CDKL5 Deficiency Disorder: Revealing the Molecular Mechanism of Pathogenic Variants
Abstract
1. Introduction
2. Results
2.1. CDKL5 Variants, Structure, and Binding Partners
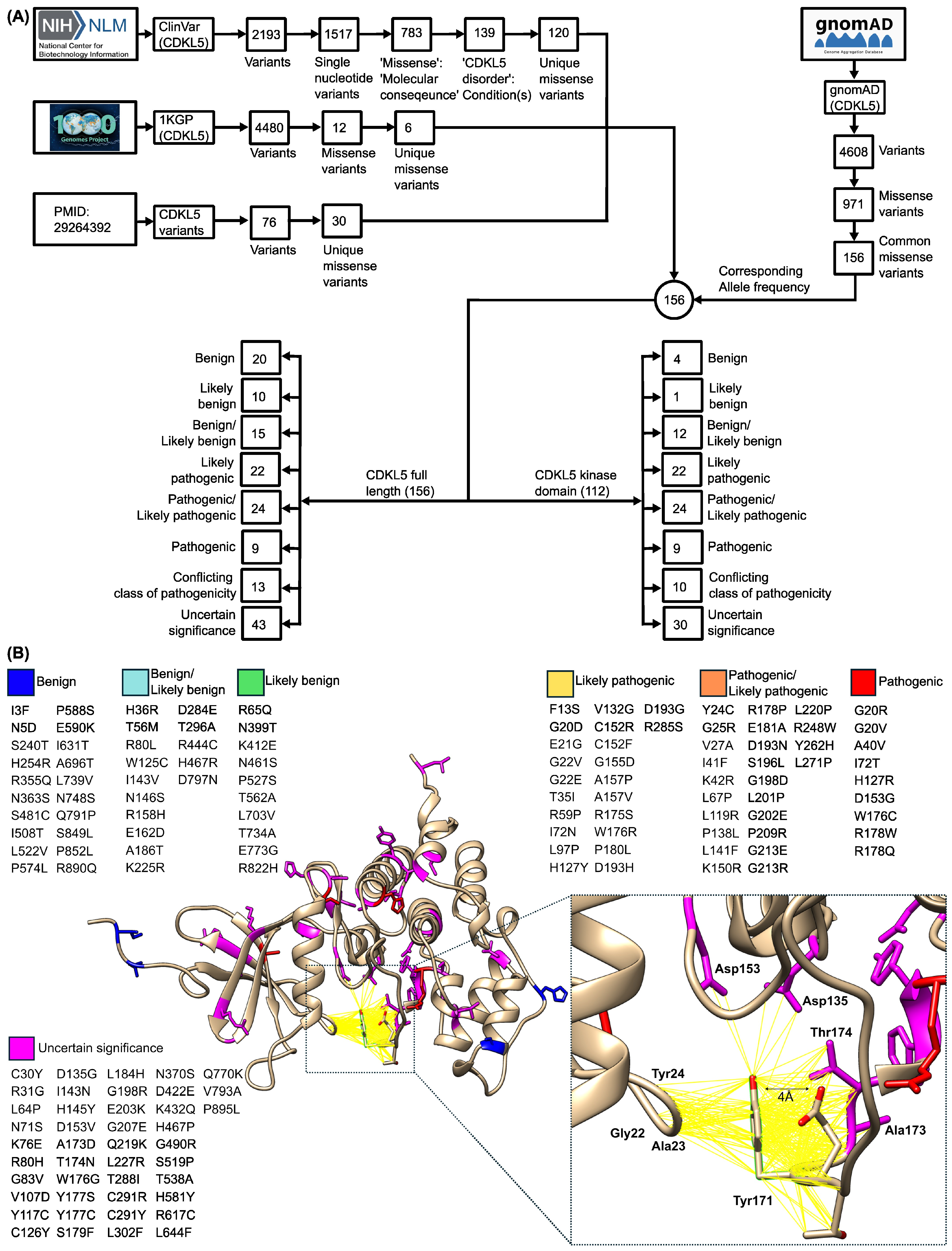
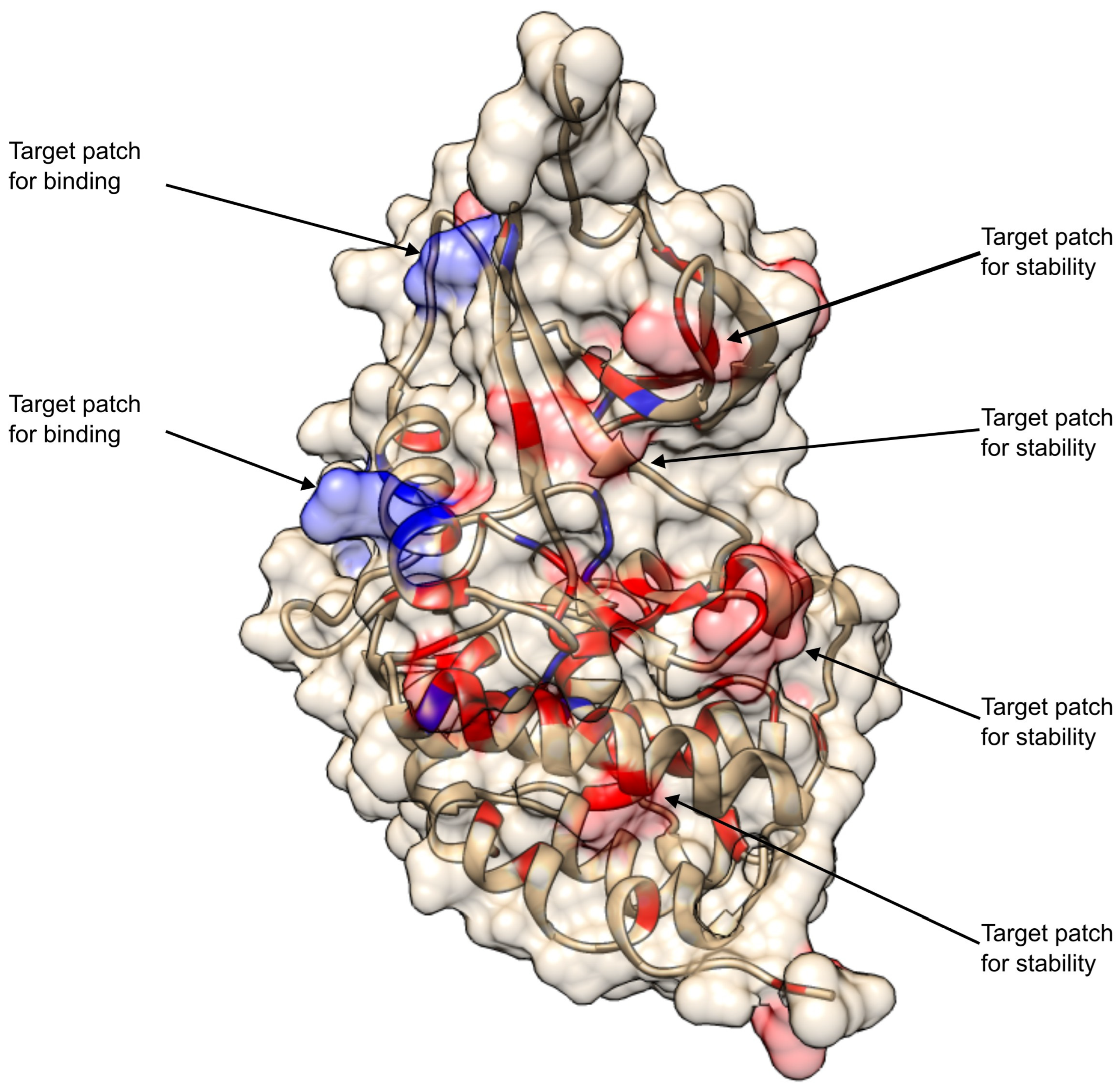
2.1.1. Curation and Structural Mapping of Missense CDKL5 Variants Associated with CDD
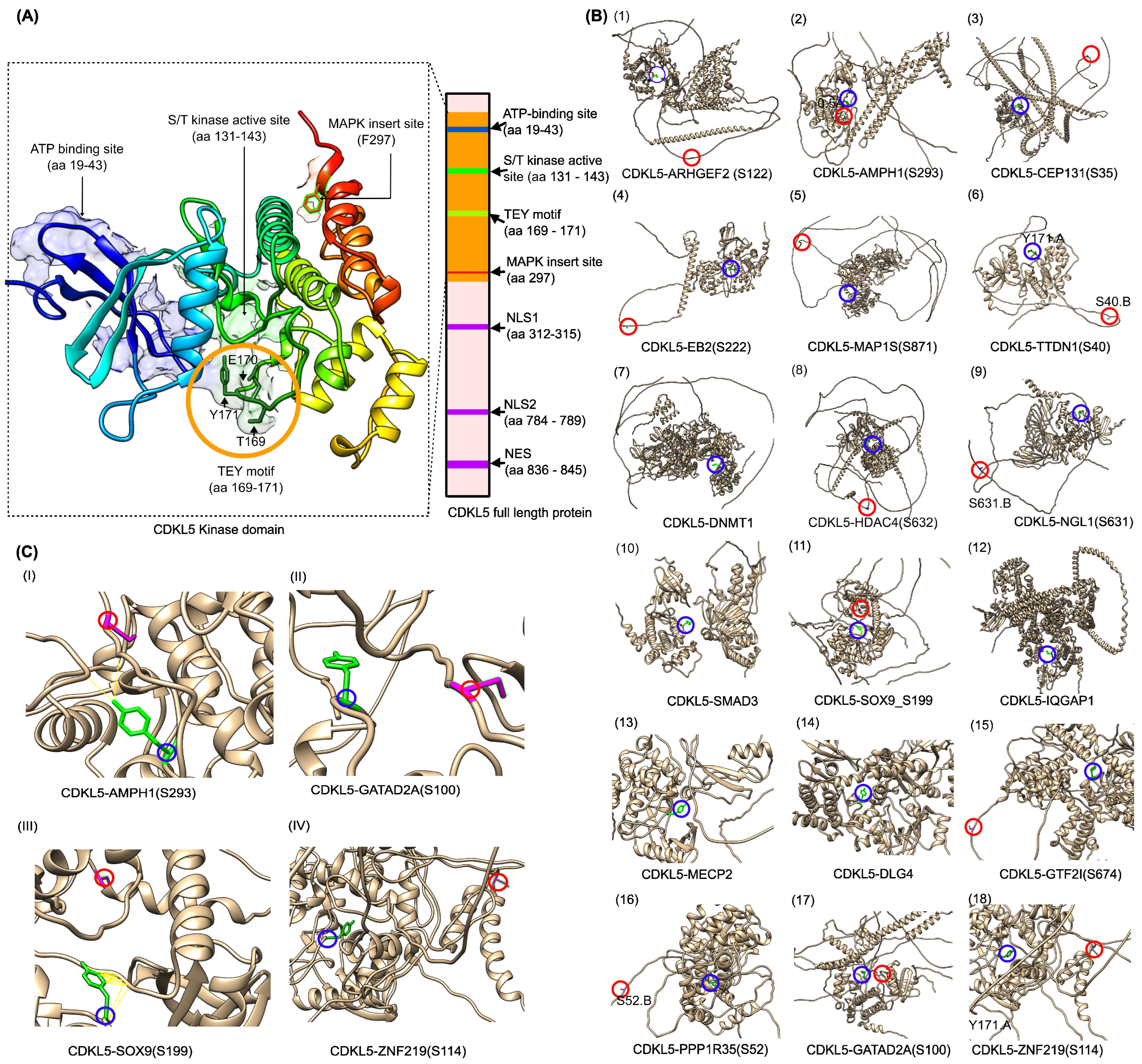
2.1.2. CDKL5 Partners

| SL | UniProt | Gene | pSite | Consensus Motif (RPX[S/T][A/G/P/S]) | Protein | Evidence | Source |
|---|---|---|---|---|---|---|---|
| 1 | Q92974 | ARHGEF2 | S122 | TIRERPSsAIYPS | Rho guanine nucleotide exchange factor 2 | Motif and biochemical | [32] |
| 2 | P49418 | AMPH1 | S293 | PAPARPRsPSQTR | Amphiphysin1 | Motif and biochemical | [32] |
| 3 | Q9UPN4 | CEP131 | S35 | PVSRRPGsAATTK | Centrosomal protein of 131 kDa | Motif and biochemical | [32] |
| 4 | Q8TDM6 | DLG5 | S1115 | QKRRRPKsAPSFR | Disks large homolog 5 | Motif and biochemical | [32] |
| 5 | Q14241 | ELOA | S311 | EENRRPPsGDNAR | Elongin A | Motif and biochemical | [32] |
| 6 | Q96L91 | EP400 | S729 | SPVNRPSsATNKA | EE1A-binding protein p400 | Motif and biochemical | [32] |
| 7 | Q66K74 | MAP1S | S871, S900 | KAPARPSsASATP, DRASRPLsARSEP | Microtubule-associated protein 1S | Motif and biochemical | [32] |
| 8 | Q15555 | EB2/MAPRE2 | S222 | STPSRPSsAKRAS | Microtubule-associated protein RP/EB family member 2 | Motif and biochemical | [32] |
| 9 | Q8TAP9 | TTDN1 | S40 | GGGPRPPsPRDGY | TTD non-photosensitive 1 protein | Motif and biochemical | [32] |
| 10 | P26358 | DNMT1 | N/A | DNA methyltransferase 1 | Biochemical | [32] | |
| 11 | P56524 | HDAC4 | S632 | RPLSRAQsSPASAtF | Histone deacetylase 4 | Motif and biochemical | [32] |
| 12 | Q9HCJ2 | NGL-1/KIAA1580/LRRC4C | S631 | PLLIRMNsKDNVQET | Netrin-G ligand-1 | Motif and biochemical | [32] |
| 13 | P84022 | SMAD3 | N/A | N/A | Mothers against decapentaplegic homolog 3 | Biochemical | [32] |
| 14 | P48436 | SOX9 | S199 | ATEQTHIsPNAIFKA | Transcription factor SOX-9 | Biochemical | [32] |
| 15 | P46940 | IQGAP1 | N/A | N/A | IQ Motif Containing GTPase Activating Protein 1 | Biochemical | [32] |
| 16 | P51608 | MeCP2 | N/A | N/A | Methyl-CpG binding protein 2 | Biochemical | [32] |
| 17 | P78352 | PSD95/DLG4 | N/A | N/A | Postsynaptic density protein 95 | Biochemical | [32] |
| 18 | A0MZ66 | SHTN1/SHOT1 | N/A | N/A | Shootin1 | Biochemical | [32] |
| 19 | P78347 | GTF2I | S674 | QSPKRPRsPGSNS | General transcription factor II-I | Motif and biochemical | [42] |
| 20 | Q8TAP8 | PPP1R35 | S52 | SLSPRPDsPQPRH | Protein phosphatase 1 regulatory subunit 35 | Motif and biochemical | [42] |
| 21 | Q86YP4 | GATAD2A | S100 | KSERRPPsPDVIV | GATA zinc finger domain containing 2A | Motif and biochemical | [42] |
| 22 | Q9P2Y4 | ZNF219 | S114 | HQPERPRsPAARL | Zinc finger protein 219 | Motif and biochemical | [42] |
| 23 | Q15878 | CACNA1E/Cav2.3 | S14 | AVVARPGsGDGD | Voltage-dependent R-type calcium channel subunit alpha-1E (Cav2.3) | Motif and biochemical | [62] |
| 24 | Q13501 | SQSTM1/p62 | T269/S272 | RSRLTPVsPESS, GGKRSRLtPVSP | Sequestosome-1(p62) | Biochemical | [63] |
2.2. Homology Modeling of the CDKL5 Kinase Domain and CDKL5-Target Complex Prediction Using ColabFold and CDKL5-Target Protein–Protein Docking Using HADDOCK3
2.3. Folding, Docking ΔΔGfolding and ΔΔGbinding Analysis, and Variant Reclassification
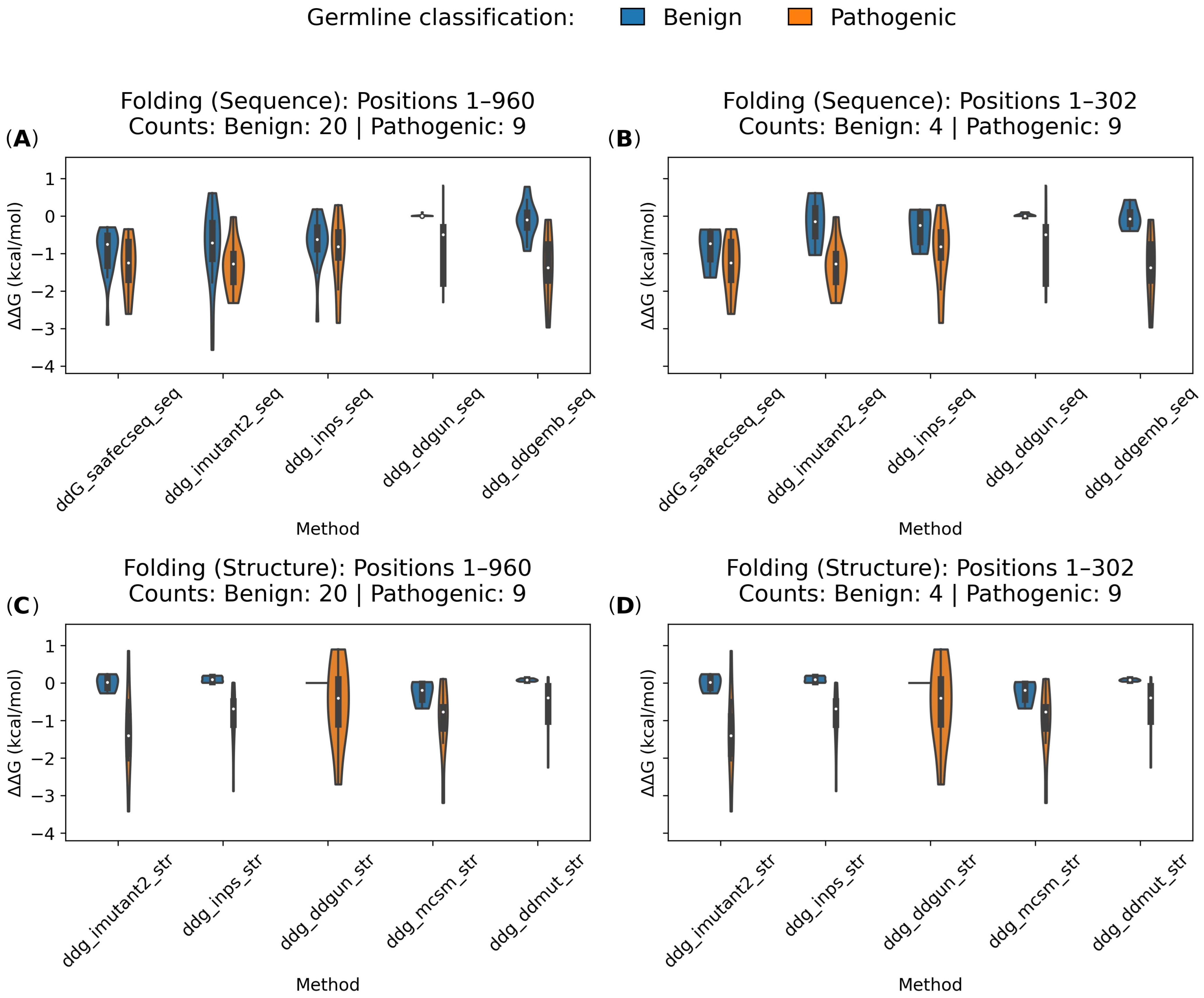
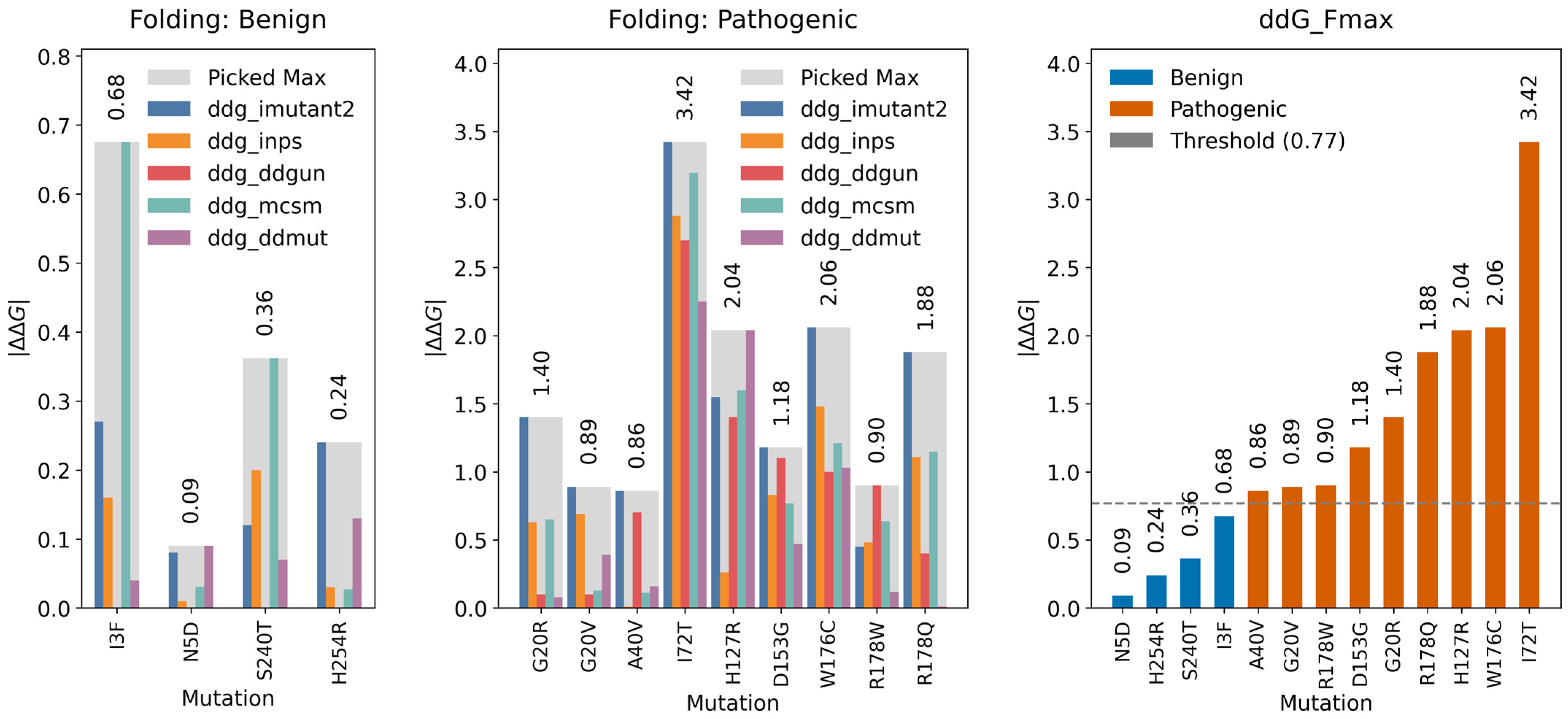
2.3.1. Folding Free Energy Change (ΔΔGfolding)
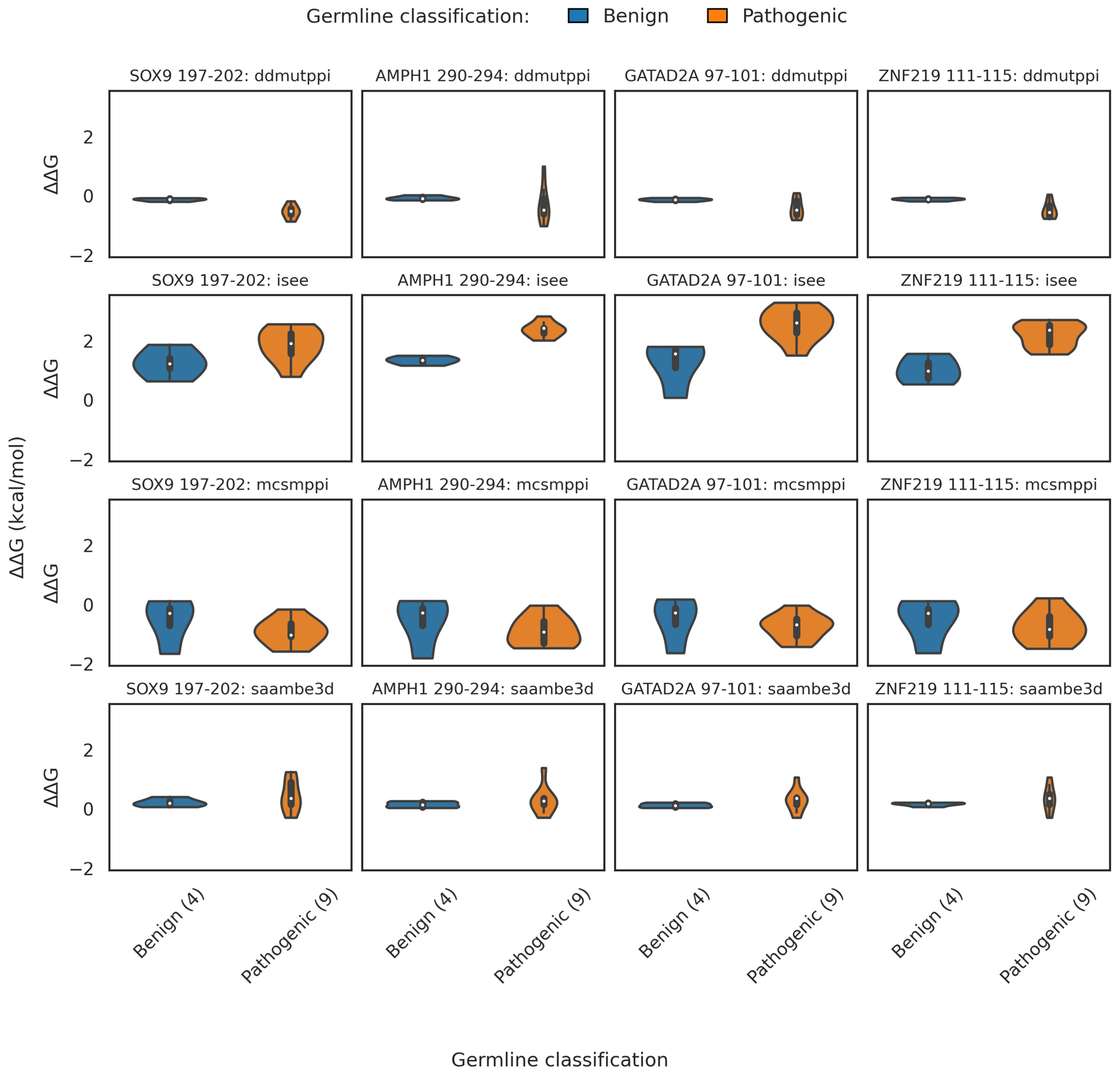
