Community Ecology and Phylogeography of Bats in the Guianan Savannas of Northern South America
Abstract
1. Introduction
2. Materials and Methods
3. Results
3.1. Biodiversity Measures
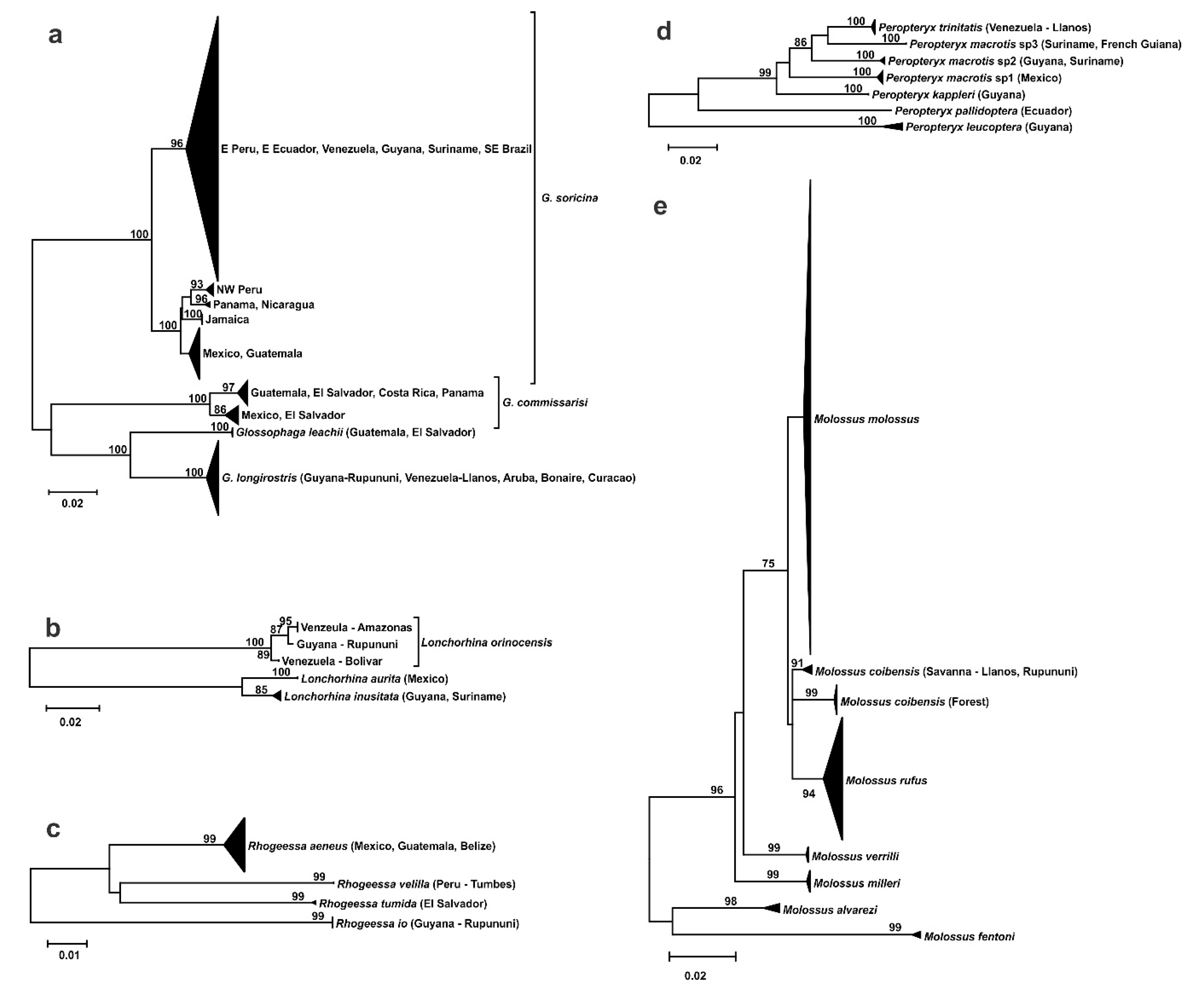
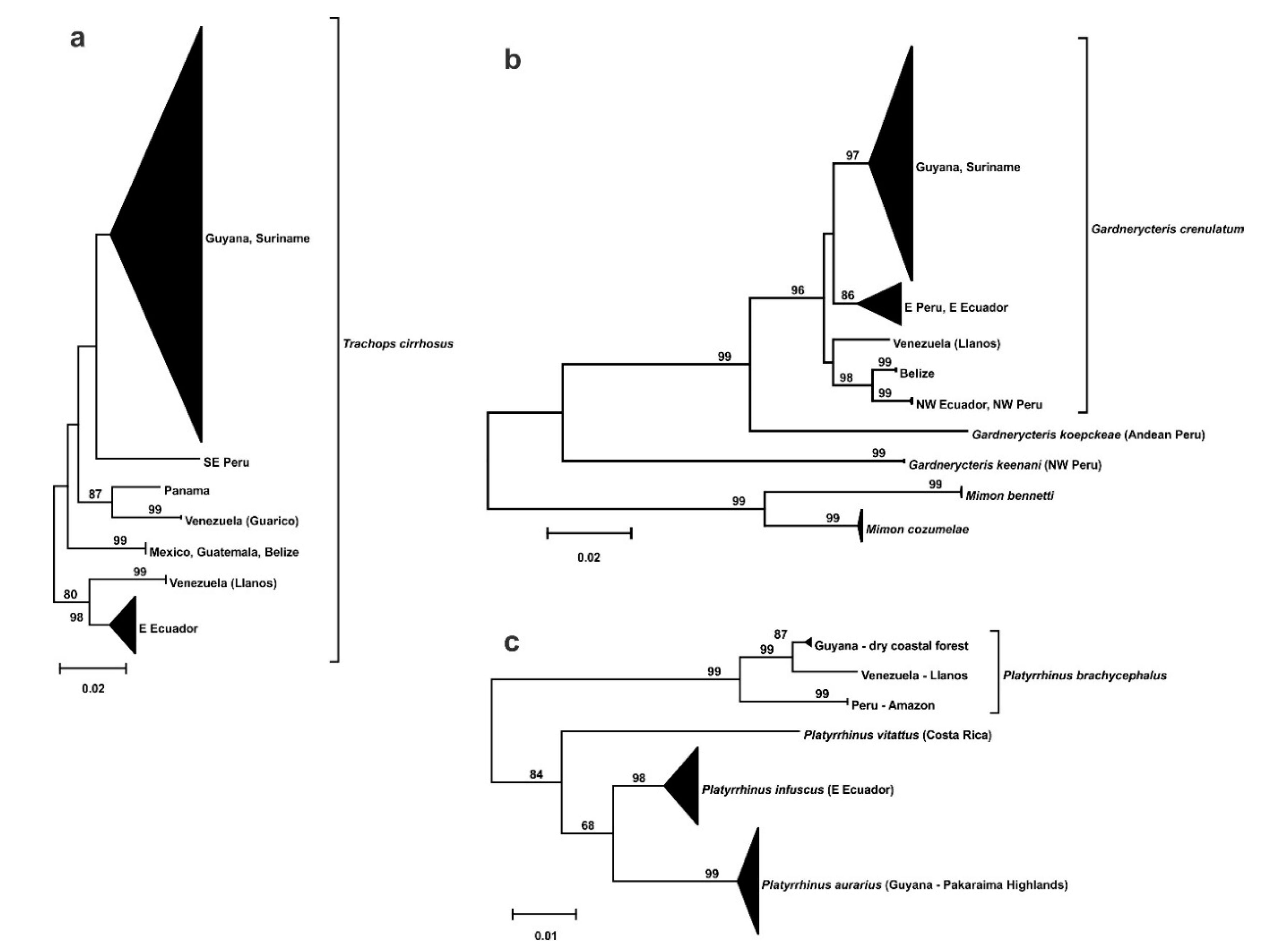
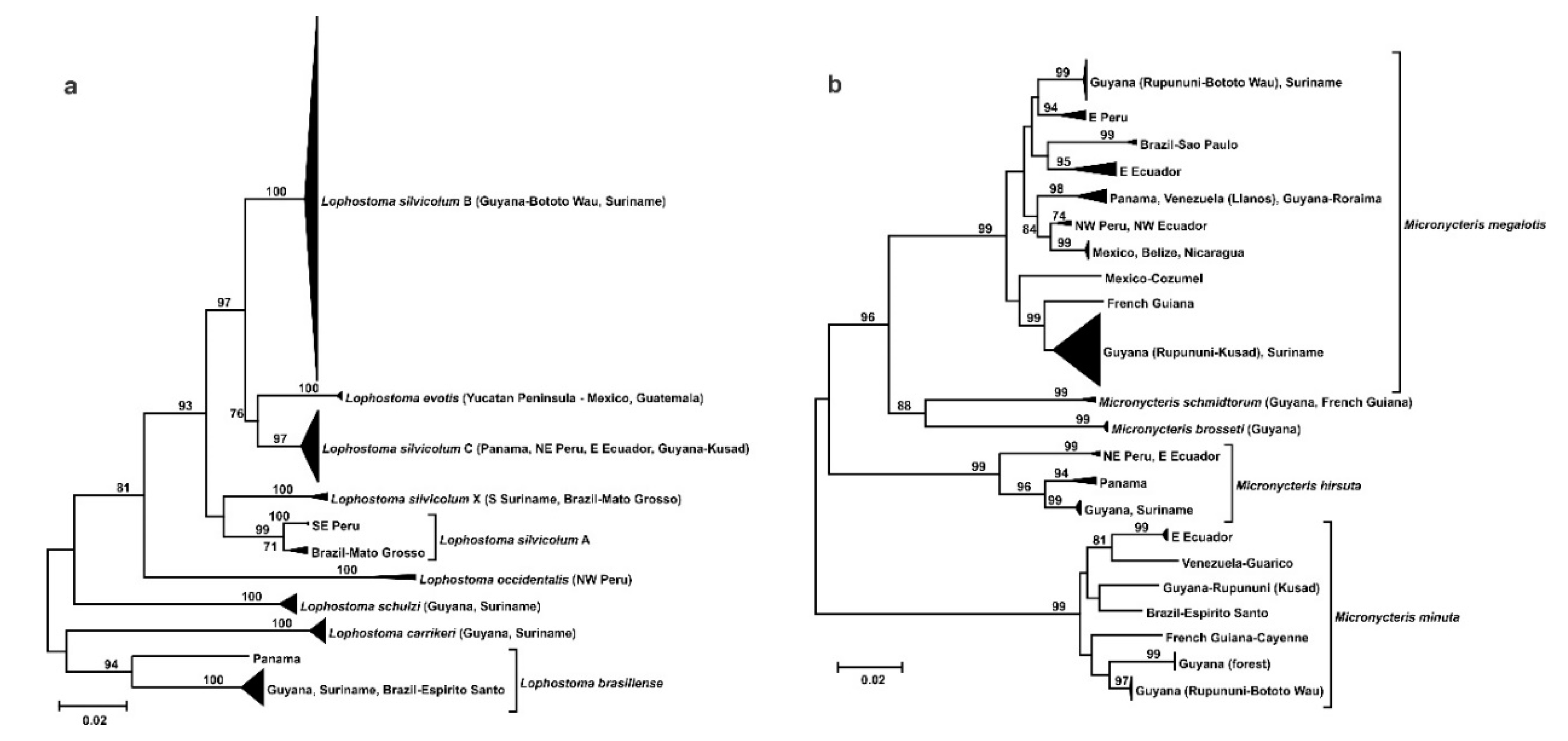
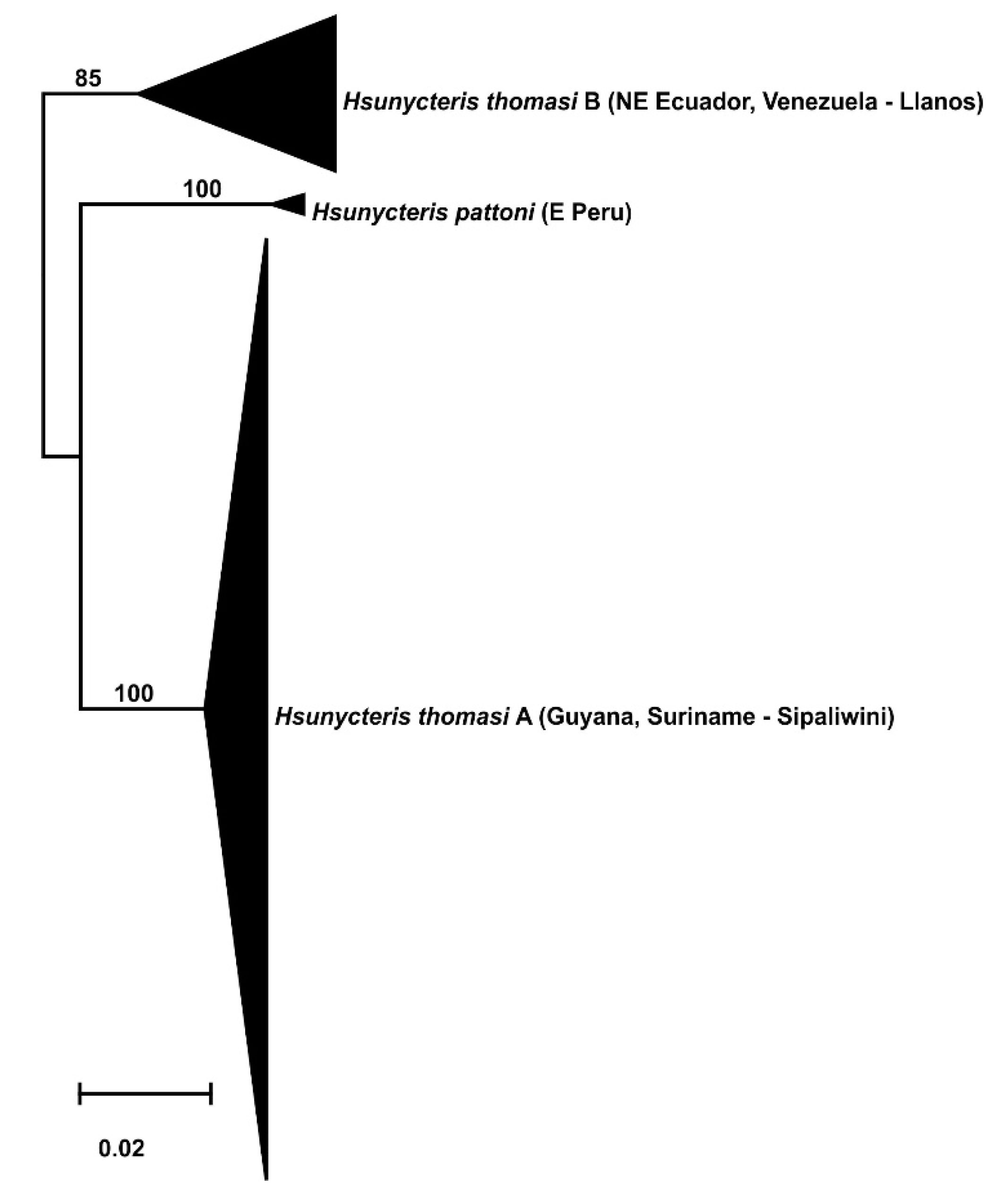
3.2. Phylogeography
4. Discussion
5. Conclusions
- Endemism in the savannas of the Guianas at the species or subspecies level.
- Divergent population from the Llanos of Venezuela.
- Split within the southern Rupununi of Guyana between Kusad and Bototo Wau.
