Methods and Guidelines for Metabolism Studies: Applications to Cancer Research
Abstract
1. Introduction
2. Untargeted Metabolomics/Lipidomics
Drawbacks and Considerations for Untargeted Metabolomics/Lipidomics
3. Seahorse Real-Time Metabolic Flux Assays
Drawbacks and Considerations for Seahorse Real-Time Metabolic Flux Assays
4. Isotope Tracing and Metabolic Flux Analysis
Drawbacks and Considerations for Isotope Tracing and Metabolic Flux Analysis
5. Fluorescent Dyes
5.1. Glucose Import
5.2. Mitochondrial Function
5.3. Lipid Metabolism
5.4. Iron Metabolism
5.5. Drawbacks and Considerations for Fluorescent Dyes
| Metabolism | Fluorescent Dye | Mechanism of Action | Compatible with Fixation | Reference |
|---|---|---|---|---|
| Glucose | 2NBDG | Enters the cell via GLUT transporters and becomes phosphorylated | No | [162,163] |
| 6AzGal + BDP-DBCO | 6AzGal enters the cell via GLUT transporters → click-chemistry to tag fluorophore to 6AzGal | No | [168] | |
| Mitochondrial | Tetramethyl rhodamine methyl ester (TMRM) | Accumulates into mitochondria via negative membrane potential; sensitive to changes in membrane potential after staining | No | [180,187] |
| Tetramethyl rhodamine ethyl ester (TMRE) | Accumulates into mitochondria via negative membrane potential; sensitive to changes in membrane potential after staining | No | [180,240] | |
| JC-1 | Forms aggregates (red emission) in mitochondria with negative membrane potential; reverts back to monomeric state (green emission) upon mitochondrial depolarization | No | [190] | |
| MitoTracker Orange/Red/Deep Red | Accumulates into mitochondria via negative membrane potential and forms covalent bond with proteins within the mitochondrial matrix with a thiol-reactive functional group | Yes | [239] | |
| MitoTracker Green | Passive diffusion into mitochondria and forms covalent bond with proteins within mitochondrial matrix with a thiol-reactive functional group | No | [239] | |
| 10-N-nonyl acridine orange (NAO) | Binds to cardiolipin in the mitochondria | No | [183] | |
| Lipid | BODIPY 558/568 C12 | Diffuses through the plasma membrane and is trafficked or incorporated into various organelles. Mainly used to trace where fatty acids localize to in the cell | Yes | [204] |
| Nile Red | Solvatochromic dye that only becomes fluorescent in a hydrophobic environment, differential emission when bound to polar lipids (i.e., phospholipids) and neutral lipids (lipid droplets) | Yes | [208,211,212] | |
| Lipi-Blue/Red/Green | Only becomes fluorescent in hydrophobic environments through pyrene and perylene ring structures, but it is specific for neutral lipids in lipid droplets and no other lipid species | Yes | [210] | |
| BODIPY 493/503 | Diffuses through the plasma membrane and incorporates into lipid droplets by interacting with neutral lipids, which are most prevalent in lipid droplets | Yes | [209,210] | |
| Probe 10 | Coumarin attached to a fatty acid; when fully cleaved by all steps of the FAO pathway in the mitochondria, coumarin fluorescence is activated. Can be used to assess FAO flux by taking timelapse images and observing increasing fluorescence intensity over time | No | [215] | |
| Diphenylhexatriene (DPH) | Fluorescent lipid moiety that is quenched upon lipid peroxidation; increase in fluorescence decay indicates an increase in the rate of lipid peroxidation | No | [241] | |
| Liperfluo | Becomes fluorescent when oxidized by lipid hydroperoxides and peroxyl radicals | No | [219] | |
| BODIPY-C11 | Ratiometric dye to detect undamaged lipids and oxidized lipids. Fluorescence emission changes from red to green as lipid peroxidation occurs | Yes | [219,220,221,242] | |
| Iron | Calcein-AM | Fluorescence quenching when bound to Fe2+, Fe3+, Ni2+, Cu2+, Co2+ | No | [233,234] |
| Phen Green SK | Fluorescence quenching when bound to Fe2+, Fe3+, Ca2+, Zn2+ | No | [233,234] | |
| CP655 | Fluorescence quenching when bound to Fe2+, Fe3+, Cu2+ | No | [233,238] | |
| FerroOrange | Specifically binds to Fe2+ irreversibly. Does not react with Fe3+ or chelated iron | No | [235] | |
| BDP-Cy-Tpy | Ratiometric probe; when bound to Fe2+, fluorescence intensity of Cy-Tpy fluorophore is quenched while BDP fluorescence remains the same. Increase in ratio of BDP to Cy-Tpy intensity indicates the presence of Fe2+ | No | [236] | |
| RhoNox-1 | Not fluorescent in oxidized state, but exhibits rhodamine-based fluorescence when reduced by Fe2+ | No | [237] |
6. Genetically Encoded Fluorescent Biosensors
6.1. Glucose Sensors
6.2. Glycolysis Sensors
6.3. Acetyl-CoA Sensors
6.4. Redox Metabolism Sensors
6.5. Adenosine Triphosphate (ATP) Sensors
6.6. Drawbacks and Considerations for Genetically Encoded Fluorescent Biosensors
| Type of Sensor | Sensor Name | Fluorescent Protein(s) | Ratiometric, Intensiometric, or Lifetime; Method of Detection | Reference |
|---|---|---|---|---|
| Glucose | FLIPglu-600µ | eCFP & eYFP | Ratiometric; FRET | [254] |
| FLII12Pglu-700µδ6 | eCFP & Citrine-YFP | Ratiometric; FRET | [256] | |
| Green Glifon | Citrine-GFP | Intensiometric; fluorescence microscopy | [259] | |
| Red Glifon | mApple | Intensiometric; fluorescence microscopy | [260] | |
| qmTQ2-glucose | mTurquoise2 | Intensiometric with fluorescence microscopy; lifetime with FLIM | [253] | |
| FBP | HYlight | cpGFP | Intensiometric with fluorescence microscopy | [264] |
| Pyruvate | Pyronic | mTFP & Venus | Ratiometric; FRET | [269] |
| PyronicSF | cpGFP | Intensiometric with fluorescence microscopy | [270] | |
| Lactate | eLACCO1.1 | cpGFP | Intensiometric with fluorescence microscopy; ratiometric with two-photon microscopy | [272] |
| eLACCO2.1 | cpGFP | Intensiometric with fluorescence microscopy; ratiometric with two-photon microscopy | [273] | |
| R-iLACCO1 | cpmApple | Intensiometric with fluorescence microscopy; ratiometric with two-photon microscopy | [273] | |
| FiLa | cpYFP | Ratiometric; fluorescence microscopy | [276] | |
| FiLa-Red | cpmApple | Ratiometric; fluorescence microscopy | [275] | |
| LiLac | mTurquoise2 | Intensiometric with fluorescence microscopy; lifetime with FLIM | [244] | |
| Acetyl-CoA | PancACe | cpGFP | Ratiometric; fluorescence microscopy | [280] |
| NADH/NAD+ | Peredox | cpT-Sapphire & mCherry | Ratiometric: fluorescence microscopy | [286] |
| SoNar | cpYFP | Ratiometric; fluorescence microscopy | [290] | |
| Reduced/Oxidized Glutathione (GSH/GSSG) | Grx1-roGFP2 | roGFP | Ratiometric; fluorescence microscopy | [294] |
| Grx1-roGFP2.iL | roGFP | Ratiometric; fluorescence microscopy | [295] | |
| ATP | ATeam | mseCFP & cpmVenus | Ratiometric; FRET | [300] |
| iATPSnFR | cpGFP | Intensiometric with fluorescence microscopy | [301] | |
| PercevalHR | cpmVenus | Ratiometric; fluorescence microscopy | [304] |
7. Concluding Remarks
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| FRET | Förster resonance energy transfer |
| BRET | Bioluminescence resonance energy transfer |
| GSH | Reduced glutathione |
| GSSG | Oxidized glutathione |
| TMRE | Tetramethyl rhodamine ethyl ester |
| TMRM | Tetramethyl rhodamine methyl ester |
| LC | Liquid chromatography |
| GC | Gas chromatography |
| LC-MS | Liquid chromatography–mass spectrometry |
| GC-MS | Gas chromatography–mass spectrometry |
| MID | Mass isotopomer distribution |
| NMR | Nuclear magnetic resonance |
| 13C-MFA | 13C-metabolic flux analysis |
| INST-MFA | Isotopic non-stationary metabolic flux analysis |
| PLS-DA | Partial least squares discriminant analysis |
| ER+ | Estrogen receptor positive |
| ER− | Estrogen receptor negative |
| MSEA | Metabolite set enrichment analysis |
| OCR | Oxygen consumption rate |
| ECAR | Extracellular acidification rate |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| GSEA | Gene set enrichment analysis |
| FOLFIRINOX | Fluorouracil, leucovorin, irinotecan, and oxaliplatin |
| CA 19-9 | Carbohydrate antigen 19-9 |
| NSCLC | Non-small-cell lung cancer |
| PCA | Principal component analysis |
| FCCP | Carbonyl cyanide 4-(trifluoromethoxy)phenylhydrazone |
| ETC | Electron transport chain |
| 2-DG | 2-deoxyglucose |
| TNBC | Triple-negative breast cancer |
| 6-AN | 6-aminonicotinamide |
| TMPD | N,N,N’,N’-tetramethyl-p-phenylenediamine |
| TCA cycle | Tricarboxylic acid cycle |
| m/z | Mass to charge ratio |
| PPP | Pentose phosphate pathway |
| TKT | Transketolase |
| TALDO1 | Transaldolase |
| PC | Pyruvate carboxylase |
| PDH | Pyruvate dehydrogenase |
| VHL | Von Hippel–Lindau |
| ccRCC | Clear cell renal cell carcinoma |
| PET | Positron emission tomography |
| 18F-FDG | 18F-Fluorodeoxyglucose |
| 2-NBGD | 2-(N-(7-nitrobenz-2-oxa-1,3-diazol-4-yl)amino)-2-deoxyglucose |
| TME | Tumor microenvironment |
| CTC | Circulating tumor cell |
| MTG | MitoTracker Green |
| NAO | 10-N-nonyl acridine orange |
| PHB1 | Prohibitin-1 |
| 2D | 2-dimensional |
| 3D | 3-dimensional |
| FAO | Fatty acid oxidation |
| DPH | Diphenylhexatriene |
| Fe-S | Iron-sulfer |
| Fe2+ | Labile iron |
| Fe3+ | Ferric iron |
| GPX4 | Glutathione peroxidase 4 |
| Acetyl-CoA | Acetyl Coenzyme A |
| FLIM | Fluorescence lifetime imaging microscopy |
| CFP | Cyan fluorescent protein |
| YFP | Yellow fluorescent protein |
| MglB | Bacterial D-glucose-galactose binding periplasmic protein |
| CDK7 | Cyclin-dependent kinase 7 |
| GFP | Green fluorescent protein |
| MCT1 | Monocarboxylate transporter 1 |
| MCT4 | Monocarboxylate transporter 4 |
| NADH | Nicotinamide adenine dinucleotide (reduced) |
| NAD+ | Nicotinamde adenine dinucleotide (oxidized) |
| 3PG | 3-phosphoglycerate |
| CMV | Human cytomegalovirus |
| cpGFP | Circularly permuted GFP |
| cpYFP | Circularly permuted YFP |
| cpmVenus | Circularly permuted monomeric Venus |
| mseCFP | Monomeric super enhanced cyan fluorescent protein |
| cpmApple | Circular permuted monomeric Apple |
| DHAP | Dihydroxyacetone phosphate |
| APC/C | Anaphase promoting complex/cyclosome |
| MPC | Mitochondrial pyruvate carrier |
| PhdR | Pyruvate dehydrogenase complex repressor |
| mTFP | Monomeric teal fluorescent protein |
References
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cells 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Racker, E. Bioenergetics and the problem of tumor growth. Am. Sci. 1972, 60, 56–63. [Google Scholar] [PubMed]
- Warburg, O. Über den Stoffwechsel der Carcinomzelle. Naturwissenschaften 1924, 12, 1131–1137. [Google Scholar] [CrossRef]
- Warburg, O.; Minami, S. Versuche an Überlebendem Carcinom-gewebe. Klin. Wochenschr. 1923, 2, 776–777. [Google Scholar] [CrossRef]
- Vander Heiden, M.G.; Cantley, L.C.; Thompson, C.B. Understanding the Warburg effect: The metabolic requirements of cell proliferation. Science 2009, 324, 1029–1033. [Google Scholar] [CrossRef] [PubMed]
- Faubert, B.; Solmonson, A.; DeBerardinis, R.J. Metabolic reprogramming and cancer progression. Science 2020, 368, eaaw5473. [Google Scholar] [CrossRef]
- Shimoni-Sebag, A.; Abramovich, I.; Agranovich, B.; Massri, R.; Stossel, C.; Atias, D.; Raites-Gurevich, M.; Yizhak, K.; Golan, T.; Gottlieb, E.; et al. A metabolic switch to the pentose-phosphate pathway induces radiation resistance in pancreatic cancer. Radiother. Oncol. 2025, 202, 110606. [Google Scholar] [CrossRef]
- Morandi, A.; Indraccolo, S. Linking metabolic reprogramming to therapy resistance in cancer. Biochim. Biophys. Acta (BBA)—Rev. Cancer 2017, 1868, 1–6. [Google Scholar] [CrossRef]
- Liu, S.; Zhang, X.; Wang, W.; Li, X.; Sun, X.; Zhao, Y.; Wang, Q.; Li, Y.; Hu, F.; Ren, H. Metabolic reprogramming and therapeutic resistance in primary and metastatic breast cancer. Mol. Cancer 2024, 23, 261. [Google Scholar] [CrossRef]
- Tilsed, C.M.; Fisher, S.A.; Nowak, A.K.; Lake, R.A.; Lesterhuis, W.J. Cancer chemotherapy: Insights into cellular and tumor microenvironmental mechanisms of action. Front. Oncol. 2022, 12, 960317. [Google Scholar] [CrossRef]
- Li, M. Available online: https://BioRender.com/h3z5wfo (accessed on 17 July 2025).