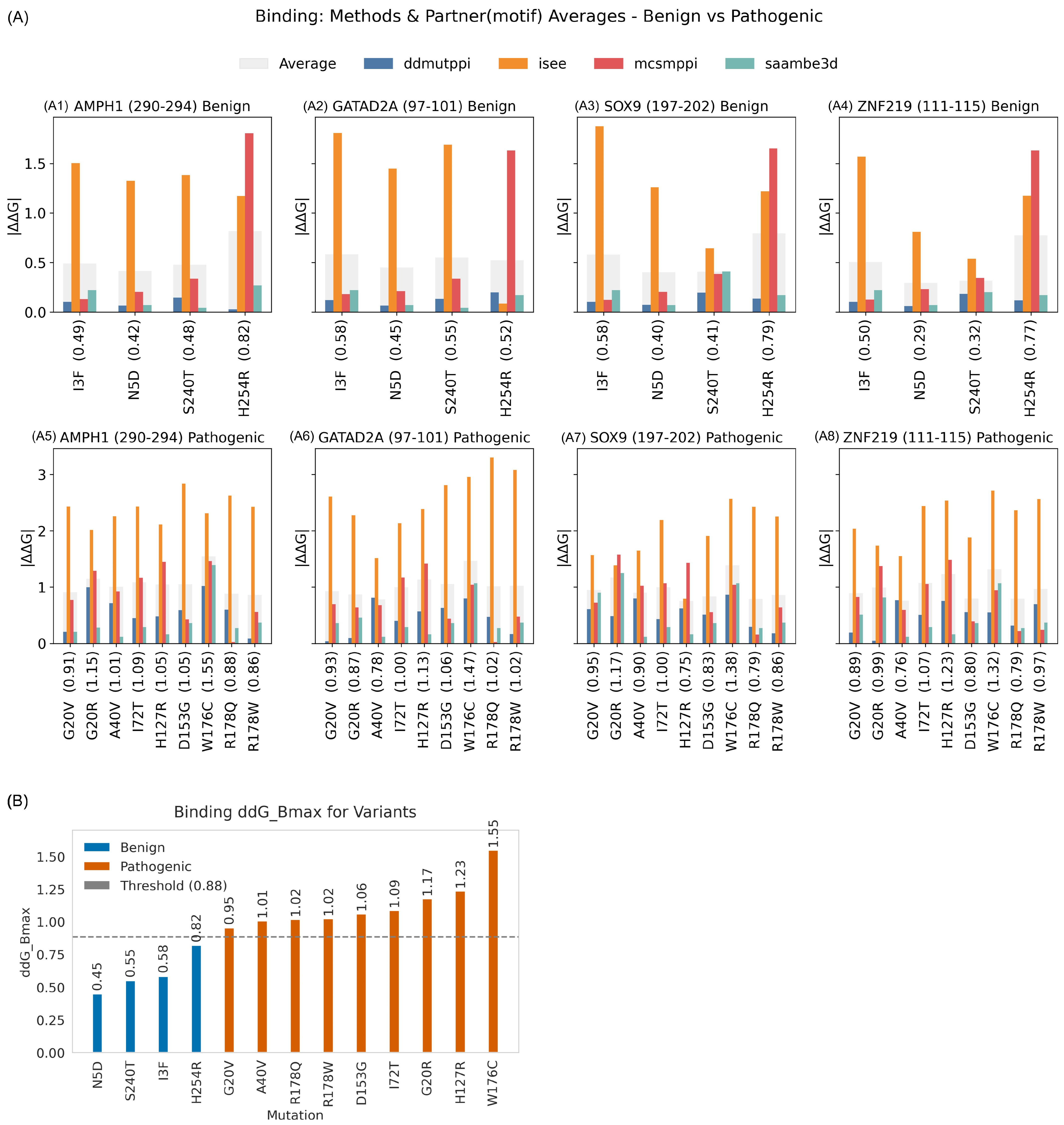
2.3.2. Binding Free Energy
2.3.3. Folding Threshold
2.3.4. Binding Threshold
2.3.5. Variants Reclassification Based on ΔΔGfolding and ΔΔGbinding Thresholds
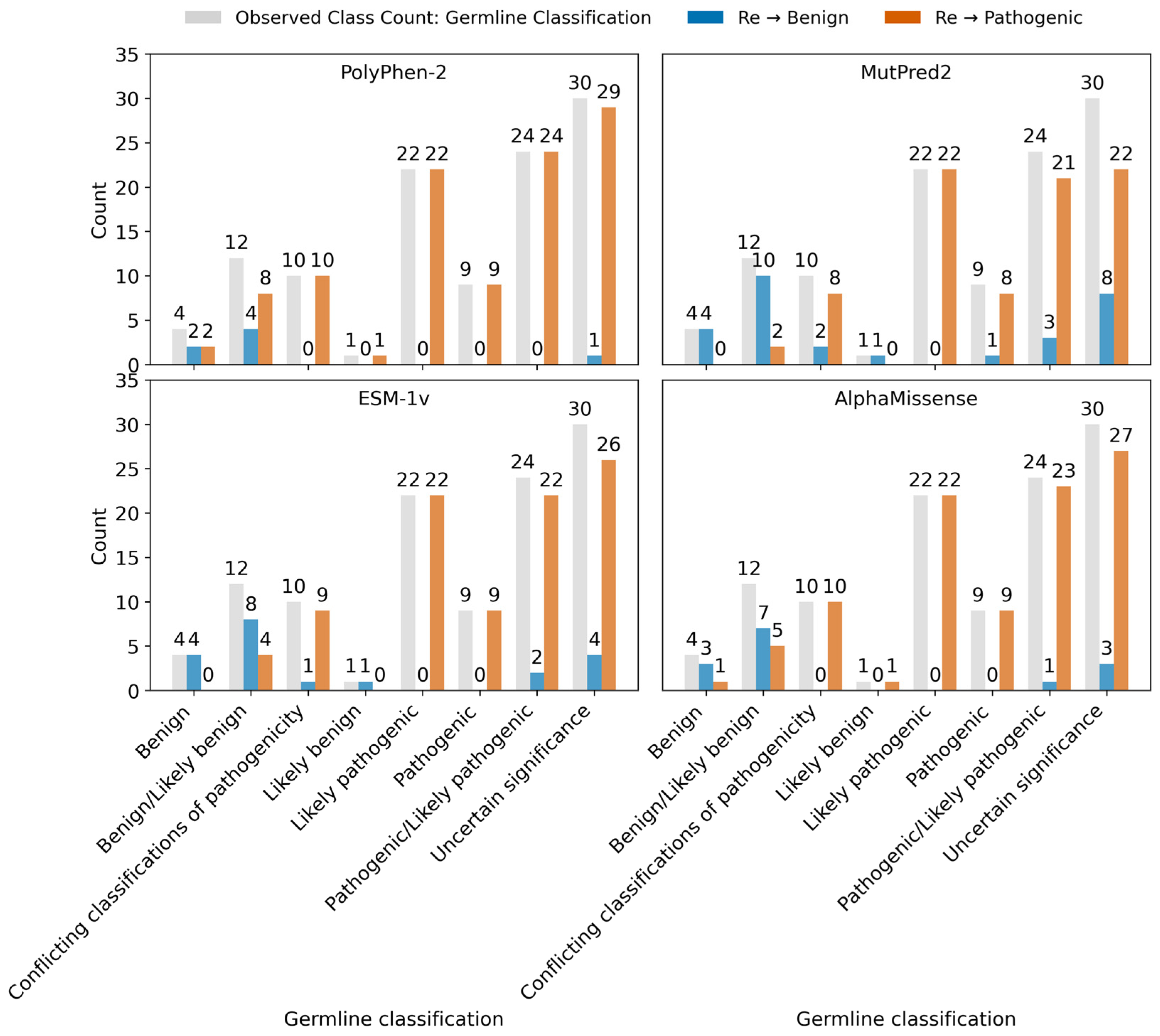
2.3.6. Variants Reclassification Based on Pathogenicity Score
3. Discussion
4. Materials and Methods
4.1. Data Collection
4.2. CDKL5 Structure Preparation and Prediction of Complex with Its Binding Partners
4.3. Folding Free Energy Calculations
4.4. HADDOCK3 Protein–Protein Docking
4.5. Binding Free Energy Calculations
4.6. CDKL5 Variant Reclassification
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 1KGP | The 1000 Genomes Project |
| AMPH1 | Amphiphysin1 |
| ARHGEF2 | Rho guanine nucleotide exchange factor 2 |
| CACNA1E/Cav2.3 | Voltage-dependent R-type calcium channel subunit alpha-1E (Cav2.3) |
| CDD | CDKL5 Deficiency Disorder |
| CDKL5 | Cyclin-Dependent Kinase-Like 5 |
| CEP131 | Centrosomal protein of 131 kDa |
| CVI | Cortical/cerebral visual impairment |
| DLG5 | Disks large homolog 5 |
| DNMT1 | DNA methyltransferase 1 |
| EB2/MAPRE2 | Microtubule-associated protein RP/EB family member 2 |
| ELOA | Elongin A |
| EP400 | EE1A-binding protein p400 |
| GATAD2A | GATA zinc finger domain containing 2A |
| GRCh38 | The Genome Reference Consortium Human Build 38 |
| GTF2I | General transcription factor II-I |
| HDAC4 | Histone deacetylase 4 |
| IGSR | The International Genome Sample Resource |
| IQGAP1 | IQ Motif Containing GTPase Activating Protein 1 |
| MAP1S | Microtubule-associated protein 1S |
| MeCP2 | Methyl-CpG binding protein 2 |
| NGL-1/KIAA1580/LRRC4C | Netrin-G ligand-1 |
| OMIM | Online Mendelian Inheritance in Man |
| PPP1R35 | Protein phosphatase 1 regulatory subunit 35 |
| PSD95/DLG4 | Postsynaptic density protein 95 |
| SHTN1/SHOT1 | Shootin1 |
| SMAD3 | Mothers against decapentaplegic homolog 3 |
| SOX9 | Transcription factor SOX-9 |
| SQSTM1/p62 | Sequestosome-1(p62) |
| STK9 | Serine threonine kinase 9 |
| TTDN1 | TTD non-photosensitive 1 protein |
| VDW | Van der Waals |
| XCI | X-chromosome inactivation |
| ZNF219 | Zinc finger protein 219 |
| ΔΔGbinding | Binding free energy change |
| ΔΔGfolding | Folding free energy changes |
| ΔΔGBmax | Maximum binding free energy change: For each variant, the maximum complex-averaged |ΔΔGbinding| across selected CDKL5-target protein–protein interactions. |
| ΔΔGFmax | Maximum folding free energy change: For each variant, the highest ΔΔGfolding value across several computational methods. |
References
- Amberger, J.S.; Bocchini, C.A.; Scott, A.F.; Hamosh, A. OMIM.org: Leveraging knowledge across phenotype-gene relationships. Nucleic Acids Res. 2019, 47, D1038–D1043. [Google Scholar] [CrossRef]
- Olson, H.E.; Demarest, S.T.; Pestana-Knight, E.M.; Swanson, L.C.; Iqbal, S.; Lal, D.; Leonard, H.; Cross, J.H.; Devinsky, O.; Benke, T.A. Cyclin-Dependent Kinase-Like 5 Deficiency Disorder: Clinical Review. Pediatr. Neurol. 2019, 97, 18–25. [Google Scholar] [CrossRef]
- Akiba, T.; Shimada, S.; Imai, K.; Takahashi, S. A case of CDKL5 deficiency disorder with a novel intragenic multi-exonic duplication. Hum. Genome Var. 2024, 11, 40. [Google Scholar] [CrossRef]
- Jakimiec, M.; Paprocka, J.; Śmigiel, R. CDKL5 Deficiency Disorder-A Complex Epileptic Encephalopathy. Brain Sci. 2020, 10, 107. [Google Scholar] [CrossRef]
- Leonard, H.; Downs, J.; Benke, T.A.; Swanson, L.; Olson, H.; Demarest, S. CDKL5 deficiency disorder: Clinical features, diagnosis, and management. Lancet Neurol. 2022, 21, 563–576. [Google Scholar] [CrossRef]