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Species | Llanos | Rupununi | Sipaliwini | Total |
|---|---|---|---|---|
| Carollia perspicillata | 60 | 239 | 201 | 500 |
| Artibeus planirostris | 7 | 103 | 77 | 187 |
| Artibeus cinereus | 0 | 94 | 9 | 103 |
| Glossophaga soricina | 6 | 51 | 22 | 79 |
| Artibeus lituratus | 0 | 30 | 46 | 76 |
| Glossophaga longirostris | 36 | 34 | 0 | 70 |
| Trachops cirrhosus | 13 | 32 | 1 | 46 |
| Peropteryx trinitatis | 46 | 0 | 0 | 46 |
| Desmodus rotundus | 5 | 36 | 2 | 43 |
| Sturnira lilium | 3 | 31 | 8 | 42 |
| Artibeus obscurus | 5 | 7 | 20 | 32 |
| Phyllostomus discolor | 2 | 19 | 9 | 30 |
| Ametrida centurio | 1 | 10 | 18 | 29 |
| Artibeus bogotensis | 2 | 17 | 8 | 27 |
| Phyllostomus elongatus | 3 | 17 | 6 | 26 |
| Pteronotus fuscus | 26 | 0 | 0 | 26 |
| Platyrrhinus lineatus | 0 | 0 | 22 | 22 |
| Lonchorhina orinocensis | 20 | 1 | 0 | 21 |
| Molossus molossus | 0 | 12 | 8 | 20 |
| Noctilio albiventris | 0 | 20 | 0 | 20 |
| Rhinophylla pumilio | 0 | 3 | 15 | 18 |
| Saccopteryx bilineata | 1 | 14 | 1 | 16 |
| Artibeus concolor | 1 | 8 | 7 | 16 |
| Lophostoma silvicolum | 0 | 12 | 4 | 16 |
| Phyllostomus hastatus | 10 | 6 | 0 | 16 |
| Myotis nigricans | 0 | 14 | 0 | 14 |
| Pteronotus rubiginosus | 0 | 13 | 1 | 14 |
| Myotis albescens | 0 | 10 | 3 | 13 |
| Rhynchonycteris naso | 2 | 10 | 0 | 12 |
| Gardnerycteris crenulatum | 2 | 7 | 2 | 11 |
| Micronycteris megalotis | 5 | 5 | 0 | 10 |
| Hsunycteris thomasi | 2 | 0 | 8 | 10 |
| Phylloderma stenops | 1 | 3 | 5 | 9 |
| Rhogeessa io | 1 | 8 | 0 | 9 |
| Chiroderma villosum | 1 | 2 | 5 | 8 |
| Carollia brevicauda | 0 | 5 | 2 | 7 |
| Molossus coibensis | 6 | 1 | 0 | 7 |
| Mesophylla macconnelli | 0 | 5 | 2 | 7 |
| Artibeus gnomus | 1 | 1 | 5 | 7 |
| Micronycteris minuta | 1 | 5 | 0 | 6 |
| Eumops auripendulus | 0 | 6 | 0 | 6 |
| Eptesicus furinalis | 0 | 3 | 2 | 5 |
| Trinycteris nicefori | 1 | 3 | 1 | 5 |
| Vampyriscus bidens | 1 | 0 | 4 | 5 |
| Uroderma bilobatum | 0 | 4 | 0 | 4 |
| Platyrrhinus guianensis | 0 | 2 | 2 | 4 |
| Platyrrhinus incarum | 0 | 1 | 3 | 4 |
| Vampyriscus brocki | 0 | 0 | 4 | 4 |
| Pteronotus alitonos | 0 | 0 | 4 | 4 |
| Sturnira tildae | 0 | 0 | 4 | 4 |
| Saccopteryx canescens | 2 | 1 | 0 | 3 |
| Saccopteryx leptura | 0 | 2 | 1 | 3 |
| Lophostoma brasiliense | 0 | 2 | 1 | 3 |
| Choeroniscus minor | 0 | 2 | 1 | 3 |
| Chrotopterus auritus | 0 | 0 | 3 | 3 |
| Chiroderma trinitatum | 0 | 0 | 3 | 3 |
| Anoura geoffroyi | 1 | 0 | 2 | 3 |
| Pteronotus personatus | 0 | 2 | 0 | 2 |
| Cormura brevirostris | 0 | 1 | 1 | 2 |
| Lionycteris spurrelli | 0 | 0 | 2 | 2 |
| Myotis riparius | 0 | 0 | 2 | 2 |
| Platyrrhinus fusciventris | 0 | 0 | 2 | 2 |
| Artibeus amplus | 2 | 0 | 0 | 2 |
| Platyrrhinus brachycephalus | 2 | 0 | 0 | 2 |
| Cynomops planirostris | 0 | 1 | 0 | 1 |
| Lasiurus blossevillii | 0 | 1 | 0 | 1 |
| Noctilio leporinus | 0 | 1 | 0 | 1 |
