1. Introduction
Erythropoiesis, the process of forming red blood cells, is crucial for oxygen delivery during fetal development and throughout adult life. During human embryogenesis, hematopoiesis occurs in multiple waves: starting from the yolk sac, followed by migration to the fetal liver, and finally localizing to the bone marrow, which remains the lifelong hematopoietic site after birth [
1]. In the fetal liver, hematopoietic stem cells (HSCs) significantly expand due to a unique microenvironment that supports extensive proliferation, contrasting with the relatively quiescent state of adult bone marrow HSCs [
1]. Specifically, fetal liver-derived CD34
+ hematopoietic stem cells exhibit dramatic proliferation in vitro, reaching a 100-fold higher expansion capacity compared to adult bone marrow-derived CD34
+ cells [
2]. This distinct proliferative potential highlights fundamental differences in the regulatory mechanism between fetal and adult hematopoietic tissues.
Transcriptomic analyses comparing definitive erythropoiesis in fetal liver and adult bone marrow have identified several key molecular differences. For example, erythroid progenitor cells, burst-forming unit-erythroid (BFU-E), from fetal liver exhibit increased expression of genes associated with cell division and proliferation, and display distinct transcriptomic signatures compared to their bone marrow counterparts significantly higher expression of genes associated with cell cycle progression and proliferation, suggesting a robust cell division program to support rapid fetal growth [
3]. Specifically, fetal liver erythroid progenitors display enhanced expression of key cell-cycle regulators such as glucocorticoid receptor (Nr3c1), Myc, and cyclin A2 (Ccna2), which promote rapid cell proliferation and expansion of the erythroid lineage during fetal development. During later stages of differentiation, terminal erythropoiesis, fetal liver erythroblasts upregulate biological processes associated with translation, indicating increased protein synthesis capacity. Moreover, pathways related to the tricarboxylic acid (TCA) cycle and hypoxia response are notably enriched, highlighting the metabolic adaptations of fetal erythroid cells necessary for efficient proliferation and hemoglobin production within the developing fetal liver environment. Additionally, fetal liver-derived erythroid cells predominantly express fetal and embryonic hemoglobin, in contrast to bone marrow-derived cells, which shift toward adult hemoglobin during maturation. This expression pattern is linked to decreased BCL11A, a key repressor of γ-globin, and elevated levels of LIN28B and IGF2BP1 in fetal liver erythroblasts, thereby sustaining high HbF production [
2,
4,
5].
Despite these known differences at the transcriptomic and cellular level, the regulatory roles of non-coding RNAs, particularly circular RNAs (circRNAs), have not been systematically explored in fetal versus adult human erythropoiesis. Given the emerging evidence that circRNAs play significant roles in gene regulation and hematopoietic differentiation, exploring circRNA expression and associated regulatory networks in erythroblasts differentiated from fetal liver and adult bone marrow-derived CD34+ hematopoietic stem cells can provide important new insights into developmental erythropoiesis.
circRNAs are endogenous RNA molecules characterized by a covalently closed loop structure. They are primarily generated through a non-canonical splicing process known as backsplicing, in which a downstream splice donor site is joined to an upstream splice acceptor site, resulting in the formation of a circular RNA transcript [
6,
7]. Due to their closed-loop structure, circRNAs are more stable than linear RNAs, resisting degradation by exonucleases [
7,
8,
9]. CircRNAs are generated through spliceosome-mediated backsplicing of exonic, intronic, or exon–intron sequences, a process influenced by specific RNA-binding proteins such as QKI, FUS, NF90/NF110, and ADAR, which can promote or inhibit their formation [
6,
7,
8,
10]. Functionally, circRNAs often act as miRNA sponges, binding and sequestering miRNAs to indirectly regulate gene expression [
6,
7]. Additionally, circRNAs interact with RBPs, contributing to post-transcriptional regulation, RNA stability, localization, and even facilitating circRNA translation into functional proteins when internal ribosome entry sites (IRES) are present [
6,
8]. circRNAs exhibit tissue-specific expression patterns and can be differentially regulated in various diseases, including cancers, neurological disorders, cardiovascular diseases, and inflammatory conditions [
6,
7]. For example, circRNAs such as circPVT1 and circHIPK3 have been implicated in leukemogenesis and other cancers, highlighting their potential as biomarkers or therapeutic targets [
3]. Detection and quantification of circRNAs typically rely on RNA-sequencing methods using ribosomal RNA depletion strategies, complemented by bioinformatic algorithms to identify backsplice junctions, while experimental validation includes RNase R treatment and quantitative PCR (qPCR) with circRNA-specific primers [
6,
7].
