Molecular Detection and Genetic Diversity of Cytomegaloviruses and Lymphocryptoviruses in Free-Roaming and Captive African Green Monkeys (Chlorocebus sabaeus)
Abstract
1. Introduction
2. Results
2.1. Detection of CMVs and LCVs in AGMs
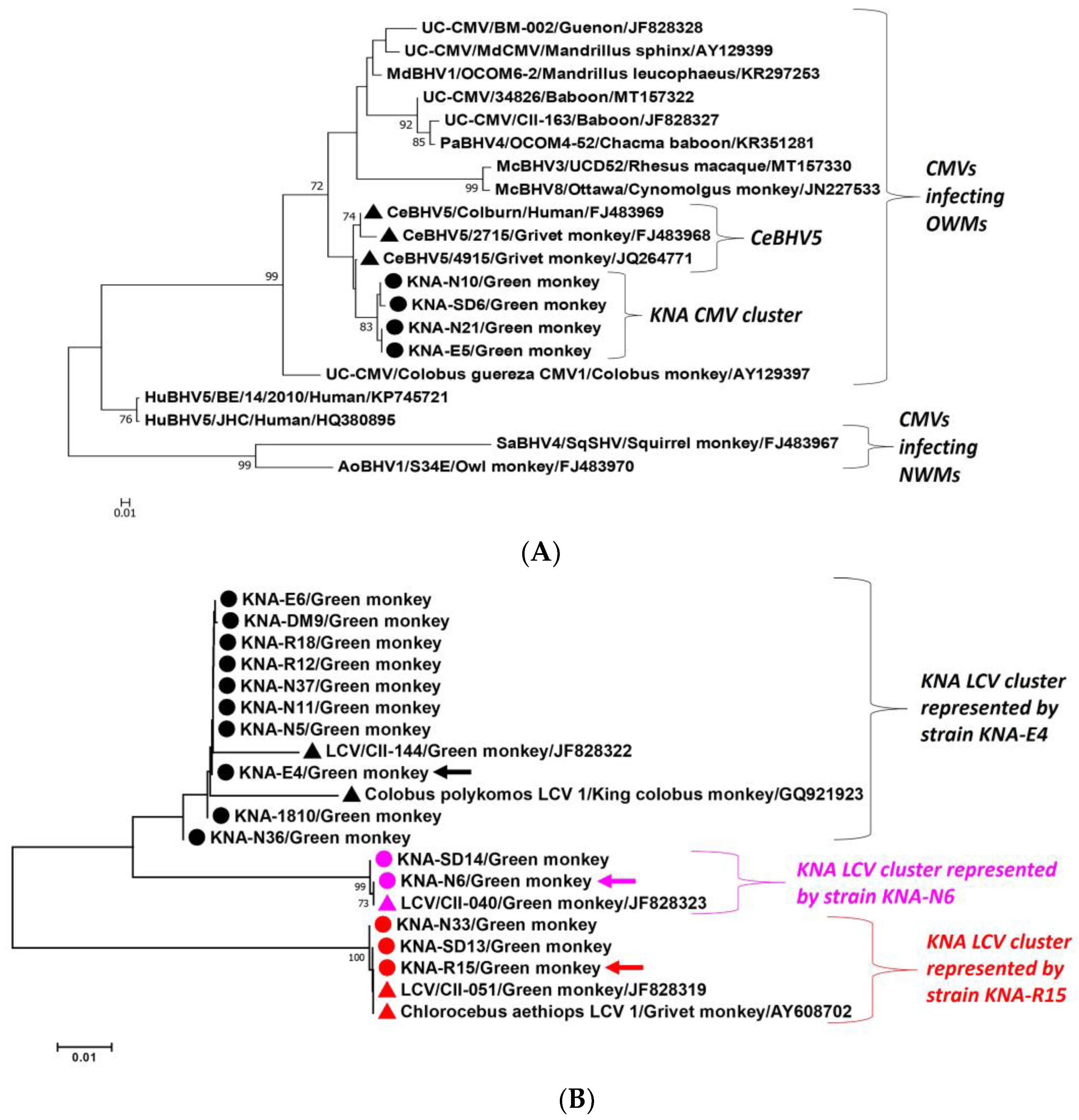
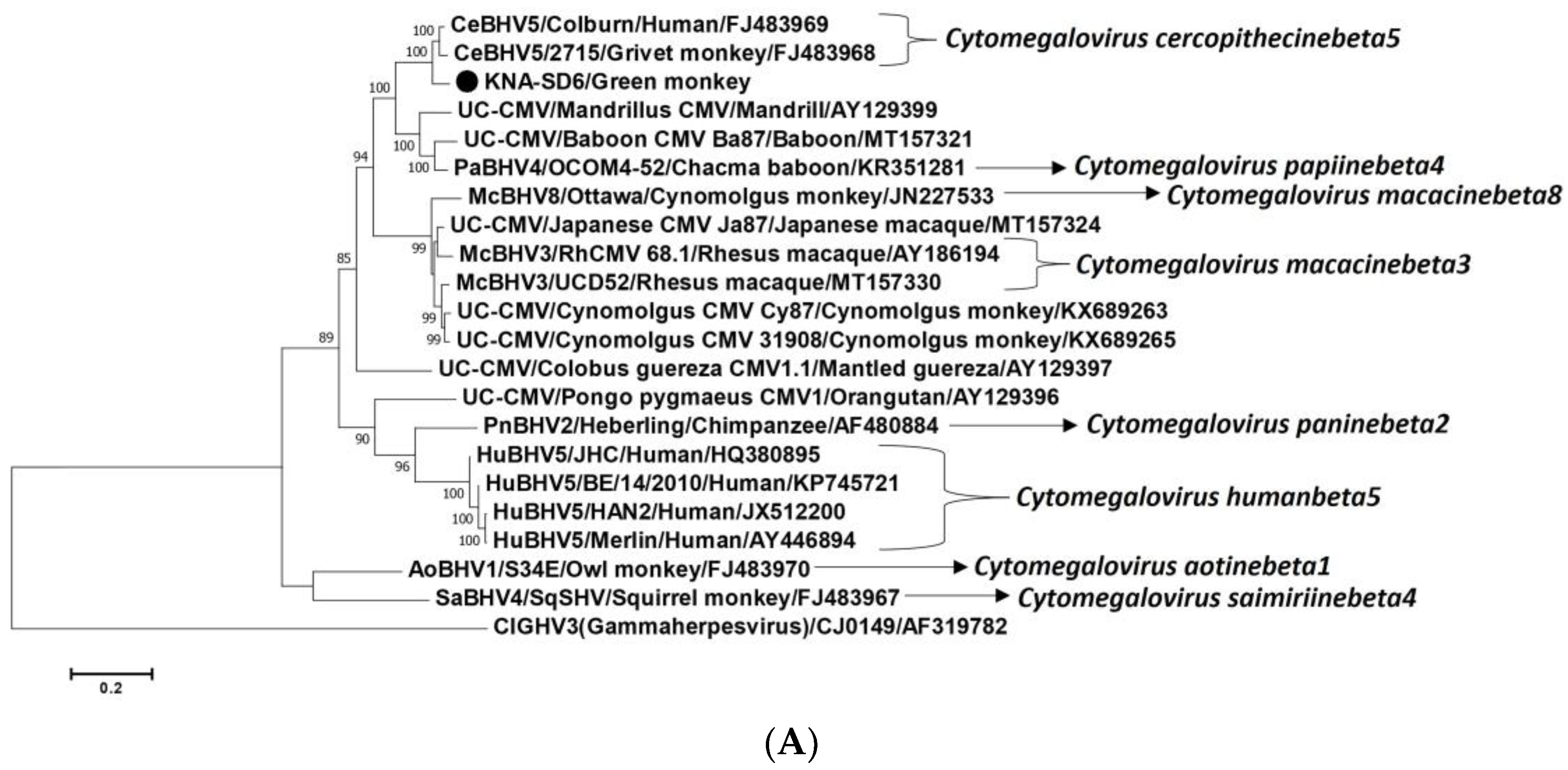
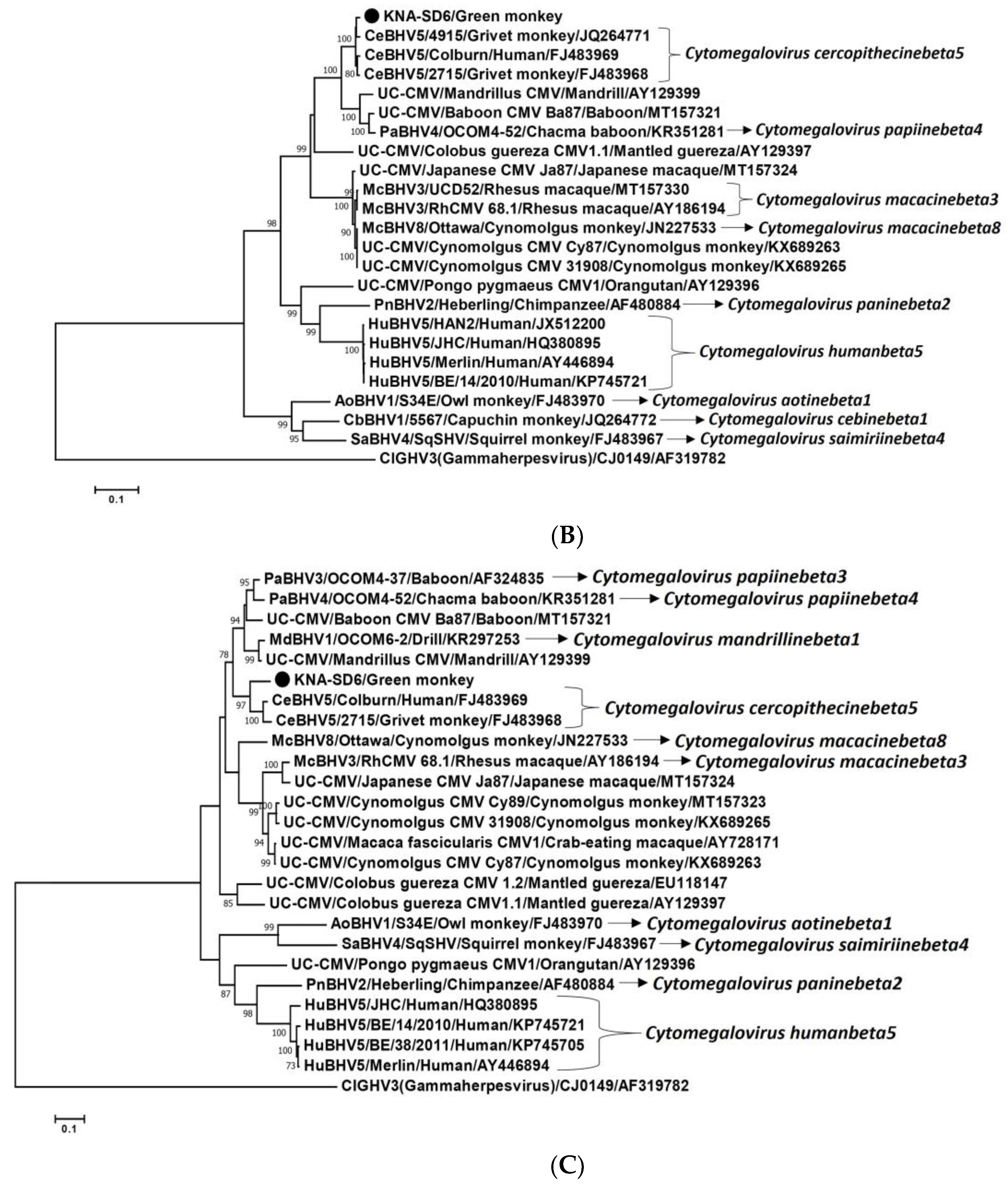
2.2. Analysis of the CMV DPOL and gB Coding Sequences from AGM Sample KNA-SD6