- Buescher, J.M.; Antoniewicz, M.R.; Boros, L.G.; Burgess, S.C.; Brunengraber, H.; Clish, C.B.; DeBerardinis, R.J.; Feron, O.; Frezza, C.; Ghesquiere, B.; et al. A roadmap for interpreting 13C metabolite labeling patterns from cells. Curr. Opin. Biotechnol. 2015, 34, 189–201. [Google Scholar] [CrossRef]
- Jang, C.; Chen, L.; Rabinowitz, J.D. Metabolomics and Isotope Tracing. Cell 2018, 173, 822–827. [Google Scholar] [CrossRef]
- Danzi, F.; Pacchiana, R.; Mafficini, A.; Scupoli, M.T.; Scarpa, A.; Donadelli, M.; Fiore, A. To metabolomics and beyond: A technological portfolio to investigate cancer metabolism. Signal Transduct. Target. Ther. 2023, 8, 137. [Google Scholar] [CrossRef]
- Rusli, H.; Putri, R.M.; Alni, A. Recent Developments of Liquid Chromatography Stationary Phases for Compound Separation: From Proteins to Small Organic Compounds. Molecules 2022, 27, 907. [Google Scholar] [CrossRef] [PubMed]
- Piechocka, J.; Wieczorek, M.; Głowacki, R. Gas Chromatography-Mass Spectrometry Based Approach for the Determination of Methionine-Related Sulfur-Containing Compounds in Human Saliva. Int. J. Mol. Sci. 2020, 21, 9252. [Google Scholar] [CrossRef] [PubMed]
- Chong, J.; Soufan, O.; Li, C.; Caraus, I.; Li, S.; Bourque, G.; Wishart, D.S.; Xia, J. MetaboAnalyst 4.0: Towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 2018, 46, W486–W494. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Lin, C.-C.; Spasojevic, I.; Iversen, E.S.; Chi, J.-T.; Marks, J.R. A joint analysis of metabolomics and genetics of breast cancer. Breast Cancer Res. 2014, 16, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Furumichi, M.; Tanabe, M.; Sato, Y.; Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017, 45, D353–D361. [Google Scholar] [CrossRef]
- Chong, J.; Xia, J. MetaboAnalystR: An R package for flexible and reproducible analysis of metabolomics data. Bioinformatics 2018, 34, 4313–4314. [Google Scholar] [CrossRef]
- Benedetti, E.; Liu, E.M.; Tang, C.; Kuo, F.; Buyukozkan, M.; Park, T.; Park, J.; Correa, F.; Hakimi, A.A.; Intlekofer, A.M.; et al. A multimodal atlas of tumour metabolism reveals the architecture of gene–metabolite covariation. Nat. Metab. 2023, 5, 1029–1044. [Google Scholar] [CrossRef]
- Chen, Y.; Li, E.M.; Xu, L.Y. Guide to Metabolomics Analysis: A Bioinformatics Workflow. Metabolites 2022, 12, 357. [Google Scholar] [CrossRef] [PubMed]
- Amrutkar, M.; Guttorm, S.J.T.; Finstadsveen, A.V.; Labori, K.J.; Eide, L.; Rootwelt, H.; Elgstøen, K.B.P.; Gladhaug, I.P.; Verbeke, C.S. Global metabolomic profiling of tumor tissue and paired serum samples to identify biomarkers for response to neoadjuvant FOLFIRINOX treatment of human pancreatic cancer. Mol. Oncol. 2024, 19, 391–411. [Google Scholar] [CrossRef] [PubMed]
- Lyu, X.; Wang, Y.; Xu, Y.; Zhao, Z.; Liu, H.; Hu, Z. Metabolomic Profiling of Tumor Tissues Unveils Metabolic Shifts in Non-Small Cell Lung Cancer Patients with Concurrent Diabetes Mellitus. J. Proteome Res. 2024, 23, 3746–3753. [Google Scholar] [CrossRef] [PubMed]
- Moreno, P.; Jiménez-Jiménez, C.; Garrido-Rodríguez, M.; Calderón-Santiago, M.; Molina, S.; Lara-Chica, M.; Priego-Capote, F.; Salvatierra, Á.; Muñoz, E.; Calzado, M.A. Metabolomic profiling of human lung tumor tissues—Nucleotide metabolism as a candidate for therapeutic interventions and biomarkers. Mol. Oncol. 2018, 12, 1778–1796. [Google Scholar] [CrossRef]
- Nizioł, J.; Ossoliński, K.; Płaza-Altamer, A.; Kołodziej, A.; Ossolińska, A.; Ossoliński, T.; Nieczaj, A.; Ruman, T. Untargeted urinary metabolomics for bladder cancer biomarker screening with ultrahigh-resolution mass spectrometry. Sci. Rep. 2023, 13, 9802. [Google Scholar] [CrossRef] [PubMed]
- Nizioł, J.; Ossoliński, K.; Płaza-Altamer, A.; Kołodziej, A.; Ossolińska, A.; Ossoliński, T.; Ruman, T. Untargeted ultra-high-resolution mass spectrometry metabolomic profiling of blood serum in bladder cancer. Sci. Rep. 2022, 12, 15156. [Google Scholar] [CrossRef]
- Monteiro, M.S.; Carvalho, M.; Bastos, M.L.; Guedes de Pinho, P. Metabolomics analysis for biomarker discovery: Advances and challenges. Curr. Med. Chem. 2013, 20, 253–271. [Google Scholar] [CrossRef]
- Schrimpe-Rutledge, A.C.; Codreanu, S.G.; Sherrod, S.D.; McLean, J.A. Untargeted Metabolomics Strategies—Challenges and Emerging Directions. J. Am. Soc. Mass Spectrom. 2016, 27, 1897–1905. [Google Scholar] [CrossRef]
- Lardy, H.A.; Johnson, D.; McMurray, W.C.; Mc, M.W. Antibiotics as tools for metabolic studies. I. A survey of toxic antibiotics in respiratory, phosphorylative and glycolytic systems. Arch. Biochem. Biophys. 1958, 78, 587–597. [Google Scholar] [CrossRef]
- Divakaruni, A.S.; Paradyse, A.; Ferrick, D.A.; Murphy, A.N.; Jastroch, M. Chapter Sixteen—Analysis and Interpretation of Microplate-Based Oxygen Consumption and pH Data. In Methods in Enzymology; Murphy, A.N., Chan, D.C., Eds.; Academic Press: Cambridge, MA, USA, 2014; Volume 547, pp. 309–354. [Google Scholar]
- Divakaruni, A.S.; Jastroch, M. A practical guide for the analysis, standardization and interpretation of oxygen consumption measurements. Nat. Metab. 2022, 4, 978–994. [Google Scholar] [CrossRef]
- Benz, R.; McLaughlin, S. The molecular mechanism of action of the proton ionophore FCCP (carbonylcyanide p-trifluoromethoxyphenylhydrazone). Biophys. J. 1983, 41, 381–398. [Google Scholar] [CrossRef]
- Li, M. Available online: https://BioRender.com/ebh7b9c (accessed on 17 July 2025).
- Li, M. Available online: https://BioRender.com/t56aq0a (accessed on 17 July 2025).
- Li, M. Available online: https://BioRender.com/3izv8nl (accessed on 17 July 2025).
- Aft, R.L.; Zhang, F.W.; Gius, D. Evaluation of 2-deoxy-D-glucose as a chemotherapeutic agent: Mechanism of cell death. Br. J. Cancer 2002, 87, 805–812. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Ma, Y.; Zhang, Y.; Fu, B.; Wu, X.; Li, Q.; Cai, G.; Chen, X.; Bai, X.-Y. Low-dose 2-deoxyglucose and metformin synergically inhibit proliferation of human polycystic kidney cells by modulating glucose metabolism. Cell Death Discov. 2019, 5, 76. [Google Scholar] [CrossRef] [PubMed]
- Brunton, H.; Caligiuri, G.; Cunningham, R.; Upstill-Goddard, R.; Bailey, U.-M.; Garner, I.M.; Nourse, C.; Dreyer, S.; Jones, M.; Moran-Jones, K.; et al. HNF4A and GATA6 Loss Reveals Therapeutically Actionable Subtypes in Pancreatic Cancer. Cell Rep. 2020, 31, 107625. [Google Scholar] [CrossRef] [PubMed]
- Lanning, N.J.; Castle, J.P.; Singh, S.J.; Leon, A.N.; Tovar, E.A.; Sanghera, A.; MacKeigan, J.P.; Filipp, F.V.; Graveel, C.R. Metabolic profiling of triple-negative breast cancer cells reveals metabolic vulnerabilities. Cancer Metab. 2017, 5, 6. [Google Scholar] [CrossRef] [PubMed]
- Guha, M.; Srinivasan, S.; Raman, P.; Jiang, Y.; Kaufman, B.A.; Taylor, D.; Dong, D.; Chakrabarti, R.; Picard, M.; Carstens, R.P.; et al. Aggressive triple negative breast cancers have unique molecular signature on the basis of mitochondrial genetic and functional defects. Biochim. Biophys. Acta (BBA)—Mol. Basis Dis. 2018, 1864, 1060–1071. [Google Scholar] [CrossRef]
- Little, A.C.; Kovalenko, I.; Goo, L.E.; Hong, H.S.; Kerk, S.A.; Yates, J.A.; Purohit, V.; Lombard, D.B.; Merajver, S.D.; Lyssiotis, C.A. High-content fluorescence imaging with the metabolic flux assay reveals insights into mitochondrial properties and functions. Commun. Biol. 2020, 3, 271. [Google Scholar] [CrossRef]
- Xia, W.; Veeragandham, P.; Cao, Y.; Xu, Y.; Rhyne, T.E.; Qian, J.; Hung, C.-W.; Zhao, P.; Jones, Y.; Gao, H.; et al. Obesity causes mitochondrial fragmentation and dysfunction in white adipocytes due to RalA activation. Nat. Metab. 2024, 6, 273–289. [Google Scholar] [CrossRef]
- Desousa, B.R.; Kim, K.K.O.; Jones, A.E.; Ball, A.B.; Hsieh, W.Y.; Swain, P.; Morrow, D.H.; Brownstein, A.J.; Ferrick, D.A.; Shirihai, O.S.; et al. Calculation of ATP production rates using the Seahorse XF Analyzer. EMBO Rep. 2023, 24, e56380. [Google Scholar] [CrossRef]
- Street, J.C.; Mahmood, U.; Ballon, D.; Alfieri, A.A.; Koutcher, J.A. 13C and 31P NMR investigation of effect of 6-aminonicotinamide on metabolism of RIF-1 tumor cells in vitro. J. Biol. Chem. 1996, 271, 4113–4119. [Google Scholar] [CrossRef]
- Liberman, E.A.; Topaly, V.P.; Tsofina, L.M.; Jasaitis, A.A.; Skulachev, V.P. Mechanism of Coupling of Oxidative Phosphorylation and the Membrane Potential of Mitochondria. Nature 1969, 222, 1076–1078. [Google Scholar] [CrossRef]
- Trotta, A.P.; Gelles, J.D.; Serasinghe, M.N.; Loi, P.; Arbiser, J.L.; Chipuk, J.E. Disruption of mitochondrial electron transport chain function potentiates the pro-apoptotic effects of MAPK inhibition. J. Biol. Chem. 2017, 292, 11727–11739. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Wang, D.; Li, B.; Wang, J.; Xu, L.; Sun, X.; Ji, K.; Yan, C.; Liu, F.; Zhao, Y. Targeting DRP1 with Mdivi-1 to correct mitochondrial abnormalities in ADOA+ syndrome. JCI Insight 2024, 9, e180582. [Google Scholar] [CrossRef] [PubMed]
- Son, J.M.; Sarsour, E.H.; Balaraju, A.K.; Fussell, J.; Kalen, A.L.; Wagner, B.A.; Buettner, G.R.; Goswami, P.C. Mitofusin 1 and optic atrophy 1 shift metabolism to mitochondrial respiration during aging. Aging Cell 2017, 16, 1136–1145. [Google Scholar] [CrossRef]
- Thoudam, T.; Chanda, D.; Sinam, I.S.; Kim, B.-G.; Kim, M.-J.; Oh, C.J.; Lee, J.Y.; Kim, M.-J.; Park, S.Y.; Lee, S.Y.; et al. Noncanonical PDK4 action alters mitochondrial dynamics to affect the cellular respiratory status. Proc. Natl. Acad. Sci. USA 2022, 119, e2120157119. [Google Scholar] [CrossRef] [PubMed]
- Caines, J.K.; Barnes, D.A.; Berry, M.D. The Use of Seahorse XF Assays to Interrogate Real-Time Energy Metabolism in Cancer Cell Lines. In Cancer Cell Biology: Methods and Protocols; Christian, S.L., Ed.; Springer: New York, NY, USA, 2022; pp. 225–234. [Google Scholar]
- Gu, X.; Ma, Y.; Liu, Y.; Wan, Q. Measurement of mitochondrial respiration in adherent cells by Seahorse XF96 Cell Mito Stress Test. STAR Protoc. 2021, 2, 100245. [Google Scholar] [CrossRef]
- Yoo, I.; Ahn, I.; Lee, J.; Lee, N. Extracellular flux assay (Seahorse assay): Diverse applications in metabolic research across biological disciplines. Mol. Cells 2024, 47, 100095. [Google Scholar] [CrossRef]
- Campioni, G.; Pasquale, V.; Busti, S.; Ducci, G.; Sacco, E.; Vanoni, M. An Optimized Workflow for the Analysis of Metabolic Fluxes in Cancer Spheroids Using Seahorse Technology. Cells 2022, 11, 866. [Google Scholar] [CrossRef]
- Tidwell, T.R.; Røsland, G.V.; Tronstad, K.J.; Søreide, K.; Hagland, H.R. Metabolic flux analysis of 3D spheroids reveals significant differences in glucose metabolism from matched 2D cultures of colorectal cancer and pancreatic ductal adenocarcinoma cell lines. Cancer Metab. 2022, 10, 9. [Google Scholar] [CrossRef]
- Russell, S.; Wojtkowiak, J.; Neilson, A.; Gillies, R.J. Metabolic Profiling of healthy and cancerous tissues in 2D and 3D. Sci. Rep. 2017, 7, 15285. [Google Scholar] [CrossRef]
- Frederick, M.I.; Nassef, M.Z.; Borrelli, M.J.; Kuang, S.; Buensuceso, A.; More, T.; Cordes, T.; O’Donoghue, P.; Shepherd, T.G.; Hiller, K.; et al. Metabolic adaptation in epithelial ovarian cancer metastasis. Biochim. Biophys. Acta (BBA)—Mol. Basis Dis. 2024, 1870, 167312. [Google Scholar] [CrossRef]
- Compton, S.L.E.; Grieco, J.P.; Gollamudi, B.; Bae, E.; Van Mullekom, J.H.; Schmelz, E.M. Metabolic Reprogramming of Ovarian Cancer Spheroids during Adhesion. Cancers 2022, 14, 1399. [Google Scholar] [CrossRef] [PubMed]
- Buckley, A.M.; Dunne, M.R.; Morrissey, M.E.; Kennedy, S.A.; Nolan, A.; Davern, M.; Foley, E.K.; Clarke, N.; Lysaght, J.; Ravi, N.; et al. Real-time metabolic profiling of oesophageal tumours reveals an altered metabolic phenotype to different oxygen tensions and to treatment with Pyrazinib. Sci. Rep. 2020, 10, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Ludikhuize, M.C.; Meerlo, M.; Burgering, B.M.T.; Colman, M.J.R. Protocol to profile the bioenergetics of organoids using Seahorse. STAR Protoc. 2021, 2, 100386. [Google Scholar] [CrossRef] [PubMed]
- Underwood, E.; Redell, J.B.; Zhao, J.; Moore, A.N.; Dash, P.K. A method for assessing tissue respiration in anatomically defined brain regions. Sci. Rep. 2020, 10, 13179. [Google Scholar] [CrossRef]
- Jiang, K.; Nellissery, J.; Swaroop, A. Determination of Mitochondrial Respiration and Glycolysis in Ex Vivo Retinal Tissue Samples. J. Vis. Exp. 2021, 174, e62914. [Google Scholar] [CrossRef] [PubMed]
- Neville, K.E.; Bosse, T.L.; Klekos, M.; Mills, J.F.; Weicksel, S.E.; Waters, J.S.; Tipping, M. A novel ex vivo method for measuring whole brain metabolism in model systems. J. Neurosci. Methods 2018, 296, 32–43. [Google Scholar] [CrossRef] [PubMed]
- Mackert, O.; Wirth, E.K.; Sun, R.; Winkler, J.; Liu, A.; Renko, K.; Kunz, S.; Spranger, J.; Brachs, S. Impact of metabolic stress induced by diets, aging and fasting on tissue oxygen consumption. Mol. Metab. 2022, 64, 101563. [Google Scholar] [CrossRef] [PubMed]
- Kluza, J.; Peugnet, V.; Daunou, B.; Laine, W.; Kervoaze, G.; Rémy, G.; Loyens, A.; Maboudou, P.; Fovez, Q.; Grangette, C.; et al. A New Strategy to Preserve and Assess Oxygen Consumption in Murine Tissues. Int. J. Mol. Sci. 2021, 23, 109. [Google Scholar] [CrossRef] [PubMed]
- Acin-Perez, R.; Benador, I.Y.; Petcherski, A.; Veliova, M.; Benavides, G.A.; Lagarrigue, S.; Caudal, A.; Vergnes, L.; Murphy, A.N.; Karamanlidis, G.; et al. A novel approach to measure mitochondrial respiration in frozen biological samples. EMBO J. 2020, 39, e104073. [Google Scholar] [CrossRef]
- Sarver, D.C.; Saqib, M.; Chen, F.; Wong, G.W. Mitochondrial respiration atlas reveals differential changes in mitochondrial function across sex and age. eLife 2024, 13, RP96926. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Reyes, I.; Chandel, N.S. Mitochondrial TCA cycle metabolites control physiology and disease. Nat. Commun. 2020, 11, 102. [Google Scholar] [CrossRef] [PubMed]
- Mookerjee, S.A.; Goncalves, R.L.S.; Gerencser, A.A.; Nicholls, D.G.; Brand, M.D. The contributions of respiration and glycolysis to extracellular acid production. Biochim. Biophys. Acta 2015, 1847, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Nicholas, D.; Proctor, E.A.; Raval, F.M.; Ip, B.C.; Habib, C.; Ritou, E.; Grammatopoulos, T.N.; Steenkamp, D.; Dooms, H.; Apovian, C.M.; et al. Advances in the quantification of mitochondrial function in primary human immune cells through extracellular flux analysis. PLOS ONE 2017, 12, e0170975. [Google Scholar] [CrossRef] [PubMed]
