Coiled-Coil Structures Mediate the Intercellular Propagation of Huntingtin
Abstract
1. Introduction
2. Results
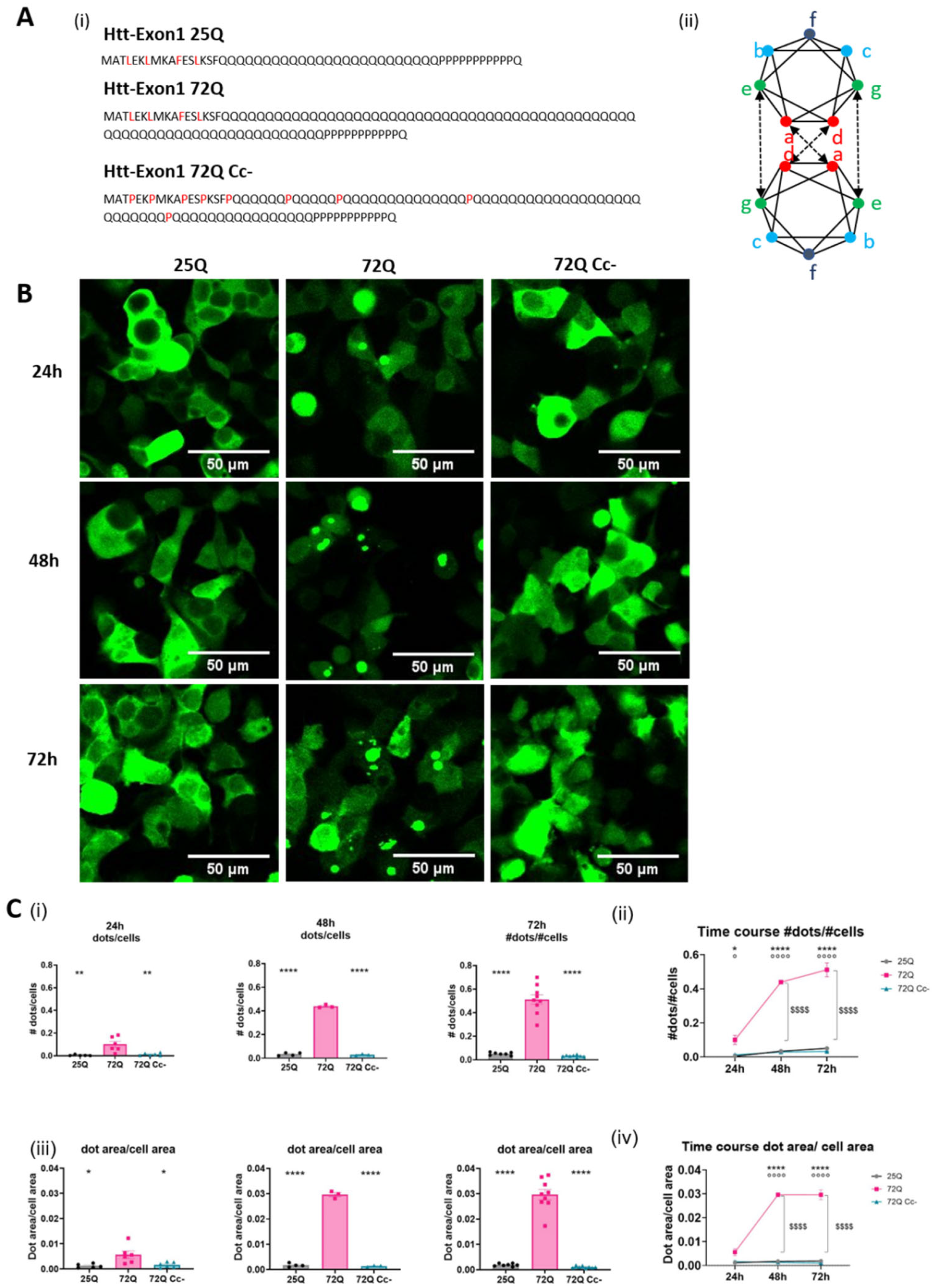
2.1. PolyQ Expanded Huntingtin Achieves the Aggregation Plateau at 48 H Following Transfection
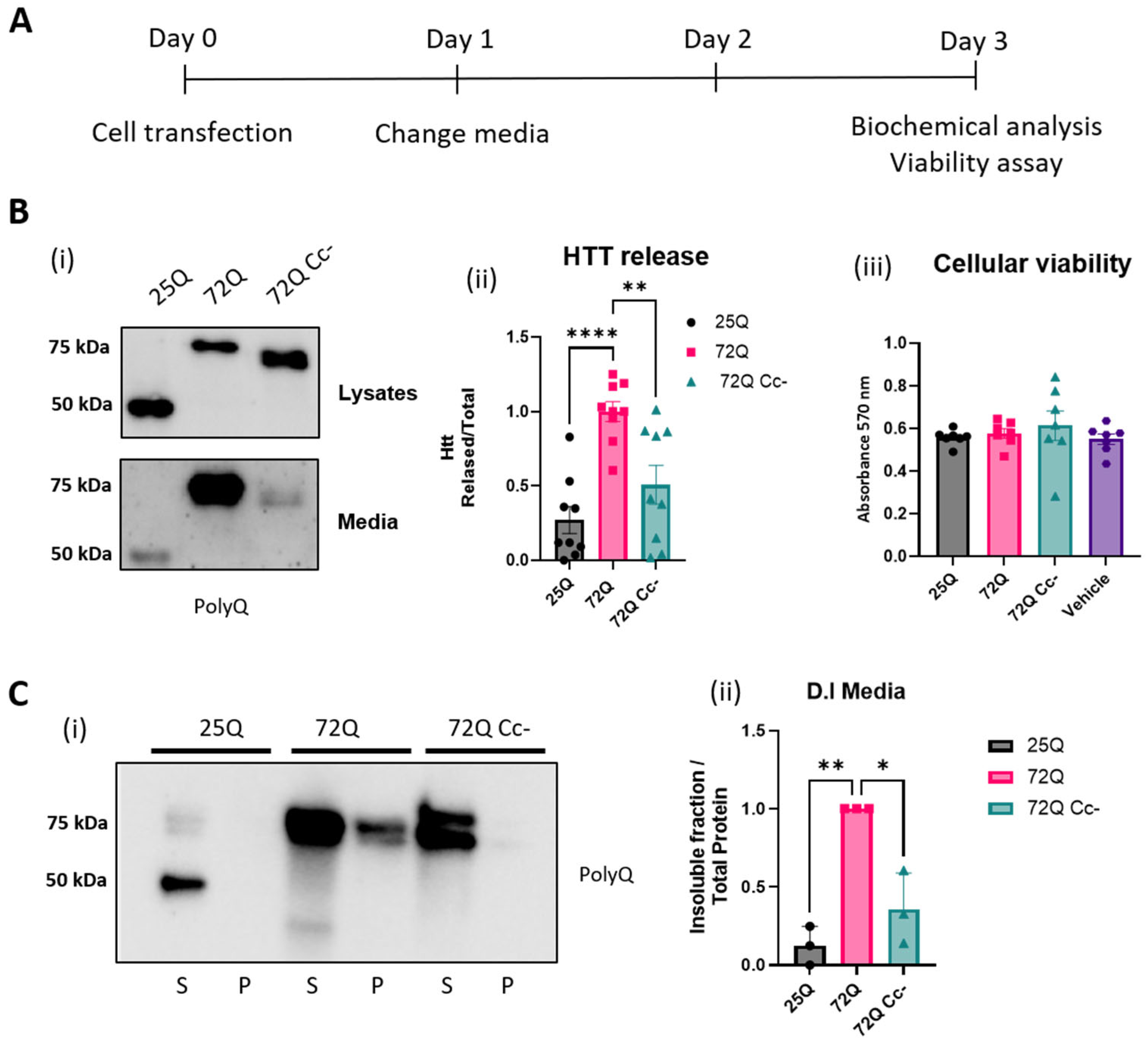
2.2. The Coiled-Coil Structure of Htt Exon 1 Mediates the Formation of Htt Insoluble Oligomers
2.3. Htt Exon 1 Colocalizes with Alix
2.4. Htt 72Q Secretion Is Reduced When the Coiled-Coil Structure Is Hampered
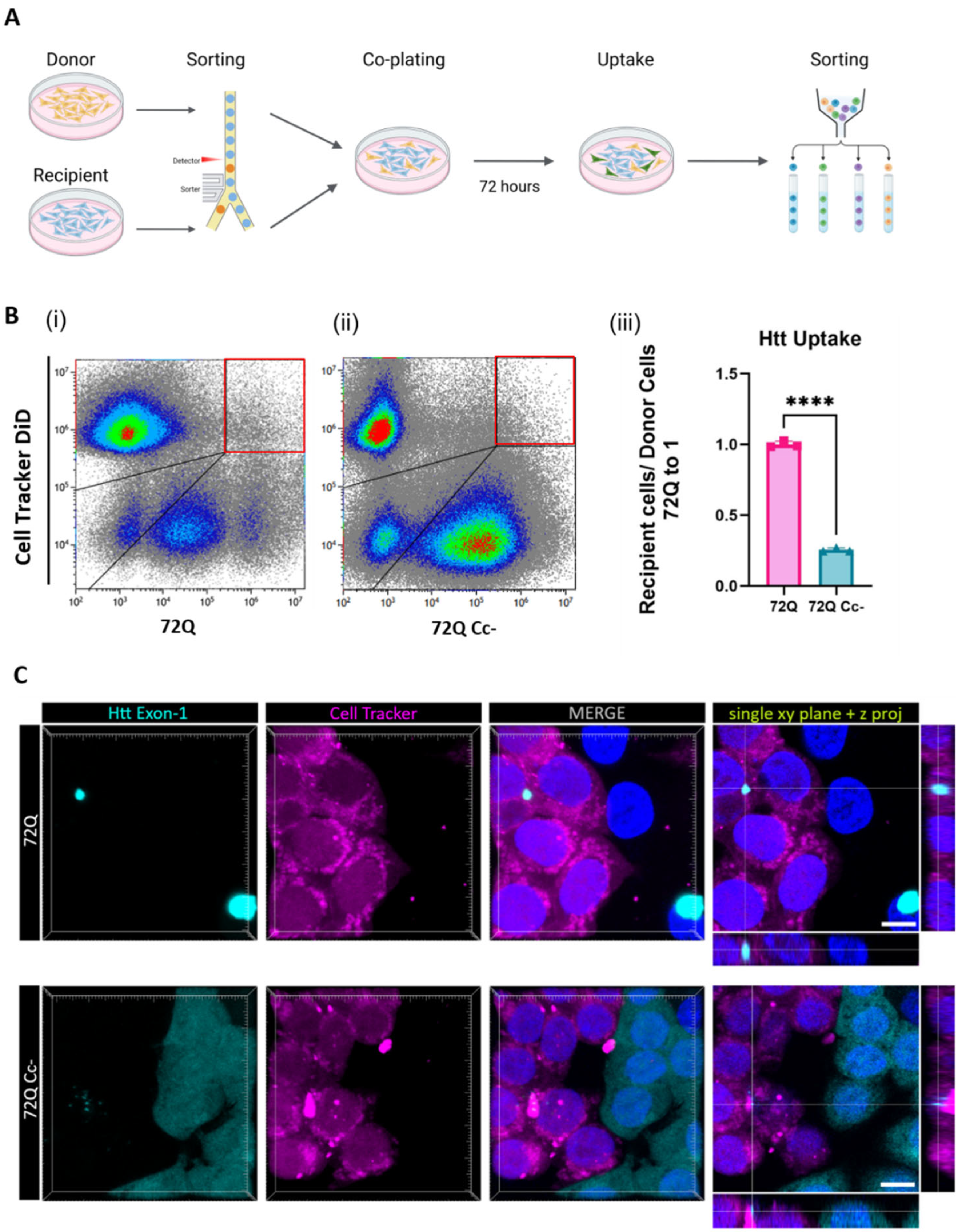
2.5. Htt 72Q Uptake Is Reduced When the Coiled-Coil Structure Is Hampered in Cell-to-Cell Contact Conditions
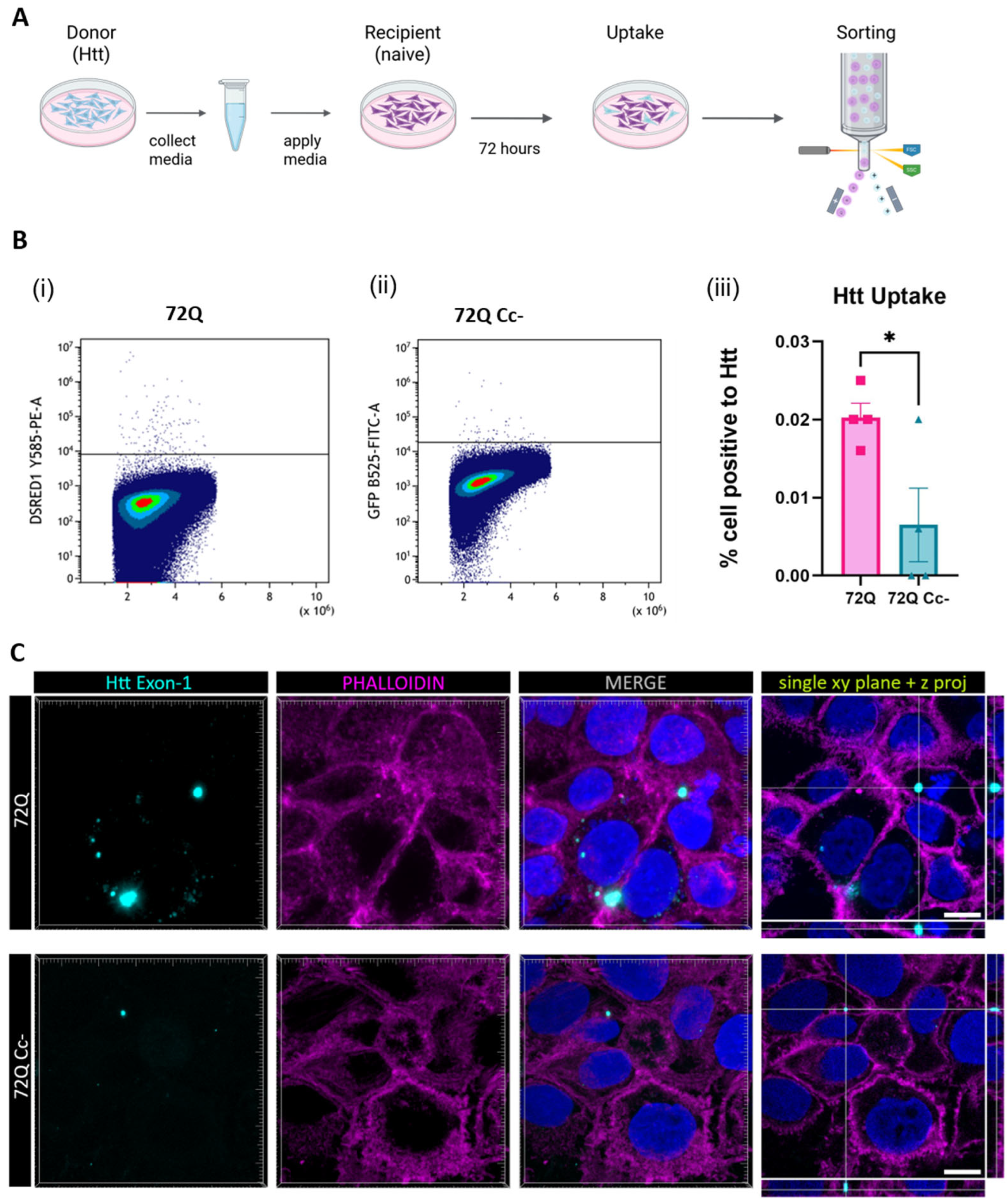
2.6. Coiled-Coil Structure Promotes the Uptake of 72Q Htt Present in the Medium
3. Discussion
4. Materials and Methods
4.1. HEK 293 and SH-SY5Y Cell Culture
4.2. PC12 Cell Culture
