Recently Naturalized Paraserianthes lophantha subsp. lophantha Displays Contrasting Genetic Diversity and Climate Relationships Compared to Native Populations
Abstract
1. Introduction
2. Materials and Methods
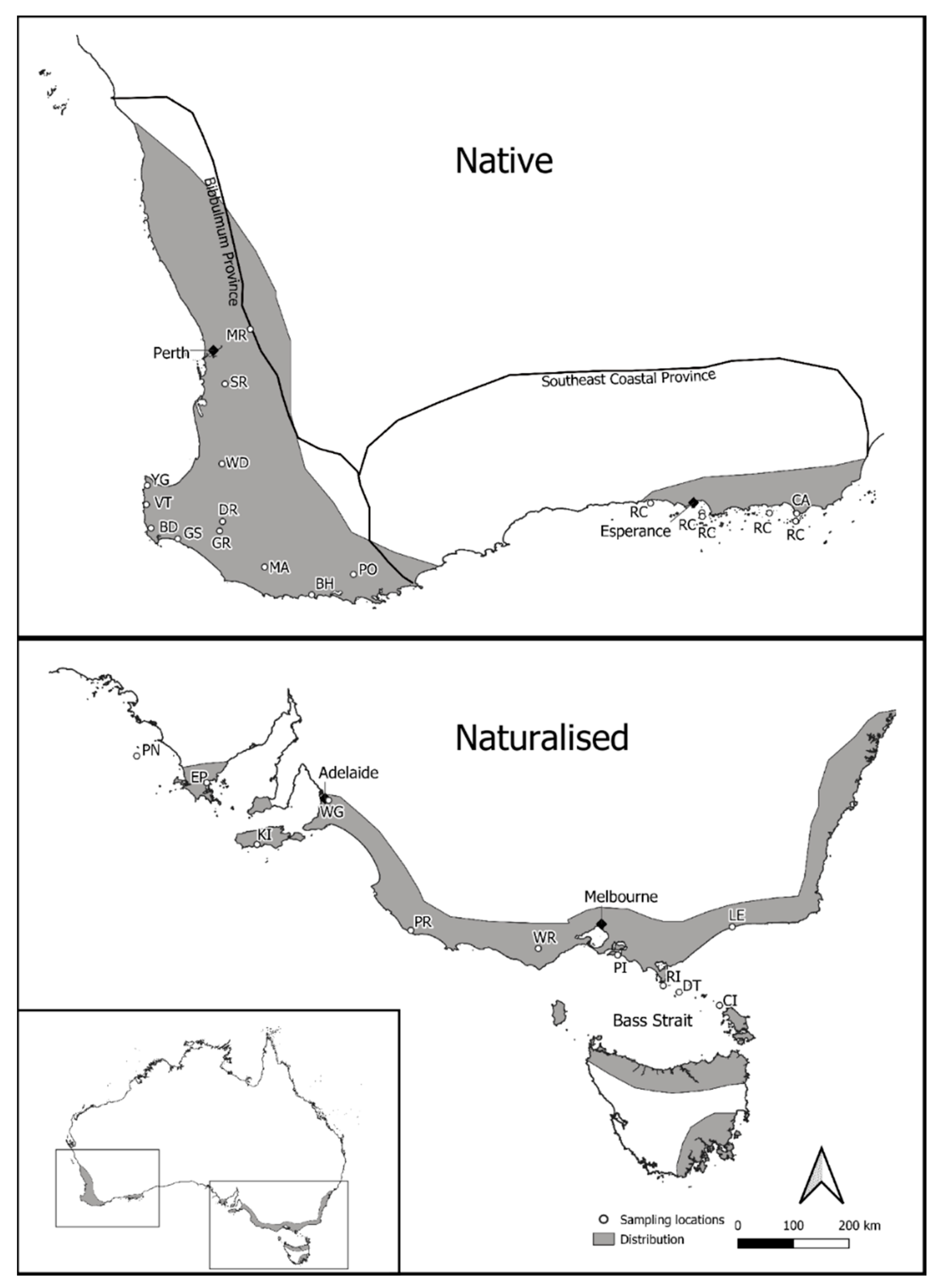
2.1. Sampling
2.2. DNA Isolation
2.3. cpDNA Sequencing and Phylogeny
2.4. Microsatellite Amplification and Screening
2.5. Genetic Analyses
2.6. Assignment of Naturalized Individuals
2.7. Relationship between the Genetic Structure and Climate for Natural and Naturalized Populations
3. Results
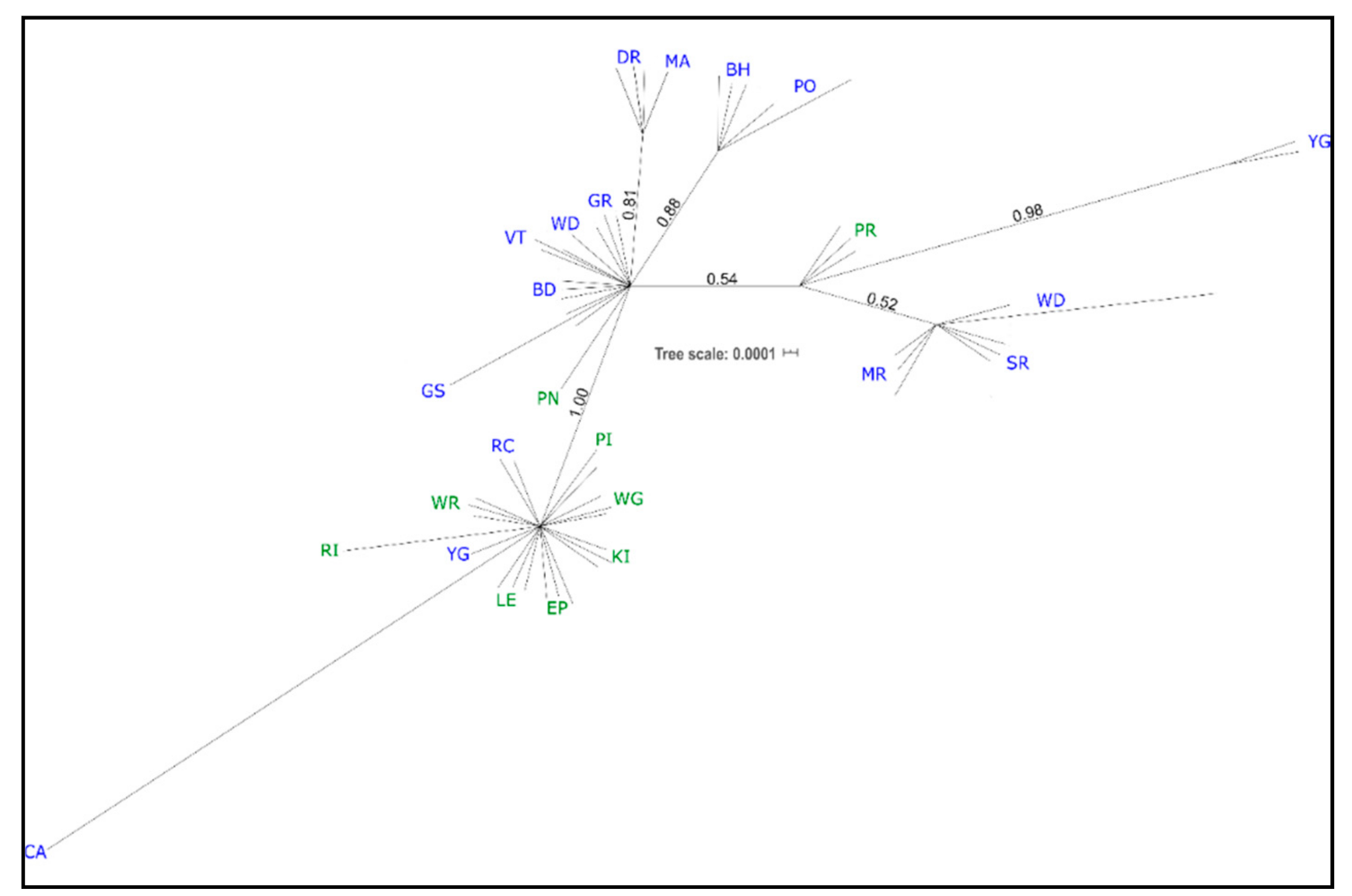
3.1. cpDNA Phylogeny
3.2. Assessment of the Genetic Diversity
3.3. Sibling Assessment
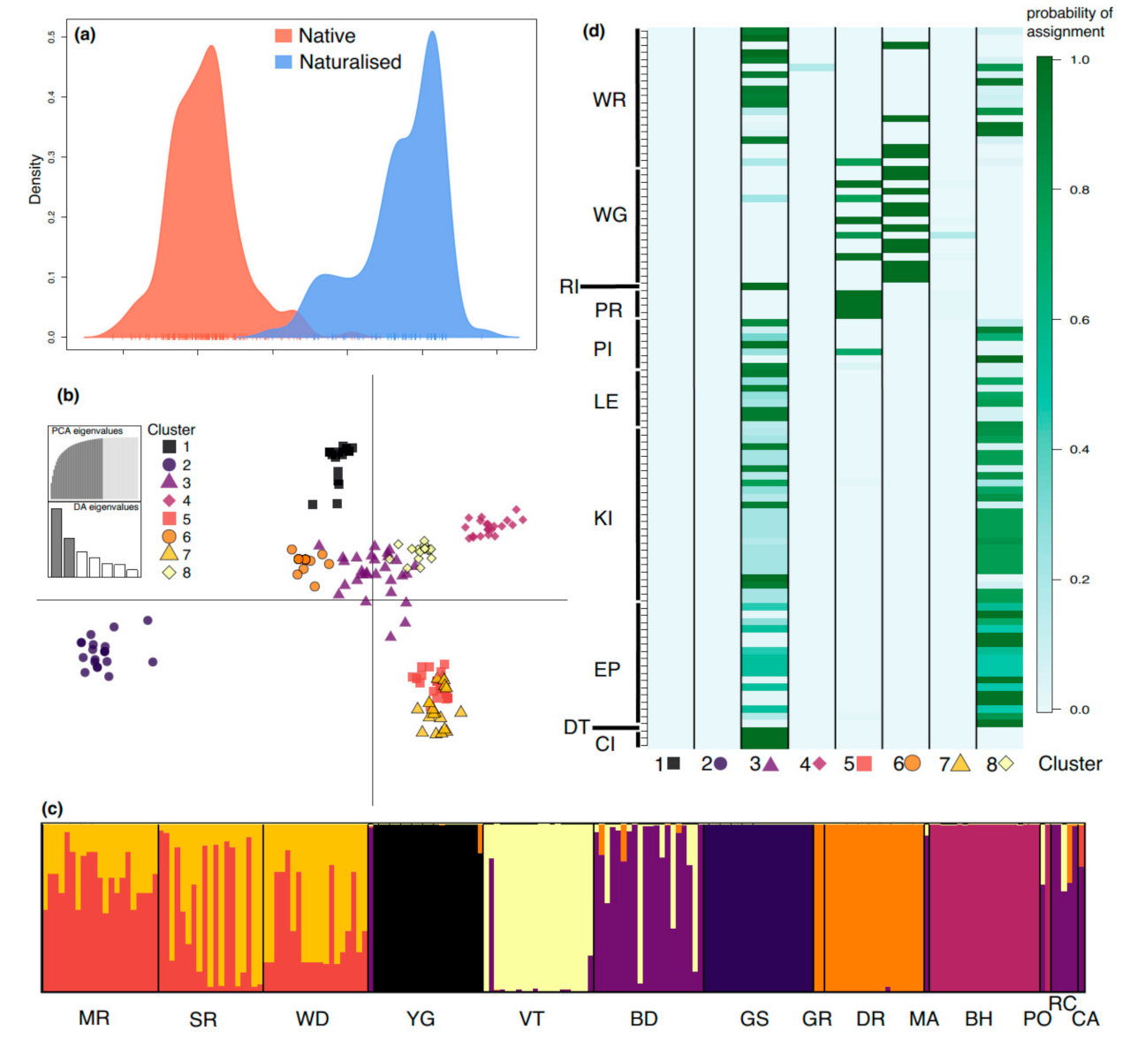
3.4. Cluster Analysis and Assignment of Naturalized Individuals
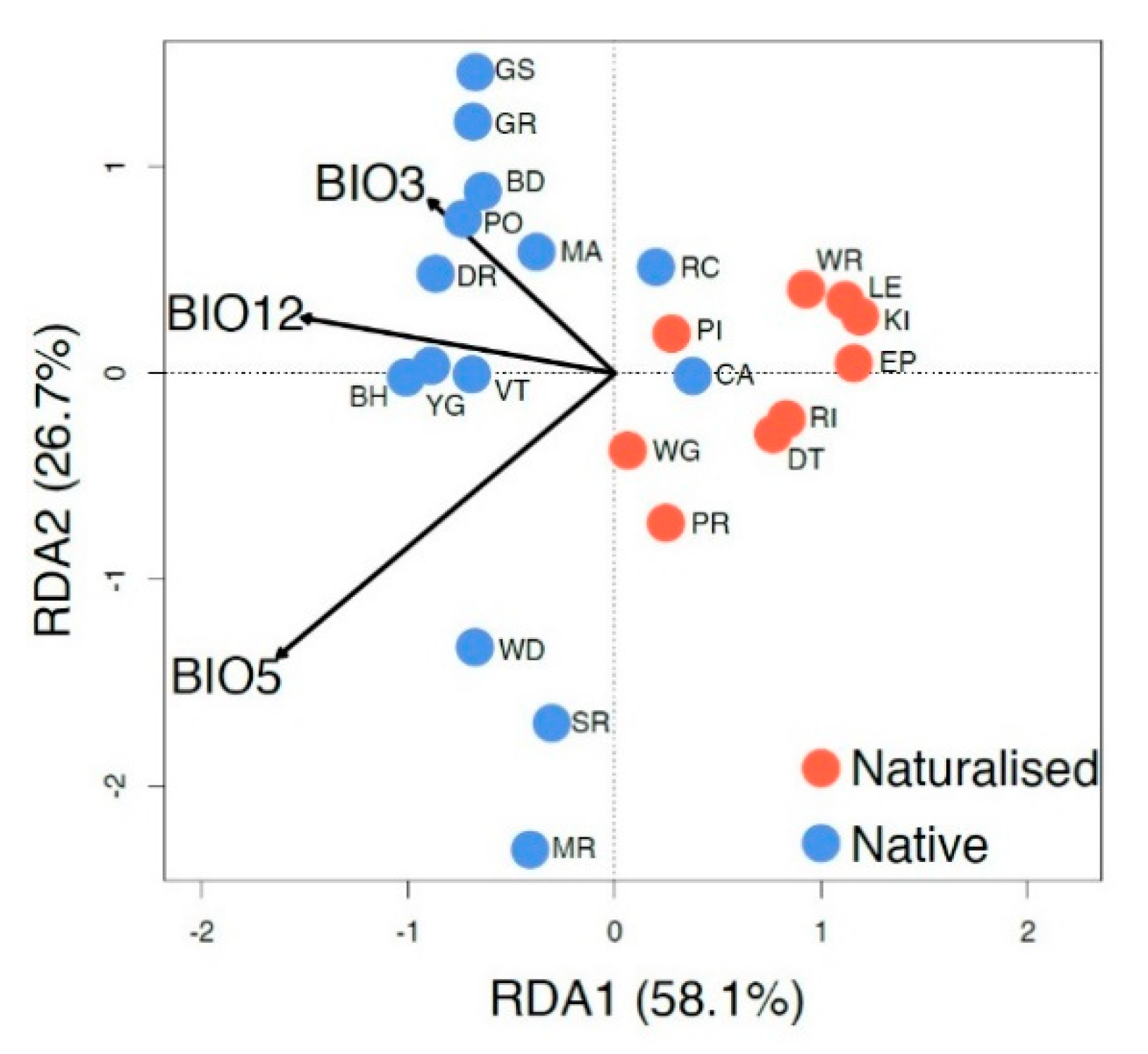
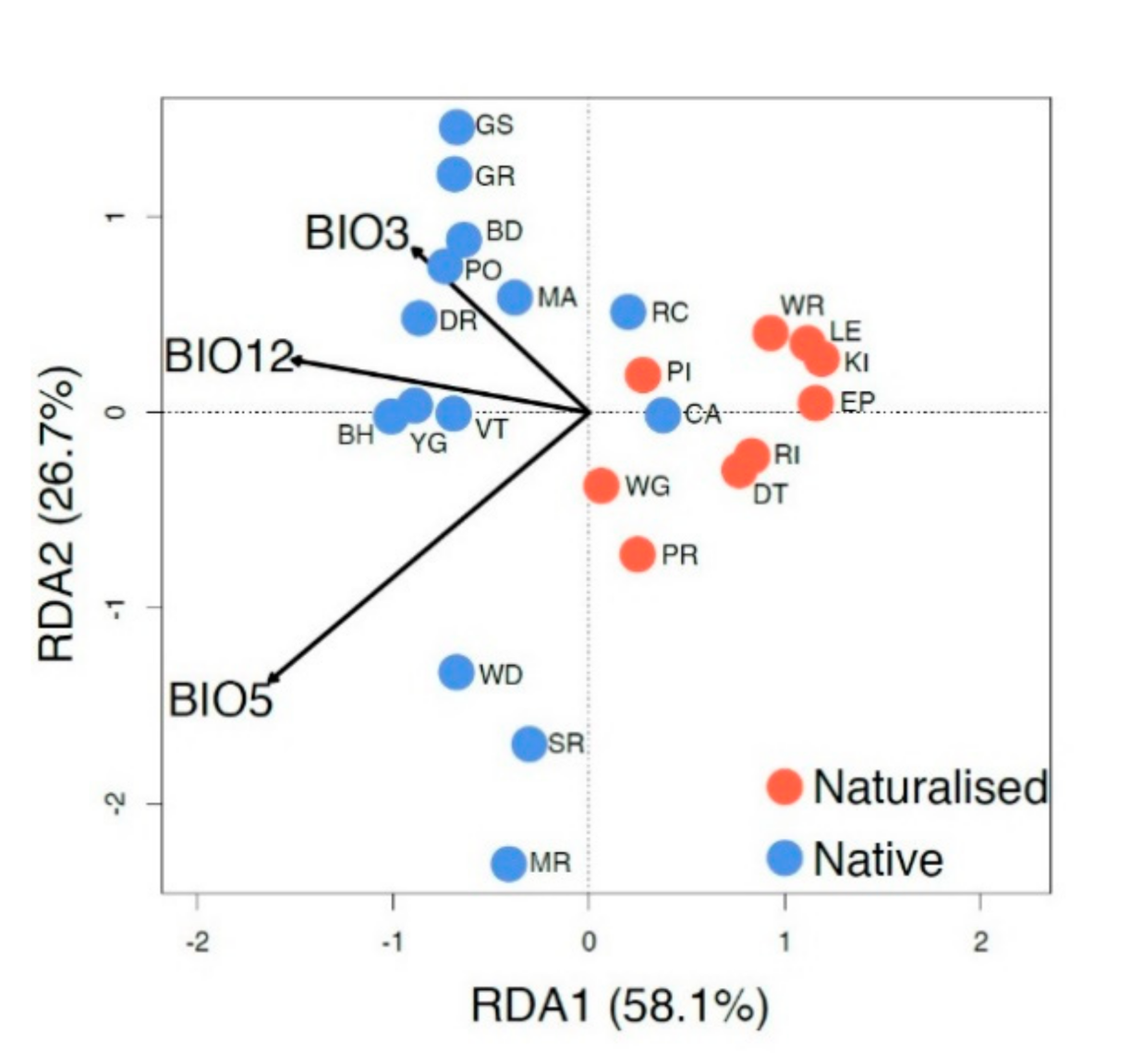
3.5. Relationship between the Genetic Structure and Climate
4. Discussion
4.1. Inbreeding and Allelic Diversity
4.2. Phylogeographic Patterns of P. lophantha within Its Native Range in Australia
4.3. Genetic Diversity of P. lophantha in the Naturalized Range in Australia
4.4. Genetic Implications for Conservation and Management
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Levin, D.A. The Origin, Expansion and Demise of Plant Species; Oxford University Press: New York, NY, USA, 2000. [Google Scholar]
- Anderson, J.T.; Willis, J.H.; Mitchell-Olds, T. Evolutionary genetics of plant adaptation. Trends Genet. 2011, 27, 258–266. [Google Scholar] [CrossRef] [PubMed]
- Kireta, D.; Christmas, M.J.; Lowe, A.J.; Breed, M.F. Disentangling the evolutionary history of three related shrub species using genome-wide molecular markers. Conserv. Genet. 2019, 20, 1101–1112. [Google Scholar] [CrossRef]
