Classification of Heterotic Groups and Prediction of Heterosis in Sorghum Based on Whole-Genome Resequencing
Abstract
1. Introduction
2. Results
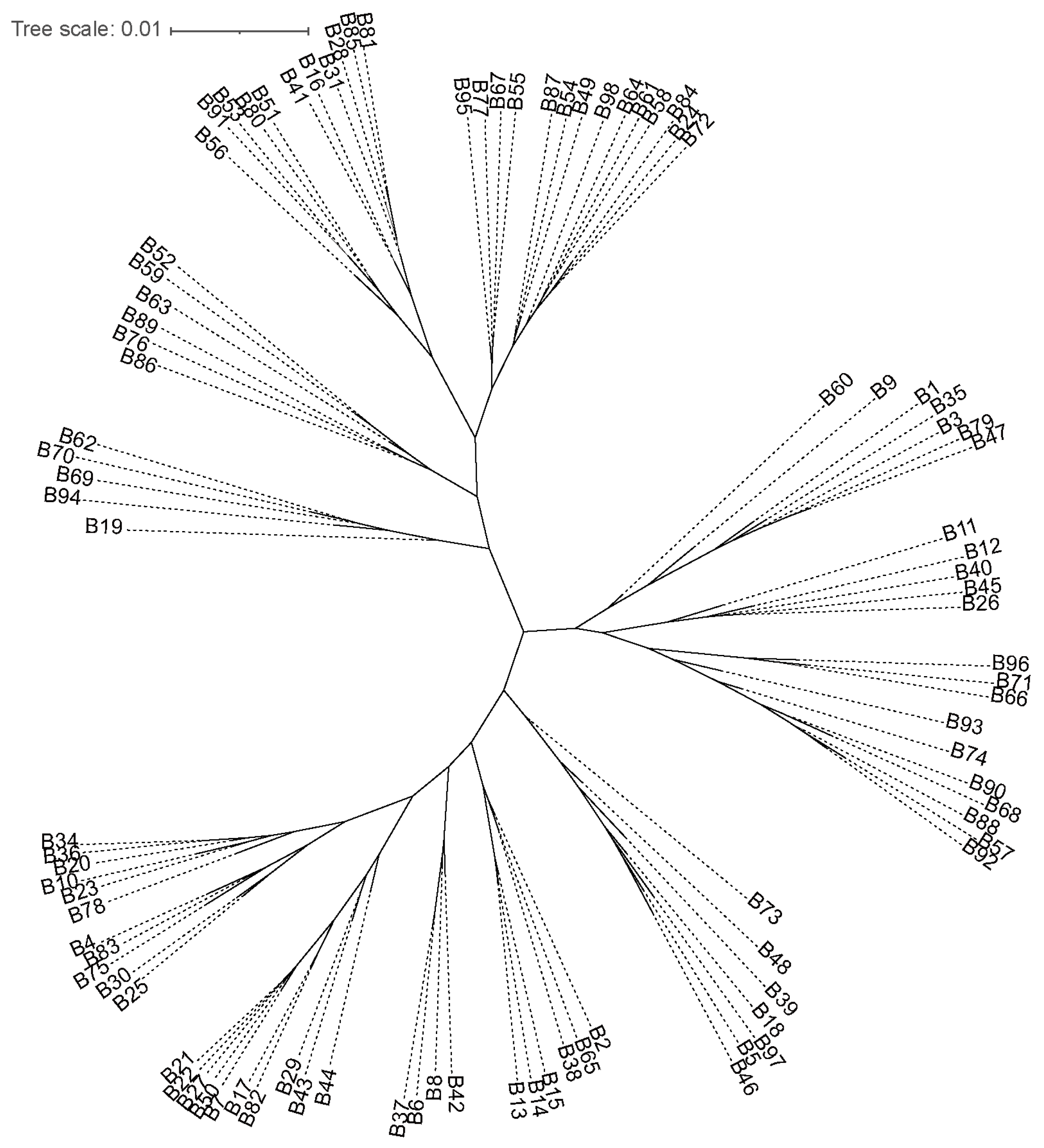
2.1. Genetic Diversity
2.2. Heterosis Analysis
2.2.1. Heterosis Performance for Various Traits
2.2.2. Analysis of Strong Heterosis Combinations
2.2.3. Genetic Basis of Heterosis
2.3. Combining Ability
2.4. Heterosis Prediction Effects
3. Discussion
3.1. Application of Whole-Genome Resequencing in the Division of Heterosis Groups
3.2. Comparison of Heterosis Prediction Methods
4. Materials and Methods
4.1. Experimental Materials
4.2. Field Trial Design
4.3. Hybridization Experiment Design
4.4. Trait Measurement
4.5. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BIY | Fresh biomass yield |
| DM | Days to maturity |
| DF | Days from emergence to flowering |
| GCA | General combining ability |
| GY | Grain yield |
| HP | High-parent |
| LD | Linkage disequilibrium |
| MP | Mid-parent |
| PGD | Phenotypic genetic distance |
| PH | Plant height |
| PL | Panicle length |
| PWT | Panicle weight |
| QTL | Quantitative trait loci |
| SCA | Specific combining ability |
| SD | Stem diameter |
| SNP | Single-nucleotide polymorphism |
| TKW | Thousand-kernel weight |
| TL | Tillering |
| WGRS | Whole-genome resequencing |
References
- Boatwright, J.L.; Sapkota, S.; Jin, H.; Schnable, J.C.; Brenton, Z.; Boyles, R.; Kresovich, S. Sorghum Association Panel whole-genome sequencing establishes cornerstone resource for dissecting genomic diversity. Plant J. 2022, 111, 888–904. [Google Scholar] [CrossRef]
- da Silva, K.J.; Pastina, M.M.; Guimarães, C.T.; Magalhães, J.V.; Pimentel, L.D.; Schaffert, R.E.; de Oliveira Pinto, M.; de Souza, V.F.; da Costa Bernardino, K.; da Silva, M.J. Genetic diversity and heterotic grouping of sorghum lines using SNP markers. Sci. Agric. 2021, 78, 9. [Google Scholar] [CrossRef]
- Maulana, F.; Perumal, R.; Serba, D.D.; Tesso, T. Genomic prediction of hybrid performance in grain sorghum (Sorghum bicolor L.). Front. Plant Sci. 2023, 14, 1139896. [Google Scholar] [CrossRef]
- Liu, F.; Wodajo, B.; Zhao, K.; Tang, S.; Xie, Q.; Xie, P. Unravelling sorghum functional genomics and molecular breeding: Past achievements and future prospects. J. Genet. Genom. 2025, 52, 719–732. [Google Scholar] [CrossRef]
- Enyew, M.; Feyissa, T.; Carlsson, A.S.; Tesfaye, K.; Hammenhag, C.; Seyoum, A.; Geleta, M. Genome-wide analyses using multi-locus models revealed marker-trait associations for major agronomic traits in Sorghum bicolor. Front. Plant Sci. 2022, 13, 999692. [Google Scholar] [CrossRef]