CircAtlas 3.0 is a comprehensive resource containing more than three million curated circRNAs from 10 vertebrate species and 33 tissues. By integrating data from both Illumina and Nanopore sequencing platforms, CircAtlas 3.0 enables reconstruction of full-length circRNA isoforms and provides extensive functional annotations, including predicted miRNA- and RBP-binding sites, IRES/ORF predictions, and disease associations. This breadth of information makes CircAtlas 3.0 an essential tool for investigating circRNA biology and its clinical relevance [
10,
11].
Collectively, circRNAs represent a potential regulatory layer in hemoglobin and erythropoiesis regulation, with emerging roles across normal physiology and various diseases.
3. Discussion
Despite the growing recognition of circRNAs as important regulators in various biological processes, relatively few studies have explored their roles in erythropoiesis and hemoglobin regulation, particularly during human development [
10,
12,
13,
14].
In β-thalassemia, transcriptome profiling has identified more than 2000 dysregulated circRNAs, many of which are linked to erythroid differentiation and globin gene regulation [
13]. Among these, hsa_circRNA_100466, also known as hsa-LBR_0002 in circAtlas and circLBR(7,8).1 as its uniform ID [
10], emerged as a key regulator of HbF through a circRNA-100466/miR-19b-3p/SOX6 axis, where SOX6 functions as a known repressor of γ-globin, thereby directly connecting circRNA activity to HbF silencing [
13].
Beyond mechanistic networks, circRNAs also hold diagnostic potential. For instance, circ_0008102, also known as hsa-LCOR_0002 in circAtlas and circLCOR(5,6,7).1 as its uniform ID [
10], was found to be significantly downregulated in the peripheral blood of β-thalassemia patients, with levels correlating to disease severity [
13]. Diagnostic performance was evaluated for circ_0008102 expression in peripheral blood of pediatric β-thalassemia patients. circ_0008102 distinguished transfusion-independent patients from those requiring transfusion with an AUC of 0.733 (95% CI: 0.590–0.877; sensitivity 72.2%; specificity 68.6%;
p = 0.006) [
13]. Additionally, it differentiated transfusion-independent patients from healthy controls with an AUC of 0.711 (95% CI: 0.554–0.868; sensitivity 72.2%; specificity 69.0%;
p = 0.016) [
13]. These findings suggest that circ-0008102 could be a novel biomarker for identifying transfusion-independent pediatric β-thalassemia patients.
Furthermore, hsa_circ_0005245, also known as hsa-TBCD_0002 in circAtlas and circTBCD(18,19,20,21,22,23).1 as its uniform ID, has been experimentally validated to promote γ-globin expression by sponging miR-425-3p and thereby derepressing GATA2, a transcription factor essential for sustaining HbF production [
14]. Overexpression of hsa_circ_0005245 in K562 shows a significant increase in HbF level, confirming that hsa_circ_0005245/hsa-miR-425-3p/GATA2 enhances γ-globin levels, providing functional evidence of circRNA-mediated regulation [
14]. Collectively, these findings illustrate that circRNAs contribute both to the molecular control of γ-globin switching and to the pathophysiology of hemoglobinopathies, underscoring their significance as potential therapeutic targets and biomarkers in disorders of erythropoiesis.
Although these few recent studies have demonstrated that circRNAs can regulate γ-globin expression or serve as potential biomarkers [
12,
13,
14], the available studies remain limited, and circRNA contributions to hemoglobin regulation and erythropoiesis are only beginning to be defined.
