Proteomic Insights into Bacterial Responses to Antibiotics: A Narrative Review
Abstract
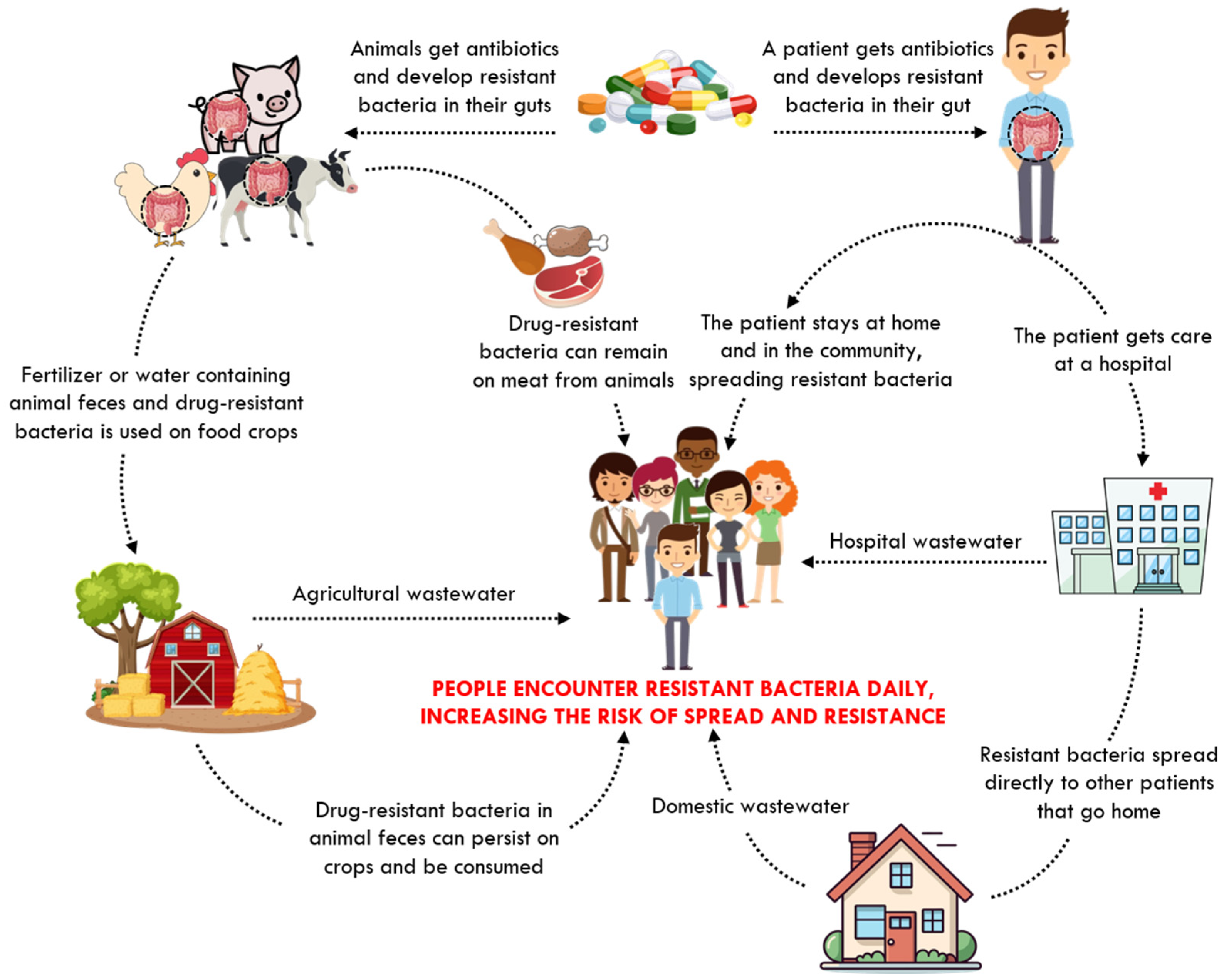
1. Introduction
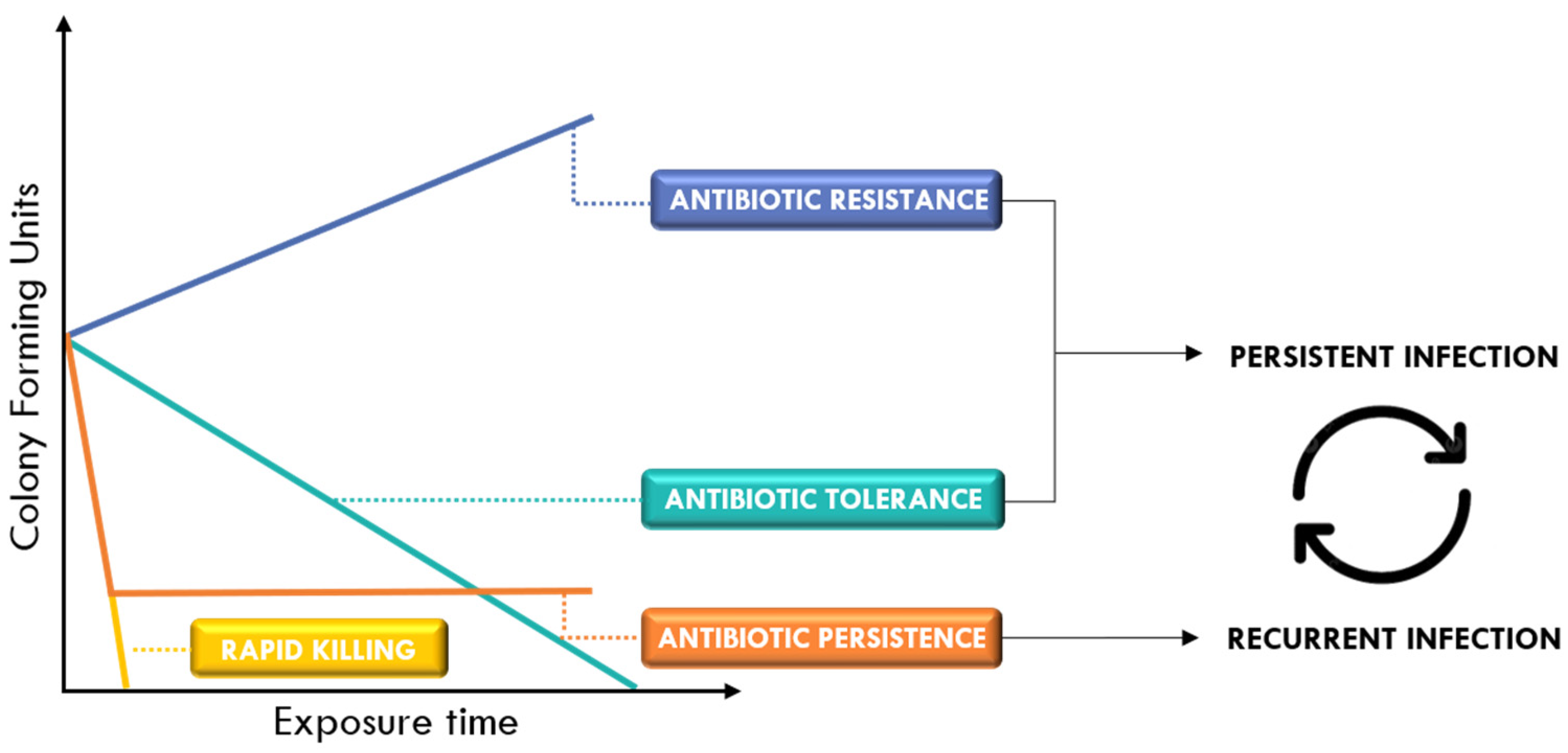
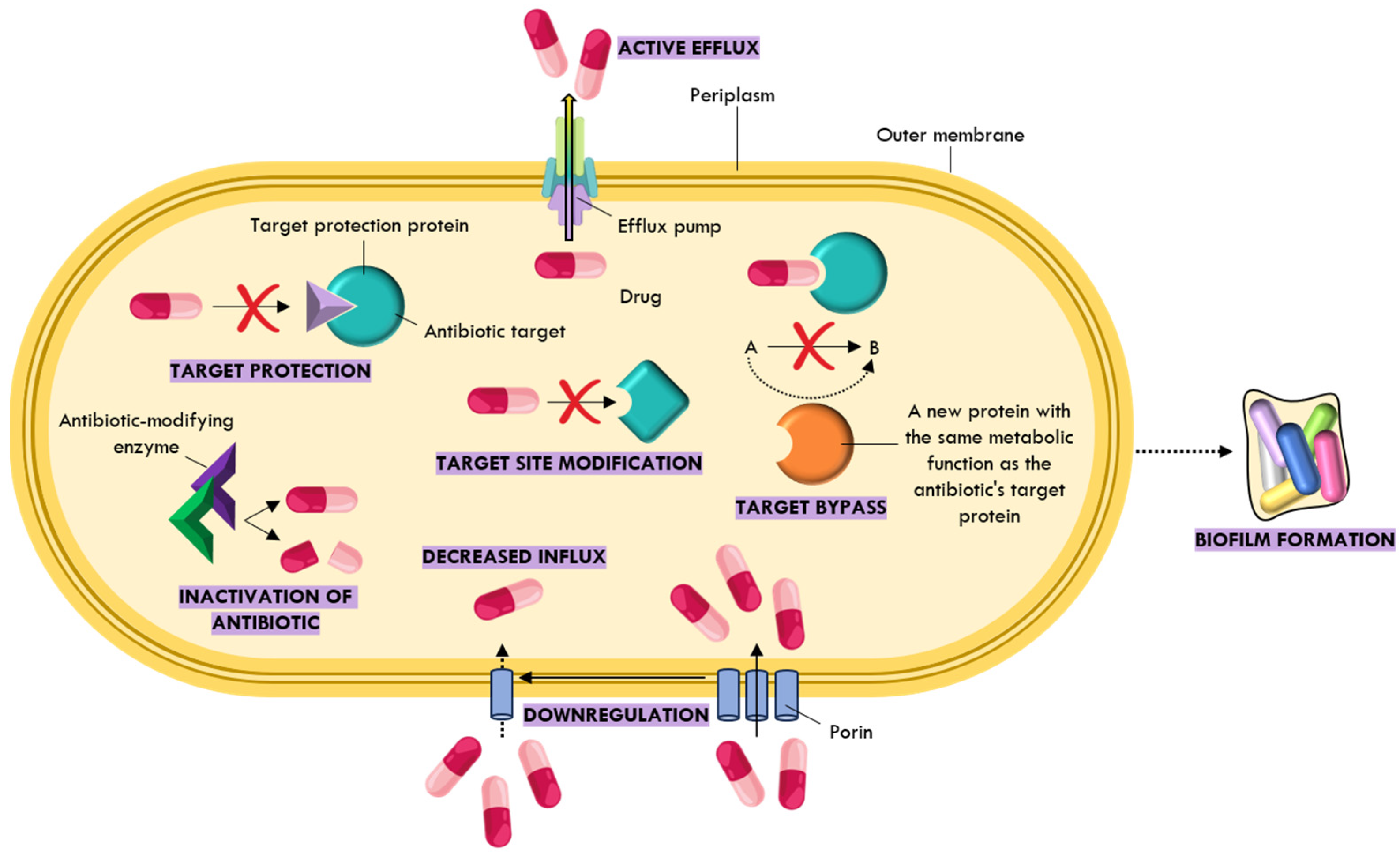
2. Different Survival Strategies of Bacteria Against Antibiotics
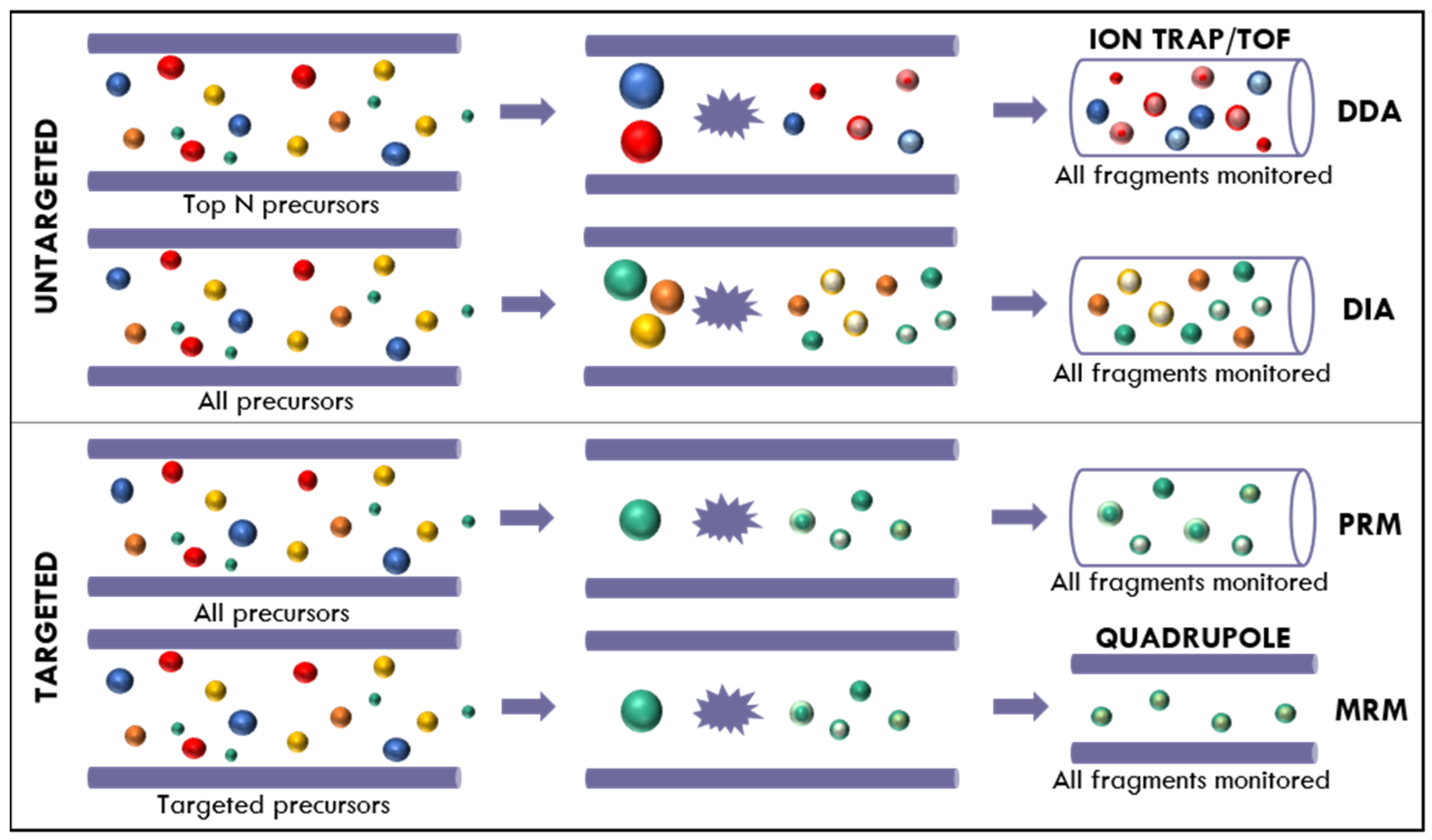
3. Proteomics by Mass Spectrometry as a Tool for Understanding AMR
3.1. Antimicrobial Resistance in ESKAPE Pathogens
3.1.1. Resistance to β-Lactams
3.1.2. Resistance to Aminoglycosides
3.1.3. Resistance to Quinolones
4. Advancing AMR Prediction: Future Directions and Challenges
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Founou, R.C.; Founou, L.L.; Essack, S.Y. Clinical and Economic Impact of Antibiotic Resistance in Developing Countries: A Systematic Review and Meta-Analysis. PLoS ONE 2017, 12, e0189621. [Google Scholar] [CrossRef]
- Tang, K.W.K.; Millar, B.C.; Moore, J.E. Antimicrobial Resistance (AMR). Br. J. Biomed. Sci. 2023, 80, 11387. [Google Scholar] [CrossRef]
- Ahmed, S.K.; Hussein, S.; Qurbani, K.; Ibrahim, R.H.; Fareeq, A.; Mahmood, K.A.; Mohamed, M.G. Antimicrobial Resistance: Impacts, Challenges, and Future Prospects. J. Med. Surg. Public Health 2024, 2, 100081. [Google Scholar] [CrossRef]
- Prestinaci, F.; Pezzotti, P.; Pantosti, A. Antimicrobial Resistance: A Global Multifaceted Phenomenon. Pathog. Glob. Health 2015, 109, 309–318. [Google Scholar] [CrossRef]
- Founou, R.C.; Blocker, A.J.; Noubom, M.; Tsayem, C.; Choukem, S.P.; Van Dongen, M.; Founou, L.L. The COVID-19 Pandemic: A Threat to Antimicrobial Resistance Containment. Future Sci. OA 2021, 7, FSO736. [Google Scholar] [CrossRef] [PubMed]
- Shrestha, P.; Cooper, B.S.; Coast, J.; Oppong, R.; Do Thi Thuy, N.; Phodha, T.; Celhay, O.; Guerin, P.J.; Wertheim, H.; Lubell, Y. Enumerating the Economic Cost of Antimicrobial Resistance per Antibiotic Consumed to Inform the Evaluation of Interventions Affecting Their Use. Antimicrob. Resist. Infect. Control 2018, 7, 98. [Google Scholar] [CrossRef]
- World Health Organization. Antimicrobial Resistance. Available online: www.who.int/europe/health-topics/antimicrobial-resistance (accessed on 22 July 2025).
- Salam, M.A.; Al-Amin, M.Y.; Salam, M.T.; Pawar, J.S.; Akhter, N.; Rabaan, A.A.; Alqumber, M.A.A. Antimicrobial Resistance: A Growing Serious Threat for Global Public Health. Healthcare 2023, 11, 1946. [Google Scholar] [CrossRef]
- Murray, C.J.L.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global Burden of Bacterial Antimicrobial Resistance in 2019: A Systematic Analysis. Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef] [PubMed]