- Sapkota, S.; Boatwright, J.L.; Kumar, N.; Myers, M.; Cox, A.; Ackerman, A.; Caughman, W.; Brenton, Z.W.; Boyles, R.E.; Kresovich, S. Genomic prediction of hybrid performance for agronomic traits in sorghum. G3 2023, 13, jkac311. [Google Scholar] [CrossRef] [PubMed]
- Faye, J.M.; Maina, F.; Akata, E.A.; Sine, B.; Diatta, C.; Mamadou, A.; Marla, S.; Bouchet, S.; Teme, N.; Rami, J.-F.; et al. A genomics resource for genetics, physiology, and breeding of West African sorghum. Plant Genome 2021, 14, e20075. [Google Scholar] [CrossRef] [PubMed]
- Baggett, J.P.; Tillett, R.L.; Cooper, E.A.; Yerka, M.K. De novo identification and targeted sequencing of SSRs efficiently fingerprints Sorghum bicolor sub-population identity. PLoS ONE 2021, 16, e0248213. [Google Scholar] [CrossRef]
- Zhang, Y.; Chen, J.; Gao, Z.; Wang, H.; Liang, D.; Guo, Q.; Zhang, X.; Fan, X.; Wu, Y.; Liu, Q. Identification of heterosis and combining ability in the hybrids of male sterile and restorer sorghum [Sorghum bicolor (L.) Moench] lines. PLoS ONE 2024, 19, e0296416. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, J.M.O.; Klein, P.E.; Crossa, J.; Pacheco, A.; Perez-Rodriguez, P.; Ramasamy, P.; Klein, R.; Rooney, W.L. Assessing combining abilities, genomic data, and genotype × environment interactions to predict hybrid grain sorghum performance. Plant Genome 2021, 14, e20127. [Google Scholar] [CrossRef]
- Thakur, N.R.; Gorthy, S.; Vemula, A.; Odeny, D.A.; Ruperao, P.; Sargar, P.R.; Mehtre, S.P.; Kalpande, H.V.; Habyarimana, E. Genome-wide association study and expression of candidate genes for Fe and Zn concentration in sorghum grains. Sci. Rep. 2024, 14, 12729. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Tu, W.; Jin, P.; Liu, Y.; Du, J.; Zheng, J.; Wang, Y.-H.; Li, J. Genome-wide association study of plant color in Sorghum bicolor. Front. Plant Sci. 2024, 15, 1320844. [Google Scholar] [CrossRef] [PubMed]
- Enyew, M.; Geleta, M.; Tesfaye, K.; Seyoum, A.; Feyissa, T.; Alemu, A.; Hammenhag, C.; Carlsson, A.S. Genome-wide association study and genomic prediction of root system architecture traits in Sorghum (Sorghum bicolor (L.) Moench) at the seedling stage. BMC Plant Biol. 2025, 25, 69. [Google Scholar] [CrossRef]
- Lee, Y.-J.; Yang, B.; Kim, W.J.; Kim, J.; Kwon, S.-J.; Kim, J.H.; Ahn, J.-W.; Kim, S.H.; Rha, E.-S.; Ha, B.-K.; et al. Genome-wide association study (GWAS) of the agronomic traits and phenolic content in sorghum (Sorghum bicolor L.) genotypes. Agronomy 2023, 13, 1449. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, Y.; Zhao, R.; Wang, W.; Lü, P.; Zheng, D.; Zhou, Z. Analysis of molecular genetic diversity and division of heterosis groups in sorghum. Mol. Plant Breed. 2025, 23, 3267–3274. [Google Scholar]