- Daniels, C.; Greene, C.; Smith, L.; Pestana-Knight, E.; Demarest, S.; Zhang, B.; Benke, T.A.; Poduri, A.; Olson, H. CDKL5 deficiency disorder and other infantile-onset genetic epilepsies. Dev. Med. Child. Neurol. 2024, 66, 456–468. [Google Scholar] [CrossRef]
- Siri, B.; Varesio, C.; Freri, E.; Darra, F.; Gana, S.; Mei, D.; Porta, F.; Fontana, E.; Galati, G.; Solazzi, R.; et al. CDKL5 deficiency disorder in males: Five new variants and review of the literature. Eur. J. Paediatr. Neurol. 2021, 33, 9–20. [Google Scholar] [CrossRef]
- Dell’Isola, G.B.; Portwood, K.E.; Consing, K.; Fattorusso, A.; Bartocci, A.; Ferrara, P.; Di Cara, G.; Verrotti, A.; Lodolo, M. Current Overview of CDKL-5 Deficiency Disorder Treatment. Pediatr. Rep. 2024, 16, 21–25. [Google Scholar] [CrossRef]
- Lombardo, A.; Sinibaldi, L.; Genovese, S.; Catino, G.; Mei, V.; Pompili, D.; Sallicandro, E.; Falasca, R.; Liambo, M.T.; Faggiano, M.V.; et al. A Case of CDKL5 Deficiency Due to an X Chromosome Pericentric Inversion: Delineation of Structural Rearrangements as an Overlooked Recurrent Pathological Mechanism. Int. J. Mol. Sci. 2024, 25, 6912. [Google Scholar] [CrossRef]
- Evans, J.C.; Archer, H.L.; Colley, J.P.; Ravn, K.; Nielsen, J.B.; Kerr, A.; Williams, E.; Christodoulou, J.; Gécz, J.; Jardine, P.E.; et al. Early onset seizures and Rett-like features associated with mutations in CDKL5. Eur. J. Hum. Genet. 2005, 13, 1113–1120. [Google Scholar] [CrossRef]
- Martinez, D.; Jiang, E.; Zhou, Z. Overcoming genetic and cellular complexity to study the pathophysiology of X-linked intellectual disabilities. J. Neurodev. Disord. 2024, 16, 5. [Google Scholar] [CrossRef]
- Adhikari, A.; Buchanan, F.K.B.; Fenton, T.A.; Cameron, D.L.; Halmai, J.A.N.M.; Copping, N.A.; Fink, K.D.; Silverman, J.L. Touchscreen cognitive deficits, hyperexcitability and hyperactivity in males and females using two models of Cdkl5 deficiency. Hum. Mol. Genet. 2022, 31, 3032–3050. [Google Scholar] [CrossRef]
- Galvani, G.; Mottolese, N.; Gennaccaro, L.; Loi, M.; Medici, G.; Tassinari, M.; Fuchs, C.; Ciani, E.; Trazzi, S. Inhibition of microglia overactivation restores neuronal survival in a mouse model of CDKL5 deficiency disorder. J. Neuroinflamm. 2021, 18, 155. [Google Scholar] [CrossRef]
- Benke, T.A.; Demarest, S.; Angione, K.; Downs, J.; Leonard, H.; Saldaris, J.; Marsh, E.D.; Olson, H.; Haviland, I. CDKL5 Deficiency Disorder. In GeneReviews®; Adam, M.P., Feldman, J., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Olson, H.E.; Daniels, C.I.; Haviland, I.; Swanson, L.C.; Greene, C.A.; Denny, A.M.M.; Demarest, S.T.; Pestana-Knight, E.; Zhang, X.; Moosa, A.N.; et al. Current neurologic treatment and emerging therapies in CDKL5 deficiency disorder. J. Neurodev. Disord. 2021, 13, 40. [Google Scholar] [CrossRef]
- Dell’Isola, G.B.; Perinelli, M.G.; Frulli, A.; D’Onofrio, G.; Fattorusso, A.; Siciliano, M.; Ferrara, P.; Striano, P.; Verrotti, A. Exploring neurodevelopment in CDKL5 deficiency disorder: Current insights and future directions. Epilepsy Behav. 2025, 171, 110504. [Google Scholar] [CrossRef]
- Melikishvili, G.; Sharkov, A.; Gachechiladze, T.; Tomenko, T.; Pivovarova, A.; Volkov, I.; Andrade, M.-T.; Castellanos, A.; Bienvenu, T.; Dulac, O.; et al. Epileptic spasms with terror during sleep in CDKL5 encephalopathy. Sleep Adv. 2022, 3, zpac010. [Google Scholar] [CrossRef]
- Müller, A.; Helbig, I.; Jansen, C.; Bast, T.; Guerrini, R.; Jähn, J.; Muhle, H.; Auvin, S.; Korenke, G.C.; Philip, S.; et al. Retrospective evaluation of low long-term efficacy of antiepileptic drugs and ketogenic diet in 39 patients with CDKL5-related epilepsy. Eur. J. Paediatr. Neurol. 2016, 20, 147–151. [Google Scholar] [CrossRef]
