Bacteria Associated with Marine Benthic Invertebrates from Polar Environments: Unexplored Frontiers for Biodiscovery?
Abstract
1. Introduction
2. Polar Marine Environments
3. Cold-Adapted Bacteria
3.1. Definition
3.2. Survival Strategies
3.3. Biotechnological Relevance
4. Polar Marine Benthic Invertebrates as Bacterial Hosts
4.1. Bacteria Associated with Porifera
4.2. Bacteria Associated with Other Benthic Invertebrates
5. Polar Benthic Invertebrates as a Source for Bacterial Biodiscovery
5.1. Antimicrobials
5.2. Extracellular Polymeric Substances
5.3. Cold-Enzymes
6. Concluding Remarks
Funding
Conflicts of Interest
References
- Frias-Lopez, J.; Klaus, J.S.; Bonheyo, G.T.; Fouket, B.W. Bacterial community associated with black band disease in corals. Appl. Environ. Microb. 2004, 70, 5955–5962. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, K.B. Regulation of microbial populations by coral surface mucus and mucus-associated bacteria. Mar. Ecol.-Prog. Ser. 2006, 322, 1–14. [Google Scholar] [CrossRef]
- Egan, S.; Thomas, T.; Kjelleberg, S. Unlocking the diversity and biotechnological potential of marine surface associated microbial communities. Curr. Op. Microbiol. 2008, 11, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Webster, N.S.; Taylor, M.W.; Behnam, F.; Lücker, S.; Rattei, T.; Whalan, S.; Horn, M.; Wagner, M. Deep sequencing reveals exceptional diversity and modes of transmission for bacterial sponge symbionts. Environ. Microbiol. 2010, 12, 2070–2082. [Google Scholar] [CrossRef] [PubMed]
- Porporato, E.M.D.; Lo Giudice, A.; Michaud, L.; De Domenico, E.; Spanò, N. Diversity and antibacterial activity of the bacterial communities associated with two Mediterranean sea pens, Pennatula phosphorea and Pteroeides spinosum (Anthozoa: Octocorallia). Microb. Ecol. 2013, 66, 701–714. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.W.; Schupp, P.J.; Dahllöf, I.; Kjelleberg, S.; Steinberg, P.D. Host specificity in marine sponge-associated bacteria, and potential implications for marine microbial diversity. Environ. Microbiol. 2004, 6, 121–130. [Google Scholar] [CrossRef] [PubMed]
- Wichels, A.; Würtz, S.; Döpke, H.; Schütt, C.; Gerdts, G. Bacterial diversity in the breadcrumb sponge Halichondria panicea (Pallas). FEMS Microbiol. Ecol. 2006, 56, 102–118. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, C.; Lo Giudice, A. Marine invertebrates: Underexplored sources of bacteria producing biologically active molecules. Diversity 2018, 10, 52. [Google Scholar] [CrossRef]
- Nyholm, S.V.; McFall-Ngai, M. The winnowing: Establishing the squid-vibrio symbiosis. Nat. Rev. Microbiol. 2004, 2, 632–642. [Google Scholar] [CrossRef] [PubMed]
- Pearce, D.A. Extremophiles in Antarctica: Life at low temperatures. In Adaption of Microbial Life to Environmental Extremes; Stan-Lotter, H., Fendrihan, S., Eds.; Springer: Vienna, Austria, 2012; pp. 87–218. [Google Scholar]
- Everatt, M.J.; Convey, P.; Bale, J.S.; Worland, M.R.; Hayward, S.A. Responses of invertebrates to temperature and water stress: A polar perspective. J. Therm. Biol. 2015, 54, 118–132. [Google Scholar] [CrossRef] [PubMed]
- Dayton, P.K.; Robilliard, G.A.; Devries, A.L. Anchor ice formation in McMurdo Sound, Antarctica, and its biological effects. Science 1969, 163, 273–274. [Google Scholar] [CrossRef] [PubMed]
- Dayton, P.K.; Robilliard, G.A.; Paine, R.T.; Dayton, L.B. Biological accommodation in the benthic community at McMurdo Sound, Antarctica. Ecol. Monogr. 1974, 44, 105–128. [Google Scholar] [CrossRef]
- McClintock, J.B.; Amsler, C.D.; Baker, B.J. Overview of the chemical ecology of benthic marine invertebrates along the western Antarctic peninsula. Integr. Comp. Biol. 2010, 50, 967–980. [Google Scholar] [CrossRef] [PubMed]
- Radjasa, O.K. Bioprospecting of marine microbial symbionts: Exploitation of underexplored marine microorganisms. In Marine Microbiology. Bioactive Compounds and Biotechnological Applications; Kim, S.K., Ed.; Wiley-VCH Verlag GmbH & Co. KGaA: Weinheim, Germany, 2013; pp. 369–378. [Google Scholar]