4.3. Plasmids and Transient Transfection
4.4. Viability Assays
4.5. Biochemical Assays
4.5.1. Cell Lysis and Protein Extraction
4.5.2. Detergent Insolubility (D.I)
4.5.3. Semi-Denaturing Detergent Agarose Gel Electrophoresis (SDD-AGE)
4.6. Huntingtin Uptake in Co-Plating Conditions
4.7. Huntingtin Uptake Through Conditioned Media
4.8. Cell Sorting
4.9. Flow Cytometry Analysis
4.10. Immunofluorescence Assays
4.11. Confocal Microscope Acquisition
4.12. Huntingtin Granules Quantification
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Aziz, N.A.; Jurgens, C.K.; Landwehrmeyer, G.B.; van Roon-Mom, W.M.C.; van Ommen, G.J.B.; Stijnen, T.; Roos, R.A.C. Normal and mutant HTT interact to affect clinical severity and progression in Huntington disease. Neurology 2009, 73, 1280–1285. [Google Scholar] [CrossRef]
- Bates, G.P.; Dorsey, R.; Gusella, J.F.; Hayden, M.R.; Kay, C.; Leavitt, B.R.; Nance, M.; Ross, C.A.; Scahill, R.I.; Wetzel, R.; et al. Huntington disease. Nat. Rev. Dis. Primers 2015, 1, 15005. [Google Scholar] [CrossRef]
- Hoffner, G.; Souès, S.; Djian, P. Aggregation of Expanded Huntingtin in the Brains of Patients with Huntington Disease. Prion 2007, 1, 26–31. [Google Scholar] [CrossRef]
- DiFiglia, M.; Sapp, E.; Chase, K.O.; Davies, S.W.; Bates, G.P.; Vonsattel, J.P.; Aronin, N. Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science 1997, 277, 1990–1993. [Google Scholar] [CrossRef]
- Alpaugh, M.; Cicchetti, F. Huntington’s disease: Lessons from prion disorders. J. Neurol. 2021, 268, 3493–3504. [Google Scholar] [CrossRef]
- Babcock, D.T.; Ganetzky, B. Transcellular spreading of huntingtin aggregates in the Drosophila brain. Proc. Natl. Acad. Sci. USA 2015, 112, E5427–E5433. [Google Scholar] [CrossRef]
