The Biogeography of Great Salt Lake Halophilic Archaea: Testing the Hypothesis of Avian Mechanical Carriers
Abstract
1. Introduction
2. Materials and Methods
2.1. Collection, Cultures, and DNA Extraction
2.2. PCR Amplification, DNA Sequencing and Genbank Comparisons
2.3. Geographic and Climate Data
2.4. Statistical Methods
3. Results
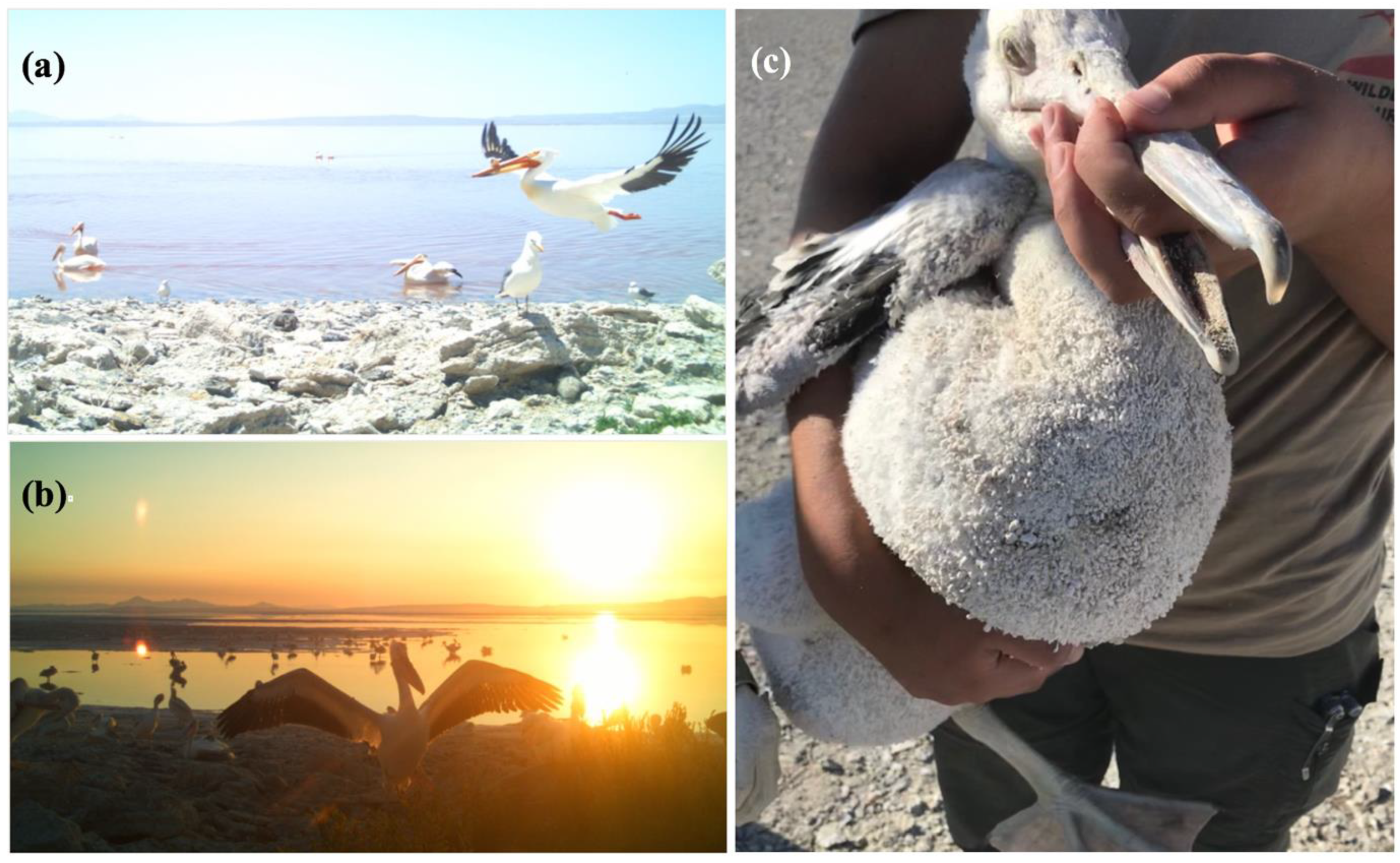
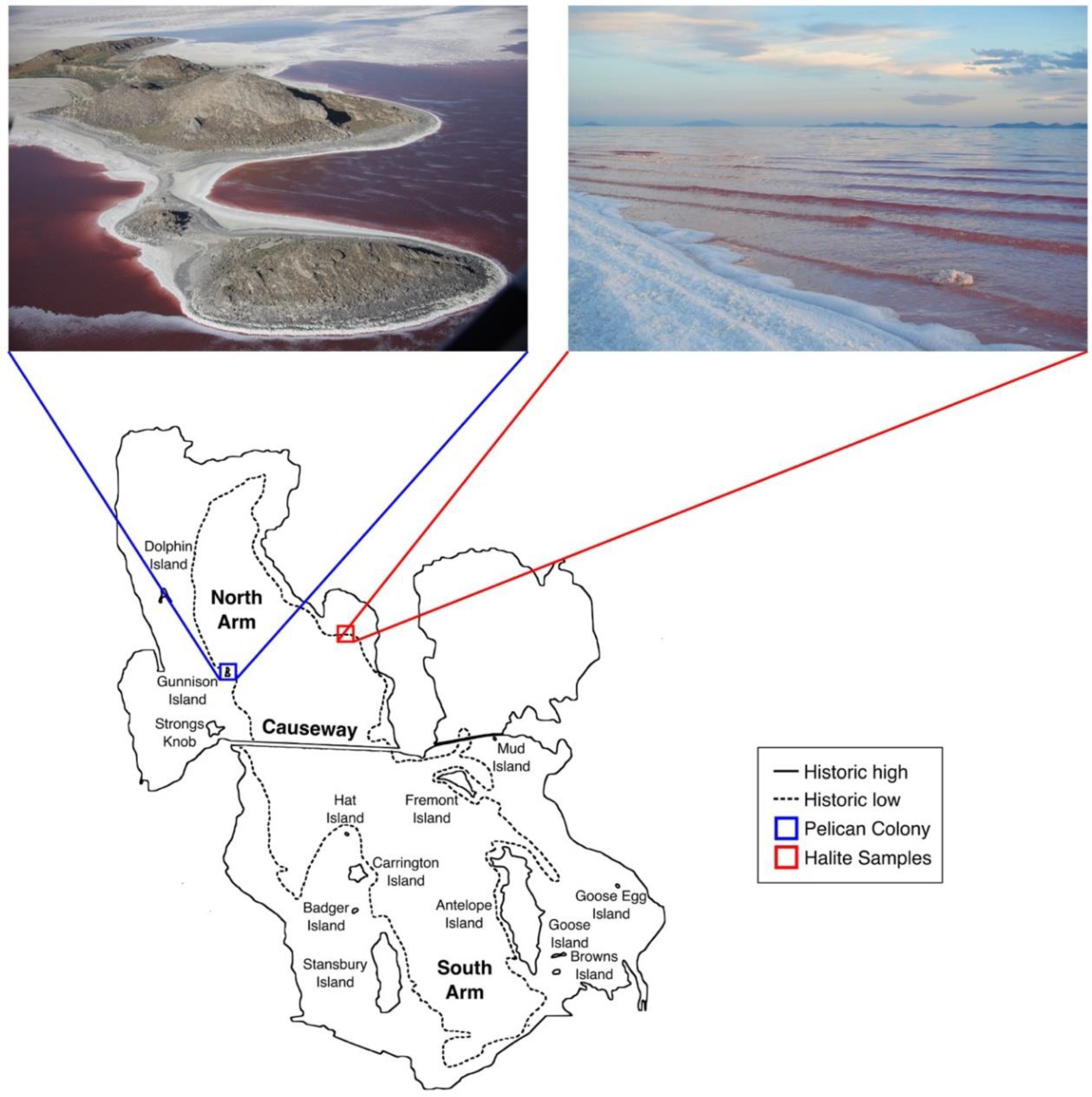
3.1. Great Salt Lake American White Pelicans, Feathers, and Halite Crystals
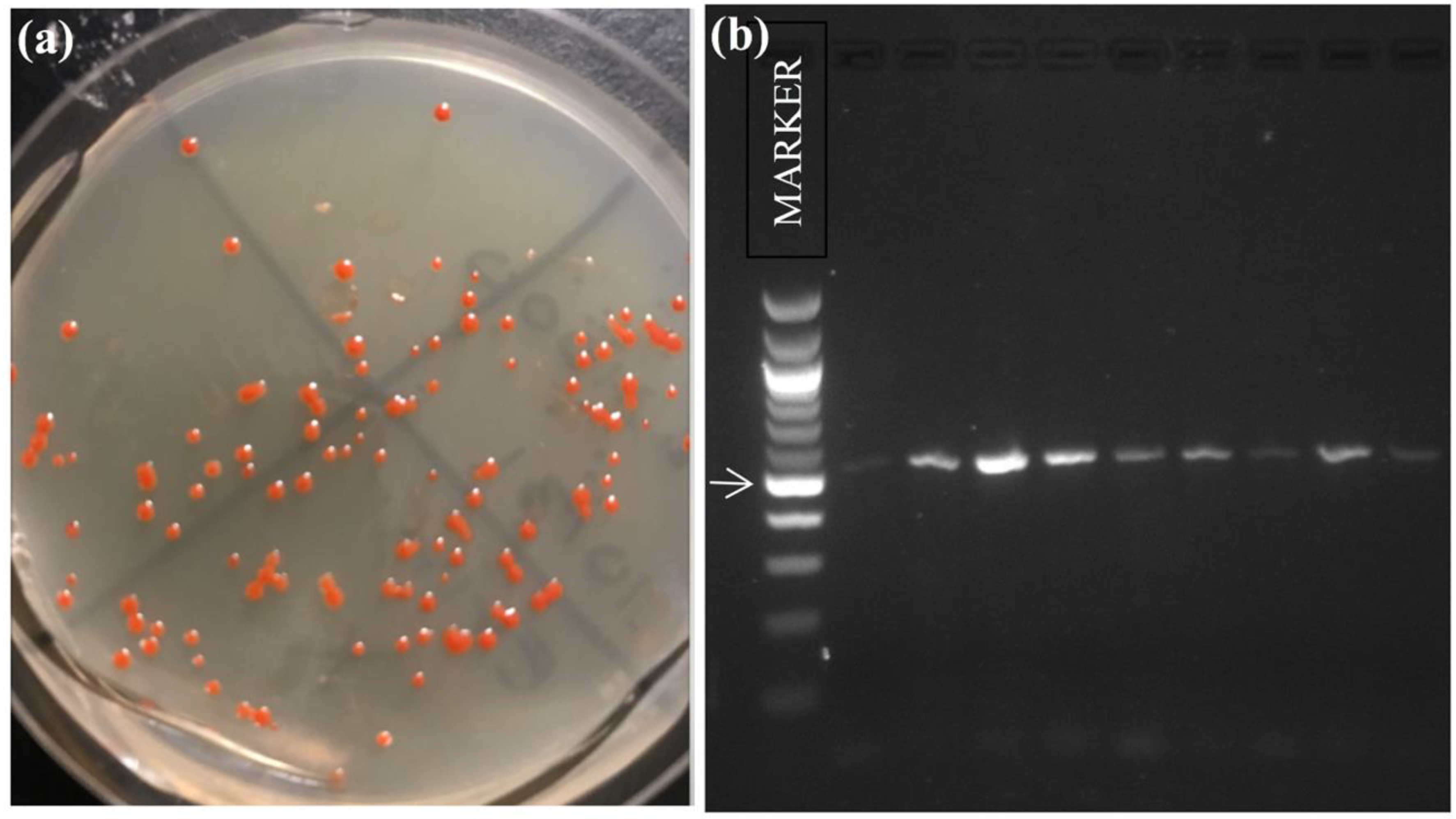
3.2. Halophilic Archaea Diversity of American White Pelican Feathers and Halite Crystal Cultivars
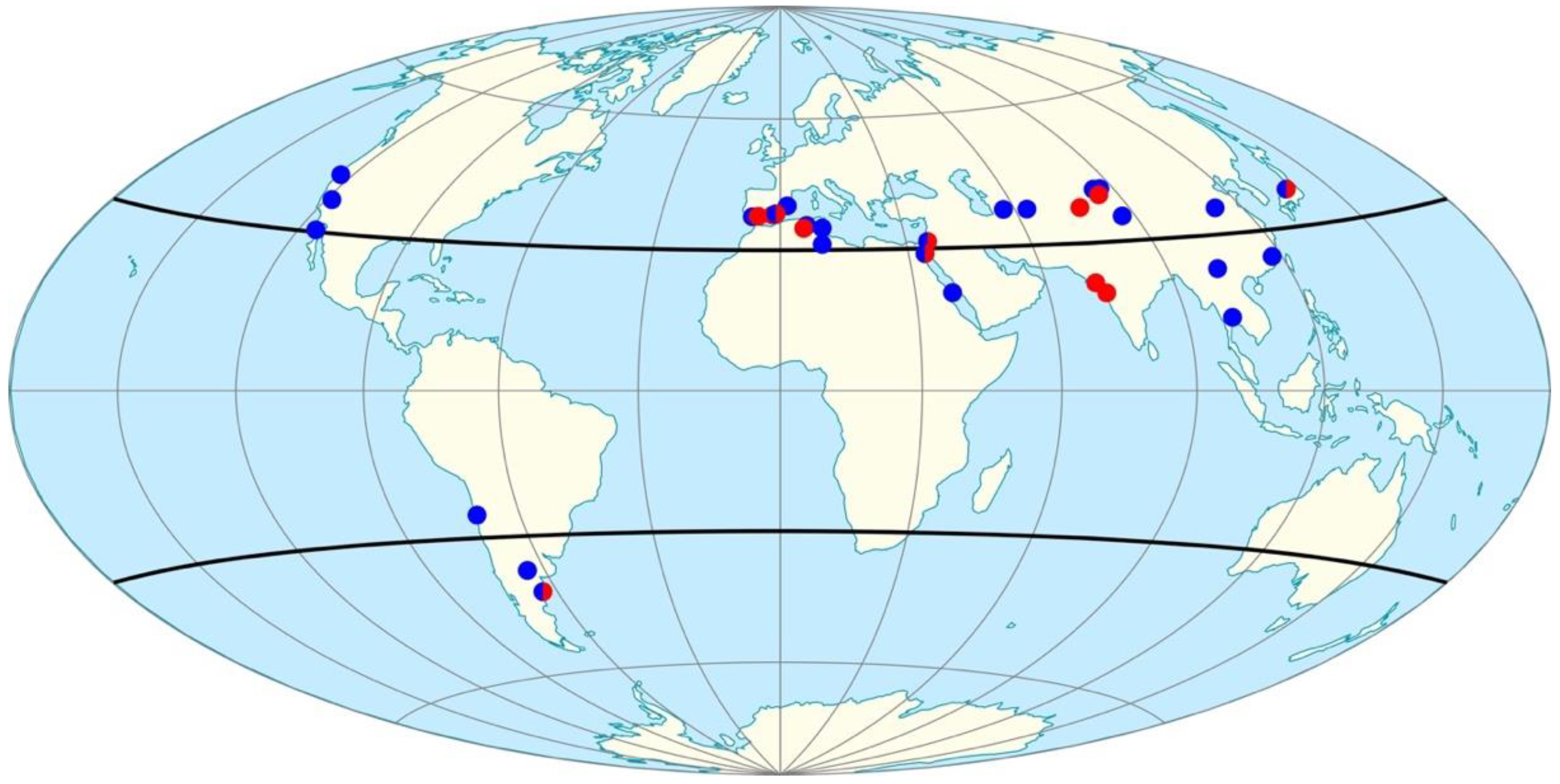
3.3. Geographic Distribution Patterns of Closest-Matched Strains
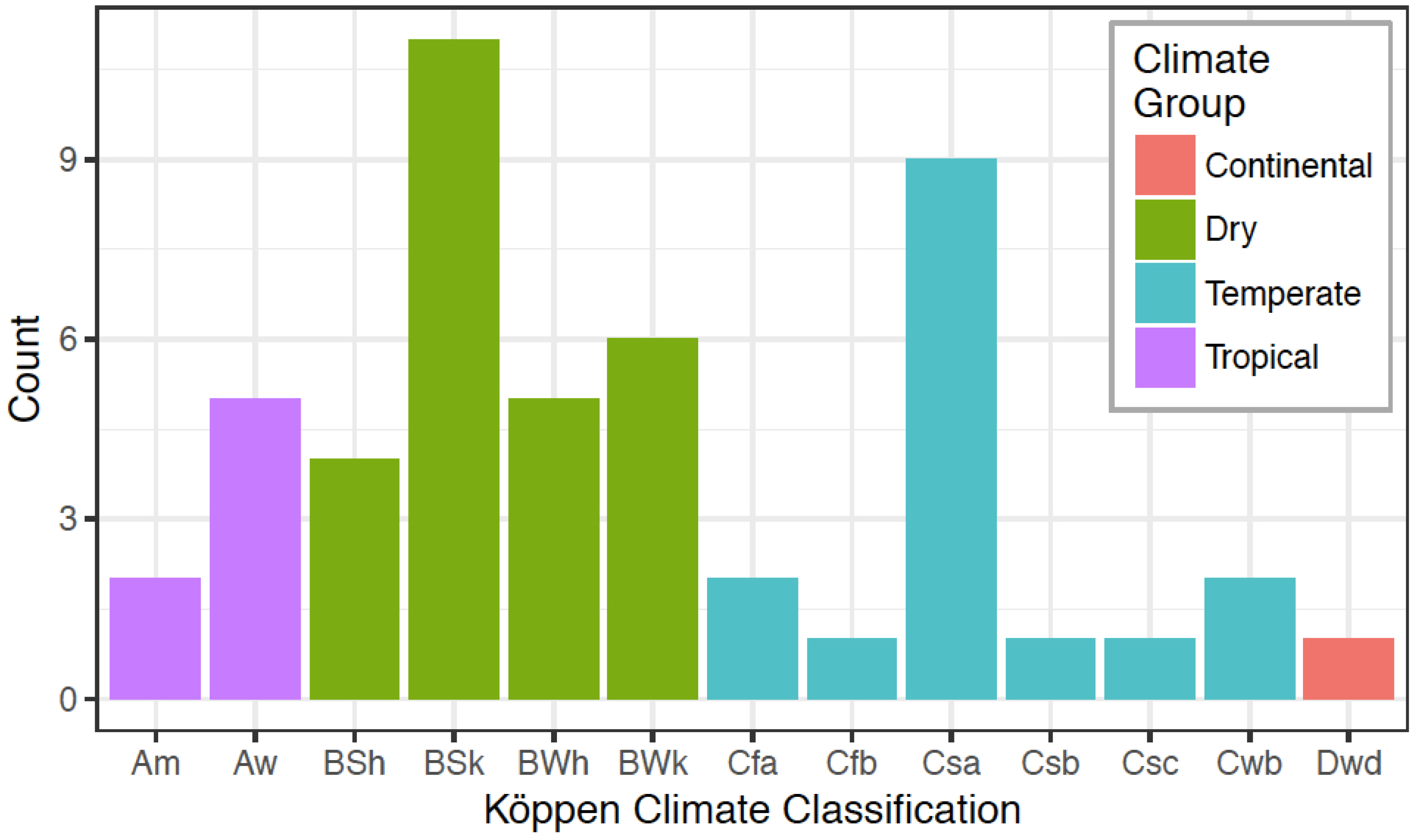
3.4. Environmental Factors as Variables in Distribution and Selection
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Data Availability Statement
Conflicts of Interest
References
- Antón, J.; Llobet-Brossa, E.; Rodriguez-Valera, F.; Amann, R. Fluorescence in situ hybridization analysis of the prokaryotic community inhabiting crystallizer ponds. Environ. Microbiol. 1999, 1, 517–523. [Google Scholar] [CrossRef] [PubMed]
- McGenity, T.J.; Gemmell, R.T.; Grant, W.D.; Stan-Lotter, H. Origins of halophilic microorganisms in ancient salt deposits. Environ. Microbiol. 2000, 2, 243–250. [Google Scholar] [CrossRef] [PubMed]
- Ghai, R.; Pašić, L.; Fernández, A.B.; Martin-Cuadrado, A.B.; Mizuno, C.M.; McMahon, K.D.; Papke, R.T.; Stepanauskas, R.; Rodriguez-Brito, B.; Rohwer, F.; et al. New abundant microbial groups in aquatic hypersaline environments. Sci. Rep. 2011, 1, 135. [Google Scholar] [CrossRef] [PubMed]
- Andrei, A.-Ş.; Banciu, H.L.; Oren, A. Living with salt: Metabolic and phylogenetic diversity of archaea inhabiting saline ecosystems. FEMS Microbiol. Lett. 2012, 330, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Almeida-Dalmet, S.; Sikaroodi, M.; Gillevet, P.M.; Litchfield, C.D.; Baxter, B.K. Temporal study of the microbial diversity of the North Arm of Great Salt Lake, Utah, US. Microorganisms 2015, 3, 310–326. [Google Scholar] [CrossRef] [PubMed]
- Fendrihan, S.; Dornmayr-Pfaffenhuemer, M.; Gerbl, F.W.; Holzinger, A.; Grösbacher, M.; Briza, P.; Erler, A.; Gruber, C.; Plätzer, K.; Stan-Lotter, H. Spherical particles of halophilic archaea correlate with exposure to low water activity–implications for microbial survival in fluid inclusions of ancient halite. Geobiology 2012, 10, 424–433. [Google Scholar] [CrossRef] [PubMed]
- Norton, C.F.; Grant, W.D. Survival of Halobacteria within fluid inclusions in salt crystals. J. Gen. Microbiol. 1988, 134, 1365–1373. [Google Scholar] [CrossRef]
- Norton, C.F.; McGenity, T.J.; Grant, W.D. Archaeal halophiles (halobacteria) from two British salt mines. Microbiology 1993, 139, 1077–1081. [Google Scholar] [CrossRef]
- Denner, E.B.M.; McGenity, T.J.; Busse, H.-J.; Grant, W.D.; Wanner, G.; Stan-Lotter, H. Halococcus salifodinae sp.nov., an archaeal isolate from an Austrian salt mine. Int. J. Syst. Evol. Microbiol. 1994, 44, 774–780. [Google Scholar]
- Grant, W.D.; Gemmell, R.T.; McGenity, T.J. Halobacteria: The evidence for longevity. Extremophiles 1998, 2, 279–287. [Google Scholar] [CrossRef] [PubMed]
