The Molecular Landscape of Colorectal Laterally Spreading Tumors: From Endoscopic Subtypes to Molecular Targets
Abstract
1. Introduction
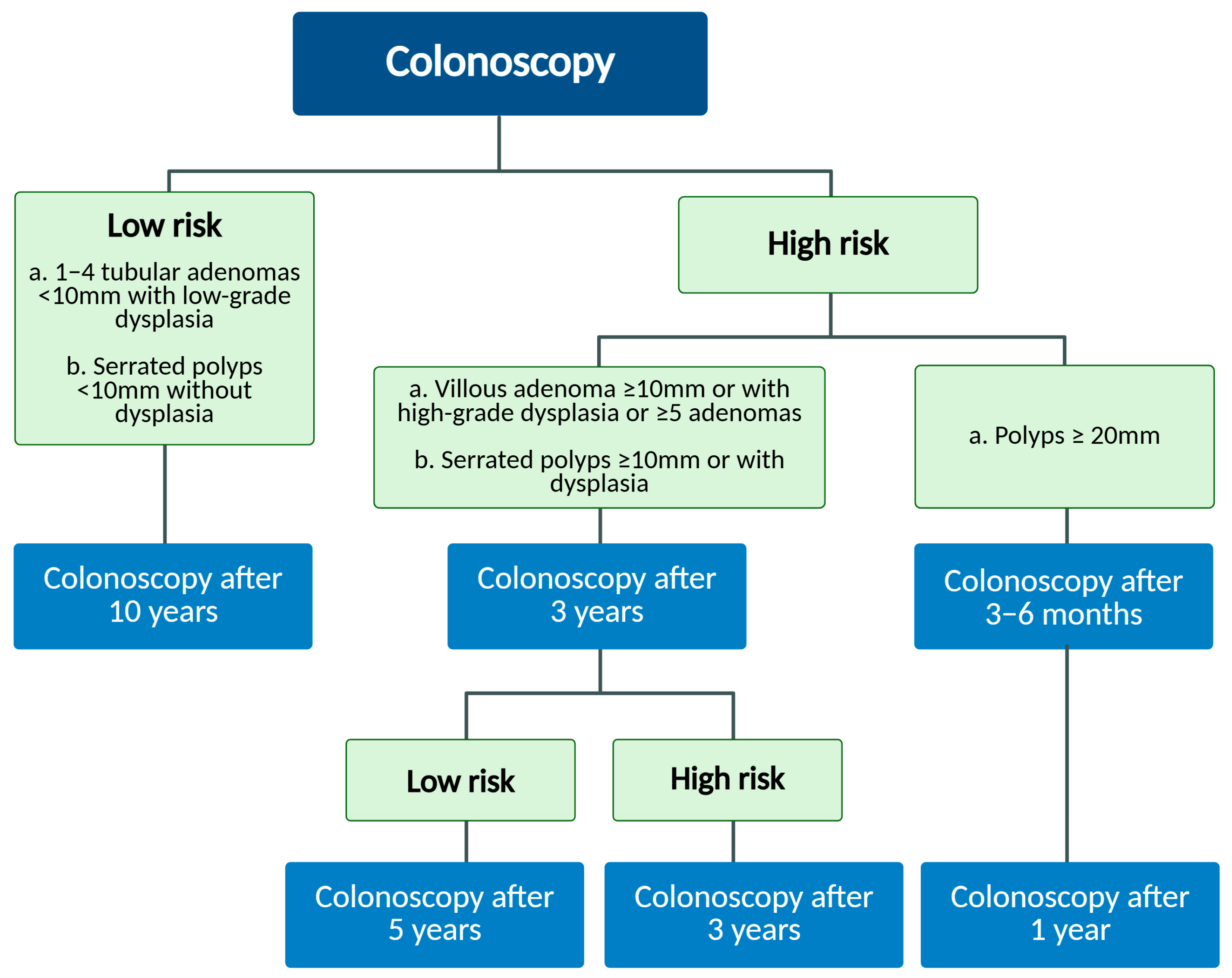
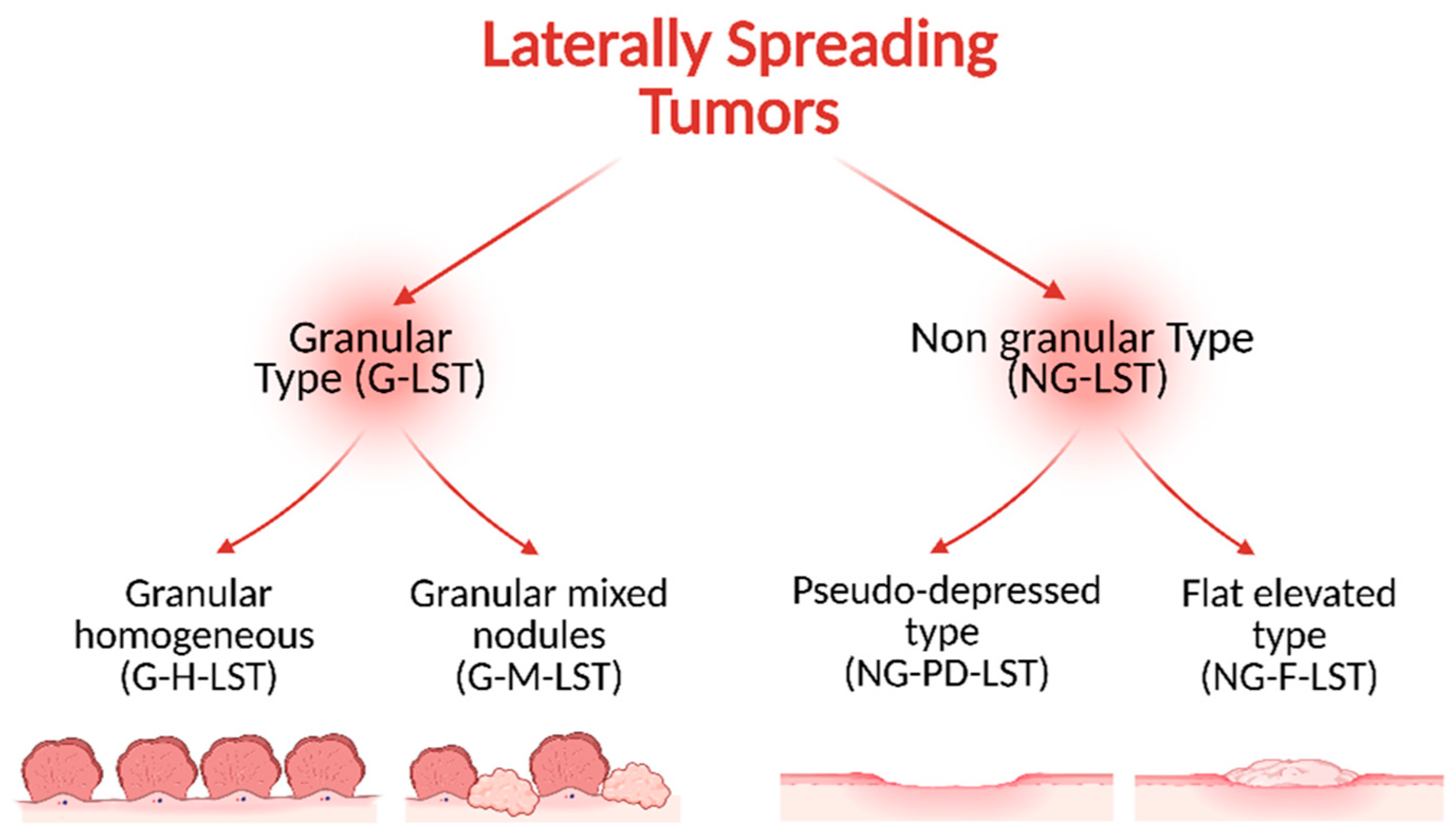
2. Morphological Characteristics of Laterally Spreading Tumors and Risk of Submucosal Invasion
3. Genetic and Molecular Features of LSTs
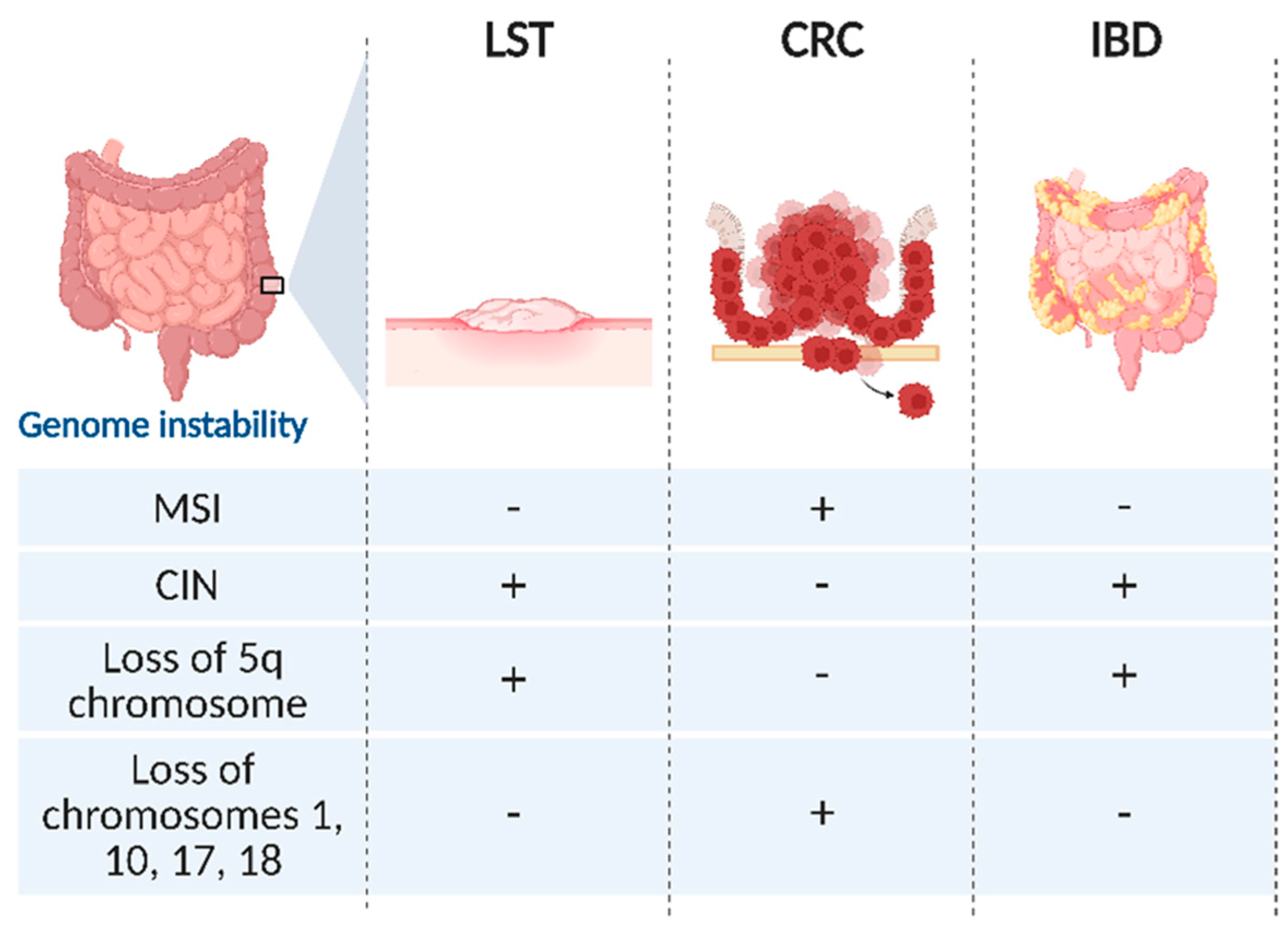
3.1. Microsatellite (MSI) and Chromosome Instability (CIN)
3.2. Genome-Wide Expression Analysis
3.3. Epigenetic Features of LSTs
4. Conclusions and Future Perspectives
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| CIMP | CpG Island Methylator Phenotype |
| CIN | Chromosomal instability |
| COBRA | COmbined Bisulfite Restriction Analysis |
| CRC | Colorectal Cancer |
| DMRs | Differentially methylated probes |
| ESGE | European Society Gastrointestinal Endoscopy |
| G-H-LST | Granular homogeneous LST |
| G-LST | Granular LST |
| G-M-LST | Granular mixed LST |
| IBDs | Inflammatory Bowel Diseases |
| IGRs | Intergenic regions |
| LOH | Loss of heterozygosity |
| LSTs | Laterally Spreading Tumors |
| MMR | Mismatch repair |
| MSI | Microsatellite instability |
| MSI-H | Microsatellite instability-High |
| NG-LST | Nongranular LST |
| NG-F-LST | Nongranular flat-elevated LST |
| NG-PD-LST | Nongranular pseudo-depressed LST |
| RNA-Seq | RNA sequencing |
| SMI | Submucosal invasion |
References
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2024, 74, 229–263. [Google Scholar] [CrossRef]
- Morgan, E.; Arnold, M.; Gini, A.; Lorenzoni, V.; Cabasag, C.J.; Laversanne, M.; Vignat, J.; Ferlay, J.; Murphy, N.; Bray, F. Global burden of colorectal cancer in 2020 and 2040: Incidence and mortality estimates from GLOBOCAN. Gut 2023, 72, 338–344. [Google Scholar] [CrossRef] [PubMed]
- Fearon, E.R.; Vogelstein, B. A Genetic Model for Colorectal Tumorigenesis. Cell 1990, 61, 759–767. [Google Scholar] [CrossRef] [PubMed]
- Mathews, A.A.; Draganov, P.V.; Yang, D. Endoscopic Management of Colorectal Polyps: From Benign to Malignant Polyps. World J. Gastrointest. Endosc. 2021, 13, 356–370. [Google Scholar] [CrossRef]
- Kuo, E.; Wang, K.; Liu, X. A Focused Review on Advances in Risk Stratification of Malignant Polyps. Gastroenterol. Res. 2020, 13, 163–183. [Google Scholar] [CrossRef]
- Nguyen, L.H.; Goel, A.; Chung, D.C. Pathways of Colorectal Carcinogenesis. Gastroenterology 2020, 158, 291–302. [Google Scholar] [CrossRef]
- Pan, J.; Xin, L.; Ma, Y.-F.; Hu, L.-H.; Li, Z.-S. Colonoscopy Reduces Colorectal Cancer Incidence and Mortality in Patients With Non-Malignant Findings: A Meta-Analysis. Am. J. Gastroenterol. 2016, 111, 355–365. [Google Scholar] [CrossRef] [PubMed]
- Kudo, S.E.; Kashida, H. Flat and Depressed Lesions of the Colorectum. Clin. Gastroenterol. Hepatol. 2005, 3, 33–36. [Google Scholar] [CrossRef]
- Matsuda, T.; Saito, Y.; Fujii, T.; Uraoka, T.; Nakajima, T.; Kobayashi, N.; Emura, F.; Ono, A.; Shimoda, T.; Ikematsu, H.; et al. Size Does Not Determine the Grade of Malignancy of Early Invasive Colorectal Cancer. World J. Gastroenterol. 2009, 15, 2708. [Google Scholar] [CrossRef]
- Saitoh, Y.; Waxman, I.; West, A.B.; Popnikolov, N.K.; Gatalica, Z.; Watari, J.; Obara, T.; Kohgo, Y.; Pasricha, P.J. Prevalence and Distinctive Biologic Features of Flat Colorectal Adenomas in a North American Population. Gastroenterology 2001, 120, 1657–1665. [Google Scholar] [CrossRef]
- Hurlstone, D.P.; Cross, S.S.; Adam, I.; Shorthouse, A.J.; Brown, S.; Sanders, D.S.; Lobo, A.J. A Prospective Clinicopathological and Endoscopic Evaluation of Flat and Depressed Colorectal Lesions in the United Kingdom. Am. J. Gastroenterol. 2003, 98, 2543–2549. [Google Scholar] [CrossRef]
- Ross, A.S.; Waxman, I. Flat and Depressed Neoplasms of the Colon in Western Populations. Am. J. Gastroenterol. 2006, 101, 172–180. [Google Scholar] [CrossRef]
- Rotondano, G.; Bianco, M.; Buffoli, F.; Gizzi, G.; Tessari, F.; Cipolletta, L. The Cooperative Italian FLIN Study Group: Prevalence and Clinico-Pathological Features of Colorectal Laterally Spreading Tumors. Endoscopy 2011, 43, 856–861. [Google Scholar] [CrossRef]