| Micronycteris schmidtorum | 0 | 1 | 0 | 1 |
| Natalus tumidirostris | 0 | 1 | 0 | 1 |
| Glyphonycteris sylvestris | 0 | 1 | 0 | 1 |
| Uroderma magnirostrum | 0 | 1 | 0 | 1 |
| Molossus fentoni | 0 | 1 | 0 | 1 |
| Lophostoma carrikeri | 0 | 0 | 1 | 1 |
| Diaemus youngi | 0 | 0 | 1 | 1 |
| Glyphonycteris daviesi | 0 | 0 | 1 | 1 |
| Pteronotus gymnonotus | 1 | 0 | 0 | 1 |
| Total | 279 | 922 | 562 | 1763 |
| First Sample | Second Sample | Shared Species | Jaccard Classic | Sorensen Classic | Chao-Jaccard-Raw | Chao-Jaccard-Est | Chao-Sorensen-Raw | Chao-Sorensen-Est | Morisita-Horn | Bray-Curtis |
|---|---|---|---|---|---|---|---|---|---|---|
| Rupununi | Sipaliwini | 34 | 0.479 | 0.648 | 0.779 | 0.824 | 0.876 | 0.903 | 0.886 | 0.592 |
| Rupununi | Llanos | 27 | 0.422 | 0.593 | 0.561 | 0.729 | 0.719 | 0.843 | 0.642 | 0.278 |
| Sipaliwini | Llanos | 21 | 0.333 | 0.5 | 0.369 | 0.415 | 0.539 | 0.586 | 0.608 | 0.247 |
References
- Mittermeier, R.A.; Myers, N.; Thomsen, J.B.; da Fonseca, G.A.B.; Olivieri, S. Biodiversity hotspots and major tropical wilderness areas: Approaches to setting conservation priorities. Conserv. Biol. 1998, 12, 516–520. [Google Scholar] [CrossRef]
- Aguirre, L.F. Structure of a Neotropical savanna bat community. J. Mammal. 2002, 83, 775–784. [Google Scholar] [CrossRef]
- Zortéa, M.; Alho, C.J.R. Bat diversity of a Cerrado habitat in central Brazil. Biodivers. Conserv. 2008, 17, 791–805. [Google Scholar] [CrossRef]
- Morales-Martínez, D.M.; Rodríguez-Posada, M.E.; Fernández-Rodríguez, C.; Calderón-Capote, M.C.; Gutiérrez-Sanabria, D.R. Spatial variation of bat diversity between three floodplain-savanna ecosystems of the Colombian Llanos. Therya 2018, 9, 41–52. [Google Scholar] [CrossRef]
- Sarmiento, G.; Monasterio, M. Studies on the savanna vegetation of the Venezuelan Llanos: I. The use of association analysis. J. Ecol. 1969, 57, 579–598. [Google Scholar] [CrossRef]
- Jansen-Jacobs, M.; ter Steege, H. Southwest Guyana: A complex mosaic of savannahs and forest. In Plant Diversity in Guyana: With Recommendations for a Protected Areas Strategy; Ter Steege, H., Ed.; Tropenbos Series; Tropenbos International: Wageningen, The Netherlands, 2000; Volume 18, pp. 147–157. [Google Scholar]
- Van Donselaar, J. Observations on savanna vegetation-types in the Guianas. Plant Ecol. 1969, 17, 271–312. [Google Scholar] [CrossRef]
- Lee, T.E., Jr.; Lim, B.K.; Hanson, J.D. Noteworthy records of mammals from the Orinoco River drainage of Venezuela. Tex. J. Sci. 2000, 52, 264–266. [Google Scholar]
- Lim, B.K.; Osborne, C.; Ignace, A. Small mammals of the South Rupununi region, Guyana. In Biodiversity Assessment Survey of the South Rupununi Savannah, Guyana; Alonso, L.E., Persaud, J., Williams, A., Eds.; BAT Survey Report No. 1; WWF-Guianas: Georgetown, Guyana, 2016; pp. 103–118. [Google Scholar]