- Givnish, T.J. The rise and fall of plant species: A population biologist’s perspective. Am. J. Bot. 2001, 88, 1928–1934. [Google Scholar] [CrossRef]
- Byrne, M. Phylogeography provides an evolutionary context for the conservation of a diverse and ancient flora. Aust. J. Bot. 2007, 55, 316–325. [Google Scholar] [CrossRef]
- Hoffmann, A.A.; Sgró, C.M. Climate change and evolutionary adaptation. Nature 2011, 470, 479–485. [Google Scholar] [CrossRef]
- Hoffmann, A.; Griffin, P.; Dillon, S.; Catullo, R.; Rane, R.; Byrne, M.; Jordan, R.; Oakeshott, J.; Weeks, A.; Joseph, L.; et al. A framework for incorporating evolutionary genomics into biodiversity conservation and management. Clim. Chang. Responses 2015, 2. [Google Scholar] [CrossRef]
- Leimu, R.; Fischer, M. A Meta-Analysis of Local Adaptation in Plants. PLoS ONE 2008, 3, e4010. [Google Scholar] [CrossRef]
- Abbott, R.J.; Brennan, A.C.; James, J.K.; Forbes, D.G.; Hegarty, M.J.; Hiscock, S.J. Recent hybrid origin and invasion of the British Isles by a self-incompatible species, Oxford ragwort (Senecio squalidus L., Asteraceae). Biol. Invasions 2009, 11, 1145–1158. [Google Scholar] [CrossRef]
- Antonelli, A.; Fry, C.; Smith, R.J.; Simmonds, M.S.J.; Kersey, P.J.; Pritchard, H.W.; Abbo, M.S.; Acedo, C.; Adams, J.; Ainsworth, A.M.; et al. State of the World’s Plants and Fungi 2020; Royal Botanic Gardens, Kew: Richmond, UK, 2020; p. 100. [Google Scholar] [CrossRef]
- Commander, L.E.; Coates, D.; Broadhurst, L.; Offord, C.A.; Makinson, R.O.; Matthes, M. Guidelines for the Translocation of Threatened Plants in Australia, 3rd ed.; Australian Network for Plant Conservation: Canberra, Australia, 2018; ISBN 9780975219133. [Google Scholar]