- Sanduleanu, S.; Siersema, P. Laterally Spreading Tumor through the Magnifying Glass: We Only See What We Know. Endoscopy 2016, 48, 421–423. [Google Scholar] [CrossRef]
- Soetikno, R.M. Prevalence of Nonpolypoid (Flat and Depressed) Colorectal Neoplasms in Asymptomatic and Symptomatic Adults. JAMA 2008, 299, 1027. [Google Scholar] [CrossRef]
- Holme, Ø.; Løberg, M.; Kalager, M.; Bretthauer, M.; Hernán, M.A.; Aas, E.; Eide, T.J.; Skovlund, E.; Lekven, J.; Schneede, J.; et al. Long-Term Effectiveness of Sigmoidoscopy Screening on Colorectal Cancer Incidence and Mortality in Women and Men. Ann. Intern. Med. 2018, 168, 775–782. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wang, Q.; Jia, C.; Li, D.; Lv, Z.; Yang, J. Role of Atrial Natriuretic Peptide Receptor in Inhibition of Laterally Spreading Tumors via Wnt/β-Catenin Signaling. Arch. Med. Sci. Atheroscler. Dis. 2022, 7, 104–108. [Google Scholar] [CrossRef] [PubMed]
- Yamada, M.; Saito, Y.; Sakamoto, T.; Nakajima, T.; Kushima, R.; Parra-Blanco, A.; Matsuda, T. Endoscopic Predictors of Deep Submucosal Invasion in Colorectal Laterally Spreading Tumors. Endoscopy 2016, 48, 456–464. [Google Scholar] [CrossRef] [PubMed]
- Ikematsu, H.; Murano, T.; Shinmura, K. Detection of Colorectal Lesions during Colonoscopy. DEN Open 2022, 2, e68. [Google Scholar] [CrossRef]
- Shahsavari, D.; Waqar, M.; Thoguluva Chandrasekar, V. Image Enhanced Colonoscopy: Updates and Prospects—A Review. Transl. Gastroenterol. Hepatol. 2023, 8, 26. [Google Scholar] [CrossRef]
- Hassan, C.; Antonelli, G.; Dumonceau, J.M.; Regula, J.; Bretthauer, M.; Chaussade, S.; Dekker, E.; Ferlitsch, M.; Gimeno-García, A.; Jover, R.; et al. Post-Polypectomy Colonoscopy Surveillance: European Society of Gastrointestinal Endoscopy (ESGE) Guideline—Update 2020. Endoscopy 2020, 52, 687–700. [Google Scholar] [CrossRef]
- Sugimoto, T.; Ohta, M.; Ikenoue, T.; Yamada, A.; Tada, M.; Fujishiro, M.; Ogura, K.; Yamaji, Y.; Okamoto, M.; Kanai, F.; et al. Macroscopic Morphologic Subtypes of Laterally Spreading Colorectal Tumors Showing Distinct Molecular Alterations. Int. J. Cancer 2010, 127, 1562–1569. [Google Scholar] [CrossRef]
- Kudo, S. Endoscopic Mucosal Resection of Flat and Depressed Types of Early Colorectal Cancer. Endoscopy 1993, 25, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Kudo, S.-E.; Takemura, O.; Ohtsuka, K. Flat and Depressed Types of Early Colorectal Cancers: From East to West. Gastrointest. Endosc. Clin. N. Am. 2008, 18, 581–593. [Google Scholar] [CrossRef]
- Lambert, R.L.C. The Paris Endoscopic Classification of Superficial Neoplastic Lesions. Gastrointest. Endosc. 2003, 58, 3–43. [Google Scholar] [CrossRef]
- Moss, A.; Bourke, M.J.; Williams, S.J.; Hourigan, L.F.; Brown, G.; Tam, W.; Singh, R.; Zanati, S.; Chen, R.Y.; Byth, K. Endoscopic Mucosal Resection Outcomes and Prediction of Submucosal Cancer from Advanced Colonic Mucosal Neoplasia. Gastroenterology 2011, 140, 1909–1918. [Google Scholar] [CrossRef]
- Metz, A.J.; Bourke, M.J.; Moss, A.; Dower, A.; Zarzour, P.; Hawkins, N.J.; Ward, R.L.; Hesson, L.B. A Correlation of the Endoscopic Characteristics of Colonic Laterally Spreading Tumours with Genetic Alterations. Eur. J. Gastroenterol. Hepatol. 2013, 25, 319–326. [Google Scholar] [CrossRef]
- Bogie, R.M.M.; Veldman, M.H.J.; Snijders, L.A.R.S.; Winkens, B.; Kaltenbach, T.; Masclee, A.A.M.; Matsuda, T.; Rondagh, E.J.A.; Soetikno, R.; Tanaka, S.; et al. Endoscopic Subtypes of Colorectal Laterally Spreading Tumors (LSTs) and the Risk of Submucosal Invasion: A Meta-Analysis. Endoscopy 2018, 50, 263–282, Correction in Endoscopy 2018, 50, 4. [Google Scholar] [CrossRef] [PubMed]
- Snover, D.C. Update on the Serrated Pathway to Colorectal Carcinoma. Hum. Pathol. 2011, 42, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Kinzler, K.W.; Vogelstein, B. Lessons from Hereditary Colorectal Cancer. Cell 1996, 87, 159–170. [Google Scholar] [CrossRef] [PubMed]
- Testa, U.; Pelosi, E.; Castelli, G. Colorectal Cancer: Genetic Abnormalities, Tumor Progression, Tumor Heterogeneity, Clonal Evolution and Tumor-Initiating Cells. Med. Sci. 2018, 6, 31. [Google Scholar] [CrossRef] [PubMed]
