Characterization and Transcriptome Analysis Reveal Abnormal Pollen Germination in Cytoplasmic Male Sterile Tomato
Abstract
1. Introduction
2. Results
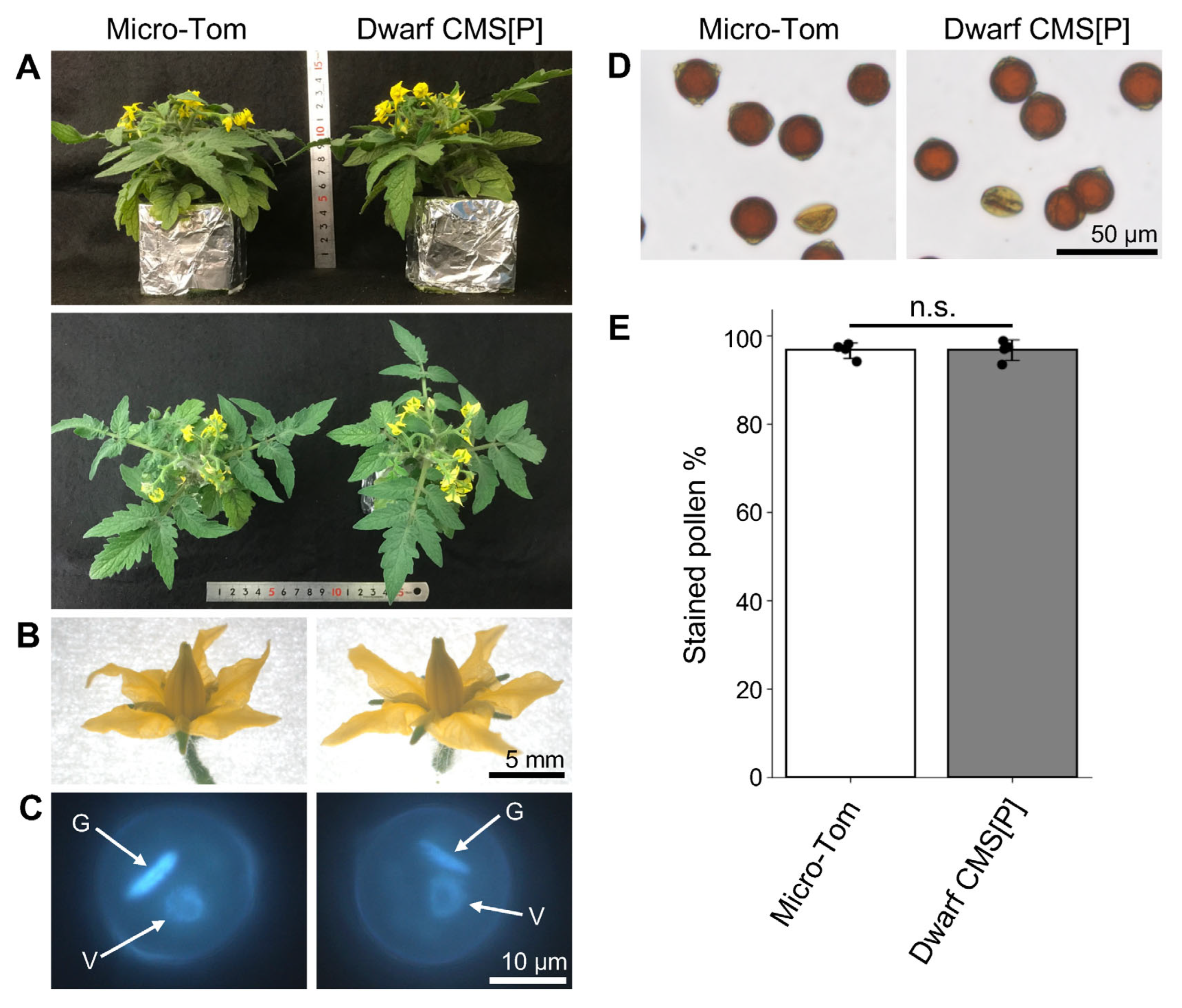
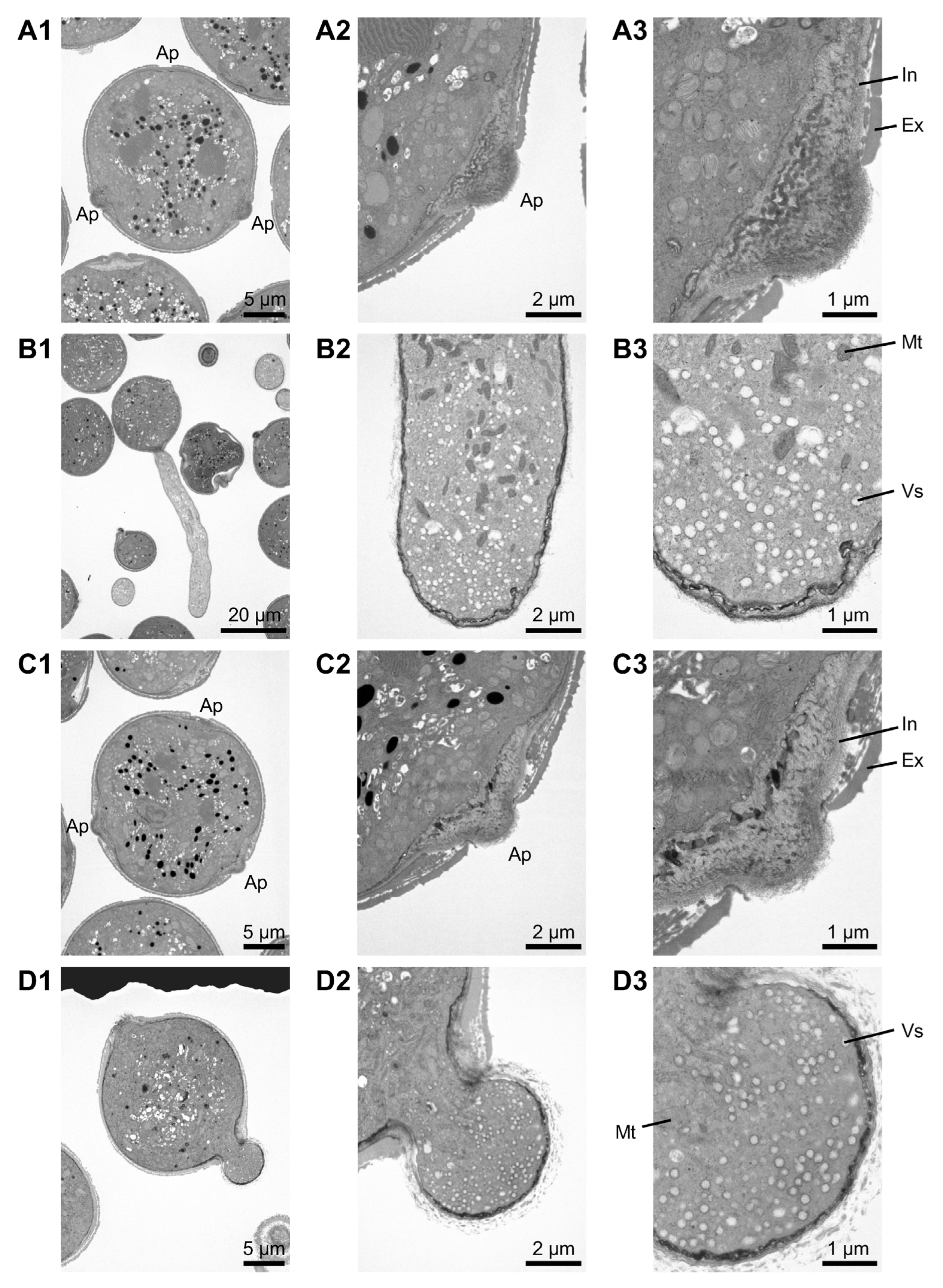
2.1. Abnormal Phenotypes During the Pollen Germination Stage in the Tomato CMS Line
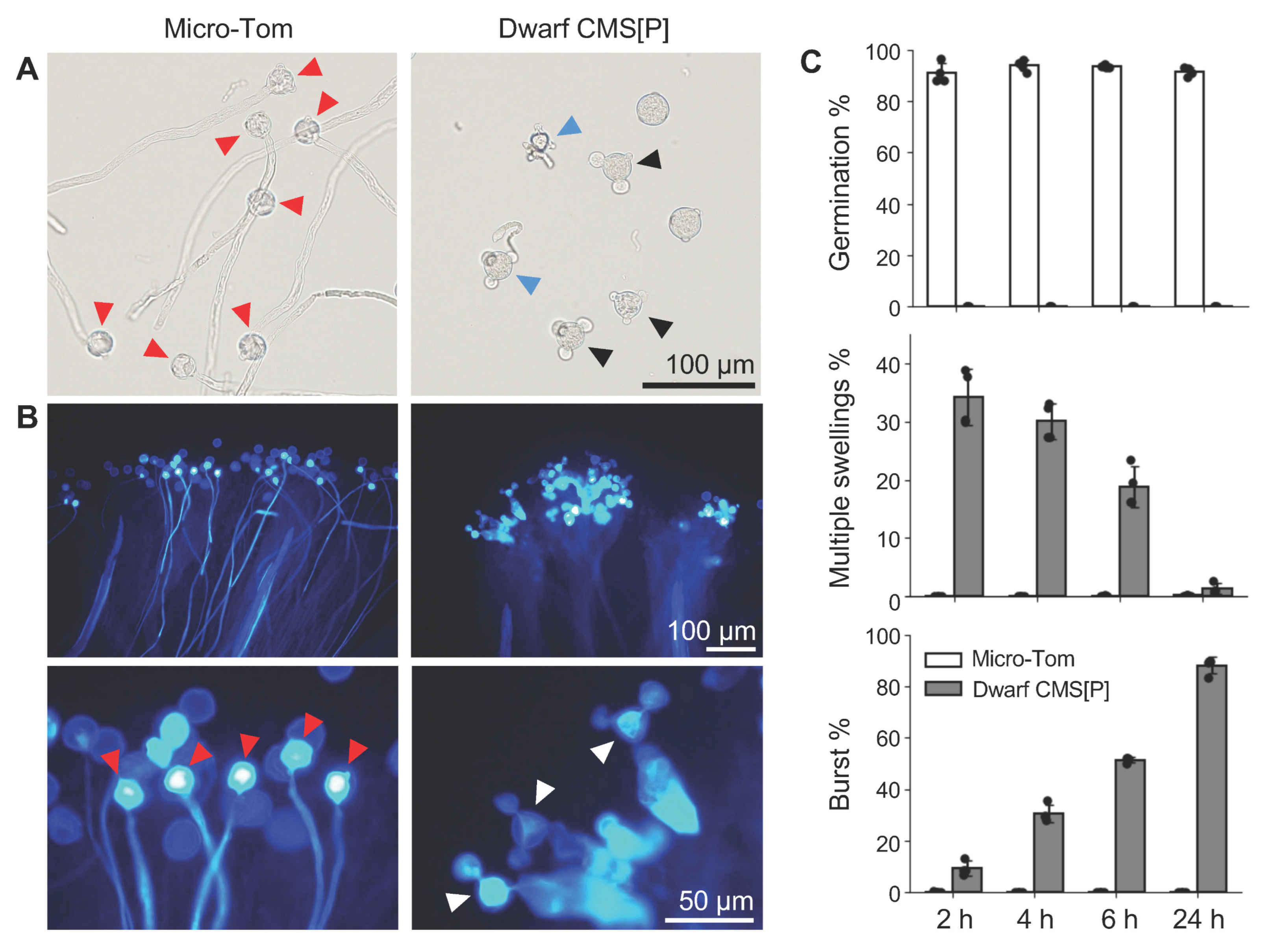
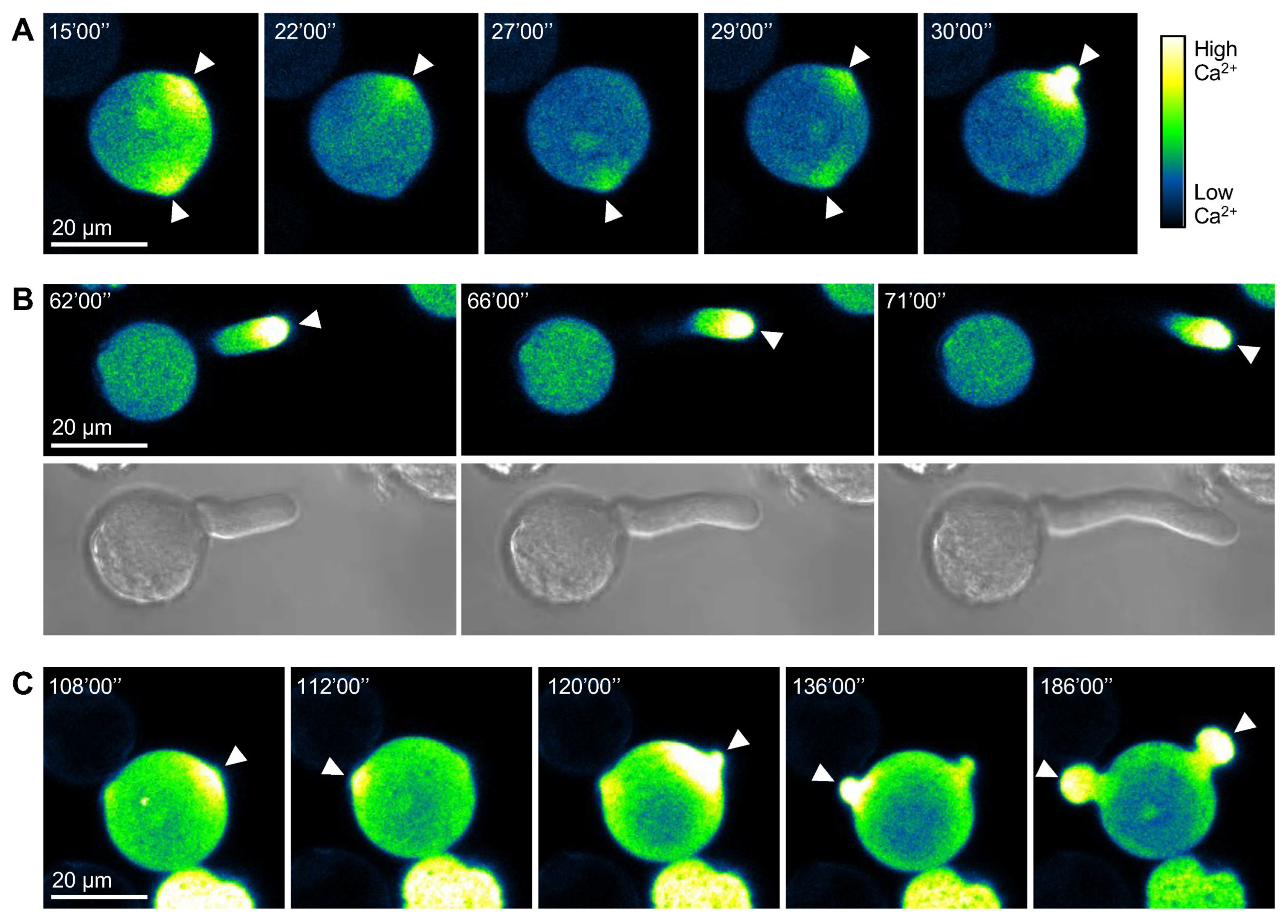
2.2. Pollen Germination at Multiple Sites in Tomato CMS Line
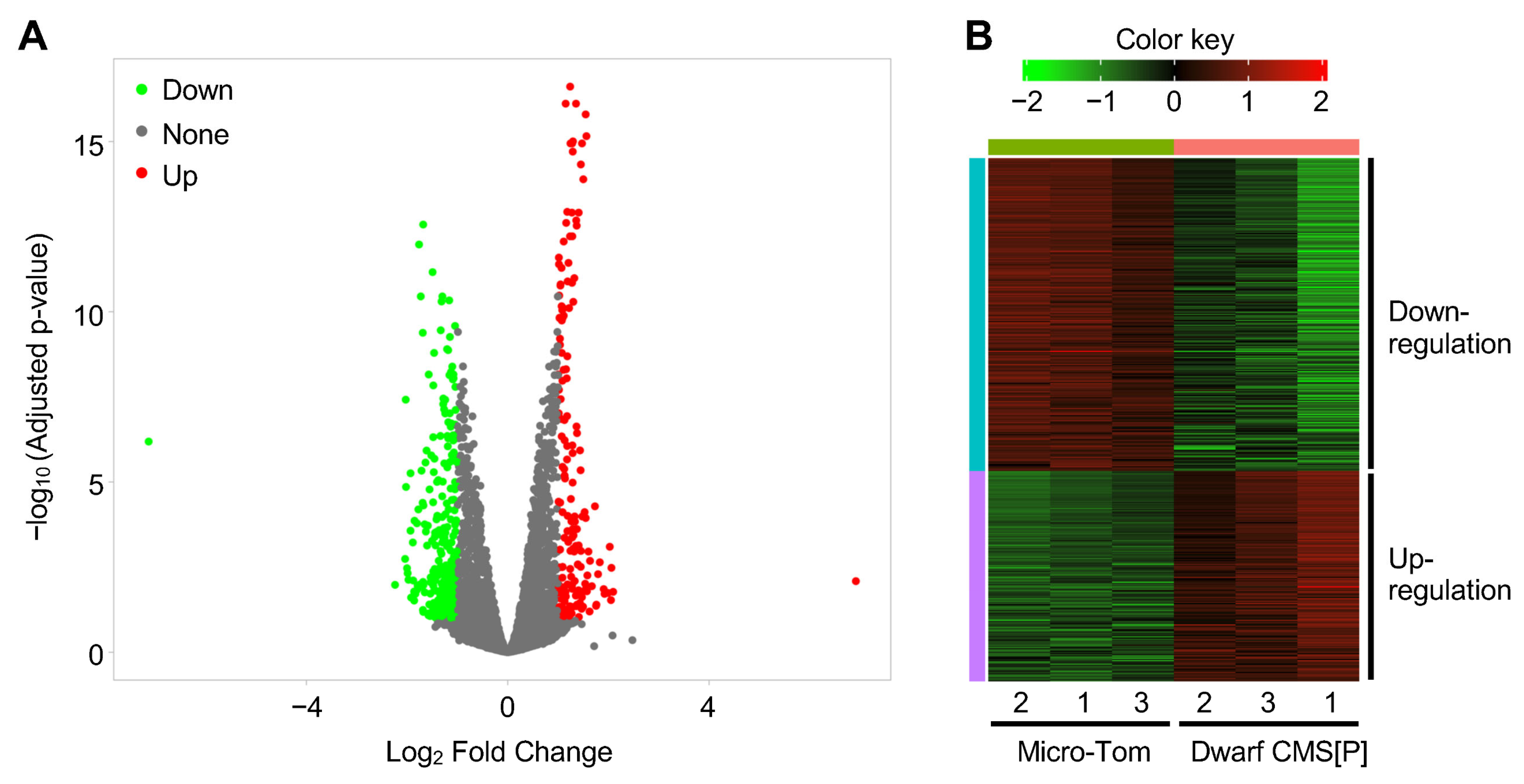
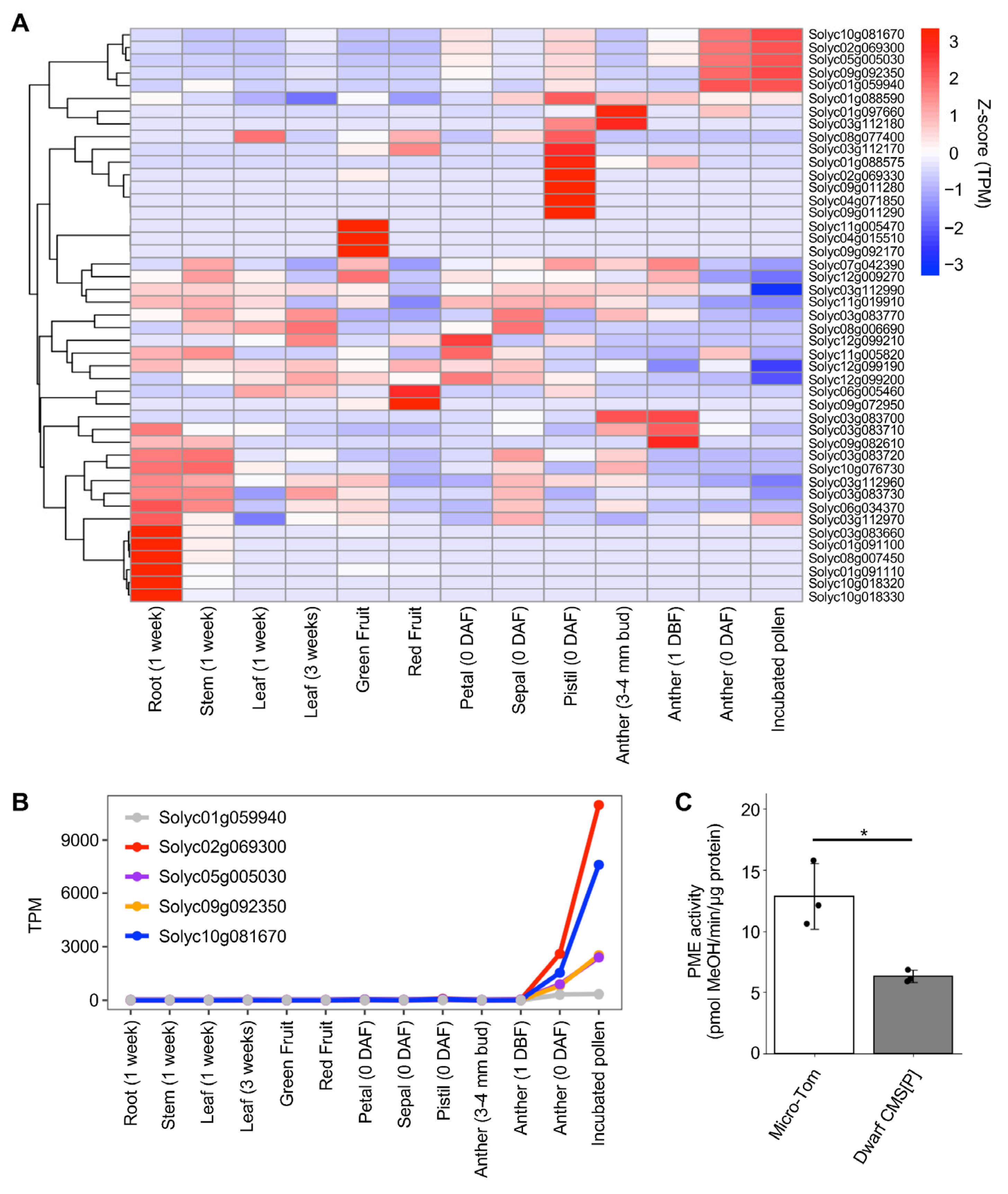
2.3. Transcriptome Analysis in Pollen of Tomato CMS Line
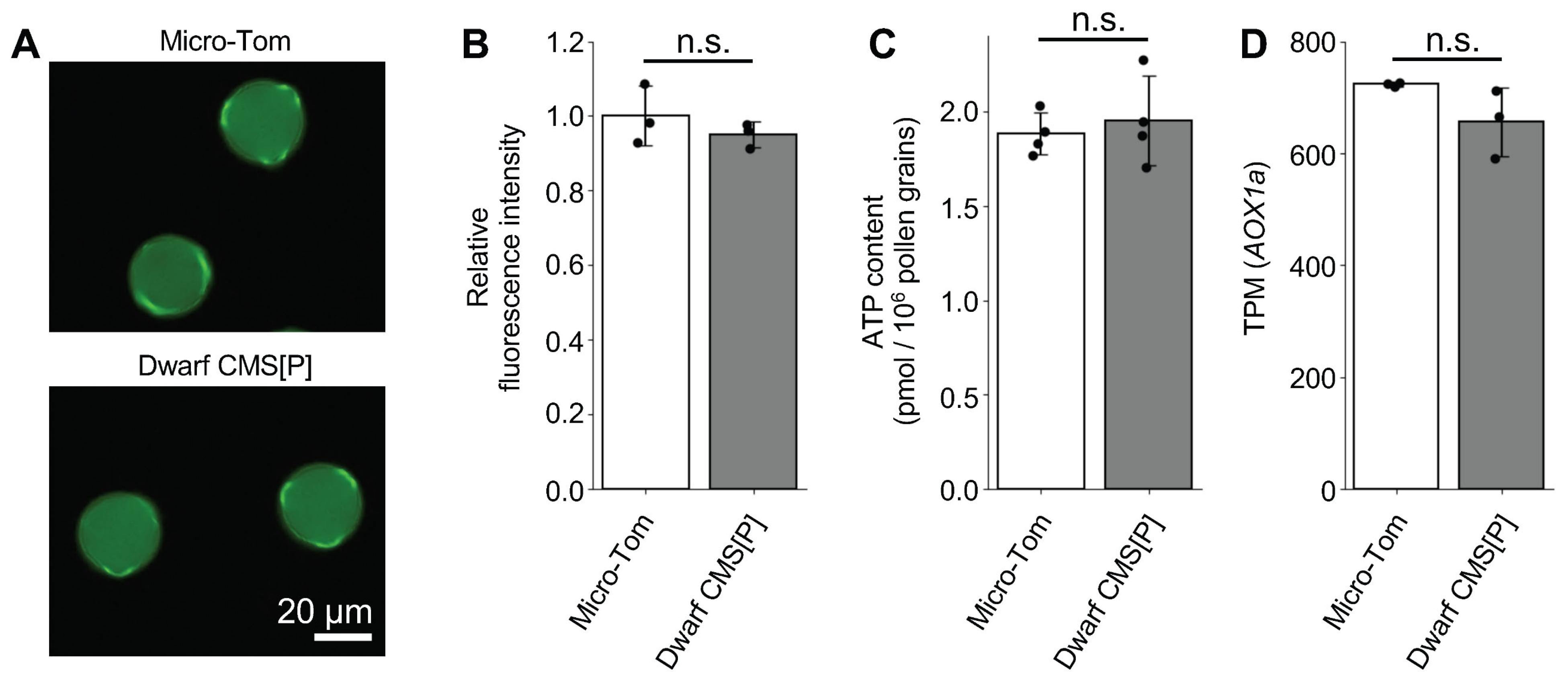
2.4. Assessment of ATP and ROS Production in Tomato CMS Pollen
3. Discussion
3.1. Tomato CMS Lines Exhibit a Unique Male Sterility Phenotype
3.2. Upregulated PMEI Gene Expression Is Associated with Male Sterility in the Tomato CMS Lines
3.3. Normal ATP and ROS Production in Tomato CMS Line
4. Materials and Methods
4.1. Plant Materials
4.2. Phenotypic Analysis of Pollen
4.3. Ca2+ Imaging in Pollen
4.4. RNA Expression Analyses
4.5. PME Activity
4.6. ATP Quantification Assay in Pollen
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CMS | Cytoplasmic male sterility |
| ROS | Reactive oxygen species |
| MRS | Mitochondrial retrograde signaling |
| PCD | Programmed cell death |
| TEM | Transmission electron microscope |