- Ren, P.-H.; Lauckner, J.E.; Kachirskaia, I.; Heuser, J.E.; Melki, R.; Kopito, R.R. Cytoplasmic penetration and persistent infection of mammalian cells by polyglutamine aggregates. Nat. Cell Biol. 2009, 11, 219–225. [Google Scholar] [CrossRef]
- Cicchetti, F.; Soulet, D.; Freeman, T.B. Neuronal degeneration in striatal transplants and Huntington’s disease: Potential mechanisms and clinical implications. Brain 2011, 134, 641–652. [Google Scholar] [CrossRef]
- Ehrlich, M.E. Huntington’s Disease and the Striatal Medium Spiny Neuron: Cell-Autonomous and Non-Cell-Autonomous Mechanisms of Disease. Neurotherapeutics 2012, 9, 270–284. [Google Scholar] [CrossRef]
- Pecho-Vrieseling, E.; Rieker, C.; Fuchs, S.; Bleckmann, D.; Esposito, M.S.; Botta, P.; Goldstein, C.; Bernhard, M.; Galimberti, I.; Müller, M.; et al. Transneuronal propagation of mutant huntingtin contributes to non-cell autonomous pathology in neurons. Nat. Neurosci. 2014, 17, 1064–1072, Erratum in Nat. Neurosci. 2018, 21, 1291. [Google Scholar] [CrossRef]
- Dinamarca, M.C.; Colombo, L.; Brykczynska, U.; Grimm, A.; Fruh, I.; Hossain, I.; Gabriel, D.; Eckert, A.; Müller, M.; Pecho-Vrieseling, E. Transmission-selective muscle pathology induced by the active propagation of mutant huntingtin across the human neuromuscular synapse. Front. Mol. Neurosci. 2023, 16, 1287510. [Google Scholar] [CrossRef] [PubMed]
- Abounit, S.; Delage, E.; Zurzolo, C. Identification and Characterization of Tunneling Nanotubes for Intercellular Trafficking. In Current Protocols in Cell Biology; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2001; Available online: http://onlinelibrary.wiley.com.ezproxy.cul.columbia.edu/doi/10.1002/0471143030.cb1210s67/abstract (accessed on 25 June 2015).