- Umetani, N.; Sasaki, S.; Masaki, T.; Watanabe, T.; Matsuda, K.; Muto, T. Involvement of APC and K-Ras Mutation in Non-Polypoid Colorectal Tumorigenesis. Br. J. Cancer 2000, 82, 9–15. [Google Scholar] [CrossRef]
- Mikami, M.; Nosho, K.; Yamamoto, H.; Takahashi, T.; Maehata, T.; Taniguchi, H.; Adachi, Y.; Imamura, A.; Fujita, M.; Hosokawa, M.; et al. Mutational Analysis of β-Catenin and the RAS-RAF Signalling Pathway in Early Flat-Type Colorectal Tumours. Eur. J. Cancer 2006, 42, 3065–3072. [Google Scholar] [CrossRef]
- Lynch, H.T.; Smyrk, T.C.; Watson, P.; Lanspa, S.J.; Lynch, J.F.; Lynch, P.M.; Cavalieri, R.J.; Boland, C.R. Genetics, Natural History, Tumor Spectrum, and Pathology of Hereditary Nonpolyposis Colorectal Cancer: An Updated Review. Gastroenterology 1993, 104, 1535–1549. [Google Scholar] [CrossRef]
- Minamoto, T.; Sawaguchi, K.; Mai, M.; Yamashita, N.; Sugimura, T.; Esumi, H. Infrequent K-Ras Activation in Superficial-Type (Flat) Colorectal Adenomas and Adenocarcinomas. Cancer Res. 1994, 54, 2841–2844. [Google Scholar]
- Yukawa, M.; Fujimori, T.; Maeda, S.; Tabuchi, M.; Nagasako, K. Comparative Clinicopathological and Immunohistochemical Study of Ras and P53 in Flat and Polypoid Type Colorectal Tumours. Gut 1994, 35, 1258–1261. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Quinlan, M.P.; Settleman, J. Isoform-Specific Ras Functions in Development and Cancer. Futur. Oncol. 2009, 5, 105–116. [Google Scholar] [CrossRef]
- Chiang, J.M.; Chou, Y.H.W.; Chou, T. Bin K-Ras Codon 12 Mutation Determines the Polypoid Growth of Colorectal Cancer. Cancer Res. 1998, 58, 3289–3293. [Google Scholar]
- Maltzman, T.; Knoll, K.; Martinez, M.E.; Byers, T.; Stevens, B.R.; Marshall, J.R.; Reid, M.E.; Einspahr, J.; Hart, N.; Bhattacharyya, A.K.; et al. Ki-Ras Proto-Oncogene Mutations in Sporadic Colorectal Adenomas: Relationship to Histologic and Clinical Characteristics. Gastroenterology 2001, 121, 302–309. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Lefferts, J.A.; Schwab, M.C.; Suriawinata, A.A.; Tsongalis, G.J. Correlation of Polypoid Colorectal Adenocarcinoma with Pre-Existing Adenomatous Polyps and KRAS Mutation. Cancer Genet. 2011, 204, 245–251. [Google Scholar] [CrossRef]
- Meng, M.; Zhong, K.; Jiang, T.; Liu, Z.; Kwan, H.Y.; Su, T. The Current Understanding on the Impact of KRAS on Colorectal Cancer. Biomed. Pharmacother. 2021, 140, 111717. [Google Scholar] [CrossRef] [PubMed]
- Voorham, Q.J.M.; Rondagh, E.J.A.; Knol, D.L.; Van Engeland, M.; Carvalho, B.; Meijer, G.A.; Sanduleanu, S. Tracking the Molecular Features of Nonpolypoid Colorectal Neoplasms: A Systematic Review and Meta-Analysis. Am. J. Gastroenterol. 2013, 108, 1042–1056. [Google Scholar] [CrossRef]
- Kaji, E.; Kato, J.; Suzuki, H.; Akita, M.; Horii, J.; Saito, S.; Higashi, R.; Ishikawa, S.; Kuriyama, M.; Hiraoka, S.; et al. Analysis of K-Ras, BRAF, and PIK3CA Mutations in Laterally-Spreading Tumors of the Colorectum. J. Gastroenterol. Hepatol. 2011, 26, 599–607. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.O.; Jang, B.I.; Jang, W.J.; Lee, S.H. Laterally Spreading Tumors of the Colorectum: Clinicopathologic Features and Malignant Potential by Macroscopic Morphology. Int. J. Color. Dis. 2013, 28, 1661–1666. [Google Scholar] [CrossRef] [PubMed]
- Nakae, K.; Mitomi, H.; Saito, T.; Takahashi, M.; Morimoto, T.; Hidaka, Y.; Sakamoto, N.; Yao, T.; Watanabe, S. MUC5AC/β-Catenin Expression and KRAS Gene Alteration in Laterally Spreading Colorectal Tumors. World J. Gastroenterol. 2012, 18, 5551–5559. [Google Scholar] [CrossRef]
- Kargman, S.L.; O’Neill, G.P.; Vickers, P.J.; Evans, J.F.; Mancini, J.A.; Jothy, S. Expression of Prostaglandin G/H Synthase-1 and -2 Protein in Human Colon Cancer. Cancer Res. 1995, 55, 2556–2559. [Google Scholar] [CrossRef]
- Singh, P.; Velasco, M.; Given, R.; Varro, A.; Wang, T.C. Progastrin Expression Predisposes Mice to Colon Carcinomas and Adenomas in Response to a Chemical Carcinogen. Gastroenterology 2000, 119, 162–171. [Google Scholar] [CrossRef]
- Masferrer, J.L.; Seibert, K.; Zweifel, B.; Needleman, P. Endogenous Glucocorticoids Regulate an Inducible Cyclooxygenase Enzyme. Proc. Natl. Acad. Sci. USA 1992, 89, 3917–3921. [Google Scholar] [CrossRef]
