Hedwig. 2018, 107, 337–347. [Google Scholar] [CrossRef]
- Khalid, N.A.; Hanif, M. Thelephora iqbalii sp. nov. from the himalayan moist temperate forests of Pakistan. Mycotaxon 2018, 132, 943–950. [Google Scholar] [CrossRef]
- Index Fungorum. 2021. Available online: http://www.indexfungorum.org/Names/Names.asp (accessed on 3 December 2021).
- Stalpers, J.A. The Aphyllophoraceous Fungi I. Keys to the species of the Thelephorales. Stud. Mycol. 1993, 35, 1–168. [Google Scholar]
- Yorou, N.S.; Agerer, R. Tomentella furcata, a new species from Benin (West Africa) with basidia forming internal hyphae. Mycol. Prog. 2007, 6, 239–247. [Google Scholar] [CrossRef]
- Larsson, K.H.; Larsson, E.; Kljalg, U. High phylogenetic diversity among corticioid homobasidiomycetes. Mycol. Res. 2004, 108, 983–1002. [Google Scholar] [CrossRef]
- Kuhar, F.; Barroetaveña, C.; Rajchenberg, M. New species of Tomentella (Thelephorales) from the patagonian andes forests. Mycologia 2016, 108, 780–790. [Google Scholar] [CrossRef]
- Kõljalg, U. Tomentella (Basidiomycota) and related genera in temperate Eurasia. Synop. Fungorum 1996, 9, 1–213. [Google Scholar]
- Patouillard, N. Les Hyménomycètes d’Europe; Paul Klincksieck: Paris, France, 1887; p. 166. [Google Scholar]
- Petersen, J.H. Farvekort. The Danish Mycological Society’s Colour-Chart; Foreningen til Svampekundskabens Fremme: Greve, Denmark, 1996; pp. 1–6. [Google Scholar]
- Dai, Y.C. Polypore diversity in China with an annotated checklist of Chinese polypores. Mycoscience 2012, 53, 49–80. [Google Scholar] [CrossRef]
- Zhao, C.L.; Wu, Z.Q. Ceriporiopsis kunmingensis sp. nov. (Polyporales, Basidiomycota) evidenced by morphological characters and phylogenetic analysis. Mycol. Prog. 2017, 16, 93–100. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protoc. Guide Methods Appl. 1990, 18, 315–322. [Google Scholar] [CrossRef]
- Vilgalys, R.; Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 1990, 172, 4239–4246. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar] [CrossRef]
- Kernaghan, G.; Sigler, L.; Khasa, D. Mycorrhizal and root endophytic fungi of containerized Picea glauca seedlings assessed by rDNA sequence analysis. Microb. Ecol. 2003, 45, 128–136. [Google Scholar] [CrossRef]
- Larsson, K.H.; Svantesson, S.; Miscevic, D.; Kljalg, U.; Larsson, E. Reassessment of the generic limits for Hydnellum and Sarcodon (thelephorales, basidiomycota). MycoKeys 2019, 54, 31–47. [Google Scholar] [CrossRef]
- Zhao, R.L.; Li, G.J.; Sánchez-Ramírez, S.; Stata, M.; Yang, Z.L.; Wu, G.; Dai, Y.C.; He, S.H.; Cui, B.K.; Zhou, J.L.; et al. A six-gene phylogenetic overview of Basidiomycota and allied phyla with estimated divergence times of higher taxa and a phyloproteomics perspective. Fungal Divers. 2017, 84, 1–32. [Google Scholar] [CrossRef]
- Palmer, J.M.; Lindner, D.L.; Volk, T.J. Ectomycorrhizal characterization of an american chestnut (Castanea dentata)-dominated community in western wisconsin. Mycorrhiza 2008, 19, 27–36. [Google Scholar] [CrossRef]
