Anandamide Alters Glycolytic Activity in Streptococcus mutans: Metabolomics and Stable Isotope Labeling Study
Abstract
1. Introduction
2. Results
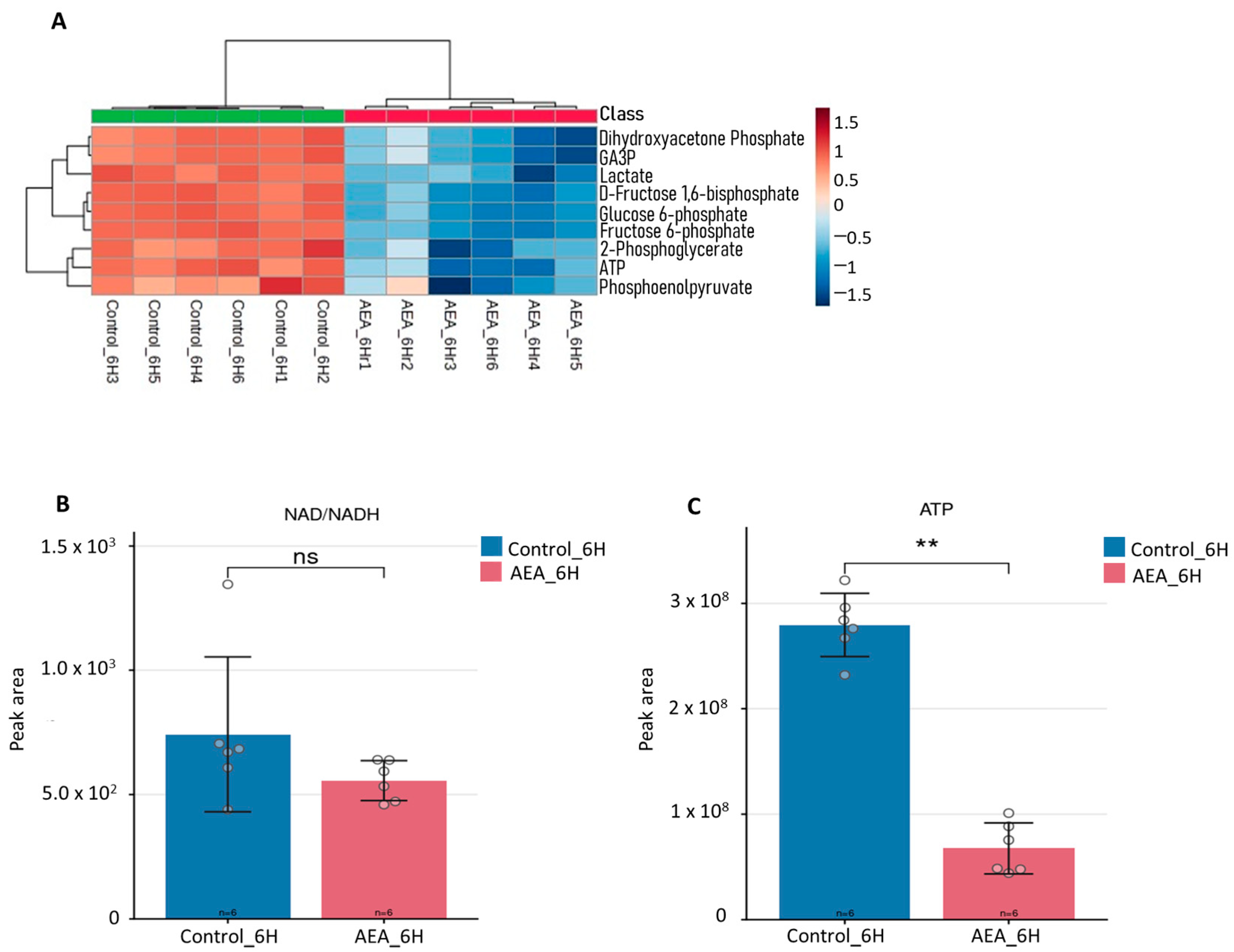
2.1. Metabolic Profiling of S. mutans Treated with AEA
2.2. Key Glycolytic Intermediates Are Suppressed
2.2.1. NAD+/NADH Imbalance Reflects Redox Disruption
2.2.2. Compromised Adenosine Triphosphate (ATP) Production
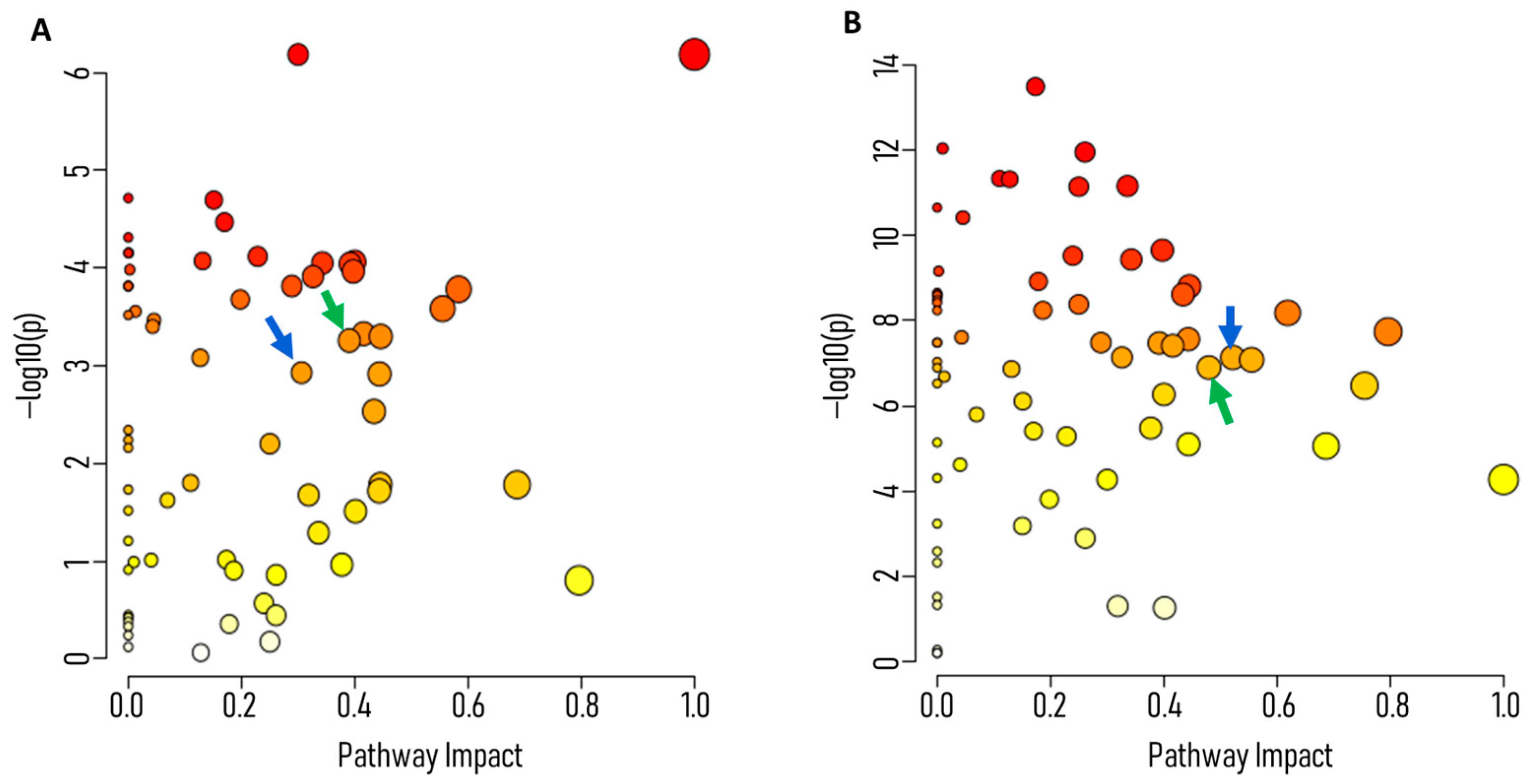
2.3. Pathway Enrichment and Impact Analysis
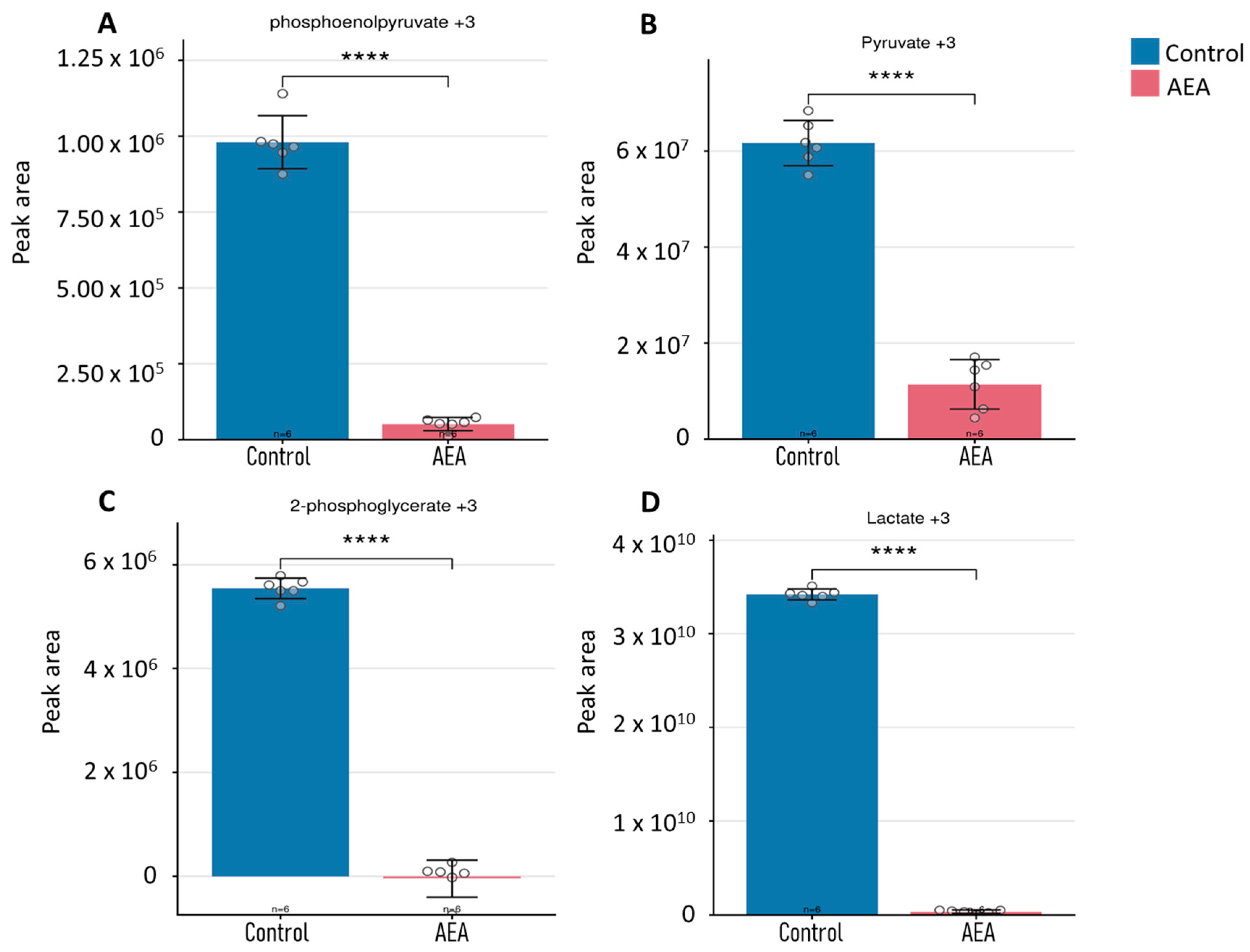
2.4. Tracing of Intracellular Metabolites
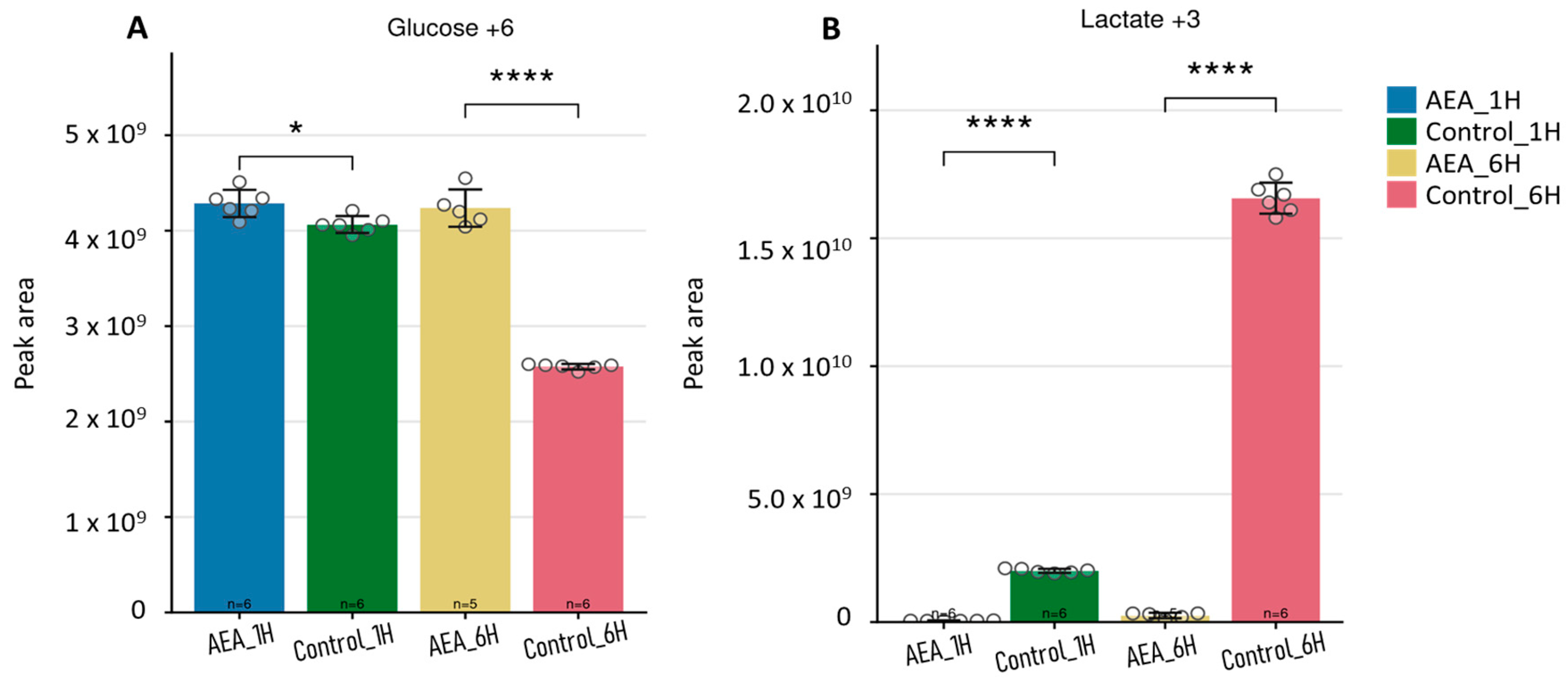
2.5. Tracing Analysis of Glucose and Lactate in the Growth Media
2.6. Flux Analysis of Intracellular Glycolytic Intermediates in the Growth Media
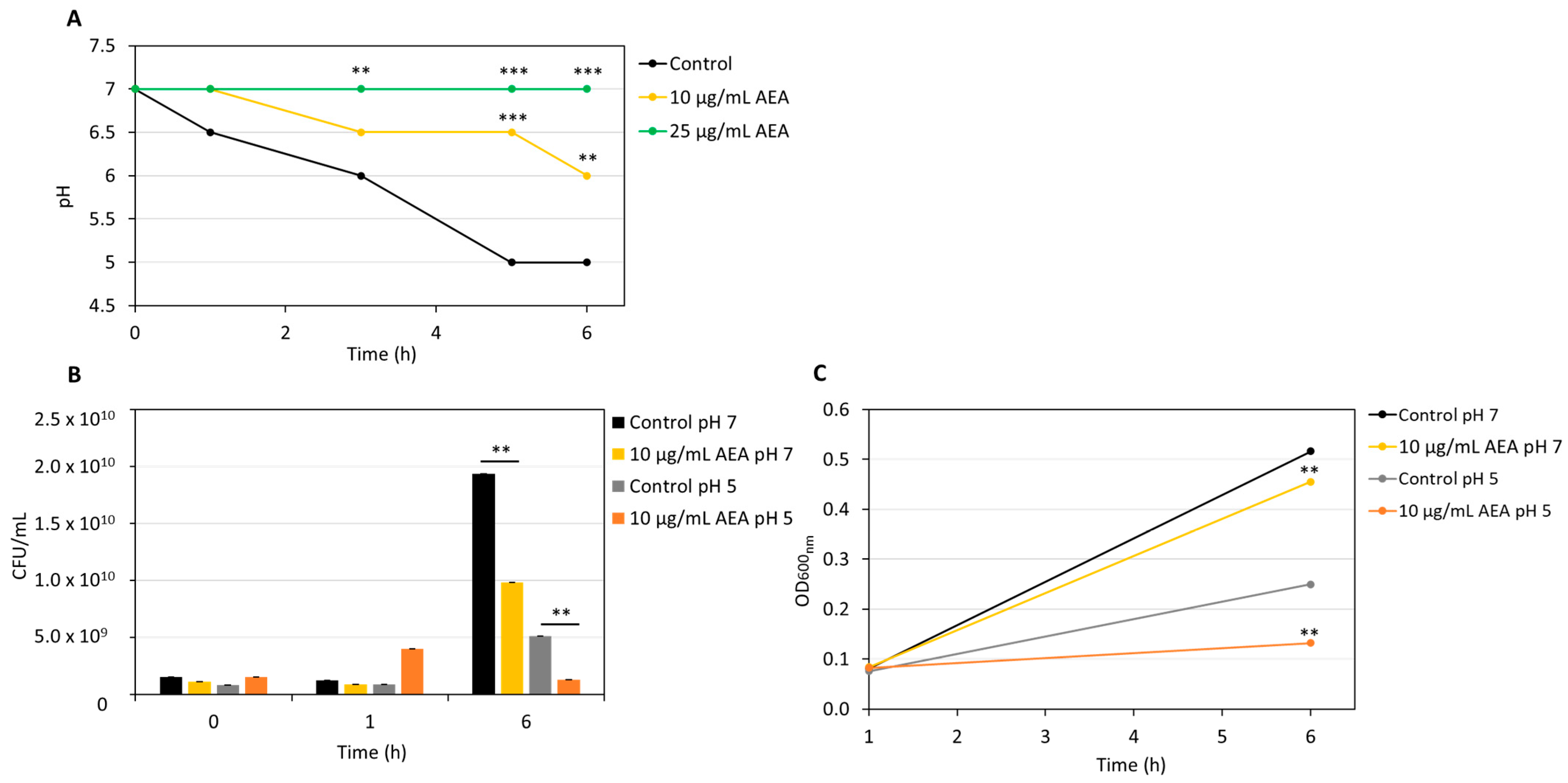
2.7. AEA Treatment Reduces Growth Rate, Aciduricity, and Acidogenicity
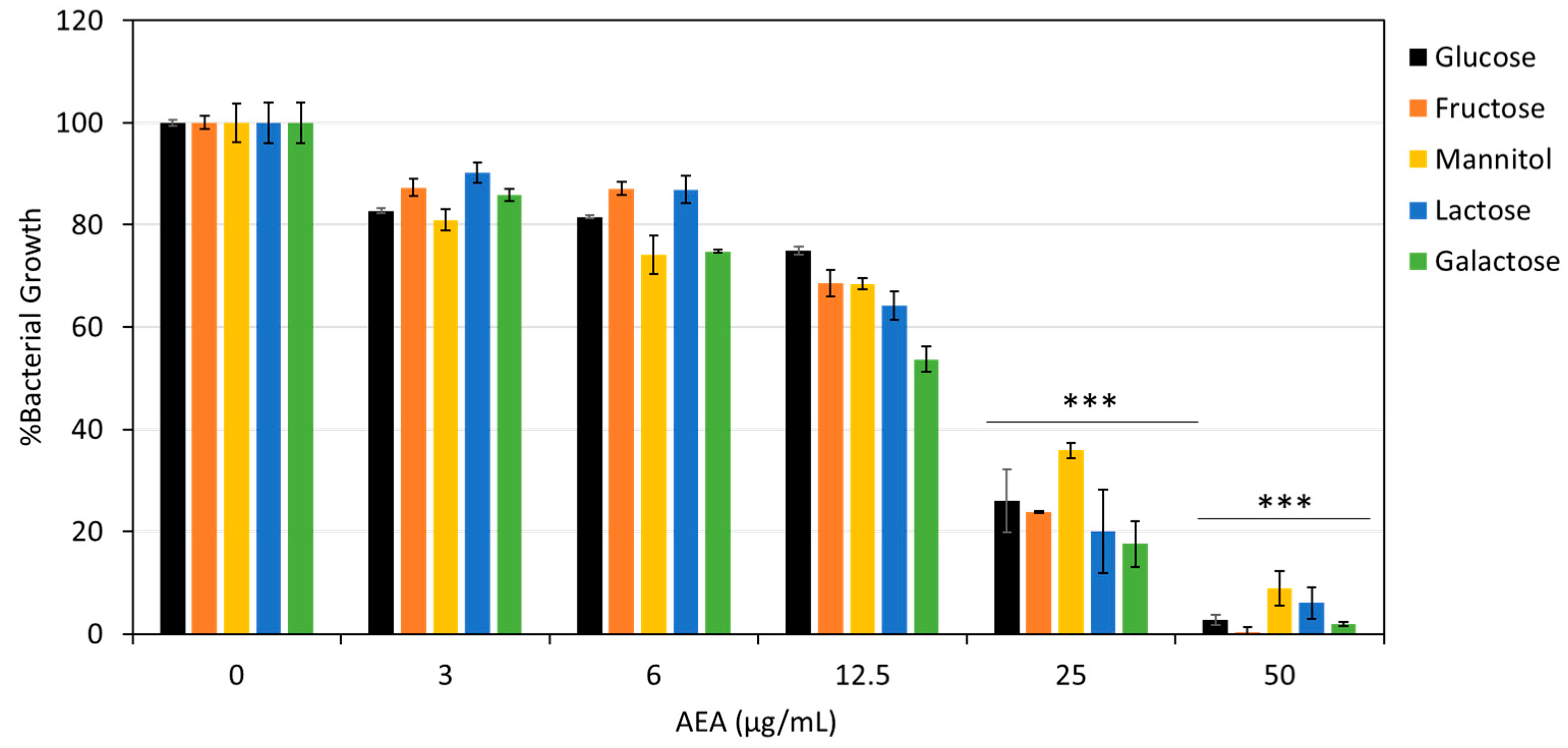
2.8. The Effect of AEA on Carbohydrate Uptake and Glycolysis
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Bacterial Growth Media
4.3. Bacterial Growth
4.4. LC-MS-Based Metabolomics
4.4.1. Sample Preparation
4.4.2. LC-MS Data Acquisition
4.4.3. Metabolomics Data Analysis
4.4.4. Metabolomics Statistical Analysis
4.4.5. Metabolomics Outlier Identification and Quality Control
4.5. Kinetic Analysis of Planktonic Bacteria and pH Measurements
4.6. Acid Tolerance Assay
4.7. Carbohydrate Rescue Assay
4.8. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| S. mutans | Streptococcus mutans |
| AEA | Anandamide |
| MIC | Minimum inhibitory concentration |
| ATP | Adenosine triphosphate |
| LC-MS | Liquid chromatography–mass spectrometry |