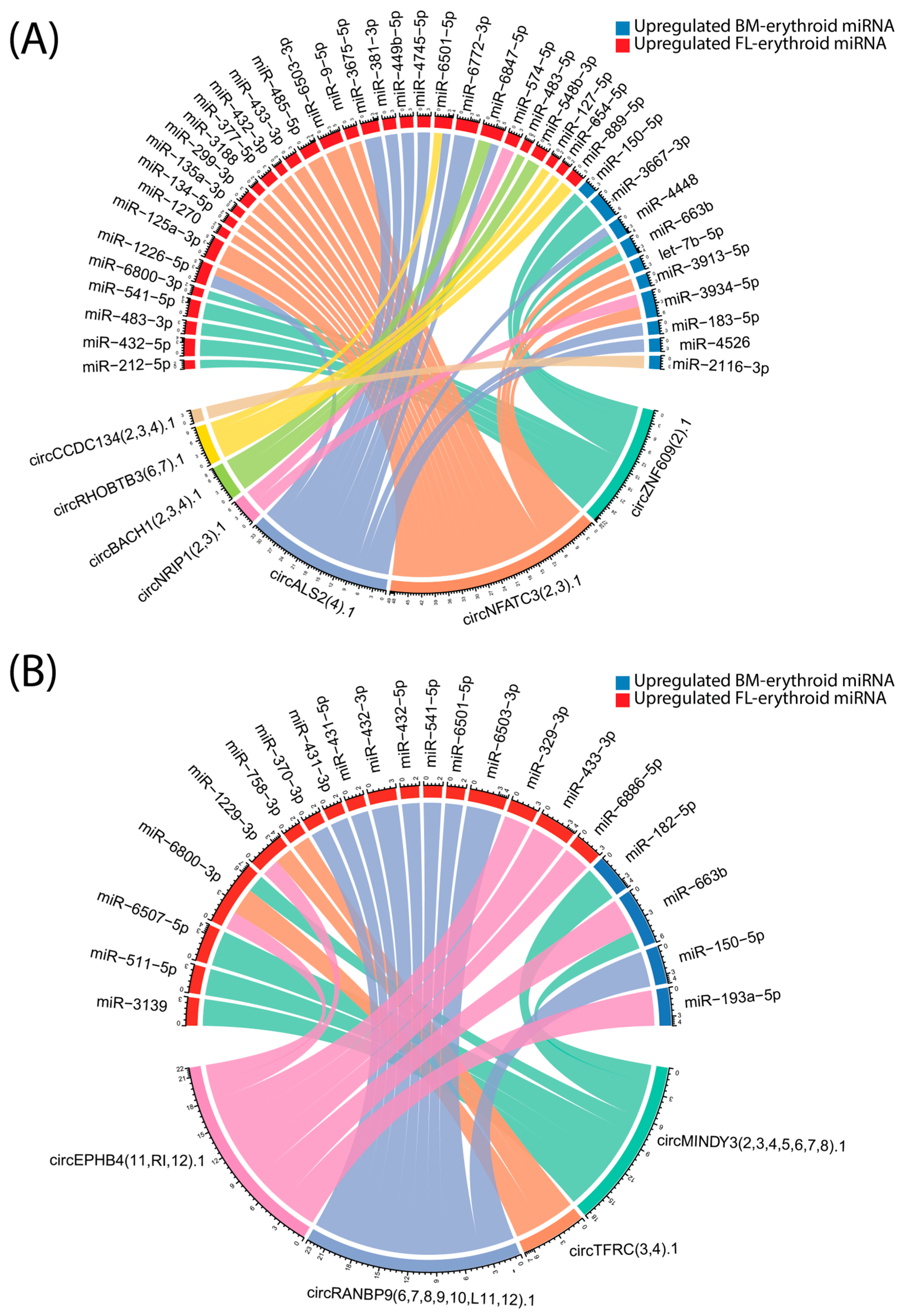
By contrast, most existing research in erythropoiesis has focused on linear transcripts, leaving the broader contribution of circRNAs to erythroid differentiation and globin gene regulation largely unexplored. To address this gap, the present study profiled circRNA expression and predicted interaction networks in erythroblasts derived from fetal liver and bone marrow hematopoietic stem cells, utilizing publicly available total RNA-seq data. While the overall numbers of circRNAs detected per sample were not significantly different between erythroblasts derived from fetal liver and those derived from bone marrow, PCA revealed statistically significant group separation based on circRNA expression profiles, underscoring the existence of distinct circRNA landscapes associated with each developmental origin; however, the effect size was small, indicating that the biological separation between these samples group is marginal. The circRNA–miRNA interaction network analysis highlighted both tissue-restricted and cross-regulatory relationships, with several circRNAs upregulated in bone marrow showing predicted interactions with miRNAs preferentially expressed in either bone marrow or fetal liver. For example, circALS2 and circZNF609 interact with both BM and FL upregulated miRNAs, suggesting a potential role in integrating developmental signals during erythroid maturation. The existence of such cross-regulation may reflect mechanisms that fine-tune gene expression transitions between fetal and adult erythropoiesis.