- Pye, C.R.; Bertin, M.J.; Lokeya, R.S.; Gerwick, W.H.; Linington, R.G. Retrospective analysis of natural products provides insights for future discovery trends. Proc. Natl. Acad. Sci. USA 2017, 114, 5601–5606. [Google Scholar] [CrossRef] [PubMed]
- Thomas, T.R.; Kavlekar, D.P.; LokaBharathi, P.A. Marine drugs from sponge-microbe association-a review. Mar. Drugs 2010, 8, 1417–1468. [Google Scholar] [CrossRef] [PubMed]
- Poli, A.; Anzelmo, G.; Nicolaus, B. Bacterial exopolysaccharides from extreme marine habitats: Production, characterization and biological activities. Mar. Drugs 2010, 8, 1779–1802. [Google Scholar] [CrossRef] [PubMed]
- Margesin, R.; Neuner, G.; Storey, K.B. Cold-loving microbes, plants, and animals—Fundamental and applied aspects. Naturwissenschaften 2007, 94, 77–99. [Google Scholar] [CrossRef] [PubMed]
- Tytgat, B.; Verleyen, E.; Sweetlove, M.; D’hondt, S.; Clercx, P.; Van Ranst, E.; Peeters, K.; Roberts, S.; Namsaraev, Z.; Wilmotte, A.; et al. Bacterial community composition in relation to bedrock type and macrobiota in soils from the SørRondane Mountains, East Antarctica. FEMS Microbiol. Ecol. 2016, 92. [Google Scholar] [CrossRef] [PubMed]
- Lo Giudice, A.; Bruni, V.; De Domenico, M.; Michaud, L. Psychrophiles-Cold-adapted hydrocarbon-degrading microorganisms. In Handbook of Hydrocarbon and Lipid Microbiology; Timmis, K.N., Ed.; Springer: Berlin, Heidelberg, 2010; pp. 1897–1921. ISBN 978-3-540-77584-3. [Google Scholar]
- Dinniman, M.S.; Klinck, J.M. A model study of circulation and cross-shelf exchange on the west Antarctic Peninsula continental shelf. Deep-Sea Res. II 2004, 51, 2003–2022. [Google Scholar] [CrossRef]
- Morita, R.Y. Psychrophilic bacteria. Bacteriol. Rev. 1975, 39, 144–167. [Google Scholar] [PubMed]
- Feller, G.; Gerday, C. Psychrophilic enzymes: Hot topics in cold adaptation. Nat. Rev. Microbiol. 2003, 1, 200–208. [Google Scholar] [CrossRef] [PubMed]
- Cavicchioli, R. On the concept of a psychrophile. ISME J. 2016, 10, 793–795. [Google Scholar] [CrossRef] [PubMed]
- Chintalapati, S.; Kiran, M.D.; Shivaji, S. Role of membrane lipid fatty acids in cold adaptation. Cell. Mol. Biol. 2004, 50, 631–642. [Google Scholar] [PubMed]
- Poli, A.; Finore, I.; Romano, I.; Gioiello, A.; Lama, L.; Nicolaus, B. Microbial diversity in extreme marine habitats and their biomolecules. Microorganisms 2017, 5, 25. [Google Scholar] [CrossRef] [PubMed]
- Tribelli, P.M.; López, N.I. Reporting key features in cold-adapted bacteria. Life 2018, 13, 8. [Google Scholar] [CrossRef] [PubMed]
- Chattopadhyay, M.K.; Jagannadham, M.V.; Vairamani, M.; Shivaji, S. Carotenoid pigments of an Antarctic psychrotrophic bacterium Micrococcus roseus: Temperature dependent biosynthesis, structure and interaction with synthetic membranes. Biochem. Biophys. Res. Comm. 1997, 239, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Jagannadham, M.V.; Chattopadhyay, M.K.; Subbalakshmi, C.; Vairamani, M.; Narayanan, K.; Rao, C.M.; Shivaji, S. Carotenoids of an Antarctic psychrotolerant bacterium, Sphingobacterium antarcticus, and a mesophilic bacterium, Sphingobacterium multivorum. Arch. Microbiol. 2000, 173, 418–424. [Google Scholar] [CrossRef] [PubMed]
- Médigue, C.; Krin, E.; Pascal, G.; Barbe, V.; Bernsel, A.; Bertin, P.N.; Cheung, F.; Cruveiller, S.; D’Amico, S.; Duilio, A.; et al. Coping with cold: The genome of the versatile marine Antarctica bacterium Pseudoalteromonas haloplanktis TAC125. Genome Res. 2005, 15, 1325–1335. [Google Scholar] [CrossRef] [PubMed]
- Bergholz, P.W.; Bakermans, C.; Tiedje, J.M. Psychrobacter arcticus 273-4 uses resource efficiency and molecular motion adaptations for subzero temperature growth. J. Bacteriol. 2009, 191, 2340–2352. [Google Scholar] [CrossRef] [PubMed]
- Benforte, F.C.; Colonnella, M.A.; Ricardi, M.M.; Venero, E.C.S.; Lizarraga, L.; López, N.I.; Tribelli, P.M. Novel role of the LPS core glycosyltransferase WapH for cold adaptation in the Antarctic bacterium Pseudomonas extremaustralis. PLoS ONE 2018, 13, e0192559. [Google Scholar] [CrossRef] [PubMed]