- Jaber, S.M.; Yadava, N.; Polster, B.M. Mapping mitochondrial respiratory chain deficiencies by respirometry: Beyond the Mito Stress Test. Exp. Neurol. 2020, 328, 113282. [Google Scholar] [CrossRef] [PubMed]
- Chacko, J.V.; Eliceiri, K.W. Autofluorescence lifetime imaging of cellular metabolism: Sensitivity toward cell density, pH, intracellular, and intercellular heterogeneity. Cytom. Part A 2018, 95, 56–69. [Google Scholar] [CrossRef]
- Jeremy, J.Y.; Ballard, S.A.; Naylor, A.M.; Miller, M.A.; Angelini, G.D. Effects of sildenafil, a type-5 cGMP phosphodiesterase inhibitor, and papaverine on cyclic GMP and cyclic AMP levels in the rabbit corpus cavernosum in vitro. Br. J. Urol. 1997, 79, 958–963. [Google Scholar] [CrossRef]
- Xu, Y.-F.; Zhao, X.; Glass, D.S.; Absalan, F.; Perlman, D.H.; Broach, J.R.; Rabinowitz, J.D. Regulation of Yeast Pyruvate Kinase by Ultrasensitive Allostery Independent of Phosphorylation. Mol. Cell 2012, 48, 52–62. [Google Scholar] [CrossRef]
- Lowry, O.H.; Carter, J.; Ward, J.B.; Glaser, L. The Effect of Carbon and Nitrogen Sources on the Level of Metabolic Intermediates in Escherichia coli. J. Biol. Chem. 1971, 246, 6511–6521. [Google Scholar] [CrossRef]
- Saborano, R.; Eraslan, Z.; Roberts, J.; Khanim, F.L.; Lalor, P.F.; Reed, M.A.C.; Günther, U.L. A framework for tracer-based metabolism in mammalian cells by NMR. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef]
- Costello, L.C.; Franklin, R.B. Prostate epithelial cells utilize glucose and aspartate as the carbon sources for net citrate production. Prostate 1989, 15, 335–342. [Google Scholar] [CrossRef] [PubMed]
- Icard, P.; Coquerel, A.; Wu, Z.; Gligorov, J.; Fuks, D.; Fournel, L.; Lincet, H.; Simula, L. Understanding the Central Role of Citrate in the Metabolism of Cancer Cells and Tumors: An Update. Int. J. Mol. Sci. 2021, 22, 6587. [Google Scholar] [CrossRef] [PubMed]
- Cappel, D.A.; Deja, S.; Duarte, J.A.G.; Kucejova, B.; Iñigo, M.; Fletcher, J.A.; Fu, X.; Berglund, E.D.; Liu, T.; Elmquist, J.K.; et al. Pyruvate-Carboxylase-Mediated Anaplerosis Promotes Antioxidant Capacity by Sustaining TCA Cycle and Redox Metabolism in Liver. Cell Metab. 2019, 29, 1291–1305.e8. [Google Scholar] [CrossRef] [PubMed]
- Rattigan, K.M.; Brabcova, Z.; Sarnello, D.; Zarou, M.; Roy, K.; Kwan, R.; de Beauchamp, L.; Dawson, A.; Ianniciello, A.; Khalaf, A.; et al. Pyruvate anaplerosis is a targetable vulnerability in persistent leukaemic stem cells. Nat. Commun. 2023, 14, 4634. [Google Scholar] [CrossRef] [PubMed]
- Sellers, K.; Fox, M.P.; Bousamra, M., 2nd; Slone, S.P.; Higashi, R.M.; Miller, D.M.; Wang, Y.; Yan, J.; Yuneva, M.O.; Deshpande, R.; et al. Pyruvate carboxylase is critical for non-small-cell lung cancer proliferation. J. Clin. Invest. 2015, 125, 687–698. [Google Scholar] [CrossRef] [PubMed]
- van Heijster, F.H.A.; Breukels, V.; Jansen, K.F.J.; Schalken, J.A.; Heerschap, A. Carbon sources and pathways for citrate secreted by human prostate cancer cells determined by NMR tracing and metabolic modeling. Proc. Natl. Acad. Sci. USA 2022, 119, e2024357119. [Google Scholar] [CrossRef]
- Metallo, C.M.; Gameiro, P.A.; Bell, E.L.; Mattaini, K.R.; Yang, J.; Hiller, K.; Jewell, C.M.; Johnson, Z.R.; Irvine, D.J.; Guarente, L.; et al. Reductive glutamine metabolism by IDH1 mediates lipogenesis under hypoxia. Nature 2012, 481, 380–384. [Google Scholar] [CrossRef] [PubMed]
- Mullen, A.R.; Wheaton, W.W.; Jin, E.S.; Chen, P.-H.; Sullivan, L.B.; Cheng, T.; Yang, Y.; Linehan, W.M.; Chandel, N.S.; DeBerardinis, R.J. Reductive carboxylation supports growth in tumour cells with defective mitochondria. Nature 2012, 481, 385–388. [Google Scholar] [CrossRef]
- Yoo, H.; Antoniewicz, M.R.; Stephanopoulos, G.; Kelleher, J.K. Quantifying reductive carboxylation flux of glutamine to lipid in a brown adipocyte cell line. J. Biol. Chem. 2008, 283, 20621–20627. [Google Scholar] [CrossRef]
- Panic, V.; Pearson, S.; Banks, J.; Tippetts, T.; Velasco-Silva, J.N.; Lee, S.; Simcox, J.; Geoghegan, G.; Bensard, C.L.; van Ry, T.; et al. Mitochondrial pyruvate carrier is required for optimal brown fat thermogenesis. eLife 2020, 9, 52558. [Google Scholar] [CrossRef]
- Britt, E.C.; Lika, J.; Giese, M.A.; Schoen, T.J.; Seim, G.L.; Huang, Z.; Lee, P.Y.; Huttenlocher, A.; Fan, J. Switching to the cyclic pentose phosphate pathway powers the oxidative burst in activated neutrophils. Nat. Metab. 2022, 4, 389–403. [Google Scholar] [CrossRef]
- Antoniewicz, M.R. A guide to 13C metabolic flux analysis for the cancer biologist. Exp. Mol. Med. 2018, 50, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.-S.; Dennis, C.; Naqvi, I.; Dailey, L.; Lorzadeh, A.; Ye, G.; Zaytouni, T.; Adler, A.; Hitchcock, D.S.; Lin, L.; et al. Ornithine aminotransferase supports polyamine synthesis in pancreatic cancer. Nature 2023, 616, 339–347. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Lee, H.M.; Cai, F.; Ko, B.; Yang, C.; Lieu, E.L.; Muhammad, N.; Rhyne, S.; Li, K.; Haloul, M.; et al. The hexosamine biosynthesis pathway is a targetable liability in KRAS/LKB1 mutant lung cancer. Nat. Metab. 2020, 2, 1401–1412. [Google Scholar] [CrossRef] [PubMed]
- Tran, D.H.; Kim, D.; Kesavan, R.; Brown, H.; Dey, T.; Soflaee, M.H.; Vu, H.S.; Tasdogan, A.; Guo, J.; Bezwada, D.; et al. De novo and salvage purine synthesis pathways across tissues and tumors. Cell 2024, 187, 3602–3618.e20. [Google Scholar] [CrossRef] [PubMed]
- Spinelli, J.B.; Yoon, H.; Ringel, A.E.; Jeanfavre, S.; Clish, C.B.; Haigis, M.C. Metabolic recycling of ammonia via glutamate dehydrogenase supports breast cancer biomass. Science 2017, 358, 941–946. [Google Scholar] [CrossRef] [PubMed]
- Lewis, C.A.; Parker, S.J.; Fiske, B.P.; McCloskey, D.; Gui, D.Y.; Green, C.R.; Vokes, N.I.; Feist, A.M.; Heiden, M.G.V.; Metallo, C.M. Tracing compartmentalized NADPH metabolism in the cytosol and mitochondria of mammalian cells. Mol. Cell 2014, 55, 253–263. [Google Scholar] [CrossRef] [PubMed]
- Li, M. Available online: https://BioRender.com/pzpxjht (accessed on 17 July 2025).
- Li, M. Available online: https://BioRender.com/mjqgkdz (accessed on 17 July 2025).
- Hilovsky, D.; Hartsell, J.; Young, J.D.; Liu, X. Stable Isotope Tracing Analysis in Cancer Research: Advancements and Challenges in Identifying Dysregulated Cancer Metabolism and Treatment Strategies. Metabolites 2024, 14, 318. [Google Scholar] [CrossRef]
- Moco, S. Studying Metabolism by NMR-Based Metabolomics. Front. Mol. Biosci. 2022, 9, 882487. [Google Scholar] [CrossRef]
- Lane, A.N.; Fan, T.W. NMR-based Stable Isotope Resolved Metabolomics in systems biochemistry. Arch Biochem. Biophys 2017, 628, 123–131. [Google Scholar] [CrossRef]
- Lin, P.; Lane, A.N.; Fan, T.W.M. NMR-Based Stable Isotope Tracing of Cancer Metabolism. In Clinical Metabolomics: Methods and Protocols; Giera, M., Sánchez-López, E., Eds.; Springer: New York, NY, USA, 2025; pp. 457–504. [Google Scholar]
- Midani, F.S.; Wynn, M.L.; Schnell, S. The importance of accurately correcting for the natural abundance of stable isotopes. Anal. Biochem. 2017, 520, 27–43. [Google Scholar] [CrossRef]
- Rosenblatt, J.; Chinkes, D.; Wolfe, M.; Wolfe, R.R. Stable isotope tracer analysis by GC-MS, including quantification of isotopomer effects. Am. J. Physiol. Metab. 1992, 263, E584–E596. [Google Scholar] [CrossRef]
- Moiz, B.; Sriram, G.; Clyne, A.M. Interpreting metabolic complexity via isotope-assisted metabolic flux analysis. Trends Biochem. Sci. 2023, 48, 553–567. [Google Scholar] [CrossRef] [PubMed]
- Heinrich, P.; Kohler, C.; Ellmann, L.; Kuerner, P.; Spang, R.; Oefner, P.J.; Dettmer, K. Correcting for natural isotope abundance and tracer impurity in MS-, MS/MS- and high-resolution-multiple-tracer-data from stable isotope labeling experiments with IsoCorrectoR. Sci. Rep. 2018, 8, 17910. [Google Scholar] [CrossRef] [PubMed]
- Jeong, H.; Yu, Y.; Johansson, H.J.; Schroeder, F.C.; Lehtiö, J.; Vacanti, N.M. Correcting for Naturally Occurring Mass Isotopologue Abundances in Stable-Isotope Tracing Experiments with PolyMID. Metabolites 2021, 11, 310. [Google Scholar] [CrossRef] [PubMed]
- Millard, P.; Delépine, B.; Guionnet, M.; Heuillet, M.; Bellvert, F.; Létisse, F. IsoCor: Isotope correction for high-resolution MS labeling experiments. Bioinformatics 2019, 35, 4484–4487. [Google Scholar] [CrossRef]
- Carreer, W.J.; Flight, R.M.; Moseley, H.N. A Computational Framework for High-Throughput Isotopic Natural Abundance Correction of Omics-Level Ultra-High Resolution FT-MS Datasets. Metabolites 2013, 3, 853–866. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Parsons, L.R.; Su, X. AccuCor2: Isotope natural abundance correction for dual-isotope tracer experiments. Mod. Pathol. 2021, 101, 1403–1410. [Google Scholar] [CrossRef] [PubMed]
- Selivanov, V.A.; Benito, A.; Miranda, A.; Aguilar, E.; Polat, I.H.; Centelles, J.J.; Jayaraman, A.; Lee, P.W.N.; Marin, S.; Cascante, M. MIDcor, an R-program for deciphering mass interferences in mass spectra of metabolites enriched in stable isotopes. BMC Bioinform. 2017, 18, 88. [Google Scholar] [CrossRef]
- Trefely, S.; Ashwell, P.; Snyder, N.W. FluxFix: Automatic isotopologue normalization for metabolic tracer analysis. BMC Bioinform. 2016, 17, 485. [Google Scholar] [CrossRef]
- Du, D.; Tan, L.; Wang, Y.; Peng, B.; Weinstein, J.N.; Wondisford, F.E.; Su, X.; Lorenzi, P.L. ElemCor: Accurate data analysis and enrichment calculation for high-resolution LC-MS stable isotope labeling experiments. BMC Bioinform. 2019, 20, 89. [Google Scholar] [CrossRef] [PubMed]
- Long, C.P.; Antoniewicz, M.R. High-resolution 13C metabolic flux analysis. Nat. Protoc. 2019, 14, 2856–2877. [Google Scholar] [CrossRef] [PubMed]
- Maiorano, F.; Ambrosino, L.; Guarracino, M.R. The MetaboX Library: Building Metabolic Networks from KEGG Database. In Bioinformatics and Biomedical Engineering; Springer International Publishing: Cham, Switzerland, 2015; pp. 565–576. [Google Scholar]
- Caspi, R.; Billington, R.; Keseler, I.M.; Kothari, A.; Krummenacker, M.; Midford, P.E.; Ong, W.K.; Paley, S.; Subhraveti, P.; Karp, P.D. The MetaCyc database of metabolic pathways and enzymes—A 2019 update. Nucleic Acids Res. 2020, 48, D445–D453. [Google Scholar] [CrossRef] [PubMed]
- Norsigian, C.J.; Pusarla, N.; McConn, J.L.; Yurkovich, J.T.; Dräger, A.; Palsson, B.O.; King, Z. BiGG Models 2020: Multi-strain genome-scale models and expansion across the phylogenetic tree. Nucleic Acids Res. 2020, 48, D402–D406. [Google Scholar] [CrossRef] [PubMed]
- TeSlaa, T.; Ralser, M.; Fan, J.; Rabinowitz, J.D. The pentose phosphate pathway in health and disease. Nat. Metab. 2023, 5, 1275–1289. [Google Scholar] [CrossRef]
- Shupletsov, M.S.; Golubeva, L.I.; Rubina, S.S.; Podvyaznikov, D.A.; Iwatani, S.; Mashko, S.V. OpenFLUX2: 13C-MFA modeling software package adjusted for the comprehensive analysis of single and parallel labeling experiments. Microb. Cell Factories 2014, 13, 152. [Google Scholar] [CrossRef]
- Weitzel, M.; Nöh, K.; Dalman, T.; Niedenführ, S.; Stute, B.; Wiechert, W. 13CFLUX2—High-performance software suite for 13C-metabolic flux analysis. Bioinformatics 2012, 29, 143–145. [Google Scholar] [CrossRef]
- Young, J.D. INCA: A computational platform for isotopically non-stationary metabolic flux analysis. Bioinformatics 2014, 30, 1333–1335. [Google Scholar] [CrossRef]
- Matsuda, F.; Maeda, K.; Taniguchi, T.; Kondo, Y.; Yatabe, F.; Okahashi, N.; Shimizu, H. mfapy: An open-source Python package for 13C-based metabolic flux analysis. Metab. Eng. Commun. 2021, 13, e00177. [Google Scholar] [CrossRef]
- Wu, C.; Guarnieri, M.; Xiong, W. FreeFlux: A Python Package for Time-Efficient Isotopically Nonstationary Metabolic Flux Analysis. ACS Synth. Biol. 2023, 12, 2707–2714. [Google Scholar] [CrossRef]
- Quek, L.-E.; Liu, M.; Joshi, S.; Turner, N. Fast exchange fluxes around the pyruvate node: A leaky cell model to explain the gain and loss of unlabelled and labelled metabolites in a tracer experiment. Cancer Metab. 2016, 4, 13. [Google Scholar] [CrossRef] [PubMed]
- Ahn, W.S.; Antoniewicz, M.R. Metabolic flux analysis of CHO cells at growth and non-growth phases using isotopic tracers and mass spectrometry. Metab. Eng. 2011, 13, 598–609. [Google Scholar] [CrossRef] [PubMed]
- Bosshart, P.D.; Kalbermatter, D.; Bonetti, S.; Fotiadis, D. Mechanistic basis of L-lactate transport in the SLC16 solute carrier family. Nat. Commun. 2019, 10, 2649. [Google Scholar] [CrossRef] [PubMed]
- Cheah, Y.E.; Young, J.D. Isotopically nonstationary metabolic flux analysis (INST-MFA): Putting theory into practice. Curr. Opin. Biotechnol. 2018, 54, 80–87. [Google Scholar] [CrossRef] [PubMed]
- Kajihata, S.; Furusawa, C.; Matsuda, F.; Shimizu, H. OpenMebius: An open source software for isotopically nonstationary 13C-based metabolic flux analysis. BioMed Res. Int. 2014, 2014, 627014. [Google Scholar] [CrossRef]
- Davidson, S.M.; Papagiannakopoulos, T.; Olenchock, B.A.; Heyman, J.E.; Keibler, M.A.; Luengo, A.; Bauer, M.R.; Jha, A.K.; O’Brien, J.P.; Pierce, K.A.; et al. Environment Impacts the Metabolic Dependencies of Ras-Driven Non-Small Cell Lung Cancer. Cell Metab. 2016, 23, 517–528. [Google Scholar] [CrossRef]
- Kaushik, A.K.; Tarangelo, A.; Boroughs, L.K.; Ragavan, M.; Zhang, Y.; Wu, C.-Y.; Li, X.; Ahumada, K.; Chiang, J.C.; Tcheuyap, V.T.; et al. In vivo characterization of glutamine metabolism identifies therapeutic targets in clear cell renal cell carcinoma. Sci. Adv. 2022, 8, eabp8293. [Google Scholar] [CrossRef]
- Bartman, C.R.; TeSlaa, T.; Rabinowitz, J.D. Quantitative flux analysis in mammals. Nat. Metab. 2021, 3, 896–908. [Google Scholar] [CrossRef]
- Faubert, B.; Tasdogan, A.; Morrison, S.J.; Mathews, T.P.; DeBerardinis, R.J. Stable isotope tracing to assess tumor metabolism in vivo. Nat. Protoc. 2021, 16, 5123–5145. [Google Scholar] [CrossRef]
- Mahher, E. An Investigation of Brain Tumor Metabolism in Patients Undergoing Surgical Resection. 2012. Clinical Trial: NCT01668082. University of Texas Southwestern Medical Center. Available online: https://clinicaltrials.gov/study/NCT01668082 (accessed on 14 May 2025).