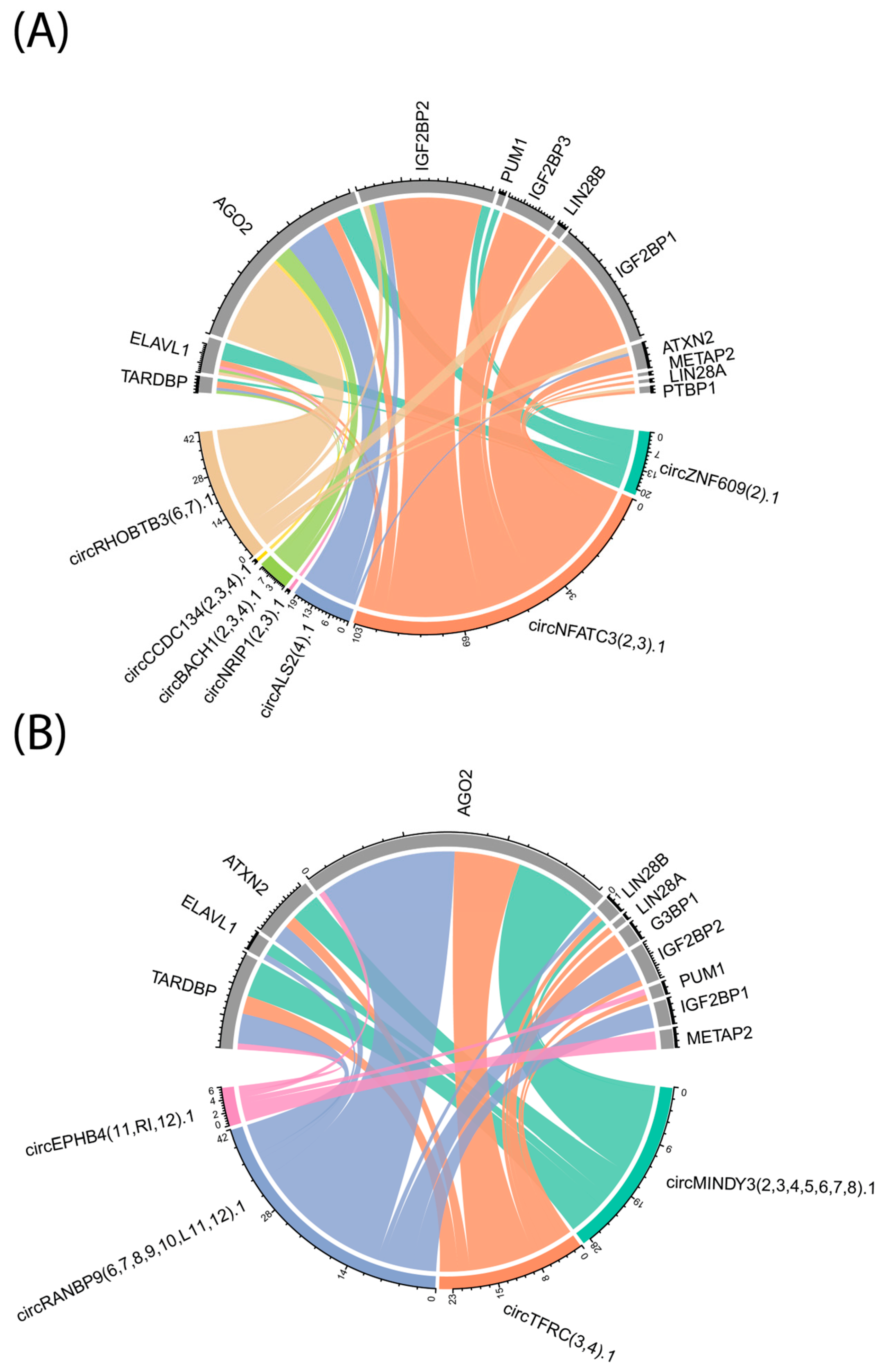
Predicted circRNA–RBP interaction networks revealed extensive binding potential for several RBPs with established roles in erythropoiesis and hemoglobin regulation, including AGO2, IGF2BP1, IGF2BP2, and LIN28B. Notably, certain circRNAs were predicted to bind multiple RBPs, while individual RBPs displayed broad connectivity with circRNAs across both developmental contexts. These networks suggest that circRNAs may serve as important modulators of post-transcriptional gene regulation during erythroid differentiation, either by sequestering RBPs or facilitating the formation of ribonucleoprotein complexes.