| 2-AG | 2-arachidonoylglycerol |
| CB1/CB2 | Cannabinoid receptor 1/2 |
| NAPE | N-acyl-phosphatidylethanolamie |
| PIP2 | Phosphatidylinositol-4,5-bisphosphate |
| GPCRs | G-protein-coupled receptors |
| MRSA | Methicillin-resistant Staphylococcus aureus |
| TSB | Tryptic soy broth |
| TSBG | Tryptic soy broth with 1% glucose |
| OD | Optical density |
| DDW | Double-distilled water |
| PBS | Phosphate-buffered saline |
| HPLC | High-performance liquid chromatography |
| QC | Quality control |
| UPLC | Ultimate 300 high-performance liquid chromatography |
| m/z | Mass/charge ratio |
| PCA | Principal component analysis |
| BHI | Brain heart infusion |
| CFUs | Colony-forming units |
| G6P | Glucose-6-phosphate |
| F6P | Fructose-6-phosphate |
| F1,6BP | Fructose-1,6-bisphosphate |
| 2-PG | 2-phosphoglycerate |
| PEP | Phosphoenolpyruvate |
| GA3P | Glyceraldehyde-3-phosphate |
| PPP | Pentose phosphate pathway |
| EPS | Extracellular polysaccharides |
| X5P | Xylitol-5-phosphate |
| PTS | Phosphotransferase system |
| EI | Enzyme I |
| EII | Enzyme II |
| HPr | Heat-stable phosphocarrier protein |
| ABCs | ATP-binding cassettes |
| PDH | Pyruvate dehydrogenase |
References
- Sionov, R.V.; Steinberg, D. Anti-Microbial Activity of Phytocannabinoids and Endocannabinoids in the Light of Their Physiological and Pathophysiological Roles. Biomedicines 2022, 10, 631. [Google Scholar] [CrossRef]
- Devane, W.A.; Hanuš, L.; Breuer, A.; Pertwee, R.G.; Stevenson, L.A.; Griffin, G.; Gibson, D.; Mandelbaum, A.; Etinger, A.; Mechoulam, R. Isolation and Structure of a Brain Constituent That Binds to the Cannabinoid Receptor. Science 1992, 258, 1946–1949. [Google Scholar] [CrossRef]