- Casa, A.M.; Pressoir, G.; Brown, P.J.; Mitchell, S.E.; Rooney, W.L.; Tuinstra, M.R.; Franks, C.D.; Kresovich, S. Community resources and strategies for association mapping in sorghum. Crop Sci. 2008, 48, 30–40. [Google Scholar] [CrossRef]
- Brown, P.J.; Rooney, W.L.; Franks, C.; Kresovich, S. Efficient mapping of plant height quantitative trait loci in a sorghum association population with introgressed dwarfing genes. Genetics 2008, 180, 629–637. [Google Scholar] [CrossRef]
- Reddy, R.N.; Madhusudhana, R.; Mohan, S.M.; Chakravarthi, D.V.N.; Seetharama, N. Characterization, development and mapping of unigene-derived microsatellite markers in sorghum [Sorghum bicolor (L.) Moench]. Mol. Breed. 2011, 29, 543–564. [Google Scholar] [CrossRef]
- Rooney, W.L.; Blumenthal, J.; Bean, B.; Mullet, J.E. Designing sorghum as a dedicated bioenergy feedstock. Biofuels Bioprod. Biorefin. 2007, 1, 147–157. [Google Scholar] [CrossRef]
- Mullet, J.; Morishige, D.; McCormick, R.; Truong, S.; Hilley, J.; McKinley, B.; Anderson, R.; Olson, S.N.; Rooney, W. Energy sorghum—a genetic model for the design of C4 grass bioenergy crops. J. Exp. Bot. 2014, 65, 3479–3489. [Google Scholar] [CrossRef]
- Sage, R.F.; Sage, T.L.; Kocacinar, F. Photorespiration and the evolution of C4 photosynthesis. Annu. Rev. Plant Biol. 2012, 63, 19–47. [Google Scholar] [CrossRef] [PubMed]
- Bernardo, R. Relationship between single-cross performance and molecular marker heterozygosity. Theor. Appl. Genet. 1992, 83, 628–634. [Google Scholar] [CrossRef] [PubMed]
- Lippman, Z.B.; Zamir, D. Heterosis: Revisiting the magic. Trends Genet. 2007, 23, 60–66. [Google Scholar] [CrossRef]
- Zhao, Y.; Mette, M.F.; Gowda, M.; Longin, C.F.H.; Reif, J.C. Bridging the gap between marker-assisted and genomic selection of heading time and plant height in hybrid wheat. Heredity 2014, 112, 638–645. [Google Scholar] [CrossRef]
- Flint-Garcia, S.A.; Buckler, E.S.; Tiffin, P.; Ersoz, E.; Springer, N.M. Heterosis is prevalent for multiple traits in diverse maize germplasm. PLoS ONE 2009, 4, e7433. [Google Scholar] [CrossRef]
- Conley, A.B.; Rishishwar, L.; Ahmad, M.; Sharma, S.; Norris, E.T.; Jordan, I.K.; Mariño-Ramírez, L. Rye: Genetic ancestry inference at biobank scale. Nucleic Acids Res. 2023, 51, e44. [Google Scholar] [CrossRef]
- Reif, J.C.; Melchinger, A.E.; Frisch, M. Genetical and mathematical properties of similarity and dissimilarity coefficients applied in plant breeding and seed bank management. Crop Sci. 2025, 45, 1–7. [Google Scholar] [CrossRef]