- Aminov, R.I. A Brief History of the Antibiotic Era: Lessons Learned and Challenges for the Future. Front. Microbiol. 2010, 1, 134. [Google Scholar] [CrossRef] [PubMed]
- Hutchings, M.I.; Truman, A.W.; Wilkinson, B. Antibiotics: Past, Present and Future. Curr. Opin. Microbiol. 2019, 51, 72–80. [Google Scholar] [CrossRef]
- Lobanovska, M.; Pilla, G. Penicillin’s Discovery and Antibiotic Resistance: Lessons for the Future? Yale J. Biol. Med. 2017, 90, 135–145. [Google Scholar] [PubMed]
- Correia, J.; Borges, A.; Simões, M.; Simões, L. Beyond Penicillin: The Potential of Filamentous Fungi for Drug Discovery in the Age of Antibiotic Resistance. Antibiotics 2023, 12, 1250. [Google Scholar] [CrossRef]
- Alanis, A.J. Resistance to Antibiotics: Are We in the Post-Antibiotic Era? Arch. Med. Res. 2005, 36, 697–705. [Google Scholar] [CrossRef] [PubMed]
- Tommasi, R.; Brown, D.G.; Walkup, G.K.; Manchester, J.I.; Miller, A.A. ESKAPEing the Labyrinth of Antibacterial Discovery. Nat. Rev. Drug Discov. 2015, 14, 529–542. [Google Scholar] [CrossRef]
- Fisher, J.F.; Mobashery, S. Endless Resistance. Endless Antibiotics? Medchemcomm 2016, 7, 37–49. [Google Scholar] [CrossRef]
- Boyd, N.K.; Teng, C.; Frei, C.R. Brief Overview of Approaches and Challenges in New Antibiotic Development: A Focus On Drug Repurposing. Front. Cell. Infect. Microbiol. 2021, 11, 684515. [Google Scholar] [CrossRef]
- Martin, M.J.; Thottathil, S.E.; Newman, T.B. Antibiotics Overuse in Animal Agriculture: A Call to Action for Health Care Providers. Am. J. Public Health 2015, 105, 2409–2410. [Google Scholar] [CrossRef]
- Auta, A.; Hadi, M.A.; Oga, E.; Adewuyi, E.O.; Abdu-Aguye, S.N.; Adeloye, D.; Strickland-Hodge, B.; Morgan, D.J. Global Access to Antibiotics without Prescription in Community Pharmacies: A Systematic Review and Meta-Analysis. J. Infect. 2019, 78, 8–18. [Google Scholar] [CrossRef]
- Milani, R.V.; Wilt, J.K.; Entwisle, J.; Hand, J.; Cazabon, P.; Bohan, J.G. Reducing Inappropriate Outpatient Antibiotic Prescribing: Normative Comparison Using Unblinded Provider Reports. BMJ Open Qual. 2019, 8, e000351. [Google Scholar] [CrossRef]
- Ierano, C.; Thursky, K.; Marshall, C.; Koning, S.; James, R.; Johnson, S.; Imam, N.; Worth, L.J.; Peel, T. Appropriateness of Surgical Antimicrobial Prophylaxis Practices in Australia. JAMA Netw. Open 2019, 2, e1915003. [Google Scholar] [CrossRef] [PubMed]
- Browne, K.; Chakraborty, S.; Chen, R.; Willcox, M.D.; Black, D.S.; Walsh, W.R.; Kumar, N. A New Era of Antibiotics: The Clinical Potential of Antimicrobial Peptides. Int. J. Mol. Sci. 2020, 21, 7047. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Available online: www.cdc.gov/narms/resistance/ (accessed on 22 July 2025).