- Weiß, M.; Oberwinkler, F. Phylogenetic relationships in auriculariales and related groups-hypotheses derived from nuclear ribosomal dna sequences. Mycol. Res. 2001, 105, 403–415. [Google Scholar] [CrossRef]
- Taylor, D.L.; Bruns, T.D. Independent, specialized invasions of ectomycorrhizal mutualism by two nonphotosyntheti corchids. Proc. Natl. Acad. Sci. USA 1997, 94, 4510–4515. [Google Scholar] [CrossRef]
- Vu, D.; Groenewald, M.; De, V.M.; Gehrmann, T.; Stielow, B.; Eberhardt, U.; Al-Hatmi, A.; Groenewald, J.Z.; Cardinali, G.; Houbraken, J.; et al. Large-Scale generation and analysis of filamentous fungal dna barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Stud. Mycol. 2019, 92, 135–154. [Google Scholar] [CrossRef]
- Swofford, D.L. PAUP*: Phylogenetic Analysis Using Parsimony (* and Other Methods); Version 4.0b10; Sinauer Associates: Sunderland, MA, USA, 2002. [Google Scholar]
- Felsenstein, J. Confidence intervals on phylogenetics: An approach using bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Miller, M.A.; Holder, M.T.; Vos, R.; Midford, P.E.; Liebowitz, T.; Chan, L.; Hoover, P.; Warnow, T. The CIPRES Portals. CIPRES. 2009. Available online: http://www.phylo.org/sub_sections/portal (accessed on 3 December 2021).
- Nylander, J.A.A. MrModeltest v2. Program Distributed by the Author; Evolutionary Biology Centre, Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef]
- Bruen, T.C.; Philippe, H.; Bryant, D. A simple and robust statistical test for detecting the presence of recombination. Genetics 2006, 172, 2665–2681. [Google Scholar] [CrossRef]
- Huson, D.H.; Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 2006, 23, 254–267. [Google Scholar] [CrossRef]
- Quaedvlieg, W.; Binder, M.; Groenewald, J.Z.; Summerell, B.A.; Carnegie, A.J.; Burgess, T.I.; Crous, P.W. Introducing the consolidated species concept to resolve species in the Teratosphaeriaceae. Persoonia 2014, 33, 1–40. [Google Scholar] [CrossRef]
- Coker, W.C. Notes on the Thelephoraceae of North Carolina. J. Elisha Mitchell Sci. Soc. 1921, 36, 146–196. [Google Scholar]
- Lentz, P.L. The genus Thelephora in Iowa. Proc. Iowa Acad. Sci. 1942, 49, 175–184. [Google Scholar]
- Global Biodiversity Information Facility (GBIF). 2021. Available online: https://www.gbif.org/species/search?q=fungi&dataset_key (accessed on 3 December 2021).
- Vidal, J.M.; Siquier, J.L.; Constantino, C. Alguns macromicets nous o interessants de l’illa de Mallorca (Balears). Rev. Catalana Micol. 1994, 16, 135–144. [Google Scholar]
- Dai, Y.C. A revised checklist of corticioid and hydnoid fungi in China for 2010. Mycoscience 2011, 52, 69–79. [Google Scholar] [CrossRef]
- Landi, M.; Salerni, E.; Ambrosio, E.; D’Aguanno, M.; Nucci, A.; Saveri, C.; Perini, C.; Angiolini, C. Concordance between vascular plant and macrofungal community composition in broadleaf deciduous forests in central Italy. Iforest Biogeosci. For. 2014, 3, 279–286. [Google Scholar] [CrossRef]
- Czernyadjeva, I.V.; Afonina, O.M.; Davydov, E.A.; Doroshina, G.Y.; Zyatnina, V. New cryptogamic records. 5. Nov. Sist. Nizshikh Rastenii 2020, 54, 261–286. [Google Scholar] [CrossRef]