- Stan-Lotter, H.; McGenity, T.J.; Legat, A.; Denner, E.B.; Glaser, K.; Stetter, K.O.; Wanner, G. Very similar strains of Halococcus salifodinae are found in geographically separated Permo-Triassic salt deposits. Microbiology 1999, 145, 3565–3574. [Google Scholar] [CrossRef] [PubMed]
- Vreeland, R.H.; Rosenzweig, W.D.; Powers, D.W. Isolation of a 250 million-year-old halotolerant bacterium from a primary salt crystal. Nature 2000, 407, 897–900. [Google Scholar] [CrossRef] [PubMed]
- Stan-Lotter, H.; Pfaffenhuemer, M.; Legat, A.; Busse, H.-J.; Radax, C.; Gruber, C. Halococcus dombrowskii sp. nov., an archaeal isolate from a Permo-Triassic alpine salt deposit. Int. J. Syst. Bacteriol. 2002, 52, 1807–1814. [Google Scholar]
- Kminek, G.; Bada, J.L.; Pogliano, K.; Ward, J.F. Radiation-Dependent Limit for the Viability of Bacterial Spores in Halite Fluid Inclusions and on Mars. Radiat. Res. 2003, 159, 722–729. [Google Scholar] [CrossRef]
- Mormile, M.R.; Biesen, M.A.; Gutierrez, M.C.; Ventosa, A.; Pavlovich, J.B.; Onstott, T.C.; Fredrickson, J.K. Isolation of Halobacterium salinarum retrieved directly from halite brine inclusions. Environ. Microbiol. 2003, 5, 1094–1102. [Google Scholar] [CrossRef] [PubMed]
- Gruber, C.; Legat, A.; Pfaffenhuemer, M.; Radax, C.; Weidler, G.; Busse, H.J.; Stan-Lotter, H. Halobacterium noricense sp. nov., an archaeal isolate from a bore core of an alpine Permian salt deposit, classification of Halobacterium sp. NRC-1 as a strain of H. salinarum and emended description of H. salinarum. Extremophiles 2004, 8, 431–439. [Google Scholar] [CrossRef] [PubMed]
- Park, J.S.; Vreeland, R.H.; Cho, B.C.; Lowenstein, T.K.; Timofeeff, M.N.; Rosenzweig, W.D. Haloarchaeal diversity in 23, 121 and 419 MYA salts. Geobiology 2009, 7, 515–523. [Google Scholar] [CrossRef] [PubMed]
- Schubert, B.A.; Lowenstein, T.K.; Timofeeff, M.N. Microscopic identification of prokaryotes in modern and ancient halite, Saline Valley and Death Valley, California. Astrobiology 2009, 9, 467–482. [Google Scholar] [CrossRef] [PubMed]
- Schubert, B.A.; Lowenstein, T.K.; Timofeeff, M.N.; Parker, M.A. Halophilic Archaea cultured from ancient halite, Death Valley, California. Environ. Microbiol. 2010, 12, 440–454. [Google Scholar] [CrossRef] [PubMed]
- Lowenstein, T.K.; Schubert, B.A.; Timofeeff, M.N. Microbial communities in fluid inclusions and long-term survival in halite. Geol. Soc. Am. Today 2011, 21, 4–9. [Google Scholar] [CrossRef]
- Sankaranarayanan, K.; Timofeeff, M.N.; Spathis, R.; Lowenstein, T.K.; Lum, J.K. Ancient Microbes from Halite Fluid Inclusions: Optimized Surface Sterilization and DNA Extraction. PLoS ONE 2011, 6, e20683. [Google Scholar] [CrossRef] [PubMed]
- Mancinelli, R.L.; White, M.R.; Rothschild, L.J. Biopan-survival I: Exposure of the osmophiles Synechococcus sp. (Nageli) and Haloarcula sp. to the space environment. Adv. Space Res. 1998, 22, 327–334. [Google Scholar] [CrossRef]
- Mancinelli, R.L. The effect of the space environment on the survival of Halorubrum chaoviator and Synechococcus (Nägeli): Data from the Space Experiment OSMO on EXPOSE-R. Int. J. Astrobiol. 2015, 14, 123–128. [Google Scholar] [CrossRef]