- Costanzo, M.; Zurzolo, C. The cell biology of prion-like spread of protein aggregates: Mechanisms and implication in neurodegeneration. Biochem. J. 2013, 452, 1–17. [Google Scholar] [CrossRef]
- Costanzo, M.; Abounit, S.; Marzo, L.; Danckaert, A.; Chamoun, Z.; Roux, P.; Zurzolo, C. Transfer of polyglutamine aggregates in neuronal cells occurs in tunneling nanotubes. J. Cell. Sci. 2013, 126 Pt 16, 3678–3685. [Google Scholar] [CrossRef]
- Jeon, I.; Cicchetti, F.; Cisbani, G.; Lee, S.; Li, E.; Bae, J.; Lee, N.; Li, L.; Im, W.; Kim, M.; et al. Human-to-mouse prion-like propagation of mutant huntingtin protein. Acta Neuropathol. 2016, 132, 577–592. [Google Scholar] [CrossRef]
- Fiumara, F.; Fioriti, L.; Kandel, E.R.; Hendrickson, W.A. Essential role of coiled coils for aggregation and activity of Q/N-rich prions and PolyQ proteins. Cell 2010, 143, 1121–1135. [Google Scholar] [CrossRef] [PubMed]
- Mason, J.M.; Arndt, K.M. Coiled Coil Domains: Stability, Specificity, and Biological Implications. ChemBioChem 2004, 5, 170–176. [Google Scholar] [CrossRef]
- Parry, D.A.D.; Fraser, R.D.B.; Squire, J.M. Fifty years of coiled-coils and α-helical bundles: A close relationship between sequence and structure. J. Struct. Biol. 2008, 163, 258–269. [Google Scholar] [CrossRef]
- Kwok, S.C.; Hodges, R.S. Stabilizing and Destabilizing Clusters in the Hydrophobic Core of Long Two-stranded α-Helical Coiled-coils. J. Biol. Chem. 2004, 279, 21576–21588. [Google Scholar] [CrossRef]
- Suzuki, K.; Mizuno, T.; Tanaka, T. Regulation of protein-protein interaction via assembly of coiled-coil domain. Nucleic Acids Symp. Ser. 2008, 52, 461. [Google Scholar] [CrossRef] [PubMed]
- Kokona, B.; Rosenthal, Z.P.; Fairman, R. Role of the Coiled-Coil Structural Motif in Polyglutamine Aggregation. Biochemistry 2014, 53, 6738–6746. [Google Scholar] [CrossRef]
- Petrakis, S.; Schaefer, M.H.; Wanker, E.E.; Andrade-Navarro, M.A. Aggregation of polyQ-extended proteins is promoted by interaction with their natural coiled-coil partners: Insights & Perspectives. BioEssays 2013, 35, 503–507. [Google Scholar] [CrossRef]
- Batlle, C.; Calvo, I.; Iglesias, V.; Lynch, C.J.; Gil-Garcia, M.; Serrano, M.; Ventura, S. MED15 prion-like domain forms a coiled-coil responsible for its amyloid conversion and propagation. Commun. Biol. 2021, 4, 414. [Google Scholar] [CrossRef]
- Versluis, F.; Voskuhl, J.; Vos, J.; Friedrich, H.; Ravoo, B.J.; Bomans, P.H.H.; Stuart, M.C.A.; Sommerdijk, N.A.J.M.; Kros, A. Coiled coil driven membrane fusion between cyclodextrin vesicles and liposomes. Soft Matter 2014, 10, 9746–9751. [Google Scholar] [CrossRef]
- Zhuang, M.; Wang, W.; De Feo, C.J.; Vassell, R.; Weiss, C.D. Trimeric, coiled-coil extension on peptide fusion inhibitor of HIV-1 influences selection of resistance pathways. J. Biol. Chem. 2012, 287, 8297–8309. [Google Scholar] [CrossRef]
- Kowal, J.; Tkach, M.; Théry, C. Biogenesis and Secretion of Exosomes. Curr. Opin. Cell Biol. 2014, 29, 116–125. [Google Scholar] [CrossRef]
- Chang, D.-K.; Cheng, S.-F.; Trivedi, V.D.; Lin, K.-L. Proline Affects Oligomerization of a Coiled Coil by Inducing a Kink in a Long Helix. J. Struct. Biol. 1999, 128, 270–279. [Google Scholar] [CrossRef]