To evaluate whether the selected RBPs are expressed in our samples, mRNA expression analysis was performed, which confirmed that the corresponding mRNAs for these RBPs are indeed present. Notably, the mRNA expression levels of LIN28B, IGF2BP1, and IGF2BP3 were significantly higher in fetal liver-derived erythroblasts compared to bone marrow-derived erythroblasts. However, as circRNAs may directly bind RBPs, acting as molecular sponges or scaffolds to modulate RBP localization, stability, and function, this could not be further assessed due to the absence of proteomic data for these samples. Nevertheless, circRNAs can also regulate RBPs indirectly via miRNA–mRNA interaction networks, as illustrated in
Figure 6.
Taken together, these findings demonstrate that circRNA–miRNA–(RBP)mRNA interaction networks are developmentally regulated, likely contributing to the distinct molecular identities observed between fetal and adult erythroid cells. The identification of specific circRNAs and their predicted interactions with regulatory miRNAs and RBPs suggests these molecules may serve as critical modulators in key developmental processes, including hemoglobin gene regulation.
However, several limitations should be considered when interpreting these results. First, the circRNA detection in this study was conducted using total RNA-seq libraries rather than circRNA-enriched preparations. Consequently, this approach may have reduced sensitivity in detecting low-abundance or tissue-specific circRNAs, potentially omitting circRNAs that might have been identified with enrichment protocols. Second, all circRNA–miRNA and circRNA–RBP interactions were predicted in silico using computational tools, primarily circAtlas 3.0 and miRTarBase [
11,
15]. Thus, experimental validation is essential to confirm the biological relevance and specificity of these interactions.
Additionally, the current study did not perform co-expression analyses. Therefore, proposed regulatory roles for circRNAs upregulated in one tissue (e.g., bone marrow) that interact with miRNAs or RBPs upregulated in another tissue (e.g., fetal liver) remain speculative. Without experimental evidence, cross-tissue regulatory effects cannot be conclusively inferred from computational predictions alone. Finally, while selected RBPs with known roles in erythropoiesis and hemoglobin regulation were examined, the lack of comprehensive proteomic profiling of RBPs in erythroid cells derived from fetal liver and bone marrow limits a complete understanding of circRNA–RBP dynamics.
Despite these limitations, the findings underscore the potential role of circRNAs as regulatory elements in hemoglobin expression and erythroid differentiation, both in normal physiology and disease states. Systematic profiling and in silico analyses of circRNA expression and interactions in fetal liver and bone marrow-derived erythroblasts offer new insights into erythroid molecular genetics. These insights may facilitate the discovery of novel biomarkers or therapeutic targets for hemoglobinopathies and related erythroid disorders.
Future studies employing experimental validation methods, such as circRNA knockdown, overexpression, or pulldown assays, will be critical for establishing the mechanistic significance of the predicted interactions. Such work may identify promising targets for therapeutic modulation of erythropoiesis or reactivation of fetal hemoglobin, offering potential advances in the treatment of hemoglobinopathies.