- Rabbow, E.; Horneck, G.; Rettberg, P.; Schott, J.U.; Panitz, C.; L’Afflitto, A.; von Heise-Rotenburg, R.; Willnecker, R.; Baglioni, P.; Hatton, J.; et al. EXPOSE, an astrobiological exposure facility on the international space station-from proposal to flight. Orig. Life Evol. Biospheres 2009, 39, 581–598. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.L.; Baxter, B.K. DNA Repair and Photoprotection: Mechanisms of Overcoming Environmental Ultraviolet Radiation Exposure in Halophilic Archaea. Front. Microbiol. 2017, 8, 1882. [Google Scholar] [CrossRef] [PubMed]
- da Costa, M.S.; Santos, H.; Galinski, E.A. An overview of the role and diversity of compatible solutes in bacteria and archaea. Adv. Biochem. Eng. Biotechnol. 1998, 61, 117–153. [Google Scholar] [PubMed]
- Roberts, M.F. Organic compatible solutes of halotolerant and halophilic microorganisms. Saline Syst. 2005, 1, 1–30. [Google Scholar] [CrossRef] [PubMed]
- Baxter, B.K.; Eddington, B.; Riddle, M.R.; Webster, T.N.; Avery, B.J. Great Salt Lake Halophilic Microorganisms as Models for Astrobiology: Evidence for Desiccation Tolerance and Ultraviolet Radiation Resistance. In Proceedings of the SPIE 6694, Instruments, Methods, and Missions for Astrobiology X, 669415, Bellingham, WA, USA, 3 October 2007. [Google Scholar]
- Kish, A.; DiRuggiero, J. DNA replication and repair in Halophiles. In Advances in Understanding the Biology of Halophilic Microorganisms; Vreeland, H.R., Ed.; Springer: Dordrecht, The Netherlands, 2012; pp. 163–198. ISBN 978-94-007-5538-3. [Google Scholar]
- Jones, D.L.; Baxter, B.K. Bipyrimidine Signatures as a Photoprotective Genome Strategy in G+ C-rich Halophilic Archaea. Life 2016, 6, 37. [Google Scholar] [CrossRef] [PubMed]
- Oren, A. Taxonomy of halophilic Archaea: Current status and future challenges. Extremophiles 2014, 18, 825–834. [Google Scholar] [CrossRef] [PubMed]
- Benlloch, S.; Martinez-Murcia, A.J.; Rodriguez-Valera, F. Sequencing of bacterial and archaeal 16S rRNA genes directly amplified from a hypersaline environment. Syst. Appl. Microbiol. 1995, 18, 574–581. [Google Scholar] [CrossRef]
- Benlloch, S.; Acinas, S.G.; Antón, J.; Lopez-Lopez, A.; Luz, S.P.; Rodríguez-Valera, F. Archaeal biodiversity in crystallizer ponds from a solar saltern: Culture versus PCR. Microb. Ecol. 2001, 41, 12–19. [Google Scholar] [PubMed]
- McGenity, T.J.; Grant, W.D. Transfer of Halobacterium saccharovorum, Halobacterium sodomense, Halobacterium trapanicum NRC 34021 and Halobacterium lacusprofundi to the genus Halorubrum gen. nov., as Halorubrum saccharovorum comb. nov., Halorubrum sodomense comb. nov., Halorubrum trapanicum comb. nov., and Halorubrum lacusprofundi comb. nov. Syst. Appl. Microbiol. 1995, 18, 237–243. [Google Scholar]
- Papke, R.T.; Koenig, J.E.; Rodríguez-Valera, F.; Doolittle, W.F. Frequent recombination in a saltern population of Halorubrum. Science 2004, 306, 1928–1929. [Google Scholar] [PubMed]
- Mancinelli, R.L.; Landheim, R.; Sanchez-Porro, C.; Dornmayr-Pfaffenhuemer, M.; Gruber, C.; Legat, A.; Ventosa, A.; Radax, C.; Ihara, K.; White, M.R.; et al. Halorubrum chaoviator sp. nov., a haloarchaeon isolated from sea salt in Baja California, Mexico, Western Australia and Naxos, Greece. Int. J. Syst. Evol. Microbiol. 2009, 59, 1908–1913. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.X.; Zhao, Z.W.; Zeng, C.; Yang, Z.L. Phylogenetic analysis of 16S rRNA gene reveals high species diversity of Halorubrum in China. Afr. J. Microbiol. Res. 2013, 7, 3009–3017. [Google Scholar]
