When Genes Wear Marks: Epigenomic Modulation in the Development and Progression of Obesity
Abstract
1. Introduction
2. DNA Methylation
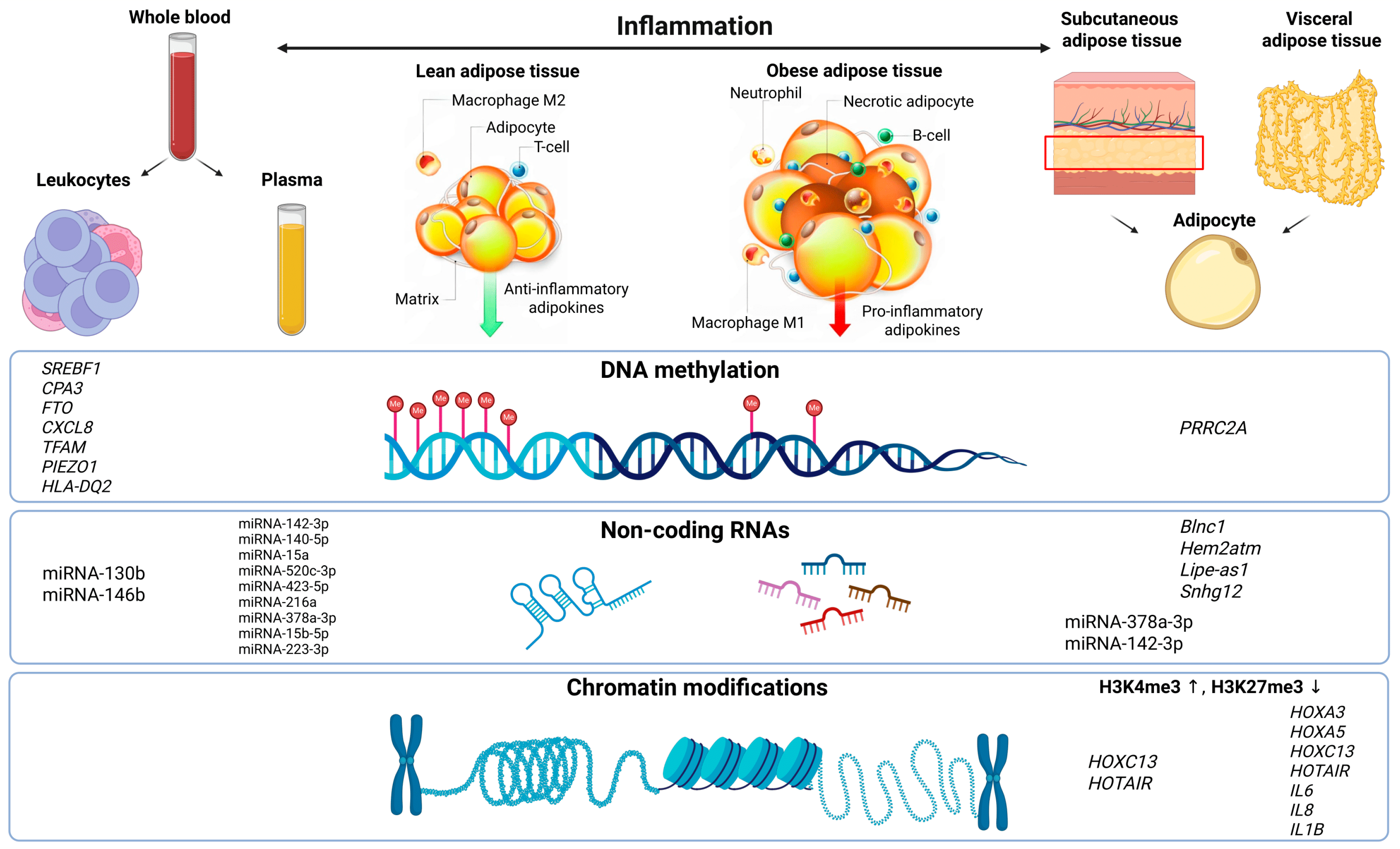
2.1. Abnormal DNA Methylation in Immune System and Peripheral Blood Cells in Obesity
2.2. Tissue-Specific Features of Abnormal DNA Methylation in Severe Obesity
| Sample, n | BMI, Percentile | Detection Method | Material | Biomarkers | Source |
|---|---|---|---|---|---|
| 28 | ≥95 | HumanMethylation 450K BeadChip | Saliva | NRF1 (cg01307483) | [53] |
| 94 | ≥99 | MethylationEPIC BeadChip | Peripheral blood | SREBF1 (cg11024682) CPA3 (cg13424229) | [37] |
| 16 | ≥99 | MSRE-qPCR | Peripheral blood mononuclear cells | FTO | [39] |
| 31 | ≥95 | Methylation-specific PCR | Peripheral blood | CXCL8 | [38] |
| 190 | ≥99 | HumanMethylation 450K BeadChip, MethylationEPIC BeadChip | Adipocytes of visceral and subcutaneous adipose tissue | PRRC2A | [14] |
| 41 | ≥97 | MethylationEPIC BeadChip, Pyrosequencing | Peripheral blood | TFAM (cg05831083), PIEZO1 (cg14926485) | [36] |
3. The Role of Non-Coding RNAs in the Mechanisms of Obesity Development
3.1. Circulating MicroRNAs Expression in Plasma
3.2. Tissue-Specific Changes in MicroRNA Expression
3.3. Long Non-Coding RNAs in the Pathogenesis of Obesity
3.4. Methylation of MiRNA Genes in Obesity
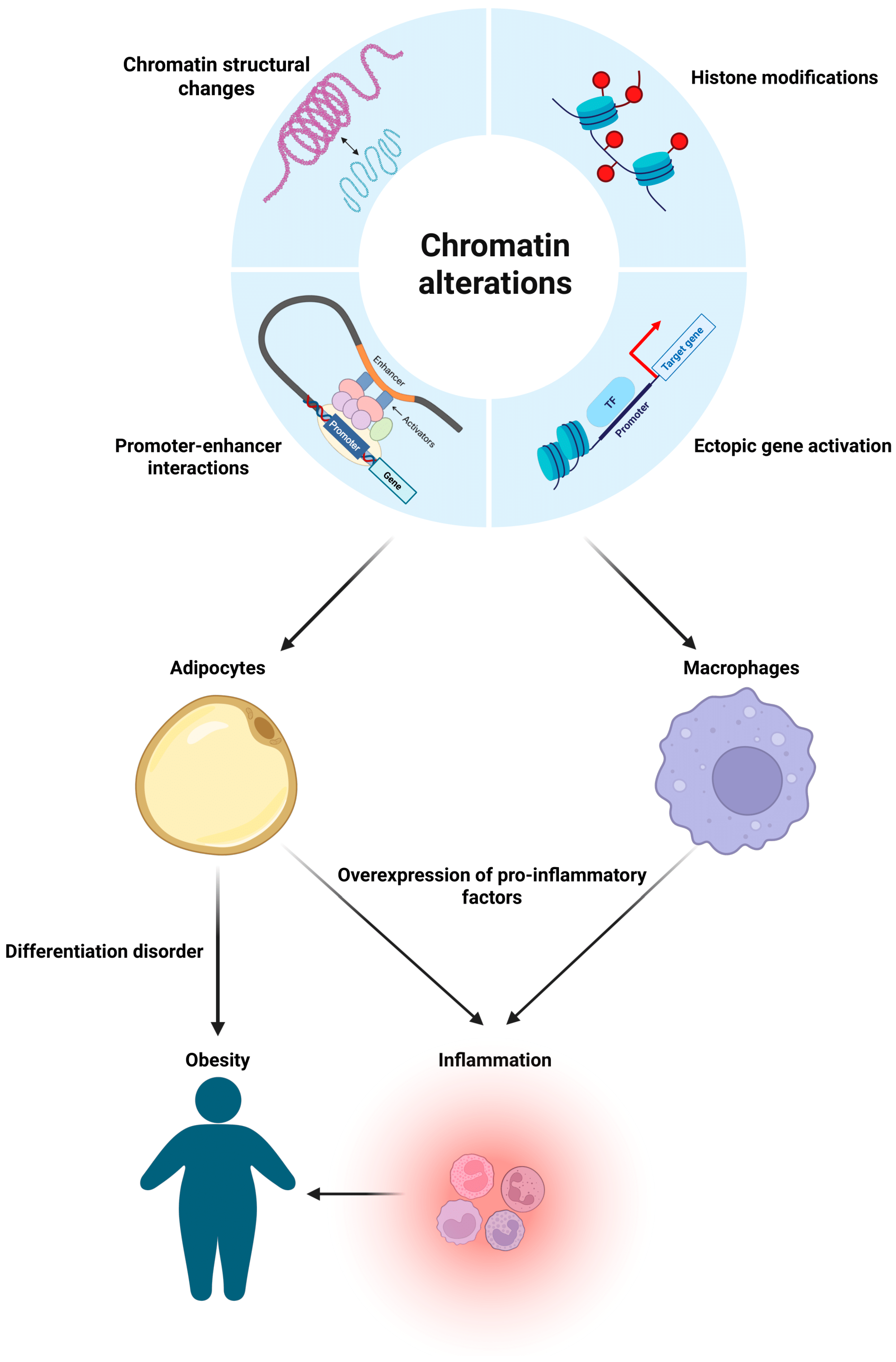
4. Chromatin Conformation and Obesity
Tissue-Specific Changes of Chromatin Organization in Obesity