- Mykytczuk, N.C.S.; Lawrence, J.R.; Omelon, C.R.; Southam, G.; Whyte, L.G. Microscopic characterization of the bacterial cell envelope of Planococcus halocryophilus Or1 during subzero growth at −15°C. Polar Biol. 2016, 39, 701–712. [Google Scholar] [CrossRef]
- Mykytczuk, N.C.S.; Foote, S.J.; Omelon, C.R.; Southam, G.; Greer, C.W.; Whyte, L.G. Bacterial growth at −15°C; molecular insights from the permafrost bacterium Planococcus halocryophilus Or1. ISME J. 2013, 7, 1211–1226. [Google Scholar] [CrossRef] [PubMed]
- Ronholm, J.; Raymond-Bouchard, I.; Creskey, M.; Cyr, T.; Cloutis, E.A.; Whyte, L.G. Characterizing the surface-exposed proteome of Planococcus halocryophilus during cryophilic growth. Extremophiles 2015, 19, 619–629. [Google Scholar] [CrossRef] [PubMed]
- D’Amico, S.; Collins, T.; Marx, J.-C.; Feller, G.; Gerday, C.; Gerday, C. Psychrophilic microorganisms: Challenges for life. EMBO Rep. 2006, 7, 385–389. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.; Yang, Z.K.; Wu, L.; Thompson, D.K.; Zhou, J. Global transcriptome analysis of the cold shock response of Shewanella oneidensis MR-1 and mutational analysis of its classical cold shock proteins. J. Bacteriol. 2006, 188, 4560–4569. [Google Scholar] [CrossRef] [PubMed]
- Rosen, R.; Ron, E.Z. Proteome analysis in the study of the bacterial heat-shock response. Mass Spectrom. Rev. 2002, 21, 244–265. [Google Scholar] [CrossRef] [PubMed]
- Piette, F.; D’Amico, S.; Mazzucchelli, G.; Danchin, A.; Leprince, P.; Feller, G. Life in the cold: A proteomic study of cold-repressed proteins in the Antarctic bacterium Pseudoalteromonas haloplanktis TAC125. Appl. Environ. Microbiol. 2011, 77, 3881–3883. [Google Scholar] [CrossRef] [PubMed]
- De Maayer, P.; Anderson, D.; Cary, G.; Cowan, D.A. Some like it cold: Understanding the survival strategies of psychrophiles. EMBO Rep. 2014, 19, e201338170. [Google Scholar] [CrossRef] [PubMed]
- Celik, Y.; Drori, R.; Petraya-Braun, N.; Altan, A.; Barton, T.; Bar-Dolev, M.; Groisman, A.; Davies, P.L.; Braslavsky, I. Microfluidic experiments reveal that antifreeze proteins bound to ice crystals suffice to prevent their growth. Proc. Natl. Acad. Sci. USA 2013, 110, 1309–1314. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, H. The structures and functions of ice crystal controlling proteins from bacteria. J. Biosci. Bioeng. 2002, 94, 492–496. [Google Scholar] [CrossRef]
- Nichols, C.A.; Guezzenec, J.; Bowman, J.P. Bacterial exopolysaccharides from extreme environments with special consideration of the southern ocean, sea ice, and deep-sea hydrothermal vents: A review. Mar. Biotechnol. 2005, 7, 253–271. [Google Scholar] [CrossRef] [PubMed]
- Mancuso Nichols, C.A.; Bowman, J.P.; Guezennec, J. Effects of incubation temperature on growth and production of exopolysaccharides by an Antarctic sea ice bacterium grown in batch culture. Appl. Environ. Microbiol. 2005, 71, 3519–3523. [Google Scholar] [CrossRef] [PubMed]
- Caruso, C.; Rizzo, C.; Mangano, S.; Poli, A.; Di Donato, P.; Nicolaus, B.; Di Marco, G.; Michaud, L.; Lo Giudice, A. Extracellular polymeric substances with metal adsorption capacity produced by Pseudoalteromonas sp. MER144 from Antarctic seawater. Environ. Sci. Poll. Res. 2018, 25, 4667–4677. [Google Scholar] [CrossRef] [PubMed]
- Ayub, N.D.; Tribelli, P.M.; López, N.I. Polyhydroxyalkanoates are essential for maintenance of redox state in the Antarctic bacterium Pseudomonas sp. 14–3 during low temperature adaptation. Extremophiles 2009, 13, 59–66. [Google Scholar] [CrossRef] [PubMed]
- Tribelli, P.M.; López, N.I. Poly(3-hydroxybutyrate) influences biofilm formation and motility in the novel Antarctic species Pseudomonas extremaustralis under cold conditions. Extremophiles 2011, 15, 541. [Google Scholar] [CrossRef] [PubMed]
- Lo Giudice, A.; Fani, R. Cold-adapted bacteria from the coastal Ross Sea (Antarctica): Linking microbial ecology to biotechnology. Hydrobiologia 2015, 761, 417–441. [Google Scholar] [CrossRef]
