News from the Sea: A New Genus and Seven New Species in the Pleosporalean Families Roussoellaceae and Thyridariaceae
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungal Isolates
2.2. Morphological Analysis
2.3. DNA Extraction, PCR Amplification, and Data Assembling
2.4. Sequence Alignment and Phylogenetic Analysis
3. Results
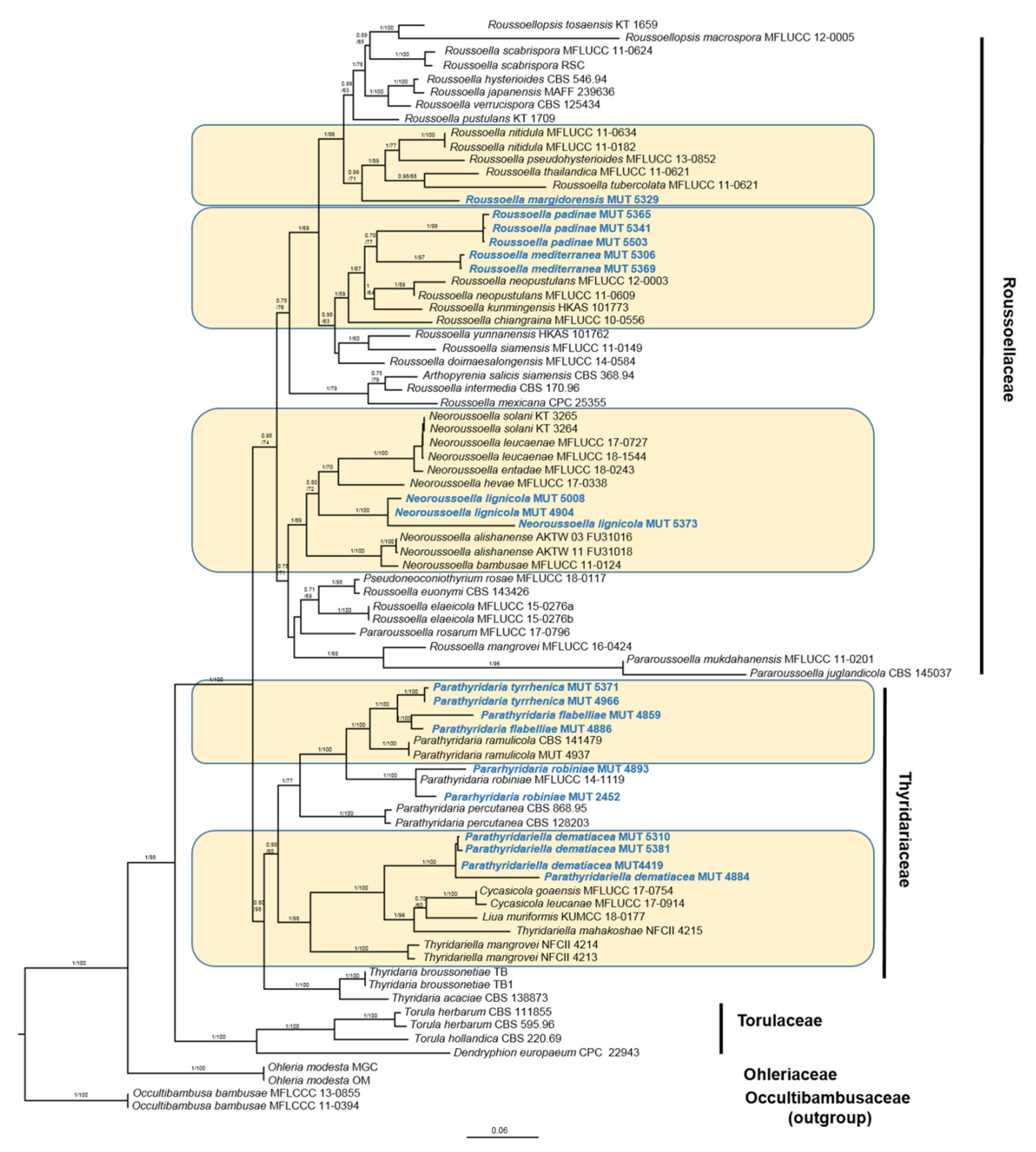
3.1. Phylogenetic Inference
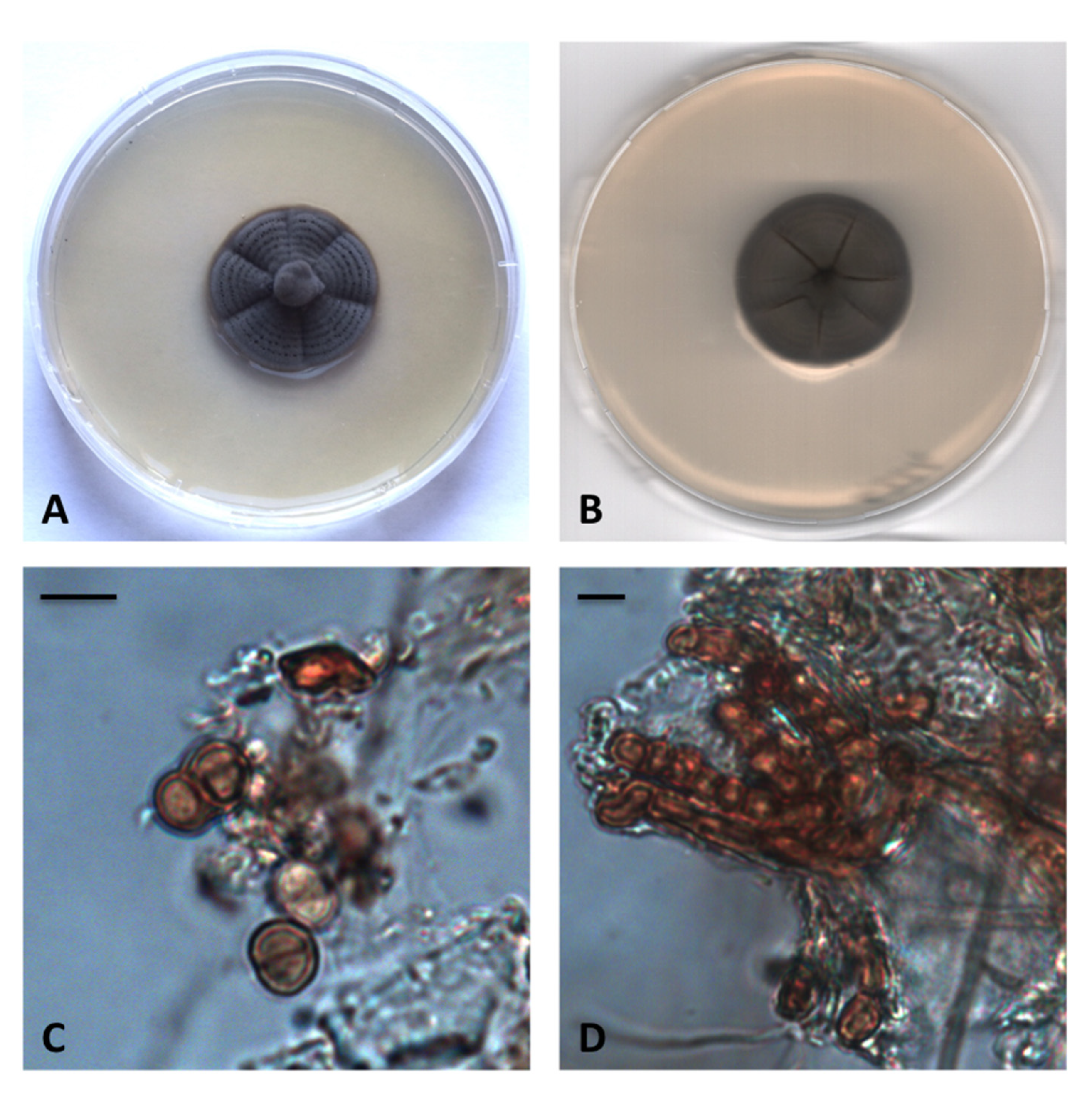
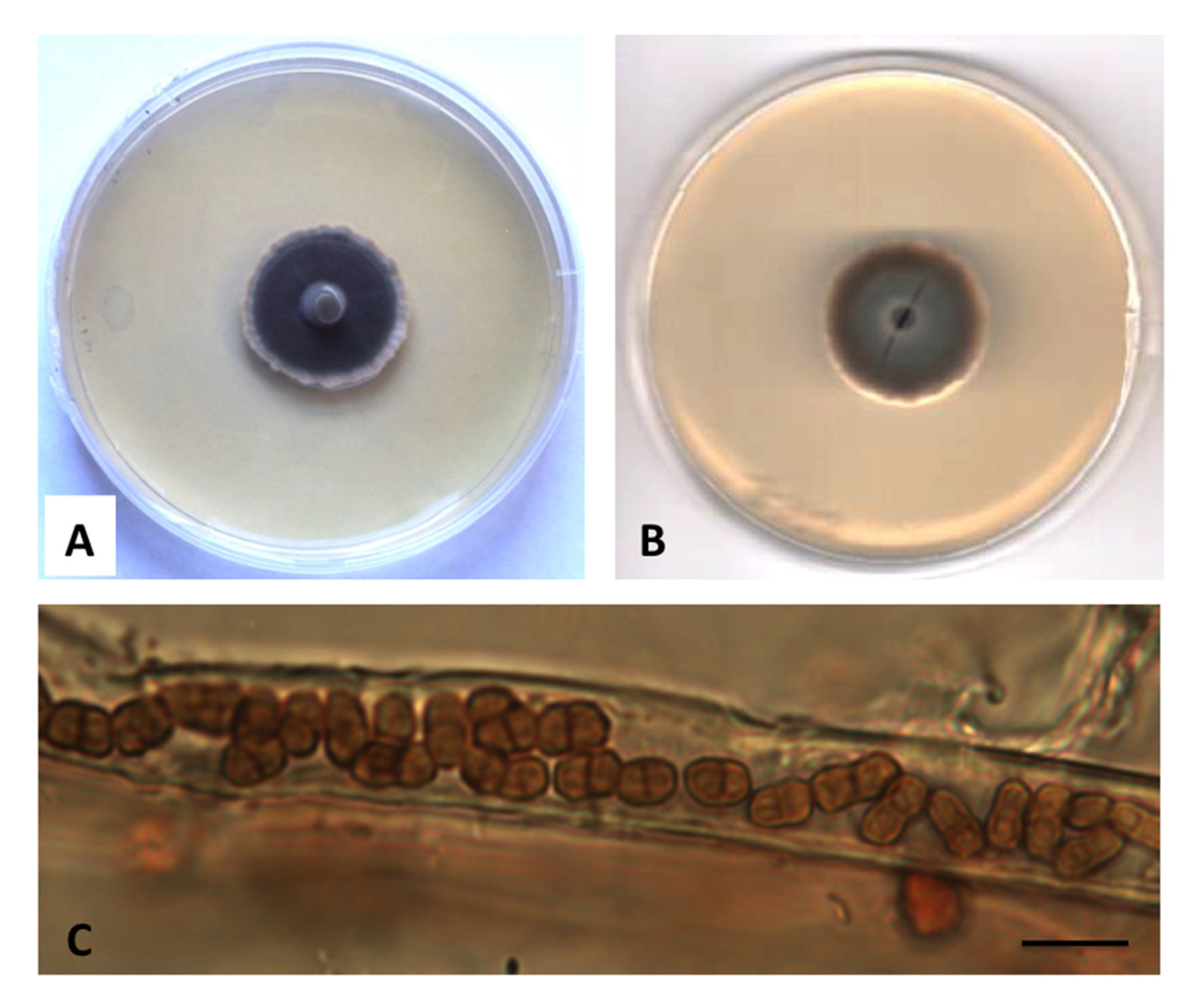
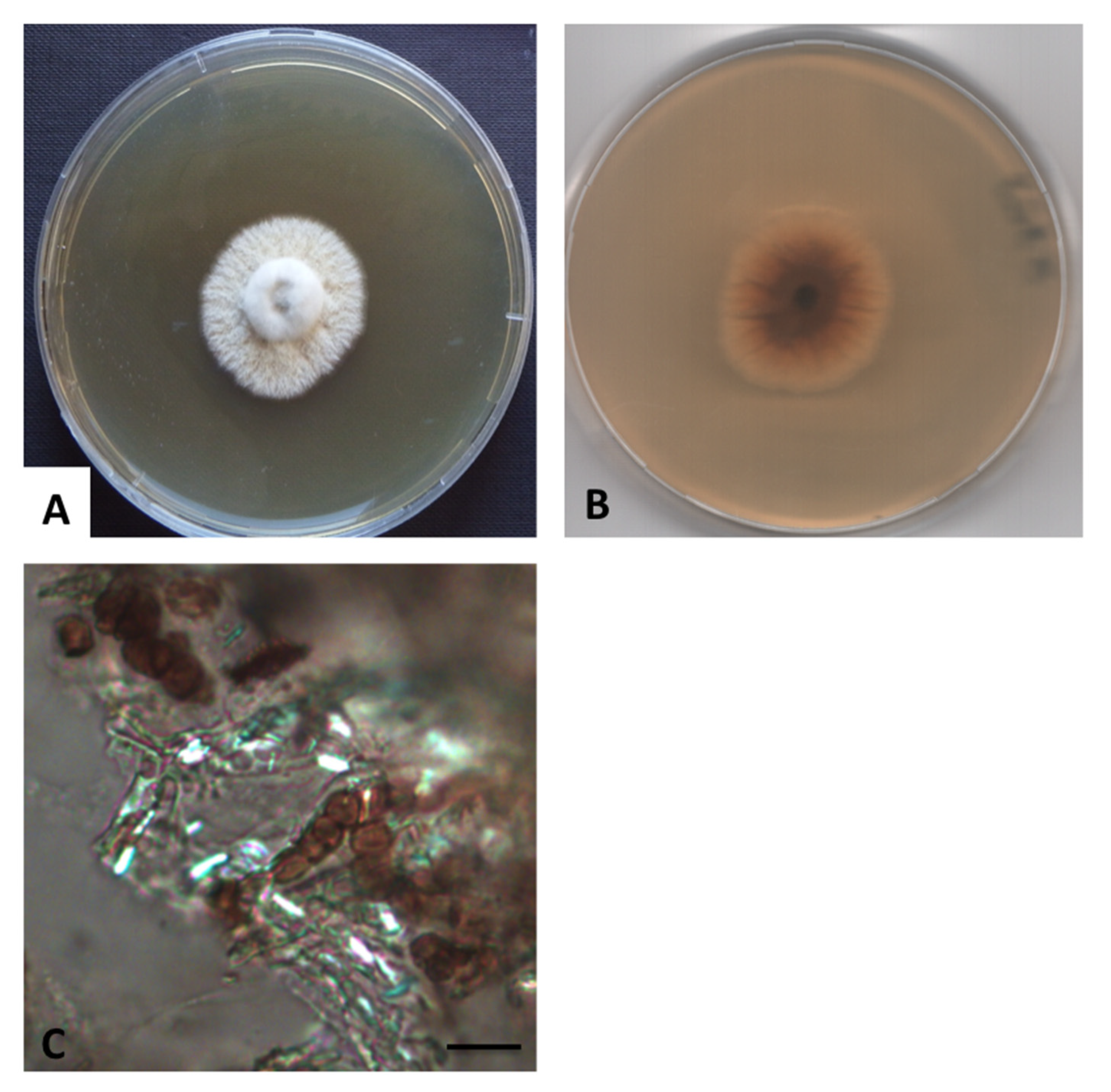
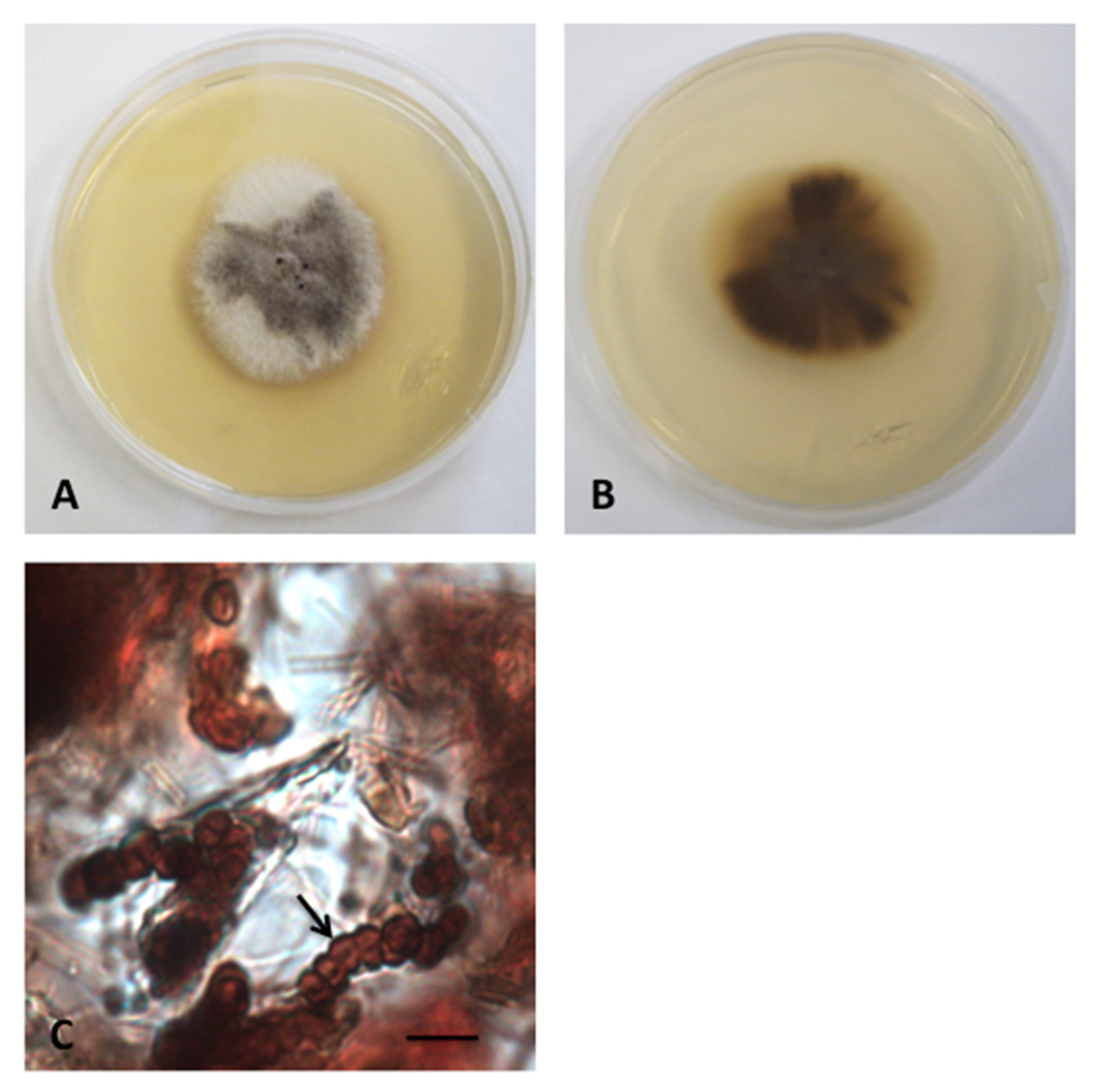
3.2. Taxonomy
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Richards, T.A.; Jones, M.D.; Leonard, G.; Bass, D. Marine fungi: Their ecology and molecular diversity. Ann. Rev. Mar. Sci. 2012, 4, 495–522. [Google Scholar] [CrossRef] [PubMed]
- Amend, A.; Burgaud, G.; Cunliffe, M.; Edgcomb, V.P.; Ettinger, C.L.; Gutierrez, M.H.; Heitman, J.; Hom, E.F.Y.; Ianiri, G.; Jones, A.C.; et al. Fungi in the marine environment: Open questions and unsolved problems. Mbio 2019, 10, 15. [Google Scholar] [CrossRef] [PubMed]
- Carroll, A.R.; Copp, B.R.; Davis, R.A.; Keyzers, R.A.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2019, 36, 122–173. [Google Scholar] [CrossRef] [PubMed]
- Jones, E.G.; Pang, K.-L. Marine fungi and fungal-like organisms; Jones, E.G., Pang, K.-L., Eds.; Walter de Gruyter: Berlin, Germany, 2012. [Google Scholar]
- Raghukumar, S. Fungi in Coastal and Oceanic Marine Ecosystems: Marine Fungi; Springer: Cham, Switzerland, 2017. [Google Scholar]
- Pang, K.L.; Overy, D.P.; Jones, E.B.G.; Calado, M.D.; Burgaud, G.; Walker, A.K.; Johnson, J.A.; Kerr, R.G.; Cha, H.J.; Bills, G.F. ‘Marine fungi’ and ‘marine-derived fungi’ in natural product chemistry research: Toward a new consensual definition. Fungal Biol. Rev. 2016, 30, 163–175. [Google Scholar] [CrossRef]
- Jones, E.B.G.; Pang, K.-L.; Abdel-Wahab, M.A.; Scholz, B.; Hyde, K.D.; Boekhout, T.; Ebel, R.; Rateb, M.E.; Henderson, L.; Sakayaroj, J.; et al. An online resource for marine fungi. Fungal Divers. 2019, 96, 347–433. [Google Scholar] [CrossRef]
- Jones, E.B.G. Fifty years of marine mycology. Fungal Divers. 2011, 50, 73–112. [Google Scholar] [CrossRef]
- Jones, E.B.G.; Suetrong, S.; Sakayaroj, J.; Bahkali, A.H.; Abdel-Wahab, M.A.; Boekhout, T.; Pang, K.L. Classification of marine Ascomycota, Basidiomycota, Blastocladiomycota and Chytridiomycota. Fungal Divers. 2015, 73, 1–72. [Google Scholar] [CrossRef]