- Wei, D.; Allsop, S.; Tye, K.; Piomelli, D. Endocannabinoid Signaling in the Control of Social Behavior. Trends Neurosci. 2017, 40, 385–396. [Google Scholar] [CrossRef]
- Vickstrom, C.R.; Liu, X.; Liu, S.; Hu, M.M.; Mu, L.; Hu, Y.; Yu, H.; Love, S.L.; Hillard, C.J.; Liu, Q.S. Role of Endocannabinoid Signaling in a Septohabenular Pathway in the Regulation of Anxiety- and Depressive-like Behavior. Mol. Psychiatry 2021, 26, 3178–3191. [Google Scholar] [CrossRef]
- Yin, A.Q.; Wang, F.; Zhang, X. Integrating Endocannabinoid Signaling in the Regulation of Anxiety and Depression. Acta Pharmacol. Sin. 2019, 40, 336–341. [Google Scholar] [CrossRef]
- Banerjee, S.; Sionov, R.V.; Feldman, M.; Smoum, R.; Mechoulam, R.; Steinberg, D. Anandamide Alters the Membrane Properties, Halts the Cell Division and Prevents Drug Efflux in Multidrug Resistant Staphylococcus aureus. Sci. Rep. 2021, 11, 8690. [Google Scholar] [CrossRef] [PubMed]
- Feldman, M.; Smoum, R.; Mechoulam, R.; Steinberg, D. Antimicrobial Potential of Endocannabinoid and Endocannabinoid-like Compounds against Methicillin-Resistant Staphylococcus aureus. Sci. Rep. 2018, 8, 17696. [Google Scholar] [CrossRef]
- Sionov, R.V.; Banerjee, S.; Bogomolov, S.; Smoum, R.; Mechoulam, R.; Steinberg, D. Targeting the Achilles’ Heel of Multidrug-Resistant Staphylococcus aureus by the Endocannabinoid Anandamide. Int. J. Mol. Sci. 2022, 23, 7798. [Google Scholar] [CrossRef] [PubMed]
- Cravatt, B.F.; Demarest, K.; Patricelli, M.P.; Bracey, M.H.; Giang, D.K.; Martin, B.R.; Lichtman, A.H. Supersensitivity to Anandamide and Enhanced Endogenous Cannabinoid Signaling in Mice Lacking Fatty Acid Amide Hydrolase. Proc. Natl. Acad. Sci. USA 2001, 98, 9371–9376. [Google Scholar] [CrossRef]