- Oren, A.; Hallsworth, J.E. Microbial weeds in hypersaline habitats: The enigma of the weed-like Haloferax mediterranei. FEMS Microbiol. Lett. 2014, 359, 134–142. [Google Scholar] [CrossRef] [PubMed]
- Papke, R.T.; Zhaxybayeva, O.; Feil, E.J.; Sommerfeld, K.; Muise, D.; Doolittle, W.F. Searching for species in haloarchaea. Proc. Natl. Acad. Sci. USA 2007, 104, 14092–14097. [Google Scholar] [CrossRef] [PubMed]
- Fullmer, M.S.; Soucy, S.M.; Swithers, K.S.; Makkay, A.M.; Wheeler, R.; Ventosa, A.; Gogarten, J.P.; Papke, R.T. Population and genomic analysis of the genus Halorubrum. Front. Microbiol. 2014, 5, 140. [Google Scholar] [CrossRef] [PubMed]
- Ram Mohan, N.; Fullmer, M.S.; Makkay, A.M.; Wheeler, R.; Ventosa, A.; Naor, A.; Gogarten, J.P.; Papke, R.T. Evidence from phylogenetic and genome fingerprinting analyses suggests rapidly changing variation in Halorubrum and Haloarcula populations. Front. Microbiol. 2014, 5, 143. [Google Scholar] [CrossRef] [PubMed]
- Oren, A.; Ginzburg, M.; Ginzburg, B.Z.; Hochstein, L.I.; Volcani, B.E. Haloarcula marismortui (Volcani) sp. nov., nom. rev., an extremely halophilic bacterium from the Dead Sea. Int. J. Syst. Evol. Microbiol. 1990, 40, 209–210. [Google Scholar] [CrossRef] [PubMed]
- Takashina, T.; Hamamoto, T.; Otozai, K.; Grant, W.D.; Horikoshi, K. Haloarcula japonica sp. nov., a new triangular halophilic archaebacterium. Syst. Appl. Microbiol. 1990, 13, 177–181. [Google Scholar] [CrossRef]
- Juez, G.; Rodriguez-Valera, F.; Ventosa, A.; Kushner, D.J. Haloarcula hispanica spec. nov. and Haloferax gibbonsii spec, nov., two new species of extremely halophilic archaebacteria. Syst. Appl. Microbial. 1986, 8, 75–79. [Google Scholar] [CrossRef]
- Almeida-Dalmet, S.; Litchfield, C.D.; Gillevet, P.; Baxter, B.K. Differential gene expression in response to salinity and temperature in a Haloarcula Strain from Great Salt Lake, Utah. Genes 2018, 9, 52. [Google Scholar] [CrossRef] [PubMed]
- Riddle, M.R.; Baxter, B.K.; Avery, B.J. Molecular identification of microorganisms associated with the brine shrimp Artemia franciscana. Aquat. Biosyst. 2013, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Keck, W.; Hassibe, W. The Great Salt Lake; U.S. Geological Survey: Reston, VA, USA, 1978; pp. 1–16.
- Bellrose, F.C. Ducks, Geese, and Swans of North America; Stackpole Books: Harrisburg, PA, USA, 1980; ISBN 978-1421407517. [Google Scholar]
- Oring, L.W.; Neel, L.; Oring, K.E. Intermountain West Regional Shorebird Plan. Available online: https://www.shorebirdplan.org/wp-content/uploads/2013/01/IMWEST4.pdf (accessed on 3 October 2018).
- Paul, D.S.; Manning, A.E. Great Salt Lake Waterbird Survey Five-Year Report (1997–2001); Publication Number 08–38; Utah Division of Wildlife Resources: Salt Lake City, UT, USA, 2002.
- Aldrich, T.W.; Paul, D.S. Avian ecology of Great Salt Lake. In Great Salt Lake: An Overview of Change; Gwynn, J.W., Ed.; Utah Department of Natural Resources: Salt Lake City, Utah, USA, 2002; pp. 343–374. ISBN 1557916675. [Google Scholar]
- Neill, J.; Leite, B.; Gonzales, J.; Sanchez, K.; Luft, J. 2015 Great Salt Lake Eared Grebe Aerial Photo Survey: Annual Report; Utah Division of Wildlife Resources: Salt Lake City, UT, USA, 2016.