- Simon, L.S. Role and Regulation of Cyclooxygenase-2 during Inflammation. Am. J. Med. 1999, 106, 37S–42S. [Google Scholar] [CrossRef]
- Lee, E.J.; Choi, E.M.; Kim, S.R.; Park, J.H.; Kim, H.; Ha, K.S.; Kim, Y.M.; Kim, S.S.; Choe, M.; Kim, J.-I.; et al. Cyclooxygenase-2 Promotes Cell Proliferation, Migration and Invasion in U2OS Human Osteosarcoma Cells. Exp. Mol. Med. 2007, 39, 469–476. [Google Scholar] [CrossRef]
- Iwase, K.; Evers, B.; Hellmich, M.; Guo, Y.; Higashide, S.; Kim, H.; Townsend, C. Regulation of Growth of Human Gastric Cancer by Gastrin and Glycine-Extended Progastrin. Gastroenterology 1997, 113, 782–790. [Google Scholar] [CrossRef] [PubMed]
- Fujita, M.; Fukui, H.; Morita, K.; Fujii, S.; Ueda, Y.; Chiba, T.; Sakamoto, C.; Kawamata, H.; Fujimori, T. Relationship between Cyclooxygenase-2 Expression and K-Ras Gene Mutation in Colorectal Adenomas. J. Gastroenterol. Hepatol. 2000, 15, 1277–1281. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.T.; Lawson, K.R.; Ignatenko, N.A.; Marek, S.E.; Stringer, D.E.; Skovan, B.A.; Gerner, E.W. Sulindac Sulfone Inhibits K-Ras-Dependent Cyclooxygenase-2 Expression in Human Colon Cancer Cells. Cancer Res. 2000, 60, 6607–6610. [Google Scholar]
- Mukawa, K.; Fujii, S.; Takeda, J.; Kitajima, K.; Tominaga, K.; Chibana, Y.; Fujita, M.; Ichikawa, K.; Tomita, S.; Ono, Y.; et al. Analysis of K-ras Mutations and Expression of Cyclooxygenase-2 and Gastrin Protein in Laterally Spreading Tumors. J. Gastroenterol. Hepatol. 2005, 20, 1584–1590. [Google Scholar] [CrossRef]
- Goss, K.H.; Groden, J. Biology of the adenomatous polyposis coli tumor suppressor. J. Clin. Oncol. 2000, 18, 1967–1979. [Google Scholar] [CrossRef] [PubMed]
- Sena, P.; Saviano, M.; Monni, S.; Losi, L.; Roncucci, L.; Marzona, L.; De Pol, A. Subcellular Localization of β-Catenin and APC Proteins in Colorectal Preneoplastic and Neoplastic Lesions. Cancer Lett. 2006, 241, 203–212. [Google Scholar] [CrossRef]
- Wang, J.; Wang, X.; Gong, W.; Mi, B.; Liu, S.; Jiang, B. Increased Expression of β-Catenin, Phosphorylated Glycogen Synthase Kinase 3β, Cyclin D1, and c-Myc in Laterally Spreading Colorectal Tumors. J. Histochem. Cytochem. 2009, 57, 363–371. [Google Scholar] [CrossRef]
- Shi, L.; Wu, Y.-X.; Yu, J.-H.; Chen, X.; Luo, X.-J.; Yin, Y.-R. Research of the Relationship between β-Catenin and c-Myc-Mediated Wnt Pathway and LSTs Occurrence. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 252–257. [Google Scholar]
- Tighe, A.; Johnson, V.L.; Taylor, S.S. Truncating APC Mutations Have Dominant Effects on Proliferation, Spindle Checkpoint Control, Survival and Chromosome Stability. J. Cell Sci. 2004, 117, 6339–6353. [Google Scholar] [CrossRef]
- Lu, S.; Jia, C.-Y.; Yang, J.-S. Future Therapeutic Implications of New Molecular Mechanism of Colorectal Cancer. World J. Gastroenterol. 2023, 29, 2359–2368. [Google Scholar] [CrossRef]
- Hirsch, D.; Hardt, J.; Sauer, C.; Heselmeyer-Hadded, K.; Witt, S.H.; Kienle, P.; Ried, T.; Gaiser, T. Molecular Characterization of Ulcerative Colitis-Associated Colorectal Carcinomas. Mod. Pathol. 2021, 34, 1153–1166. [Google Scholar] [CrossRef]
- Esteller, M.; Sparks, A.; Toyota, M.; Sanchez-Cespedes, M.; Capella, G.; Peinado, M.A.; Gonzalez, S.; Tarafa, G.; Sidransky, D.; Meltzer, S.J.; et al. Analysis of Adenomatous Polyposis Coli Promoter Hypermethylation in Human Cancer. Cancer Res. 2000, 60, 4366–4371. [Google Scholar] [PubMed]
- Huang, K.; Zhang, J.-X.; Han, L.; You, Y.-P.; Jiang, T.; Pu, P.-Y.; Kang, C.-S. MicroRNA Roles in Beta-Catenin Pathway. Mol. Cancer 2010, 9, 252. [Google Scholar] [CrossRef]
- Fearnhead, N.S. The ABC of APC. Hum. Mol. Genet. 2001, 10, 721–733. [Google Scholar] [CrossRef]
- Voorham, Q.J.M.; Carvalho, B.; Spiertz, A.J.; Claes, B.; Mongera, S.; van Grieken, N.C.T.; Grabsch, H.; Kliment, M.; Rembacken, B.; van de Wiel, M.A.; et al. Comprehensive Mutation Analysis in Colorectal Flat Adenomas. PLoS ONE 2012, 7, e41963. [Google Scholar] [CrossRef]
- Sakai, E.; Ohata, K.; Chiba, H.; Matsuhashi, N.; Doi, N.; Fukushima, J.; Endo, H.; Takahashi, H.; Tsuji, S.; Yagi, K.; et al. Methylation Epigenotypes and Genetic Features in Colorectal Laterally Spreading Tumors. Int. J. Cancer 2014, 135, 1586–1595. [Google Scholar] [CrossRef]