- Garzoli, L.; Poli, A.; Prigione, V.; Gnavi, G.; Varese, G.C. Peacock’s tail with a fungal cocktail: First assessment of the mycobiota associated with the brown alga Padina pavonica. Fungal Ecol. 2018, 35, 87–97. [Google Scholar] [CrossRef]
- Gnavi, G.; Garzoli, L.; Polil, A.; Prigione, V.; Burgaud, G.; Varese, G.C. The culturable mycobiota of Flabellia petiolata: First survey of marine fungi associated to a Mediterranean green alga. PLoS ONE 2017, 12. [Google Scholar] [CrossRef]
- Panno, L.; Bruno, M.; Voyron, S.; Anastasi, A.; Gnavi, G.; Miserere, L.; Varese, G.C. Diversity, ecological role and potential biotechnological applications of marine fungi associated to the seagrass Posidonia oceanica. New Biotechnol. 2013, 30, 685–694. [Google Scholar] [CrossRef]
- Bovio, E.; Garzoli, L.; Poli, A.; Prigione, V.; Firsova, D.; McCormack, G.; Varese, G. The culturable mycobiota associated with three Atlantic sponges, including two new species: Thelebolus balaustiformis and T. spongiae. Fungal Syst. Evol. 2018, 1, 141–167. [Google Scholar] [CrossRef]
- Liu, J.-K.; Phookamsak, R.; Dai, D.-Q.; Tanaka, K.; Jones, E.; Xu, J.-C.; Chukeatirote, E.; Hyde, K.D. Roussoellaceae, a new pleosporalean family to accommodate the genera Neoroussoella gen. nov., Roussoella and Roussoellopsis. Phytotaxa 2014, 181, 1–33. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Voglmayr, H. Hidden diversity in Thyridaria and a new circumscription of the Thyridariaceae. Stud. Mycol. 2016, 85, 35–64. [Google Scholar] [CrossRef]
- Tibpromma, S.; Hyde, K.D.; Jeewon, R.; Maharachchikumbura, S.S.N.; Liu, J.K.; Bhat, D.J.; Jones, E.B.G.; McKenzie, E.H.C.; Camporesi, E.; Bulgakov, T.S.; et al. Fungal diversity notes 491–602: Taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers. 2017, 83, 1–261. [Google Scholar] [CrossRef]
- Devadatha, B.; Sarma, V.V.; Jeewon, R.; Wanasinghe, D.N.; Hyde, K.D.; Jones, E.B.G. Thyridariella, a novel marine fungal genus from India: Morphological characterization and phylogeny inferred from multigene DNA sequence analyses. Mycol. Prog. 2018, 17, 791–804. [Google Scholar] [CrossRef]
- Jayasiri, S.; Hyde, K.; Jones, E.; McKenzie, E.; Jeewon, R.; Phillips, A.; Bhat, D.; Wanasinghe, D.; Liu, J.; Lu, Y.; et al. Diversity, morphology and molecular phylogeny of Dothideomycetes on decaying wild seed pods and fruits. Mycosphere 2019, 10, 1–186. [Google Scholar] [CrossRef]