- Packard, A.S., Jr. On insects inhabiting salt water. Am. J. Sci. 1871, 3, 100–110. [Google Scholar] [CrossRef]
- Aldrich, J.M. The biology of some western species of the dipterous genus Ephydra. J. N. Y. Entomol. Soc. 1912, 20, 77–102. [Google Scholar]
- Collins, N. Population ecology of Ephydra cinerea Jones (Diptera: Ephydridae), the only benthic metazoan of the Great Salt Lake, USA. Hydrobiologia 1980, 68, 99–112. [Google Scholar] [CrossRef]
- Wurtsbaugh, W.A.; Gliwicz, Z.M. Limnological control of brine shrimp population dynamics and cyst production in the Great Salt Lake, Utah. Hydrobiologia 2001, 466, 119–132. [Google Scholar] [CrossRef]
- Roberts, A.J. Avian diets in a saline ecosystem: Great Salt Lake, Utah, USA. Hum. Wildl. Interact. 2013, 7, 15. [Google Scholar]
- Ochsenreiter, T.; Pfeifer, F.; Schleper, C. Diversity of Archaea in hypersaline environments characterized by molecular-phylogenetic and cultivation studies. Extremophiles 2002, 6, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Gill, F.B. Migration. In Ornithology, 2nd ed.; W.H. Freeman and Co.: New York, NY, USA, 1995; pp. 287–309. [Google Scholar]
- Hubálek, Z. An annotated checklist of pathogenic microorganisms associated with migratory birds. J. Wildl. Dis. 2004, 40, 639–659. [Google Scholar] [CrossRef] [PubMed]
- Georgopoulou, I.; Tsiouris, V. The potential role of migratory birds in the transmission of zoonoses. Vet. Ital. 2008, 44, 671–677. [Google Scholar] [PubMed]
- Warner, G.M.; French, D.W. Dissemination of fungi by migratory birds: Survival and recovery of fungi from birds. Can. J. Bot. 1970, 48, 907–910. [Google Scholar] [CrossRef]
- Johnson, M.L.; Speare, R. Possible modes of dissemination of the amphibian chytrid Batrachochytrium dendrobatidis in the environment. Dis. Aquat. Org. 2005, 65, 181–186. [Google Scholar] [CrossRef] [PubMed]
- Oh, D.; Porter, K.; Russ, B.; Burns, D.; Dyall-Smith, M. Diversity of Haloquadratum and other haloarchaea in three, geographically distant, Australian saltern crystallizer ponds. Extremophiles 2010, 14, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Baricz, A.; Cristea, A.; Muntean, V.; Teodosiu, G.; Andrei, A.Ş.; Molnár, I.; Alexe, M.; Rakosy-Tican, E.; Banciu, H.L. Culturable diversity of aerobic halophilic archaea (Fam. Halobacteriaceae) from hypersaline, meromictic Transylvanian lakes. Extremophiles 2015, 19, 525–537. [Google Scholar] [CrossRef] [PubMed]
- Brito-Echeverría, J.; López-López, A.; Yarza, P.; Antón, J.; Rosselló-Mora, R. Occurrence of Halococcus spp. in the nostrils salt glands of the seabird Calonectris diomedea. Extremophiles 2009, 13, 557–565. [Google Scholar] [CrossRef] [PubMed]
- Yim, K.J.; Kwon, J.; Cha, I.T.; Oh, K.S.; Song, H.S.; Lee, H.W.; Rhee, J.K.; Song, E.J.; Rho, J.R.; Seo, M.L.; et al. Occurrence of viable, red-pigmented haloarchaea in the plumage of captive flamingoes. Sci. Rep. 2015, 5, 16425. [Google Scholar] [CrossRef] [PubMed]
- Dyall-Smith, M. The Halohandbook, Version 7.2. 2009. Available online: http://www.haloarchaea.com/resources/halohandbook/ (accessed on 3 October 2018).
- Litchfield, C.D.; Sikaroodi, M.; Gillevet, P.M. Characterization of natural communities of halophilic microorganisms. Methods Microbiol. 2006, 35, 513–533. [Google Scholar]
- Mankin, A.S.; Kagramanova, V.K.; Teterina, N.L.; Rubtsov, P.M.; Belova, E.N.; Kopylov, A.M.; Baratova, L.A.; Bogdanov, A.A. The nucleotide sequence of the gene coding for the 16S rRNA from the archaebacterium Halobacterium halobium. Gene 1985, 37, 181–189. [Google Scholar] [CrossRef]
- Martinez-Murcia, A.J.; Acinas, S.G.; Rodriguez-Valera, F. Evaluation of prokaryotic diversity by restrictase digestion of 16S rDNA directly amplified from hypersaline environments. FEMS Microbiol. Ecol. 1995, 17, 247–255. [Google Scholar] [CrossRef]
- DeLong, E.F.; Wu, K.Y.; Prézelin, B.B.; Jovine, R.V. High abundance of Archaea in Antarctic marine picoplankton. Nature 1994, 371, 695. [Google Scholar] [CrossRef] [PubMed]
- National Library of Medicine, National Center for Biotechnology Information. Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE_TYPE=BlastSearch (accessed on 15 April 2018).
- Peel, M.C.; Finlayson, B.L.; McMahon, T.A. Updated world map of the Köppen-Geiger climate classification. Hydrol. Earth Syst. Sci. Discuss. 2007, 4, 439–473. [Google Scholar] [CrossRef]
- Climate-Data. Available online: https://en.climate-data.org/ (accessed on 17 March 2018).
- State of Utah. Pelican Management Act. Available online: https://le.utah.gov/xcode/Title23/Chapter21A/C23-21a_1800010118000101.pdf (accessed on 3 October 2018).
- Great Salt Lake Institute PELIcam. Available online: https://westminstercollege.edu/campus-life/centers-and-institutes/great-salt-lake-institute/pelicam (accessed on 24 May 2018).
- Neill, J.; Brewerton, A.; Davidson, M.; Leite, B.; Gonzalez, J.; Sanchez, K. 2016 American White Pelican Census, Gunnison Island Utah; Utah Division of Wildlife Resources: Salt Lake City, UT, USA, 2018; Unpublished report; 23p.
- Knopf, F.L. Spatial and temporal aspects of colonial nesting of White Pelicans. Condor 1979, 81, 353–363. [Google Scholar] [CrossRef]
- State of Utah, Division of Wildlife Resources PeliTrack. Available online: https://wildlife.utah.gov/pelican_webmap/ (accessed on 13 July 2018).
- Pal, K.K.; Dey, R.; Thomas, M.; Dave, S.R.; Ghorai, S. Diversity of Archaebacteria of Natural and Man-Made Salt Pan of Kachchh Region of Gujarat. Direct Submission to GenBank; 2011. Available online: https://www.ncbi.nlm.nih.gov/nuccore/JF802147.1?report=GenBank (accessed on 22 May 2018).
- Yang, Y.; Cui, H.L.; Zhou, P.J.; Liu, S.J. Haloarcula amylolytica sp. nov., an extremely halophilic archaeon isolated from Aibi salt lake in Xin-Jiang, China. Int. J. Syst. Evol. Microbiol. 2007, 57, 103–106. [Google Scholar] [CrossRef] [PubMed]
- Yadav, A.N.; Sachan, S.G.; Verma, P.; Kaushik, R.; Saxena, A.K. Diversity of Psychrotolerant Archaea from Cold Desert of Leh-Ladakh and Rohtang Pass (India). Direct Submission to GenBank; 2013. Available online: https://www.ncbi.nlm.nih.gov/nuccore/KF650693.1?report=GenBank (accessed on 22 May 2018).
- Ihara, K.; Watanabe, S.; Tamura, T. Haloarcula argentinensis sp. nov. and Haloarcula mukohataei sp. nov., two new extremely halophilic archaea collected in Argentina. Int. J. Syst. Evol. Microbiol. 1997, 47, 73–77. [Google Scholar] [CrossRef] [PubMed]
- Fukushima, T.; Usami, R.; Kamekura, M. A traditional Japanese-style salt field is a niche for haloarchaeal strains that can survive in 0.5% salt solution. Saline Syst. 2007, 3, 2. [Google Scholar] [CrossRef] [PubMed]
- Thombre, R.S.; Shinde, V.D.; Oke, R.S.; Dhar, S.K.; Shouche, Y.S. Biology and survival of extremely halophilic archaeon Haloarcula marismortui RR12 isolated from Mumbai salterns, India in response to salinity stress. Sci. Rep. 2016, 6, 25642. [Google Scholar] [CrossRef] [PubMed]
- Baliga, N.S.; Bonneau, R.; Facciotti, M.T.; Pan, M.; Glusman, G.; Deutsch, E.W.; Shannon, P.; Chiu, Y.; Weng, R.S.; Gan, R.R.; et al. Genome sequence of Haloarcula marismortui: A halophilic archaeon from the Dead Sea. Genome Res. 2004, 14, 2221–2234. [Google Scholar] [CrossRef] [PubMed]
- Luque, R.; González-Domenech, C.M.; Llamas, I.; Quesada, E.; Béjar, V. Diversity of culturable halophilic archaea isolated from Rambla Salada, Murcia (Spain). Extremophiles 2012, 16, 205–213. [Google Scholar] [CrossRef] [PubMed]
- Oren, A.; Ventosa, A.; Gutiérrez, M.C.; Kamekura, M. Haloarcula quadrata sp. nov., a square, motile archaeon isolated from a brine pool in Sinai (Egypt). Int. J. Syst. Evol. Microbiol. 1999, 49, 1149–1155. [Google Scholar] [CrossRef] [PubMed]
- Namwong, S.; Tanasupawat, S.; Kudo, T.; Itoh, T. Haloarcula salaria sp. nov. and Haloarcula tradensis sp. nov., isolated from salt in Thai fish sauce. Int. J. Syst. Evol. Microbiol. 2011, 61, 231–236. [Google Scholar] [CrossRef] [PubMed]
- Akolkar, AV.; Deshpande, GM.; Desai, A.J. Isolation and Characterization of Halophilic Archaea from Indian Salterns. Direct Submission to GenBank; 2006. Available online: https://www.ncbi.nlm.nih.gov/nuccore/DQ899739.1?report=GenBank (accessed on 22 May 2018).