- Fornelos, N.; Franzosa, E.A.; Bishai, J.; Annand, J.W.; Oka, A.; Lloyd-Price, J.; Arthur, T.D.; Garner, A.; Avila-Pacheco, J.; Haiser, H.J.; et al. Growth Effects of N-Acylethanolamines on Gut Bacteria Reflect Altered Bacterial Abundances in Inflammatory Bowel Disease. Nat. Microbiol. 2020, 5, 486–497. [Google Scholar] [CrossRef] [PubMed]
- Sionov, R.V.; Feldman, M.; Smoum, R.; Mechoulam, R.; Steinberg, D. Anandamide Prevents the Adhesion of Filamentous Candida albicans to Cervical Epithelial Cells. Sci. Rep. 2020, 10, 13728. [Google Scholar] [CrossRef]
- Wolfson, G.; Sionov, R.V.; Smoum, R.; Korem, M.; Polacheck, I.; Steinberg, D. Anti-Bacterial and Anti-Biofilm Activities of Anandamide against the Cariogenic Streptococcus mutans. Int. J. Mol. Sci. 2023, 24, 6177. [Google Scholar] [CrossRef]
- Baker, J.L.; Faustoferri, R.C.; Quivey, R.G. Acid-Adaptive Mechanisms of Streptococcus mutans–the More We Know, the More We Don’t. Mol. Oral Microbiol. 2017, 32, 107–117, Corrigendum in: Mol. Oral Microbiol. 2017, 32, 354. [Google Scholar] [CrossRef] [PubMed]
- Dashper, S.G.; Reynolds, E.C. Lactic Acid Excretion by Streptococcus mutans. Microbiology 1996, 142, 33–39. [Google Scholar] [CrossRef] [PubMed]
- Lemos, J.A.; Palmer, S.R.; Zeng, L.; Wen, Z.T.; Kajfasz, J.K.; Freires, I.A.; Abranches, J.; Brady, L.J. The Biology of Streptococcus Mutans. Microbiol. Spectr. 2018, 7, 10–1128. [Google Scholar] [CrossRef]