- Kannan, E.P.; Gopal, J.; Muthu, M. Analytical Techniques for Assessing Antimicrobial Resistance: Conventional Solutions, Contemporary Problems and Futuristic Outlooks. TrAC Trends Anal. Chem. 2024, 178, 117843. [Google Scholar] [CrossRef]
- McArthur, A.G.; Tsang, K.K. Antimicrobial Resistance Surveillance in the Genomic Age. Ann. N. Y. Acad. Sci. 2017, 1388, 78–91. [Google Scholar] [CrossRef]
- Jia, B.; Raphenya, A.R.; Alcock, B.; Waglechner, N.; Guo, P.; Tsang, K.K.; Lago, B.A.; Dave, B.M.; Pereira, S.; Sharma, A.N.; et al. CARD 2017: Expansion and Model-Centric Curation of the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 2017, 45, D566–D573. [Google Scholar] [CrossRef]
- Ellington, M.J.; Ekelund, O.; Aarestrup, F.M.; Canton, R.; Doumith, M.; Giske, C.; Grundman, H.; Hasman, H.; Holden, M.T.G.; Hopkins, K.L.; et al. The Role of Whole Genome Sequencing in Antimicrobial Susceptibility Testing of Bacteria: Report from the EUCAST Subcommittee. Clin. Microbiol. Infect. 2017, 23, 2–22. [Google Scholar] [CrossRef]
- Liu, Y.; Beyer, A.; Aebersold, R. On the Dependency of Cellular Protein Levels on MRNA Abundance. Cell 2016, 165, 535–550. [Google Scholar] [CrossRef]
- D’Esposito, M.; Ciccodicola, A.; Gianfrancesco, F.; Esposito, T.; Flagiello, L.; Mazzarella, R.; Schlessinger, D.; D’Urso, M. A Synaptobrevin–like Gene in the Xq28 Pseudoautosomal Region Undergoes X Inactivation. Nat. Genet. 1996, 13, 227–229. [Google Scholar] [CrossRef] [PubMed]
- Vogel, C.; Marcotte, E.M. Insights into the Regulation of Protein Abundance from Proteomic and Transcriptomic Analyses. Nat. Rev. Genet. 2012, 13, 227–232. [Google Scholar] [CrossRef] [PubMed]
- Fluit, A.C.; Visser, M.R.; Schmitz, F.-J. Molecular Detection of Antimicrobial Resistance. Clin. Microbiol. Rev. 2001, 14, 836–871. [Google Scholar] [CrossRef] [PubMed]
- Aslam, B.; Basit, M.; Nisar, M.A.; Khurshid, M.; Rasool, M.H. Proteomics: Technologies and Their Applications. J. Chromatogr. Sci. 2017, 55, 182–196. [Google Scholar] [CrossRef]
- Jadot, M.; Boonen, M.; Thirion, J.; Wang, N.; Xing, J.; Zhao, C.; Tannous, A.; Qian, M.; Zheng, H.; Everett, J.K.; et al. Accounting for Protein Subcellular Localization: A Compartmental Map of the Rat Liver Proteome. Mol. Cell. Proteom. 2017, 16, 194–212. [Google Scholar] [CrossRef]
- Torres-Sangiao, E.; Giddey, A.D.; Leal Rodriguez, C.; Tang, Z.; Liu, X.; Soares, N.C. Proteomic Approaches to Unravel Mechanisms of Antibiotic Resistance and Immune Evasion of Bacterial Pathogens. Front. Med. 2022, 9, 850374. [Google Scholar] [CrossRef]
- Sulaiman, J.E.; Lam, H. Evolution of Bacterial Tolerance Under Antibiotic Treatment and Its Implications on the Development of Resistance. Front. Microbiol. 2021, 12, 617412. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.M.; Nielsen, K.M. Mechanisms of, and Barriers to, Horizontal Gene Transfer between Bacteria. Nat. Rev. Microbiol. 2005, 3, 711–721. [Google Scholar] [CrossRef]
- Chiș, A.A.; Rus, L.L.; Morgovan, C.; Arseniu, A.M.; Frum, A.; Vonica-Țincu, A.L.; Gligor, F.G.; Mureșan, M.L.; Dobrea, C.M. Microbial Resistance to Antibiotics and Effective Antibiotherapy. Biomedicines 2022, 10, 1121. [Google Scholar] [CrossRef]
- Mohanty, H.; Pachpute, S.; Yadav, R.P. Mechanism of Drug Resistance in Bacteria: Efflux Pump Modulation for Designing of New Antibiotic Enhancers. Folia Microbiol. 2021, 66, 727–739. [Google Scholar] [CrossRef]
- De Pascale, G.; Wright, G.D. Antibiotic Resistance by Enzyme Inactivation: From Mechanisms to Solutions. ChemBioChem 2010, 11, 1325–1334. [Google Scholar] [CrossRef]
- Wright, G. Bacterial Resistance to Antibiotics: Enzymatic Degradation and Modification. Adv. Drug Deliv. Rev. 2005, 57, 1451–1470. [Google Scholar] [CrossRef] [PubMed]
- Lambert, P. Bacterial Resistance to Antibiotics: Modified Target Sites. Adv. Drug Deliv. Rev. 2005, 57, 1471–1485. [Google Scholar] [CrossRef]
- Schaenzer, A.J.; Wright, G.D. Antibiotic Resistance by Enzymatic Modification of Antibiotic Targets. Trends Mol. Med. 2020, 26, 768–782. [Google Scholar] [CrossRef] [PubMed]
- Then, R.L. Mechanisms of Resistance to Trimethoprim, the Sulfonamides, and Trimethoprim-Sulfamethoxazole. Clin. Infect. Dis. 1982, 4, 261–269. [Google Scholar] [CrossRef]
- Fernández, L.; Hancock, R.E.W. Adaptive and Mutational Resistance: Role of Porins and Efflux Pumps in Drug Resistance. Clin. Microbiol. Rev. 2012, 25, 661–681. [Google Scholar] [CrossRef] [PubMed]
- Webber, M.A. The Importance of Efflux Pumps in Bacterial Antibiotic Resistance. J. Antimicrob. Chemother. 2003, 51, 9–11. [Google Scholar] [CrossRef]
- Wilson, D.N.; Hauryliuk, V.; Atkinson, G.C.; O’Neill, A.J. Target Protection as a Key Antibiotic Resistance Mechanism. Nat. Rev. Microbiol. 2020, 18, 637–648. [Google Scholar] [CrossRef]
- Melnyk, A.H.; Wong, A.; Kassen, R. The Fitness Costs of Antibiotic Resistance Mutations. Evol. Appl. 2015, 8, 273–283. [Google Scholar] [CrossRef]
- Vogwill, T.; MacLean, R.C. The Genetic Basis of the Fitness Costs of Antimicrobial Resistance: A Meta-analysis Approach. Evol. Appl. 2015, 8, 284–295. [Google Scholar] [CrossRef]
- Alonso, A. Overexpression of the Multidrug Efflux Pump SmeDEF Impairs Stenotrophomonas Maltophilia Physiology. J. Antimicrob. Chemother. 2004, 53, 432–434. [Google Scholar] [CrossRef] [PubMed]
- Olivares, J.; Alvarez-Ortega, C.; Linares, J.F.; Rojo, F.; Köhler, T.; Martínez, J.L. Overproduction of the Multidrug Efflux Pump MexEF-OprN Does Not Impair Pseudomonas aeruginosa Fitness in Competition Tests, but Produces Specific Changes in Bacterial Regulatory Networks. Environ. Microbiol. 2012, 14, 1968–1981. [Google Scholar] [CrossRef] [PubMed]
- Bagel, S.; Hüllen, V.; Wiedemann, B.; Heisig, P. Impact of GyrA and ParC Mutations on Quinolone Resistance, Doubling Time, and Supercoiling Degree of Escherichia coli. Antimicrob. Agents Chemother. 1999, 43, 868–875. [Google Scholar] [CrossRef]
- Enne, V.I. Reducing Antimicrobial Resistance in the Community by Restricting Prescribing: Can It Be Done? J. Antimicrob. Chemother. 2010, 65, 179–182. [Google Scholar] [CrossRef]
- Andersson, D.I. The Biological Cost of Mutational Antibiotic Resistance: Any Practical Conclusions? Curr. Opin. Microbiol. 2006, 9, 461–465. [Google Scholar] [CrossRef]
- Uruén, C.; Chopo-Escuin, G.; Tommassen, J.; Mainar-Jaime, R.C.; Arenas, J. Biofilms as Promoters of Bacterial Antibiotic Resistance and Tolerance. Antibiotics 2020, 10, 3. [Google Scholar] [CrossRef]
- van Duin, D.; Paterson, D.L. Multidrug-Resistant Bacteria in the Community. Infect. Dis. Clin. N. Am. 2020, 34, 709–722. [Google Scholar] [CrossRef]
- Peacock, S.J.; Paterson, G.K. Mechanisms of Methicillin Resistance in Staphylococcus aureus. Annu. Rev. Biochem. 2015, 84, 577–601. [Google Scholar] [CrossRef]
- Baroud, M.; Dandache, I.; Araj, G.F.; Wakim, R.; Kanj, S.; Kanafani, Z.; Khairallah, M.; Sabra, A.; Shehab, M.; Dbaibo, G.; et al. Underlying Mechanisms of Carbapenem Resistance in Extended-Spectrum β-Lactamase-Producing Klebsiella Pneumoniae and Escherichia coli Isolates at a Tertiary Care Centre in Lebanon: Role of OXA-48 and NDM-1 Carbapenemases. Int. J. Antimicrob. Agents 2013, 41, 75–79. [Google Scholar] [CrossRef] [PubMed]
- Mishra, R.; Shukla, P.; Huang, W.; Hu, N. Gene Mutations in Mycobacterium Tuberculosis: Multidrug-Resistant TB as an Emerging Global Public Health Crisis. Tuberculosis 2015, 95, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Espinosa, K.; Park, J.A.; Gerrity, J.J.; Buono, S.; Shearer, A.; Dick, C.; Mak, M.L.; Teramoto, K.; Klausner, J.D.; Pandori, M.; et al. Fluoroquinolone Resistance in Neisseria Gonorrhoeae After Cessation of Ciprofloxacin Usage in San Francisco. Sex. Transm. Dis. 2015, 42, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Kumar, A.; Roudbary, M.; Mohammadi, R.; Černáková, L.; Rodrigues, C.F. Overview on the Infections Related to Rare Candida Species. Pathogens 2022, 11, 963. [Google Scholar] [CrossRef]