- Finger, A.; Kettle, C.J.; Kaiser-Bunbury, C.N.; Valentin, T.; Doudee, D.; Matatiken, D.; Ghazoul, J. Back from the brink: Potential for genetic rescue in a critically endangered tree. Mol. Ecol. 2011, 20, 3773–3784. [Google Scholar] [CrossRef]
- Gleadow, R.M.; Ashton, D.H. Invasion by Pittosporum undulatum of the Forests of Central Victoria. I. Invasion patterns and plant morphology. Aust. J. Bot. 1981, 29, 705–720. [Google Scholar] [CrossRef]
- Bean, A.R. A new system for determining which plant species are indigenous in Australia. Aust. Syst. Bot. 2007, 20, 1–43. [Google Scholar] [CrossRef]
- Olden, J.D.; Comte, L.; Giam, X. Biotic Homogenisation. eLS 2016, 1–8. [Google Scholar] [CrossRef]
- Dodd, A.J.; Burgman, M.A.; McCarthy, M.A.; Ainsworth, N. The changing patterns of plant naturalization in Australia. Divers. Distrib. 2015, 21, 1038–1050. [Google Scholar] [CrossRef]
- Cook, G.D.; Dias, L. It was no accident: Deliberate plant introductions by Australian government agencies during the 20th century. Aust. J. Bot. 2006, 54, 601. [Google Scholar] [CrossRef]
- Pyšek, P.; Richardson, D.M. Traits Associated with Invasiveness in Alien Plants: Where Do we Stand? In Biological Invasions; Nentwig, W., Ed.; Springer: Berlin/Heidelberg, Germany, 2008; pp. 97–125. [Google Scholar]
- Webber, B.L.; Scott, J.K. Rapid global change: Implications for defining natives and aliens. Glob. Ecol. Biogeogr. 2012, 21, 305–311. [Google Scholar] [CrossRef]
- Duxbury, K. Out of the garden and into the bush. Aust. Gard. Hist. 2002, 14, 2–4. [Google Scholar]