- Hernández-López, L.; Rodríguez-Alcántar, O.; Figueroa-García, D.; Reynoso-Dueñas, J.J.; Arias, A. Threatened and endemic flora and mycobiota in the municipality San Sebastián del Oeste, Jalisco, Mexico. Acta Bot. Mex. 2021, 128, 2448–7589. [Google Scholar] [CrossRef]
- Vila, J.; Angel, F.; Mayoral, A.; Hoyo, P. Segona aportació al coneixement dels macromicets de la zona de Garraf. Rev. Catalana Micol. 1998, 21, 45–61. [Google Scholar]
- Mleczko, P. Mycorrhizal and saprobic macrofungi of two zinc wastes in southern poland. Acta Biol. Crac. Ser. Bot. 2004, 46, 25–38. [Google Scholar] [CrossRef]
- Abrahão, M.C.; Gugliotta, A.D.M.; Bononi, V.L.R. Xylophilous Agaricomycetes (Basidiomycota) of the Brazilian Cerrado. Check List 2012, 8, 1102–1116. [Google Scholar] [CrossRef]
- Liu, T.Z.; Li, G.L. Atlas of Macrofungi in Saihanwula National Nature Reserve of Inner Mongolia; Inner Mongolia Science Press: Hohhot, China, 2019; p. 341. [Google Scholar]
- Chanonagómez, F.; Andradegallegos, R.H.; Castellanosalbores, J.; Sánchez, J.E. Macromicetos del parque educativo laguna bélgica, muncipio de ocozocoautla de espinosa, chiapas, méxico. Rev. Mex. Biodivers. 2007, 78, 369–381. [Google Scholar] [CrossRef]
- Chacón, S.; Tapia, F.; Jarvio, D. Four interesting aphyllophoroid species in the tropical northern region of Veracruz, Mexico. Mycotaxon 2018, 133, 153–163. [Google Scholar] [CrossRef]
- Budathoki, U.S.H.A. A new species of Periconiella from Kathmandu valley, Nepal. Recent Mycol. Res. 2009, 77–80. [Google Scholar]
- Roberts, P.J.; Spooner, B.M. Cantharelloid, clavarioid and thelephoroid fungi from Brunei Darussalam. Kew Bull. 2000, 55, 843–851. [Google Scholar] [CrossRef]
- Welden, A.L. West Indian species of dark-spored Thelephoraceae. Sydowia 1968, 22, 269–273. [Google Scholar]
- An, G. The excellent edible mushrooms in Yunnan—The exploitation and utilization of Thelephora vialis schw and T. aurantiotincta Corner. J. Southwest For. Coll. 1998, 18, 105–108. [Google Scholar]
- Kõljalg, U.; Dahlberg, A.; Taylor, A.F.S.; Larsson, E.; Hallenberg, J.; Stenlid, K.H.; Larsson, P.M.; Fransson, O.; Kårén, L.; Jonsson, L. Diversity and abundance of resupinate thelephoroid fungi as ectomycorrhizal symbionts in swedish boreal forests. Mol. Ecol. 2000, 9, 1985–1996. [Google Scholar] [CrossRef]
- Cunningham, G.H. Thelephoraceae of New Zealand (parts XII and XIII). Part XII: The genera Thelephora and Tomentella. Trans. R. Soc. N. Z. 1957, 84, 479–496. [Google Scholar]
- Ramirez-Lópezl, I.; Villegas-Ríos, M.; Cano-Santana, Z. Phenotypic plasticity of the basidiomata of Thelephora sp. (Thelephoraceae) in tropical forest habitats. Rev. Biol. Trop. 2013, 61, 343–350. [Google Scholar] [CrossRef][Green Version]
- Erland, S.; Taylor, A.F.S. Resupinate ectomycorrhizal fungal genera. In Ectomycorrhizal Fungi: Key Genera in Profile; Springer: Berlin/Heidelberg, Germany, 1999; p. 369. [Google Scholar] [CrossRef]
- Zmitrovich, I.V.; Wasser, S.P.; Ţura, D. Wood-Inhabiting fungi. In Fungi from Different Substrates; Misra, J.K., Tewari, J.P., Deshmukh, S.K., Vágvölgyi, C., Eds.; CRC Press, Taylor and Francis Group: New York, NY, USA, 2015; pp. 17–74. [Google Scholar]