- Ichikawa, Y.; Nagashima, Y.; Morioka, K.; Akimoto, K.; Kojima, Y.; Ishikawa, T.; Goto, A.; Kobayashi, N.; Watanabe, K.; Ota, M.; et al. Colorectal Laterally Spreading Tumors Show Characteristic Expression of Cell Polarity Factors, Including Atypical Protein Kinase C λ/ι, E-Cadherin, β-Catenin and Basement Membrane Component. Oncol. Lett. 2014, 8, 977–984. [Google Scholar] [CrossRef]
- Wang, X.-Y.; Lai, Z.-S.; Yeung, C.-M.; Wang, J.-D.; Deng, W.; Li, H.Y.; Han, Y.-J.; Kung, H.-F.; Jiang, B.; Lin, M.C. Establishment and Characterization of a New Cell Line Derived from Human Colorectal Laterally Spreading Tumor. World J. Gastroenterol. 2008, 14, 1204–1211. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chen, J. The Cell-Cycle Arrest and Apoptotic and Progression. Cold Spring Harb. Perspect. Med. 2016, 6, a026104. [Google Scholar] [CrossRef]
- Sugai, T.; Habano, W.; Uesugi, N.; Jiao, Y.F.; Nakamura, S.I.; Yoshida, T.; Higuchi, T. Frequent Allelic Imbalance at the ATM Locus in DNA Multiploid Colorectal Carcinomas. Oncogene 2001, 20, 6095–6101. [Google Scholar] [CrossRef] [PubMed]
- Noro, A.; Sugai, T.; Habano, W.; Nakamura, S.I. Analysis of Ki-Ras and P53 Gene Mutations in Laterally Spreading Tumors of the Colorectum. Pathol. Int. 2003, 53, 828–836. [Google Scholar] [CrossRef]
- Sakai, E.; Fukuyo, M.; Matsusaka, K.; Ohata, K.; Doi, N.; Takane, K.; Matsuhashi, N.; Fukushima, J.; Nakajima, A.; Kaneda, A. TP53 Mutation at Early Stage of Colorectal Cancer Progression from Two Types of Laterally Spreading Tumors. Cancer Sci. 2016, 107, 820–827. [Google Scholar] [CrossRef]
- Orita, H.; Sakamoto, N.; Ajioka, Y.; Terai, T.; Hino, O.; Sato, N.; Shimoda, T.; Kamano, T.; Tsurumaru, M.; Fujii, H. Allelic Loss Analysis of Early-Stage Flat-Type Colorectal Tumors. Ann. Oncol. 2006, 17, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Barault, L.; Veyrie, N.; Jooste, V.; Lecorre, D.; Chapusot, C.; Ferraz, J.; Lièvre, A.; Cortet, M.; Bouvier, A.; Rat, P.; et al. Mutations in the RAS-MAPK, PI(3)K (Phosphatidylinositol-3-OH Kinase) Signaling Network Correlate with Poor Survival in a Population-based Series of Colon Cancers. Int. J. Cancer 2008, 122, 2255–2259. [Google Scholar] [CrossRef]
- Itatani, Y.; Kawada, K.; Sakai, Y. Transforming Growth Factor-β Signaling Pathway in Colorectal Cancer and Its Tumor Microenvironment. Int. J. Mol. Sci. 2019, 20, 5822. [Google Scholar] [CrossRef]
- Chang, L.C.; Chiu, H.M.; Shun, C.T.; Liang, J.T.; Lin, J.T.; Chen, C.C.; Lee, Y.C.; Wu, M.S. Mutational Profiles of Different Macroscopic Subtypes of Colorectal Adenoma Reveal Distinct Pathogenetic Roles for KRAS, BRAF and PIK3CA. BMC Gastroenterol. 2014, 14, 221. [Google Scholar] [CrossRef][Green Version]
- Velho, S.; Oliveira, C.; Ferreira, A.; Ferreira, A.C.; Suriano, G.; Schwartz, S., Jr.; Duval, A.; Carneiro, F.; Machado, J.C.; Hamelin, R.; et al. The Prevalence of PIK3CA Mutations in Gastric and Colon Cancer. Eur. J. Cancer 2005, 41, 1649–1654. [Google Scholar] [CrossRef]
- Kato, S.; Iida, S.; Higuchi, T.; Ishikawa, T.; Takagi, Y.; Yasuno, M.; Enomoto, M.; Uetake, H.; Sugihara, K. PIK3CA Mutation Is Predictive of Poor Survival in Patients with Colorectal Cancer. Int. J. Cancer 2007, 121, 1771–1778. [Google Scholar] [CrossRef]
- Konda, K.; Konishi, K.; Yamochi, T.; Ito, Y.M.; Nozawa, H.; Tojo, M.; Shinmura, K.; Kogo, M.; Katagiri, A.; Kubota, Y.; et al. Distinct Molecular Features of Different Macroscopic Subtypes of Colorectal Neoplasms. PLoS ONE 2014, 9, e103822. [Google Scholar] [CrossRef] [PubMed]
- Hesson, L.B.; Ng, B.; Zarzour, P.; Srivastava, S.; Kwok, C.; Packham, D.; Nunez, A.C.; Beck, D.; Ryan, R.; Dower, A.; et al. Integrated Genetic, Epigenetic, and Transcriptional Profiling Identifies Molecular Pathways in the Development of Laterally Spreading Tumors. Mol. Cancer Res. 2016, 14, 1217–1228. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Ding, M.; Qian, N.; Song, B.; Yu, J.; Tang, J.; Wang, J. Decreased Expression of Semaphorin 3D Is Associated with Genesis and Development in Colorectal Cancer. World J. Surg. Oncol. 2017, 15, 67. [Google Scholar] [CrossRef]