- Nielsen, I.; Guinet, P.; Baretta-Kuipers, T. Studies in the Malesian, Australian and Pacific Ingeae (Leguminosae-Mimosoideae): The genera Archidendropsis, Wallaceodendron, Paraserianthes, Pararchidendron and Serianthes (part 2). Bull. Muséum Natl. d’Histoire Nat. Sect. B Adansonia Bot. Phytochim. 1983, 5, 335–360. [Google Scholar]
- Brown, G.K. Systematics of the tribe Ingeae (Leguminosae-Mimosiodeae) over the last 25 years. Muelleria 2008, 26, 27–42. [Google Scholar]
- Brown, G.K.; Murphy, D.J.; Miller, J.T.; Ladiges, P.Y. Acacia s.s. and its Relationship Among Tropical Legumes, Tribe Ingeae (Leguminosae: Mimosoideae). Syst. Bot. 2008, 33, 739–751. [Google Scholar] [CrossRef]
- GBIF Paraserianthes lophantha subsp. montana GBIF Occurrence Records. Available online: https://www.gbif.org/occurrence/search?country=ID&taxon_key=2943721 (accessed on 27 October 2020).
- Nielsen, I. Mimosaceae (Leguminosae-Mimosoideae). Flora Males. Ser. I Spermatophyta 1992, 11, 1–226. [Google Scholar]
- Hopper, S.D. Biogeographical aspects of speciation in the southwest Australian flora. Annu. Rev. Ecol. Syst. 1979, 10, 399–422. [Google Scholar] [CrossRef]
- Gioia, P.; Hopper, S.D. A new phytogeographic map for the Southwest Australian Floristic Region after an exceptional decade of collection and discovery. Bot. J. Linn. Soc. 2017, 184, 1–15. [Google Scholar] [CrossRef]
- Cowan, R.S. Mimosaceae (excl. Acacia), Caesalpiniaceae. In Flora of Australia; CSIRO: Melbourne, Australia, 1998. [Google Scholar]
- Hopper, S.D.; Gioia, P. The Southwest Australian Floristic Region: Evolution and Conservation of a Global Hot Spot of Biodiversity. Annu. Rev. Ecol. Evol. Syst. 2004, 35, 623–650. [Google Scholar] [CrossRef]
- Characteristic Soils of South Western Australia. Geogr. Inf. Serv. Maps 2012. Available online: https://researchlibrary.agric.wa.gov.au/gis_maps/13/ (accessed on 1 November 2020).