- Dysregulation of Glutamine Utilization in the Pathogenesis of Multiple Myeloma. 2017. Clinical Trial: NCT03119883. Mayo Clinic. Available online: https://clinicaltrials.gov/study/NCT03119883 (accessed on 14 May 2025).
- Pilot Study To Investigate Targetable Metabolic Pathways Sustaining Triple Negative (TN) Breast Cancer and Associated Genomic Alterations. 2018. Clinical Trial: NCT03457779. Baylor Research Institute. Available online: https://clinicaltrials.gov/study/NCT03457779 (accessed on 14 May 2025).
- Deen, S. A Physiological Study of the Metabolism of Pyruvate in High Grade Serous Ovarian Cancer Using Hyperpolarised Carbon-13 Magnetic Resonance Spectroscopic Imaging. 2018. Clinical Trial: NCT03526809. Cambridge University Hospitals NHS Foundation Trust. Available online: https://clinicaltrials.gov/study/NCT03526809 (accessed on 14 May 2025).
- Margulis, V. An Investigation of Kidney and Urothelial Tumor Metabolism in Patients Undergoing Surgical Resection and/or Biopsy. 2020. Clinical Trial: NCT04623502. University of Texas Southwestern Medical Center. Available online: https://clinicaltrials.gov/study/NCT04623502 (accessed on 14 May 2025).
- Metabolic Profiling of Leukemic Cells Through Isotope Tracing in Patients With CLL. 2021. Clinical Trial: NCT04785989. University of Wisconsin, Madison. Available online: https://clinicaltrials.gov/study/NCT04785989 (accessed on 14 May 2025).
- Hochster, H. Pilot Study to Investigate Targetable Metabolic Pathways Sustaining Pancreatic Cancer and Associated Genomic Alterations. 2022. Clinical Trial: NCT05296421. Rutgers, The State University of New Jersey. Available online: https://clinicaltrials.gov/study/NCT05296421 (accessed on 14 May 2025).
- Omene, C. Pilot Study to Investigate the Metabolic Pathways in Hormone Receptor Positive/HER2 Negative Breast Cancer. 2023. Clinical Trial: NCT05736367. Rutgers, The State University of New Jersey. Available online: https://clinicaltrials.gov/study/NCT05736367 (accessed on 14 May 2025).
- Ghergurovich, J.M.; Lang, J.D.; Levin, M.K.; Briones, N.; Facista, S.J.; Mueller, C.; Cowan, A.J.; McBride, M.J.; Rodriguez, E.S.R.; Killian, A.; et al. Local production of lactate, ribose phosphate, and amino acids within human triple-negative breast cancer. Med 2021, 2, 736–754.e6. [Google Scholar] [CrossRef]
- Faubert, B.; Li, K.Y.; Cai, L.; Hensley, C.T.; Kim, J.; Zacharias, L.G.; Yang, C.; Do, Q.N.; Doucette, S.; Burguete, D.; et al. Lactate Metabolism in Human Lung Tumors. Cell 2017, 171, 358–371.e9. [Google Scholar] [CrossRef]
- Courtney, K.D.; Bezwada, D.; Mashimo, T.; Pichumani, K.; Vemireddy, V.; Funk, A.M.; Wimberly, J.; McNeil, S.S.; Kapur, P.; Lotan, Y.; et al. Isotope Tracing of Human Clear Cell Renal Cell Carcinomas Demonstrates Suppressed Glucose Oxidation In Vivo. Cell Metab. 2018, 28, 793–800.e2. [Google Scholar] [CrossRef]
- Kim, I.-Y.; Park, S.; Kim, Y.; Kim, H.-J.; Wolfe, R.R. Tracing metabolic flux in vivo: Basic model structures of tracer methodology. Exp. Mol. Med. 2022, 54, 1311–1322. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Yin, Y.; Li, J.; Wang, H.; Lv, W.; Gao, Y.; Wang, T.; Zhong, Y.; Zhou, Z.; Cai, Y.; et al. Global stable-isotope tracing metabolomics reveals system-wide metabolic alternations in aging Drosophila. Nat. Commun. 2022, 13, 3518. [Google Scholar] [CrossRef] [PubMed]
- Grankvist, N.; Jönsson, C.; Hedin, K.; Sundqvist, N.; Sandström, P.; Björnsson, B.; Begzati, A.; Mickols, E.; Artursson, P.; Jain, M.; et al. Global 13C tracing and metabolic flux analysis of intact human liver tissue ex vivo. Nat. Metab. 2024, 6, 1963–1975. [Google Scholar] [CrossRef] [PubMed]
- Gebreselassie, N.A.; Antoniewicz, M.R. 13C-metabolic flux analysis of co-cultures: A novel approach. Metab. Eng. 2015, 31, 132–139. [Google Scholar] [CrossRef]
- Kang, S.; Antoniewicz, M.R.; Hay, N. Metabolic and transcriptomic reprogramming during contact inhibition-induced quiescence is mediated by YAP-dependent and YAP-independent mechanisms. Nat. Commun. 2024, 15, 6777. [Google Scholar] [CrossRef]
- Oates, E.H.; Antoniewicz, M.R. 13C-metabolic flux analysis of 3T3-L1 adipocytes illuminates its core metabolism under hypoxia. Metab. Eng. 2023, 76, 158–166. [Google Scholar] [CrossRef]
- Backman, T.W.H.; Schenk, C.; Radivojevic, T.; Ando, D.; Singh, J.; Czajka, J.J.; Costello, Z.; Keasling, J.D.; Tang, Y.; Akhmatskaya, E.; et al. BayFlux: A Bayesian method to quantify metabolic Fluxes and their uncertainty at the genome scale. PLOS Comput. Biol. 2023, 19, e1011111. [Google Scholar] [CrossRef]
- Hogg, M.; Wolfschmitt, E.-M.; Wachter, U.; Zink, F.; Radermacher, P.; Vogt, J.A. Bayesian 13C-metabolic Flux Analysis of Parallel Tracer Experiments in Granulocytes: A Directional Shift within the Non-Oxidative Pentose Phosphate Pathway Supports Phagocytosis. Metabolites 2024, 14, 24. [Google Scholar] [CrossRef]
- Theorell, A.; Jadebeck, J.F.; Wiechert, W.; McFadden, J.; Nöh, K. Rethinking 13C-metabolic flux analysis—The Bayesian way of flux inference. Metab. Eng. 2024, 83, 137–149. [Google Scholar] [CrossRef]
- Theorell, A.; Leweke, S.; Wiechert, W.; Nöh, K. To be certain about the uncertainty: Bayesian statistics for 13C metabolic flux analysis. Biotechnol. Bioeng. 2017, 114, 2668–2684. [Google Scholar] [CrossRef]
- Specht, E.A.; Braselmann, E.; Palmer, A.E. A Critical and Comparative Review of Fluorescent Tools for Live-Cell Imaging. Annu. Rev. Physiol. 2017, 79, 93–117. [Google Scholar] [CrossRef] [PubMed]
- Nikolova, K.; Kaloyanova, S.; Mihaylova, N.; Stoitsova, S.; Chausheva, S.; Vasilev, A.; Lesev, N.; Dimitrova, P.; Deligeorgiev, T.; Tchorbanov, A. New fluorogenic dyes for analysis of cellular processes by flow cytometry and confocal microscopy. J. Photochem. Photobiol. B Biol. 2013, 129, 125–134. [Google Scholar] [CrossRef] [PubMed]
- Robinson, J.P.; Ostafe, R.; Iyengar, S.N.; Rajwa, B.; Fischer, R. Flow Cytometry: The Next Revolution. Cells 2023, 12, 1875. [Google Scholar] [CrossRef] [PubMed]
- McGranahan, N.; Swanton, C. Clonal Heterogeneity and Tumor Evolution: Past, Present, and the Future. Cell 2017, 168, 613–628. [Google Scholar] [CrossRef] [PubMed]
- Semba, T.; Ishimoto, T. Spatial analysis by current multiplexed imaging technologies for the molecular characterisation of cancer tissues. Br. J. Cancer 2024, 131, 1737–1747. [Google Scholar] [CrossRef]
- Xiao, Z.; Dai, Z.; Locasale, J.W. Metabolic landscape of the tumor microenvironment at single cell resolution. Nat. Commun. 2019, 10, 3763. [Google Scholar] [CrossRef]
- Bar-Peled, L.; Kory, N. Principles and functions of metabolic compartmentalization. Nat. Metab. 2022, 4, 1232–1244. [Google Scholar] [CrossRef]
- Endo, K.; Oriuchi, N.; Higuchi, T.; Iida, Y.; Hanaoka, H.; Miyakubo, M.; Ishikita, T.; Koyama, K. PET and PET/CT using 18F-FDG in the diagnosis and management of cancer patients. Int. J. Clin. Oncol. 2006, 11, 286–296. [Google Scholar] [CrossRef]
- Flavahan, W.A.; Wu, Q.; Hitomi, M.; Rahim, N.; Kim, Y.; Sloan, A.E.; Weil, R.J.; Nakano, I.; Sarkaria, J.N.; Stringer, B.W.; et al. Brain tumor initiating cells adapt to restricted nutrition through preferential glucose uptake. Nat. Neurosci. 2013, 16, 1373–1382. [Google Scholar] [CrossRef]
- Masin, M.; Vazquez, J.; Rossi, S.; Groeneveld, S.; Samson, N.; Schwalie, P.C.; Deplancke, B.; Frawley, L.E.; Gouttenoire, J.; Moradpour, D.; et al. GLUT3 is induced during epithelial-mesenchymal transition and promotes tumor cell proliferation in non-small cell lung cancer. Cancer Metab. 2014, 2, 11. [Google Scholar] [CrossRef] [PubMed]
- Varghese, E.; Samuel, S.M.; Líšková, A.; Samec, M.; Kubatka, P.; Büsselberg, D. Targeting Glucose Metabolism to Overcome Resistance to Anticancer Chemotherapy in Breast Cancer. Cancers 2020, 12, 2252. [Google Scholar] [CrossRef] [PubMed]
- Yoshioka, K.; Takahashi, H.; Homma, T.; Saito, M.; Oh, K.B.; Nemoto, Y.; Matsuoka, H. A novel fluorescent derivative of glucose applicable to the assessment of glucose uptake activity of Escherichia coli. Biochim. Biophys. Acta (BBA)—Gen. Subj. 1996, 1289, 5–9. [Google Scholar] [CrossRef]
- Zou, C.; Wang, Y.; Shen, Z. 2-NBDG as a fluorescent indicator for direct glucose uptake measurement. J. Biochem. Biophys. Methods 2005, 64, 207–215. [Google Scholar] [CrossRef]
- Kim, W.H.; Lee, J.; Jung, D.-W.; Williams, D.R. Visualizing Sweetness: Increasingly Diverse Applications for Fluorescent-Tagged Glucose Bioprobes and Their Recent Structural Modifications. Sensors 2012, 12, 5005–5027. [Google Scholar] [CrossRef]
- Millon, S.R.; Ostrander, J.H.; Brown, J.Q.; Raheja, A.; Seewaldt, V.L.; Ramanujam, N. Uptake of 2-NBDG as a method to monitor therapy response in breast cancer cell lines. Breast Cancer Res. Treat. 2011, 126, 55–62. [Google Scholar] [CrossRef]
- Chang, C.H.; Qiu, J.; O’Sullivan, D.; Buck, M.D.; Noguchi, T.; Curtis, J.D.; Chen, Q.; Gindin, M.; Gubin, M.M.; van der Windt, G.J.; et al. Metabolic Competition in the Tumor Microenvironment Is a Driver of Cancer Progression. Cell 2015, 162, 1229–1241. [Google Scholar] [CrossRef]
- Cai, H.; Peng, F. 2-NBDG Fluorescence Imaging of Hypermetabolic Circulating Tumor Cells in Mouse Xenograft model of Breast Cancer. J. Fluoresc. 2013, 23, 213–220. [Google Scholar] [CrossRef]