- Arakawa, H. Netrin-1 and Its Receptors in Tumorigenesis. Nat. Rev. Cancer 2004, 4, 978–987. [Google Scholar] [CrossRef]
- Shin, S.K.; Nagasaka, T.; Jung, B.H.; Matsubara, N.; Kim, W.H.; Carethers, J.M.; Boland, C.R.; Goel, A. Epigenetic and Genetic Alterations in Netrin-1 Receptors UNC5C and DCC in Human Colon Cancer. Gastroenterology 2007, 133, 1849–1857. [Google Scholar] [CrossRef]
- Cortina, C.; Palomo-Ponce, S.; Iglesias, M.; Fernández-Masip, J.L.; Vivancos, A.; Whissell, G.; Humà, M.; Peiró, N.; Gallego, L.; Jonkheer, S.; et al. EphB–Ephrin-B Interactions Suppress Colorectal Cancer Progression by Compartmentalizing Tumor Cells. Nat. Genet. 2007, 39, 1376–1383. [Google Scholar] [CrossRef]
- Chédotal, A.; Kerjan, G.; Moreau-Fauvarque, C. The Brain within the Tumor: New Roles for Axon Guidance Molecules in Cancers. Cell Death Differ. 2005, 12, 1044–1056. [Google Scholar] [CrossRef] [PubMed]
- Dallol, A.; Morton, D.; Maher, E.R.; Latif, F. SLIT2 axon guidance molecule is frequently inactivated in colorectal cancer and suppresses growth of colorectal carcinoma cells. Cancer Res. 2003, 63, 1054–1058. [Google Scholar] [PubMed]
- Li, J.; Ye, L.; Sanders, A.J.; Jiang, W.G. Repulsive Guidance Molecule B (RGMB) Plays Negative Roles in Breast Cancer by Coordinating BMP Signaling. J. Cell Biochem. 2012, 113, 2523–2531. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, H.; Ikematsu, H.; Fujii, S.; Osera, S.; Odagaki, T.; Oono, Y.; Yano, T.; Ochiai, A.; Sasaki, Y.; Kaneko, K. Clinicopathological Differences of Laterally Spreading Tumors Arising in the Colon and Rectum. Int. J. Color. Dis. 2014, 29, 1069–1075. [Google Scholar] [CrossRef]
- Osera, S.; Fujii, S.; Ikematsu, H.; Miyamoto, H.; Oono, Y.; Yano, T.; Ochiai, A.; Yoshino, T.; Ohtsu, A.; Kaneko, K. Clinicopathological, Endoscopic, and Molecular Characteristics of the “Skirt”—A New Entity of Lesions at the Margin of Laterally Spreading Tumors. Endoscopy 2016, 48, 448–455. [Google Scholar] [CrossRef]
- Markowitz, S.D.; Bertagnolli, M.M. Molecular Basis of Colorectal Cancer. N. Engl. J. Med. 2009, 361, 2449–2460. [Google Scholar] [CrossRef]
- De La Chapelle, A.; Hampel, H. Clinical Relevance of Microsatellite Instability in Colorectal Cancer. J. Clin. Oncol. 2010, 28, 3380–3387. [Google Scholar] [CrossRef]
- Pino, M.S.; Chung, D.C. The Chromosomal Instability Pathway in Colon Cancer. Gastroenterology 2010, 138, 2059–2072. [Google Scholar] [CrossRef] [PubMed]
- Hosea, R.; Hillary, S.; Naqvi, S.; Wu, S.; Kasim, V. The two sides of chromosomal instability: Drivers and brakes in cancer. Signal Transduct. Target. Ther. 2024, 9, 75. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Luo, H.; Huang, L.; Luo, H.; Zhu, X. Microsatellite instability: A review of what the oncologist should know. Cancer Cell Int. 2020, 20, 16. [Google Scholar] [CrossRef]
- Sugai, T.; Habano, W.; Takagi, R.; Yamano, H.; Eizuka, M.; Arakawa, N.; Takahashi, Y.; Yamamoto, E.; Kawasaki, K.; Yanai, S.; et al. Analysis of Molecular Alterations in Laterally Spreading Tumors of the Colorectum. J. Gastroenterol. 2017, 52, 715–723. [Google Scholar] [CrossRef]
- Rioux, J.D.; Silverberg, M.S.; Daly, M.J.; Steinhart, A.H.; McLeod, R.S.; Griffiths, A.M.; Green, T.; Brettin, T.S.; Stone, V.; Bull, S.B.; et al. Genomewide Search in Canadian Families with Inflammatory Bowel Disease Reveals Two Novel Susceptibility Loci. Am. J. Hum. Genet. 2000, 66, 1863–1870. [Google Scholar] [CrossRef]
- Diep, C.B.; Kleivi, K.; Ribeiro, F.R.; Teixeira, M.R.; Lindgjærde, O.C.; Lothe, R.A. The Order of Genetic Events Associated with Colorectal Cancer Progression Inferred from Meta-Analysis of Copy Number Changes. Genes, Chromosom. Cancer 2005, 45, 31–41. [Google Scholar] [CrossRef]
- Fijneman, R.J.A.; Carvalho, B.; Postma, C.; Mongera, S.; van Hinsbergh, V.W.M.; Meijer, G.A. Loss of 1p36, Gain of 8q24, and Loss of 9q34 Are Associated with Stroma Percentage of Colorectal Cancer. Cancer Lett. 2007, 258, 223–229. [Google Scholar] [CrossRef]