- Liu, J.T.; Lu, X.L.; Liu, X.Y.; Gao, Y.; Hu, B.; Jiao, B.H.; Zheng, H. Bioactive natural products from the Antarctic and Arctic organisms. Mini Rev. Med. Chem. 2013, 13, 617–626. [Google Scholar] [CrossRef] [PubMed]
- Nichols, D.S.; Hart, P.; Nichols, P.D.; McMeekin, T.A. Enrichment of the rotifer Brachionus plicatilis fed an Antarctic bacterium containing polyunsaturated fatty acids. Aquaculture 1996, 147, 115–125. [Google Scholar] [CrossRef]
- Gentile, G.; Bonasera, V.; Amico, C.; Giuliano, L.; Yakimov, M.M. Shewanella sp. GA-22, a psychrophilic hydrocarbonoclastic Antarctic bacterium producing polyunsaturated fatty acids. J. Appl. Microbiol. 2003, 95, 1124–1133. [Google Scholar] [CrossRef] [PubMed]
- Russell, N.J.; Nichols, D.S. Polyunsaturated fatty acids in marine bacteria—A dogma rewritten. Microbiology 1999, 145, 767–779. [Google Scholar] [CrossRef] [PubMed]
- Bowman, J.P.; McCammon, S.A.; Nichols, D.S.; Skerratt, J.H.; Rea, S.M.; Rea Nichols, P.D.; McMeekin, T.A. Shewanella gelidimarina sp. nov. and Shewanella frigidimarina sp. nov., novel Antarctic species with the ability to produce eicosapentaenoic acid (20:5x3) and grow anaerobically by dissimilatoty Fe (III) reduction. Int. J. Syst. Bacteriol. 1997, 47, 1040–1047. [Google Scholar] [CrossRef] [PubMed]
- Sarmiento, F.; Peralta, R.; Blamey, J.M. Cold and Hot Extremozymes: Industrial Relevance and Current Trends. Front. Bioeng. Biotechnol. 2015, 3, 148. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.S.; Mody, K.; Jha, B. Bacterial exopolysaccharides—A perception. J. Basic Microb. 2007, 47, 103–117. [Google Scholar] [CrossRef] [PubMed]
- Freitas, F.; Alves, V.D.; Reis, M.A. Advances in bacterial exopolysaccharides: From production to biotechnological applications. Trends Biotechnol. 2011, 29, 388–398. [Google Scholar] [CrossRef] [PubMed]
- Angulo-Preckler, C.; Spurkland, T.; Avila, C.; Iken, K. Antimicrobial activity of selected benthic Arctic invertebrates. Polar Biol. 2015, 38, 1941. [Google Scholar] [CrossRef]
- Gili, J.-M.; Arntz, W.E.; Palanques, A.; Orejas, C.; Clarke, A.; Dayton, P.K.; Isla, E.; Teixidó, N.; Rossi, S.; López-González, P.J. A unique assemblage of epibenthic sessile suspension feeders with archaic features in the high-Antarctic. Deep Sea Res. II 2006, 53, 1029–1052. [Google Scholar] [CrossRef]
- Clark, G.F.; Raymond, B.; Riddle, M.J.; Stark, J.S.; Johnston, E.L. Vulnerability of Antarctic shallow invertebrate-dominated ecosystems. Austral. Ecol. 2015, 40, 482–491. [Google Scholar] [CrossRef]
- Nichols, D.; Lewis, K.; Orjala, J.; Mo, S.; Ortenberg, R.; O’Connor, P.; Zhao, C.; Vouros, P.; Kaeberlein, T.; Epstein, S.S. Short peptide induces an “uncultivable” microorganism to grow in vitro. Appl. Environ. Microbiol. 2008, 74, 4889–4897. [Google Scholar] [CrossRef] [PubMed]
- Vartoukian, S.R.; Palmer, R.M.; Wade, W.G. Strategies for culture of ‘unculturable’ bacteria. FEMS Microbiol. Lett. 2010, 309, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Papaleo, M.C.; Fondi, M.; Maida, I.; Perrin, E.; Lo Giudice, A.; Michaud, L.; Mangano, S.; Bartolucci, G.; Romoli, R.; Fani, R. Sponge-associated microbial Antarctic communities exhibiting antimicrobial activity against Burkholderia cepacia complex bacteria. Biotechnol. Adv. 2012, 30, 272–293. [Google Scholar] [CrossRef] [PubMed]
- Mangano, S.; Michaud, L.; Caruso, C.; Brilli, M.; Bruni, V.; Fani, R.; Lo Giudice, A. Antagonistic interactions among psychrotrophic cultivable bacteria isolated from Antarctic sponges: A preliminary analysis. Res. Microbiol. 2009, 160, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Verhoeven, J.T.P.; Dufour, S.C. Microbiomes of the Arctic carnivorous sponges Chondrocladiagrandis and Cladorhizaoxeata suggest a specific, but differential involvement of bacterial associates. Arctic Sci. 2017, 4, 186–204. [Google Scholar] [CrossRef]
- Rodríguez-Marconi, S.; De la Iglesia, R.; Díez, B.; Fonseca, C.A.; Hajdu, E.; Trefault, N. Characterization of bacterial, archaeal and eukaryote symbionts from Antarctic sponges reveals a high diversity at a three-domain level and a particular signature for this ecosystem. PLoS ONE 2015, 10, e0138837. [Google Scholar] [CrossRef] [PubMed]