- Sikes, R.S.; Animal Care and Use Committee of the American Society of Mammalogists. 2016 guidelines of the American Society of Mammalogists for the use of wild mammals in research and education. J. Mammal. 2016, 97, 663–688. [Google Scholar] [CrossRef]
- Colwell, R.K. EstimateS: Statistical Estimation of Species Richness and Shared Species from Samples, Version 9. 2013. Available online: http://purl.oclc.org/estimates (accessed on 1 December 2017).
- Clare, E.L.; Lim, B.K.; Engstrom, M.D.; Eger, J.L.; Hebert, P.D.N. DNA barcoding of Neotropical bats: Species identification and discovery within Guyana. Mol. Ecol. Notes 2007, 7, 184–190. [Google Scholar] [CrossRef]
- Lim, B.K. Review of genetic diversification of bats in the Caribbean and biogeographic relationships to Neotropical species based on DNA barcodes. Genome 2017, 60, 65–73. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D.N. BOLD: The Barcode of Life Data System (www.barcodinglife.org). Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef] [PubMed]
- LaVal, R.K.; Fitch, H.S. Structure, movements and reproduction in three Costa Rican bat communities. Occas. Pap. 1977, 69, 1–28. [Google Scholar]
- Lim, B.K.; Engstrom, M.D. Bat community structure at Iwokrama Forest, Guyana. J. Trop. Ecol. 2001, 17, 647–665. [Google Scholar] [CrossRef]
- Wilson, D.E. Mammals of La Selva, Costa Rica. In Four Neotropical Rainforests; Gentry, A.H., Ed.; Yale University Press: New Haven, CT, USA, 1990; pp. 273–286. ISBN 9780300054484. [Google Scholar]
- Lim, B.K.; Engstrom, M.D. Mammals of Iwokrama Forest. Proc. Acad. Nat. Sci. Phila. 2005, 154, 71–108. [Google Scholar] [CrossRef]
- Rex, K.; Kelm, D.H.; Wiesner, K.; Kunz, T.H.; Voigt, C.C. Species richness and structure of three Neotropical bat assemblages. Biol. J. Linn. Soc. 2008, 94, 617–629. [Google Scholar] [CrossRef]
- Lim, B.K. Environmental assessment at the Bakhuis bauxite concession: Small-sized mammal diversity and abundance in the lowland humid forests of Suriname. Open Biol. J. 2009, 2, 42–53. [Google Scholar] [CrossRef]
- Wilson, D.E.; Ascorra, C.F.; Solari, S. Bats as indicators of habitat disturbance. In Manu: The Biodiversity of Southeastern Peru; Wilson, D.E., Sandoval, A., Eds.; Smithsonian Institution and Editorial Horizonte: Washington, DC, USA; Lima, Peru, 1996; pp. 613–625. ISBN 1560987103. [Google Scholar]
- Arteaga, L.L.; Aguirre, L.F.; Moya, M.I. Seed rain produced by bats and birds in forest islands in a Neotropical savanna. Biotropica 2006, 38, 718–724. [Google Scholar] [CrossRef]
- Rivero-Blanco, C.; Dixon, J.R. Origin and distribution of the herpetofauna of the dry lowland regions of northern South America. In The South America Herpetofauna: Its Origin, Evolution, and Dispersal; Duellman, W.E., Ed.; Monograph of the Museum of Kansas: Lawrence, Kansas, 1979; Volume 7, pp. 281–298. ISBN 0-89338-008-3. [Google Scholar]