- Tjio, J.H. The somatic chromosomes of some tropical plants. Hereditas 1948, 34, 135–146. [Google Scholar] [CrossRef]
- Fedorov, A.A. (Ed.) Chromosome Numbers of Flowering Plants; Komarov Botanical Institute: Leningrad, Russia, 1969. [Google Scholar]
- Kenrick, J.; Knox, R.B. Function of the polyad in reproduction of Acacia. Ann. Bot. 1982, 50, 721–727. [Google Scholar] [CrossRef]
- Dunphy, B.K.K.; Hamrick, J.L.L. Gene flow among established Puerto Rican populations of the exotic tree species, Albizia lebbeck. Heredity 2005, 94, 418–425. [Google Scholar] [CrossRef]
- Ruiz Guajardo, J.C.; Schnabel, A.; Ennos, R.; Preuss, S.; Otero-Arnaiz, A.; Stone, G. Landscape genetics of the key African acacia species Senegalia mellifera (Vahl)- the importance of the Kenyan Rift Valley. Mol. Ecol. 2010, 19, 5126–5139. [Google Scholar] [CrossRef]
- García-Duro, J.; Cruz, O.; Casal, M.; Reyes, O. Fire as driver of the expansion of Paraserianthes lophantha (Willd.) I. C. Nielsen in SW Europe. Biol. Invasions 2019, 21, 1427–1438. [Google Scholar] [CrossRef]
- Thompson, G.D.; Richardson, D.M.; Wilson, J.R.U.U.; Bellstedt, D.U.; Le Roux, J.J. Genetic diversity and structure of the globally invasive tree, Paraserianthes lophantha subspecies lophantha, suggest an introduction history characterised by varying propagule pressure. Tree Genet. Genomes 2016, 12. [Google Scholar] [CrossRef]
- Harris, C.J.; Dormontt, E.E.; Le Roux, J.J.; Lowe, A.; Leishman, M.R. No consistent association between changes in genetic diversity and adaptive responses of Australian acacias in novel ranges. Evol. Ecol. 2012, 26, 1345–1360. [Google Scholar] [CrossRef]
- Harris, S.; Davis, G. The vegetation and flor of Deal Island, Kent Group. Pap. Proc. R. Soc. Tasman. 1995, 129, 43–51. [Google Scholar]
- Carlyon, K.; Visoiu, M.; Hawkins, C.; Richards, K.; Alderman, R. Rodondo Island, Bass Strait: Biodiversity & Oil Spill Response Survey, January 2015. Nature Conservation Report Series 15/04; Natural and Cultural Heritage Division, DPIPWE: Hobart, Australia, 2015. [Google Scholar]
- Santamarina, S.; Alfaro-Saiz, E.; Llamas, F.; Acedo, C. Different approaches to assess the local invasion risk on a threatened species: Opportunities of using high-resolution species distribution models by selecting the optimal model complexity. Glob. Ecol. Conserv. 2019, 20. [Google Scholar] [CrossRef]
- Australian Weeds Committee Cape Leeuwin Wattle Weeds Australia. Available online: https://web.archive.org/web/20080804001332/http://www.weeds.org.au/cgi-bin/weedident.cgi?tpl=plant.tpl&ibra=all&card=E10 (accessed on 13 July 2020).
- Eurobodalla Shire Council South Coast Weeds—Cape Wattle. Available online: https://www.erbg.org.au/education-in-the-botanic-gardens/identify-your-weed/a-to-z-south-coast-weeds/ (accessed on 13 July 2020).
- Le Roux, J.J.; Brown, G.K.; Byrne, M.; Ndlovu, J.; Richardson, D.M.; Thompson, G.D.; Wilson, J.R.U. Phylogeographic consequences of different introduction histories of invasive Australian Acacia species and Paraserianthes lophantha (Fabaceae) in South Africa. Divers. Distrib. 2011, 17, 861–871. [Google Scholar] [CrossRef]
- Stirton, C. Plant Invaders: Beautiful, but Dangerous: A Guide to the Identification and Control of Twenty-Six Plant Invaders of the Province of the Cape of Good Hope; The Department of Nature and Environment Conservation of the Provincial Administration of the Cape of Good Hope, ABC Press: Cape Town, South Africa, 1978; ISBN 0798400927. [Google Scholar]
- Dennill, G.B.; Donnelly, D. Biological control of Acacia longifolia and related weed species (Fabaceae) in South Africa. Agric. Ecosyst. Environ. 1991, 37, 115–135. [Google Scholar] [CrossRef]
- Carr, G.W.; Yugovic, J.V.; Robinson, K.E. Environmental Weed Invasions in Victoria: Conservation and Management Implications; Dept. of Conservation and Environment and Ecological Horticulture Pty Ltd.: Melbourne, Australia, 1992. [Google Scholar]
- Schmidt, F.; Hoffmann, J.H.; Donnelly, D. Levels of damage caused by Melanterius servulus Pascoe (Coleoptera: Curculionidae), a seed-feeding weevil introduced into South Africa for biological control of Paraserianates lophantha (Fabaceae). Afr. Entomol. 1999, 7, 107–112. [Google Scholar]
- Henderson, L. Alien Weeds and Invasive Plants. A Complete Guide to Declared Weeds and Invaders in South Africa; Agricultural Research Council: Pretoria, South Africa, 2001; ISBN 1868491927. [Google Scholar]
- Muyt, A. Bush Invaders of South-East Australia: A Guide to the Identification and Control of Environmental Weeds Found in South-East Australia; R.G. and F.J. Richardson: Meredith, Australia, 2001; ISBN 0958743975. [Google Scholar]
- Lewis, G.P.; Rico Arce, M.L. Tribe Ingeae. In Legumes of the World; RBG Kew: Richmond, UK, 2005; pp. 193–213. [Google Scholar]
- Cunze, S.; Leiblein, M.C.; Tackenberg, O. Range Expansion of Ambrosia artemisiifolia in Europe Is Promoted by Climate Change. ISRN Ecol. 2013, 2013, 1–9. [Google Scholar] [CrossRef]
- Brown, G.K.; Murphy, D.J.; Ladiges, P.Y. Relationships of the Australo-Malesian genus Paraserianthes (Mimosoideae: Leguminosae) identifies the sister group of Acacia sensu stricto and two biogeographical tracks. Cladistics 2011, 27, 380–390. [Google Scholar] [CrossRef]
- Wheeler, M.A.; Byrne, M. Congruence between phylogeographic patterns in cpDNA variation in Eucalyptus marginata (Myrtaceae) and geomorphology of the Darling Plateau, south-west of Western Australia. Aust. J. Bot. 2006, 54, 17–26. [Google Scholar] [CrossRef]
- Byrne, M.; MacDonald, B.; Coates, D.J. Phylogeographical patterns in chloroplast DNA variation within the Acacia acuminata (Leguminosae: Mimosoideae) complex in Western Australia. J. Evol. Biol. 2002, 15, 576–587. [Google Scholar] [CrossRef]