- Mason, S.; Devincenzo, J.P.; Toovey, S.; Wu, J.Z.; Whitley, R.J. Comparison of Antiviral Resistance across Acute and Chronic Viral Infections. Antivir. Res. 2018, 158, 103–112. [Google Scholar] [CrossRef]
- Gupta, N.; Tanner, S.; Jaitly, N.; Adkins, J.N.; Lipton, M.; Edwards, R.; Romine, M.; Osterman, A.; Bafna, V.; Smith, R.D.; et al. Whole Proteome Analysis of Post-Translational Modifications: Applications of Mass-Spectrometry for Proteogenomic Annotation. Genome Res. 2007, 17, 1362–1377. [Google Scholar] [CrossRef][Green Version]
- Macklin, A.; Khan, S.; Kislinger, T. Recent Advances in Mass Spectrometry Based Clinical Proteomics: Applications to Cancer Research. Clin. Proteom. 2020, 17, 17. [Google Scholar] [CrossRef]
- Noor, Z.; Ahn, S.B.; Baker, M.S.; Ranganathan, S.; Mohamedali, A. Mass Spectrometry–Based Protein Identification in Proteomics—A Review. Brief. Bioinform. 2021, 22, 1620–1638. [Google Scholar] [CrossRef]
- Han, X.; Aslanian, A.; Yates, J.R. Mass Spectrometry for Proteomics. Curr. Opin. Chem. Biol. 2008, 12, 483–490. [Google Scholar] [CrossRef]
- Cumeras, R.; Figueras, E.; Davis, C.E.; Baumbach, J.I.; Gràcia, I. Review on Ion Mobility Spectrometry. Part 2: Hyphenated Methods and Effects of Experimental Parameters. Analyst 2015, 140, 1391–1410. [Google Scholar] [CrossRef] [PubMed]
- Eiceman, G.A.; Karpas, Z. Ion Mobility Spectrometry; CRC Press: Boca Raton, FL, USA, 2005; ISBN 9780429195891. [Google Scholar]
- Helm, D.; Vissers, J.P.C.; Hughes, C.J.; Hahne, H.; Ruprecht, B.; Pachl, F.; Grzyb, A.; Richardson, K.; Wildgoose, J.; Maier, S.K.; et al. Ion Mobility Tandem Mass Spectrometry Enhances Performance of Bottom-up Proteomics. Mol. Cell. Proteom. 2014, 13, 3709–3715. [Google Scholar] [CrossRef]
- Lanucara, F.; Holman, S.W.; Gray, C.J.; Eyers, C.E. The Power of Ion Mobility-Mass Spectrometry for Structural Characterization and the Study of Conformational Dynamics. Nat. Chem. 2014, 6, 281–294. [Google Scholar] [CrossRef]
- Ewing, M.A.; Glover, M.S.; Clemmer, D.E. Hybrid Ion Mobility and Mass Spectrometry as a Separation Tool. J. Chromatogr. A 2016, 1439, 3–25. [Google Scholar] [CrossRef]
- Chi, H.; Chen, H.; He, K.; Wu, L.; Yang, B.; Sun, R.-X.; Liu, J.; Zeng, W.-F.; Song, C.-Q.; He, S.-M.; et al. PNovo+: De Novo Peptide Sequencing Using Complementary HCD and ETD Tandem Mass Spectra. J. Proteome Res. 2013, 12, 615–625. [Google Scholar] [CrossRef]
- Ma, B. Novor: Real-Time Peptide de Novo Sequencing Software. J. Am. Soc. Mass. Spectrom. 2015, 26, 1885–1894. [Google Scholar] [CrossRef] [PubMed]
- Muth, T.; Renard, B.Y. Evaluating de Novo Sequencing in Proteomics: Already an Accurate Alternative to Database-Driven Peptide Identification? Brief. Bioinform. 2018, 19, 954–970. [Google Scholar] [CrossRef] [PubMed]
- Mann, M.; Wilm, M. Error-Tolerant Identification of Peptides in Sequence Databases by Peptide Sequence Tags. Anal. Chem. 1994, 66, 4390–4399. [Google Scholar] [CrossRef]
- Eng, J.K.; McCormack, A.L.; Yates, J.R. An Approach to Correlate Tandem Mass Spectral Data of Peptides with Amino Acid Sequences in a Protein Database. J. Am. Soc. Mass. Spectrom. 1994, 5, 976–989. [Google Scholar] [CrossRef]
- Park, C.Y.; Klammer, A.A.; Käll, L.; MacCoss, M.J.; Noble, W.S. Rapid and Accurate Peptide Identification from Tandem Mass Spectra. J. Proteome Res. 2008, 7, 3022–3027. [Google Scholar] [CrossRef]
- Keller, A.; Eng, J.; Zhang, N.; Li, X.; Aebersold, R. A Uniform Proteomics MS/MS Analysis Platform Utilizing Open XML File Formats. Mol. Syst. Biol. 2005, 1, 2005.0017. [Google Scholar] [CrossRef]
- Cox, J.; Neuhauser, N.; Michalski, A.; Scheltema, R.A.; Olsen, J.V.; Mann, M. Andromeda: A Peptide Search Engine Integrated into the MaxQuant Environment. J. Proteome Res. 2011, 10, 1794–1805. [Google Scholar] [CrossRef] [PubMed]
- Stewart, I.I.; Thomson, T.; Figeys, D. 18O Labeling: A Tool for Proteomics. Rapid Commun. Mass. Spectrom. 2001, 15, 2456–2465. [Google Scholar] [CrossRef] [PubMed]
- Boersema, P.J.; Raijmakers, R.; Lemeer, S.; Mohammed, S.; Heck, A.J.R. Multiplex Peptide Stable Isotope Dimethyl Labeling for Quantitative Proteomics. Nat. Protoc. 2009, 4, 484–494. [Google Scholar] [CrossRef]
- Ong, S.-E.; Blagoev, B.; Kratchmarova, I.; Kristensen, D.B.; Steen, H.; Pandey, A.; Mann, M. Stable Isotope Labeling by Amino Acids in Cell Culture, SILAC, as a Simple and Accurate Approach to Expression Proteomics. Mol. Cell. Proteom. 2002, 1, 376–386. [Google Scholar] [CrossRef]
- Ong, S.-E.; Kratchmarova, I.; Mann, M. Properties of 13C-Substituted Arginine in Stable Isotope Labeling by Amino Acids in Cell Culture (SILAC). J. Proteome Res. 2003, 2, 173–181. [Google Scholar] [CrossRef] [PubMed]
- Megger, D.A.; Pott, L.L.; Ahrens, M.; Padden, J.; Bracht, T.; Kuhlmann, K.; Eisenacher, M.; Meyer, H.E.; Sitek, B. Comparison of Label-Free and Label-Based Strategies for Proteome Analysis of Hepatoma Cell Lines. Biochim. Biophys. Acta (BBA) Proteins Proteom. 2014, 1844, 967–976. [Google Scholar] [CrossRef]
- Patel, V.J.; Thalassinos, K.; Slade, S.E.; Connolly, J.B.; Crombie, A.; Murrell, J.C.; Scrivens, J.H. A Comparison of Labeling and Label-Free Mass Spectrometry-Based Proteomics Approaches. J. Proteome Res. 2009, 8, 3752–3759. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Mann, M. MaxQuant Enables High Peptide Identification Rates, Individualized p.p.b.-Range Mass Accuracies and Proteome-Wide Protein Quantification. Nat. Biotechnol. 2008, 26, 1367–1372. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Hein, M.Y.; Luber, C.A.; Paron, I.; Nagaraj, N.; Mann, M. Accurate Proteome-Wide Label-Free Quantification by Delayed Normalization and Maximal Peptide Ratio Extraction, Termed MaxLFQ. Mol. Cell. Proteom. 2014, 13, 2513–2526. [Google Scholar] [CrossRef]
- Simanjuntak, Y.; Schamoni-Kast, K.; Grün, A.; Uetrecht, C.; Scaturro, P. Top-Down and Bottom-Up Proteomics Methods to Study RNA Virus Biology. Viruses 2021, 13, 668. [Google Scholar] [CrossRef]
- Zhou, M.; Lantz, C.; Brown, K.A.; Ge, Y.; Paša-Tolić, L.; Loo, J.A.; Lermyte, F. Higher-Order Structural Characterisation of Native Proteins and Complexes by Top-down Mass Spectrometry. Chem. Sci. 2020, 11, 12918–12936. [Google Scholar] [CrossRef] [PubMed]
- Catherman, A.D.; Skinner, O.S.; Kelleher, N.L. Top Down Proteomics: Facts and Perspectives. Biochem. Biophys. Res. Commun. 2014, 445, 683–693. [Google Scholar] [CrossRef]
- Kalantari, S.; Jafari, A.; Moradpoor, R.; Ghasemi, E.; Khalkhal, E. Human Urine Proteomics: Analytical Techniques and Clinical Applications in Renal Diseases. Int. J. Proteom. 2015, 2015, 782798. [Google Scholar] [CrossRef]
- Park, J.; Kim, H.; Kim, S.Y.; Kim, Y.; Lee, J.-S.; Dan, K.; Seong, M.-W.; Han, D. In-Depth Blood Proteome Profiling Analysis Revealed Distinct Functional Characteristics of Plasma Proteins between Severe and Non-Severe COVID-19 Patients. Sci. Rep. 2020, 10, 22418. [Google Scholar] [CrossRef]
- Bader, J.M.; Geyer, P.E.; Müller, J.B.; Strauss, M.T.; Koch, M.; Leypoldt, F.; Koertvelyessy, P.; Bittner, D.; Schipke, C.G.; Incesoy, E.I.; et al. Proteome Profiling in Cerebrospinal Fluid Reveals Novel Biomarkers of Alzheimer’s Disease. Mol. Syst. Biol. 2020, 16, e9356. [Google Scholar] [CrossRef]
- Erdmann, J.; Thöming, J.G.; Pohl, S.; Pich, A.; Lenz, C.; Häussler, S. The Core Proteome of Biofilm-Grown Clinical Pseudomonas Aeruginosa Isolates. Cells 2019, 8, 1129. [Google Scholar] [CrossRef]