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
Abbreviations
| 3′-UTR | 3′-untranslated region |
| 5′-UTR | 5′-untranslated region |
| AF | Abdominal fat |
| AMPK | AMP-activated protein kinase |
| ATAC-seq | Assay for transposase-accessible chromatin with high-throughput sequencing |
| BMI | Body mass index |
| ChIP | Chromatin immunoprecipitation |
| CpG | Cytosine-phosphate-guanine dinucleotide |
| DNA | Deoxyribonucleic acid |
| DNMT | DNA methyltransferase |
| GF | Gluteofemoral fat |
| HAD | HAT1-dependent Accessibility Domain |
| HDACi | Histone deacetylase inhibitors |
| HFD | High-fat diet |
| IL | Interleukin |
| LAD | Lamina-associated domain |
| lncRNA | Long non-coding RNA |
| LRI | Long-range chromosomal interactions |
| MBD | Methyl-CpG-binding domain |
| miRNA | MicroRNA |
| MSRE-qPCR | Methylation-sensitive restriction enzyme quantitative PCR |
| NAFLD | Non-alcoholic fatty liver disease |
| pCHi-C | Promoter capture Hi-C |
| PcG | Polycomb group proteins |
| RNA | Ribonucleic acid |
| RIP | RNA immunoprecipitation |
| RT-qPCR | Reverse transcription quantitative PCR |
| SAT | Subcutaneous adipose tissue |
| TAD | Topologically associating domain |
| TNF | Tumor necrosis factor |
| VAT | Visceral adipose tissue |
| WAT | White adipose tissue |
References
- Obesity and Overweight. Available online: https://www.who.int/news-room/fact-sheets/detail/obesity-and-overweight (accessed on 23 July 2025).