- Phookamsak, R.; Hyde, K.D.; Jeewon, R.; Bhat, D.J.; Jones, E.B.G.; Maharachchikumbura, S.S.N.; Raspe, O.; Karunarathna, S.C.; Wanasinghe, D.N.; Hongsanan, S.; et al. Fungal diversity notes 929–1035: Taxonomic and phylogenetic contributions on genera and species of fungi. Fungal Divers. 2019, 95, 1–273. [Google Scholar] [CrossRef]
- Jiang, H.B.; Hyde, K.D.; Jayawardena, R.S.; Doilom, M.; Xu, J.C.; Phookamsak, R. Taxonomic and phylogenetic characterizations reveal two new species and two new records of Roussoella (Roussoellaceae, Pleosporales) from Yunnan, China. Mycol. Prog. 2019, 18, 577–591. [Google Scholar] [CrossRef]
- Dai, D.Q.; Phookamsak, R.; Wijayawardene, N.N.; Li, W.J.; Bhat, D.J.; Xu, J.C.; Taylor, J.E.; Hyde, K.D.; Chukeatirote, E. Bambusicolous fungi. Fungal Divers. 2017, 82, 1–105. [Google Scholar] [CrossRef]
- Panebianco, C.; Tam, W.Y.; Jones, E.B.G. The effect of pre-inoculation of balsa wood by selected marine fungi and their effect on subsequent colonisation in the sea. Fungal Divers. 2002, 10, 77–88. [Google Scholar]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR protocols: A guide to methods and applications; Academic Press: New York, NY, USA, 1990; Volume 18, pp. 315–322. [Google Scholar]
- Vilgalys, R.; Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 1990, 172, 4238–4246. [Google Scholar] [CrossRef]
- Stielow, J.; Lévesque, C.; Seifert, K.; Meyer, W.; Iriny, L.; Smits, D.; Renfurm, R.; Verkley, G.; Groenewald, M.; Chaduli, D. One fungus, which genes? Development and assessment of universal primers for potential secondary fungal DNA barcodes. Persoonia 2015, 35, 242–263. [Google Scholar] [CrossRef]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerse II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef]
- Vaidya, G.; Lohman, D.J.; Meier, R. SequenceMatrix: Concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 2011, 27, 171–180. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Hohna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian Phylogenetic Inference and Model Choice Across a Large Model Space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Lieckfeldt, E.; Meyer, W.; Borner, T. Rapid identification and differentiation of yeasts by DNA and PCR fingerprinting. J. Basic Microbiol. 1993, 33, 413–426. [Google Scholar] [CrossRef]