- Baros-Steyl, S.S.; Nakedi, K.C.; Ganief, T.A.; Okendo, J.O.; Tabb, D.L.; Soares, N.C.; Blackburn, J.M. Phosphoproteomics Reveals New Insights into the Role of PknG during the Persistence of Pathogenic Mycobacteria in Host Macrophages. bioRxiv 2021. [Google Scholar] [CrossRef]
- Zampieri, E.; Chiapello, M.; Daghino, S.; Bonfante, P.; Mello, A. Soil Metaproteomics Reveals an Inter-Kingdom Stress Response to the Presence of Black Truffles. Sci. Rep. 2016, 6, 25773. [Google Scholar] [CrossRef]
- Heyer, R.; Schallert, K.; Büdel, A.; Zoun, R.; Dorl, S.; Behne, A.; Kohrs, F.; Püttker, S.; Siewert, C.; Muth, T.; et al. A Robust and Universal Metaproteomics Workflow for Research Studies and Routine Diagnostics Within 24 h Using Phenol Extraction, FASP Digest, and the MetaProteomeAnalyzer. Front. Microbiol. 2019, 10, 1883. [Google Scholar] [CrossRef] [PubMed]
- Pak, H.; Michaux, J.; Huber, F.; Chong, C.; Stevenson, B.J.; Müller, M.; Coukos, G.; Bassani-Sternberg, M. Sensitive Immunopeptidomics by Leveraging Available Large-Scale Multi-HLA Spectral Libraries, Data-Independent Acquisition, and MS/MS Prediction. Mol. Cell. Proteom. 2021, 20, 100080. [Google Scholar] [CrossRef]
- Vidova, V.; Spacil, Z. A Review on Mass Spectrometry-Based Quantitative Proteomics: Targeted and Data Independent Acquisition. Anal. Chim. Acta 2017, 964, 7–23. [Google Scholar] [CrossRef] [PubMed]
- Castro-Gamero, A.M.; Izumi, C.; Rosa, J.C. Biomarker Verification Using Selected Reaction Monitoring and Shotgun Proteomics. Methods Mol. Biol. 2014, 1156, 295–306. [Google Scholar]
- Hristova, V.A.; Chan, D.W. Cancer Biomarker Discovery and Translation: Proteomics and Beyond. Expert Rev. Proteom. 2019, 16, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Keane, R.E.; Tidy, R.J.; Parker, G.J.; Gummer, J.P.A.; Priddis, C. Mass Spectrometry Based Proteomics: Changing the Impact of Protein Analysis in Forensic Science. WIREs Forensic Sci. 2024, 6, e1516. [Google Scholar] [CrossRef]
- Ludwig, C.; Gillet, L.; Rosenberger, G.; Amon, S.; Collins, B.C.; Aebersold, R. Data-independent Acquisition-based SWATH-MS for Quantitative Proteomics: A Tutorial. Mol. Syst. Biol. 2018, 14, e8126. [Google Scholar] [CrossRef]
- Sajic, T.; Liu, Y.; Aebersold, R. Using Data-independent, High-resolution Mass Spectrometry in Protein Biomarker Research: Perspectives and Clinical Applications. Proteom. Clin. Appl. 2015, 9, 307–321. [Google Scholar] [CrossRef]
- Aebersold, R.; Bensimon, A.; Collins, B.C.; Ludwig, C.; Sabido, E. Applications and Developments in Targeted Proteomics: From SRM to DIA/SWATH. Proteomics 2016, 16, 2065–2067. [Google Scholar] [CrossRef]
- Nakamura, K.; Hirayama-Kurogi, M.; Ito, S.; Kuno, T.; Yoneyama, T.; Obuchi, W.; Terasaki, T.; Ohtsuki, S. Large-scale Multiplex Absolute Protein Quantification of Drug-metabolizing Enzymes and Transporters in Human Intestine, Liver, and Kidney Microsomes by SWATH-MS: Comparison with MRM/SRM and HR-MRM/PRM. Proteomics 2016, 16, 2106–2117. [Google Scholar] [CrossRef] [PubMed]
- Belshan, M.; Holbrook, A.; George, J.W.; Durant, H.E.; Callahan, M.; Jaquet, S.; West, J.T.; Siedlik, J.; Ciborowski, P. Discovery of Candidate HIV-1 Latency Biomarkers Using an OMICs Approach. Virology 2021, 558, 86–95. [Google Scholar] [CrossRef]
- Malmström, E.; Kilsgård, O.; Hauri, S.; Smeds, E.; Herwald, H.; Malmström, L.; Malmström, J. Large-Scale Inference of Protein Tissue Origin in Gram-Positive Sepsis Plasma Using Quantitative Targeted Proteomics. Nat. Commun. 2016, 7, 10261. [Google Scholar] [CrossRef]
- Sidjabat, H.E.; Gien, J.; Kvaskoff, D.; Ashman, K.; Vaswani, K.; Reed, S.; McGeary, R.P.; Paterson, D.L.; Bordin, A.; Schenk, G. The Use of SWATH to Analyse the Dynamic Changes of Bacterial Proteome of Carbapanemase-Producing Escherichia coli under Antibiotic Pressure. Sci. Rep. 2018, 8, 3871. [Google Scholar] [CrossRef]
- Bakochi, A.; Mohanty, T.; Pyl, P.T.; Gueto-Tettay, C.A.; Malmström, L.; Linder, A.; Malmström, J. Cerebrospinal Fluid Proteome Maps Detect Pathogen-Specific Host Response Patterns in Meningitis. eLife 2021, 10, e64159. [Google Scholar] [CrossRef]
- Dülfer, J.; Kadek, A.; Kopicki, J.-D.; Krichel, B.; Uetrecht, C. Structural Mass Spectrometry Goes Viral. Adv. Virus Res. 2019, 105, 189–238. [Google Scholar]
- Limpikirati, P.; Liu, T.; Vachet, R.W. Covalent Labeling-Mass Spectrometry with Non-Specific Reagents for Studying Protein Structure and Interactions. Methods 2018, 144, 79–93. [Google Scholar] [CrossRef] [PubMed]
- Teichmann, L.; Pasman, R.; Luitwieler, S.; Varriale, C.; Bengtsson-Palme, J.; Ter Kuile, B. Adaptation of Escherichia coli to Ciprofloxacin and Enrofloxacin: Differential Proteomics of the SOS Response and RecA-Independent Mechanisms. Int. J. Antimicrob. Agents 2025, 65, 107420. [Google Scholar] [CrossRef] [PubMed]
- Aswal, M.; Singh, N.; Singhal, N.; Kumar, M. An Integrated Proteo-Transcriptomics Approach Reveals Novel Drug Targets against Multidrug Resistant Escherichia coli. Front. Microbiol. 2025, 16, 1531739. [Google Scholar] [CrossRef]
- Boucherabine, S.; Giddey, A.; Nassar, R.; Al-Hroub, H.M.; Mohamed, L.; Harb, M.; Soares, N.C.; Senok, A. Proteomic and Metabolomic Profiling of Methicillin Resistant versus Methicillin Sensitive Staphylococcus aureus Using a Simultaneous Extraction Protocol. Front. Microbiol. 2024, 15, 1402796. [Google Scholar] [CrossRef]
- Wang, Z.; Sun, X.; Chen, X.; Wang, H.; He, H. The Mass Spectrometry Identification, Antimicrobial Genes Detection, and Proteomics Analysis of Stutzerimonas stutzeri Strain Was Isolated from Industrial Wastewater. Separations 2023, 10, 461. [Google Scholar] [CrossRef]
- Goodyear, M.C.; Seidel, L.; Krieger, J.R.; Geddes-McAlister, J.; Levesque, R.C.; Khursigara, C.M. Quantitative Proteomics Reveals Unique Responses to Antimicrobial Treatments in Clinical Pseudomonas aeruginosa Isolates. mSystems 2023, 8, e0049123. [Google Scholar] [CrossRef]
- Li, X.; Feng, D.; Zhou, J.; Wu, W.; Zheng, W.; Gan, W.; Jiang, M.; Li, H.; Peng, X.; Zhang, T. Metabolomics Method in Understanding and Sensitizing Carbapenem-Resistant Acinetobacter baumannii to Meropenem. ACS Infect. Dis. 2024, 10, 184–195. [Google Scholar] [CrossRef]
- Deatherage Kaiser, B.L.; Birdsell, D.N.; Hutchison, J.R.; Thelaus, J.; Jenson, S.C.; Andrianaivoarimanana, V.; Byström, M.; Myrtennäs, K.; McDonough, R.F.; Nottingham, R.D.; et al. Proteomic Signatures of Antimicrobial Resistance in Yersinia Pestis and Francisella tularensis. Front. Med. 2022, 9, 821071. [Google Scholar] [CrossRef]
- Kou, T.; Wu, J.; Chen, X.; Peng, B. Functional Proteomics Identify Mannitol Metabolism in Serum Resistance and Therapeutic Implications in Vibrio alginolyticus. Front. Immunol. 2022, 13, 1010526. [Google Scholar] [CrossRef]
- Bakker, A.T.; Kotsogianni, I.; Mirenda, L.; Straub, V.M.; Avalos, M.; van den Berg, R.J.B.H.N.; Florea, B.I.; van Wezel, G.P.; Janssen, A.P.A.; Martin, N.I.; et al. Chemical Proteomics Reveals Antibiotic Targets of Oxadiazolones in MRSA. J. Am. Chem. Soc. 2023, 145, 1136–1143. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, L.; Song, Q.; Wang, G.; Yang, W.; Tang, H.; Srinivasan, R.; Lin, L.; Lin, X. Proteomics Analysis Reveals Bacterial Antibiotics Resistance Mechanism Mediated by AhslyA Against Enoxacin in Aeromonas hydrophila. Front. Microbiol. 2021, 12, 699415. [Google Scholar] [CrossRef]
- Shen, C.; Shen, Y.; Zhang, H.; Xu, M.; He, L.; Qie, J. Comparative Proteomics Demonstrates Altered Metabolism Pathways in Cotrimoxazole-Resistant and Amikacin-Resistant Klebsiella pneumoniae Isolates. Front. Microbiol. 2021, 12, 773829. [Google Scholar] [CrossRef]
- Ma, H.; Lai, B.; Jin, Y.; Tian, C.; Liu, J.; Wang, K. Proteomics and Metabolomics Analysis Reveal Potential Mechanism of Extended-Spectrum β-Lactamase Production in Escherichia coli. RSC Adv. 2020, 10, 26862–26873. [Google Scholar] [CrossRef]