- Klein, K.M.; Yendle, S.C.; Harvey, A.S.; Antony, J.H.; Wallace, G.; Bienvenu, T.; Scheffer, I.E. A distinctive seizure type in patients with CDKL5 mutations: Hypermotor-tonic-spasms sequence. Neurology 2011, 76, 1436–1438. [Google Scholar] [CrossRef]
- Massey, S.; Quigley, A.; Rochfort, S.; Christodoulou, J.; Van Bergen, N.J. Cannabinoids and Genetic Epilepsy Models: A Review with Focus on CDKL5 Deficiency Disorder. Int. J. Mol. Sci. 2024, 25, 10768. [Google Scholar] [CrossRef]
- Sun, X.; Wang, T. Research progress on the pathogenesis of CDKL5 pathogenic variants and related encephalopathy. Eur. J. Pediatr. 2023, 182, 3049–3056. [Google Scholar] [CrossRef]
- Silvestre, M.; Dempster, K.; Mihaylov, S.R.; Claxton, S.; Ultanir, S.K. Cell type-specific expression, regulation and compensation of CDKL5 activity in mouse brain. Mol. Psychiatry 2024, 29, 1844–1856. [Google Scholar] [CrossRef]
- Saldaris, J.M.; Jacoby, P.; Marsh, E.D.; Suter, B.; Leonard, H.; Olson, H.E.; Rajaraman, R.; Pestana-Knight, E.; Weisenberg, J.; Price, D.; et al. Adapting a measure of gross motor skills for individuals with CDKL5 deficiency disorder: A psychometric study. Epilepsy Res. 2024, 200, 107287. [Google Scholar] [CrossRef] [PubMed]
- Wong, K.; Junaid, M.; Demarest, S.; Saldaris, J.; Benke, T.A.; Marsh, E.D.; Downs, J.; Leonard, H. Factors influencing the attainment of major motor milestones in CDKL5 deficiency disorder. Eur. J. Hum. Genet. 2023, 31, 169–178. [Google Scholar] [CrossRef]
- Brock, D.; Fidell, A.; Thomas, J.; Juarez-Colunga, E.; Benke, T.A.; Demarest, S. Cerebral Visual Impairment in CDKL5 Deficiency Disorder Correlates With Developmental Achievement. J. Child. Neurol. 2021, 36, 974–980. [Google Scholar] [CrossRef]
- Quintiliani, M.; Ricci, D.; Petrianni, M.; Leone, S.; Orazi, L.; Amore, F.; Gambardella, M.L.; Contaldo, I.; Veredice, C.; Perulli, M.; et al. Cortical Visual Impairment in CDKL5 Deficiency Disorder. Front. Neurol. 2021, 12, 805745. [Google Scholar] [CrossRef] [PubMed]
- Peikes, T.; Hartley, J.N.; Mhanni, A.A.; Greenberg, C.R.; Appendino, J.P. Reflex Seizures in a Patient with CDKL5 Deficiency Disorder. Can. J. Neurol. Sci. 2019, 46, 482–485. [Google Scholar] [CrossRef] [PubMed]
- Downs, J.; Jacoby, P.; Saldaris, J.; Leonard, H.; Benke, T.; Marsh, E.; Demarest, S. Negative impact of insomnia and daytime sleepiness on quality of life in individuals with the cyclin-dependent kinase-like 5 deficiency disorder. J. Sleep. Res. 2022, 31, e13600. [Google Scholar] [CrossRef] [PubMed]
- Amin, S.; Monaghan, M.; Aledo-Serrano, A.; Bahi-Buisson, N.; Chin, R.F.; Clarke, A.J.; Cross, J.H.; Demarest, S.; Devinsky, O.; Downs, J.; et al. International Consensus Recommendations for the Assessment and Management of Individuals With CDKL5 Deficiency Disorder. Front. Neurol. 2022, 13, 874695. [Google Scholar] [CrossRef]
- La Montanara, P.; Hervera, A.; Baltussen, L.L.; Hutson, T.H.; Palmisano, I.; De Virgiliis, F.; Kong, G.; Chadwick, J.; Gao, Y.; Bartus, K.; et al. Cyclin-dependent-like kinase 5 is required for pain signaling in human sensory neurons and mouse models. Sci. Transl. Med. 2020, 12, eaax4846. [Google Scholar] [CrossRef]
- Specchio, N.; Trivisano, M.; Lenge, M.; Ferretti, A.; Mei, D.; Parrini, E.; Napolitano, A.; Rossi-Espagnet, C.; Talenti, G.; Longo, D.; et al. CDKL5 deficiency disorder: Progressive brain atrophy may be part of the syndrome. Cereb. Cortex 2023, 33, 9709–9717. [Google Scholar] [CrossRef]