- de Lourdes Moreno, M.; García, M.T.; Ventosa, A.; Mellado, E. Characterization of Salicola sp. IC10, a lipase-and protease-producing extreme halophile. FEMS Microbiol. Ecol. 2009, 68, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Kebbouche-Gana, S.; Gana, M.L.; Khemili, S.; Fazouane-Naimi, F.; Bouanane, N.A.; Penninckx, M.; Hacene, H. Isolation and characterization of halophilic Archaea able to produce biosurfactants. J. Ind. Microbiol. Biotechnol. 2009, 36, 727–738. [Google Scholar] [CrossRef] [PubMed]
- Nercessian, D.; Di Meglio, L.; De Castro, R.; Paggi, R. Exploring the multiple biotechnological potential of halophilic microorganisms isolated from two Argentinean salterns. Extremophiles 2015, 19, 1133–1143. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wu, Z.; Li, M.; Zhang, F.; Zheng, H.; Han, J.; Liu, J.; Zhou, J.; Wang, S.; Xiang, H. Complete genome sequence of Haloarcula hispanica, a model haloarchaeon for studying genetics, metabolism, and virus-host interaction. J. Bacteriol. 2011, 193, 6086–6087. [Google Scholar] [CrossRef] [PubMed]
- Javor, B.; Requadt, C.; Stoeckenius, W. Box-shaped halophilic bacteria. J. Bacteriol. 1982, 151, 1532–1542. [Google Scholar] [PubMed]
- Gonzalez, C.; Gutierrez, C.; Ramirez, C. Halobacterium vallismortis sp. nov. An amylolytic and carbohydrate-metabolizing, extremely halophilic bacterium. Can. J. Microbiol. 1978, 24, 710–715. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.W.; Wu, Y.H.; Zhang, H.B.; Wu, M. Halorubrum arcis sp. nov., an extremely halophilic archaeon isolated from a saline lake on the Qinghai–Tibet Plateau, China. Int. J. Syst. Evol. Microbiol. 2007, 57, 1069–1072. [Google Scholar] [CrossRef] [PubMed]
- Viver, T.; Cifuentes, A.; Díaz, S.; Rodríguez-Valdecantos, G.; González, B.; Antón, J.; Rosselló-Móra, R. Diversity of extremely halophilic cultivable prokaryotes in Mediterranean, Atlantic and Pacific solar salterns: Evidence that unexplored sites constitute sources of cultivable novelty. Syst. Appl. Microbiol. 2015, 38, 266–275. [Google Scholar] [CrossRef] [PubMed]
- Pesenti, P.T.; Sikaroodi, M.; Gillevet, P.M.; Sanchez-Porro, C.; Ventosa, A.; Litchfield, C.D. Halorubrum californiense sp. nov., an extreme archaeal halophile isolated from a crystallizer pond at a solar salt plant in California, USA. Int. J. Syst. Evol. Microbiol. 2008, 58, 2710–2715. [Google Scholar] [CrossRef] [PubMed]
- Baati, H.; Guermazi, S.; Gharsallah, N.; Sghir, A.; Ammar, E. Novel prokaryotic diversity in sediments of Tunisian multipond solar saltern. Res. Microbiol. 2010, 161, 573–582. [Google Scholar] [CrossRef] [PubMed]
- Boujelben, I.; Martínez-García, M.; van Pelt, J.; Maalej, S. Diversity of cultivable halophilic archaea and bacteria from superficial hypersaline sediments of Tunisian solar salterns. Antonie Leeuwenhoek 2014, 106, 675–692. [Google Scholar] [CrossRef] [PubMed]
- Dilbar, T.; Yinipigul, H. Isolation and phylogenetic analysis of haloalkalophilic bacteria in the black lake of Yuli County in Xinjiang. Microbiol. Bull. 2016, 43, 2601–2608. [Google Scholar]
- Castillo, A.M.; Gutiérrez, M.C.; Kamekura, M.; Xue, Y.; Ma, Y.; Cowan, D.A.; Jones, B.E.; Grant, W.D.; Ventosa, A. Halorubrum ejinorense sp. nov., isolated from Lake Ejinor, Inner Mongolia, China. Int. J. Syst. Evol. Microb. 2007, 757, 2538–2542. [Google Scholar] [CrossRef] [PubMed]
- Kharroub, K.; Quesada, T.; Ferrer, R.; Fuentes, S.; Aguilera, M.; Boulahrouf, A.; Ramos-Cormenzana, A.; Monteoliva-Sanchez, M. Halorubrum ezzemoulense sp. nov., a halophilic archaeon isolated from Ezzemoul sabkha, Algeria. Int. J. Syst. Evol. Microbiol. 2006, 56, 1583–1588. [Google Scholar] [CrossRef] [PubMed]
- Dammak, D.F.; Smaoui, S.M.; Ghanmi, F.; Boujelben, I.; Maalej, S. Characterization of halo-alkaline and thermostable protease from Halorubrum ezzemoulense strain ETR14 isolated from Sfax solar saltern in Tunisia. J. Basic Microbiol. 2016, 56, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.L.; Lin, Z.Y.; Dong, Y.; Zhou, P.J.; Liu, S.J. Halorubrum litoreum sp. nov., an extremely halophilic archaeon from a solar saltern. Int. J. Syst. Evol. Microbiol. 2007, 57, 2204–2206. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, R.O.; Higa, L.H.; Cutrullis, R.A.; Bilen, M.; Morelli, I.; Roncaglia, D.I.; Corral, R.S.; Morilla, M.J.; Petray, P.B.; Romero, E.L. Archaeosomes made of Halorubrum tebenquichense total polar lipids: A new source of adjuvancy. BMC Biotechnol. 2009, 9, 71. [Google Scholar] [CrossRef] [PubMed]
- Lizama, C.; Monteoliva-Sánchez, M.; Suárez-García, A.; Roselló-Mora, R.; Aguilera, M.; Campos, V.; Ramos-Cormenzana, A. Halorubrum tebenquichense sp. nov., a novel halophilic archaeon isolated from the Atacama Saltern, Chile. Int. J. Syst. Evol. Microbiol. 2002, 52, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Zvyagintseva, I.S.; Tarasov, A.L. Extreme halophilic bacteria from saline soils. Mikrobiologiya 1988, 56, 839–844. (In Russian) [Google Scholar]
- Ventosa, A.; Gutierrez, M.C.; Kamekura, M.; Zvyagintseva, I.S.; Oren, A. Taxonomic study of Halorubrum distributum and proposal of Halorubrum terrestre sp. nov. Int. J. Syst. Evol. Microbiol. 2004, 54, 389–392. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Zhou, P.J.; Liu, S.J. Halorubrum xinjiangense sp. nov., a novel halophile isolated from saline lakes in China. Int. J. Syst. Evol. Microbiol. 2004, 54, 1789–1791. [Google Scholar] [CrossRef] [PubMed]
- Atanasova, N.S.; Roine, E.; Oren, A.; Bamford, D.H.; Oksanen, H.M. Global network of specific virus–host interactions in hypersaline environments. Environ. Microbiol. 2012, 14, 426–440. [Google Scholar] [CrossRef] [PubMed]
- Atanasova, N.S.; Demina, T.A.; Buivydas, A.; Bamford, D.H.; Oksanen, H.M. Archaeal viruses multiply: Temporal screening in a solar saltern. Viruses 2015, 7, 1902–1926. [Google Scholar] [CrossRef] [PubMed]
- China, S. Species Diversity of Yunnan Salt Mine, China. Direct Submission to GenBank; 2016. Available online: https://www.ncbi.nlm.nih.gov/nuccore/KX376714.1 (accessed on 22 May 2018).