- Krzyściak, W.; Jurczak, A.; Kościelniak, D.; Bystrowska, B.; Skalniak, A. The Virulence of Streptococcus mutans and the Ability to Form Biofilms. Eur. J. Clin. Microbiol. Infect. Dis. 2014, 33, 499–515. [Google Scholar] [CrossRef]
- Zhang, H.; Mu, R.; Wang, Z.; Peng, S.; Yang, X.Y.; Qin, X. Trans-Cinnamaldehyde Inhibition of Pyruvate Dehydrogenase: Effects on Streptococcus mutans Carbohydrate Metabolism. J. Proteome Res. 2024, 23, 3682–3695. [Google Scholar] [CrossRef]
- Miyasawa, H.; Iwami, Y.; Mayanagi, H.; Takahashi, N. Xylitol Inhibition of Anaerobic Acid Production by Streptococcus mutans at Various PH Levels. Oral Microbiol. Immunol. 2003, 18, 215–219. [Google Scholar] [CrossRef]
- Lin, Y.; Gong, T.; Ma, Q.; Jing, M.; Zheng, T.; Yan, J.; Chen, J.; Pan, Y.; Sun, Q.; Zhou, X.; et al. Nicotinamide Could Reduce Growth and Cariogenic Virulence of Streptococcus mutans. J. Oral Microbiol. 2022, 14, 2056291. [Google Scholar] [CrossRef] [PubMed]
- Maehara, H.; Iwami, Y.; Mayanagi, H.; Takahashi, N. Synergistic Inhibition by Combination of Fluoride and Xylitol on Glycolysis by Mutans Streptococci and Its Biochemical Mechanism. Caries Res. 2005, 39, 521–528. [Google Scholar] [CrossRef]
- Daboor, S.M.; Masood, F.S.S.; Al-Azab, M.S.; Nori, E.E. A Review On Streptococcus mutans With Its Diseases Dental Caries, Dental Plaque And Endocarditis. Indian J. Microbiol. Res. 2015, 2, 76–82. [Google Scholar]
- Queiroz, V.S.D.; Ccahuana-Vásquez, R.A.; Tedesco, A.F.; Lyra, L.; Cury, J.A.; Schreiber, A.Z. Influence of the Culture Medium in Dose-Response Effect of the Chlorhexidine on Streptococcus mutans Biofilms. Scientifica 2016, 2016, 2816812. [Google Scholar] [CrossRef] [PubMed]