- Schrag, T.A.; Möhring, J.; Maurer, H.P.; Dhillon, B.S.; Melchinger, A.E.; Piepho, H.-P.; Sørensen, A.P.; Frisch, M. Molecular marker-based prediction of hybrid performance in maize using unbalanced data from multiple experiments with factorial crosses. Theor. Appl. Genet. 2008, 118, 741–751. [Google Scholar] [CrossRef]
- Xu, S.; Zhu, D.; Zhang, Q. Predicting hybrid performance in rice using genomic best linear unbiased prediction. Proc. Natl. Acad. Sci. USA 2014, 111, 12456–12461. [Google Scholar] [CrossRef] [PubMed]
- Davey, J.W.; Hohenlohe, P.A.; Etter, P.D.; Boone, J.Q.; Catchen, J.M.; Blaxter, M.L. Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat. Rev. Genet. 2011, 12, 499–510. [Google Scholar] [CrossRef]
- Crossa, J.; Perez, P.; Hickey, J.; Burgueno, J.; Ornella, L.; Ceron-Rojas, J.; Zhang, X.; Dreisigacker, S.; Babu, R.; Li, Y.; et al. Genomic prediction in CIMMYT maize and wheat breeding programs. Heredity 2014, 112, 48–60. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Kurata, N.; Wei, X.; Wang, Z.X.; Wang, A.; Zhao, Q.; Zhao, Y.; Liu, K.; Lu, H.; Li, W.; et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 2012, 490, 497–501. [Google Scholar] [CrossRef]
- Mace, E.S.; Tai, S.; Gilding, E.K.; Li, Y.; Prentis, P.J.; Bian, L.; Campbell, B.C.; Hu, W.; Innes, D.J.; Han, X.; et al. Whole-genome sequencing reveals untapped genetic potential in Africa’s indigenous cereal crop sorghum. Nat. Commun. 2013, 4, 2320. [Google Scholar] [CrossRef]
- Lin, T.; Zhu, G.; Zhang, J.; Xu, X.; Yu, Q.; Zheng, Z.; Zhang, Z.; Lun, Y.; Li, S.; Wang, X.; et al. Genomic analyses provide insights into the history of tomato breeding. Nat. Genet. 2014, 46, 1220–1226. [Google Scholar] [CrossRef]
- Li, Y.; Ruperao, P.; Batley, J.; Edwards, D.; Khan, T.; Colmer, T.D.; Pang, J.; Siddique, K.H.M.; Sutton, T. Investigating drought tolerance in chickpea using genome-wide association mapping and genomic selection based on whole-genome resequencing data. Front. Plant Sci. 2018, 9, 190. [Google Scholar] [CrossRef]
- Raj, A.; Stephens, M.; Pritchard, J.K. fastSTRUCTURE: Variational inference of population structure in large SNP data sets. Genetics 2014, 197, 573–589. [Google Scholar] [CrossRef] [PubMed]
- Price, A.L.; Patterson, N.J.; Plenge, R.M.; Weinblatt, M.E.; Shadick, N.A.; Reich, D. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 2006, 38, 904–909. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.-H.; Guo, H.; Wang, X.; Kim, C.; Paterson, A.H. SNPhylo: A pipeline to construct a phylogenetic tree from huge SNP data. BMC Genom. 2014, 15, 162. [Google Scholar] [CrossRef]