- Van Bergen, N.J.; Massey, S.; Quigley, A.; Rollo, B.; Harris, A.R.; Kapsa, R.M.I.; Christodoulou, J. CDKL5 deficiency disorder: Molecular insights and mechanisms of pathogenicity to fast-track therapeutic development. Biochem. Soc. Trans. 2022, 50, 1207–1224. [Google Scholar] [CrossRef]
- Chowdhury, I.; Dashi, G.; Keskitalo, S. CMGC Kinases in Health and Cancer. Cancers 2023, 15, 3838. [Google Scholar] [CrossRef]
- Quadalti, C.; Sannia, M.; Humphreys, N.E.; Baldassarro, V.A.; Gurgone, A.; Ascolani, M.; Zanella, L.; Giardino, L.; Gross, C.T.; Croci, S.; et al. A new knockin mouse carrying the E364X patient mutation for CDKL5 deficiency disorder: Neurological, behavioral and molecular profiling. Heliyon 2024, 10, e40165. [Google Scholar] [CrossRef] [PubMed]
- Hamdan, F.F.; Myers, C.T.; Cossette, P.; Lemay, P.; Spiegelman, D.; Laporte, A.D.; Nassif, C.; Diallo, O.; Monlong, J.; Cadieux-Dion, M.; et al. High Rate of Recurrent De Novo Mutations in Developmental and Epileptic Encephalopathies. Am. J. Hum. Genet. 2017, 101, 664–685. [Google Scholar] [CrossRef]
- Katayama, S.; Sueyoshi, N.; Inazu, T.; Kameshita, I. Cyclin-Dependent Kinase-Like 5 (CDKL5): Possible Cellular Signalling Targets and Involvement in CDKL5 Deficiency Disorder. Neural Plast. 2020, 2020, 6970190. [Google Scholar] [CrossRef]
- Rusconi, L.; Salvatoni, L.; Giudici, L.; Bertani, I.; Kilstrup-Nielsen, C.; Broccoli, V.; Landsberger, N. CDKL5 expression is modulated during neuronal development and its subcellular distribution is tightly regulated by the C-terminal tail. J. Biol. Chem. 2008, 283, 30101–30111. [Google Scholar] [CrossRef]
- Muñoz, I.M.; Morgan, M.E.; Peltier, J.; Weiland, F.; Gregorczyk, M.; Brown, F.C.; Macartney, T.; Toth, R.; Trost, M.; Rouse, J. Phosphoproteomic screening identifies physiological substrates of the CDKL5 kinase. EMBO J. 2018, 37, e99559. [Google Scholar] [CrossRef]
- Baltussen, L.L.; Negraes, P.D.; Silvestre, M.; Claxton, S.; Moeskops, M.; Christodoulou, E.; Flynn, H.R.; Snijders, A.P.; Muotri, A.R.; Ultanir, S.K. Chemical genetic identification of CDKL5 substrates reveals its role in neuronal microtubule dynamics. EMBO J. 2018, 37, e99763. [Google Scholar] [CrossRef]
- Eyers, P.A. A new consensus for evaluating CDKL5/STK9-dependent signalling mechanisms. EMBO J. 2018, 37, e100848. [Google Scholar] [CrossRef]
- Li, C.; Liu, Y.; Luo, S.; Yang, M.; Li, L.; Sun, L. A review of CDKL: An underestimated protein kinase family. Int. J. Biol. Macromol. 2024, 277, 133604. [Google Scholar] [CrossRef]
- Massey, S.; Ang, C.-S.; Davidson, N.M.; Quigley, A.; Rollo, B.; Harris, A.R.; Kapsa, R.M.I.; Christodoulou, J.; Van Bergen, N.J. Novel CDKL5 targets identified in human iPSC-derived neurons. Cell Mol. Life Sci. 2024, 81, 347. [Google Scholar] [CrossRef]
- Hart, R.K.; Fokkema, I.F.A.C.; DiStefano, M.; Hastings, R.; Laros, J.F.J.; Taylor, R.; Wagner, A.H.; den Dunnen, J.T. HGVS Nomenclature 2024: Improvements to community engagement, usability, and computability. Genome Med. 2024, 16, 149. [Google Scholar] [CrossRef] [PubMed]
- Landrum, M.J.; Lee, J.M.; Riley, G.R.; Jang, W.; Rubinstein, W.S.; Church, D.M.; Maglott, D.R. ClinVar: Public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014, 42, D980–D985. [Google Scholar] [CrossRef]
- 1000 Genomes Project Consortium; Auton, A.; Brooks, L.D.; Durbin, R.M.; Garrison, E.P.; Kang, H.M.; Korbel, J.O.; Marchini, J.L.; McCarthy, S.; McVean, G.A.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [CrossRef]