| PME | Pectin methylesterase |
| PMEI | Pectin methylesterase inhibitor |
References
- Bohra, A.; Jha, U.C.; Adhimoolam, P.; Bisht, D.; Singh, N.P. Cytoplasmic Male Sterility (CMS) in Hybrid Breeding in Field Crops. Plant Cell Rep. 2016, 35, 967–993. [Google Scholar] [CrossRef]
- Chen, L.; Liu, Y.-G. Male Sterility and Fertility Restoration in Crops. Annu. Rev. Plant Biol. 2014, 65, 579–606. [Google Scholar] [CrossRef]
- Bhattacharya, J.; Nitnavare, R.; Bhatnagar-Mathur, P.; Palakolanu, S.R. Cytoplasmic Male Sterility-Based Hybrids: Mechanistic Insights. Planta 2024, 260, 100. [Google Scholar] [CrossRef]
- Hu, J.; Huang, W.; Huang, Q.; Qin, X.; Yu, C.; Wang, L.; Li, S.; Zhu, R.; Zhu, Y. Mitochondria and Cytoplasmic Male Sterility in Plants. Mitochondrion 2014, 19, 282–288. [Google Scholar] [CrossRef]
- Biswas, R.; Chaudhuri, S. The Tale of Tapetum: From Anther Walls to Pollen Wall. Nucleus 2024, 67, 611–630. [Google Scholar] [CrossRef]
- Fujii, S.; Toriyama, K. Suppressed Expression of RETROGRADE-REGULATED MALE STERILITY Restores Pollen Fertility in Cytoplasmic Male Sterile Rice Plants. Proc. Natl. Acad. Sci. USA 2009, 106, 9513–9518. [Google Scholar] [CrossRef] [PubMed]
- Suketomo, C.; Kazama, T.; Toriyama, K. Fertility Restoration of Chinese Wild Rice-Type Cytoplasmic Male Sterility by CRISPR/Cas9-Mediated Genome Editing of Nuclear-Encoded RETROGRADE-REGULATED MALE STERILITY. Plant Biotechnol. 2020, 37, 285–292. [Google Scholar] [CrossRef]