- Pakhare, M.; Anjankar, A. Critical Correlation Between Obesity and Cardiovascular Diseases and Recent Advancements in Obesity. Cureus 2024, 16, e51681. [Google Scholar] [CrossRef]
- Sivakumar, S.; Lama, D.; Rabhi, N. Childhood Obesity from the Genes to the Epigenome. Front. Endocrinol. 2024, 15, 1393250. [Google Scholar] [CrossRef]
- Mǎrginean, C.O.; Mǎrginean, C.; Meliţ, L.E. New Insights Regarding Genetic Aspects of Childhood Obesity: A Minireview. Front. Pediatr. 2018, 6, 271. [Google Scholar] [CrossRef] [PubMed]
- Furse, S.; Fernandez-Twinn, D.S.; Chiarugi, D.; Koulman, A.; Ozanne, S.E. Lipid Metabolism Is Dysregulated Before, During and After Pregnancy in a Mouse Model of Gestational Diabetes. Int. J. Mol. Sci. 2021, 22, 7452. [Google Scholar] [CrossRef] [PubMed]
- Lim, L.-L.; Chow, E.; Chan, J.C.N. Cardiorenal Diseases in Type 2 Diabetes Mellitus: Clinical Trials and Real-World Practice. Nat. Rev. Endocrinol. 2023, 19, 151–163. [Google Scholar] [CrossRef] [PubMed]
- Klaus, V.S.; Schriever, S.C.; Monroy Kuhn, J.M.; Peter, A.; Irmler, M.; Tokarz, J.; Prehn, C.; Kastenmüller, G.; Beckers, J.; Adamski, J.; et al. Correlation Guided Network Integration (CoNI) Reveals Novel Genes Affecting Hepatic Metabolism. Mol. Metab. 2021, 53, 101295. [Google Scholar] [CrossRef]
- He, F.; Berg, A.; Imamura Kawasawa, Y.; Bixler, E.O.; Fernandez-Mendoza, J.; Whitsel, E.A.; Liao, D. Association Between DNA Methylation in Obesity-Related Genes and Body Mass Index Percentile in Adolescents. Sci. Rep. 2019, 9, 2079. [Google Scholar] [CrossRef]
- Xu, X.; Su, S.; Barnes, V.A.; De Miguel, C.; Pollock, J.; Ownby, D.; Shi, H.; Zhu, H.; Snieder, H.; Wang, X. A Genome-Wide Methylation Study on Obesity. Epigenetics 2013, 8, 522–533. [Google Scholar] [CrossRef]
- Künzel, R.; Faust, H.; Bundalian, L.; Blüher, M.; Jasaszwili, M.; Kirstein, A.; Kobelt, A.; Körner, A.; Popp, D.; Wenzel, E.; et al. Detecting Monogenic Obesity: A Systematic Exome-Wide Workup of over 500 Individuals. Int. J. Obes. 2025, 49, 1400–1411. [Google Scholar] [CrossRef]
- Samblas, M.; Milagro, F.I.; Martínez, A. DNA Methylation Markers in Obesity, Metabolic Syndrome, and Weight Loss. Epigenetics 2019, 14, 421–444. [Google Scholar] [CrossRef]
- Avni, B.; Neiman, D.; Shaked, E.; Gal-Rosenberg, O.; Grisariu, S.; Kuzli, M.; Avni, I.; Fracchia, A.; Stepensky, P.; Zuckerman, T.; et al. Chronic Graft-Versus-Host Disease Detected by Tissue-Specific Cell-Free DNA Methylation Biomarkers. J. Clin. Investig. 2024, 134, e163541. [Google Scholar] [CrossRef]
- Tóth, D.M.; Szeri, F.; Ashaber, M.; Muazu, M.; Székvölgyi, L.; Arányi, T. Tissue-Specific Roles of De Novo DNA Methyltransferases. Epigenet. Chromatin 2025, 18, 5. [Google Scholar] [CrossRef]
- McAllan, L.; Baranasic, D.; Villicaña, S.; Brown, S.; Zhang, W.; Lehne, B.; Adamo, M.; Jenkinson, A.; Elkalaawy, M.; Mohammadi, B.; et al. Integrative Genomic Analyses in Adipocytes Implicate DNA Methylation in Human Obesity and Diabetes. Nat. Commun. 2023, 14, 2784. [Google Scholar] [CrossRef]
- Tonyan, Z.N.; Barbitoff, Y.A.; Nasykhova, Y.A.; Danilova, M.M.; Kozyulina, P.Y.; Mikhailova, A.A.; Bulgakova, O.L.; Vlasova, M.E.; Golovkin, N.V.; Glotov, A.S. Plasma MicroRNA Profiling in Type 2 Diabetes Mellitus: A Pilot Study. Int. J. Mol. Sci. 2023, 24, 17406. [Google Scholar] [CrossRef] [PubMed]
- Maixner, N.; Haim, Y.; Blüher, M.; Chalifa-Caspi, V.; Veksler-Lublinsky, I.; Makarenkov, N.; Yoel, U.; Bashan, N.; Liberty, I.F.; Kukeev, I.; et al. Visceral Adipose Tissue E2F1-MiRNA206/210 Pathway Associates with Type 2 Diabetes in Humans with Extreme Obesity. Cells 2022, 11, 3046. [Google Scholar] [CrossRef] [PubMed]
- Elemam, N.M.; Hasswan, H.; Aljaibeji, H.; Sulaiman, N. Circulating Soluble ACE2 and Upstream MicroRNA Expressions in Serum of Type 2 Diabetes Mellitus Patients. Int. J. Mol. Sci. 2021, 22, 5263. [Google Scholar] [CrossRef] [PubMed]
- Ji, C.; Guo, X. The Clinical Potential of Circulating MicroRNAs in Obesity. Nat. Rev. Endocrinol. 2019, 15, 731–743. [Google Scholar] [CrossRef]
- Metzinger-Le Meuth, V.; Metzinger, L. The Roles of MicroRNAs in Obesity: Emphasizing Links with Chronic Kidney Disease and Cardiovascular Disorders. Obesities 2023, 3, 243–252. [Google Scholar] [CrossRef]
- Kiran, S.; Kumar, V.; Kumar, S.; Price, R.L.; Singh, U.P. Adipocyte, Immune Cells, and MiRNA Crosstalk: A Novel Regulator of Metabolic Dysfunction and Obesity. Cells 2021, 10, 1004. [Google Scholar] [CrossRef]
- Roos, J.; Dahlhaus, M.; Funcke, J.-B.; Kustermann, M.; Strauss, G.; Halbgebauer, D.; Boldrin, E.; Holzmann, K.; Möller, P.; Trojanowski, B.M.; et al. MiR-146a Regulates Insulin Sensitivity via NPR3. Cell. Mol. Life Sci. 2021, 78, 2987–3003. [Google Scholar] [CrossRef]