- Hector, R.D.; Kalscheuer, V.M.; Hennig, F.; Leonard, H.; Downs, J.; Clarke, A.; Benke, T.A.; Armstrong, J.; Pineda, M.; Bailey, M.E.S.; et al. CDKL5 variants: Improving our understanding of a rare neurologic disorder. Neurol. Genet. 2017, 3, e200. [Google Scholar] [CrossRef]
- Chen, S.; Francioli, L.C.; Goodrich, J.K.; Collins, R.L.; Kanai, M.; Wang, Q.; Alföldi, J.; Watts, N.A.; Vittal, C.; Gauthier, L.D.; et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 2024, 625, 92–100. [Google Scholar] [CrossRef]
- McLaren, W.; Gil, L.; Hunt, S.E.; Riat, H.S.; Ritchie, G.R.S.; Thormann, A.; Flicek, P.; Cunningham, F. The Ensembl Variant Effect Predictor. Genome Biol. 2016, 17, 122. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Fong, C.S.; Ozaki, K.; Tsou, M.-F.B. PPP1R35 ensures centriole homeostasis by promoting centriole-to-centrosome conversion. Mol. Biol. Cell 2018, 29, 2801–2808. [Google Scholar] [CrossRef]
- Archambault, D.; Cheong, A.; Iverson, E.; Tremblay, K.D.; Mager, J. Protein phosphatase 1 regulatory subunit 35 is required for ciliogenesis, notochord morphogenesis, and cell-cycle progression during murine development. Dev. Biol. 2020, 465, 1–10. [Google Scholar] [CrossRef]
- Panda, P.; Kovacs, L.; Dzhindzhev, N.; Fatalska, A.; Persico, V.; Geymonat, M.; Riparbelli, M.G.; Callaini, G.; Glover, D.M. Tissue specific requirement of Drosophila Rcd4 for centriole duplication and ciliogenesis. J. Cell Biol. 2020, 219, e201912154. [Google Scholar] [CrossRef]
- Lepanto, P.; Badano, J.L.; Zolessi, F.R. Neuron’s little helper: The role of primary cilia in neurogenesis. Neurogenesis 2016, 3, e1253363. [Google Scholar] [CrossRef]
- Adams, J.W.; Vinokur, A.; de Souza, J.S.; Austria, C.; Guerra, B.S.; Herai, R.H.; Wahlin, K.J.; Muotri, A.R. Loss of GTF2I promotes neuronal apoptosis and synaptic reduction in human cellular models of neurodevelopment. Cell Rep. 2024, 43, 113867. [Google Scholar] [CrossRef]
- López-Tobón, A.; Shyti, R.; Villa, C.E.; Cheroni, C.; Fuentes-Bravo, P.; Trattaro, S.; Caporale, N.; Troglio, F.; Tenderini, E.; Mihailovich, M.; et al. GTF2I dosage regulates neuronal differentiation and social behavior in 7q11.23 neurodevelopmental disorders. Sci. Adv. 2023, 9, eadh2726. [Google Scholar] [CrossRef]
- Barak, B.; Feng, G. Neurobiology of social behavior abnormalities in autism and Williams syndrome. Nat. Neurosci. 2016, 19, 647–655. [Google Scholar] [CrossRef]
- Chimge, N.-O.; Makeyev, A.V.; Ruddle, F.H.; Bayarsaihan, D. Identification of the TFII-I family target genes in the vertebrate genome. Proc. Natl. Acad. Sci. USA 2008, 105, 9006–9010. [Google Scholar] [CrossRef]
- Hakre, S.; Tussie-Luna, M.I.; Ashworth, T.; Novina, C.D.; Settleman, J.; Sharp, P.A.; Roy, A.L. Opposing functions of TFII-I spliced isoforms in growth factor-induced gene expression. Mol. Cell 2006, 24, 301–308. [Google Scholar] [CrossRef]
- Ricciardi, S.; Ungaro, F.; Hambrock, M.; Rademacher, N.; Stefanelli, G.; Brambilla, D.; Sessa, A.; Magagnotti, C.; Bachi, A.; Giarda, E.; et al. CDKL5 ensures excitatory synapse stability by reinforcing NGL-1-PSD95 interaction in the postsynaptic compartment and is impaired in patient iPSC-derived neurons. Nat. Cell Biol. 2012, 14, 911–923. [Google Scholar] [CrossRef]
- Yan, M.; Guo, X.; Xu, C. Revealing the complex role of CDKL5 in developmental epilepsy through a calcium channel related vision. Acta Epileptol. 2024, 6, 15. [Google Scholar] [CrossRef]
- Inkscape Project Inkscape, Version 1.3.2. Available online: https://inkscape.org (accessed on 25 August 2025).