- Byrne, M.; Hines, B. Phylogeographical analysis of cpDNA variation in Eucalyptus loxophleba (Myrtaceae). Aust. J. Bot. 2004, 52, 459–470. [Google Scholar] [CrossRef]
- Gouws, G.; Stewart, B.A.; Daniels, S.R. Phylogeographic structure of a freshwater crayfish (Decapoda: Parastacidae: Cherax preissii) in south-western Australia. Mar. Freshw. Res. 2006, 57, 837–848. [Google Scholar] [CrossRef]
- Edwards, D.L.; Dale Roberts, J.; Scott Keogh, J. Climatic fluctuations shape the phylogeography of a mesic direct-developing frog from the south-western Australian biodiversity hotspot. J. Biogeogr. 2008, 35, 1803–1815. [Google Scholar] [CrossRef]
- Cooper, S.J.B.; Harvey, M.S.; Saint, K.M.; Main, B.Y. Deep phylogeographic structuring of populations of the trapdoor spider Moggridgea tingle (Migidae) from southwestern Australia: Evidence for long-term refugia within refugia. Mol. Ecol. 2011, 20, 3219–3236. [Google Scholar] [CrossRef] [PubMed]
- Levy, E.; Kennington, W.J.; Tomkins, J.L.; Lebas, N.R. Phylogeography and population genetic structure of the Ornate Dragon Lizard, Ctenophorus ornatus. PLoS ONE 2012, 7, e46351. [Google Scholar] [CrossRef] [PubMed]
- Kay, G.M.; Keogh, J.S. Molecular phylogeny and morphological revision of the Ctenotus labillardieri (Reptilia: Swuamata: Scincidae) species group and a new species of immediate conservation concern in the southwestern Australian biodiversity hotspot. Zootaxa 2012, 3390, 1–18. [Google Scholar] [CrossRef]
- Shaw, J.; Lickey, E.B.; Schilling, E.E.; Small, R.L. Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: The Tortoise and the hare III. Am. J. Bot. 2007, 94, 275–288. [Google Scholar] [CrossRef] [PubMed]
- Johnson, L.A.; Soltis, D.E. matK DNA Sequences and Phylogenetic Reconstruction in Saxifragaceae s. str. Syst. Bot. 1994, 19, 143–156. [Google Scholar] [CrossRef]
- Miller, J.T.; Bayer, R.J. Molecular phylogenetics of Acacia (Fabaceae: Mimosoideae) based on the chloroplast matK coding sequence and flanking trnK intro spacer regions. Am. J. Bot. 2001, 88, 697–705. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Nylander, J.A.A. MrModeltest v2. Program distributed by the author. Evol. Biol. Cent. Upps. Univ. 2004, 2, 1–2. [Google Scholar]
- Simmons, M.P.; Ochoterena, H. Gaps as characters in sequence-based phylogenetic analyses. Syst. Biol. 2000, 49, 369–381. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.K.; Gardner, M.G. Isolation, characterisation and transferability of microsatellites for Paraserianthes lophantha, Cape Wattle (Leguminosae: Mimosoideae): A significant weed worldwide. Muelleria 2011, 29, 87–92. [Google Scholar]
- Schuelke, M. An economic method for the fluorescent labeling of PCR fragments. Nat. Biotechnol. 2000, 18, 233–234. [Google Scholar] [CrossRef] [PubMed]
- Butcher, P.A.; Decroocq, S.; Gray, Y.; Moran, G.F. Development, inheritance and cross-species amplification of microsatellite markers from Acacia mangium. Theor. Appl. Genet. 2000, 101, 1282–1290. [Google Scholar] [CrossRef]
- Millar, M.A.; Byrne, M. Characterization of polymorphic microsatellite DNA markers for Acacia saligna (Labill.) H.L.Wendl. (Mimosaceae). Mol. Ecol. Notes 2007, 7, 1372–1374. [Google Scholar] [CrossRef]
- Grimes, J. Inflorescence morphology, heterochrony, and phylogeny in the mimosoid tribes Ingeae and Acacieae (Leguminosae: Mimosoideae). Bot. Rev. 1999, 65, 317–347. [Google Scholar] [CrossRef]
- Kress, W.J. Sibling Competition and Evolution of Pollen Unit, Ovule Number, and Pollen Vector in Angiosperms. Syst. Bot. 1981, 6, 101. [Google Scholar] [CrossRef]
- Muona, O.; Moran, G.F.; Bell, J.C. Hierarchical patterns of correlated mating in Acacia melanoxylon. Genetics 1991, 127, 619–626. [Google Scholar]
- Casiva, P.V.; Vilardi, J.C.; Cialdella, A.M.; Saidman, B.O. Mating system and population structure of Acacia aroma and A. macracantha (Fabaceae). Am. J. Bot. 2004, 91, 58–64. [Google Scholar] [CrossRef]
- Waples, R.S.; Anderson, E.C. Purging putative siblings from population genetic data sets: A cautionary view. Mol. Ecol. 2017, 26, 1211–1224. [Google Scholar] [CrossRef] [PubMed]
- Jones, O.R.; Wang, J. COLONY: A program for parentage and sibship inference from multilocus genotype data. Mol. Ecol. Resour. 2010, 10, 551–555. [Google Scholar] [CrossRef] [PubMed]
- Meimans, P.G.; Van Tienderen, P.H.; Meirmans, P.G.; Van Tienderen, P.H. Genotype and Genodive: Two programs for the analysis of genetic diversity of asexual organisms. Mol. Ecol. Notes 2004, 4, 792–794. [Google Scholar] [CrossRef]
- Jombart, T. Adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef] [PubMed]
- R Core Development Team. R: A Language and Environment for Statistical Computing; R foundation for Statistical Computing: Vienna, Austria, 2015. [Google Scholar]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of Population Structure Using Multilocus Genotype Data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef]
- Falush, D.; Stephens, M.; Pritchard, J.K. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 2003, 164, 1567–1587. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Earl, D.A.; VonHoldt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Kopelman, N.M.; Mayzel, J.; Jakobsson, M.; Rosenberg, N.A.; Mayrose, I. CLUMPAK: A program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 2015, 15, 1179–1191. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. R Package Version 2.5-3. 2018. Available online: https://cran.r-project.org/web/packages/vegan/ (accessed on 1 November 2020).