5. Conclusions
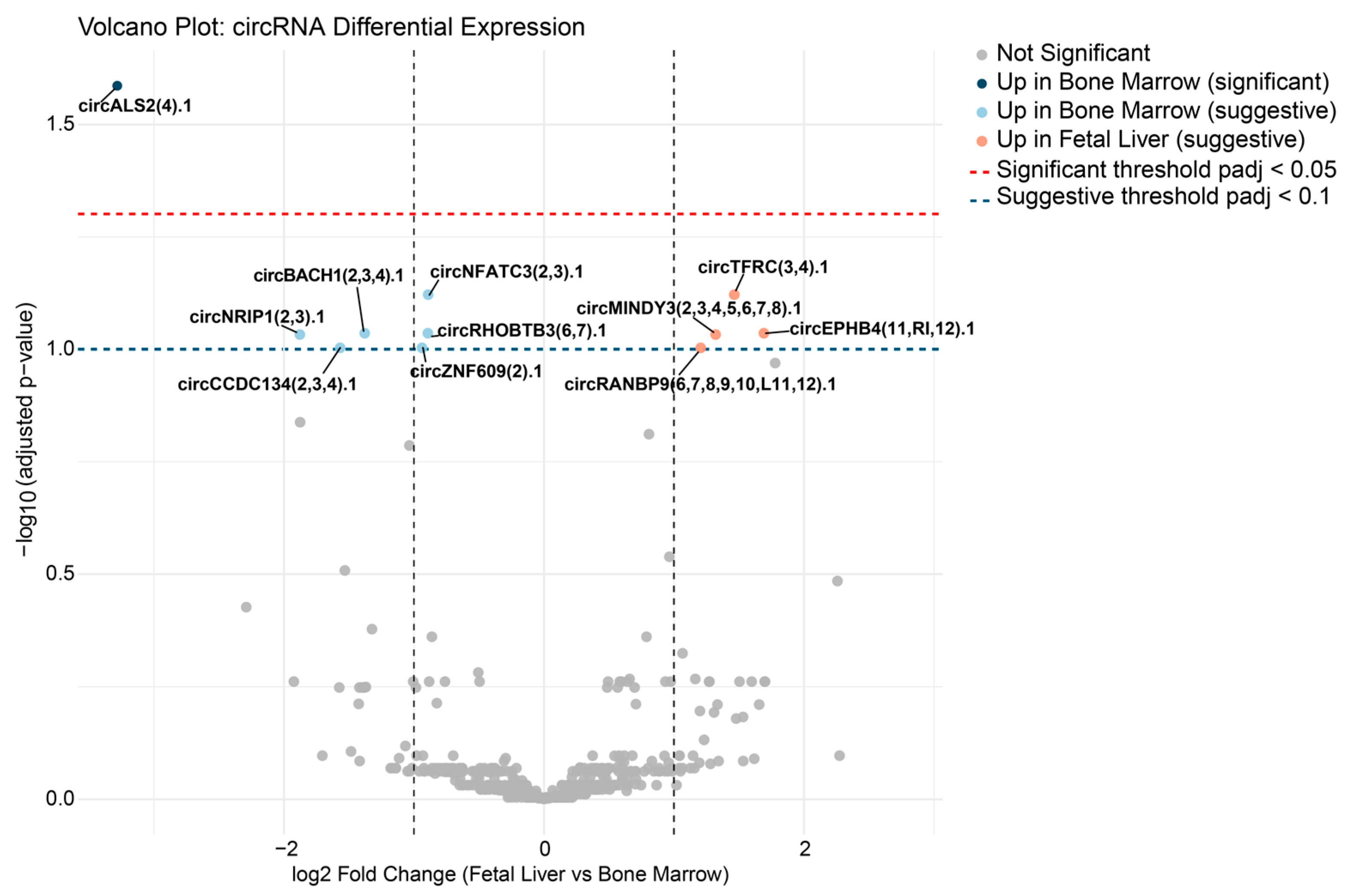
In summary, this study provides a comparative analysis of circRNA expression and predicted regulatory networks, including circRNA–miRNA and circRNA–RBP interactions, in erythroblasts derived from human fetal liver and bone marrow. One circRNA, circALS2(4).1, was found to be significantly upregulated in bone marrow-derived erythroblasts, with ten additional circRNAs showing suggestive developmental regulation. While the overall separation between developmental origins was statistically significant but modest in effect size, distinct interaction networks involving circRNAs, miRNAs, and RBPs were uncovered. These findings highlight a potential novel regulatory circuit that may contribute to developmental differences in human erythropoiesis.
Although the physiological roles of the identified circRNAs, their associated miRNAs, and RBPs remain to be established, the results suggest their potential involvement in regulating erythroid differentiation and hemoglobin expression. Future studies should focus on experimental validation of key circRNA candidates, such as circALS2(4).1, and their interactions with relevant miRNAs and RBPs, using approaches such as knockdown, overexpression, or pulldown assays. This will be essential to elucidate the specific functions and regulatory mechanisms of these molecules in erythropoiesis, and may identify new therapeutic targets for red blood cell disorders and hemoglobinopathies.