- Sampedro-Castañeda, M.; Baltussen, L.L.; Lopes, A.T.; Qiu, Y.; Sirvio, L.; Mihaylov, S.R.; Claxton, S.; Richardson, J.C.; Lignani, G.; Ultanir, S.K. Epilepsy-linked kinase CDKL5 phosphorylates voltage-gated calcium channel Cav2.3, altering inactivation kinetics and neuronal excitability. Nat. Commun. 2023, 14, 7830. [Google Scholar] [CrossRef]
- Thinwa, J.W.; Zou, Z.; Parks, E.; Sebti, S.; Hui, K.; Wei, Y.; Goodarzi, M.; Singh, V.; Urquhart, G.; Jewell, J.L.; et al. CDKL5 regulates p62-mediated selective autophagy and confers protection against neurotropic viruses. J. Clin. Investig. 2024, 134, e168544. [Google Scholar] [CrossRef]
- Canning, P.; Park, K.; Gonçalves, J.; Li, C.; Howard, C.J.; Sharpe, T.D.; Holt, L.J.; Pelletier, L.; Bullock, A.N.; Leroux, M.R. CDKL Family Kinases Have Evolved Distinct Structural Features and Ciliary Function. Cell Rep. 2018, 22, 885–894. [Google Scholar] [CrossRef]
- Fiser, A.; Sali, A. Modeller: Generation and refinement of homology-based protein structure models. Methods Enzymol. 2003, 374, 461–491. [Google Scholar] [CrossRef]
- Mirdita, M.; Schütze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: Making protein folding accessible to all. Nat. Methods 2022, 19, 679–682. [Google Scholar] [CrossRef]
- Pandey, P.; Alexov, E. Most Monogenic Disorders Are Caused by Mutations Altering Protein Folding Free Energy. Int. J. Mol. Sci. 2024, 25, 1963. [Google Scholar] [CrossRef]
- Pandey, P.; Ghimire, S.; Wu, B.; Alexov, E. On the linkage of thermodynamics and pathogenicity. Curr. Opin. Struct. Biol. 2023, 80, 102572. [Google Scholar] [CrossRef]
- Pandey, P.; Panday, S.K.; Rimal, P.; Ancona, N.; Alexov, E. Predicting the Effect of Single Mutations on Protein Stability and Binding with Respect to Types of Mutations. Int. J. Mol. Sci. 2023, 24, 12073. [Google Scholar] [CrossRef]
- Poudel, P.; Shapovalov, I.; Panday, S.K.; Nouri, K.; Davies, P.L.; Greer, P.A.; Alexov, E. In Silico Screening for Small Molecules to Alter Calpain Proteolysis through Modulating Conformation Changes Induced by Heterodimerization. J. Chem. Inf. Model. 2025, 65, 5528–5543. [Google Scholar] [CrossRef]
- Zhang, Z.; Witham, S.; Petukh, M.; Moroy, G.; Miteva, M.; Ikeguchi, Y.; Alexov, E. A rational free energy-based approach to understanding and targeting disease-causing missense mutations. J. Am. Med. Inform. Assoc. 2013, 20, 643–651. [Google Scholar] [CrossRef]
- Poudel, P.; Miteva, M.A.; Alexov, E. Strategies for in Silico Drug Discovery to Modulate Macromolecular Interactions Altered by Mutations. FBL 2025, 30, 26339. [Google Scholar] [CrossRef]
- Fairley, S.; Lowy-Gallego, E.; Perry, E.; Flicek, P. The International Genome Sample Resource (IGSR) collection of open human genomic variation resources. Nucleic Acids Res. 2020, 48, D941–D947. [Google Scholar] [CrossRef]
- Cock, P.J.A.; Antao, T.; Chang, J.T.; Chapman, B.A.; Cox, C.J.; Dalke, A.; Friedberg, I.; Hamelryck, T.; Kauff, F.; Wilczynski, B.; et al. Biopython: Freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 2009, 25, 1422–1423. [Google Scholar] [CrossRef]
- Savojardo, C.; Manfredi, M.; Martelli, P.L.; Casadio, R. DDGemb: Predicting protein stability change upon single- and multi-point variations with embeddings and deep learning. Bioinformatics 2024, 41, btaf019. [Google Scholar] [CrossRef]
- Lin, Z.; Akin, H.; Rao, R.; Hie, B.; Zhu, Z.; Lu, W.; Smetanin, N.; Verkuil, R.; Kabeli, O.; Shmueli, Y.; et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 2023, 379, 1123–1130. [Google Scholar] [CrossRef]
- Kumar, M.D.S. ProTherm and ProNIT: Thermodynamic databases for proteins and protein-nucleic acid interactions. Nucleic Acids Res. 2006, 34, D204–D206. [Google Scholar] [CrossRef]
- Stourac, J.; Dubrava, J.; Musil, M.; Horackova, J.; Damborsky, J.; Mazurenko, S.; Bednar, D. FireProtDB: Database of manually curated protein stability data. Nucleic Acids Res. 2021, 49, D319–D324. [Google Scholar] [CrossRef]
- Dehouck, Y.; Kwasigroch, J.M.; Gilis, D.; Rooman, M. PoPMuSiC 2.1: A web server for the estimation of protein stability changes upon mutation and sequence optimality. BMC Bioinform. 2011, 12, 151. [Google Scholar] [CrossRef]
- Zhou, Y.; Pan, Q.; Pires, D.E.V.; Rodrigues, C.H.M.; Ascher, D.B. DDMut: Predicting effects of mutations on protein stability using deep learning. Nucleic Acids Res. 2023, 51, W122–W128. [Google Scholar] [CrossRef]
- Rodrigues, C.H.M.; Pires, D.E.V.; Ascher, D.B. DynaMut2: Assessing changes in stability and flexibility upon single and multiple point missense mutations. Protein Sci. 2021, 30, 60–69. [Google Scholar] [CrossRef]
- Li, G.; Panday, S.K.; Alexov, E. SAAFEC-SEQ: A Sequence-Based Method for Predicting the Effect of Single Point Mutations on Protein Thermodynamic Stability. Int. J. Mol. Sci. 2021, 22, 606. [Google Scholar] [CrossRef]
- Montanucci, L.; Capriotti, E.; Frank, Y.; Ben-Tal, N.; Fariselli, P. DDGun: An untrained method for the prediction of protein stability changes upon single and multiple point variations. BMC Bioinform. 2019, 20, 335. [Google Scholar] [CrossRef]
- Eddy, S.R. Where did the BLOSUM62 alignment score matrix come from? Nat. Biotechnol. 2004, 22, 1035–1036. [Google Scholar] [CrossRef]
- Sasidharan Nair, P.; Vihinen, M. VariBench: A benchmark database for variations. Hum. Mutat. 2013, 34, 42–49. [Google Scholar] [CrossRef]
- Savojardo, C.; Fariselli, P.; Martelli, P.L.; Casadio, R. INPS-MD: A web server to predict stability of protein variants from sequence and structure. Bioinformatics 2016, 32, 2542–2544. [Google Scholar] [CrossRef]
- Pires, D.E.V.; Ascher, D.B.; Blundell, T.L. mCSM: Predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics 2014, 30, 335–342. [Google Scholar] [CrossRef]
- Capriotti, E.; Fariselli, P.; Casadio, R. I-Mutant2.0: Predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res. 2005, 33, W306–W310. [Google Scholar] [CrossRef]
- Giulini, M.; Reys, V.; Teixeira, J.M.C.; Jiménez-García, B.; Honorato, R.V.; Kravchenko, A.; Xu, X.; Versini, R.; Engel, A.; Verhoeven, S.; et al. HADDOCK3: A Modular and Versatile Platform for Integrative Modeling of Biomolecular Complexes. J. Chem. Inf. Model. 2025, 65, 7315–7324. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Pahari, S.; Li, G.; Murthy, A.K.; Liang, S.; Fragoza, R.; Yu, H.; Alexov, E. SAAMBE-3D: Predicting Effect of Mutations on Protein-Protein Interactions. Int. J. Mol. Sci. 2020, 21, 2563. [Google Scholar] [CrossRef]
- Geng, C.; Vangone, A.; Folkers, G.E.; Xue, L.C.; Bonvin, A.M.J.J. iSEE: Interface structure, evolution, and energy-based machine learning predictor of binding affinity changes upon mutations. Proteins 2019, 87, 110–119. [Google Scholar] [CrossRef]
- Gribskov, M.; McLachlan, A.D.; Eisenberg, D. Profile analysis: Detection of distantly related proteins. Proc. Natl. Acad. Sci. USA 1987, 84, 4355–4358. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Adzhubei, I.; Jordan, D.M.; Sunyaev, S.R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Genet. 2013, 76, 7–20. [Google Scholar] [CrossRef]
- Pejaver, V.; Urresti, J.; Lugo-Martinez, J.; Pagel, K.A.; Lin, G.N.; Nam, H.-J.; Mort, M.; Cooper, D.N.; Sebat, J.; Iakoucheva, L.M.; et al. Inferring the molecular and phenotypic impact of amino acid variants with MutPred2. Nat. Commun. 2020, 11, 5918. [Google Scholar] [CrossRef]