- Elias, T.S. The genera of Mimosoideae (Leguminosae) in the Southeastern United States. J. Arnold Arbor. 1974, 1, 67–118. [Google Scholar]
- Broadhurst, L.; Coates, D.J. Genetic diversity within and divergence between rare and geographically widespread taxa of the Acacia acuminata Benth. (Mimosaceae) complex. Heredity 2002, 88, 250–257. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Butcher, P.A.; Moran, G.F.; Perkins, H.D. RFLP diversity in the nuclear genome of Acacia mangium. Heredity 1998, 81, 205–213. [Google Scholar] [CrossRef]
- Khasa, P.D.; Cheliak, W.M.; Bousquet, J. Mating system of Racosperma auriculiforme in a seed production area in Zaire. Can. J. Bot. 1993, 71, 779–785. [Google Scholar] [CrossRef]
- McDonald, M.W.; Butcher, P.A.; Bell, J.C.; Nguyen, C. V Clinal variation and genetic divergence in Acacia tumida (Mimosaceae). Aust. Syst. Bot. 2003, 16, 57–68. [Google Scholar] [CrossRef]
- Millar, M.A.; Byrne, M. Biogeographic origins and reproductive mode of naturalised populations of Acacia saligna. Aust. J. Bot. 2012, 60, 383–395. [Google Scholar] [CrossRef]
- Millar, M.A.; Coates, D.J.; Byrne, M. Genetic connectivity and diversity in inselberg populations of Acacia woodmaniorum, a rare endemic of the Yilgarn Craton banded iron formations. Heredity 2013, 111, 437. [Google Scholar] [CrossRef][Green Version]
- Playford, J.; Bell, J.C.; Moran, G.F. A Major Disjunction in Genetic Diversity over the Geographic Range of Acacia melanoxylon R. Br. Aust. J. Bot. 1993, 41, 355–368. [Google Scholar] [CrossRef]
- Searle, S.D.; Bell, J.C.; Moran, G.F. Genetic diversity in natural populations of Acacia mearnsii. Aust. J. Bot. 2000, 48, 279–286. [Google Scholar] [CrossRef]
- Kenrick, J.; Knox, R.B. Quantitative Analysis of Self-Incompatibility in Trees of Seven Species of Acacia. J. Hered. 1989, 80, 240–245. [Google Scholar] [CrossRef]
- Endress, P.K. Diversity and Evolutionary Biology of Tropical Flowers; Cambridge University Press: Cambridge, UK; New York, NY, USA, 1996; ISBN 9780521565103. [Google Scholar]
- Bumrungsri, S.; Harbit, A.; Benzie, C.; Carmouche, K.; Sridith, K.; Racey, P. The pollination ecology of two species of Parkia (Mimosaceae) in southern Thailand. J. Trop. Ecol. 2008, 24, 467–475. [Google Scholar] [CrossRef]
- Pometti, C.L.; Bessega, C.F.; Vilardi, J.C.; Saidman, B.O. Comparison of mating system parameters and genetic structure in three natural scenarios of Acacia visco (Leguminosae, Mimosoideae). Plant. Syst. Evol. 2013, 299, 761–771. [Google Scholar] [CrossRef]
- Vilas, A.; Pérez-Figueroa, A.; Quesada, H.; Caballero, A. Allelic diversity for neutral markers retains a higher adaptive potential for quantitative traits than expected heterozygosity. Mol. Ecol. 2015, 24, 4419–4432. [Google Scholar] [CrossRef] [PubMed]
- Harris, C.J.; Manea, A.; Moles, A.T.; Murray, B.R.; Leishman, M.R. Differences in life-cycle stage components between native and introduced ranges of five woody Fabaceae species. Austral. Ecol. 2017, 42, 404–413. [Google Scholar] [CrossRef]
- Colautti, R.I.; Barrett, S.C.H. Rapid adaptation to climate facilitates range expansion of an invasive plant. Science 2013, 342, 364–366. [Google Scholar] [CrossRef] [PubMed]
- Hallett, S.G. Dislocation from coevolved relationships: A unifying theory for plant invasion and naturalization? Weed Sci. 2006, 54, 282–290. [Google Scholar] [CrossRef]
- Commonwealth of Australia IBRA7 Interim Biogeographic Regionalisation for Australia 2012, 02. Available online: https://www.environment.gov.au/system/files/pages/5b3d2d31-2355-4b60-820c-e370572b2520/files/bioregions-new.pdf (accessed on 1 November 2020).
- Rajora, O.P.; Eckert, A.J.; Zinck, J.W.R. Single-locus versus multilocus patterns of local adaptation to climate in eastern white pine (Pinus strobus, Pinaceae). PLoS ONE 2016, 11, 1–26. [Google Scholar] [CrossRef]
- Nistelberger, H.; Gibson, N.; Macdonald, B.; Tapper, S.; Byrne, M. Phylogeographic evidence for two mesic refugia in a biodiversity hotspot. Heredity 2014, 113, 454–463. [Google Scholar] [CrossRef]
- Walters, S.J.; Robinson, T.P.; Byrne, M.; Wardell-Johnson, G.W.; Nevill, P. Contrasting patterns of local adaptation along climatic gradients between a sympatric parasitic and autotrophic tree species. Mol. Ecol. 2020, 1–16. [Google Scholar] [CrossRef]
- Sampson, J.; Tapper, S.; Coates, D.J.; Hankinson, M.; Mcarthur, S.; Byrne, M. Persistence with episodic range expansion from the early Pleistocene: The distribution of genetic variation in the forest tree Corymbia calophylla (Myrtaceae) in south-western Australia. Biol. J. Linn. Soc. 2018, 123, 545–560. [Google Scholar] [CrossRef]
- Broadhurst, L.; Breed, M.F.; Lowe, A.; Bragg, J.; Catullo, R.; Coates, D.J.; Encinas-viso, F.; Gellie, N.; James, E.; Krauss, S.; et al. Genetic diversity and structure of the Australian flora. Divers. Distrib. 2017, 23, 41–52. [Google Scholar] [CrossRef]
- Ahrens, C.W.; Byrne, M.; Rymer, P.D. Standing genomic variation within coding and regulatory regions contributes to the adaptive capacity to climate in a foundation tree species. Mol. Ecol. 2019, 28, 2502–2516. [Google Scholar] [CrossRef] [PubMed]
- Raymond, O.L.; Gallagher, R.; Shaw, R.; Yeates, A.N.; Doutch, H.F.; Palfreyman, W.D.; Blake, D.H.; Highet, L. Surface Geology of Australia 1:2.5 Million Scale Dataset, 2012 ed.; Commonwealth of Australia (Geoscience Australia): Canberra, Australia, 2012. [Google Scholar] [CrossRef]
- Atlas Of Living Australia ALA Paraserianthes Occurrence Records. Available online: https://doi.ala.org.au/doi/10.26197/5d8c4e6842581 (accessed on 26 September 2019).