- Mangano, S.; Michaud, L.; Caruso, C.; Lo Giudice, A. Metal and antibiotic-resistance in psychrotrophic bacteria associated with the Antarctic sponge Hemigellius pilosus (Kirkpatrick, 1907). Polar Biol. 2014, 37, 227–235. [Google Scholar] [CrossRef]
- Webster, N.S.; Negri, A.P.; Munro, M.M.H.G.; Battershill, C.N. Diverse microbial communities inhabit Antarctic sponges. Environ. Microbiol. 2004, 6, 288–300. [Google Scholar] [CrossRef] [PubMed]
- Xin, Y.; Kanagasabhapathy, M.; Janussen, D.; Xue, S.; Zhang, W. Phylogenetic diversity of Gram-positive bacteria cultured from Antarctic deep-sea sponges. Polar Biol. 2011, 34, 1501–1512. [Google Scholar] [CrossRef]
- Mangano, S.; Caruso, C.; Michaud, L.; Lo Giudice, A. First evidence of quorum sensing activity in bacteria associated with Antarctic sponges. Polar Biol. 2018, 41, 1435–1445. [Google Scholar] [CrossRef]
- González-Aravena, M.; Urtubia, R.; Del Campo, K.; Lavín, P.; Wong, C.M.V.L.; Cárdenas, C.A.; González-Rocha, G. Antibiotic and metal resistance of cultivable bacteria in the Antarctic sea urchin. Ant. Sci. 2016, 28, 261–268. [Google Scholar] [CrossRef]
- Webster, N.; Bourne, D. Bacterial community structure associated with the Antarctic soft coral, Alcyoniumantarcticum. FEMS Microbiol. Ecol. 2007, 59, 81–94. [Google Scholar] [CrossRef] [PubMed]
- Herrera, L.M.; García-Laviña, C.X.; Marizcurrena, J.J.; Volonterio, O.; Ponce de León, R.; Castro-Sowinski, S. Hydrolytic enzyme-producing microbes in the Antarctic oligochaete Grania sp. (Annelida). Polar Biol. 2017, 40, 947–953. [Google Scholar] [CrossRef]
- Lo Giudice, A.; Caruso, C.; Mangano, S.; Bruni, V.; De Domenico, M.; Michaud, L. Marine bacterioplankton diversity and community composition in an Antarctic coastal environment. Microb. Ecol. 2012, 63, 210–223. [Google Scholar] [CrossRef] [PubMed]
- Soldatou, S.; Baker, B.J. Cold-water marine natural products, 2006 to 2016. Nat. Prod. Rep. 2017, 34, 585–626. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, E.; Zilber-Rosenberg, I. Microbes drive evolution of animals and plants: The hologenome concept. MBio 2016, 7, e01395-15. [Google Scholar] [CrossRef] [PubMed]
- Díaz, A.; Gérard, K.; González-Wevar, C.; Maturana, C.; Féral, J.-P.; David, B.; Saucède, T.; Poulin, E. Genetic structure and demographic inference of the regular sea urchin Sterechinus neumayeri (Meissner, 1900) in the Southern Ocean: The role of the last glaciation. PLoS ONE 2018, 13, e0197611. [Google Scholar] [CrossRef] [PubMed]
- Peters, K.J.; Amsler, C.D.; McClintock, J.B.; Baker, B.J. Potential chemical defenses of Antarctic sponges against sympatric microorganisms. Polar Biol. 2010, 33, 649–658. [Google Scholar] [CrossRef]
- Piel, J. Metabolites from symbiotic bacteria. Nat. Prod. Rep. 2004, 21, 519–538. [Google Scholar] [CrossRef] [PubMed]
- Piel, J. Metabolites from symbiotic bacteria. Nat. Prod. Rep. 2009, 26, 338–362. [Google Scholar] [CrossRef] [PubMed]
- Piel, J.; Hui, D.; Wen, G.; Butzke, D.; Platzer, M.; Fusetani, N.; Matsunaga, S. Antitumor polyketide biosynthesis by an uncultivated bacterial symbiont of the marine sponge Theonella swinhoei. Proc. Natl. Acad. Sci. USA 2004, 101, 16222–16227. [Google Scholar] [CrossRef] [PubMed]
- Leal, M.C.; Puga, J.; Serodio, J.; Gomes, N.C.M.; Calado, R. Trends in the discovery of new marine natural products from invertebrates over the last two decades—Where and what are we bioprospecting? PLoS ONE 2012, 7, e30580. [Google Scholar] [CrossRef] [PubMed]
- Abbas, S.; Kelly, M.; Bowling, J.; Sims, J.; Waters, A.; Hamann, M. Advancement into the Arctic region for bioactive sponge secondary metabolites. Mar. Drugs 2011, 9, 2423–2437. [Google Scholar] [CrossRef] [PubMed]
- Slattery, M.; McClintock, J.B. Population-structure and feeding deterrence in 3 shallow-water Antarctic soft corals. Mar. Biol. 1995, 122, 461–470. [Google Scholar] [CrossRef]