- Kuwabara, K.; Harada, I.; Matsuzawa, Y.; Ariizumi, T.; Shirasawa, K. Organelle Genome Assembly Uncovers the Dynamic Genome Reorganization and Cytoplasmic Male Sterility Associated Genes in Tomato. Hortic. Res. 2021, 8, 250. [Google Scholar] [CrossRef] [PubMed]
- Gulyas, G.; Shin, Y.; Kim, H.; Lee, J.-S.; Hirata, Y. Altered Transcript Reveals an Orf507 Sterility-Related Gene in Chili Pepper (Capsicum annuum L.). Plant Mol. Biol. Rep. 2010, 28, 605–612. [Google Scholar] [CrossRef]
- Kuwabara, K.; Arimura, S.-I.; Shirasawa, K.; Ariizumi, T. Orf137 Triggers Cytoplasmic Male Sterility in Tomato. Plant Physiol. 2022, 189, 465–468. [Google Scholar] [CrossRef]
- Iwano, M.; Shiba, H.; Miwa, T.; Che, F.-S.; Takayama, S.; Nagai, T.; Miyawaki, A.; Isogai, A. Ca2+ Dynamics in a Pollen Grain and Papilla Cell during Pollination of Arabidopsis. Plant Physiol. 2004, 136, 3562–3571. [Google Scholar] [CrossRef]
- Diao, M.; Qu, X.; Huang, S. Calcium Imaging in Arabidopsis Pollen Cells Using G-CaMP5. J. Integr. Plant Biol. 2018, 60, 897–906. [Google Scholar] [CrossRef] [PubMed]
- Leroux, C.; Bouton, S.; Kiefer-Meyer, M.-C.; Fabrice, T.N.; Mareck, A.; Guénin, S.; Fournet, F.; Ringli, C.; Pelloux, J.; Driouich, A.; et al. PECTIN METHYLESTERASE48 Is Involved in Arabidopsis Pollen Grain Germination. Plant Physiol. 2015, 167, 367–380. [Google Scholar] [CrossRef] [PubMed]