- Latorre, J.; Ortega, F.J.; Liñares-Pose, L.; Moreno-Navarrete, J.M.; Lluch, A.; Comas, F.; Oliveras-Cañellas, N.; Ricart, W.; Höring, M.; Zhou, Y.; et al. Compounds That Modulate AMPK Activity and Hepatic Steatosis Impact the Biosynthesis of MicroRNAs Required to Maintain Lipid Homeostasis in Hepatocytes. eBioMedicine 2020, 53, 102697. [Google Scholar] [CrossRef]
- Zhang, Y.-L.; Liu, L.; Peymanfar, Y.; Anderson, P.; Xian, C.J. Roles of MicroRNAs in Osteogenesis or Adipogenesis Differentiation of Bone Marrow Stromal Progenitor Cells. Int. J. Mol. Sci. 2021, 22, 7210. [Google Scholar] [CrossRef]
- Handoko, L.; Xu, H.; Li, G.; Ngan, C.Y.; Chew, E.; Schnapp, M.; Lee, C.W.H.; Ye, C.; Ping, J.L.H.; Mulawadi, F.; et al. CTCF-Mediated Functional Chromatin Interactome in Pluripotent Cells. Nat. Genet. 2011, 43, 630–638. [Google Scholar] [CrossRef] [PubMed]
- Lieberman-Aiden, E.; Van Berkum, N.L.; Williams, L.; Imakaev, M.; Ragoczy, T.; Telling, A.; Amit, I.; Lajoie, B.R.; Sabo, P.J.; Dorschner, M.O.; et al. Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. Science 2009, 326, 289–293. [Google Scholar] [CrossRef]
- Hinte, L.C.; Castellano-Castillo, D.; Ghosh, A.; Melrose, K.; Gasser, E.; Noé, F.; Massier, L.; Dong, H.; Sun, W.; Hoffmann, A.; et al. Adipose Tissue Retains an Epigenetic Memory of Obesity After Weight Loss. Nature 2024, 636, 457–465. [Google Scholar] [CrossRef] [PubMed]
- Saeed, S.; la Cour Poulsen, L.; Visnovska, T.; Hoffmann, A.; Ghosh, A.; Wolfrum, C.; Rønningen, T.; Dahl, M.B.; Wang, J.; Cayir, A.; et al. Chromatin Landscape in Paired Human Visceral and Subcutaneous Adipose Tissue and Its Impact on Clinical Variables in Obesity. eBioMedicine 2025, 114, 105653. [Google Scholar] [CrossRef] [PubMed]
- Divoux, A.; Sandor, K.; Bojcsuk, D.; Talukder, A.; Li, X.; Balint, B.L.; Osborne, T.F.; Smith, S.R. Differential Open Chromatin Profile and Transcriptomic Signature Define Depot-Specific Human Subcutaneous Preadipocytes: Primary Outcomes. Clin. Epigenet. 2018, 10, 148. [Google Scholar] [CrossRef]
- Hata, M.; Andriessen, E.M.M.A.; Hata, M.; Diaz-Marin, R.; Fournier, F.; Crespo-Garcia, S.; Blot, G.; Juneau, R.; Pilon, F.; Dejda, A.; et al. Past History of Obesity Triggers Persistent Epigenetic Changes in Innate Immunity and Exacerbates Neuroinflammation. Science 2023, 379, 45–62. [Google Scholar] [CrossRef]
- Asenso, J.; Wang, L.; Du, Y.; Liu, Q.; Xu, B.; Guo, M.; Tang, D. Advances in Detection and Quantification of Methylcytosine and Its Derivatives. J. Sep. Sci. 2019, 42, 1105–1116. [Google Scholar] [CrossRef]
- Dor, Y.; Cedar, H. Principles of DNA Methylation and Their Implications for Biology and Medicine. Lancet 2018, 392, 777–786. [Google Scholar] [CrossRef]
- Chen, P.; Wang, Y.; Chen, F.; Zhou, B. Epigenetics in Obesity: Mechanisms and Advances in Therapies Based on Natural Products. Pharmacol. Res. Perspect. 2024, 12, e1171. [Google Scholar] [CrossRef]
- Younesian, S.; Mohammadi, M.H.; Younesian, O.; Momeny, M.; Ghaffari, S.H.; Bashash, D. DNA Methylation in Human Diseases. Heliyon 2024, 10, e32366. [Google Scholar] [CrossRef] [PubMed]
- Dunnet, M.J.; Ortega-Recalde, O.J.; Waters, S.A.; Weeks, R.J.; Morison, I.M.; Hore, T.A. Leukocyte-Specific DNA Methylation Biomarkers and Their Implication for Pathological Epigenetic Analysis. Epigenet. Commun. 2022, 2, 5. [Google Scholar] [CrossRef]
- Le Thuc, O.; García-Cáceres, C. Obesity-Induced Inflammation: Connecting the Periphery to the Brain. Nat. Metab. 2024, 6, 1237–1252. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Huang, P.; Huang, X.; Zhang, L.; Liu, L.; Xiang, W.; Liu, L.; He, X. Alterations of DNA Methylation Profile in Peripheral Blood of Children with Simple Obesity. Health Inf. Sci. Syst. 2024, 12, 26. [Google Scholar] [CrossRef]
- Koh, I.-U.; Choi, N.-H.; Lee, K.; Yu, H.-Y.; Yun, J.H.; Kong, J.-H.; Kim, H.J.; Lee, S.; Kim, S.C.; Kim, B.-J.; et al. Obesity Susceptible Novel DNA Methylation Marker on Regulatory Region of Inflammation Gene: Results from the Korea Epigenome Study (KES). BMJ Open Diabetes Res. Care 2020, 8, e001338. [Google Scholar] [CrossRef]
- Lima, R.S.; Mattos, R.T.; Medeiros, N.I.; Kattah, F.M.; Nascimento, J.R.S.; Menezes, C.A.; Rios-Santos, F.; Dutra, W.O.; Gomes, J.A.S.; Moreira, P.R. CXCL8 Expression and Methylation Are Correlated with Anthropometric and Metabolic Parameters in Childhood Obesity. Cytokine 2021, 143, 155538. [Google Scholar] [CrossRef]
- Czogała, W.; Czogała, M.; Strojny, W.; Wątor, G.; Wołkow, P.; Wójcik, M.; Bik Multanowski, M.; Tomasik, P.; Wędrychowicz, A.; Kowalczyk, W.; et al. Methylation and Expression of FTO and PLAG1 Genes in Childhood Obesity: Insight into Anthropometric Parameters and Glucose–Lipid Metabolism. Nutrients 2021, 13, 1683. [Google Scholar] [CrossRef]