- McClintock, J.; Baker, B.J. A review of the chemical ecology of Antarctic marine invertebrates. Am. Zool. 1997, 37, 329–342. [Google Scholar] [CrossRef]
- McClintock, J.; Gauthier, J.J. Antimicrobial activities of Antarctic sponges. Antarct. Sci. 1992, 4, 179–183. [Google Scholar] [CrossRef]
- Lebar, M.D.; Heimbegner, J.L.; Baker, B.J. Cold-water marine natural products. Nat. Prod. Rep. 2007, 24, 774–797. [Google Scholar] [CrossRef] [PubMed]
- Avila, C.; Taboada, S.; Nunez-Pons, L. Marine Antarctic chemical ecology: What is next? Mar. Ecol. 2008, 29, 1–70. [Google Scholar] [CrossRef]
- Figuerola, B.; Sala-Comorera, L.; Angulo-Preckler, C.; Vázquez, J.; Jesús Montes, M.; García-Aljaro, C.; Mercadé, E.; Blanch, A.R.; Avila, C. Antimicrobial activity of Antarctic bryozoans: An ecological perspective with potential for clinical applications. Mar. Environ. Res. 2014, 101, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Berne, S.; Kalauz, M.; Lapat, M.; Savin, L.; Janussen, D.; Kersken, D.; Ambrožič, J.; Špela, A.; Jokhadar, Z.; Jaklič, D.; et al. Screening of the Antarctic marine sponges (Porifera) as a source of bioactive compounds. Polar Biol. 2016, 39, 947–959. [Google Scholar] [CrossRef]
- Sacristán-Soriano, O.; Angulo-Preckler, C.; Vázquez, J.; Avila, C. Potential chemical defenses of Antarctic benthic organisms against marine bacteria. Polar Res. 2017, 36, 1390385. [Google Scholar] [CrossRef]
- Fuerst, J.A. Diversity and biotechnological potential of microorganisms associated with marine sponges. Appl. Microbiol. Biotechnol. 2014, 98, 7331–7347. [Google Scholar] [CrossRef] [PubMed]
- Blockley, A.; Elliott, D.R.; Roberts, A.P.; Sweet, M. Symbiotic microbes from marine invertebrates: Driving a new era of natural product drug discovery. Diversity 2017, 9, 49. [Google Scholar] [CrossRef]
- Maida, I.; Fondi, M.; Papaleo, M.C.; Perrin, E.; Orlandini, V.; Emiliani, G.; de Pascale, D.; Parrilli, E.; Tutino, M.L.; Michaud, L.; et al. Phenotypic and genomic characterization of the Antarctic bacterium Gillisia sp. CAL575, a producer of antimicrobial compounds. Extremophiles 2014, 18, 35–49. [Google Scholar] [CrossRef] [PubMed]
- Bosi, E.; Fondi, M.; Maida, I.; Perrin, E.; de Pascale, D.; Tutino, M.L.; Parrilli, E.; Lo Giudice, A.; Filloux, A.; Fani, R. Genome-scale phylogenetic and DNA composition analyses of Antarctic Pseudoalteromonas bacteria reveal inconsistencies in current taxonomic affiliation. Hydrobiologia 2015, 761, 85–95. [Google Scholar] [CrossRef]
- Maida, I.; Bosi, E.; Fondi, M.; Perrin, E.; Orlandini, V.; Papaleo, M.C.; Mengoni, A.; de Pascale, D.; Tutino, M.L.; Michaud, L.; et al. Antimicrobial activity of Pseudoalteromonas strains isolated from the Ross Sea (Antarctica) vs Cystic Fibrosis opportunistic pathogens. Hydrobiologia 2015, 761, 443–457. [Google Scholar] [CrossRef]
- Jayatilake, G.S.; Thornton, M.P.; Leonard, A.C.; Grimwade, J.E.; Baker, B.J. Metabolites from an Antarctic sponge-associated bacterium, Pseudomonas aeruginosa. J. Nat. Prod. 1996, 59, 293–296. [Google Scholar] [CrossRef] [PubMed]
- Romoli, R.; Papaleo, M.C.; de Pascale, D.; Tutino, M.L.; Michaud, L.; Lo Giudice, A.; Fani, R.; Bartolucci, G. Characterization of the volatile profile of Antarctic bacteriaby using solid-phase microextraction-gas chromatography-mass spectrometry. J. Mass. Spectr. 2011, 46, 1051–1059. [Google Scholar] [CrossRef] [PubMed]
- Romoli, R.; Papaleo, M.C.; de Pascale, D.; Tutino, M.L.; Michaud, L.; Lo Giudice, A.; Fani, R.; Bartolucci, G. GC–MS volatolomic approach to study the antimicrobial activity of the Antarctic bacterium Pseudoalteromonas sp.TB41. Metabolomics 2014, 10, 42–51. [Google Scholar] [CrossRef]
- Papaleo, M.C.; Romoli, R.; Bartolucci, G.; Maida, I.; Perrin, E.; Fondi, M.; Orlandini, V.; Mengoni, A.; Emiliani, G.; Tutino, M.L.; et al. Bioactive volatile organic compounds from Antarctic (sponges) bacteria. New Biotechnol. 2013, 30, 824–838. [Google Scholar] [CrossRef] [PubMed]