- Tsuchiya, M.; Tachibana, N.; Hamachi, I. Post-click labeling enables highly accurate single cell analyses of glucose uptake ex vivo and in vivo. Commun. Biol. 2024, 7, 459. [Google Scholar] [CrossRef]
- De Jesus, A.; Keyhani-Nejad, F.; Pusec, C.M.; Goodman, L.; Geier, J.A.; Stoolman, J.S.; Stanczyk, P.J.; Nguyen, T.; Xu, K.; Suresh, K.V.; et al. Hexokinase 1 cellular localization regulates the metabolic fate of glucose. Mol. Cell 2022, 82, 1261–1277.e1269. [Google Scholar] [CrossRef] [PubMed]
- Bhagavan, N.V.; Ha, C.-E. Chapter 14—Carbohydrate Metabolism II: Gluconeogenesis, Glycogen Synthesis and Breakdown, and Alternative Pathways. In Essentials of Medical Biochemistry, 2nd ed.; Bhagavan, N.V., Ha, C.-E., Eds.; Academic Press: San Diego, CA, USA, 2015; pp. 205–225. [Google Scholar]
- Yazdani, S.; Bilan, P.J.; Jaldin-Fincati, J.R.; Pang, J.; Ceban, F.; Saran, E.; Brumell, J.H.; Freeman, S.A.; Klip, A. Dynamic glucose uptake, storage, and release by human microvascular endothelial cells. Mol. Biol. Cell 2022, 33, ar106. [Google Scholar] [CrossRef] [PubMed]
- Lane, R.H.; Crawford, S.E.; Flozak, A.S.; Simmons, R.A. Localization and quantification of glucose transporters in liver of growth-retarded fetal and neonatal rats. Am. J. Physiol. Metab. 1999, 276, E135–E142. [Google Scholar] [CrossRef] [PubMed]
- Sinclair, L.V.; Barthelemy, C.; Cantrell, D.A. Single Cell Glucose Uptake Assays: A Cautionary Tale. Immunometabolism 2020, 2, e200029. [Google Scholar] [CrossRef] [PubMed]
- Chakrabarty, R.P.; Chandel, N.S. Beyond ATP, new roles of mitochondria. Biochemist 2022, 44, 2–8. [Google Scholar] [CrossRef]
- Wallace, D.C. Mitochondria and cancer. Nat. Rev. Cancer 2012, 12, 685–698. [Google Scholar] [CrossRef]
- Vyas, S.; Zaganjor, E.; Haigis, M.C. Mitochondria and Cancer. Cell 2016, 166, 555–566. [Google Scholar] [CrossRef]
- Jin, P.; Jiang, J.; Zhou, L.; Huang, Z.; Nice, E.C.; Huang, C.; Fu, L. Mitochondrial adaptation in cancer drug resistance: Prevalence, mechanisms, and management. J. Hematol. Oncol. 2022, 15, 1–42. [Google Scholar] [CrossRef]
- Zorova, L.D.; Popkov, V.A.; Plotnikov, E.Y.; Silachev, D.N.; Pevzner, I.B.; Jankauskas, S.S.; Babenko, V.A.; Zorov, S.D.; Balakireva, A.V.; Juhaszova, M.; et al. Mitochondrial membrane potential. Anal. Biochem. 2018, 552, 50–59. [Google Scholar] [CrossRef]
- Nicholls, D.G.; Ward, M.W. Mitochondrial membrane potential and neuronal glutamate excitotoxicity: Mortality and millivolts. Trends Neurosci. 2000, 23, 166–174. [Google Scholar] [CrossRef]
- Gooz, M.; Maldonado, E.N. Fluorescence microscopy imaging of mitochondrial metabolism in cancer cells. Front. Oncol. 2023, 13, 1152553. [Google Scholar] [CrossRef] [PubMed]
- Pendergrass, W.; Wolf, N.; Poot, M. Efficacy of MitoTracker Green and CMXrosamine to measure changes in mitochondrial membrane potentials in living cells and tissues. Cytom. Part A 2004, 61A, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Mancini, M.; Anderson, B.O.; Caldwell, E.; Sedghinasab, M.; Paty, P.B.; Hockenbery, D.M. Mitochondrial proliferation and paradoxical membrane depolarization during terminal differentiation and apoptosis in a human colon carcinoma cell line. J. Cell Biol. 1997, 138, 449–469. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, M.E.; Azizuddin, K.; Zhang, P.; Chiu, S.-M.; Lam, M.; Kenney, M.E.; Burda, C.; Oleinick, N.L. Targeting of mitochondria by 10-N-alkyl acridine orange analogues: Role of alkyl chain length in determining cellular uptake and localization. Mitochondrion 2008, 8, 237–246. [Google Scholar] [CrossRef]
- Widlansky, M.E.; Wang, J.; Shenouda, S.M.; Hagen, T.M.; Smith, A.R.; Kizhakekuttu, T.J.; Kluge, M.A.; Weihrauch, D.; Gutterman, D.D.; Vita, J.A. Altered mitochondrial membrane potential, mass, and morphology in the mononuclear cells of humans with type 2 diabetes. Transl. Res. 2010, 156, 15–25. [Google Scholar] [CrossRef]
- Bonora, M.; Ito, K.; Morganti, C.; Pinton, P.; Ito, K. Membrane-potential compensation reveals mitochondrial volume expansion during HSC commitment. Exp. Hematol. 2018, 68, 30–37.e1. [Google Scholar] [CrossRef] [PubMed]
- Jacobson, J.; Duchen, M.R.; Heales, S.J.R. Intracellular distribution of the fluorescent dye nonyl acridine orange responds to the mitochondrial membrane potential: Implications for assays of cardiolipin and mitochondrial mass. J. Neurochem. 2002, 82, 224–233. [Google Scholar] [CrossRef] [PubMed]
- Desai, S.; Grefte, S.; van de Westerlo, E.; Lauwen, S.; Paters, A.; Prehn, J.H.M.; Gan, Z.; Keijer, J.; Adjobo-Hermans, M.J.W.; Koopman, W.J.H. Performance of TMRM and Mitotrackers in mitochondrial morphofunctional analysis of primary human skin fibroblasts. Biochim. Biophys. Acta (BBA)—Bioenerg. 2024, 1865, 149027. [Google Scholar] [CrossRef] [PubMed]
- Mot, A.I.; Liddell, J.R.; White, A.R.; Crouch, P.J. Circumventing the Crabtree Effect: A method to induce lactate consumption and increase oxidative phosphorylation in cell culture. Int. J. Biochem. Cell Biol. 2016, 79, 128–138. [Google Scholar] [CrossRef]
- Zhou, B.; Zhang, J.-y.; Liu, X.-s.; Chen, H.-z.; Ai, Y.-l.; Cheng, K.; Sun, R.-y.; Zhou, D.; Han, J.; Wu, Q. Tom20 senses iron-activated ROS signaling to promote melanoma cell pyroptosis. Cell Res. 2018, 28, 1171–1185. [Google Scholar] [CrossRef]
- Sivandzade, F.; Bhalerao, A.; Cucullo, L. Analysis of the Mitochondrial Membrane Potential Using the Cationic JC-1 Dye as a Sensitive Fluorescent Probe. Bio-Protocol 2019, 9, e3128. [Google Scholar] [CrossRef]
- Al-Zubaidi, U.; Liu, J.; Cinar, O.; Robker, R.L.; Adhikari, D.; Carroll, J. The spatio-temporal dynamics of mitochondrial membrane potential during oocyte maturation. Mol. Hum. Reprod. 2019, 25, 695–705. [Google Scholar] [CrossRef]
- Kuwahara, Y.; Tomita, K.; Roudkenar, M.H.; Roushandeh, A.M.; Urushihara, Y.; Igarashi, K.; Kurimasa, A.; Sato, T. Decreased mitochondrial membrane potential is an indicator of radioresistant cancer cells. Life Sci. 2021, 286, 120051. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekharan, A.; Varadarajan, S.N.; Lekshmi, A.; Lupitha, S.S.; Darvin, P.; Chandrasekhar, L.; Pillai, P.R.; Santhoshkumar, T.R.; Pillai, M.R. A high-throughput real-time in vitro assay using mitochondrial targeted roGFP for screening of drugs targeting mitochondria. Redox Biol. 2019, 20, 379–389. [Google Scholar] [CrossRef] [PubMed]
- Vianello, C.; Bello, F.D.; Shin, S.H.; Schiavon, S.; Bean, C.; Rebelo, A.P.M.; Knedlík, T.; Esfahani, E.N.; Costiniti, V.; Lacruz, R.S.; et al. High-Throughput Microscopy Analysis of Mitochondrial Membrane Potential in 2D and 3D Models. Cells 2023, 12, 1089. [Google Scholar] [CrossRef] [PubMed]
- Snaebjornsson, M.T.; Janaki-Raman, S.; Schulze, A. Greasing the Wheels of the Cancer Machine: The Role of Lipid Metabolism in Cancer. Cell Metab. 2020, 31, 62–76. [Google Scholar] [CrossRef] [PubMed]
- Broadfield, L.A.; Pane, A.A.; Talebi, A.; Swinnen, J.V.; Fendt, S.-M. Lipid metabolism in cancer: New perspectives and emerging mechanisms. Dev. Cell 2021, 56, 1363–1393. [Google Scholar] [CrossRef]
- Fu, Y.; Zou, T.; Shen, X.; Nelson, P.J.; Li, J.; Wu, C.; Yang, J.; Zheng, Y.; Bruns, C.; Zhao, Y.; et al. Lipid metabolism in cancer progression and therapeutic strategies. Medcomm 2021, 2, 27–59. [Google Scholar] [CrossRef]
- Bastiaanse, E.M.L.; Höld, K.M.; Van der Laarse, A. The effect of membrane cholesterol content on ion transport processes in plasma membranes. Cardiovasc. Res. 1997, 33, 272–283. [Google Scholar] [CrossRef]
- Casares, D.; Escribá, P.V.; Rosselló, C.A. Membrane Lipid Composition: Effect on Membrane and Organelle Structure, Function and Compartmentalization and Therapeutic Avenues. Int. J. Mol. Sci. 2019, 20, 2167. [Google Scholar] [CrossRef]
- Pöhnl, M.; Trollmann, M.F.W.; Böckmann, R.A. Nonuniversal impact of cholesterol on membranes mobility, curvature sensing and elasticity. Nat. Commun. 2023, 14, 8038. [Google Scholar] [CrossRef]
- Dierge, E.; Debock, E.; Guilbaud, C.; Corbet, C.; Mignolet, E.; Mignard, L.; Bastien, E.; Dessy, C.; Larondelle, Y.; Feron, O. Peroxidation of n-3 and n-6 polyunsaturated fatty acids in the acidic tumor environment leads to ferroptosis-mediated anticancer effects. Cell Metab. 2021, 33, 1701–1715.e5. [Google Scholar] [CrossRef] [PubMed]
- Jarc, E.; Kump, A.; Malavašič, P.; Eichmann, T.O.; Zimmermann, R.; Petan, T. Lipid droplets induced by secreted phospholipase A(2) and unsaturated fatty acids protect breast cancer cells from nutrient and lipotoxic stress. Biochim. Biophys. Acta (BBA)—Mol. Cell Biol. Lipids 2018, 1863, 247–265. [Google Scholar] [CrossRef] [PubMed]
- Zadoorian, A.; Du, X.; Yang, H. Lipid droplet biogenesis and functions in health and disease. Nat. Rev. Endocrinol. 2023, 19, 443–459. [Google Scholar] [CrossRef] [PubMed]
- Kajiwara, K.; Osaki, H.; Greßies, S.; Kuwata, K.; Kim, J.H.; Gensch, T.; Sato, Y.; Glorius, F.; Yamaguchi, S.; Taki, M. A negative-solvatochromic fluorescent probe for visualizing intracellular distributions of fatty acid metabolites. Nat. Commun. 2022, 13, 2533. [Google Scholar] [CrossRef]
- Kolahi, K.A.-O.; Louey, S.; Varlamov, O.; Thornburg, K. Real-Time Tracking of BODIPY-C12 Long-Chain Fatty Acid in Human Term Placenta Reveals Unique Lipid Dynamics in Cytotrophoblast Cells. PLOS ONE 2016, 11, e0153522. [Google Scholar] [CrossRef]