- Chamlagain, M.; Hu, J.; Sionov, R.V.; Steinberg, D. Anti-Bacterial and Anti-Biofilm Activities of Arachidonic Acid against the Cariogenic Bacterium Streptococcus Mutans. Front. Microbiol. 2024, 15, 1333274. [Google Scholar] [CrossRef] [PubMed]
- Rewak-Soroczynska, J.; Dorotkiewicz-Jach, A.; Drulis-Kawa, Z.; Wiglusz, R.J. Culture Media Composition Influences the Antibacterial Effect of Silver, Cupric, and Zinc Ions against Pseudomonas aeruginosa. Biomolecules 2022, 12, 963. [Google Scholar] [CrossRef]
- Bock, L.J.; Hind, C.K.; Sutton, J.M.; Wand, M.E. Growth Media and Assay Plate Material Can Impact on the Effectiveness of Cationic Biocides and Antibiotics against Different Bacterial Species. Lett. Appl. Microbiol. 2018, 66, 368–377. [Google Scholar] [CrossRef]
- Dashper, S.G.; Reynolds, E.C. pH Regulation by Streptococcus mutans. J. Dent. Res. 1992, 71, 1159–1165. [Google Scholar] [CrossRef]
- Thevenot, T.; Brochu, D.; Vadeboncoeur, C.; Hamilton, I.R. Regulation of ATP-Dependent P-(Ser)-HPr Formation in Streptococcus mutans and Streptococcus salivarius. J. Bacteriol. 1995, 177, 2751–2759. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, M.; Yu, W.; Zhou, S.; Zhang, B.; Lo, E.C.M.; Xu, X.; Zhang, D. In Vitro Antibacterial Activity of an FDA-Approved H+-ATPase Inhibitor, Bedaquiline, Against Streptococcus mutans in Acidic Milieus. Front. Microbiol. 2021, 12, 647611. [Google Scholar] [CrossRef]
- Sekiya, M.; Izumisawa, S.; Iwamoto-Kihara, A.; Fan, Y.; Shimoyama, Y.; Sasaki, M.; Nakanishi-Matsui, M. Proton-Pumping F-ATPase Plays an Important Role in Streptococcus mutans under Acidic Conditions. Arch. Biochem. Biophys. 2019, 666, 46–51. [Google Scholar] [CrossRef]