- Ma, Y.; Galinski, E.A.; Grant, W.D.; Oren, A.; Ventosa, A. Halophiles 2010: Life in Saline Environments. Appl. Environ. Microbiol. 2010, 76, 6971–6981. [Google Scholar] [CrossRef] [PubMed]
- Crosman, E.T.; Horel, J.D. MODIS-derived surface temperature of the Great Salt Lake. Remote Sens. Environ. 2009, 113, 73–81. [Google Scholar] [CrossRef]
- Post, F.J. The microbial ecology of the Great Salt Lake. Microbiol Ecol. 1977, 3, 143–165. [Google Scholar] [CrossRef] [PubMed]
- Baas-Becking, L.G.M. Geobiologie of Inleiding tot the Milieukunde; W. P. van Stockum and Zoon: The Hague, The Netherlands, 1934; ISBN 44189670. [Google Scholar]
- Finlay, B.J. Global dispersal of free-living microbial eukaryote species. Science 2002, 296, 1061–1063. [Google Scholar] [CrossRef] [PubMed]
- Whitfield, J. Biogeography: Is everything everywhere? Science 2005, 310, 960–962. [Google Scholar] [CrossRef] [PubMed]
- Martiny, J.B.; Bohannan, B.J.; Brown, J.H.; Colwell, R.K.; Fuhrman, J.A.; Green, J.L.; Horner-Devine, M.C.; Kane, M.; Krumins, J.A.; Kuske, C.R.; et al. Microbial biogeography: Putting microorganisms on the map. Nat. Rev. Microbiol. 2006, 4, 102–112. [Google Scholar] [CrossRef] [PubMed]
- Ryšánek, D.; Hrčková, K.; Škaloud, P. Global ubiquity and local endemism of free-living terrestrial protists: Phylogeographic assessment of the streptophyte alga Klebsormidium. Environ. Microbiol. 2014, 17, 689–698. [Google Scholar] [CrossRef] [PubMed]
- Van der Gast, C.J. Microbial biogeography: The end of the ubiquitous dispersal hypothesis? Environ. Microbiol. 2015, 17, 544–546. [Google Scholar] [CrossRef] [PubMed]
- Pagaling, E.; Wang, H.; Venables, M.; Wallace, A.; Grant, W.D.; Cowan, D.A.; Jones, B.E.; Ma, Y.; Ventosa, A.; Heaphy, S. Microbial biogeography of six salt lakes in Inner Mongolia, China, and a salt lake in Argentina. Appl. Environ. Microbiol. 2009, 75, 5750–5760. [Google Scholar] [CrossRef] [PubMed]
- Schiaffino, M.R.; Unrein, F.; Gasol, J.M.; Massana, R.; Balague, V.; Izaguirre, I. Bacterial community structure in a latitudinal gradient of lakes: The roles of spatial versus environmental factors. Freshw. Biol. 2011, 56, 1973–1991. [Google Scholar] [CrossRef]
- Bowers, K.J.; Wiegel, J. Temperature and pH optima of extremely halophilic archaea: A mini-review. Extremophiles 2011, 15, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Siroosi, M.; Amoozegar, M.A.; Khajeh, K.; Fazeli, M.; Rezaei, M.H. Purification and characterization of a novel extracellular halophilic and organic solvent-tolerant amylopullulanase from the haloarchaeon, Halorubrum sp. strain Ha25. Extremophiles 2014, 18, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Papke, R.T.; Ward, D.M. The importance of physical isolation to microbial diversification. FEMS Microbiol. Ecol. 2005, 48, 293–303. [Google Scholar] [CrossRef] [PubMed]
- Rossello-Mora, R.; Lucio, M.; Peña, A.; Brito-Echeverría, J.; Lopez-Lopez, A.; Valens-Vadell, M.; Frommberger, M.; Antón, J.; Schmitt-Kopplin, P. Metabolic evidence for biogeographic isolation of the extremophilic bacterium Salinibacter ruber. ISME J. 2008, 2, 242. [Google Scholar] [CrossRef] [PubMed]
- Woese, C.R. Bacterial evolution. Microbiol. Rev. 1987, 51, 221. [Google Scholar] [PubMed]
- Hedlund, B.P.; Staley, J.T. Microbial endemism and biogeography. In Microbial Diversity and Bioprospecting; Bull, A.T., Ed.; American Society for Microbiology Press: Washington, DC, USA, 2004; pp. 225–231. ISBN 1-55581-267-8. [Google Scholar]
- Mylvaganam, S.; Dennis, P.P. Sequence heterogeneity between the two genes encoding 16S rRNA from the halophilic archaebacterium Haloarcula marismortui. Genetics 1992, 130, 399–410. [Google Scholar] [PubMed]
- Boucher, Y.; Douady, C.J.; Sharma, A.K.; Kamekura, M.; Doolittle, W.F. Intragenomic heterogeneity and intergenomic recombination among haloarchaeal rRNA genes. J. Bacteriol. 2004, 186, 3980–3990. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.L.; Zhou, P.J.; Oren, A.; Liu, S.J. Intraspecific polymorphism of 16S rRNA genes in two halophilic archaeal genera. Haloarcula and Halomicrobium. Extremophiles 2009, 13, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Sun, D.L.; Jiang, X.; Wu, Q.L.; Zhou, N.Y. Intragenomic heterogeneity of 16S rRNA genes causes overestimation of prokaryotic diversity. Appl. Environ. Microbiol. 2013, 79, 5962–5969. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Oh, H.S.; Park, S.C.; Chun, J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int. J. Syst. Evol. Microbiol. 2014, 64, 346–351. [Google Scholar] [CrossRef] [PubMed]
- Head, I.M.; Saunders, J.R.; Pickup, R.W. Microbial evolution, diversity, and ecology: A decade of ribosomal RNA analysis of uncultivated microorganisms. Microb. Ecol. 1998, 35, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Pyatibratov, M.G.; Beznosov, S.N.; Rachel, R.; Tiktopulo, E.I.; Surin, A.K.; Syutkin, A.S.; Fedorov, O.V. Alternative flagellar filament types in the haloarchaeon Haloarcula marismortui. Can. J. Microbiol. 2008, 54, 835–844. [Google Scholar] [CrossRef] [PubMed]
- Oren, A. The enigma of square and triangular halophilic Archaea. In Enigmatic Microorganisms and Life in Extreme Environments; Seckbach, J., Ed.; Springer: Dordrecht, The Netherlands, 1999; pp. 337–355. ISBN 978-94-011-4838-2. [Google Scholar]
- Lin, Y.C.; Fu, H.Y.; Yang, C.S. Phototaxis of Haloarcula marismortui revealed through a novel microbial motion analysis algorithm. Photochem. Photobiol. 2010, 86, 1084–1090. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Valera, F.; Ruiz-Berraquero, F.; Ramos-Cormenzana, A. Isolation of extreme halophiles from seawater. Appl. Environ. Microbiol. 1979, 38, 164–165. [Google Scholar] [PubMed]
- Munson, M.A.; Nedwell, D.B.; Embley, T.M. Phylogenetic diversity of Archaea in sediment samples from a coastal salt marsh. Appl. Environ. Microbiol. 1997, 63, 4729–4733. [Google Scholar] [PubMed]
- Lavens, P.; Sorgeloos, P. The history, present status and prospects of the availability of Artemia cysts for aquaculture. Aquaculture 2000, 181, 397–403. [Google Scholar] [CrossRef]
- Kellogg, C.A.; Griffin, D.W. Aerobiology and the global transport of desert dust. Trends Ecol. Evol. 2006, 21, 638–644. [Google Scholar] [CrossRef] [PubMed]
- Hirst, J.M.; Stedman, O.J.; Hurst, G.W. Long-distance spore transport: Vertical sections of spore clouds over the sea. J. Gen. Microbiol. 1957, 48, 357–377. [Google Scholar] [CrossRef] [PubMed]
- Echigo, A.; Hino, M.; Fukushima, T.; Mizuki, T.; Kamekura, M.; Usami, R. Endospores of halophilic bacteria of the family Bacillaceae isolated from non-saline Japanese soil may be transported by Kosa event (Asian dust storm). Saline Syst. 2005, 1, 8. [Google Scholar] [CrossRef] [PubMed]
- Duce, R.A.; Unni, C.K.; Ray, B.J.; Prospero, J.M.; Merrill, J.T. Long-range atmospheric transport of soil dust from Asia to the tropical north Pacific: Temporal variability. Science 1980, 209, 1522–1524. [Google Scholar] [CrossRef] [PubMed]