- Stojanova, B.; Šurinová, M.; Zeisek, V.; Münzbergová, Z.; Pánková, H. Low genetic differentiation despite high fragmentation in the endemic serpentinophyte Minuartia smejkalii (M. verna agg., Caryophyllaceae) revealed by RADSeq SNP markers. Conserv. Genet. 2020, 21, 187–198. [Google Scholar] [CrossRef]
- Cornuet, J.M.; Luikart, G. Description and Power Analysis of Two Tests for Detecting Recent Population Bottlenecks From Allele Frequency Data. Genetics 1996, 144, 2001–2014. [Google Scholar] [PubMed]
- Cumming, D.A.; Garratt, D.; McCarthy, M.; Wolfe, A. Port Related Structures on the Coast of Western Australia; No.98; Department of Maritime Archaeology, Western Australian Maritime Museum: Fremantle, Australia, 1995. [Google Scholar]
- Cambage, R.H. Acacia seedlings Part I. J. R. Soc. New South Wales 1915, 49, 81–121. [Google Scholar]
- Godfrey, J.S.; Ridgway, K.R. The Large-Scale Environment of the Poleward-Flowing Leeuwin Current, Western Australia: Longshore Steric Height Gradients, Wind Stresses and Geostrophic Flow. J. Phys. Oceanogr. 1985, 15, 481–495. [Google Scholar] [CrossRef]
- Sgrò, C.M.; Lowe, A.J.; Hoffmann, A.A. Building evolutionary resilience for conserving biodiversity under climate change. Evol. Appl. 2011, 4, 326–337. [Google Scholar] [CrossRef]
- Rossetto, M.; Wilson, P.D.; Bragg, J.; Cohen, J.; Fahey, M.; Yap, J.S.; van der Merwe, M. Perceptions of Similarity Can Mislead Provenancing Strategies—An Example from Five Co-Distributed Acacia Species. Diversity 2020, 12, 306. [Google Scholar] [CrossRef]
- Brouwers, N.; Matusick, G.; Ruthrof, K.; Lyons, T.; Hardy, G. Landscape-scale assessment of tree crown dieback following extreme drought and heat in a Mediterranean eucalypt forest ecosystem. Landsc. Ecol. 2013, 28, 69–80. [Google Scholar] [CrossRef]
| All Individuals | No Siblings in Populations | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Popn | Location | N | Ne | Ho | He | FIS | N | Ne | Ho | He | FIS |
| Native | |||||||||||
| BD | Boranup Drive | 20 | 2.305 | 0.389 | 0.469 | 0.178 | 18 | 2.308 | 0.382 | 0.476 | 0.192 |
| BH | Boat Harbour | 20 | 1.457 | 0.211 | 0.28 | 0.245 | 3 | 1.441 | 0.241 | 0.333 | 0.195 |
| CA | Cape Arid | 1 | - | 0.444 | - | - | 1 | - | 0.444 | - | - |
| DR | Donnelly River | 18 | 1.148 | 0 | 0.106 | 1 | 18 | 1.148 | 0 | 0.106 | 1 |
| GR | Graphit Rd | 2 | 1.267 | 0.056 | 0.25 | 0.5 | 2 | 1.267 | 0.056 | 0.25 | 0.5 |
| GS | Gingilup Swamp | 20 | 1.31 | 0.081 | 0.176 | 0.35 | 1 | - | 0.222 | - | - |
| MA | Manjimup | 1 | - | 0.22 | - | - | 1 | - | 0.222 | - | - |
| MR | Morangup Reserve | 21 | 1.167 | 0.073 | 0.078 | 0.023 | 21 | 1.167 | 0.073 | 0.078 | 0.023 |
| PO | Porongorups | 2 | 1.667 | 0 | 0.667 | 1 | 2 | 1.667 | 0 | 0.667 | 1 |
| RC | Recherche Islands | 5 | 3.277 | 0.444 | 0.658 | 0.35 | 5 | 3.277 | 0.444 | 0.658 | 0.35 |
| SR | Serpentine River National Park | 19 | 2.098 | 0.195 | 0.341 | 0.429 | 19 | 2.098 | 0.195 | 0.341 | 0.429 |
| VT | Van Tripp Rd | 20 | 1.303 | 0.119 | 0.196 | 0.376 | 1 | - | 0.111 | - | - |
| WD | Wellington Dam | 19 | 1.302 | 0.093 | 0.187 | 0.313 | 19 | 1.302 | 0.093 | 0.187 | 0.313 |
| YG | Yallingup | 21 | 1.293 | 0.071 | 0.191 | 0.745 | 2 | 1.881 | 0.167 | 0.722 | 0.714 |
| Naturalized | |||||||||||
| CI | Craggy Island | 2 | 1.125 | 0.125 | 0.063 | −1 | 1 | - | 0.125 | - | - |
| DT | Devil’s Tower | 1 | - | 0.25 | - | - | 1 | - | 0.25 | - | - |
| EP | Eyre Peninsula | 17 | 1.097 | 0.059 | 0.074 | 0.43 | 17 | 1.097 | 0.059 | 0.074 | 0.43 |
| KI | Kangaroo Island | 24 | 1.243 | 0.218 | 0.137 | −0.245 | 24 | 1.243 | 0.218 | 0.137 | −0.245 |
| LE | Lakes Entrance | 8 | 1.352 | 0.278 | 0.173 | −0.591 | 8 | 1.352 | 0.278 | 0.173 | −0.591 |
| PI | Phillip Island | 7 | 1.521 | 0.238 | 0.294 | 0.068 | 7 | 1.521 | 0.238 | 0.294 | 0.068 |
| PR | Pudney’s Rd | 4 | 1.258 | 0.188 | 0.141 | −0.3 | 1 | - | 0.143 | - | - |
| RI | Rodondo Island | 1 | - | 0.125 | - | - | 1 | - | 0.125 | - | - |
| WG | Waterfall Gully | 16 | 1.705 | 0.22 | 0.353 | 0.3 | 6 | 1.665 | 0.259 | 0.439 | 0.278 |
| WR | Wye River | 20 | 1.479 | 0.157 | 0.271 | 0.498 | 20 | 1.479 | 0.157 | 0.271 | 0.498 |
| Source | df | Sums of Squares | Mean Sum of Squares | Estimated Variation | % Variation | p Value |
|---|---|---|---|---|---|---|
| all samples (n = 289) | ||||||
| Among Regions | 1 | 425.66 | 425.66 | 2.897 | 27.7 | 0.001 |
| Among Pops | 22 | 1262.504 | 57.387 | 4.578 | 43.7 | 0.001 |
| Within Pops | 265 | 792.864 | 2.992 | 2.992 | 28.6 | |
| reduced dataset (n = 199) | ||||||
| Among Regions | 1 | 355.749 | 355.749 | 3.089 | 31.3 | 0.001 |
| Among Pops | 22 | 697.671 | 31.712 | 1.921 | 37.9 | 0.001 |
| Within Pops | 175 | 532.444 | 3.043 | 2.863 | 30.8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brown, G.K.; James, E.A.; Simmons, C.L.; Ahrens, C.W. Recently Naturalized Paraserianthes lophantha subsp. lophantha Displays Contrasting Genetic Diversity and Climate Relationships Compared to Native Populations. Diversity 2020, 12, 422. https://doi.org/10.3390/d12110422
Brown GK, James EA, Simmons CL, Ahrens CW. Recently Naturalized Paraserianthes lophantha subsp. lophantha Displays Contrasting Genetic Diversity and Climate Relationships Compared to Native Populations. Diversity. 2020; 12(11):422. https://doi.org/10.3390/d12110422
Chicago/Turabian StyleBrown, Gillian K., Elizabeth A. James, Catherine L. Simmons, and Collin W. Ahrens. 2020. "Recently Naturalized Paraserianthes lophantha subsp. lophantha Displays Contrasting Genetic Diversity and Climate Relationships Compared to Native Populations" Diversity 12, no. 11: 422. https://doi.org/10.3390/d12110422
APA StyleBrown, G. K., James, E. A., Simmons, C. L., & Ahrens, C. W. (2020). Recently Naturalized Paraserianthes lophantha subsp. lophantha Displays Contrasting Genetic Diversity and Climate Relationships Compared to Native Populations. Diversity, 12(11), 422. https://doi.org/10.3390/d12110422