- Ansari, M.Z.; Yadav, G.; Gokhale, R.S.; Mohanty, D. NRPS-PKS: A knowledge-based resource for analysis of NRPS/PKS megasynthases. Nucleic Acids Res. 2004, 32, W405–W413. [Google Scholar] [CrossRef] [PubMed]
- Orlandini, V.; Maida, I.; Fondi, M.; Perrin, E.; Papaleo, M.C.; Bosi, E.; de Pascale, D.; Tutino, M.L.; Michaud, L.; Lo Giudice, A.; et al. Genomic analysis of three sponge-associated Arthrobacter Antarctic strains, inhibiting the growth of Burkholderia cepacia complex bacteria by synthesizing volatile organic compounds. Microbiol. Res. 2014, 169, 593–601. [Google Scholar] [CrossRef] [PubMed]
- Decho, A.W.; Gutierrez, T. Microbial extracellular polymeric substances (EPSs) in Ocean Systems. Front. Microbiol. 2017, 8, 92. [Google Scholar] [CrossRef] [PubMed]
- Caruso, C.; Rizzo, C.; Mangano, S.; Poli, A.; Di Donato, P.; Finore, I.; Nicolaus, B.; Di Marco, G.; Michaud, L.; Lo Giudice, A. Production and biotechnological potential of extracellular polymeric substances from sponge-associated Antarctic bacteria. Appl. Environ. Microbiol. 2018, 84, e01624-17. [Google Scholar] [CrossRef] [PubMed]
- Panwar, A.S.; Molpa, D.; Joshi, G.K. An overview of the biotechnological applications of bacterial cold active enzymes. Environ. Cons. J. 2015, 16, 59–66. [Google Scholar]
- Gurung, N.; Ray, S.; Bose, S.; Rai, V. A broader view: Microbial enzymes and their relevance in industries, medicine, and beyond. Biomed. Res. Int. 2013, 2013, 329121. [Google Scholar] [CrossRef] [PubMed]
- Marx, J.C.; Collins, T.; D’Amico, S.; Feller, G.; Gerday, C. Cold-adapted enzymes from marine Antarctic microorganisms. Mar. Biotechnol. 2007, 9, 293–304. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, V.C.; Satish, S.; Horam, S.; Raj, S.; Lal, A.; Arockiaraj, J.; Pasupuleti, M.; Dikshit, D.K. Natural products from polar organisms: Structural diversity, bioactivities and potential pharmaceutical applications. Polar Sci. 2018, in press. [Google Scholar] [CrossRef]
- Wilson, Z.E.; Brimble, M.A. Molecules derived from the extremes of life. Nat. Prod. Rep. 2009, 26, 44–71. [Google Scholar] [CrossRef] [PubMed]
- Turk, T.; Ambrožič, A.J.; Batista, U.; Strugar, G.; Kosmina, R.; Čivović, S.; Janussen, D.; Kauferstein, S.; Mebs, D.; Sepčić, K. Biological activities of ethanolic extracts from deep-sea Antarctic marine sponges. Mar. Drugs 2013, 11, 1126–1139. [Google Scholar] [CrossRef] [PubMed]
- Avila, C. Ecological and pharmacological activities of Antarctic marine natural products. Planta Med. 2016, 82, 767–774. [Google Scholar] [CrossRef] [PubMed]
- Guyomard, A.-I. Ethics and bioprospecting in Antarctica. Ethics Sci. Environ. Polit. 2010, 10, 31–44. [Google Scholar] [CrossRef]
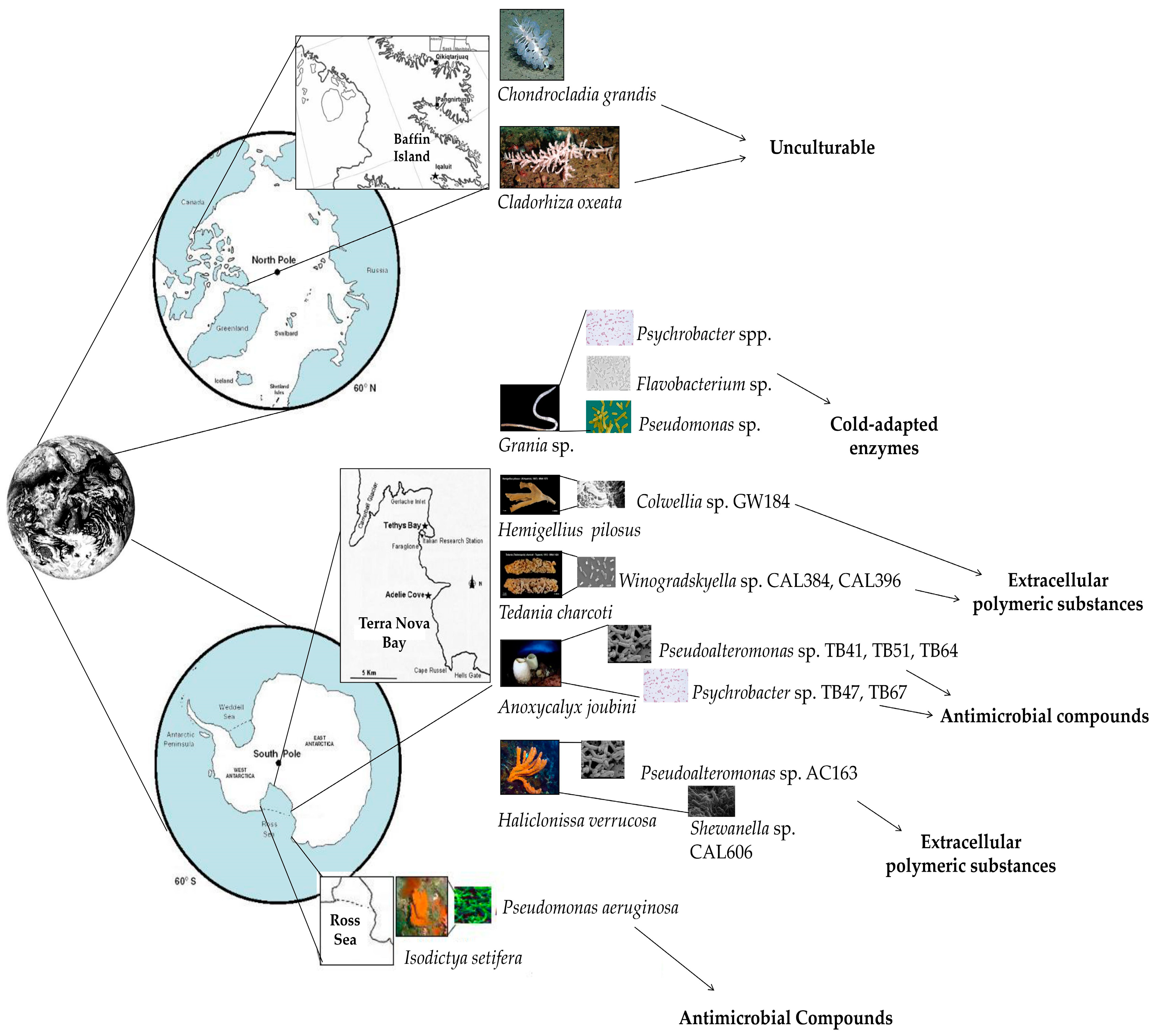
| Phylum | Invertebrate Host | Culturable/Unculturable Bacterial Community | Sampling Site | Reference(s) |
|---|---|---|---|---|
| Porifera | Anoxycalyx (Scolymastra) joubini | Culturable | Terra Nova Bay (Ross Sea, Antarctica) | [63,64] |
| Chondrocladia grandis | Unculturable | Baffin Island (Qikiqtaaluk Region, Nunavut, Canadian Arctic) | [65] | |
| Cladorhiza oxeata | Unculturable | Baffin Island (Qikiqtaaluk Region, Nunavut, Canadian Arctic) | [65] | |
| Clathria sp. | Unculturable | Fildes Bay (King George Island, South Shetlands, Antarctica) | [66] | |
| Haliclona (Gellius) sp. | Unculturable | Fildes Bay (King George Island, South Shetlands, Antarctica) | [66] | |
| Haliclonissa verrucosa | Culturable | Terra Nova Bay (Ross Sea, Antarctica) | [63] | |
| Hemigellius pilosus | Culturable | Terra Nova Bay (Ross Sea, Antarctica) | [67] | |
| Homaxinella balfourensis | Culturable/Unculturable | McMurdo Sound (Ross Sea, Antarctica); Weddell Sea (Antarctica) | [68,69] | |
| Hymeniacidon torquata | Unculturable | Fildes Bay (King George Island, South Shetlands, Antarctica) | [66] | |
| Kirkpatrickia variolosa | Unculturable | McMurdo Sound (Ross Sea, Antarctica); Fildes Bay (Antarctica) | [66,68] | |
| Latrunculia apicalis | Unculturable | McMurdo Sound (Ross Sea, Antarctica) | [68] | |
| Leucetta antarctica | Unculturable | Fildes Bay (King George Island, South Shetlands, Antarctica) | [66] | |