- Poli, A.; Lazzari, A.; Prigione, V.; Voyron, S.; Spadaro, D.; Varese, G.C. Influence of plant genotype on the cultivable fungi associated to tomato rhizosphere and roots in different soils. Fungal Biol. 2016, 120, 862–872. [Google Scholar] [CrossRef]
- Poli, A.; Vizzini, A.; Prigione, V.; Varese, G.C. Basidiomycota isolated from the Mediterranean Sea-Phylogeny and putative ecological roles. Fungal Ecol. 2018, 36, 51–62. [Google Scholar] [CrossRef]
- Oses, R.; Valenzuela, S.; Freer, J.; Sanfuentes, E.; Rodriguez, J. Fungal endophytes in xylem of healthy Chilean trees and their possible role in early wood decay. Fungal Divers. 2008, 33, 77–86. [Google Scholar]
- Vohnik, M.; Borovce, O.; Kolarikova, Z.; Sudova, R.; Reblova, M. Extensive sampling and high-throughput sequencing reveal Posidoniomyces atricolor gen. et sp. nov. (Aigialaceae, Pleosporales) as the dominant root mycobiont of the dominant Mediterranean seagrass Posidoniaoceanica. Mycokeys 2019, 55, 59–86. [Google Scholar] [CrossRef] [PubMed]



| Species | Strain Code | Source | nrITS | nrSSU | nrLSU | TEF-1α | RPB2 |
|---|---|---|---|---|---|---|---|
| Roussoellaceae | |||||||
| Arthopyrenia salicis Massal | CBS 368.94 | Salix bark | KF443410 | AY538333 | AY538339 | KF443404 | KF443397 |
| Neoroussoella bambusae Liu and Hyde | MFLUCC 11-0124 | Dead branch of Bambusa | KJ474827 | -- | KJ474839 | KJ474848 | KJ474856 |
| N. alishanense Karunarathna, Kuo, Phookamsak and Hyde | AKTW 03 FU31016 | Pennisetum purpureum | MK503816 | MK503828 | MK503822 | MK336181 | MN037756 |
| AKTW 11 FU31018 | Pennisetum purpureum | MK503818 | MK503830 | MK503824 | MK336182 | MN037757 | |
| N. entadae Jones and Hyde | MFLUCC 18-0243 | Leucaena sp. | MK347786 | MK347893 | MK348004 | MK360065 | MK434866 |
| N. heveae Senwanna, Phookamsak and Hyde | MFLUCC 17-1983 | Twig of Hevea brasiliensis | MH590693 | -- | MH590689 | -- | -- |
| N. leucaenae Jones and Hyde | MFLUCC 18-1544 | Decaying pod of Leucaena | MK347767 | MK347874 | MK347984 | MK360067 | MK434876 |
| MFLUCC 17-0927 | Pterocarpus sp. | MK347733 | MK347841 | MK347950 | MK360066 | MK434896 | |
| Neoroussoella lignicolasp. nov. | MUT 4904 | P. pavonica | KT699129 | MN556307* | MN556319* | MN605894* | MN605914* |
| MUT 5008 | P. oceanica leaves | MN556317* | MN556308* | MN556320* | MN605895* | MN605915* | |
| MUT 5373 | P. pavonica | KU314953 | KU314954 | MN556321* | MN605896* | MN605916* | |