- Baker, J.L.; Derr, A.M.; Faustoferri, R.C.; Quivey, R.G. Loss of NADH Oxidase Activity in Streptococcus mutans Leads to Rex-Mediated Overcompensation in NAD+ Regeneration by Lactate Dehydrogenase. J. Bacteriol. 2015, 197, 3645–3657. [Google Scholar] [CrossRef]
- Abranches, J.; Candella, M.M.; Wen, Z.T.; Baker, H.V.; Burne, R.A. Different Roles of EIIABMan and EIIGlc in Regulation of Energy Metabolism, Biofilm Development, and Competence in Streptococcus mutans. J. Bacteriol. 2006, 188, 3748–3756. [Google Scholar] [CrossRef] [PubMed]
- Iwami, Y.; Schachtele, C.F.; Yamada, T. Mechanism of Inhibition of Glycolysis in Streptococcus mutans NCIB 11723 by Chlorhexidine. Oral Microbiol. Immunol. 1995, 10, 360–364. [Google Scholar] [CrossRef] [PubMed]
- Phan, T.N.; Marquis, R.E. Triclosan Inhibition of Membrane Enzymes and Glycolysis of Streptococcus mutans in Suspensions and Biofilms. Can. J. Microbiol. 2006, 52, 977–983. [Google Scholar] [CrossRef] [PubMed]
- Mielko, K.A.; Jabłoński, S.J.; Łukaszewicz, M.; Młynarz, P. Comparison of Bacteria Disintegration Methods and Their Influence on Data Analysis in Metabolomics. Sci. Rep. 2021, 11, 20859. [Google Scholar] [CrossRef] [PubMed]
- MacKay, G.M.; Zheng, L.; Van Den Broek, N.J.F.; Gottlieb, E. Analysis of Cell Metabolism Using LC-MS and Isotope Tracers. In Methods Enzymol; Academic Press Inc.: Cambridge, MA, USA, 2015; Volume 561, pp. 171–196. [Google Scholar]
- Alam, S.; Brailsford, S.R.; Adams, S.; Allison, C.; Sheehy, E.; Zoitopoulos, L.; Kidd, E.A.; Beighton, A.D. Genotypic Heterogeneity of Streptococcus oralis and Distinct Aciduric Subpopulations in Human Dental Plaque. Appl. Environ. Microbiol. 2000, 66, 3330–3336. [Google Scholar] [CrossRef]