| Lissodendoryx (Ectyodoryx) nobilis | Culturable | Terra Nova Bay (Ross Sea, Antarctica) | [63,64,70] | |
| Megaciella annectens | Unculturable | Fildes Bay (King George Island, South Shetlands, Antarctica) | [66] | |
| Mycale acerata | Unculturable | McMurdo Sound (Ross Sea, Antarctica) | [68] | |
| Myxilla (Burtonanchora) sp. | Unculturable | Fildes Bay (King George Island, South Shetlands, Antarctica) | [66] | |
| Myxilla mollis | Culturable | Weddell Sea (Antarctica) | [69] | |
| Myxodoryx hanitschi | Culturable | Terra Nova Bay (Ross Sea, Antarctica) | [70] | |
| Phorbas glaberrimus | Culturable | Terra Nova Bay (Ross Sea, Antarctica) | [70] | |
| Radiella antarctica | Culturable | Weddell Sea (Antarctica) | [69] | |
| Rossella nuda | Culturable | Weddell Sea (Antarctica) | [69] | |
| Rossella racovitzae | Culturable | Weddell Sea (Antarctica) | [69] | |
| Sphaerotylus antarcticus | Unculturable | McMurdo Sound (Ross Sea, Antarctica) | [68] | |
| Echinodermata | Sterechinus neumayeri | Culturable | Maxwell Bay (King George Island, South Shetlands, Antarctica) | [71] |
| Cnidaria | Alcyonium antarcticum | Culturable/Unculturable | McMurdo Sound (Ross Sea, Antarctica) | [72] |
| Annelida | Grania sp. | Culturable | Maxwell Bay (King George Island, Antarctica) | [73] |
| Invertebrate Host (Species, Phylum) | Antarctic Location | Bacterial Isolate(s) | Bioactive Compound(s) Produced | Reference |
|---|---|---|---|---|
| Lissodendoryx nobilis,Porifera | Terra Nova Bay | Arthrobacter sp. TB23 | Antimicrobial compounds | [63] |
| Lissodendoryx nobilis, Porifera | Terra Nova Bay | Gillisia sp. CAL575 | [94] | |
| Haliclonissa verrucosa, Porifera | Terra Nova Bay | Pseudoalteromonas sp. AC163 | [95] | |
| Lissodendoryx nobilis, Porifera | Terra Nova Bay | Pseudoalteromonas sp. TB13 | [96] | |
| Lissodendoryx nobilis, Porifera | Terra Nova Bay | Pseudoalteromonas sp. TB25 | [96] | |
| Anoxycalyxjoubini, Porifera | Terra Nova Bay | Pseudoalteromonas sp. TB41 | [96] | |
| Anoxycalyx joubini, Porifera | Terra Nova Bay | Pseudoalteromonas sp. TB51 | [96] | |
| Anoxycalyx joubini, Porifera | Terra Nova Bay | Pseudoalteromonas sp. TB64 | [96] | |
| Isodictya setifera, Porifera | Ross Island | Pseudomonas aeruginosa | [97] | |
| Anoxycalyx joubini, Porifera | Terra Nova Bay | Psychrobacter sp. TB47 | [96] | |
| Anoxycalyx joubini, Porifera | Terra Nova Bay | Psychrobacter sp. TB67 | [96] | |
| Lissodendoryx nobilis, Porifera | Terra Nova Bay | Shewanella sp. TB4 | [98] | |
| Hemigellius pilosus, Porifera | Terra Nova Bay | Colwellia sp. GW185 | Extracellular polymeric substances | [46] |
| Haliclonissa verrucosa, Porifera | Terra Nova Bay | Shewanella sp. CAL606 | [46] | |
| Tedania charcoti, Porifera | Terra Nova Bay | Winogradskyella sp. CAL384 | [46] | |
| Tedania charcoti, Porifera | Terra Nova Bay | Winogradskyella sp. CAL3396 | [46] | |
| Grania sp., Annelida | Maxwell Bay | Flavobacterium spp. | Cold-adapted enzymes | [73] |
| Grania sp., Annelida | Maxwell Bay | Pseudomonas spp. | [73] | |
| Grania sp., Annelida | Maxwell Bay | Psychrobacter spp. | [73] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lo Giudice, A.; Rizzo, C. Bacteria Associated with Marine Benthic Invertebrates from Polar Environments: Unexplored Frontiers for Biodiscovery? Diversity 2018, 10, 80. https://doi.org/10.3390/d10030080
Lo Giudice A, Rizzo C. Bacteria Associated with Marine Benthic Invertebrates from Polar Environments: Unexplored Frontiers for Biodiscovery? Diversity. 2018; 10(3):80. https://doi.org/10.3390/d10030080
Chicago/Turabian StyleLo Giudice, Angelina, and Carmen Rizzo. 2018. "Bacteria Associated with Marine Benthic Invertebrates from Polar Environments: Unexplored Frontiers for Biodiscovery?" Diversity 10, no. 3: 80. https://doi.org/10.3390/d10030080
APA StyleLo Giudice, A., & Rizzo, C. (2018). Bacteria Associated with Marine Benthic Invertebrates from Polar Environments: Unexplored Frontiers for Biodiscovery? Diversity, 10(3), 80. https://doi.org/10.3390/d10030080