| Pararoussoella juglandicola Crous and Schumach | CBS 145037 | MK442607 | -- | MK442543 | MK442699 | MK442671 | |
| P. mukdahanensis (Phookamsak, Dai and Hyde) Crous | MFLUCC 11-0201 | Bamboo | KU940129 | KU872121 | KU863118 | -- | -- |
| P. rosarum Jones and Hyde | MFLUCC 17-0796 | Rosa sp. | MG8289391 | MG829154 | MG829048 | MG829224 | -- |
| Pseudoneoconiothyrium rosae (Phukhams., Camporesi and Hyde) Phukhams., Camporesi and Hyde | MFLUCC 15-0052 | Dead aerial spines of Rosa canina | MG828922 | MG829138 | MG829032 | -- | -- |
| Roussoella chiangraina Phookamsak, Liu and Hyde | MFLUCC 10-0556 | Dead branch of bamboo | KJ474828 | -- | KJ474840 | KJ474849 | KJ474857 |
| R. doimaesalongensis Thambug. and Hyde | MFLUCC 14–0584 | Dead branch of bamboo | KY026584 | -- | KY000659 | KY651249 | KY678394 |
| R. elaeicola Konta. and Hyde | MFLUCC 15-0276a | Dead petiole of Elaeis guineensis | MH742329 | -- | MH742326 | -- | -- |
| MFLUCC 15-0276b | Dead petiole of Elaeis guineensis | MH742330 | -- | MH742327 | -- | -- | |
| R. euonymi Crous and Akulov | CBS 143426 | Fallen branches of Euonymus europaeus | MH107915 | -- | MH107961 | -- | MH108007 |
| R. hysterioides (Ces.) Höhn. | CBS 546.94 | Phyllostachys | KF443405 | AY642528 | KF443381 | KF443399 | KF443392 |
| R. intermedia Ju, Rogers and Huhndorf | CBS 170.96 | Bamboo | KF443407 | KF443390 | KF443382 | KF443398 | KF443394 |
| R. japanensisKaz. Tanaka, Liu and Hyde | MAFF 239636 | Twigs of Sasa veitchii | KJ474829 | AB524480 | AB524621 | AB539114 | AB539101 |
| R. kunmingensis Jiang, Phookamsak and Hyde | KUMCC 18-0128 | Dead bamboo | MH453491 | -- | MH453487 | MH453480 | MH453484 |
| R. mangrovei Phukhams. and Hyde | MFLUCC 16-0424 | Dead branches of Rhizophora | MH025951 | -- | MH023318 | MH028246 | MH028250 |
| Roussoella margidorensis sp. nov. | MUT 5329 | P. pavonica | KU314944 | MN556309* | MN556322* | MN605897* | MN605917* |
| Roussoella mediterranea sp. nov. | MUT 5306 | P. pavonica | KU255054 | MN556310* | MN556323* | MN605898* | MN605918* |
| MUT 5369 | P. pavonica | KU314947 | KU314948 | MN556324* | MN605899* | MN605919* | |
| R. mexicana Crous and Yáñez-Mor. | CPC 25355 | Leaf spots of Coffea arabica | KT950848 | -- | KT950862 | -- | -- |
| R. neopustulans Dai, Liu and Hyde | MFLUCC 11-0609 | Bamboo | KJ474833 | -- | KJ474841 | KJ474850 | |
| MFLUCC 12-0003 | Bamboo | KU940130 | KU872122 | KU863119 | -- | -- | |