- Lu, Y.; Pang, J.; Wang, G.; Hu, X.; Li, X.; Li, G.; Wang, X.; Yang, X.; Li, C.; You, X. Quantitative Proteomics Approach to Investigate the Antibacterial Response of Helicobacter pylori to Daphnetin, a Traditional Chinese Medicine Monomer. RSC Adv. 2021, 11, 2185–2193. [Google Scholar] [CrossRef]
- Liu, X.; Wang, J.; Chen, M.; Che, R.; Ding, W.; Yu, F.; Zhou, Y.; Cui, W.; Xiaoxu, X.; God’spower, B.-O.; et al. Comparative Proteomic Analysis Reveals Drug Resistance of Staphylococcus xylosus ATCC700404 under Tylosin Stress. BMC Vet. Res. 2019, 15, 224. [Google Scholar] [CrossRef]
- Uddin, M.J.; Ma, C.J.; Kim, J.-C.; Ahn, J. Proteomics-Based Discrimination of Differentially Expressed Proteins in Antibiotic-Sensitive and Antibiotic-Resistant Salmonella Typhimurium, Klebsiella pneumoniae, and Staphylococcus aureus. Arch. Microbiol. 2019, 201, 1259–1275. [Google Scholar] [CrossRef]
- Ye, J.; Lin, X.; Cheng, Z.; Su, Y.; Li, W.; Ali, F.; Zheng, J.; Peng, B. Identification and Efficacy of Glycine, Serine and Threonine Metabolism in Potentiating Kanamycin-Mediated Killing of Edwardsiella piscicida. J. Proteom. 2018, 183, 34–44. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Yao, Z.; Sun, L.; Hu, W.; Cao, J.; Lin, W.; Lin, X. Proteomics Analysis Reveals a Potential Antibiotic Cocktail Therapy Strategy for Aeromonas hydrophila Infection in Biofilm. J. Proteome Res. 2016, 15, 1810–1820. [Google Scholar] [CrossRef]
- Rice, L.B. Federal Funding for the Study of Antimicrobial Resistance in Nosocomial Pathogens: No ESKAPE. J. Infect. Dis. 2008, 197, 1079–1081. [Google Scholar] [CrossRef]
- Beatson, S.A.; Walker, M.J. Tracking Antibiotic Resistance. Science (1979) 2014, 345, 1454–1455. [Google Scholar] [CrossRef] [PubMed]
- Naylor, N.R.; Atun, R.; Zhu, N.; Kulasabanathan, K.; Silva, S.; Chatterjee, A.; Knight, G.M.; Robotham, J.V. Estimating the Burden of Antimicrobial Resistance: A Systematic Literature Review. Antimicrob. Resist. Infect. Control 2018, 7, 58. [Google Scholar] [CrossRef] [PubMed]
- Jansen, K.U.; Knirsch, C.; Anderson, A.S. The Role of Vaccines in Preventing Bacterial Antimicrobial Resistance. Nat. Med. 2018, 24, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Spiller, R.C. Hidden Dangers of Antibiotic Use: Increased Gut Permeability Mediated by Increased Pancreatic Proteases Reaching the Colon. Cell Mol. Gastroenterol. Hepatol. 2018, 6, 347–348.e1. [Google Scholar] [CrossRef]
- Argudín, M.; Deplano, A.; Meghraoui, A.; Dodémont, M.; Heinrichs, A.; Denis, O.; Nonhoff, C.; Roisin, S. Bacteria from Animals as a Pool of Antimicrobial Resistance Genes. Antibiotics 2017, 6, 12. [Google Scholar] [CrossRef] [PubMed]
- Bush, K.; Bradford, P.A. β-Lactams and β-Lactamase Inhibitors: An Overview. Cold Spring Harb. Perspect. Med. 2016, 6, a025247. [Google Scholar] [CrossRef] [PubMed]
- Oates, J.A.; Wood, A.J.J.; Donowitz, G.R.; Mandell, G.L. Beta-Lactam Antibiotics. N. Engl. J. Med. 1988, 318, 419–426. [Google Scholar] [CrossRef]
- Zapun, A.; Contreras-Martel, C.; Vernet, T. Penicillin-Binding Proteins and β-Lactam Resistance. FEMS Microbiol. Rev. 2008, 32, 361–385. [Google Scholar] [CrossRef] [PubMed]
- Fisher, J.F.; Mobashery, S. β-Lactam Resistance Mechanisms: Gram-Positive Bacteria and Mycobacterium tuberculosis. Cold Spring Harb. Perspect. Med. 2016, 6, a025221. [Google Scholar] [CrossRef]
- Bush, K. Proliferation and Significance of Clinically Relevant Β-lactamases. Ann. N. Y. Acad. Sci. 2013, 1277, 84–90. [Google Scholar] [CrossRef]
- Eren, E.; Vijayaraghavan, J.; Liu, J.; Cheneke, B.R.; Touw, D.S.; Lepore, B.W.; Indic, M.; Movileanu, L.; van den Berg, B. Substrate Specificity within a Family of Outer Membrane Carboxylate Channels. PLoS Biol. 2012, 10, e1001242. [Google Scholar] [CrossRef]
- Xu, C.; Lin, X.; Ren, H.; Zhang, Y.; Wang, S.; Peng, X. Analysis of Outer Membrane Proteome of Escherichia coli Related to Resistance to Ampicillin and Tetracycline. Proteomics 2006, 6, 462–473. [Google Scholar] [CrossRef]
- Peng, X.; Xu, C.; Ren, H.; Lin, X.; Wu, L.; Wang, S. Proteomic Analysis of the Sarcosine-Insoluble Outer Membrane Fraction of Pseudomonas a Eruginosa Responding to Ampicilin, Kanamycin, and Tetracycline Resistance. J. Proteome Res. 2005, 4, 2257–2265. [Google Scholar] [CrossRef]
- Khalifa, S.M.; Abd El-Aziz, A.M.; Hassan, R.; Abdelmegeed, E.S. β-Lactam Resistance Associated with β-Lactamase Production and Porin Alteration in Clinical Isolates of E. coli and K. pneumoniae. PLoS ONE 2021, 16, e0251594. [Google Scholar] [CrossRef]
- Harder, K.J.; Nikaido, H.; Matsuhashi, M. Mutants of Escherichia coli That Are Resistant to Certain Beta-Lactam Compounds Lack the OmpF Porin. Antimicrob. Agents Chemother. 1981, 20, 549–552. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.W.; Park, S.B.; Im, S.P.; Lee, J.S.; Jung, J.W.; Gong, T.W.; Lazarte, J.M.S.; Kim, J.; Seo, J.-S.; Kim, J.-H.; et al. Outer Membrane Vesicles from β-Lactam-Resistant Escherichia coli Enable the Survival of β-Lactam-Susceptible E. coli in the Presence of β-Lactam Antibiotics. Sci. Rep. 2018, 8, 5402. [Google Scholar] [CrossRef]
- Ude, J.; Tripathi, V.; Buyck, J.M.; Söderholm, S.; Cunrath, O.; Fanous, J.; Claudi, B.; Egli, A.; Schleberger, C.; Hiller, S.; et al. Outer Membrane Permeability: Antimicrobials and Diverse Nutrients Bypass Porins in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2021, 118, e2107644118. [Google Scholar] [CrossRef]
- Wang, P.; Li, R.-Q.; Wang, L.; Yang, W.-T.; Zou, Q.-H.; Xiao, D. Proteomic Analyses of Acinetobacter Baumannii Clinical Isolates to Identify Drug Resistant Mechanism. Front. Cell. Infect. Microbiol. 2021, 11, 625430. [Google Scholar] [CrossRef]
- Hillyer, T.; Benin, B.M.; Sun, C.; Aguirre, N.; Willard, B.; Sham, Y.Y.; Shin, W.S. A Novel Strategy to Characterize the Pattern of β-Lactam Antibiotic-Induced Drug Resistance in Acinetobacter baumannii. Sci. Rep. 2023, 13, 9177. [Google Scholar] [CrossRef]
- Liu, X.; Hu, Y.; Pai, P.-J.; Chen, D.; Lam, H. Label-Free Quantitative Proteomics Analysis of Antibiotic Response in Staphylococcus aureus to Oxacillin. J. Proteome Res. 2014, 13, 1223–1233. [Google Scholar] [CrossRef]
- Li, Z.; He, M.; Dong, X.; Lin, H.; Ge, H.; Shen, S.; Li, J.; Ye, R.D.; Chen, D. New Erythromycin Derivatives Enhance β-Lactam Antibiotics against Methicillin-Resistant Staphylococcus aureus. Lett. Appl. Microbiol. 2015, 60, 352–358. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Pai, P.-J.; Zhang, W.; Hu, Y.; Dong, X.; Qian, P.; Chen, D.; Lam, H. Proteomic Response of Methicillin-Resistant S. aureus to a Synergistic Antibacterial Drug Combination: A Novel Erythromycin Derivative and Oxacillin. Sci. Rep. 2016, 6, 19841. [Google Scholar] [CrossRef]
- Sharma, D.; Garg, A.; Kumar, M.; Khan, A.U. Proteome Profiling of Carbapenem-Resistant K. Pneumoniae Clinical Isolate (NDM-4): Exploring the Mechanism of Resistance and Potential Drug Targets. J. Proteom. 2019, 200, 102–110. [Google Scholar] [CrossRef] [PubMed]
- Becker, B.; Cooper, M.A. Aminoglycoside Antibiotics in the 21st Century. ACS Chem. Biol. 2013, 8, 105–115. [Google Scholar] [CrossRef]
- Allison, K.R.; Brynildsen, M.P.; Collins, J.J. Metabolite-Enabled Eradication of Bacterial Persisters by Aminoglycosides. Nature 2011, 473, 216–220. [Google Scholar] [CrossRef]
- Davies, J.; Wright, G.D. Bacterial Resistance to Aminoglycoside Antibiotics. Trends Microbiol. 1997, 5, 234–240. [Google Scholar] [CrossRef]
- Bryan, L.E.; O’Hara, K.; Wong, S. Lipopolysaccharide Changes in Impermeability-Type Aminoglycoside Resistance in Pseudomonas Aeruginosa. Antimicrob. Agents Chemother. 1984, 26, 250–255. [Google Scholar] [CrossRef]
- Garneau-Tsodikova, S.; Labby, K.J. Mechanisms of Resistance to Aminoglycoside Antibiotics: Overview and Perspectives. Medchemcomm 2016, 7, 11–27. [Google Scholar] [CrossRef]