- Nemtsova, M.V.; Zaletaev, D.V.; Bure, I.V.; Mikhaylenko, D.S.; Kuznetsova, E.B.; Alekseeva, E.A.; Beloukhova, M.I.; Deviatkin, A.A.; Lukashev, A.N.; Zamyatnin, A.A. Epigenetic Changes in the Pathogenesis of Rheumatoid Arthritis. Front. Genet. 2019, 10, 570. [Google Scholar] [CrossRef]
- Houseman, E.A.; Accomando, W.P.; Koestler, D.C.; Christensen, B.C.; Marsit, C.J.; Nelson, H.H.; Wiencke, J.K.; Kelsey, K.T. DNA Methylation Arrays as Surrogate Measures of Cell Mixture Distribution. BMC Bioinform. 2012, 13, 86. [Google Scholar] [CrossRef]
- Baca, P.; Barajas-Olmos, F.; Mirzaeicheshmeh, E.; Zerrweck, C.; Guilbert, L.; Sánchez, E.C.; Flores-Huacuja, M.; Villafán, R.; Martínez-Hernández, A.; García-Ortiz, H.; et al. DNA methylation and gene expression analysis in adipose tissue to identify new loci associated with T2D development in obesity. Nutr. Diabetes 2022, 12, 50. [Google Scholar] [CrossRef]
- Akbar, N.; Pinnick, K.E.; Paget, D.; Choudhury, R.P. Isolation and Characterization of Human Adipocyte-Derived Extracellular Vesicles Using Filtration and Ultracentrifugation. J. Vis. Exp. 2021, 170, e61979. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhang, Y.; Ma, L.; Bao, Q.; Liang, C.; Chu, M.; Guo, X.; Bao, P.; Yan, P. DNA Methylation Dynamics During Yak Adipocyte Differentiation. Int. J. Biol. Macromol. 2024, 261, 129715. [Google Scholar] [CrossRef] [PubMed]
- Keyhan, S.; Burke, E.; Schrott, R.; Huang, Z.; Grenier, C.; Price, T.; Raburn, D.; Corcoran, D.L.; Soubry, A.; Hoyo, C.; et al. Male Obesity Impacts DNA Methylation Reprogramming in Sperm. Clin. Epigenet. 2021, 13, 17. [Google Scholar] [CrossRef] [PubMed]
- Guay, S.-P.; Brisson, D.; Lamarche, B.; Biron, S.; Lescelleur, O.; Biertho, L.; Marceau, S.; Vohl, M.-C.; Gaudet, D.; Bouchard, L. ADRB3 Gene Promoter DNA Methylation in Blood and Visceral Adipose Tissue Is Associated with Metabolic Disturbances in Men. Epigenomics 2014, 6, 33–43. [Google Scholar] [CrossRef]
- Lebovitz, H.E.; Banerji, M.A. Point: Visceral Adiposity Is Causally Related to Insulin Resistance. Diabetes Care 2005, 28, 2322–2325. [Google Scholar] [CrossRef]
- Mezzacappa, C.; Wang, Z.; Lu, L.; Risch, H.; Taddei, T.; Yu, H. Detection of Hepatocellular Carcinoma Methylation Markers in Salivary DNA. Biosci. Rep. 2024, 44, BSR20232063. [Google Scholar] [CrossRef]
- Rapado-González, Ó.; Henrique, R.; Jerónimo, C. Salivary Epigenetic Biomarkers in Oral Cancer; IntechOpen: London, UK, 2025. [Google Scholar] [CrossRef]
- Rahbar, E.; Waits, C.M.K.; Kirby, E.H.; Miller, L.R.; Ainsworth, H.C.; Cui, T.; Sergeant, S.; Howard, T.D.; Langefeld, C.D.; Chilton, F.H. Allele-Specific Methylation in the FADS Genomic Region in DNA from Human Saliva, CD4+ Cells, and Total Leukocytes. Clin. Epigenet. 2018, 10, 46. [Google Scholar] [CrossRef]
- Murata, Y.; Fujii, A.; Kanata, S.; Fujikawa, S.; Ikegame, T.; Nakachi, Y.; Zhao, Z.; Jinde, S.; Kasai, K.; Bundo, M.; et al. Evaluation of the Usefulness of Saliva for DNA Methylation Analysis in Cohort Studies. Neuropsychopharmacol. Rep. 2019, 39, 301–305. [Google Scholar] [CrossRef]
- Langie, S.A.S.; Moisse, M.; Declerck, K.; Koppen, G.; Godderis, L.; Vanden Berghe, W.; Drury, S.; De Boever, P. Salivary DNA Methylation Profiling: Aspects to Consider for Biomarker Identification. Basic Clin. Pharmacol. Toxicol. 2017, 121, 93–101. [Google Scholar] [CrossRef]
- Rushing, A.; Sommer, E.C.; Zhao, S.; Po’e, E.K.; Barkin, S.L. Salivary Epigenetic Biomarkers as Predictors of Emerging Childhood Obesity. BMC Med. Genet. 2020, 21, 34. [Google Scholar] [CrossRef]
- Patti, M.E.; Butte, A.J.; Crunkhorn, S.; Cusi, K.; Berria, R.; Kashyap, S.; Miyazaki, Y.; Kohane, I.; Costello, M.; Saccone, R.; et al. Coordinated Reduction of Genes of Oxidative Metabolism in Humans with Insulin Resistance and Diabetes: Potential Role of PGC1 and NRF1. Proc. Natl. Acad. Sci. USA 2003, 100, 8466–8471. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Yang, J.; Luo, Y.; Ran, D.; Xia, R.; Zheng, Q.; Yao, P.; Wang, H. Nrf1 Reduces COX-2 Expression and Maintains Cellular Homeostasis After Cerebral Ischemia/Reperfusion By Targeting IL-6/TNF-α Protein Production. J. Neuroimmune Pharmacol. 2024, 19, 41. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.; Ren, S.; Hou, Y.; Zuo, Z.; Liu, S.; Liu, Z.; Fu, J.; Wang, H.; Yang, B.; Zhao, R.; et al. Single-Nucleus RNA-Sequencing Reveals NRF1/NFE2L1 as a Key Factor Determining the Thermogenesis and Cellular Heterogeneity and Dynamics of Brown Adipose Tissues in Mice. Redox Biol. 2023, 67, 102879. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Zhang, J.; Lei, H.; Meng, Y.; Cheng, H.; Zhao, Y.; Geng, G.; Mu, C.; Chen, L.; Liu, Q.; et al. NRF1-mediated Mitochondrial Biogenesis Antagonizes Innate Antiviral Immunity. EMBO J. 2023, 42, e113258. [Google Scholar] [CrossRef]