- Hallauer, A.R.; Carena, M.J.; Filho, J.B.M. Quantitative Genetics in Maize Breeding; Springer: New York, NY, USA, 1988. [Google Scholar]
- Guo, M.; Liu, Q.; Yu, H.; Zhou, T.; Zou, J.; Zhang, H.; Bian, M.; Lui, X. Characterization of alkali stress-responsive genes of the CIPK family in sweet sorghum [Sorghum bicolor (L.) Moench]. Crop Sci. 2015, 55, 1254–1263. [Google Scholar] [CrossRef]
- Ropokis, A.; Ntatsi, G.; Kittas, C.; Katsoulas, N.; Savvas, D. Impact of cultivar and grafting on nutrient and water uptake by sweet pepper (Capsicum annuum L.) grown hydroponically under Mediterranean climatic conditions. Front. Plant Sci. 2018, 9, 1244. [Google Scholar] [CrossRef] [PubMed]
- Guerrero, C.G.; Robles, M.A.G.; Ortega, J.G.L.; Castillo, I.O.; Vázquez, C.V.; Carrillo, M.G.; Resendez, A.M.; Torres, A.G. Combining ability and heterosis in corn breeding lines to forage and grain. Am. J. Plant Sci. 2014, 5, 845–856. [Google Scholar] [CrossRef]
- Griffing, B. Concept of general and specific combining ability in relation to diallel crossing systems. Aust. J. Biol. Sci. 1956, 9, 463–493. [Google Scholar] [CrossRef]
- Goudet, J. HIERFSTAT, a package for R to compute and test hierarchical F-statistics. Mol. Ecol. Notes 2025, 5, 184–186. [Google Scholar] [CrossRef]
| Genotypes | Parent | GCA | SCA | PWT | Heterosis Index |
|---|---|---|---|---|---|
| 157 × 307A | 157 | 22.24 | 53.02 | 157.5 | 0.94 |
| 307A | 13.08 | ||||
| 157 × 521A | 157 | 22.24 | 48.87 | 150 | 0.84 |
| 521A | 17.83 | ||||
| 3618 × 170A | 3618 | 16.78 | 36.62 | 137.5 | 0.69 |
| 170A | −8.91 | ||||
| 157 × I15A | 157 | 22.24 | 6.46 | 137.5 | 0.69 |
| I15A | 1.55 | ||||
| 3618 × 521A | 3618 | 16.78 | 35.76 | 136.5 | 0.68 |
| 521A | 17.83 | ||||
| 157 × 170A | 157 | 22.24 | 35.37 | 136.5 | 0.68 |
| 170A | −8.91 | ||||
| E8 × 521A | E8 | −4.47 | 28.93 | 130 | 0.60 |
| 521A | 17.83 | ||||
| 3618 × 428A | 3618 | 16.78 | 25.03 | 126 | 0.55 |
| 428A | 5.47 | ||||
| 307fu × 428A | 307fu | 3.86 | 24.41 | 125 | 0.54 |
| 428A | 5.47 | ||||
| 3618 × I15A | 3618 | 16.78 | 23.67 | 124 | 0.52 |
| I15A | 1.55 | ||||
| 124fu × JinchangzaoA | 124fu | −5.17 | −17.49 | 81.5 | 0.00 |
| JinchangzaoA | −4.79 | ||||
| JinR7 × 428A | JinR7 | −21.09 | −27.82 | 73 | −0.10 |
| 428A | 5.47 | ||||
| 14T22 × 4190A | 14T22 | −9.41 | −25.75 | 72 | −0.12 |
| 4190A | −16.78 | ||||
| 14T22 × 170A | 14T22 | −9.41 | −29.41 | 70.5 | −0.13 |
| 170A | −8.91 | ||||
| JinR7 × JinchangzaoA | JinR7 | −21.09 | −30.24 | 69.5 | −0.15 |
| JinchangzaoA | −4.79 | ||||