- Doi, Y.; Wachino, J.; Arakawa, Y. Aminoglycoside Resistance. Infect. Dis. Clin. N. Am. 2016, 30, 523–537. [Google Scholar] [CrossRef]
- Sharma, D.; Gautam, S.; Srivastava, N.; Khan, A.M.; Bisht, D. Comparative Proteomic Analysis of Capsule Proteins in Aminoglycoside-Resistant and Sensitive Mycobacterium tuberculosis Clinical Isolates: Unraveling Potential Drug Targets. Int. J. Mycobacteriol. 2024, 13, 197–205. [Google Scholar] [CrossRef]
- Zhang, D.; Li, H.; Lin, X.; Peng, X. Outer Membrane Proteomics of Kanamycin-Resistant Escherichia coli Identified MipA as a Novel Antibiotic Resistance-Related Protein. FEMS Microbiol. Lett. 2015, 362, fnv074. [Google Scholar] [CrossRef]
- Machin, J.M.; Ranson, N.A.; Radford, S.E. OmpA Specifically Modulates the Activity of Enzymes That Reside in the Crowded Bacterial Outer Membrane. J. Mol. Biol. 2025, 437, 169346. [Google Scholar] [CrossRef]
- Wu, X.; Held, K.; Zheng, C.; Staudinger, B.J.; Chavez, J.D.; Weisbrod, C.R.; Eng, J.K.; Singh, P.K.; Manoil, C.; Bruce, J.E. Dynamic Proteome Response of Pseudomonas aeruginosa to Tobramycin Antibiotic Treatment. Mol. Cell. Proteom. 2015, 14, 2126–2137. [Google Scholar] [CrossRef]
- Ye, J.; Su, Y.; Lin, X.; Lai, S.; Li, W.; Ali, F.; Zheng, J.; Peng, B. Alanine Enhances Aminoglycosides-Induced ROS Production as Revealed by Proteomic Analysis. Front. Microbiol. 2018, 9, 29. [Google Scholar] [CrossRef]
- Soares, N.C.; Spät, P.; Méndez, J.A.; Nakedi, K.; Aranda, J.; Bou, G. Ser/Thr/Tyr Phosphoproteome Characterization of Acinetobacter baumannii: Comparison between a Reference Strain and a Highly Invasive Multidrug-Resistant Clinical Isolate. J. Proteom. 2014, 102, 113–124. [Google Scholar] [CrossRef]
- Pham, T.D.M.; Ziora, Z.M.; Blaskovich, M.A.T. Quinolone Antibiotics. Medchemcomm 2019, 10, 1719–1739. [Google Scholar] [CrossRef]
- Naqvi, S.A.R.; Roohi, S.; Iqbal, A.; Sherazi, T.A.; Zahoor, A.F.; Imran, M. Ciprofloxacin: From Infection Therapy to Molecular Imaging. Mol. Biol. Rep. 2018, 45, 1457–1468. [Google Scholar] [CrossRef]
- Khodursky, A.B.; Zechiedrich, E.L.; Cozzarelli, N.R. Topoisomerase IV Is a Target of Quinolones in Escherichia coli. Proc. Natl. Acad. Sci. USA 1995, 92, 11801–11805. [Google Scholar] [CrossRef]
- Aldred, K.J.; Kerns, R.J.; Osheroff, N. Mechanism of Quinolone Action and Resistance. Biochemistry 2014, 53, 1565–1574. [Google Scholar] [CrossRef]
- Hong, Y.; Li, Q.; Gao, Q.; Xie, J.; Huang, H.; Drlica, K.; Zhao, X. Reactive Oxygen Species Play a Dominant Role in All Pathways of Rapid Quinolone-Mediated Killing. J. Antimicrob. Chemother. 2020, 75, 576–585. [Google Scholar] [CrossRef]
- Kottur, J.; Nair, D.T. Reactive Oxygen Species Play an Important Role in the Bactericidal Activity of Quinolone Antibiotics. Angew. Chem. Int. Ed. 2016, 55, 2397–2400. [Google Scholar] [CrossRef]
- Jacoby, G.A. Mechanisms of Resistance to Quinolones. Clin. Infect. Dis. 2005, 41, S120–S126. [Google Scholar] [CrossRef]
- Lin, X.; Yang, M.; Li, H.; Wang, C.; Peng, X.-X. Decreased Expression of LamB and Odp1 Complex Is Crucial for Antibiotic Resistance in Escherichia coli. J. Proteom. 2014, 98, 244–253. [Google Scholar] [CrossRef]
- Li, W.; Wang, G.; Zhang, S.; Fu, Y.; Jiang, Y.; Yang, X.; Lin, X. An Integrated Quantitative Proteomic and Metabolomics Approach to Reveal the Negative Regulation Mechanism of LamB in Antibiotics Resistance. J. Proteom. 2019, 194, 148–159. [Google Scholar] [CrossRef]
- Ramos, S.; Chafsey, I.; Hebraud, M.; Sousa, M.; Poeta, P.; Igrejas, G. Ciprofloxacin Stress Proteome of the Extended-Spectrum β-Lactamase Producing Escherichia coli from Slaughtered Pigs. Curr. Proteom. 2016, 13, 285–289. [Google Scholar] [CrossRef]
- Su, H.-C.; Ramkissoon, K.; Doolittle, J.; Clark, M.; Khatun, J.; Secrest, A.; Wolfgang, M.C.; Giddings, M.C. The Development of Ciprofloxacin Resistance in Pseudomonas aeruginosa Involves Multiple Response Stages and Multiple Proteins. Antimicrob. Agents Chemother. 2010, 54, 4626–4635. [Google Scholar] [CrossRef] [PubMed]
- Peng, J.; Cao, J.; Ng, F.M.; Hill, J. Pseudomonas Aeruginosa Develops Ciprofloxacin Resistance from Low to High Level with Distinctive Proteome Changes. J. Proteom. 2017, 152, 75–87. [Google Scholar] [CrossRef]
- Machado, I.; Coquet, L. Proteomic Changes in Pseudomonas Aeruginosa Biofilm Cells after Adaptive Resistance Development. J. Proteom. Bioinform. 2016, 9, 58–62. [Google Scholar] [CrossRef]
- Cheng, Z.; Yang, M.-J.; Peng, B.; Peng, X.; Lin, X.; Li, H. The Depressed Central Carbon and Energy Metabolisms Is Associated to the Acquisition of Levofloxacin Resistance in Vibrio alginolyticus. J. Proteom. 2018, 181, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Correia, S.; Hébraud, M.; Chafsey, I.; Chambon, C.; Viala, D.; Torres, C.; de Toro, M.; Capelo, J.L.; Poeta, P.; Igrejas, G. Impacts of Experimentally Induced and Clinically Acquired Quinolone Resistance on the Membrane and Intracellular Subproteomes of Salmonella typhimurium DT104B. J. Proteom. 2016, 145, 46–59. [Google Scholar] [CrossRef]
- Vranakis, I.; De Bock, P.-J.; Papadioti, A.; Tselentis, Y.; Gevaert, K.; Tsiotis, G.; Psaroulaki, A. Identification of Potentially Involved Proteins in Levofloxacin Resistance Mechanisms in Coxiella burnetii. J. Proteome Res. 2011, 10, 756–762. [Google Scholar] [CrossRef]
- Friedman, S.M.; Lu, T.; Drlica, K. Mutation in the DNA Gyrase A Gene of Escherichia coli That Expands the Quinolone Resistance-Determining Region. Antimicrob. Agents Chemother. 2001, 45, 2378–2380. [Google Scholar] [CrossRef] [PubMed]
- Allocati, N.; Federici, L.; Masulli, M.; Di Ilio, C. Glutathione Transferases in Bacteria. FEBS J. 2009, 276, 58–75. [Google Scholar] [CrossRef]
- Li, L.; Wang, W.; Zhang, R.; Xu, J.; Wang, R.; Wang, L.; Zhao, X.; Li, J. First Acetyl-Proteome Profiling of Salmonella Typhimurium Revealed Involvement of Lysine Acetylation in Drug Resistance. Vet. Microbiol. 2018, 226, 1–8. [Google Scholar] [CrossRef]
- Tran, D.T.; Dahlin, A. Multi-Omics Approaches to Resolve Antimicrobial Resistance. In Antimicrobial Resistance: Factors to Findings; Springer International Publishing: Cham, Switzerland, 2024; pp. 275–294. [Google Scholar]
- Zampieri, M.; Szappanos, B.; Buchieri, M.V.; Trauner, A.; Piazza, I.; Picotti, P.; Gagneux, S.; Borrell, S.; Gicquel, B.; Lelievre, J.; et al. High-Throughput Metabolomic Analysis Predicts Mode of Action of Uncharacterized Antimicrobial Compounds. Sci. Transl. Med. 2018, 10, eaal3973. [Google Scholar] [CrossRef]
- Zhou, J.; He, C.; Yang, H.; Shu, W.; Liu, Q. Integrative Omics Analysis Reveals Insights into Small Colony Variants of Staphylococcus aureus Induced by Sulfamethoxazole-Trimethoprim. BMC Microbiol. 2024, 24, 212. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, Z.; Wang, H.-Y.; Chung, C.-R.; Horng, J.-T.; Lu, J.-J.; Lee, T.-Y. Rapid Antibiotic Resistance Serial Prediction in Staphylococcus aureus Based on Large-Scale MALDI-TOF Data by Applying XGBoost in Multi-Label Learning. Front. Microbiol. 2022, 13, 853775. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.-Y.; Hsieh, T.-T.; Chung, C.-R.; Chang, H.-C.; Horng, J.-T.; Lu, J.-J.; Huang, J.-H. Efficiently Predicting Vancomycin Resistance of Enterococcus Faecium From MALDI-TOF MS Spectra Using a Deep Learning-Based Approach. Front. Microbiol. 2022, 13, 821233. [Google Scholar] [CrossRef] [PubMed]
- Weis, C.; Cuénod, A.; Rieck, B.; Dubuis, O.; Graf, S.; Lang, C.; Oberle, M.; Brackmann, M.; Søgaard, K.K.; Osthoff, M.; et al. Direct Antimicrobial Resistance Prediction from Clinical MALDI-TOF Mass Spectra Using Machine Learning. Nat. Med. 2022, 28, 164–174. [Google Scholar] [CrossRef] [PubMed]