- Taylor, D.A. Dust in the wind. Environ. Health Perspect. 2002, 110, A80. [Google Scholar] [CrossRef] [PubMed]
- Fendrihan, S.; Berces, A.; Lammer, H.; Musso, M.; Ronto, G.; Polacsek, T.K.; Holzinger, A.; Kolb, C.; Stan-Lotter, H. Investigating the effects of simulated Martian ultraviolet radiation on Halococcus dombrowskii and other extremely halophilic Archaebacteria. Astrobiology 2009, 9, 104–112. [Google Scholar] [CrossRef] [PubMed]
- Stan-Lotter, H.; Fendrihan, S. Halophilic archaea: Life with desiccation, radiation and oligotrophy over geological times. Life 2015, 5, 1487–1496. [Google Scholar] [CrossRef] [PubMed]
- Alerstam, T. Bird Migration; Cambridge University Press: Cambridge, UK, 1993; ISBN 0521448220. [Google Scholar]
- Liechti, F.; Schaller, E. The use of low-level jets by migrating birds. Naturwissenschaften 1999, 86, 549–551. [Google Scholar] [CrossRef] [PubMed]
- Able, K.P. A radar study of the altitude of nocturnal passerine migration. Bird Band. 1970, 41, 282–290. [Google Scholar] [CrossRef]


| Matched Strain | % Sim | Location [Reference] | KCC | EL (m) | AR (mm) | AT (°C) |
|---|---|---|---|---|---|---|
| Haloarcula amylolytica | 99 | Kachchh Salt Pans, Gujarat, India [81] | BSh | 5 | 753 | 27.3 |
| Haloarcula amylolytica | 99 | Xinjiang, China [82] | BSk | 408 | 514 | 13.2 |
| Haloarcula argentinensis | 99 | Leh-Ladkh, Rohtang Pass, India [83] | Dwd | 3978 | 27 | 3.75 |
| Haloarcula argentinensis | 99 | Salinas Chica, Chubut, Argentina [84] | BSk | −20 | 206 | 13.5 |
| Haloarcula hispanica | 99 | Santa Pola, Alicante, Spain [44] | Csa | 10 | 66 | 26 |
| Haloarcula japonica | 99 | Noto Peninsula, Japan [85] | Cfb | 438 | 180.016 | 13.75 |
| Haloarcula marismortui | 99 | Mumbai salterns, India [86] | Am | 14 | 242.2 | 27.2 |
| Haloarcula marismortui | 98 | Dead Sea, Israel [87] | Csa | −430.5 | 50 | 25.5 |
| Haloarcula quadrata | 99 | Kachchh Salt Pans, Gujarat, India [81] | BSh | 5 | 753 | 27.3 |
| Haloarcula quadrata | 99 | Salada, Murcia, Spain [88] | BSk | 4 | 297 | 18.6 |
| Haloarcula quadrata | 99 | Sebkha Gavish, Southern Sinai, Egypt [89] | BWh | 13 | 40 | 24.4 |
| Haloarcula salaria | 99 | Kachchh Salt Pans, Gujarat, India [81] | BSh | 5 | 753 | 27.3 |
| Haloarcula salaria | 99 | Thailand (Thai fish sauce) [90] | Aw | 0 | 1184 | 25.6 |
| Haloarcula tradensis | 99 | Thailand (Thai fish sauce) [90] | Aw | 0 | 1184 | 27.3 |
| Haloarcula sp. strain SP2(1) | 100 | Kachchh Salt Pans, Gujarat, India [91] | Am | 14 | 242.2 | 27.2 |
| Haloarcula sp. strain I.C6 | 100 | Isla Cristina, Southern Spain [92] | Csa | 11 | 612 | 17.8 |
| Haloarcula sp. strain D21 | 100 | Chott Melrhir, Algeria [93] | BWh | −40 | 160 | 22.8 |
| Matched Strain | % Sim | Location [Reference] | KCC | EL (m) | AR (mm) | AT (°C) |
|---|---|---|---|---|---|---|
| * Haloarcula argentinensis | 100 | Salinas Chica, Chubut, Argentina [84,94] | BSk | −20 | 206 | 13.5 |
| *Haloarcula hispanica | 99 | Santa Pola, Alicante, Spain [44] | Csa | 10 | 66 | 26 |
| Haloarcula hispanica | 100 | Santa Pola, Alicante, Spain [95] | Csa | 10 | 66 | 26 |
| *Haloarcula japonica | 100 | Noto Peninsula, Japan [85] | Cfa | 438 | 180.016 | 13.75 |
| *Haloarcula marismortui | 100 | Dead Sea, Israel [87] | Csa | −430.5 | 50 | 25.5 |
| *Haloarcula quadrata | 100 | Sebkha Gavish, Southern Sinai, Egypt [89] | BWh | 13 | 40 | 24.4 |
| *Haloarcula salaria | 99 | Thailand (Thai fish sauce) [90] | Aw | 0 | 1184 | 27.3 |
| Haloarcula sinaiiensis | 100 | Sebkha Gavish, Southern Sinai, Egypt [96] | BWh | −999 | 60 | 26.5 |
| *Haloarcula tradensis | 100 | Thailand (Thai fish sauce) [90] | Aw | 0 | 1184 | 27.3 |
| Haloarcula vallismortis | 100 | Death Valley, CA, USA [97] | BSk | −86 | 60 | 27.2 |
| Halorubrum arcis | 100 | Tibetan Plateau, China [98] | BWk | 4500 | 43 | 4.8 |
| Halorubrum californiense | 100 | Balearic Islands, Spain [99] | BSk | 13 | 449 | 18.2 |
| Halorubrum californiense | 100 | Newark, California, USA [100] | Csc | 6 | 368 | 15 |
| Halorubrum chaoviator | 100 | Santa Pola, Alicante, Spain [33] | BSh | 6 | 317 | 18.2 |
| Halorubrum chaoviator | 100 | Saharan Saline Syst., Southern Tunisia [101,102] | Csa | −22.5 | 100 | 20 |
| Halorubrum chaoviator | 99 | Yuli, Bayingol, Xinjiang, China [103] | BWk | 900 | 30 | 11 |
| Halorubrum chaoviator | 100 | Baja California, Mexico [36] | Csb | 660 | 116.8 | 18.2 |
| Halorubrum ejinorense | 100 | Lake Ejinor, Inner Mongolia, China [37,104] | BSk | 1065 | 391 | 6.4 |
| Halorubrum ezzemoulense | 100 | Sebkhet ez Zemoul, Oum el Bouaghi, Algeria [105] | BSk | 850 | 382 | 14.5 |
| Halorubrum ezzemoulense | 100 | Sfax solar saltern, Tunisia [106] | BWh | 0 | 212 | 19 |
| Halorubrum litoreum | 99 | Xiuyu, Fujian, China [107] | Cfa | 21 | 1178 | 20.9 |
| Halorubrum tebenquichense | 100 | Patagonian salt flat, Argentina [108] | BSk | 175 | 685.8 | 19.5 |
| Halorubrum tebenquichense | 100 | Atacama saltern, Chile [109] | BWk | 2339 | 7 | 17 |
| Halorubrum terrestre | 100 | Ashgabat, Turkmenistan [110] | BSk | 219 | 228 | 15.4 |
| Halorubrum terrestre | 100 | Gyaur-Kala, Turkmenistan [110,111] | BWk | 252 | 124 | 14.8 |
| Halorubrum trapanicum | 99 | Turkmenistan [110] | BWk | 281 | 124 | 14.8 |
| Halorubrum xinjiangense | 99 | Xiao-Er-Kule Lake, Xinjiang, China [112] | BWk | 4475 | 105 | 11.5 |
| Halorubrum sp. strain SP9-2 | 98 | Red Sea [113] | Csa | −400 | 41.9 | 26.1 |
| Halorubrum sp. strain spIB11 | 98 | Isla Bacuta and Isla Cristina, Huelva, Spain [92] | Csa | 8 | 467 | 17.8 |
| Halorubrum sp. strain SS13-13 | 98 | Samut Sakhon, Thailand [114] | Aw | 7 | 1330 | 27.8 |
| Halorubrum sp. strain Y122 | 99 | Yunnan Salt Mine, China [115] | Cwb | 1892 | 570.7 | 15.2 |
| Halorubrum sp. strain Y62 | 99 | Yunnan Salt Mine, China [37] | Cwb | 1892 | 570.7 | 15.2 |
| Halorubrum sp. strain YC-11 | 98 | Yuncheng Salt Lake, Shanxi, China [37] | BSk | 353 | 74.7 | 14.05 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kemp, B.L.; Tabish, E.M.; Wolford, A.J.; Jones, D.L.; Butler, J.K.; Baxter, B.K. The Biogeography of Great Salt Lake Halophilic Archaea: Testing the Hypothesis of Avian Mechanical Carriers. Diversity 2018, 10, 124. https://doi.org/10.3390/d10040124
Kemp BL, Tabish EM, Wolford AJ, Jones DL, Butler JK, Baxter BK. The Biogeography of Great Salt Lake Halophilic Archaea: Testing the Hypothesis of Avian Mechanical Carriers. Diversity. 2018; 10(4):124. https://doi.org/10.3390/d10040124
Chicago/Turabian StyleKemp, Bex L., Erin M. Tabish, Adam J. Wolford, Daniel L. Jones, Jaimi K. Butler, and Bonnie K. Baxter. 2018. "The Biogeography of Great Salt Lake Halophilic Archaea: Testing the Hypothesis of Avian Mechanical Carriers" Diversity 10, no. 4: 124. https://doi.org/10.3390/d10040124
APA StyleKemp, B. L., Tabish, E. M., Wolford, A. J., Jones, D. L., Butler, J. K., & Baxter, B. K. (2018). The Biogeography of Great Salt Lake Halophilic Archaea: Testing the Hypothesis of Avian Mechanical Carriers. Diversity, 10(4), 124. https://doi.org/10.3390/d10040124