- Kita, H.; Hikichi, Y.; Hikami, K.; Tsuneyama, K.; Cui, Z.-G.; Osawa, H.; Ohnishi, H.; Mutoh, H.; Hoshino, H.; Bowlus, C.L.; et al. Differential Gene Expression between Flat Adenoma and Normal Mucosa in the Colon in a Microarray Analysis. J. Gastroenterol. 2006, 41, 1053–1063. [Google Scholar] [CrossRef]
- Minemura, S.; Tanaka, T.; Arai, M.; Okimoto, K.; Oyamada, A.; Saito, K.; Maruoka, D.; Matsumura, T.; Nakagawa, T.; Katsuno, T.; et al. Gene Expression Profiling of Laterally Spreading Tumors. BMC Gastroenterol. 2015, 15, 64. [Google Scholar] [CrossRef] [PubMed]
- Altieri, D.C. Validating Survivin as a Cancer Therapeutic Target. Nat. Rev. Cancer 2003, 3, 46–54. [Google Scholar] [CrossRef]
- Nong, Y.; Zhang, Y.; Zhang, Y.; Pan, L.; Chen, J.; Zhu, C.; Han, L.; Li, A.; Liu, S. Analysis of Genetic Alterations Identifies the Frequent Mutation of GNAS in Colorectal Laterally Spreading Tumors. Cancer Commun. 2020, 40, 636–640. [Google Scholar] [CrossRef] [PubMed]
- Weinstein, L.S.; Liu, J.; Sakamoto, A.; Xie, T.; Chen, M. Minireview: GNAS: Normal and Abnormal Functions. Endocrinology 2004, 145, 5459–5464. [Google Scholar] [CrossRef] [PubMed]
- Fadaka, A.O.; Bakare, O.O.; Pretorius, A.; Klein, A. Genomic Profiling of MicroRNA Target Genes in Colorectal Cancer. Tumor Biology 2020, 42, 1010428320933512. [Google Scholar] [CrossRef]
- Afolabi, H.A.; Salleh, S.M.; Zakaria, Z.; Ch’ng, E.S.; Nafi, S.N.M.; Aziz, A.A.B.A.; Irekeola, A.A.; Wada, Y.; Al-Mhanna, S.B. A GNAS Gene Mutation’s Independent Expression in the Growth of Colorectal Cancer: A Systematic Review and Meta-Analysis. Cancers 2022, 14, 5480. [Google Scholar] [CrossRef] [PubMed]
- Rai, S.; Singh, M.P.; Srivastava, S. Integrated Analysis Identifies Novel Fusion Transcripts in Laterally Spreading Tumors Suggestive of Distinct Etiology Than Colorectal Cancers. J. Gastrointest. Cancer 2023, 54, 913–926. [Google Scholar] [CrossRef]
- Kaneda, A.; Yagi, K. Two Groups of DNA Methylation Markers to Classify Colorectal Cancer into Three Epigenotypes. Cancer Sci. 2011, 102, 18–24. [Google Scholar] [CrossRef]
- Hiraoka, S.; Kato, J.; Tatsukawa, M.; Harada, K.; Fujita, H.; Morikawa, T.; Shiraha, H.; Shiratori, Y. Laterally Spreading Type of Colorectal Adenoma Exhibits a Unique Methylation Phenotype and K-Ras Mutations. Gastroenterology 2006, 131, 379–389. [Google Scholar] [CrossRef]
- Hawkins, N.J.; Ward, R.L. Sporadic Colerectal Cancers with Microsatellite Instability and Their Possible Origin in Hyperplastic Polyps and Serrated Adenomas. J. Natl. Cancer Inst. 2001, 93, 1307–1313. [Google Scholar] [CrossRef]
- Takahashi, T.; Nosho, K.; Yamamoto, H.; Mikami, M.; Taniguchi, H.; Miyamoto, N.; Adachi, Y.; Itoh, F.; Imai, K.; Shinomura, Y. Flat-Type Colorectal Advanced Adenomas (Laterally Spreading Tumors) Have Different Genetic and Epigenetic Alterations from Protruded-Type Advanced Adenomas. Mod. Pathol. 2007, 20, 139–147. [Google Scholar] [CrossRef][Green Version]
- Esteller, M.; Toyota, M.; Sanchez-Cespedes, M.; Capella, G.; Peinado, M.A.; Watkins, D.N.; Issa, J.P.J.; Sidransky, D.; Baylin, S.B.; Herman, J.G. Inactivation of the DNA Repair Gene O6-Methylguanine-DNA Methyltransferase by Promoter Hypermethylation Is Associated with G to A Mutations in K-Ras in Colorectal Tumorigenesis. Cancer Res. 2000, 60, 2368–2371. [Google Scholar]
- Sakamoto, N.; Terai, T.; Ajioka, Y.; Abe, S.; Kobayasi, O.; Hirai, S.; Hino, O.; Watanabe, H.; Sato, N.; Shimoda, T.; et al. Frequent Hypermethylation of RASSF1A in Early Flat-Type Colorectal Tumors. Oncogene 2004, 23, 8900–8907. [Google Scholar] [CrossRef] [PubMed]
- Ni, H.-B.; Wang, F.-Y.; Xu, J.; He, X.-J.; Chen, J.; Wu, Q.; Wu, J.-F.; Sun, Y.-S. Screening and Identification of a Tumor Specific Methylation Phenotype in the Colorectal Laterally Spreading Tumor. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 2611–2616. [Google Scholar] [PubMed]
- Iwaizumi, M.; Taniguchi, T.; Kurachi, K.; Osawa, S.; Sugimoto, K.; Baba, S.; Sugimura, H.; Maekawa, M. Methylation of CpG Island Promoters at ZNF625, LONRF2, SDC2 and WDR17 in a Patient with Numerous Non-Granular Type Laterally Spreading Tumors and Colorectal Cancer: A Case Report. Oncol. Lett. 2023, 25, 14. [Google Scholar] [CrossRef]