- Bosch, M.; Hepler, P.K. Pectin Methylesterases and Pectin Dynamics in Pollen Tubes. Plant Cell 2005, 17, 3219–3226. [Google Scholar] [CrossRef]
- Finn, R.D.; Clements, J.; Eddy, S.R. HMMER Web Server: Interactive Sequence Similarity Searching. Nucleic Acids Res. 2011, 39, W29–W37. [Google Scholar] [CrossRef] [PubMed]
- Khan, K.; Tran, H.C.; Mansuroglu, B.; Önsell, P.; Buratti, S.; Schwarzländer, M.; Costa, A.; Rasmusson, A.G.; Van Aken, O. Mitochondria-Derived Reactive Oxygen Species Are the Likely Primary Trigger of Mitochondrial Retrograde Signaling in Arabidopsis. Curr. Biol. 2024, 34, 327–342.e4. [Google Scholar] [CrossRef]
- Xiao, S.; Zang, J.; Pei, Y.; Liu, J.; Liu, J.; Song, W.; Shi, Z.; Su, A.; Zhao, J.; Chen, H. Activation of Mitochondrial Orf355 Gene Expression by a Nuclear-Encoded DREB Transcription Factor Causes Cytoplasmic Male Sterility in Maize. Mol. Plant 2020, 13, 1270–1283. [Google Scholar] [CrossRef]
- Zhao, L.; Rehmani, M.S.; Wang, H. Exocytosis and Endocytosis: Yin-Yang Crosstalk for Sculpting a Dynamic Growing Pollen Tube Tip. Front. Plant Sci. 2020, 11, 572848. [Google Scholar] [CrossRef]
- Ruan, H.; Li, J.; Wang, T.; Ren, H. Secretory Vesicles Targeted to Plasma Membrane During Pollen Germination and Tube Growth. Front. Cell Dev. Biol. 2021, 8, 615447. [Google Scholar] [CrossRef]