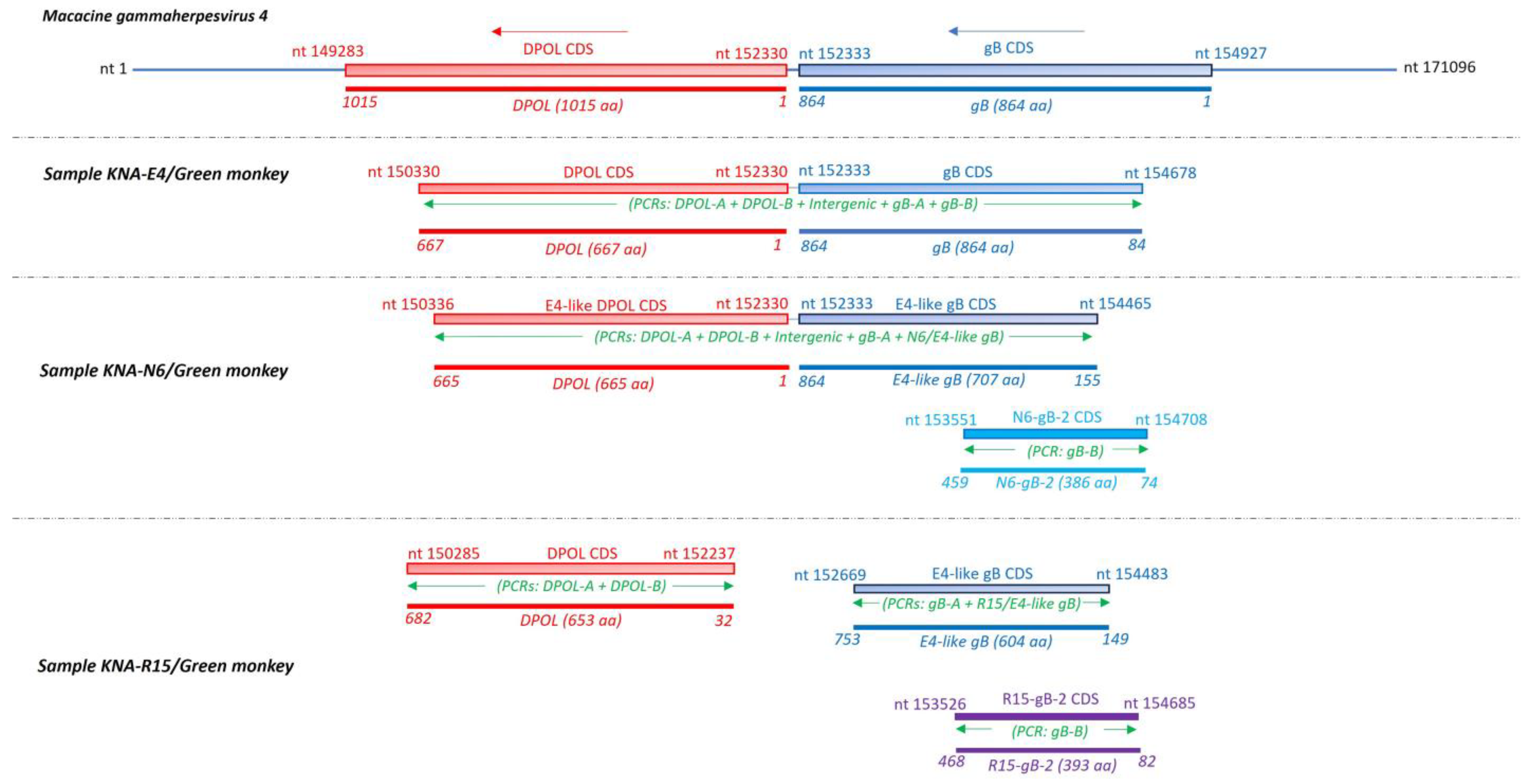
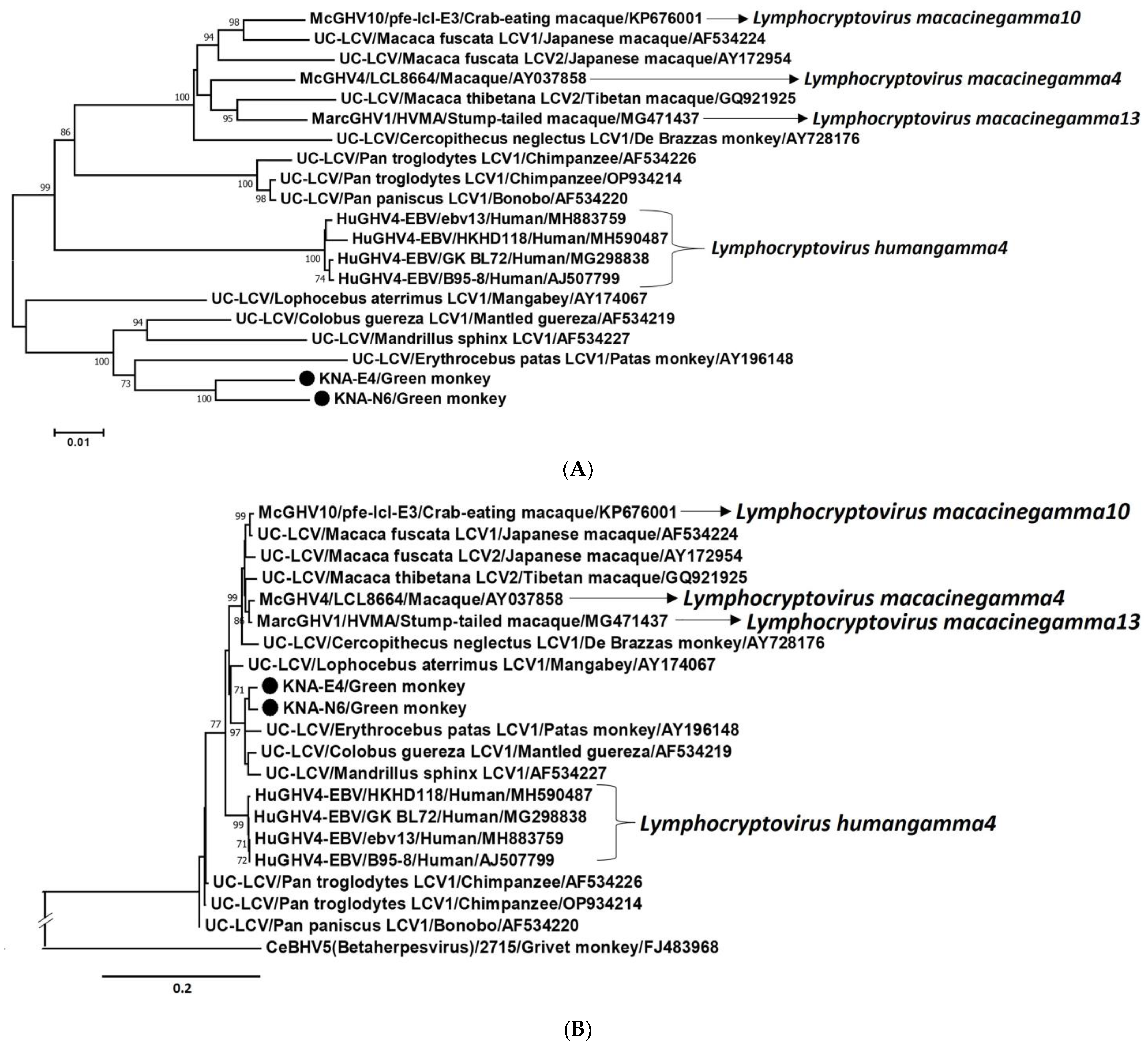
2.3. Analysis of the LCV DPOL and gB Coding Sequences from AGM Samples KNA-E4, -N6 and -R15
3. Discussion
4. Materials and Methods
4.1. Sampling
4.2. Amplification of Herpesvirus DNA
4.3. Nucleotide Sequencing
4.4. Analyses of AGM Herpesvirus Sequences
4.5. GenBank Accession Numbers
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gatherer, D.; Depledge, D.P.; Hartley, C.A.; Szpara, M.L