- Linares, O.J.; Ojasti, J. Una nueva especie de murcielago del genero Lonchorhina (Chiroptera: Phyllstomidae) del sur de Venezuela. Serie Zoológica 1971, 36, 1–8. [Google Scholar]
- Hernández-Camacho, J.; Cadena-G., A. Notas para la revision del genero Lonchorhina (Chiroptera, Phyllstomidae). Caldasia 1978, 12, 199–251. [Google Scholar]
- Lim, B.K.; Engstrom, M.D.; Bickham, J.W.; Patton, J.C. Molecular phylogeny of New World emballonurid bats (Tribe Diclidurini) based on loci from the four genetic transmission systems in mammals. Biol. J. Linn. Soc. 2008, 93, 189–209. [Google Scholar] [CrossRef]
- Handley, C.O., Jr. Mammals of the Smithsonian Venezuelan Project. Brigh. Young Univ. Sci. Bull. Biol. Ser. 1976, 20, 1–89. [Google Scholar] [CrossRef]
- Webster, W.D.; Handley, C.O., Jr. Systematics of Miller’s Long-Tongued Bat, Glossophaga Longirostris, with Description of Two New Subspecies; Occasional Papers, No. 100; The Museum, Texas Tech University: Lubbock, TX, USA, 1986; pp. 1–22. [Google Scholar]
- Griffiths, T.A.; Gardner, A.L. Subfamily Glossophaginae Bonaparte, 1845. In Mammals of South America: Volume 1: Marsupials, Xenarthrans, Shrews, and Bats; Gardner, A.L., Ed.; The University of Chicago Press: Chicago, IL, USA; London, UK, 2008; pp. 224–255. ISBN 978-0-226-28240-4. [Google Scholar]
- Loureiro, L.O.; Lim, B.K.; Engstrom, M.D. A new species of mastiff bat (Chiroptera, Molossidae, Molossus) from Guyana and Ecuador. Mammal. Biol. 2018, 90, 10–21. [Google Scholar] [CrossRef]
- Bickham, J.W.; Ruedas, L.A. Genus Rhogeessa H. Allen, 1866. In Mammals of South America: Volume 1: Marsupials, Xenarthrans, Shrews, and Bats; Gardner, A.L., Ed.; The University of Chicago Press: Chicago, IL, USA; London, UK, 2008; pp. 481–484. ISBN 978-0-226-28240-4. [Google Scholar]
- Baird, A.B.; Hillis, D.M.; Patton, J.C.; Bickham, J.W. Evolutionary history of the genus Rhogeessa (Chiroptera: Vespertilionidae) as revealed by mitochondrial DNA sequences. J. Mammal. 2008, 89, 744–754. [Google Scholar] [CrossRef]
- Clare, E.L. Cryptic species: Patterns of maternal and paternal gene flow in eight Neotropical bats. PLoS ONE 2011, 6, e21460. [Google Scholar] [CrossRef] [PubMed]
- Velazco, P.M.; Cadenillas, R. On the identify of Lophostoma silvicolum occidentalis (Davis & Carter, 1978) (Chiroptera: Phyllostomidae). Zootaxa 2011, 2962, 1–20, ISSN 1175-5326. [Google Scholar]
- Velazco, P.M.; Soto-Centeno, J.A.; Fleck, D.W.; Voss, R.S.; Simmons, N.B. A new species of nectar-feeding bat of the genus Hsunycteris (Phyllostomidae: Lonchophyllinae) from northeastern Peru. Am. Mus. Novit. 2017, 3881, 1–26, ISSN 0003-0082. [Google Scholar] [CrossRef]
- Parlos, J.A.; Timm, R.M.; Swier, V.J.; Zeballos, H.; Baker, R.J. Evaluation of Paraphyletic Assemblages within Lonchophyllinae, with Description of a New Tribe and Genus; Occasional Papers; Museum of Texas Tech University: Lubbock, TX, USA, 2014; Volume 320, pp. 1–23, ISSN 0149-175X. [Google Scholar]