- Steinhorst, L.; Kudla, J. Calcium-a Central Regulator of Pollen Germination and Tube Growth. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2013, 1833, 1573–1581. [Google Scholar] [CrossRef]
- Wormit, A.; Usadel, B. The Multifaceted Role of Pectin Methylesterase Inhibitors (PMEIs). Int. J. Mol. Sci. 2018, 19, 2878. [Google Scholar] [CrossRef]
- Yamamoto, M.; Shitsukawa, N.; Yamada, M.; Kato, K.; Takumi, S.; Kawaura, K.; Ogihara, Y.; Murai, K. Identification of a Novel Homolog for a Calmodulin-Binding Protein That Is Upregulated in Alloplasmic Wheat Showing Pistillody. Planta 2013, 237, 1001–1013. [Google Scholar] [CrossRef]
- Hama, E.; Takumi, S.; Ogihara, Y.; Murai, K. Pistillody Is Caused by Alterations to the Class-B MADS-Box Gene Expression Pattern in Alloplasmic Wheats. Planta 2004, 218, 712–720. [Google Scholar] [CrossRef]
- Melchers, G.; Mohri, Y.; Watanabe, K.; Wakabayashi, S.; Harada, K. One-Step Generation of Cytoplasmic Male Sterility by Fusion of Mitochondrial-Inactivated Tomato Protoplasts with Nuclear-Inactivated Solanum Protoplasts. Proc. Natl. Acad. Sci. USA 1992, 89, 6832–6836. [Google Scholar] [CrossRef] [PubMed]
- Scott, J.W.; University of Florida. Agricultural Experiment Station; Harbaugh, B.K. Micro-Tom: A Miniature Dwarf Tomato; Agricultural Experiment Station, Institute of Food and Agricultural Sciences, University of Florida: Gainesville, FL, USA, 1989. [Google Scholar]