| JinR7 × 4190A | JinR7 | −21.09 | −32.24 | 67 | −0.18 |
| 4190A | −16.78 | ||||
| E8 × 170A | E8 | −4.47 | −38.24 | 60.4 | −0.26 |
| 170A | −8.91 | ||||
| JinR7 × 170A | JinR7 | −21.09 | −40.05 | 60 | −0.26 |
| 170A | −8.91 | ||||
| E8 × 4190A | E8 | −4.47 | −43.56 | 54 | −0.34 |
| 4190A | −16.78 | ||||
| JinR7 × QL33A | JinR7 | −21.09 | −45.7 | 49.5 | −0.39 |
| QL33A | −7.46 |
| Variance | Agronomic Traits 1 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| GY | PWT | TKW | PL | PH | SD | DF | DM | TL | BIY | |
| General coordination variance% | 44.23 | 53.67 | 70.88 | 67.19 | 81.79 | 75.02 | 10.13 | 12.23 | 74.63 | 70.45 |
| Variance of special coordination force% | 55.77 | 46.33 | 29.12 | 32.81 | 18.21 | 24.98 | 89.87 | 87.77 | 25.37 | 29.55 |
| Generalized heritability% | 75.22 | 99.38 | 99.05 | 61.55 | 81.69 | 58.21 | 100 | 99.99 | 48.87 | 98.73 |
| Narrow heritability% | 33.27 | 53.33 | 70.21 | 41.35 | 66.81 | 43.67 | 10.13 | 12.23 | 25.33 | 69.55 |
| Parent | Agronomic Traits 1 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| GY | PWT | TKW | PL | PH | SD | DF | DM | TL | BIY | Comprehensive GCA | |
| 428A | 3.73 | 5.47 | −0.9 | −0.85 | 4.29 | 2.51 | 3.74 | 0.18 | −0.13 | 3.9 | 2.4 |
| 521A | 4.98 | 17.83 | 10.52 | −4.05 | 12.81 | 0.6 | 2.78 | −0.11 | −0.2 | 7.71 | 7.34 |
| 170A | −4.26 | −8.91 | −9.83 | −5.06 | −13.31 | 2.3 | 4.86 | −1.32 | 0.67 | −15.51 | −7.83 |
| I15A | 2 | 1.55 | 6.23 | 3.57 | 2.38 | 2.88 | −5.53 | −0.29 | −0.79 | 5.81 | 3.15 |
| 307A | 7.48 | 13.08 | 7.84 | 0.88 | 1.46 | −2.51 | −6.01 | 0.36 | −0.18 | 16.4 | 6.11 |
| JinchangzaoA | −1.73 | −4.79 | 3.37 | 8.15 | 3.12 | 0.94 | −2.98 | 0.36 | −0.1 | 12.69 | 3.2 |
| 4190A | −8.59 | −16.78 | −13.97 | 1.6 | −12.51 | −1.04 | 2.46 | 0.46 | 0.31 | −14.41 | −9.25 |
| QL33A | −3.61 | −7.46 | −3.25 | −4.24 | 1.76 | −5.69 | 0.7 | 0.36 | 0.31 | −16.58 | −5.13 |
| E8 | −2.15 | −4.47 | −7.72 | 3.1 | −13.7 | 13.81 | 5.49 | 1.77 | 0.19 | −1.72 | −2.52 |
| 307fu | −1.57 | 3.86 | 14.23 | 1.34 | 12.37 | −7.61 | −6.49 | 0.83 | −0.3 | −7.83 | 2.57 |
| 14T22 | −4.2 | −9.41 | −14 | 0.16 | −16.55 | 3.16 | −2.02 | −0.29 | 0.33 | −10.09 | −7.76 |
| 157 | 11.65 | 22.24 | 9.07 | 3.48 | 5.06 | −4.84 | 3.26 | 0.46 | −0.37 | 16.61 | 8.61 |
| 3618 | 8.65 | 16.78 | 17.19 | −10.99 | 54.92 | −15.35 | −5.21 | −1.98 | −0.79 | 12.72 | 13.44 |
| 124fu | −1.39 | −5.17 | −6.29 | 3.82 | −3.11 | 0.04 | −0.58 | −1.7 | −0.47 | 12.63 | 0.07 |
| JinR7 | −11.11 | −21.09 | −11.74 | −1.01 | −25.78 | 5.06 | 5.49 | 0.46 | 0.14 | −18.98 | −12.01 |
| Meiza | 0.12 | −2.75 | −0.74 | 0.11 | −13.21 | 5.74 | 0.06 | 0.46 | 0.28 | −3.35 | −2.45 |