- Wang, H.; Wei, E.; Quiroga, A.D.; Sun, X.; Touret, N.; Lehner, R. Altered lipid droplet dynamics in hepatocytes lacking triacylglycerol hydrolase expression. Mol. Biol. Cell 2010, 21, 1991–2000. [Google Scholar] [CrossRef]
- Rambold, A.S.; Cohen, S.; Lippincott-Schwartz, J. Fatty Acid Trafficking in Starved Cells: Regulation by Lipid Droplet Lipolysis, Autophagy, and Mitochondrial Fusion Dynamics. Dev. Cell 2015, 32, 678–692, Correction in Dev. Cell 2015, 33, 489–490. [Google Scholar] [CrossRef]
- Greenspan, P.; Mayer, E.P.; Fowler, S.D. Nile red: A selective fluorescent stain for intracellular lipid droplets. J. Cell Biol. 1985, 100, 965–973. [Google Scholar] [CrossRef]
- Qiu, B.; Simon, M.C. BODIPY 493/503 Staining of Neutral Lipid Droplets for Microscopy and Quantification by Flow Cytometry. Bio-Protocol 2016, 6, e1912. [Google Scholar] [CrossRef]
- Tatenaka, Y.A.-O.; Kato, H.; Ishiyama, M.; Sasamoto, K.; Shiga, M.; Nishitoh, H.; Ueno, Y. Monitoring Lipid Droplet Dynamics in Living Cells by Using Fluorescent Probes. Biochemistry 2019, 58, 499–503. [Google Scholar] [CrossRef] [PubMed]
- Gajo, C.; Shchepanovska, D.; Jones, J.F.; Karras, G.; Malakar, P.; Greetham, G.M.; Hawkins, O.A.; Jordan, C.J.C.; Curchod, B.F.E.; Oliver, T.A.A. Nile Red Fluorescence: Where’s the Twist? J. Phys. Chem. B 2024, 128, 11768–11775. [Google Scholar] [CrossRef] [PubMed]
- Diaz, G.; Melis, M.; Batetta, B.; Angius, F.; Falchi, A.M. Hydrophobic characterization of intracellular lipids in situ by Nile Red red/yellow emission ratio. Micron 2008, 39, 819–824. [Google Scholar] [CrossRef] [PubMed]
- Kostecka, L.G.; Mendez, S.; Li, M.; Khare, P.; Zhang, C.; Le, A.; Amend, S.R.; Pienta, K.J. Cancer cells employ lipid droplets to survive toxic stress. Prostate 2024, 84, 644–655. [Google Scholar] [CrossRef]
- Pillai, S.; Mahmud, I.; Mahar, R.; Griffith, C.; Langsen, M.; Nguyen, J.; Wojtkowiak, J.W.; Swietach, P.; Gatenby, R.A.; Bui, M.M.; et al. Lipogenesis mediated by OGR1 regulates metabolic adaptation to acid stress in cancer cells via autophagy. Cell Rep. 2022, 39, 110796. [Google Scholar] [CrossRef]
- Uchinomiya, S.; Matsunaga, N.; Kamoda, K.; Kawagoe, R.; Tsuruta, A.; Ohdo, S.; Ojida, A. Fluorescence detection of metabolic activity of the fatty acid beta oxidation pathway in living cells. Chem. Commun. 2020, 56, 3023–3026. [Google Scholar] [CrossRef]
- Loftus, L.V.; Amend, S.R.; Pienta, K.J. Interplay between Cell Death and Cell Proliferation Reveals New Strategies for Cancer Therapy. Int. J. Mol. Sci. 2022, 23, 4723. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Liu, X.; Jin, S.; Chen, Y.; Guo, R. Ferroptosis in cancer therapy: A novel approach to reversing drug resistance. Mol. Cancer 2022, 21, 47. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Conrad, M. The Metabolic Underpinnings of Ferroptosis. Cell Metab. 2020, 32, 920–937. [Google Scholar] [CrossRef]
- Jeong, H.J.; Picou, C.; Jeong, K.; Chung, J.K. Oxidation Kinetics of Fluorescent Membrane Lipid Peroxidation Indicators. ACS Chem. Biol. 2024, 19, 1786–1793. [Google Scholar] [CrossRef]
- Pap, E.H.; Drummen, G.P.; Winter, V.J.; Kooij, T.W.; Rijken, P.; Wirtz, K.W.; Op den Kamp, J.A.; Hage, W.J.; Post, J.A. Ratio-fluorescence microscopy of lipid oxidation in living cells using C11-BODIPY(581/591). FEBS Lett. 1999, 453, 278–282. [Google Scholar] [CrossRef] [PubMed]
- Drummen, G.P.; Gadella, B.M.; Post, J.A.; Brouwers, J.F. Mass spectrometric characterization of the oxidation of the fluorescent lipid peroxidation reporter molecule C11-BODIPY(581/591). Free. Radic. Biol. Med. 2004, 36, 1635–1644. [Google Scholar] [CrossRef] [PubMed]
- Loftus, L.V.; Rolle, L.T.A.; Wang, B.; Pienta, K.J.; Amend, S.R. Dysregulation of Labile Iron Predisposes Chemotherapy Resistant Cancer Cells to Ferroptosis. Int. J. Mol. Sci. 2025, 26, 4193. [Google Scholar] [CrossRef] [PubMed]
- Zeng, W.; Zhang, R.; Huang, P.; Chen, M.; Chen, H.; Zeng, X.; Liu, J.; Zhang, J.; Huang, D.; Lao, L. Ferroptotic Neutrophils Induce Immunosuppression and Chemoresistance in Breast Cancer. Cancer Res. 2024, 85, 477–496. [Google Scholar] [CrossRef]
- Brown, R.A.M.; Richardson, K.L.; Kabir, T.D.; Trinder, D.; Ganss, R.; Leedman, P.J. Altered Iron Metabolism and Impact in Cancer Biology, Metastasis, and Immunology. Front. Oncol. 2020, 10, 476. [Google Scholar] [CrossRef]
- Wang, Y.A.; Yu, L.; Ding, J.; Chen, Y. Iron Metabolism in Cancer. Int. J. Mol. Sci. 2018, 20, 95. [Google Scholar] [CrossRef]
- Yu, Y.; Kovacevic, Z.; Richardson, D.R. Tuning cell cycle regulation with an iron key. Cell Cycle 2007, 6, 1982–1994. [Google Scholar] [CrossRef]
- Read, A.D.; Bentley, R.E.; Archer, S.L.; Dunham-Snary, K.J. Mitochondrial iron-sulfur clusters: Structure, function, and an emerging role in vascular biology. Redox Biol. 2021, 47, 102164. [Google Scholar] [CrossRef]
- Shi, R.; Hou, W.; Wang, Z.Q.; Xu, X. Biogenesis of Iron-Sulfur Clusters and Their Role in DNA Metabolism. Front. Cell Dev. Biol. 2021, 9, 735678. [Google Scholar] [CrossRef]
- Ibrahim, O.; O’Sullivan, J. Iron chelators in cancer therapy. BioMetals 2020, 33, 201–215. [Google Scholar] [CrossRef]
- Dixon, S.J.; Lemberg, K.M.; Lamprecht, M.R.; Skouta, R.; Zaitsev, E.M.; Gleason, C.E.; Patel, D.N.; Bauer, A.J.; Cantley, A.M.; Yang, W.S.; et al. Ferroptosis: An iron-dependent form of nonapoptotic cell death. Cell 2012, 149, 1060–1072. [Google Scholar] [CrossRef] [PubMed]
- Bauminger, E.R.; Harrison, P.M.; Hechel, D.; Nowik, I.; Treffry, A. Iron (III) can be transferred between ferritin molecules. Proc. R. Soc. B Biol. Sci. 1991, 244, 211–217. [Google Scholar]
- Chen, X.; Yu, C.; Kang, R.; Tang, D. Iron Metabolism in Ferroptosis. Front. Cell Dev. Biol. 2020, 8, 590226. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Abbate, V.; Hider, R.C. Iron-sensitive fluorescent probes: Monitoring intracellular iron pools. Metallomics 2015, 7, 212–222. [Google Scholar] [CrossRef] [PubMed]
- Kakhlon, O.; Cabantchik, Z.I. The labile iron pool: Characterization, measurement, and participation in cellular processes. This article is part of a series of reviews on “Iron and Cellular Redox Status”. The full list of papers may be found on the homepage of the journal. Free Radic. Biol. Med. 2002, 33, 1037–1046. [Google Scholar] [CrossRef]
- Yu, F.; Zhang, Q.; Liu, H.; Liu, J.; Yang, S.; Luo, X.; Liu, W.; Zheng, H.; Liu, Q.; Cui, Y.; et al. Dynamic O-GlcNAcylation coordinates ferritinophagy and mitophagy to activate ferroptosis. Cell Discov. 2022, 8, 40. [Google Scholar] [CrossRef]
- Li, P.; Fang, L.; Zhou, H.; Zhang, W.; Wang, X.; Li, N.; Zhong, H.; Tang, B. A new ratiometric fluorescent probe for detection of Fe2+ with high sensitivity and its intracellular imaging applications. Chem.—Eur. J. 2011, 17, 10520–10523. [Google Scholar] [CrossRef]
- Hirayama, T.; Okuda, K.; Nagasawa, H. A highly selective turn-on fluorescent probe for iron(ii) to visualize labile iron in living cells. Chem. Sci. 2013, 4, 1250–1256. [Google Scholar] [CrossRef]
- Ma, Y.; Liu, Z.; Hider, R.C.; Petrat, F. Determination of the labile iron pool of human lymphocytes using the fluorescent probe, CP655. Anal. Chem. Insights 2007, 2, 61–67. [Google Scholar] [CrossRef]
- Neikirk, K.; Marshall, A.G.; Kula, B.; Smith, N.; LeBlanc, S.; Hinton, A. MitoTracker: A useful tool in need of better alternatives. Eur. J. Cell Biol. 2023, 102, 151371. [Google Scholar] [CrossRef]
- Perry, S.W.; Norman, J.P.; Barbieri, J.; Brown, E.B.; Gelbard, H.A. Mitochondrial membrane potential probes and the proton gradient: A practical usage guide. BioTechniques 2011, 50, 98–115. [Google Scholar] [CrossRef]
- Mazière, J.C.; Routier, J.D.; Mazière, C.; Santus, R.; Patterson, L.K. Diphenylhexatriene (DPH)-Labeled Lipids as a Potential Tool for Studies on Lipid Peroxidation in Monolayer Films. Free. Radic. Biol. Med. 1997, 22, 795–802. [Google Scholar] [CrossRef]
- Alarcon-Gil, J.; Sierra-Magro, A.; Morales-Garcia, J.A.; Sanz-SanCristobal, M.; Alonso-Gil, S.; Cortes-Canteli, M.; Niso-Santano, M.; Martínez-Chacón, G.; Fuentes, J.M.; Santos, A.; et al. Neuroprotective and Anti-Inflammatory Effects of Linoleic Acid in Models of Parkinson’s Disease: The Implication of Lipid Droplets and Lipophagy. Cells 2022, 11, 2297. [Google Scholar] [CrossRef] [PubMed]
- Frei, M.S.; Mehta, S.; Zhang, J. Next-Generation Genetically Encoded Fluorescent Biosensors Illuminate Cell Signaling and Metabolism. Annu. Rev. Biophys. 2024, 53, 275–297. [Google Scholar] [CrossRef] [PubMed]
- Koveal, D.; Rosen, P.C.; Meyer, D.J.; Díaz-García, C.M.; Wang, Y.; Cai, L.-H.; Chou, P.J.; Weitz, D.A.; Yellen, G. A high-throughput multiparameter screen for accelerated development and optimization of soluble genetically encoded fluorescent biosensors. Nat. Commun. 2022, 13, 2919. [Google Scholar] [CrossRef] [PubMed]
- Choe, M.; Titov, D.V. Genetically encoded tools for measuring and manipulating metabolism. Nat. Chem. Biol. 2022, 18, 451–460. [Google Scholar] [CrossRef] [PubMed]
- Trefely, S.; Liu, J.; Huber, K.; Doan, M.T.; Jiang, H.; Singh, J.; von Krusenstiern, E.; Bostwick, A.; Xu, P.; Bogner-Strauss, J.G.; et al. Subcellular metabolic pathway kinetics are revealed by correcting for artifactual post harvest metabolism. Mol. Metab. 2019, 30, 61–71. [Google Scholar] [CrossRef] [PubMed]