| R. nitidula Sacc. and Paol. | MFLUCC 11-0182 | Bamboo | KJ474835 | -- | KJ474843 | KJ474852 | KJ474859 |
| MFLUCC 11-0634 | Bamboo | KJ474834 | -- | KJ474842 | KJ474851 | KJ474858 | |
| Roussoella padinae sp. nov. | MUT 5341 | P. pavonica | KU158153 | KU158176 | MN556325* | MN605900* | MN605920* |
| MUT 5365 | P. pavonica | KU158170 | KU158179 | MN556326* | MN605901* | MN605921* | |
| MUT 5503 | P. pavonica | KU158170 | MN556312* | MN556327* | MN605902* | MN605922* | |
| R. pseudohysterioides Dai and Hyde | MFLUCC 13-0852 | Bamboo | KU940131 | KU872123 | KU863120 | KU940198 | -- |
| R. pustulans (Ellis and Everh.) Ju, Rogers and Huhndorf | KT 1709 | Culms of Sasa kurilensis | KJ474830 | AB524482 | AB524623 | AB539116 | AB539103 |
| R. scabrispora (Höhn.) Aptroot | MFLUCC 11-0624 | Bamboo | KJ474836 | -- | KJ474844 | KJ474853 | KJ474860 |
| RSC | Bamboo | KX650566 | -- | KX650566 | KX650537 | -- | |
| R. siamensis Phookamsak, Liu and Hyde | MFLUCC 11-0149 | Bamboo | KJ474837 | KU872125 | KJ474845 | KJ474854 | KJ474861 |
| R. thailandica Dai, Liu and Hyde | MFLUCC 11-0621 | Bamboo | KJ474838 | -- | KJ474846 | -- | |
| R. tuberculata Dai and Hyde | MFLUCC 13-0854 | Bamboo | KU940132 | KU872124 | KU863121 | KU940199 | |
| R. verrucispora Kaz. Tanaka, Liu and Hyde | CBS 125434 | Sasa kurilensis | KJ474832 | AB52448 | AB524622 | AB539115 | -- |
| R. yunnanensis Jiang, Phookamsak and Hyde | KUMCC 18-0115 | Dead bamboo | MH453492 | -- | MH453488 | MH453481 | -- |
| Roussoellopsis macrospora (Hino and Katum.) Hinoand Katum | MFLUCC 12-0005 | Bamboo | KJ739604 | KJ739608 | KJ474847 | KJ474855 | KJ474862 |
| Ro. tosaensis (Hino and Katum.) Hino and Katum | KT 1659 | Culms of bamboo | -- | AB524484 | AB524625 | AB539117 | AB539104 |
| Thyridariaceae | |||||||
| Cycasicola goaensis Jones and Hyde | MFLUCC 17-0754 | Cycas sp. | MG828885 | MG829112 | MG829001 | MG829198 | -- |
| C. leucaenae Jones and Hyde | MFLUCC 17-0914 | Leucaena leucocephala | MK34772 | MK347833 | MK347942 | MK360046 | MK434900 |
| Liua muriformis Phookamsak, Jiang and Hyde | KUMCC 18-0177 | Dead hanging branches of Lonicera maackii | MK433599 | MK433595 | MK433598 | MK426798 | MK426799 |
| Parathyridaria percutanea (Ahmed, Stevens, van de Sande and de Hoog) Jaklitsch and Voglmayr | CBS 868.95 | Human | KF322118 | KF366451 | KF366449 | KF407987 | KF366452 |
| CBS 128203 | Human | KF322117 | KF366450 | KF366448 | KF407988 | KF366453 | |