- Teng, S.C. Fungi of China; Science Press: Beijing, China, 1963; p. 808. [Google Scholar]
- Lin, H.; Liu, J.K. p-Terphenyls from the basidiomycete Thelephora aurantiotincta. Z. Nat. C J. Biosci. 2003, 58, 452–454. [Google Scholar] [CrossRef]
- Wang, X.H.; Liu, P.G. Resources investigation and study on the wild commercial fungi in Yunnan Province. Biodivers. Sci. 2002, 10, 318–3325. [Google Scholar] [CrossRef]
- Barad, A.A. British Columbian Wild Mushrooms as Potent Source of Novel Natural Anti-Cancer Compounds. Ph.D. Thesis, University of Northern British Columbia, Prince George, BC, Canada, 2017. [Google Scholar]
- Liu, J.K.; Lin, H.; Dong, Z.J.; Hu, Q. Dpph radical scavenging activity of ten natural p-terphenyl derivatives obtained from three edible mushrooms indigenous to china. Chem. Biodivers. 2010, 1, 601–605. [Google Scholar] [CrossRef] [PubMed]
- Norikura, T.; Fujiwara, K.; Narita, T.; Yamaguchi, S.; Morinaga, Y.; Kunihisa, I.; Matsue, H. Anticancer activities of thelephantin o and vialinin an isolated from Thelephora aurantiotincta. J. Agric. Food Chem. 2011, 59, 6974–6979. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.H.; Meng, W.W.; Liu, R.C.; Bai, Y.X.; Xu, H.Q.; Ding, R.; Shao, S.C. Complete mitochondrial genome of the edible Basidiomycete mushroom Thelephora aurantiotincta (Aphyllophorales: Thelephoraceae) from China. Mitochondrial DNA Part B 2021, 6, 606–607. [Google Scholar] [CrossRef]
- Hu, L.; Gao, J.M.; Liu, J.K. From fruiting bodies of the basidiomycete Thelephora ganbajun. Helv. Chim. Acta 2001, 84, 3342–3349. [Google Scholar] [CrossRef]
- Hu, L.; Liu, J.K. Two novel phenylacetoxylated p-terphenyls from Thelephora ganbajun Zang. Z. Nat. Sect. C J. Biosci. 2001, 56, 983–987. [Google Scholar] [CrossRef]
- Xu, D.P.; Jie, Z.; Yue, Z.; Li, Y.; Li, S.; Li, H.B. Extraction of natural antioxidants from the Thelephora ganbajun mushroom by an ultrasound-assisted extraction technique and evaluation of antiproliferative activity of the extract against human cancer cells. Int. J. Mol. Sci. 2016, 17, 1664. [Google Scholar] [CrossRef]
- Zhang, L.; Zhang, A.L.; Li, X.M.; Wang, J.; Gao, J.M. Preliminary study of the anti-microbial activities of fermentation broth of Thelephora japonica. J. Northwest For. Univ. 2007, 22, 135–137. [Google Scholar]
- Radulovic, N.; Dang, N.Q.; Hashimoto, T.; Nukada, M.; Asakawa, Y. Terrestrins A–G: Pmilerphenyl derivatives from the inedible mushroom Thelephora terrestris. Phytochemistry 2005, 66, 1052–1059. [Google Scholar] [CrossRef]
- Xie, C.; Koshino, H.; Esumi, Y.; Onose, J.; Yoshikawa, K.; Abe, N. Vialinin B, a novel potent inhibitor of tnf-alpha production, isolated from an edible mushroom, Thelephora vialis. Bioorg. Med. Chem. Lett. 2006, 16, 5424–5426. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Koshino, H.; Esumi, Y.; Takahashi, S.; Abe, N. Vialinins A and B: Novel bioactive compounds from Thelephora vialis, an edible mushroom in China. ACS Symp. 2008, 993, 465–472. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).