- Greenwald, E.C.; Mehta, S.; Zhang, J. Genetically Encoded Fluorescent Biosensors Illuminate the Spatiotemporal Regulation of Signaling Networks. Chem. Rev. 2018, 118, 11707–11794. [Google Scholar] [CrossRef]
- Fang, C.; Huang, Y.; Zhao, Y. Review of FRET biosensing and its application in biomolecular detection. Am. J. Transl. Res. 2023, 15, 694–709. [Google Scholar]
- Nasu, Y.; Shen, Y.; Kramer, L.; Campbell, R.E. Structure- and mechanism-guided design of single fluorescent protein-based biosensors. Nat. Chem. Biol. 2021, 17, 509–518. [Google Scholar] [CrossRef]
- Goedhart, J.; Hink, M.A.; Jalink, K. An Introduction to Fluorescence Imaging Techniques Geared Towards Biosensor Applications. In Fluorescent Protein-Based Biosensors: Methods and Protocols; Zhang, J., Ni, Q., Newman, R.H., Eds.; Humana Press: Totowa, NJ, USA, 2014; pp. 17–28. [Google Scholar]
- Yellen, G.; Mongeon, R. Quantitative two-photon imaging of fluorescent biosensors. Curr. Opin. Chem. Biol. 2015, 27, 24–30. [Google Scholar] [CrossRef]
- Datta, R.; Heaster, T.M.; Sharick, J.T.; Gillette, A.A.; Skala, M.C. Fluorescence lifetime imaging microscopy: Fundamentals and advances in instrumentation, analysis, and applications. J. Biomed. Opt. 2020, 25, 071203. [Google Scholar] [CrossRef] [PubMed]
- Zhong, C.; Arai, S.; Okada, Y. Development of fluorescence lifetime biosensors for ATP, cAMP, citrate, and glucose using the mTurquoise2-based platform. Cell Rep. Methods 2024, 4, 100902. [Google Scholar] [CrossRef] [PubMed]
- Fehr, M.; Lalonde, S.; Lager, I.; Wolff, M.W.; Frommer, W.B. Imaging of the Dynamics of Glucose Uptake in the Cytosol of COS-7 Cells by Fluorescent Nanosensors. J. Biol. Chem. 2003, 278, 19127–19133. [Google Scholar] [CrossRef] [PubMed]
- John, S.A.; Ottolia, M.; Weiss, J.N.; Ribalet, B. Dynamic modulation of intracellular glucose imaged in single cells using a FRET-based glucose nanosensor. Pflügers Arch. —Eur. J. Physiol. 2008, 456, 307–322. [Google Scholar] [CrossRef]
- Takanaga, H.; Chaudhuri, B.; Frommer, W.B. GLUT1 and GLUT9 as major contributors to glucose influx in HepG2 cells identified by a high sensitivity intramolecular FRET glucose sensor. Biochim. Biophys. Acta (BBA)—Biomembr. 2008, 1778, 1091–1099. [Google Scholar] [CrossRef]
- Ghezzi, C.; Wong, A.; Chen, B.Y.; Ribalet, B.; Damoiseaux, R.; Clark, P.M. A high-throughput screen identifies that CDK7 activates glucose consumption in lung cancer cells. Nat. Commun. 2019, 10, 5444. [Google Scholar] [CrossRef] [PubMed]
- Kondo, H.; Ratcliffe, C.D.H.; Hooper, S.; Ellis, J.; MacRae, J.I.; Hennequart, M.; Dunsby, C.W.; Anderson, K.I.; Sahai, E. Single-cell resolved imaging reveals intra-tumor heterogeneity in glycolysis, transitions between metabolic states, and their regulatory mechanisms. Cell Rep. 2021, 34, 108750. [Google Scholar] [CrossRef] [PubMed]
- Mita, M.; Ito, M.; Harada, K.; Sugawara, I.; Ueda, H.; Tsuboi, T.; Kitaguchi, T. Green Fluorescent Protein-Based Glucose Indicators Report Glucose Dynamics in Living Cells. Anal. Chem. 2019, 91, 4821–4830. [Google Scholar] [CrossRef] [PubMed]
- Mita, M.; Sugawara, I.; Harada, K.; Ito, M.; Takizawa, M.; Ishida, K.; Ueda, H.; Kitaguchi, T.; Tsuboi, T. Development of red genetically encoded biosensor for visualization of intracellular glucose dynamics. Cell Chem. Biol. 2022, 29, 98–108.e4. [Google Scholar] [CrossRef]
- Vyas, N.K.; Vyas, M.N.; Quiocho, F.A. Sugar and Signal-Transducer Binding Sites of the Escherichia coli Galactose Chemoreceptor Protein. Science 1988, 242, 1290–1295. [Google Scholar] [CrossRef]
- Li, X.; Wen, X.; Tang, W.; Wang, C.; Chen, Y.; Yang, Y.; Zhang, Z.; Zhao, Y. Elucidating the spatiotemporal dynamics of glucose metabolism with genetically encoded fluorescent biosensors. Cell Rep. Methods 2024, 4, 100904. [Google Scholar] [CrossRef]
- Kierans, S.J.; Taylor, C.T. Glycolysis: A multifaceted metabolic pathway and signaling hub. J. Biol. Chem. 2024, 300, 107906. [Google Scholar] [CrossRef] [PubMed]
- Koberstein, J.N.; Stewart, M.L.; Smith, C.B.; Tarasov, A.I.; Ashcroft, F.M.; Stork, P.J.S.; Goodman, R.H. Monitoring glycolytic dynamics in single cells using a fluorescent biosensor for fructose 1,6-bisphosphate. Proc. Natl. Acad. Sci. USA 2022, 119, e2204407119. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Chávez, I.; Koberstein, J.N.; Pueyo, J.M.; Gilglioni, E.H.; Vertommen, D.; Baeyens, N.; Ezeriņa, D.; Gurzov, E.N.; Messens, J. Tracking fructose 1,6-bisphosphate dynamics in liver cancer cells using a fluorescent biosensor. iScience 2024, 27, 111336. [Google Scholar] [CrossRef] [PubMed]
- Park, J.O.; Rubin, S.A.; Xu, Y.-F.; Amador-Noguez, D.; Fan, J.; Shlomi, T.; Rabinowitz, J.D. Metabolite concentrations, fluxes and free energies imply efficient enzyme usage. Nat. Chem. Biol. 2016, 12, 482–489. [Google Scholar] [CrossRef]
- Paul, D.; Bolhuis, D.L.; Yan, H.; Das, S.; Xu, X.; Abbate, C.C.; Jenkins, L.M.M.; Emanuele, M.J.; Andresson, T.; Huang, J.; et al. Transient APC/C inactivation by mTOR boosts glycolysis during cell cycle entry. Nature 2025. [Google Scholar] [CrossRef]
- Gray, L.R.; Tompkins, S.C.; Taylor, E.B. Regulation of pyruvate metabolism and human disease. Cell. Mol. Life Sci. 2014, 71, 2577–2604. [Google Scholar] [CrossRef]
- Martín, A.S.; Ceballo, S.; Baeza-Lehnert, F.; Lerchundi, R.; Valdebenito, R.; Contreras-Baeza, Y.; Alegría, K.; Barros, L.F. Imaging Mitochondrial Flux in Single Cells with a FRET Sensor for Pyruvate. PLoS ONE 2014, 9, e85780. [Google Scholar] [CrossRef]
- Arce-Molina, R.; Cortés-Molina, F.; Sandoval, P.Y.; Galaz, A.; Alegría, K.; Schirmeier, S.; Barros, L.F.; Martín, A.S. A highly responsive pyruvate sensor reveals pathway-regulatory role of the mitochondrial pyruvate carrier MPC. eLife 2020, 9, e53917. [Google Scholar] [CrossRef]
- Lee, W.D.; Weilandt, D.R.; Liang, L.; MacArthur, M.R.; Jaiswal, N.; Ong, O.; Mann, C.G.; Chu, Q.; Hunter, C.J.; Ryseck, R.-P.; et al. Lactate homeostasis is maintained through regulation of glycolysis and lipolysis. Cell Metab. 2025, 37, 758–771.e8. [Google Scholar] [CrossRef] [PubMed]
- Nasu, Y.; Murphy-Royal, C.; Wen, Y.; Haidey, J.N.; Molina, R.S.; Aggarwal, A.; Zhang, S.; Kamijo, Y.; Paquet, M.-E.; Podgorski, K.; et al. A genetically encoded fluorescent biosensor for extracellular l-lactate. Nat. Commun. 2021, 12, 7058. [Google Scholar] [CrossRef] [PubMed]
- Nasu, Y.; Aggarwal, A.; Le, G.N.T.; Vo, C.T.; Kambe, Y.; Wang, X.; Beinlich, F.R.M.; Lee, A.B.; Ram, T.R.; Wang, F.; et al. Lactate biosensors for spectrally and spatially multiplexed fluorescence imaging. Nat. Commun. 2023, 14, 6598. [Google Scholar] [CrossRef] [PubMed]
- Anzai, T.; Kijima, K.; Fujimori, M.; Nakamoto, S.; Ishihama, A.; Shimada, T. Expanded roles of lactate-sensing LldR in transcription regulation of the Escherichia coli K-12 genome: Lactate utilisation and acid resistance. Microb. Genom. 2023, 9, 001015. [Google Scholar] [CrossRef]
- Chang, X.; Chen, X.; Zhang, X.; Chen, N.; Tang, W.; Zhang, Z.; Zheng, S.; Huang, J.; Ji, Y.; Zhao, Y.; et al. A bright red fluorescent genetically encoded sensor for lactate imaging. Biochem. Biophys. Res. Commun. 2024, 734, 150449. [Google Scholar] [CrossRef]
- Wang, A.; Zou, Y.; Liu, S.; Zhang, X.; Li, T.; Zhang, L.; Wang, R.; Xia, Y.; Li, X.; Zhang, Z.; et al. Comprehensive multiscale analysis of lactate metabolic dynamics in vitro and in vivo using highly responsive biosensors. Nat. Protoc. 2024, 19, 1311–1347. [Google Scholar] [CrossRef]
- He, W.; Li, Q.; Li, X. Acetyl-CoA regulates lipid metabolism and histone acetylation modification in cancer. Biochim. Biophys. Acta (BBA)—Rev. Cancer 2023, 1878, 188837. [Google Scholar] [CrossRef]
- Pietrocola, F.; Galluzzi, L.; Bravo-San Pedro, J.M.; Madeo, F.; Kroemer, G. Acetyl Coenzyme A: A Central Metabolite and Second Messenger. Cell Metab. 2015, 21, 805–821. [Google Scholar] [CrossRef]
- Shi, L.; Tu, B.P. Acetyl-CoA and the regulation of metabolism: Mechanisms and consequences. Curr. Opin. Cell Biol. 2015, 33, 125–131. [Google Scholar] [CrossRef]
- Smith, J.J.; Valentino, T.R.; Ablicki, A.H.; Banerjee, R.; Colligan, A.R.; Eckert, D.M.; Desjardins, G.A.; Diehl, K.L. A genetically encoded fluorescent biosensor for visualization of acetyl-CoA in live cells. Cell Chem. Biol. 2025, 32, 325–337.e10. [Google Scholar] [CrossRef]
- Monteiro, D.C.F.; Rugen, M.D.; Shepherd, D.; Nozaki, S.; Niki, H.; Webb, M.E. Formation of a heterooctameric complex between aspartate α-decarboxylase and its cognate activating factor, PanZ, is CoA-dependent. Biochem. Biophys. Res. Commun. 2012, 426, 350–355. [Google Scholar] [CrossRef]
- Lieberman, W.K.; Brown, Z.A.; Kantner, D.S.; Jing, Y.; Megill, E.; Evans, N.D.; Crawford, M.C.; Jhulki, I.; Grose, C.; Jones, J.E.; et al. Chemoproteomics Yields a Selective Molecular Host for Acetyl-CoA. J. Am. Chem. Soc. 2023, 145, 16899–16905. [Google Scholar] [CrossRef] [PubMed]
- Guertin, D.A.; Wellen, K.E. Acetyl-CoA metabolism in cancer. Nat. Rev. Cancer 2023, 23, 156–172. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Jiang, J.; Lei, Y.; Zhou, S.; Wei, Y.; Huang, C. Targeting Metabolic-Redox Circuits for Cancer Therapy. Trends Biochem. Sci. 2019, 44, 401–414. [Google Scholar] [CrossRef] [PubMed]