| Parent | 428A | 521A | 170A | I15A | 307A | JinchangzaoA | 4190A | QL33A |
|---|---|---|---|---|---|---|---|---|
| E8 | 4.61 | 3.18 | 5.47 | 3.57 | 3.57 | 6.02 | 3.88 | 4.92 |
| 307fu | 3.44 | 3.5 | 4.18 | 4.13 | 1.54 | 5.52 | 3.19 | 3.8 |
| 14T22 | 4.33 | 2.63 | 5.86 | 3.15 | 3.58 | 5.64 | 4.4 | 5.18 |
| 157 | 3.64 | 4.71 | 5.39 | 5.74 | 4.65 | 6.09 | 4.98 | 6.14 |
| 3618 | 4.04 | 3.7 | 4.61 | 2.84 | 3.18 | 4.1 | 4.49 | 5.86 |
| 124fu | 3.79 | 3.76 | 4.39 | 4.13 | 2.63 | 5.86 | 3.03 | 4.6 |
| JinR7 | 3.87 | 5.07 | 2.46 | 5 | 2.79 | 5.8 | 3.65 | 6.15 |
| Parent | 428A | 521A | 170A | I15A | 307A | JinchangzaoA | 4190A | QL33A |
| Genetic Distance | Agronomic Traits 1 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| GY | PWT | TKW | PL | PH | SD | DF | DM | TL | BIY | |
| Phenotypic Genetic Distance | −0.34 | 0.66 | −0.15 | −0.18 | 0.34 | −0.4 | −0.07 | 0.46 | 0.53 | 0.42 |
| Molecular Genetic Distance | 0.61 * | 0.80 ** | 0.4 | −0.46 | −0.27 | −0.18 | −0.29 | 0.43 | −0.2 | 0.49 |
| Combining Ability | Agronomic Traits 2 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| GY | PWT | TKW | PL | PH | SD | DF | DM | TL | BIY | |
| GCA | 0.610 * | 0.560 * | 0.620 * | 0.035 | 0.170 | −0.540 * | −0.150 | −0.025 | −0.420 | 0.190 |
| SCA | 0.340 ** | 0.520 ** | 0.180 | 0.070 | 0.720 ** | −0.056 | −0.850 ** | −0.260 * | 0.310 * | 0.530 ** |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, H.; Lyu, D.; Zhang, Y.; Wang, W.; Zhao, R.; Lü, P.; Zhao, W.; Zhou, Z.; Lu, S. Classification of Heterotic Groups and Prediction of Heterosis in Sorghum Based on Whole-Genome Resequencing. Int. J. Mol. Sci. 2025, 26, 7950. https://doi.org/10.3390/ijms26167950
Zhang H, Lyu D, Zhang Y, Wang W, Zhao R, Lü P, Zhao W, Zhou Z, Lu S. Classification of Heterotic Groups and Prediction of Heterosis in Sorghum Based on Whole-Genome Resequencing. International Journal of Molecular Sciences. 2025; 26(16):7950. https://doi.org/10.3390/ijms26167950
Chicago/Turabian StyleZhang, Hongyou, Dexin Lyu, Yu Zhang, Wei Wang, Renjie Zhao, Pengfei Lü, Wenjing Zhao, Ziyang Zhou, and Shan Lu. 2025. "Classification of Heterotic Groups and Prediction of Heterosis in Sorghum Based on Whole-Genome Resequencing" International Journal of Molecular Sciences 26, no. 16: 7950. https://doi.org/10.3390/ijms26167950
APA StyleZhang, H., Lyu, D., Zhang, Y., Wang, W., Zhao, R., Lü, P., Zhao, W., Zhou, Z., & Lu, S. (2025). Classification of Heterotic Groups and Prediction of Heterosis in Sorghum Based on Whole-Genome Resequencing. International Journal of Molecular Sciences, 26(16), 7950. https://doi.org/10.3390/ijms26167950