- Gao, X.-Q.; Liu, C.Z.; Li, D.D.; Zhao, T.T.; Li, F.; Jia, X.N.; Zhao, X.-Y.; Zhang, X.S. The Arabidopsis KINβγ Subunit of the SnRK1 Complex Regulates Pollen Hydration on the Stigma by Mediating the Level of Reactive Oxygen Species in Pollen. PLoS Genet. 2016, 12, e1006228. [Google Scholar] [CrossRef] [PubMed]
- Abràmoff, M.D.; Magalhães, P.J.; Ram, S.J. Image Processing with ImageJ. Biophotonics Int. 2004, 11, 36–42. [Google Scholar]
- Sun, H.-J.; Uchii, S.; Watanabe, S.; Ezura, H. A Highly Efficient Transformation Protocol for Micro-Tom, a Model Cultivar for Tomato Functional Genomics. Plant Cell Physiol. 2006, 47, 426–431. [Google Scholar] [CrossRef]
- Ezura, K.; Ji-Seong, K.; Mori, K.; Suzuki, Y.; Kuhara, S.; Ariizumi, T.; Ezura, H. Genome-Wide Identification of Pistil-Specific Genes Expressed during Fruit Set Initiation in Tomato (Solanum lycopersicum). PLoS ONE 2017, 12, e0180003. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A Fast Spliced Aligner with Low Memory Requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Hosmani, P.S.; Flores-Gonzalez, M.; van de Geest, H.; Maumus, F.; Bakker, L.V.; Schijlen, E.; van Haarst, J.; Cordewener, J.; Sanchez-Perez, G.; Peters, S.; et al. An Improved de Novo Assembly and Annotation of the Tomato Reference Genome Using Single-Molecule Sequencing, Hi-C Proximity Ligation and Optical Maps. bioRxiv 2019, 767764. [Google Scholar] [CrossRef]