| P. ramulicola Jaklitsch, Fourn and Voglmayr | CBS 141479 | Twigs of Ribes rubrum | NR_147657 | KX650514 | KX650565 | KX650536 | KX650584 |
| MUT 4397 | P. oceanica | KC339235 | MN556311* | KF636775 | MN605913* | MN605933* | |
| P. robiniaeMapook, Camporesi and Hyde | MFLUCC 14-1119 | Dead branch of Robinia pseudoacacia | KY511142 | -- | KY511141 | KY549682 | -- |
| MUT 2452 | Dysidea fragilis | MG813183 | MN556312* | MG816491 | MN605903* | MN605923* | |
| MUT 4893 | P. pavonica | KM355998 | KM355993 | MN556328* | MN605904* | MN605924* | |
| Parathyridaria flabelliae sp. nov. | MUT 4859 | F. petiolata | KR014355 | KT587315 | KP671716 | MN605909* | MN605929* |
| MUT 4886 | F. petiolata | KR014358 | KT587317 | KP671720 | MN605910* | MN605930* | |
| Parathyridaria tyrrhenica sp. nov. | MUT 4966 | F. petiolata, | KR014366 | KT587309 | KP671740 | MN605911* | MN605931* |
| MUT 5371 | P. pavonica | KU314951 | KU314952 | MN556329* | MN605912* | MN605932* | |
| Parathyridariella dematiacea sp. nov. | MUT 4419 | P. oceanica rhizomes | KC339245 | MN556313* | KF636786 | MN605905* | MN605925* |
| MUT 4884 | F. petiolata | MN556317* | KT587329 | KP671726 | MN605906* | MN605926* | |
| MUT 5310 | P. pavonica | KU255057 | MN556314* | MN556330* | MN605907* | MN605927* | |
| MUT 5381 | P. pavonica | KU314959 | KU314960 | MN556331* | MN605908* | MN605928* | |
| Thyridaria acaciae (Crous and Wingf.) Jaklitsch and Voglmayr | CBS 138873 | Leaves of Acacia tortilis | KP004469 | -- | KP004497 | -- | -- |
| T. broussonetiae (Sacc.) Traverso | TB | Hippocrepis emerus | KX650567 | -- | KX650567 | KX650538 | KX650585 |
| TB1 | Amorpha fruticosa | KX650568 | KX650515 | KX650568 | KX650539 | KX650586 | |
| Thyridariella mahakoshae Devadatha, Sarma, Wanas., Hyde and Jones | NFCCI 4215 | Decaying wood Avicennia marina | MG020435 | MG020441 | MG020438 | MG023140 | MG020446 |
| Th. mangrovei Devadatha, Sarma, Hyde, Wanas. and Jones | NFCCI 4213 | Decaying wood Avicennia marina | MG020434 | MG020440 | MG020437 | MG020443 | MG020445 |
| NFCCI 4214 | Decaying wood Avicennia marina | MG020436 | MG020442 | MG020439 | MG020444 | MG020447 | |
| Occultibambusaceae | |||||||
| Occultibambusa bambusae Dai and Hyde | MFLUCC 11-0394 | Bamboo | KU940124 | -- | KU863113 | KU940194 | KU940171 |
| MFLUCC 13-0855 | Bamboo | KU940123 | KU872116 | KU863112 | KU940193 | KU940170 | |
| Ohleriaceae | |||||||