- Fioriti, L.; Myers, C.; Huang, Y.-Y.; Li, X.; Stephan, J.S.; Trifilieff, P.; Colnaghi, L.; Kosmidis, S.; Drisaldi, B.; Pavlopoulos, E.; et al. The Persistence of Hippocampal-Based Memory Requires Protein Synthesis Mediated by the Prion-like Protein CPEB3. Neuron 2015, 86, 1433–1448. [Google Scholar] [CrossRef] [PubMed]
- Drisaldi, B.; Colnaghi, L.; Fioriti, L.; Rao, N.; Myers, C.; Snyder, A.M.; Metzger, D.J.; Tarasoff, J.; Konstantinov, E.; Fraser, P.E.; et al. SUMOylation Is an Inhibitory Constraint that Regulates the Prion-like Aggregation and Activity of CPEB3. Cell Rep. 2015, 11, 1694–1702. [Google Scholar] [CrossRef] [PubMed]
- Stephan, J.S.; Fioriti, L.; Lamba, N.; Colnaghi, L.; Karl, K.; Derkatch, I.L.; Kandel, E.R. The CPEB3 Protein Is a Functional Prion that Interacts with the Actin Cytoskeleton. Cell Rep. 2015, 11, 1772–1785. [Google Scholar] [CrossRef]
- Weiss, A.; Abramowski, D.; Bibel, M.; Bodner, R.; Chopra, V.; DiFiglia, M.; Fox, J.; Kegel, K.; Klein, C.; Grueninger, S.; et al. Single-step detection of mutant huntingtin in animal and human tissues: A bioassay for Huntington’s disease. Anal. Biochem. 2009, 395, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Fioriti, L.; Quaglio, E.; Massignan, T.; Colombo, L.; Stewart, R.S.; Salmona, M.; Harris, D.A.; Forloni, G.; Chiesa, R. The neurotoxicity of prion protein (PrP) peptide 106-126 is independent of the expression level of PrP and is not mediated by abnormal PrP species. Mol. Cell. Neurosci. 2005, 28, 165–176. [Google Scholar] [CrossRef]
- Fioriti, L.; Dossena, S.; Stewart, L.R.; Stewart, R.S.; Harris, D.A.; Forloni, G.; Chiesa, R. Cytosolic prion protein (PrP) is not toxic in N2a cells and primary neurons expressing pathogenic PrP mutations. J. Biol. Chem. 2005, 280, 11320–11328. [Google Scholar] [CrossRef]
- Février, B.; Vilette, D.; Laude, H.; Raposo, G. Exosomes: A bubble ride for prions? Traffic 2005, 6, 10–17. [Google Scholar] [CrossRef] [PubMed]
- Chivet, M.; Hemming, F.; Pernet-Gallay, K.; Fraboulet, S.; Sadoul, R. Emerging Role of Neuronal Exosomes in the Central Nervous System. Front. Physio. 2012, 3, 145. [Google Scholar] [CrossRef]
- Chivet, M.; Javalet, C.; Hemming, F.; Pernet-Gallay, K.; Laulagnier, K.; Fraboulet, S.; Sadoul, R. Exosomes as a novel way of interneuronal communication. Biochem. Soc. Trans. 2013, 41, 241–244. [Google Scholar] [CrossRef] [PubMed]
- Abounit, S.; Wu, J.W.; Duff, K.; Victoria, G.S.; Zurzolo, C. Tunneling nanotubes: A possible highway in the spreading of tau and other prion-like proteins in neurodegenerative diseases. Prion 2016, 10, 344–351. [Google Scholar] [CrossRef] [PubMed]
- Garden, G.A.; La Spada, A.R. Intercellular (Mis)communication in Neurodegenerative Disease. Neuron 2012, 73, 886–901. [Google Scholar] [CrossRef]
- Alpaugh, M.; Denis, H.L.; Cicchetti, F. Prion-like properties of the mutant huntingtin protein in living organisms: The evidence and the relevance. Mol. Psychiatry 2022, 27, 269–280. [Google Scholar] [CrossRef]
- Donnelly, K.M.; DeLorenzo, O.R.; Zaya, A.D.; Pisano, G.E.; Thu, W.M.; Luo, L.; Kopito, R.R.; Panning Pearce, M.M. Phagocytic glia are obligatory intermediates in transmission of mutant huntingtin aggregates across neuronal synapses. Elife 2020, 9, e58499. [Google Scholar] [CrossRef]
- Pearce, M.M.P.; Spartz, E.J.; Hong, W.; Luo, L.; Kopito, R.R. Prion-like transmission of neuronal huntingtin aggregates to phagocytic glia in the Drosophila brain. Nat. Commun. 2015, 6, 6768. [Google Scholar] [CrossRef]