- Wilkins, J.C.; Homer, K.A.; Beighton, D. Analysis of Streptococcus mutans Proteins Modulated by Culture under Acidic Conditions. Appl. Environ. Microbiol. 2002, 68, 2382–2390. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wolfson, G.; Steinberg, D.; Eliassaf, A.; Morshina, A.; Enríquez-Rodríguez, C.J.; Polacheck, I.; Korem, M.; Shalev, O. Anandamide Alters Glycolytic Activity in Streptococcus mutans: Metabolomics and Stable Isotope Labeling Study. Int. J. Mol. Sci. 2025, 26, 8401. https://doi.org/10.3390/ijms26178401
Wolfson G, Steinberg D, Eliassaf A, Morshina A, Enríquez-Rodríguez CJ, Polacheck I, Korem M, Shalev O. Anandamide Alters Glycolytic Activity in Streptococcus mutans: Metabolomics and Stable Isotope Labeling Study. International Journal of Molecular Sciences. 2025; 26(17):8401. https://doi.org/10.3390/ijms26178401
Chicago/Turabian StyleWolfson, Goldie, Doron Steinberg, Alexandra Eliassaf, Anna Morshina, César Jessé Enríquez-Rodríguez, Itzhack Polacheck, Maya Korem, and Ori Shalev. 2025. "Anandamide Alters Glycolytic Activity in Streptococcus mutans: Metabolomics and Stable Isotope Labeling Study" International Journal of Molecular Sciences 26, no. 17: 8401. https://doi.org/10.3390/ijms26178401
APA StyleWolfson, G., Steinberg, D., Eliassaf, A., Morshina, A., Enríquez-Rodríguez, C. J., Polacheck, I., Korem, M., & Shalev, O. (2025). Anandamide Alters Glycolytic Activity in Streptococcus mutans: Metabolomics and Stable Isotope Labeling Study. International Journal of Molecular Sciences, 26(17), 8401. https://doi.org/10.3390/ijms26178401