- Cantarero-Cuenca, A.; Gonzalez-Jimenez, A.; Martínez-Núñez, G.M.; Garrido-Sánchez, L.; Ranea, J.A.G.; Tinahones, F.J. Epigenetic Profiles in Blood and Adipose Tissue: Identifying Strong Correlations in Morbidly Obese and Non-Obese Patients. J. Mol. Med. 2024, 102, 1315–1325. [Google Scholar] [CrossRef]
- Heydarnezhad Asl, M.; Pasban Khelejani, F.; Bahojb Mahdavi, S.Z.; Emrahi, L.; Jebelli, A.; Mokhtarzadeh, A. The Various Regulatory Functions of Long Noncoding RNAs in Apoptosis, Cell Cycle, and Cellular Senescence. J. Cell. Biochem. 2022, 123, 995–1024. [Google Scholar] [CrossRef]
- Klinge, C.M. Non-Coding RNAs: Long Non-Coding RNAs and MicroRNAs in Endocrine-Related Cancers. Endocr. Relat. Cancer 2018, 25, R259–R282. [Google Scholar] [CrossRef]
- Bofill-De Ros, X.; Vang Ørom, U.A. Recent Progress in MiRNA Biogenesis and Decay. RNA Biol. 2024, 21, 36–43. [Google Scholar] [CrossRef]
- Makarova, J.; Turchinovich, A.; Shkurnikov, M.; Tonevitsky, A. Extracellular MiRNAs and Cell–Cell Communication: Problems and Prospects. Trends Biochem. Sci. 2021, 46, 640–651. [Google Scholar] [CrossRef]
- Brandao, B.B.; Lino, M.; Kahn, C.R. Extracellular MiRNAs as Mediators of Obesity-Associated Disease. J. Physiol. 2022, 600, 1155–1169. [Google Scholar] [CrossRef]
- Mattick, J.S.; Amaral, P.P.; Carninci, P.; Carpenter, S.; Chang, H.Y.; Chen, L.-L.; Chen, R.; Dean, C.; Dinger, M.E.; Fitzgerald, K.A.; et al. Long Non-Coding RNAs: Definitions, Functions, Challenges and Recommendations. Nat. Rev. Mol. Cell Biol. 2023, 24, 430–447. [Google Scholar] [CrossRef] [PubMed]
- Llères, D.; Imaizumi, Y.; Feil, R. Exploring Chromatin Structural Roles of Non-Coding RNAs at Imprinted Domains. Biochem. Soc. Trans. 2021, 49, 1867–1879. [Google Scholar] [CrossRef]
- Lu, X.; Ding, Y.; Bai, Y.; Li, J.; Zhang, G.; Wang, S.; Gao, W.; Xu, L.; Wang, H. Detection of Allosteric Effects of LncRNA Secondary Structures Altered by SNPs in Human Diseases. Front. Cell Dev. Biol. 2020, 8, 242. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Wang, L.; Li, H.; Han, X.; Chen, S.; Yang, B.; Hu, Z.; Zhu, H.; Cai, C.; Chen, J.; et al. Characterization of LncRNA Expression Profile and Identification of Novel LncRNA Biomarkers to Diagnose Coronary Artery Disease. Atherosclerosis 2018, 275, 359–367. [Google Scholar] [CrossRef] [PubMed]
- Daneshmoghadam, J.; Omidifar, A.; Akbari Dilmaghani, N.; Karimi, Z.; Emamgholipour, S.; Shanaki, M. The Gene Expression of Long Non-Coding RNAs (LncRNAs): MEG3 and H19 in Adipose Tissues from Obese Women and Its Association with Insulin Resistance and Obesity Indices. J. Clin. Lab. Anal. 2021, 35, e23741. [Google Scholar] [CrossRef]
- La Sala, L.; Crestani, M.; Garavelli, S.; de Candia, P.; Pontiroli, A.E. Does MicroRNA Perturbation Control the Mechanisms Linking Obesity and Diabetes? Implications for Cardiovascular Risk. Int. J. Mol. Sci. 2020, 22, 143. [Google Scholar] [CrossRef]
- Heo, Y.; Kim, H.; Lim, J.; Choi, S.S. Adipocyte Differentiation Between Obese and Lean Conditions Depends on Changes in MiRNA Expression. Sci. Rep. 2022, 12, 11543. [Google Scholar] [CrossRef]
- Catanzaro, G.; Filardi, T.; Sabato, C.; Vacca, A.; Migliaccio, S.; Morano, S.; Ferretti, E. Tissue and Circulating MicroRNAs as Biomarkers of Response to Obesity Treatment Strategies. J. Endocrinol. Investig. 2021, 44, 1159–1174. [Google Scholar] [CrossRef]
- Tekcan, E.; Kara, N.; Aydın, H.M.; Abur, Ü.; Abbaszadeh, M. Evaluation of the Promoter Methylation Status of Hypoxia Factor 3A and Interleukin-6 Genes and Expression Levels of Mir-130b and Mir-146b in Childhood Obesity. Rev. Assoc. Med. Bras. 2022, 68, 1276–1281. [Google Scholar] [CrossRef]
- Wilfinger, W.W.; Eghbalnia, H.R.; Mackey, K.; Miller, R.; Chomczynski, P. Whole Blood RNA Extraction Efficiency Contributes to Variability in RNA Sequencing Data Sets. PLoS ONE 2023, 18, e0291209. [Google Scholar] [CrossRef]
- Fehlmann, T.; Lehallier, B.; Schaum, N.; Hahn, O.; Kahraman, M.; Li, Y.; Grammes, N.; Geffers, L.; Backes, C.; Balling, R.; et al. Common Diseases Alter the Physiological Age-Related Blood MicroRNA Profile. Nat. Commun. 2020, 11, 5958. [Google Scholar] [CrossRef]
- Romero-Ruiz, A.; Pineda, B.; Ovelleiro, D.; Perdices-Lopez, C.; Torres, E.; Vazquez, M.J.; Guler, I.; Jiménez, Á.; Pineda, R.; Persano, M.; et al. Molecular Diagnosis of Polycystic Ovary Syndrome in Obese and Non-Obese Women by Targeted Plasma MiRNA Profiling. Eur. J. Endocrinol. 2021, 185, 637–652. [Google Scholar] [CrossRef]
- Ortega, F.J.; Mercader, J.M.; Catalán, V.; Moreno-Navarrete, J.M.; Pueyo, N.; Sabater, M.; Gómez-Ambrosi, J.; Anglada, R.; Fernández-Formoso, J.A.; Ricart, W.; et al. Targeting the Circulating MicroRNA Signature of Obesity. Clin. Chem. 2013, 59, 781–792. [Google Scholar] [CrossRef] [PubMed]