- Bartl, S.; Oueslati, A.; Southwell, A.L.; Siddu, A.; Parth, M.; David, L.S.; Maxan, A.; Salhat, N.; Burkert, M.; Mairhofer, A.; et al. Inhibiting cellular uptake of mutant huntingtin using a monoclonal antibody: Implications for the treatment of Huntington’s disease. Neurobiol. Dis. 2020, 141, 104943. [Google Scholar] [CrossRef]
- Trajkovic, K.; Jeong, H.; Krainc, D. Mutant Huntingtin Is Secreted via a Late Endosomal/Lysosomal Unconventional Secretory Pathway. J. Neurosci. 2017, 37, 9000–9012. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Behbahanipour, M.; García-Pardo, J.; Ventura, S. Decoding the role of coiled-coil motifs in human prion-like proteins. Prion 2021, 15, 143–154. [Google Scholar] [CrossRef]
- Cheng, H.; Sablitzky, F. The coiled coil domain of DEF6 facilitates formation of large vesicle-like, cytoplasmic aggregates that trap the P-body marker DCP1 and exhibit prion-like features. bioRxiv 2019. [Google Scholar] [CrossRef]
- Gerlich, D.W. Cell organization by liquid phase separation. Nat. Rev. Mol. Cell Biol. 2017, 18, 593. [Google Scholar] [CrossRef]
- Hayashi, Y.; Ford, L.K.; Fioriti, L.; McGurk, L.; Zhang, M. Liquid-Liquid Phase Separation in Physiology and Pathophysiology of the Nervous System. J. Neurosci. 2021, 41, 834. [Google Scholar] [CrossRef]
- Ford, L.K.; Fioriti, L. Coiled-Coil Motifs of RNA-Binding Proteins: Dynamicity in RNA Regulation. Front. Cell Dev. Biol. 2020, 8, 607947. [Google Scholar] [CrossRef]
- Strauss, H.M.; Keller, S. Pharmacological interference with protein-protein interactions mediated by coiled-coil motifs. Handb. Exp. Pharmacol. 2008, 186, 461–482. [Google Scholar] [CrossRef]


| Antibody (Clone) | Host Animal | Dilution for Western Blot | Distributor, Catalog Number |
|---|---|---|---|
| PolyQ (3B5H10) | Monoclonal Mouse | 1:1000 | Sigma, P1874 |
| Huntingtin (EM-48) | Monoclonal Mouse | 1:500 | Merck Millipore, MAB5374 |
| GAPDH (FL-335) | Polyclonal Rabbit | 1:1000 | Santacruz, sc-25778 |
| Antibody (Clone) | Host Animal | Dilution for Western Blot | Distributor, Catalog Number |
|---|---|---|---|
| Anti HRP | Mouse | 1:10,000 | Jackson Immunoresearch, #115-035-174 |
| Anti HRP | Rabbit | 1:10,000 | GE Healthcare, NA934 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bosica, M.; Grasselli, C.; Panfili, A.; Orsini, F.; Fioriti, L. Coiled-Coil Structures Mediate the Intercellular Propagation of Huntingtin. Int. J. Mol. Sci. 2025, 26, 8162. https://doi.org/10.3390/ijms26178162
Bosica M, Grasselli C, Panfili A, Orsini F, Fioriti L. Coiled-Coil Structures Mediate the Intercellular Propagation of Huntingtin. International Journal of Molecular Sciences. 2025; 26(17):8162. https://doi.org/10.3390/ijms26178162
Chicago/Turabian StyleBosica, Marco, Chiara Grasselli, Andrea Panfili, Franca Orsini, and Luana Fioriti. 2025. "Coiled-Coil Structures Mediate the Intercellular Propagation of Huntingtin" International Journal of Molecular Sciences 26, no. 17: 8162. https://doi.org/10.3390/ijms26178162
APA StyleBosica, M., Grasselli, C., Panfili, A., Orsini, F., & Fioriti, L. (2025). Coiled-Coil Structures Mediate the Intercellular Propagation of Huntingtin. International Journal of Molecular Sciences, 26(17), 8162. https://doi.org/10.3390/ijms26178162