| Ohleria modesta Fuckel | MGC | KX650562 | -- | KX650562 | KX650533 | KX650582 | |
| OM | Branches of Chamaecytisus proliferus | KX650563 | KX650513 | KX650563 | KX650534 | KX650583 | |
| Torulaceae | |||||||
| Dendryphion europaeum Crous and Schumacher | CPC 22943 | Heracleum sphondylium | KJ869146 | -- | KJ869203 | -- | -- |
| Torula herbarum (Pers.) Link | CBS 111855 | n.a. | KF443409 | KF443391 | KF443386 | KF443403 | KF443396 |
| CBS 595.96 | KF443408 | KF443387 | KF443385 | KF443402 | KF443395 | ||
| Torula hollandica Crous | CBS 220.69 | Delphinium dead stem | KF443406 | -- | MH877717 | -- | -- |
| Forward and Reverse Primers | Thermocycler Conditions | References | |
|---|---|---|---|
| ITS | ITS1- ITS4 | 95 °C: 5 min, (95 °C: 40 s, 55 °C: 50 s, 72 °C: 50 sec) × 35 cycles; 72 °C: 8 min; 4 °C: ∞ | [23] |
| LSU | LR0R-LR7 | 95 °C: 5 min, (95 °C: 1 min, 50 °C: 1 min, 72 °C: 2 min) × 35 cycles; 72 °C: 10 min; 4 °C: ∞ | [24] |
| SSU | NS1-NS4 | 95 °C: 5 min, (95 °C: 1 min, 50 °C: 1 min, 72 °C: 2 min) × 35 cycles; 72 °C: 10 min; 4 °C: ∞ | [23] |
| TEF-1α | 1018F/1620R | 95 °C: 5 min, (95 °C: 1 min, 50 °C: 1 min; 72 °C: 2 min) × 40 cycles, 72 °C: 10 min; 4 °C: ∞ | [25] |
| RPB2 | fRPB2-5F/fPB2-7cR | 94 °C: 3 min, (94 °C: 30 s; 55 °C: 30 s; 72 °C: 1 min) × 40 cycles, 72 °C: 10 min; 4 °C: ∞ | [26] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Poli, A.; Bovio, E.; Ranieri, L.; Varese, G.C.; Prigione, V. News from the Sea: A New Genus and Seven New Species in the Pleosporalean Families Roussoellaceae and Thyridariaceae. Diversity 2020, 12, 144. https://doi.org/10.3390/d12040144
Poli A, Bovio E, Ranieri L, Varese GC, Prigione V. News from the Sea: A New Genus and Seven New Species in the Pleosporalean Families Roussoellaceae and Thyridariaceae. Diversity. 2020; 12(4):144. https://doi.org/10.3390/d12040144
Chicago/Turabian StylePoli, Anna, Elena Bovio, Lucrezia Ranieri, Giovanna Cristina Varese, and Valeria Prigione. 2020. "News from the Sea: A New Genus and Seven New Species in the Pleosporalean Families Roussoellaceae and Thyridariaceae" Diversity 12, no. 4: 144. https://doi.org/10.3390/d12040144
APA StylePoli, A., Bovio, E., Ranieri, L., Varese, G. C., & Prigione, V. (2020). News from the Sea: A New Genus and Seven New Species in the Pleosporalean Families Roussoellaceae and Thyridariaceae. Diversity, 12(4), 144. https://doi.org/10.3390/d12040144