- Navas, L.E.; Carnero, A. NAD+ metabolism, stemness, the immune response, and cancer. Signal Transduct. Target. Ther. 2021, 6, 2. [Google Scholar] [CrossRef]
- Hung, Y.P.; Yellen, G. Live-cell imaging of cytosolic NADH-NAD+ redox state using a genetically encoded fluorescent biosensor. Methods Mol. Biol. 2014, 1071, 83–95. [Google Scholar]
- McLaughlin, K.J.; Strain-Damerell, C.M.; Xie, K.; Brekasis, D.; Soares, A.S.; Paget, M.S.B.; Kielkopf, C.L. Structural Basis for NADH/NAD+ Redox Sensing by a Rex Family Repressor. Mol. Cell 2010, 38, 563–575. [Google Scholar] [CrossRef]
- Sickmier, E.A.; Brekasis, D.; Paranawithana, S.; Bonanno, J.B.; Paget, M.S.B.; Burley, S.K.; Kielkopf, C.L. X-Ray Structure of a Rex-Family Repressor/NADH Complex Insights into the Mechanism of Redox Sensing. Structure 2005, 13, 43–54. [Google Scholar] [CrossRef]
- Birts, C.N.; Banerjee, A.; Darley, M.; Dunlop, C.R.; Nelson, S.; Nijjar, S.K.; Parker, R.; West, J.; Tavassoli, A.; Rose-Zerilli, M.J.J.; et al. p53 is regulated by aerobic glycolysis in cancer cells by the CtBP family of NADH-dependent transcriptional regulators. Sci. Signal. 2020, 13, eaau9529. [Google Scholar] [CrossRef]
- Zhao, Y.; Hu, Q.; Cheng, F.; Su, N.; Wang, A.; Zou, Y.; Hu, H.; Chen, X.; Zhou, H.M.; Huang, X.; et al. SoNar, a Highly Responsive NAD+/NADH Sensor, Allows High-Throughput Metabolic Screening of Anti-tumor Agents. Cell Metab. 2015, 21, 777–789. [Google Scholar] [CrossRef]
- Emmert, S.; Quargnali, G.; Thallmair, S.; Rivera-Fuentes, P. A locally activatable sensor for robust quantification of organellar glutathione. Nat. Chem. 2023, 15, 1415–1421. [Google Scholar] [CrossRef]
- Jozefczak, M.; Remans, T.; Vangronsveld, J.; Cuypers, A. Glutathione is a key player in metal-induced oxidative stress defenses. Int. J. Mol. Sci. 2012, 13, 3145–3175. [Google Scholar] [CrossRef]
- Li, F.J.; Long, H.Z.; Zhou, Z.W.; Luo, H.Y.; Xu, S.G.; Gao, L.C. System X(c) (-)/GSH/GPX4 axis: An important antioxidant system for the ferroptosis in drug-resistant solid tumor therapy. Front. Pharmacol. 2022, 13, 910292. [Google Scholar] [CrossRef] [PubMed]
- Gutscher, M.; Pauleau, A.-L.; Marty, L.; Brach, T.; Wabnitz, G.H.; Samstag, Y.; Meyer, A.J.; Dick, T.P. Real-time imaging of the intracellular glutathione redox potential. Nat. Methods 2008, 5, 553–559. [Google Scholar] [CrossRef] [PubMed]
- Aller, I.; Rouhier, N.; Meyer, A.J. Development of roGFP2-derived redox probes for measurement of the glutathione redox potential in the cytosol of severely glutathione-deficient rml1 seedlings. Front. Plant Sci. 2013, 4, 506. [Google Scholar] [CrossRef] [PubMed]
- Abbas, G.; Cui, M.; Wang, D.; Li, M.; Zhang, X.-E. Construction of Genetically Encoded Biosensors to Monitor Subcellular Compartment-Specific Glutathione Response to Chemotherapeutic Drugs in Acute Myeloid Leukemia Cells. Anal. Chem. 2023, 95, 2838–2847. [Google Scholar] [CrossRef]
- Dunn, J.; Grider, M.H. Physiology, Adenosine Triphosphate; BTI-StatPearls: Treasure Island, FL, USA, 2023. [Google Scholar]
- Parlani, M.; Jorgez, C.; Friedl, P. Plasticity of cancer invasion and energy metabolism. Trends Cell Biol. 2023, 33, 388–402. [Google Scholar] [CrossRef]
- Koundouros, N.; Poulogiannis, G. Reprogramming of fatty acid metabolism in cancer. Br. J. Cancer 2020, 122, 4–22. [Google Scholar] [CrossRef]
- Imamura, H.; Huynh Nhat, K.P.; Togawa, H.; Saito, K.; Iino, R.; Kato-Yamada, Y.; Nagai, T.; Noji, H. Visualization of ATP levels inside single living cells with fluorescence resonance energy transfer-based genetically encoded indicators. Proc. Natl. Acad. Sci. USA 2009, 106, 15651–15656. [Google Scholar] [CrossRef]
- Lobas, M.A.; Tao, R.; Nagai, J.; Kronschläger, M.T.; Borden, P.M.; Marvin, J.S.; Looger, L.L.; Khakh, B.S. A genetically encoded single-wavelength sensor for imaging cytosolic and cell surface ATP. Nat. Commun. 2019, 10, 711. [Google Scholar] [CrossRef]
- Zhan, H.; Pal, D.S.; Borleis, J.; Deng, Y.; Long, Y.; Janetopoulos, C.; Huang, C.-H.; Devreotes, P.N. Self-organizing glycolytic waves tune cellular metabolic states and fuel cancer progression. Nat. Commun. 2025, 16, 5563. [Google Scholar] [CrossRef]
- Maldonado, E.N.; Lemasters, J.J. ATP/ADP ratio, the missed connection between mitochondria and the Warburg effect. Mitochondrion 2014, 19, 78–84. [Google Scholar] [CrossRef] [PubMed]
- Tantama, M.; Martínez-François, J.R.; Mongeon, R.; Yellen, G. Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio. Nat. Commun. 2013, 4, 2550. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.; Hung, Y.P.; Yellen, G. A genetically encoded fluorescent reporter of ATP:ADP ratio. Nat. Methods 2009, 6, 161–166. [Google Scholar] [CrossRef]
- Zanotelli, M.R.; Zhang, J.; Ortiz, I.; Wang, W.; Chada, N.C.; Reinhart-King, C.A. Highly motile cells are metabolically responsive to collagen density. Proc. Natl. Acad. Sci. USA 2022, 119, e2114672119. [Google Scholar] [CrossRef]
- Zanotelli, M.R.; Goldblatt, Z.E.; Miller, J.P.; Bordeleau, F.; Li, J.; VanderBurgh, J.A.; Lampi, M.C.; King, M.R.; Reinhart-King, C.A. Regulation of ATP utilization during metastatic cell migration by collagen architecture. Mol. Biol. Cell 2018, 29, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Tsien, R.Y. THE GREEN FLUORESCENT PROTEIN. Annu. Rev. Biochem. 1998, 67, 509–544. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.N.; Mehta, S.; Zhang, J. Recent advances in the use of genetically encodable optical tools to elicit and monitor signaling events. Curr. Opin. Cell Biol. 2020, 63, 114–124. [Google Scholar] [CrossRef] [PubMed]
- Zaver, S.A.; Johnson, C.J.; Berndt, A.; Simpson, C.L. Live Imaging with Genetically Encoded Physiologic Sensors and Optogenetic Tools. J. Investig. Dermatol. 2023, 143, 353–361.e4. [Google Scholar] [CrossRef]
- Emiliani, V.; Entcheva, E.; Hedrich, R.; Hegemann, P.; Konrad, K.R.; Lüscher, C.; Mahn, M.; Pan, Z.-H.; Sims, R.R.; Vierock, J.; et al. Optogenetics for light control of biological systems. Nat. Rev. Methods Primers 2022, 2, 55. [Google Scholar] [CrossRef]
- Tkatch, T.; Greotti, E.; Baranauskas, G.; Pendin, D.; Roy, S.; Nita, L.I.; Wettmarshausen, J.; Prigge, M.; Yizhar, O.; Shirihai, O.S.; et al. Optogenetic control of mitochondrial metabolism and Ca2+ signaling by mitochondria-targeted opsins. Proc. Natl. Acad. Sci. USA 2017, 114, E5167–E5176. [Google Scholar] [CrossRef]
- Wollman, A.J.M.; Kioumourtzoglou, D.; Ward, R.; Gould, G.W.; Bryant, N.J. Large scale, single-cell FRET-based glucose uptake measurements within heterogeneous populations. iScience 2022, 25, 104023. [Google Scholar] [CrossRef]
- Roszik, J.; Szöllosi, J.; Vereb, G. AccPbFRET: An ImageJ plugin for semi-automatic, fully corrected analysis of acceptor photobleaching FRET images. BMC Bioinform. 2008, 9, 346. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Tsoy, Y.; Persson, J.; Grailhe, R. FLIM-FRET analyzer: Open source software for automation of lifetime-based FRET analysis. Source Code Biol. Med. 2017, 12, 7. [Google Scholar] [CrossRef]
- McQuin, C.; Goodman, A.; Chernyshev, V.; Kamentsky, L.; Cimini, B.A.; Karhohs, K.W.; Doan, M.; Ding, L.; Rafelski, S.M.; Thirstrup, D.; et al. CellProfiler 3.0: Next-generation image processing for biology. PLOS Biol. 2018, 16, e2005970. [Google Scholar] [CrossRef]
- Yang, W.H.; Qiu, Y.; Stamatatos, O.; Janowitz, T.; Lukey, M.J. Enhancing the Efficacy of Glutamine Metabolism Inhibitors in Cancer Therapy. Trends Cancer 2021, 7, 790–804. [Google Scholar] [CrossRef]
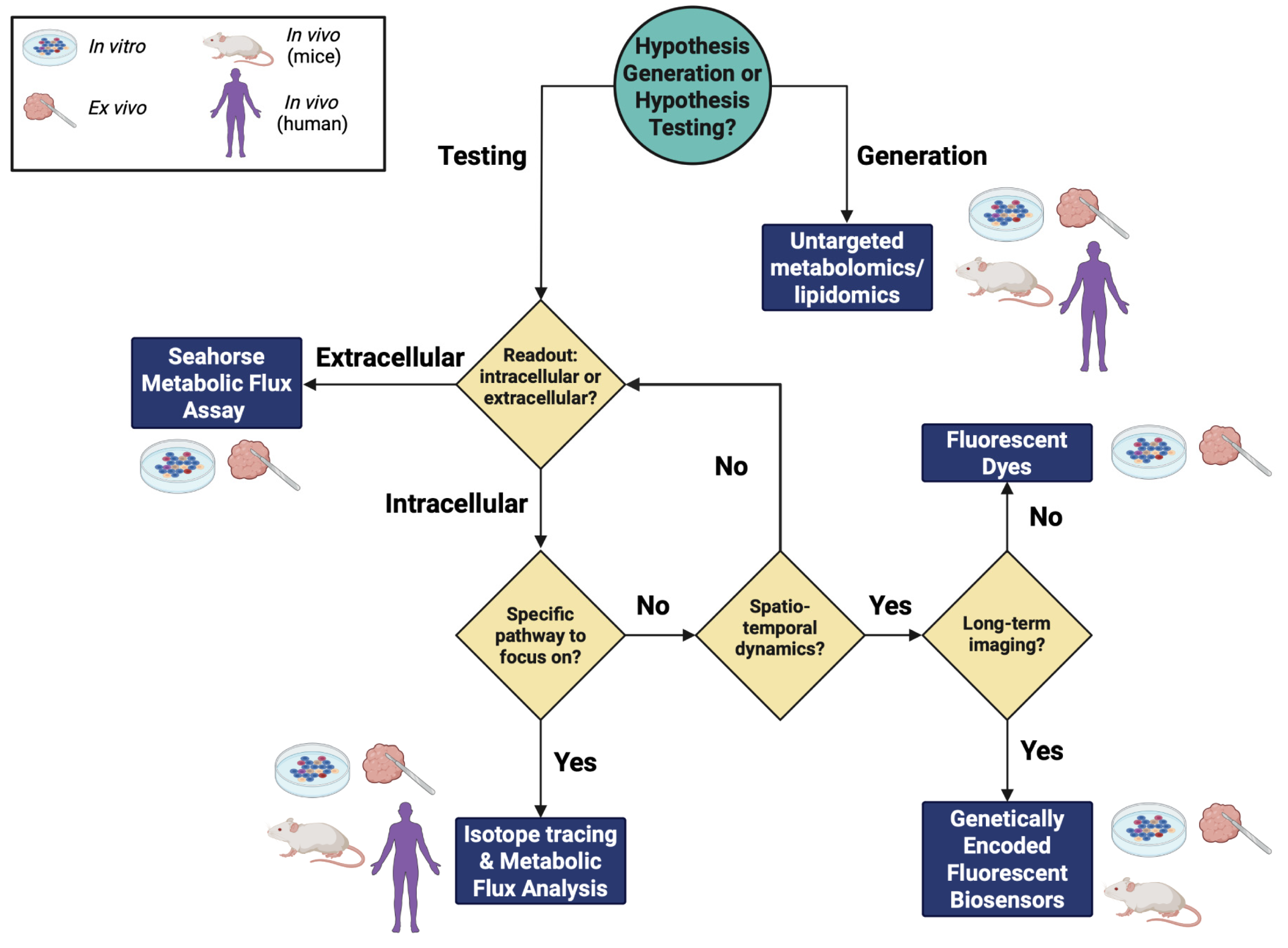
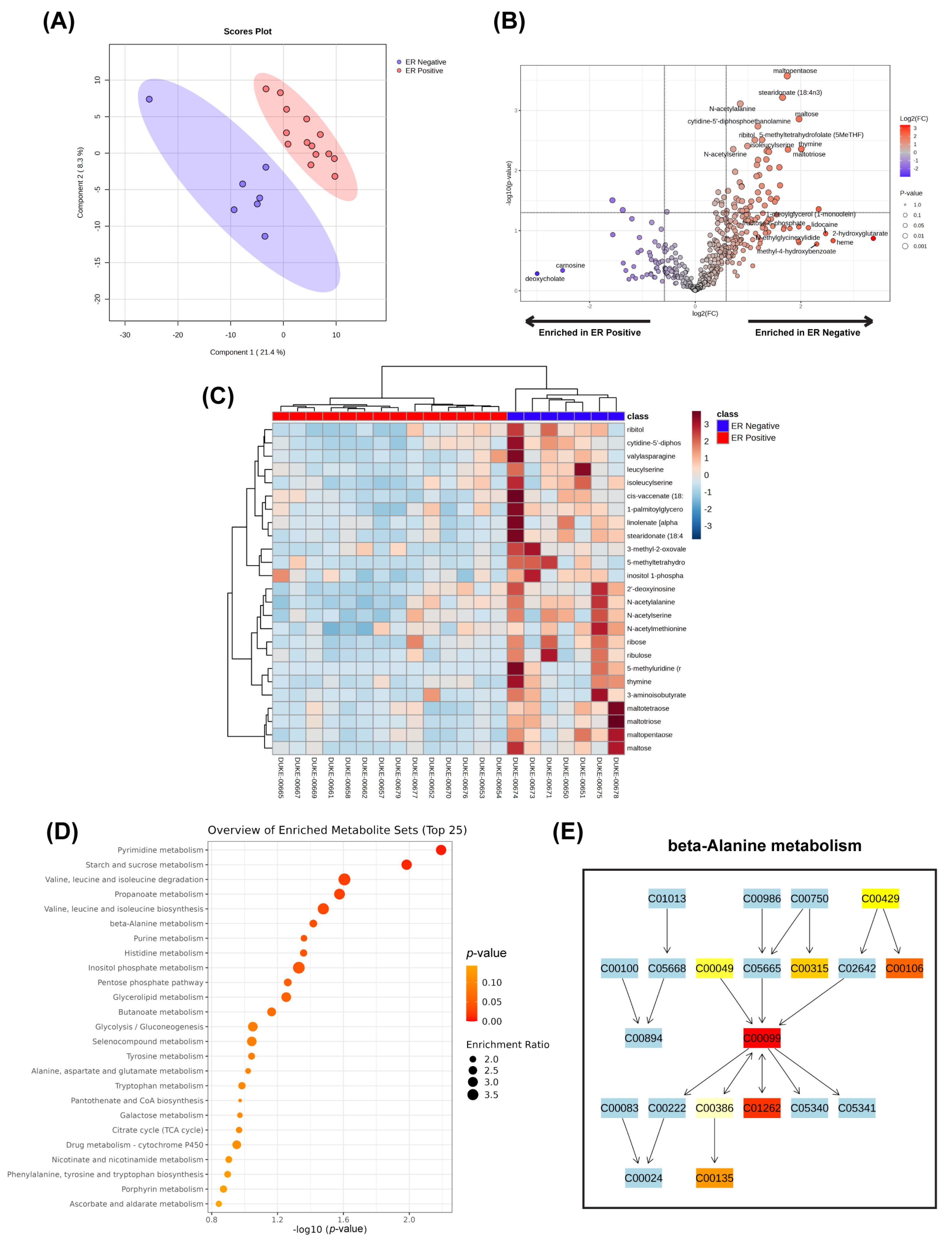
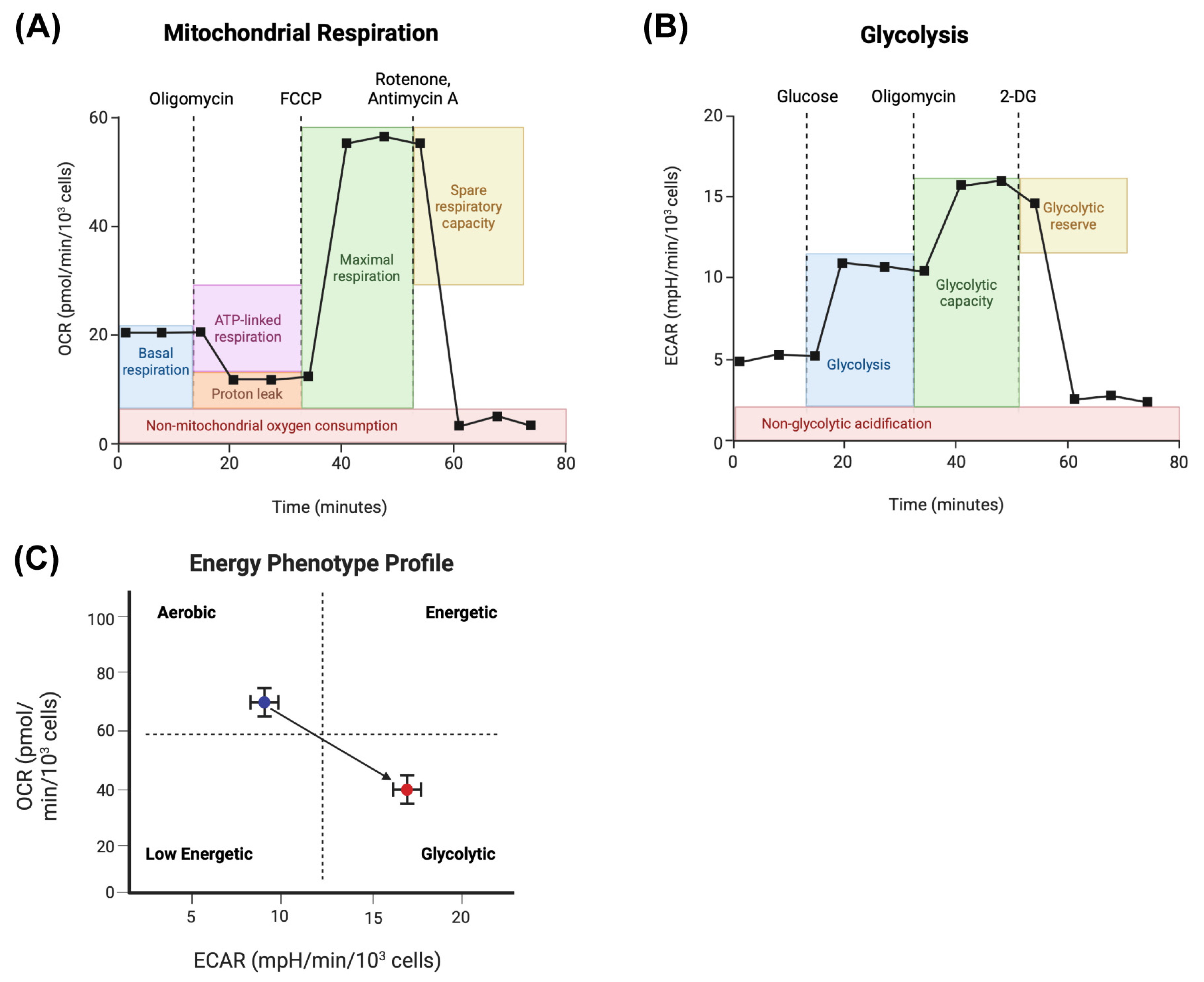
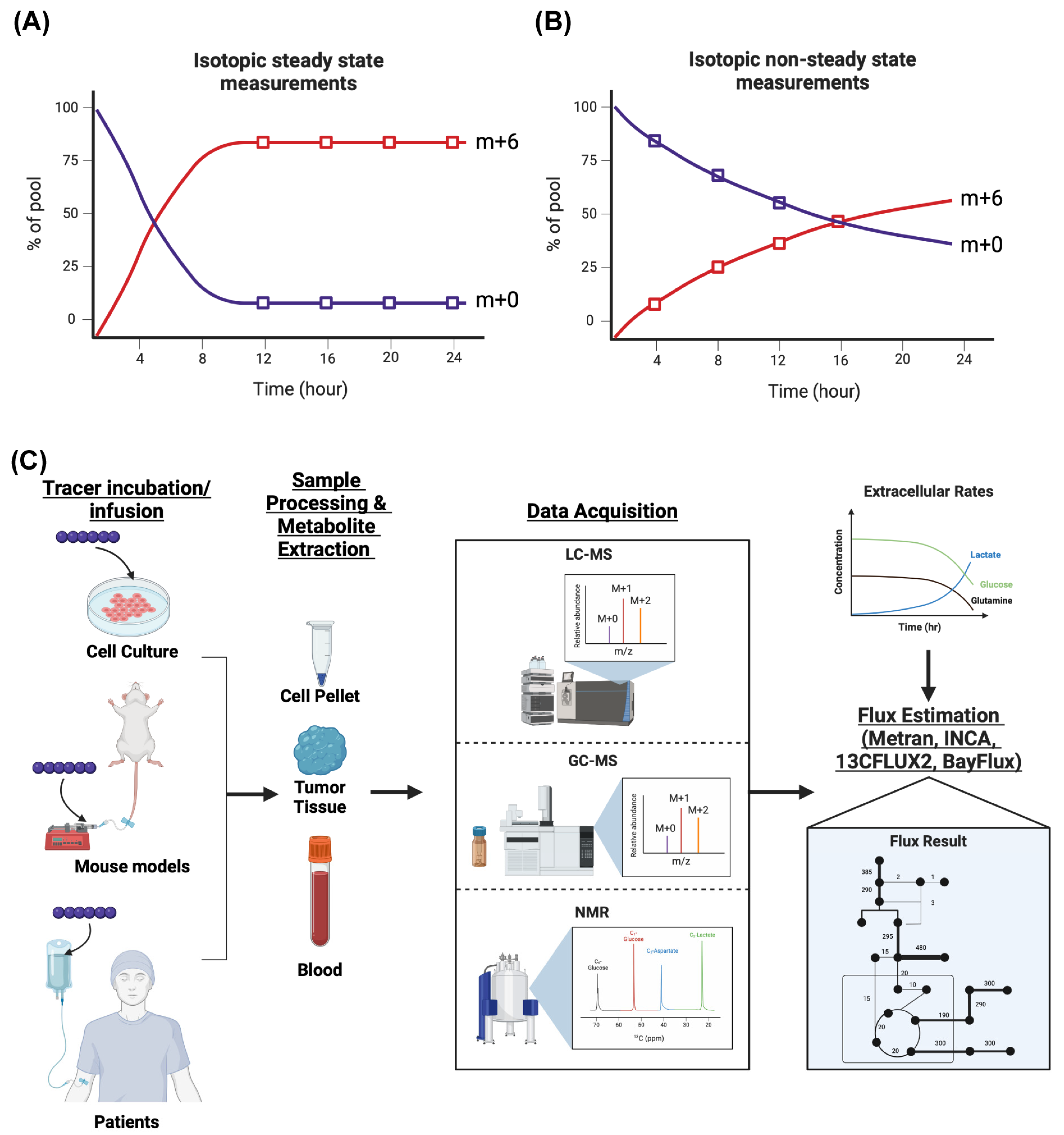
|
Natural Abundance Correction Software | Programming Language |
Limit for Number of Tracers to Use in Experiments | Reference |
|---|---|---|---|
| IsoCor (Version 2.2.3) | Python (Version 3.13.7) | 1 | [105] |
| PyNAC (Version 1.0) | Python (Version 3.13.7) | 2 | [106] |
| PolyMID-Correct (Version 1.0) | Python (Version 3.13.7) | 1 | [104] |
| IsoCorrectoR (Version 3.21) | R (Version 4.5.1) | 2 (any combination of 2H, 13C, 15N, 18O, 34S) | [103] |
| AccuCor2 (Version 0.3.1) | R (Version 4.5.1) | 2 (2H-13C or 13C-15N) | [107] |
| MIDcor (Version 1.0) | R (Version 4.5.1) | 1 | [108] |
| FluxFix (Version 0.1.0) | Web-based (Version 0.1.0) | 1 | [109] |
| ElemCor (Version 1.0) | MATLAB (R2025a) | 1 (2H, 13C, or 15N) | [110] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, M.; Amend, S.R.; Pienta, K.J. Methods and Guidelines for Metabolism Studies: Applications to Cancer Research. Int. J. Mol. Sci. 2025, 26, 8466. https://doi.org/10.3390/ijms26178466
Li M, Amend SR, Pienta KJ. Methods and Guidelines for Metabolism Studies: Applications to Cancer Research. International Journal of Molecular Sciences. 2025; 26(17):8466. https://doi.org/10.3390/ijms26178466
Chicago/Turabian StyleLi, Melvin, Sarah R. Amend, and Kenneth J. Pienta. 2025. "Methods and Guidelines for Metabolism Studies: Applications to Cancer Research" International Journal of Molecular Sciences 26, no. 17: 8466. https://doi.org/10.3390/ijms26178466
APA StyleLi, M., Amend, S. R., & Pienta, K. J. (2025). Methods and Guidelines for Metabolism Studies: Applications to Cancer Research. International Journal of Molecular Sciences, 26(17), 8466. https://doi.org/10.3390/ijms26178466