- Lim, B.K. Historical biogeography of New World emballonurid bats (Tribe Diclidurini): Taxon pulse diversification. J. Biogeogr. 2008, 35, 1385–1401. [Google Scholar] [CrossRef]
- Lim, B.K. Divergence times and origin of Neotropical sheath-tailed bats (Tribe Diclidurini) in South America. Mol. Phylogenet. Evol. 2007, 45, 777–791. [Google Scholar] [CrossRef]
- Wyss, A.R.; Flynn, J.J.; Norell, M.A.; Swisher, C.C.; Charrier, R.; Novacek, M.J.; McKenna, M.C. South America’s earliest rodent and recognition of a new interval of mammalian evolution. Nature 1993, 365, 434–437. [Google Scholar] [CrossRef]
- Lim, B.K.; Engstrom, M.D.; Patton, J.C.; Bickham, J.W. Systematic review of small fruit-eating bats (Artibeus) from the Guianas, and a re-evaluation of A. glaucus bogotensis. Acta Chiropterol. 2008, 10, 243–256. [Google Scholar] [CrossRef]
- Marques-Aguiar, S.A. Genus Artibeus Leach, 1821. In Mammals of South America: Volume 1: Marsupials, Xenarthrans, Shrews, and Bats; Gardner, A.L., Ed.; The University of Chicago Press: Chicago, IL, USA; London, UK, 2008; pp. 301–321. ISBN 978-0-226-28240-4. [Google Scholar]
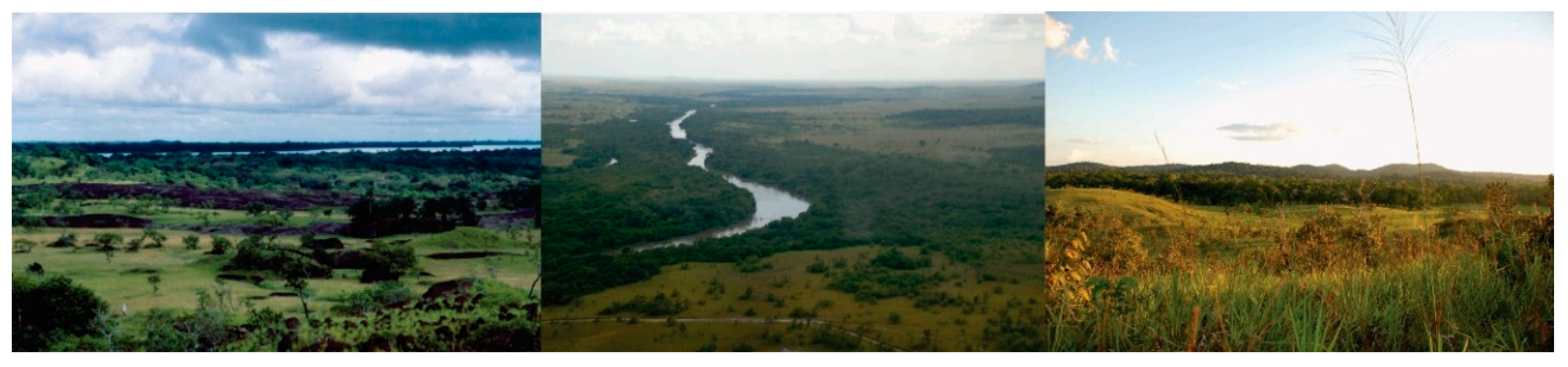
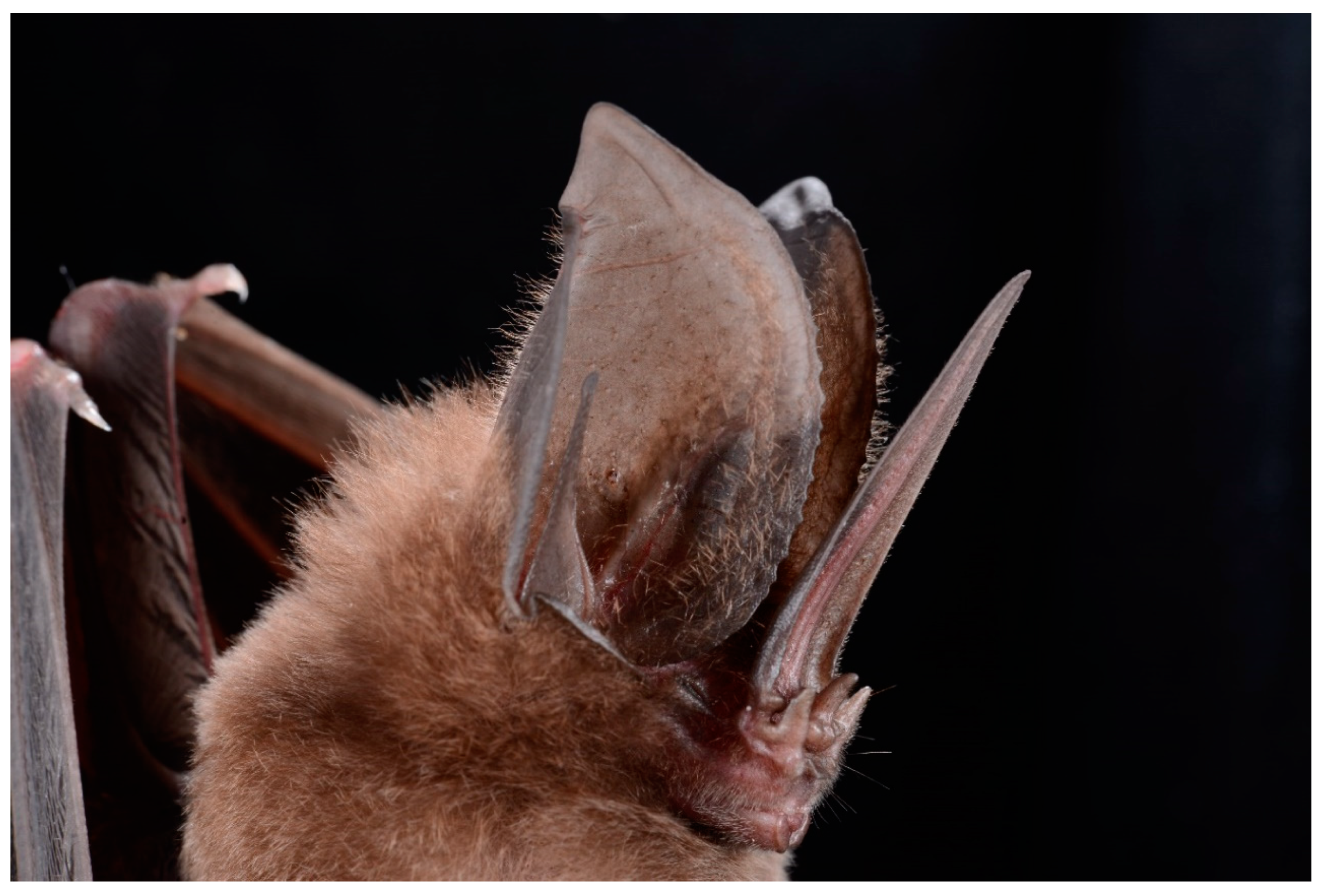

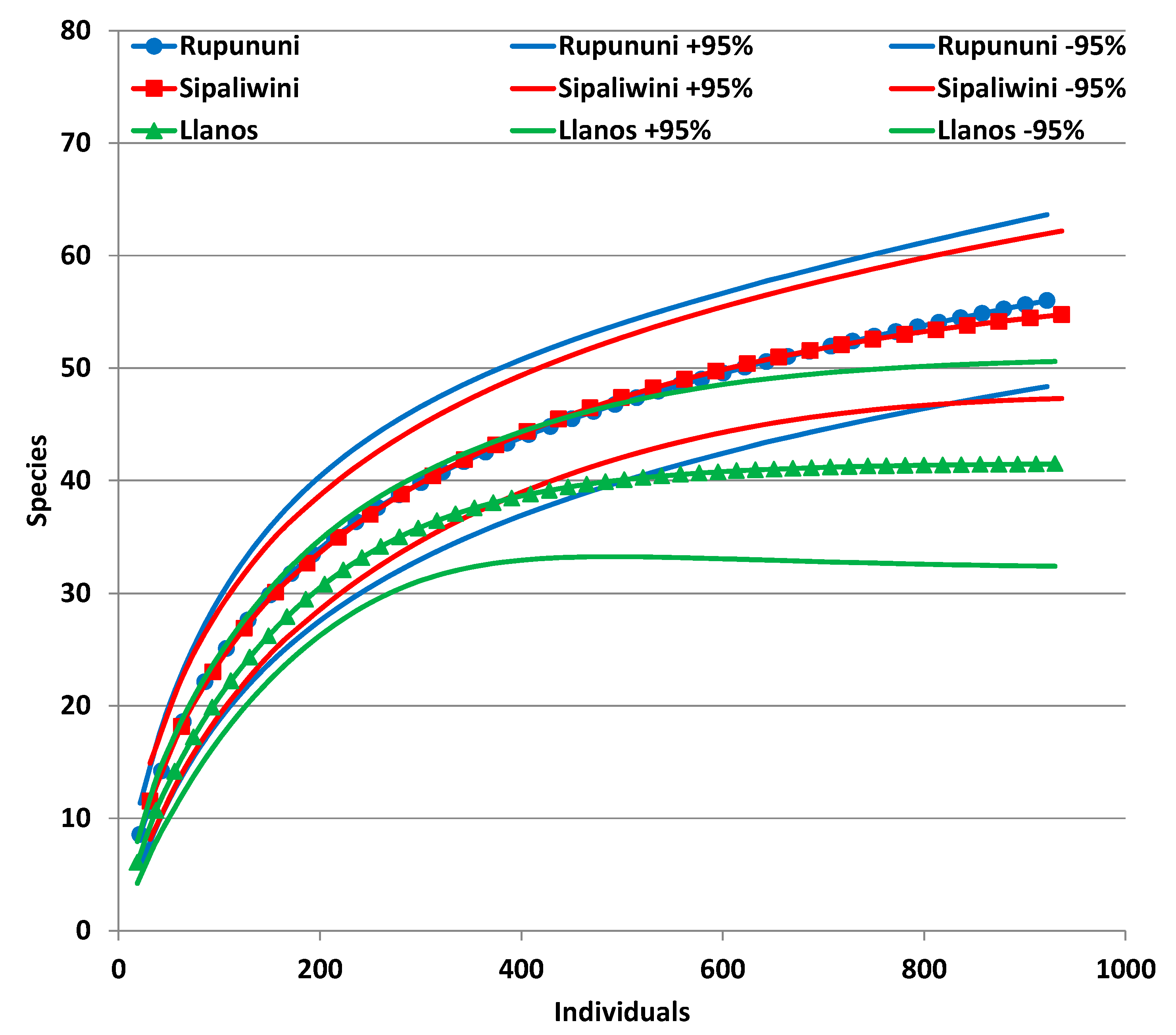

| Variable | Llanos | Rupununi | Sipaliwini | Total |
|---|---|---|---|---|
| Observed data: | ||||
| Individuals | 279 | 922 | 562 | 1763 |
| Species | 35 | 56 | 49 | 76 |
| Trap-nights | 15 | 43 | 18 | 76 |
| Richness estimators: | ||||
| ACE | 48.75 | 69.14 | 56.77 | 83.31 |
| ICE | 51.04 | 69.3 | 61.4 | 86.52 |
| Chao 1 | 42.31 | 68.99 | 53.99 | 84.25 |
| Chao 2 | 40.6 | 72.74 | 56.81 | 92.92 |
| Jack 1 | 47.13 | 71.63 | 62.22 | 91.79 |
| Jack 2 | 48.74 | 81.3 | 66.3 | 101.6 |
| Bootstrap | 41.17 | 62.81 | 55.5 | 83.16 |
| Average richness | 45.68 | 70.84 | 59 | 89.7 |
| Diversity indices: | ||||
| Alpha | 10.57 | 13.13 | 12.91 | 16.17 |
| Shannon | 2.66 | 2.95 | 2.64 | 3.11 |
| Shannon Exponential | 14.31 | 19.16 | 14.02 | 22.52 |
| Simpson | 9.04 | 9.72 | 6.18 | 9.42 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lim, B.K.; Lee, T.E., Jr. Community Ecology and Phylogeography of Bats in the Guianan Savannas of Northern South America. Diversity 2018, 10, 129. https://doi.org/10.3390/d10040129
Lim BK, Lee TE Jr. Community Ecology and Phylogeography of Bats in the Guianan Savannas of Northern South America. Diversity. 2018; 10(4):129. https://doi.org/10.3390/d10040129
Chicago/Turabian StyleLim, Burton K., and Thomas E. Lee, Jr. 2018. "Community Ecology and Phylogeography of Bats in the Guianan Savannas of Northern South America" Diversity 10, no. 4: 129. https://doi.org/10.3390/d10040129
APA StyleLim, B. K., & Lee, T. E., Jr. (2018). Community Ecology and Phylogeography of Bats in the Guianan Savannas of Northern South America. Diversity, 10(4), 129. https://doi.org/10.3390/d10040129