- Vonhögen, I.G.C.; Mohseni, Z.; Winkens, B.; Xiao, K.; Thum, T.; Calore, M.; da Costa Martins, P.A.; de Windt, L.J.; Spaanderman, M.E.A.; Ghossein-Doha, C. Circulating MiR-216a as a Biomarker of Metabolic Alterations and Obesity in Women. Noncoding RNA Res. 2020, 5, 144–152. [Google Scholar] [CrossRef] [PubMed]
- Mansego, M.L.; Garcia-Lacarte, M.; Milagro, F.I.; Marti, A.; Martinez, J.A. DNA Methylation of MiRNA Coding Sequences Putatively Associated with Childhood Obesity. Pediatr. Obes. 2017, 12, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Cao, D.; Liu, Z.; Li, Y.; Ouyang, S.; Wu, J. Identification of Novel Circulating MiRNAs Biomarkers for Healthy Obese and Lean Children. BMC Endocr. Disord. 2023, 23, 238. [Google Scholar] [CrossRef]
- Lin, H.; Tas, E.; Børsheim, E.; Mercer, K.E. Circulating MiRNA Signatures Associated with Insulin Resistance in Adolescents with Obesity. Diabetes Metab. Syndr. Obes. 2020, 13, 4929–4939. [Google Scholar] [CrossRef]
- Das, K.; Rao, L.V.M. The Role of MicroRNAs in Inflammation. Int. J. Mol. Sci. 2022, 23, 15479. [Google Scholar] [CrossRef]
- Engin, A.B.; Engin, A. Adipogenesis-Related MicroRNAs in Obesity. exRNA 2022, 4, 16. [Google Scholar] [CrossRef]
- Matveev, G.A.; Khromova, N.V.; Zasypkin, G.G.; Kononova, Y.A.; Vasilyeva, E.Y.; Babenko, A.Y.; Shlyakhto, E.V. Tissue and Circulating MicroRNAs 378 and 142 as Biomarkers of Obesity and Its Treatment Response. Int. J. Mol. Sci. 2023, 24, 13426. [Google Scholar] [CrossRef]
- Youssef, E.M.; Elfiky, A.M.; BanglySoliman; Abu-Shahba, N.; Elhefnawi, M.M. Expression Profiling and Analysis of Some MiRNAs in Subcutaneous White Adipose Tissue During Development of Obesity. Genes Nutr. 2020, 15, 8. [Google Scholar] [CrossRef] [PubMed]
- Wijesinghe, S.N.; Nicholson, T.; Tsintzas, K.; Jones, S.W. Involvements of Long Noncoding RNAs in Obesity-associated Inflammatory Diseases. Obes. Rev. 2021, 22, e13156. [Google Scholar] [CrossRef] [PubMed]
- Ghafouri-Fard, S.; Taheri, M. The Expression Profile and Role of Non-Coding RNAs in Obesity. Eur. J. Pharmacol. 2021, 892, 173809. [Google Scholar] [CrossRef] [PubMed]
- Tang, S.; Zhu, W.; Zheng, F.; Gui, W.; Zhang, W.; Lin, X.; Li, H. The Long Noncoding RNA Blnc1 Protects Against Diet-Induced Obesity by Promoting Mitochondrial Function in White Fat. Diabetes Metab. Syndr. Obes. 2020, 13, 1189–1201. [Google Scholar] [CrossRef]
- Xing, Y.; Zhang, T.; Wan, S.; Cheng, Y.; Zhou, S.; Sun, Y.; Zhang, H.; Yao, X.; Hua, Q.; Meng, X.; et al. LncRNA HEM2ATM Improves Obesity-Associated Adipose Tissues Meta-Inflammation and Insulin Resistance by Interacting with Heterogeneous Nuclear Ribonucleoprotein U. Clin. Immunol. 2023, 247, 109234. [Google Scholar] [CrossRef]
- Thunen, A.; La Placa, D.; Zhang, Z.; Shively, J.E. Role of LncRNA LIPE-AS1 in Adipogenesis. Adipocyte 2022, 11, 11–27. [Google Scholar] [CrossRef]
- Huang, X.; Chen, J.; Li, H.; Cai, Y.; Liu, L.; Dong, Q.; Li, Y.; Ren, Y.; Xiang, W.; He, X. LncRNA SNHG12 Suppresses Adipocyte Inflammation and Insulin Resistance by Regulating the HDAC9/Nrf2 Axis. FASEB J. 2024, 38, e23794. [Google Scholar] [CrossRef]
- Ling, C.; Rönn, T. Epigenetics in Human Obesity and Type 2 Diabetes. Cell Metab. 2019, 29, 1028–1044. [Google Scholar] [CrossRef]
- Dixon, J.R.; Selvaraj, S.; Yue, F.; Kim, A.; Li, Y.; Shen, Y.; Hu, M.; Liu, J.S.; Ren, B. Topological Domains in Mammalian Genomes Identified by Analysis of Chromatin Interactions. Nature 2012, 485, 376–380. [Google Scholar] [CrossRef]
- Lupiáñez, D.G.; Kraft, K.; Heinrich, V.; Krawitz, P.; Brancati, F.; Klopocki, E.; Horn, D.; Kayserili, H.; Opitz, J.M.; Laxova, R.; et al. Disruptions of Topological Chromatin Domains Cause Pathogenic Rewiring of Gene-Enhancer Interactions. Cell 2015, 161, 1012–1025. [Google Scholar] [CrossRef] [PubMed]
- Crispatzu, G.; Rehimi, R.; Pachano, T.; Bleckwehl, T.; Cruz-Molina, S.; Xiao, C.; Mahabir, E.; Bazzi, H.; Rada-Iglesias, A. The Chromatin, Topological and Regulatory Properties of Pluripotency-Associated Poised Enhancers Are Conserved In Vivo. Nat. Commun. 2021, 12, 4344. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, C.; Nissen, I.; Vincent, C.A.; Hägglund, A.-C.; Hörnblad, A.; Remeseiro, S. Rewiring of the Promoter-Enhancer Interactome and Regulatory Landscape in Glioblastoma Orchestrates Gene Expression Underlying Neurogliomal Synaptic Communication. Nat. Commun. 2023, 14, 6446. [Google Scholar] [CrossRef] [PubMed]
- Garske, K.M.; Comenho, C.; Pan, D.Z.; Alvarez, M.; Mohlke, K.; Laakso, M.; Pietiläinen, K.H.; Pajukanta, P. Long-Range Chromosomal Interactions Increase and Mark Repressed Gene Expression During Adipogenesis. Epigenetics 2022, 17, 1849–1862. [Google Scholar] [CrossRef]