- Vera Alvarez, R.; Pongor, L.S.; Mariño-Ramírez, L.; Landsman, D. TPMCalculator: One-Step Software to Quantify mRNA Abundance of Genomic Features. Bioinformatics 2019, 35, 1960–1962. [Google Scholar] [CrossRef] [PubMed]
- Ge, S.X.; Son, E.W.; Yao, R. iDEP: An Integrated Web Application for Differential Expression and Pathway Analysis of RNA-Seq Data. BMC Bioinform. 2018, 19, 534. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Yano, R.; Nonaka, S.; Ezura, H. Melonet-DB, a Grand RNA-Seq Gene Expression Atlas in Melon (Cucumis melo L.). Plant Cell Physiol. 2018, 59, e4. [Google Scholar] [CrossRef]
- Yano, R.; Ariizumi, T.; Nonaka, S.; Kawazu, Y.; Zhong, S.; Mueller, L.; Giovannoni, J.J.; Rose, J.K.C.; Ezura, H. Comparative Genomics of Muskmelon Reveals a Potential Role for Retrotransposons in the Modification of Gene Expression. Commun. Biol. 2020, 3, 432. [Google Scholar] [CrossRef]
- Klavons, J.A.; Bennett, R.D. Determination of Methanol Using Alcohol Oxidase and Its Application to Methyl Ester Content of Pectins. J. Agric. Food Chem. 1986, 34, 597–599. [Google Scholar] [CrossRef]