- Oh, T.; Kim, N.; Moon, Y.; Kim, M.S.; Hoehn, B.D.; Park, C.H.; Kim, T.S.; Kim, N.K.; Chung, H.C.; An, S. Genome-Wide Identification and Validation of a Novel Methylation Biomarker, SDC2, for Blood-Based Detection of Colorectal Cancer. J. Mol. Diagn. 2013, 15, 498–507. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Zhao, J.; Li, Z.; Pan, C.; Liu, J.; Zheng, K.; Wang, X.; Zhang, Y. Research Progress on the Role of Zinc Finger Protein in Colorectal Cancer. Cancer Rep. 2025, 8, e70123. [Google Scholar] [CrossRef]
- Lin, P.; Lin, J.; Lin, C.; Lin, H.; Yang, S. Clinical Relevance of Plasma DNA Methylation in Colorectal Cancer Patients Identified by Using a Genome-Wide High-Resolution Array. Ann. Surg. Oncol. 2015, 22 (Suppl. S3), 1419–1427. [Google Scholar] [CrossRef]
- Hua, Y.; Ma, X.; Liu, X.; Yuan, X.; Qin, H.; Zhang, X. Abnormal Expression of MRNA, MicroRNA Alteration and Aberrant DNA Methylation Patterns in Rectal Adenocarcinoma. PLoS ONE 2017, 12, e0174461. [Google Scholar] [CrossRef]
- Zhu, L.; Yan, F.; Wang, Z.; Dong, H.; Bian, C.; Wang, T.; Yu, E.; Li, J. Genome-Wide DNA Methylation Profiling of Primary Colorectal Laterally Spreading Tumors Identifies Disease-Specific Epimutations on Common Pathways. Int. J. Cancer 2018, 143, 2488–2498. [Google Scholar] [CrossRef]

| Mutation | Pathway/Function | LST |
Adenoma/Polypoid Carcinoma | Correlation | References |
|---|---|---|---|---|---|
| KRAS | MAPK signaling pathways | G-LSTs in distal colon | More frequent in early-stage carcinomas and protruded type adenomas | COX-2 overexpression Gastrin overexpression | [27,67,68,72,76,87] |
| APC (5q chromosome) | Wnt signaling pathway | LOH at 5q chromosome in NG-LTSs | Loss of function of APC | High levels of CCND1 and c-Myc Nuclear β-catenin accumulation β-catenin resistant to ubiquitination Adherence proteins expression Missense mutation of MED12L | [22,76] |
| β-catenin | High expression Low level of mutation of CTNNB1 in NG-LSTs | Low expression High levels of CTNNB1 mutations | [22,37,38,42,56] | ||
| TP53 (17p chromosome) | DNA repair/genome stability | Mostly in NG-LSTs | Undetected in adenomas, but detected in cancer derived from malignant progression of LSTs LOH 17p chromosome is crucial in progression from adenoma to cancer | [56,69,71,76] | |
| PIK3CA | AKT signaling pathway | LSTs with high pathological grades Mostly in G-LSTs | Present in CRC (10–30%), but not detected in polypoid/depressed and flat-elevated adenomas | KRAS mutation | [88] |
| BRAF | RAS/RAF/MEK/MAPK signaling pathway | Rare in LSTs | Rare in CRC | Mutually exclusive with KRAS mutation Concurrent mutation with KRAS | [76] |
| SLITRK, SLIT, NTRK genes family | Axonal guidance pathway | Frequently mutated | Not in polypoid adenomas, but mutated in CRC | [80] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martinelli, M.; Cascelli, N.; Bartolo, O.; Ciuffi, M.; Mazzoccoli, C.; Dieli, R.; Lioy, R.; Landriscina, M.; Calabrese, C.; Crispo, F. The Molecular Landscape of Colorectal Laterally Spreading Tumors: From Endoscopic Subtypes to Molecular Targets. Int. J. Mol. Sci. 2025, 26, 8445. https://doi.org/10.3390/ijms26178445
Martinelli M, Cascelli N, Bartolo O, Ciuffi M, Mazzoccoli C, Dieli R, Lioy R, Landriscina M, Calabrese C, Crispo F. The Molecular Landscape of Colorectal Laterally Spreading Tumors: From Endoscopic Subtypes to Molecular Targets. International Journal of Molecular Sciences. 2025; 26(17):8445. https://doi.org/10.3390/ijms26178445
Chicago/Turabian StyleMartinelli, Mara, Nicoletta Cascelli, Ottavia Bartolo, Mario Ciuffi, Carmela Mazzoccoli, Rosalia Dieli, Rosa Lioy, Matteo Landriscina, Carlo Calabrese, and Fabiana Crispo. 2025. "The Molecular Landscape of Colorectal Laterally Spreading Tumors: From Endoscopic Subtypes to Molecular Targets" International Journal of Molecular Sciences 26, no. 17: 8445. https://doi.org/10.3390/ijms26178445
APA StyleMartinelli, M., Cascelli, N., Bartolo, O., Ciuffi, M., Mazzoccoli, C., Dieli, R., Lioy, R., Landriscina, M., Calabrese, C., & Crispo, F. (2025). The Molecular Landscape of Colorectal Laterally Spreading Tumors: From Endoscopic Subtypes to Molecular Targets. International Journal of Molecular Sciences, 26(17), 8445. https://doi.org/10.3390/ijms26178445