- Cheng, J.; Novati, G.; Pan, J.; Bycroft, C.; Žemgulytė, A.; Applebaum, T.; Pritzel, A.; Wong, L.H.; Zielinski, M.; Sargeant, T.; et al. Accurate proteome-wide missense variant effect prediction with AlphaMissense. Science 2023, 381, eadg7492. [Google Scholar] [CrossRef]
- Graser, S.; Stierhof, Y.-D.; Lavoie, S.B.; Gassner, O.S.; Lamla, S.; Le Clech, M.; Nigg, E.A. Cep164, a Novel Centriole Appendage Protein Required for Primary Cilium Formation. J. Cell Biol. 2007, 179, 321–330. [Google Scholar] [CrossRef]
- Khanam, T.; Muñoz, I.; Weiland, F.; Carroll, T.; Morgan, M.; Borsos, B.N.; Pantazi, V.; Slean, M.; Novak, M.; Toth, R.; et al. CDKL5 Kinase Controls Transcription-Coupled Responses to DNA Damage. EMBO J. 2021, 40, e108271. [Google Scholar] [CrossRef]
- Conaway, R.C.; Conaway, J.W. The Hunt for RNA Polymerase II Elongation Factors: A Historical Perspective. Nat. Struct. Mol. Biol. 2019, 26, 771–776. [Google Scholar] [CrossRef] [PubMed]
- Weems, J.C.; Slaughter, B.D.; Unruh, J.R.; Hall, S.M.; McLaird, M.B.; Gilmore, J.M.; Washburn, M.P.; Florens, L.; Yasukawa, T.; Aso, T.; et al. Assembly of the Elongin A Ubiquitin Ligase Is Regulated by Genotoxic and Other Stresses. J. Biol. Chem. 2015, 290, 15030–15041. [Google Scholar] [CrossRef] [PubMed]
- Pradhan, S.K.; Su, T.; Yen, L.; Jacquet, K.; Huang, C.; Côté, J.; Kurdistani, S.K.; Carey, M.F. EP400 Deposits H3.3 into Promoters and Enhancers during Gene Activation. Mol. Cell 2016, 61, 27–38. [Google Scholar] [CrossRef]
- Nakabayashi, K.; Amann, D.; Ren, Y.; Saarialho-Kere, U.; Avidan, N.; Gentles, S.; MacDonald, J.R.; Puffenberger, E.G.; Christiano, A.M.; Martinez-Mir, A.; et al. Identification of C7orf11 (TTDN1) Gene Mutations and Genetic Heterogeneity in Nonphotosensitive Trichothiodystrophy. Am. J. Hum. Genet. 2005, 76, 510–516. [Google Scholar] [CrossRef]
- Heller, E.R.; Khan, S.G.; Kuschal, C.; Tamura, D.; DiGiovanna, J.J.; Kraemer, K.H. Mutations in the TTDN1 Gene Are Associated with a Distinct Trichothiodystrophy Phenotype. J. Investig. Dermatol. 2015, 135, 734–741. [Google Scholar] [CrossRef]
- Sekiguchi, M.; Katayama, S.; Hatano, N.; Shigeri, Y.; Sueyoshi, N.; Kameshita, I. Identification of Amphiphysin 1 as an Endogenous Substrate for CDKL5, a Protein Kinase Associated with X-Linked Neurodevelopmental Disorder. Arch. Biochem. Biophys. 2013, 535, 257–267. [Google Scholar] [CrossRef]
- Micheva, K.D.; Ramjaun, A.R.; Kay, B.K.; McPherson, P.S. SH3 Domain-Dependent Interactions of Endophilin with Amphiphysin. FEBS Lett. 1997, 414, 308–312. [Google Scholar] [CrossRef] [PubMed]
- Murakami, N.; Xie, W.; Lu, R.C.; Chen-Hwang, M.-C.; Wieraszko, A.; Hwang, Y.W. Phosphorylation of Amphiphysin I by Minibrain Kinase/Dual-Specificity Tyrosine Phosphorylation-Regulated Kinase, a Kinase Implicated in Down Syndrome. J. Biol. Chem. 2006, 281, 23712–23724. [Google Scholar] [CrossRef]
- Zhu, Y.-C.; Li, D.; Wang, L.; Lu, B.; Zheng, J.; Zhao, S.-L.; Zeng, R.; Xiong, Z.-Q. Palmitoylation-Dependent CDKL5-PSD-95 Interaction Regulates Synaptic Targeting of CDKL5 and Dendritic Spine Development. Proc. Natl. Acad. Sci. USA 2013, 110, 9118–9123. [Google Scholar] [CrossRef] [PubMed]
- Barbiero, I.; Peroni, D.; Tramarin, M.; Chandola, C.; Rusconi, L.; Landsberger, N.; Kilstrup-Nielsen, C. The Neurosteroid Pregnenolone Reverts Microtubule Derangement Induced by the Loss of a Functional CDKL5-IQGAP1 Complex. Hum. Mol. Genet. 2017, 26, 3520–3530. [Google Scholar] [CrossRef] [PubMed]
- Swiech, L.; Blazejczyk, M.; Urbanska, M.; Pietruszka, P.; Dortland, B.R.; Malik, A.R.; Wulf, P.S.; Hoogenraad, C.C.; Jaworski, J. CLIP-170 and IQGAP1 Cooperatively Regulate Dendrite Morphology. J. Neurosci. 2011, 31, 4555–4568. [Google Scholar] [CrossRef]
- Grohmanova, K.; Schlaepfer, D.; Hess, D.; Gutierrez, P.; Beck, M.; Kroschewski, R. Phosphorylation of IQGAP1 Modulates Its Binding to Cdc42, Revealing a New Type of Rho-GTPase Regulator. J. Biol. Chem. 2004, 279, 48495–48504. [Google Scholar] [CrossRef]
- Toriyama, M.; Shimada, T.; Kim, K.B.; Mitsuba, M.; Nomura, E.; Katsuta, K.; Sakumura, Y.; Roepstorff, P.; Inagaki, N. Shootin1: A Protein Involved in the Organization of an Asymmetric Signal for Neuronal Polarization. J. Cell Biol. 2006, 175, 147–157. [Google Scholar] [CrossRef]
- Nawaz, M.S.; Giarda, E.; Bedogni, F.; La Montanara, P.; Ricciardi, S.; Ciceri, D.; Alberio, T.; Landsberger, N.; Rusconi, L.; Kilstrup-Nielsen, C. CDKL5 and Shootin1 Interact and Concur in Regulating Neuronal Polarization. PLoS ONE 2016, 11, e0148634. [Google Scholar] [CrossRef]
- Fuchs, C.; Medici, G.; Trazzi, S.; Gennaccaro, L.; Galvani, G.; Berteotti, C.; Ren, E.; Loi, M.; Ciani, E. CDKL5 Deficiency Predisposes Neurons to Cell Death through the Deregulation of SMAD3 Signaling. Brain Pathol. 2019, 29, 658–674, Erratum in Brain Pathol. 2020, 30, 721. [Google Scholar] [CrossRef]
- Kim, J.Y.; Bai, Y.; Jayne, L.A.; Hector, R.D.; Persaud, A.K.; Ong, S.S.; Rojesh, S.; Raj, R.; Feng, M.J.H.H.; Chung, S.; et al. A Kinome-Wide Screen Identifies a CDKL5-SOX9 Regulatory Axis in Epithelial Cell Death and Kidney Injury. Nat. Commun. 2020, 11, 1924. [Google Scholar] [CrossRef] [PubMed]
- Kameshita, I.; Sekiguchi, M.; Hamasaki, D.; Sugiyama, Y.; Hatano, N.; Suetake, I.; Tajima, S.; Sueyoshi, N. Cyclin-Dependent Kinase-like 5 Binds and Phosphorylates DNA Methyltransferase 1. Biochem. Biophys. Res. Commun. 2008, 377, 1162–1167. [Google Scholar] [CrossRef] [PubMed]
- Bertani, I.; Rusconi, L.; Bolognese, F.; Forlani, G.; Conca, B.; De Monte, L.; Badaracco, G.; Landsberger, N.; Kilstrup-Nielsen, C. Functional Consequences of Mutations in CDKL5, an X-Linked Gene Involved in Infantile Spasms and Mental Retardation. J. Biol. Chem. 2006, 281, 32048–32056. [Google Scholar] [CrossRef] [PubMed]
- Trazzi, S.; Fuchs, C.; Viggiano, R.; De Franceschi, M.; Valli, E.; Jedynak, P.; Hansen, F.K.; Perini, G.; Rimondini, R.; Kurz, T.; et al. HDAC4: A Key Factor Underlying Brain Developmental Alterations in CDKL5 Disorder. Hum. Mol. Genet. 2016, 25, 3887–3907. [Google Scholar] [CrossRef]
- Antao, A.; Burton, J.D.; Dawson, D.; Gemmill, J.; Gerstener, Z.; Godfrey, B.; Groel, S.; Jordan, Z.; Ligon, B.; Smith, D.; et al. Modernizing Clemson University’s Palmetto Cluster: Lessons Learned from 17 Years of HPC Administration. In Proceedings of the Practice and Experience in Advanced Research Computing 2024: Human Powered Computing, Providence, RI, USA, 21–25 July 2024; Association for Computing Machinery: New York, NY, USA, 2024; pp. 1–9. [Google Scholar]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Paul, S.K.; Panday, S.K.; Boccuto, L.; Alexov, E. CDKL5 Deficiency Disorder: Revealing the Molecular Mechanism of Pathogenic Variants. Int. J. Mol. Sci. 2025, 26, 8399. https://doi.org/10.3390/ijms26178399
Paul SK, Panday SK, Boccuto L, Alexov E. CDKL5 Deficiency Disorder: Revealing the Molecular Mechanism of Pathogenic Variants. International Journal of Molecular Sciences. 2025; 26(17):8399. https://doi.org/10.3390/ijms26178399
Chicago/Turabian StylePaul, Shamrat Kumar, Shailesh Kumar Panday, Luigi Boccuto, and Emil Alexov. 2025. "CDKL5 Deficiency Disorder: Revealing the Molecular Mechanism of Pathogenic Variants" International Journal of Molecular Sciences 26, no. 17: 8399. https://doi.org/10.3390/ijms26178399
APA StylePaul, S. K., Panday, S. K., Boccuto, L., & Alexov, E. (2025). CDKL5 Deficiency Disorder: Revealing the Molecular Mechanism of Pathogenic Variants. International Journal of Molecular Sciences, 26(17), 8399. https://doi.org/10.3390/ijms26178399