- Bradford, M.M. A Rapid and Sensitive Method for the Quantitation of Microgram Quantities of Protein Utilizing the Principle of Protein-Dye Binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Bernard, C.; Traub, M.; Kunz, H.-H.; Hach, S.; Trentmann, O.; Möhlmann, T. Equilibrative Nucleoside Transporter 1 (ENT1) Is Critical for Pollen Germination and Vegetative Growth in Arabidopsis. J. Exp. Bot. 2011, 62, 4627–4637. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kuwabara, K.; Ariizumi, T. Characterization and Transcriptome Analysis Reveal Abnormal Pollen Germination in Cytoplasmic Male Sterile Tomato. Int. J. Mol. Sci. 2025, 26, 8337. https://doi.org/10.3390/ijms26178337
Kuwabara K, Ariizumi T. Characterization and Transcriptome Analysis Reveal Abnormal Pollen Germination in Cytoplasmic Male Sterile Tomato. International Journal of Molecular Sciences. 2025; 26(17):8337. https://doi.org/10.3390/ijms26178337
Chicago/Turabian StyleKuwabara, Kosuke, and Tohru Ariizumi. 2025. "Characterization and Transcriptome Analysis Reveal Abnormal Pollen Germination in Cytoplasmic Male Sterile Tomato" International Journal of Molecular Sciences 26, no. 17: 8337. https://doi.org/10.3390/ijms26178337
APA StyleKuwabara, K., & Ariizumi, T. (2025). Characterization and Transcriptome Analysis Reveal Abnormal Pollen Germination in Cytoplasmic Male Sterile Tomato. International Journal of Molecular Sciences, 26(17), 8337. https://doi.org/10.3390/ijms26178337