- López-Cortés, X.A.; Manríquez-Troncoso, J.M.; Hernández-García, R.; Peralta, D. MSDeepAMR: Antimicrobial Resistance Prediction Based on Deep Neural Networks and Transfer Learning. Front. Microbiol. 2024, 15, 1361795. [Google Scholar] [CrossRef] [PubMed]
- Gao, W.; Li, H.; Yang, J.; Zhang, J.; Fu, R.; Peng, J.; Hu, Y.; Liu, Y.; Wang, Y.; Li, S.; et al. Machine Learning Assisted MALDI Mass Spectrometry for Rapid Antimicrobial Resistance Prediction in Clinicals. Anal. Chem. 2024, 96, 13398–13409. [Google Scholar] [CrossRef]
- Nguyen, H.-A.; Peleg, A.Y.; Song, J.; Antony, B.; Webb, G.I.; Wisniewski, J.A.; Blakeway, L.V.; Badoordeen, G.Z.; Theegala, R.; Zisis, H.; et al. Predicting Pseudomonas aeruginosa Drug Resistance Using Artificial Intelligence and Clinical MALDI-TOF Mass Spectra. mSystems 2024, 9, e0078924. [Google Scholar] [CrossRef]
- Aytan-Aktug, D.; Clausen, P.T.L.C.; Bortolaia, V.; Aarestrup, F.M.; Lund, O. Prediction of Acquired Antimicrobial Resistance for Multiple Bacterial Species Using Neural Networks. mSystems 2020, 5, e00774-19. [Google Scholar] [CrossRef]
- Lundberg, S.; Lee, S.-I. A Unified Approach to Interpreting Model Predictions. In Proceedings of the 31st International Conference on Neural Information Processing Systems, Long Beach, CA, USA, 4–9 December 2017. [Google Scholar]
- Santos, A.; Colaço, A.R.; Nielsen, A.B.; Niu, L.; Strauss, M.; Geyer, P.E.; Coscia, F.; Albrechtsen, N.J.W.; Mundt, F.; Jensen, L.J.; et al. A Knowledge Graph to Interpret Clinical Proteomics Data. Nat. Biotechnol. 2022, 40, 692–702. [Google Scholar] [CrossRef]
- Silcox, C.; Zimlichmann, E.; Huber, K.; Rowen, N.; Saunders, R.; McClellan, M.; Kahn, C.N.; Salzberg, C.A.; Bates, D.W. The Potential for Artificial Intelligence to Transform Healthcare: Perspectives from International Health Leaders. NPJ Digit. Med. 2024, 7, 88. [Google Scholar] [CrossRef]
- Lipworth, S.; Vihta, K.-D.; Chau, K.; Barker, L.; George, S.; Kavanagh, J.; Davies, T.; Vaughan, A.; Andersson, M.; Jeffery, K.; et al. Ten-Year Longitudinal Molecular Epidemiology Study of Escherichia coli and Klebsiella Species Bloodstream Infections in Oxfordshire, UK. Genome Med. 2021, 13, 144. [Google Scholar] [CrossRef] [PubMed]
- Loo, E.X.L.; Zain, A.; Yap, G.C.; Purbojati, R.W.; Drautz-Moses, D.I.; Koh, Y.Q.; Chong, Y.S.; Tan, K.H.; Gluckman, P.D.; Yap, F.; et al. Longitudinal Assessment of Antibiotic Resistance Gene Profiles in Gut Microbiomes of Infants at Risk of Eczema. BMC Infect. Dis. 2020, 20, 312. [Google Scholar] [CrossRef] [PubMed]
- Fang, G.-Y.; Wu, F.-H.; Mu, X.-J.; Jiang, Y.-J.; Liu, X.-Q. Monitoring Longitudinal Antimicrobial Resistance Trends of Staphylococcus aureus Strains Worldwide over the Past 100 Years to Decipher Its Evolution and Transmission. J. Hazard. Mater. 2024, 465, 133136. [Google Scholar] [CrossRef] [PubMed]


| Article | Year | Bacterial Strain(s) | Antibiotic(s) Involved | Key Proteins Identified | Affected Metabolic Pathways |
|---|---|---|---|---|---|
| [112] | 2025 | Escherichia coli | Ciprofloxacin, Enrofloxacin | Proteins linked to SOS response and RecA-independent mechanisms | SOS response, DNA repair, oxidative stress response, nucleotide metabolism |
| [113] | 2025 | Escherichia coli drug sensitive and MDR | Various antibiotics | A total of 763 differentially expressed proteins | Protein biosynthesis, transcription, translation, stress adaptation |
| [114] | 2024 | Staphylococcus aureus MRSA and MSSA | Methicillin | A total of 407 differentially expressed proteins | Fatty acid degradation, glycine, serine, and threonine metabolism |
| [115] | 2023 | Stutzerimonas stutzeri | Chloramphenicol, Minocycline | Multidrug/solvent RND membrane fusion protein, MexE | Membrane transport, antibiotic efflux |
| [116] | 2023 | Pseudomonas aeruginosa | Aztreonam, Carbenicillin, Piperacillin, Tobramycin | Various stress response proteins | Oxidative stress response, protein synthesis, biofilm formation |
| [117] | 2023 | Acinetobacter baumannii | Meropenem | Various metabolic enzymes | Central carbon metabolism, energy production |
| [118] | 2022 | Yersinia pestis, Francisella tularensis | Streptomycin, Gentamicin, Doxycycline | Various differentially expressed proteins | Fatty acid biosynthesis, TCA cycle, purine biosynthesis |
| [119] | 2022 | Vibrio alginolyticus | Serum resistance factors | Mannitol transporters, glycolytic enzymes, pyruvate cycle enzymes, cAMP/CRP | Metabolic pathways involving glycine, serine, threonine, fructose, mannose, and pyruvate, alongside central carbon metabolism and the biosynthesis of amino acids |
| [120] | 2022 | Staphylococcus aureus MRSA | Oxadiazolones | FabH, FphC, AdhE, FphE | Fatty acid biosynthesis pathway, redox and energy metabolism |
| [121] | 2021 | Aeromonas hydrophila | Enoxacin | Multidrug efflux transporters, DNA repair proteins, transcriptional regulators | DNA damage, SOS response, stress response and membrane transport |
| [122] | 2021 | Klebsiella pneumoniae | Cotrimoxazole, Amikacin | GarK, uxaC, exuT, hpaB, fhuA, KPN_01492, fumA, hisC, aroE | TCA cycle, alcohol metabolic process, folate biosynthesis |
| [123] | 2020 | Escherichia coli | β-lactams | A total of 1553 differentially expressed proteins | Purine metabolism, translation, transcription |
| [124] | 2020 | Helicobacter pylori | Daphnetin | Various membrane, repair and stress related proteins | Metabolism, membrane structure, nucleic acid and protein synthesis, ion binding, stress response |
| [125] | 2019 | Staphylococcus xylosus | Tylosin | A total of 155 differentially expressed proteins | Stress response, biosynthesis of amino acids, carbon metabolism, ABC transporters |
| [126] | 2019 | Salmonella Typhimurium, Klebsiella pneumoniae, Staphylococcus aureus | Various antibiotics | PrsA, YadC, FimA, RplB, AcrB, RpoB | Efflux, stress response, energy metabolism and redox processes, translation and transcription machinery |
| [127] | 2018 | Edwardsiella piscicida | Kanamycin | Various outer membrane proteins and type III secretion system related proteins and regulators | Biosynthesis of amino acids, 2-oxicarboloxylic acid metabolism, biosynthesis of secondary metabolites and metabolic pathways |
| [128] | 2016 | Aeromonas hydrophila | Chlortetracycline | Propanoate and fatty acid biosynthesis metabolism-related proteins | Biofilm formation, fatty-acid biosynthesis, propanoate metabolism |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aita, S.E.; Ristori, M.V.; Cristiano, A.; Marfoli, T.; De Cesaris, M.; La Vaccara, V.; Cammarata, R.; Caputo, D.; Spoto, S.; Angeletti, S. Proteomic Insights into Bacterial Responses to Antibiotics: A Narrative Review. Int. J. Mol. Sci. 2025, 26, 7255. https://doi.org/10.3390/ijms26157255
Aita SE, Ristori MV, Cristiano A, Marfoli T, De Cesaris M, La Vaccara V, Cammarata R, Caputo D, Spoto S, Angeletti S. Proteomic Insights into Bacterial Responses to Antibiotics: A Narrative Review. International Journal of Molecular Sciences. 2025; 26(15):7255. https://doi.org/10.3390/ijms26157255
Chicago/Turabian StyleAita, Sara Elsa, Maria Vittoria Ristori, Antonio Cristiano, Tiziana Marfoli, Marina De Cesaris, Vincenzo La Vaccara, Roberto Cammarata, Damiano Caputo, Silvia Spoto, and Silvia Angeletti. 2025. "Proteomic Insights into Bacterial Responses to Antibiotics: A Narrative Review" International Journal of Molecular Sciences 26, no. 15: 7255. https://doi.org/10.3390/ijms26157255
APA StyleAita, S. E., Ristori, M. V., Cristiano, A., Marfoli, T., De Cesaris, M., La Vaccara, V., Cammarata, R., Caputo, D., Spoto, S., & Angeletti, S. (2025). Proteomic Insights into Bacterial Responses to Antibiotics: A Narrative Review. International Journal of Molecular Sciences, 26(15), 7255. https://doi.org/10.3390/ijms26157255