- Ouchi, N.; Parker, J.L.; Lugus, J.J.; Walsh, K. Adipokines in Inflammation and Metabolic Disease. Nat. Rev. Immunol. 2011, 11, 85–97. [Google Scholar] [CrossRef]
- Pyrina, I.; Chung, K.-J.; Michailidou, Z.; Koutsilieris, M.; Chavakis, T.; Chatzigeorgiou, A. Fate of Adipose Progenitor Cells in Obesity-Related Chronic Inflammation. Front. Cell Dev. Biol. 2020, 8, 644. [Google Scholar] [CrossRef]
- Garske, K.M.; Kar, A.; Comenho, C.; Balliu, B.; Pan, D.Z.; Bhagat, Y.V.; Rosenberg, G.; Koka, A.; Das, S.S.; Miao, Z.; et al. Increased Body Mass Index Is Linked to Systemic Inflammation Through Altered Chromatin Co-Accessibility in Human Preadipocytes. Nat. Commun. 2023, 14, 4214. [Google Scholar] [CrossRef]
- Qin, Y.; Grimm, S.A.; Roberts, J.D.; Chrysovergis, K.; Wade, P.A. Alterations in Promoter Interaction Landscape and Transcriptional Network Underlying Metabolic Adaptation to Diet. Nat. Commun. 2020, 11, 962. [Google Scholar] [CrossRef]
- Tillander, V.; Alexson, S.E.H.; Cohen, D.E. Deactivating Fatty Acids: Acyl-CoA Thioesterase-Mediated Control of Lipid Metabolism. Trends Endocrinol. Metab. 2017, 28, 473–484. [Google Scholar] [CrossRef]
- Divoux, A.; Sandor, K.; Bojcsuk, D.; Yi, F.; Hopf, M.E.; Smith, J.S.; Balint, B.L.; Osborne, T.F.; Smith, S.R. Fat Distribution in Women Is Associated with Depot-Specific Transcriptomic Signatures and Chromatin Structure. J. Endocr. Soc. 2020, 4, bvaa042. [Google Scholar] [CrossRef]
- Jin, L.; Wang, D.; Zhang, J.; Liu, P.; Wang, Y.; Lin, Y.; Liu, C.; Han, Z.; Long, K.; Li, D.; et al. Dynamic Chromatin Architecture of the Porcine Adipose Tissues with Weight Gain and Loss. Nat. Commun. 2023, 14, 3457. [Google Scholar] [CrossRef]
| Sample, n | BMI, Percentile/ kg/m2 | Detection Method | Material | Biomarkers | Source |
|---|---|---|---|---|---|
| 15 (Prepubertal children) | ≥95 | RT-qPCR | Peripheral blood | miRNA-130b, miRNA-146b | [72] |
| 60 Women | ≥30 kg/m2 | RT-qPCR | Plasm | miRNA-216a | [77] |
| 51 Adults | >30 kg/m2 | RT-qPCR | Subcutaneous adipose tissue, plasm | miRNA-378a-3p, miRNA-142-3p | [83] |
| Wister rats | - | RT-qPCR | Subcutaneous adipose tissue | miRNA-133a, let-7-5p, miRNA-107-5p, miRNA-130a-5p, miRNA-30a-5p | [84] |
| C57BL/6 mice; 3T3-L1 murine preadipocyte culture | - | ChiP, RIP, RT-qPCR | Subcutaneous adipose tissue | Blnc1 | [87] |
| HEM2ATM+/+ mice | - | RT-qPCR | Subcutaneous adipose tissue | Hem2atm | [88] |
| C57BL/6 mice, OP9 cell culture (CRL-2749) | - | RT-qPCR | Subcutaneous adipose tissue, liver | Lipe-as1 | [89] |
| RAW264.7 mice, 3T3-L1 cell culture | - | RT-qPCR | Plasm, liver, adipose tissue, biopsy of the adnexal tissue | Snhg12 | [90] |
| 12 Children | ≥97 | HumanMethylation450K BeadChip | Peripheral blood leukocytes | miRNA-1203, miRNA-412, miRNA-216A | [78] |
| 15 Children | >32 | Agilent Human miRNA Array V19.0, RT-qPCR | Plasm | miRNA-15b-5p, miRNA-223-3p | [79] |
| LncRNA | Localization | Role in Obesity | Mechanism of Action | Source |
|---|---|---|---|---|
| Blnc1 | Brown/white adipose tissue, liver | Regulation of adipogenesis Reduction in insulin resistance Reduction in adipose tissue fibrosis | Activation of PPARγ/C/EBPα Binding to hnRNPA1 → ↑ PGC1β Inhibition of TGF-β | [87,89] |
| Hem2atm | Adipose tissue macrophages (M2) | Suppression of inflammation Improvement of insulin sensitivity | Binding to hnRNP U → ↓ TNF-α/IL-6 Activation of the Nrf2 pathway | [88] |
| Lipe-as1 mlas-V | Adipose tissue, liver | Regulation of lipolysis Control of adipocyte differentiation | Suppression of LIPE Regulation of PLIN4 and TP53 | [89] |
| Snhg12 | Adipose tissue, liver | Anti-inflammatory effect Macrophage polarization toward M2 phenotype | Binding hnRNPA1 → ↓ HDAC9 Activation of Nrf2 | [90] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nikolaeva, A.F.; Nemtsova, M.V.; Pustovalova, A.V.; Sigin, V.O. When Genes Wear Marks: Epigenomic Modulation in the Development and Progression of Obesity. Int. J. Mol. Sci. 2025, 26, 8067. https://doi.org/10.3390/ijms26168067
Nikolaeva AF, Nemtsova MV, Pustovalova AV, Sigin VO. When Genes Wear Marks: Epigenomic Modulation in the Development and Progression of Obesity. International Journal of Molecular Sciences. 2025; 26(16):8067. https://doi.org/10.3390/ijms26168067
Chicago/Turabian StyleNikolaeva, Alexandra F., Marina V. Nemtsova, Anna V. Pustovalova, and Vladimir O. Sigin. 2025. "When Genes Wear Marks: Epigenomic Modulation in the Development and Progression of Obesity" International Journal of Molecular Sciences 26, no. 16: 8067. https://doi.org/10.3390/ijms26168067
APA StyleNikolaeva, A. F., Nemtsova, M. V., Pustovalova, A. V., & Sigin, V. O. (2025). When Genes Wear Marks: Epigenomic Modulation in the Development and Progression of Obesity. International Journal of Molecular Sciences, 26(16), 8067. https://doi.org/10.3390/ijms26168067