.; Vaz, P.K.; Benkő, M.; Brandt, C.R.; Bryant, N.A.; Dastjerdi, A.; Doszpoly, A.; et al. ICTV Virus Taxonomy Profile: Herpesviridae 2021. J. Gen. Virol. 2021, 102, 001673. [Google Scholar] [CrossRef]
- Cagliani, R.; Forni, D.; Mozzi, A.; Sironi, M. Evolution and Genetic Diversity of Primate Cytomegaloviruses. Microorganisms 2020, 8, 624. [Google Scholar] [CrossRef] [PubMed]
- Anoh, A.E.; Murthy, S.; Akoua-Koffi, C.; Couacy-Hymann, E.; Leendertz, F.H.; Calvignac-Spencer, S.; Ehlers, B. Cytomegaloviruses in a Community of Wild Nonhuman Primates in Taï National Park, Côte D’Ivoire. Viruses 2018, 10, 11. [Google Scholar] [CrossRef] [PubMed]
- Voevodin, A.F.; Marx, P.A., Jr. (Eds.) Cytomegaloviruses. In Simian Virology; Wiley-Blackwell: Ames, IA, USA, 2009; pp. 309–322. [Google Scholar] [CrossRef]
- Wachtman, L.; Mansfield, K. Viral Diseases of Nonhuman Primates. In Nonhuman Primates in Biomedical Research, 2nd ed.; Abee, C.R., Mansfield, K., Tardif, S., Morris, T., Eds.; Academic Press: London, UK, 2012; pp. 1–104. [Google Scholar] [CrossRef]
- Barry, P.A.; William Chang, W.L. Primate betaherpesviruses. In Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis; Arvin, A., Campadelli-Fiume, G., Mocarski, E., Moore, P.S., Roizman, B., Whitley, R., Yamanishi, K., Eds.; Cambridge University Press: Cambridge, UK, 2007; pp. 1051–1075. [Google Scholar]
- Fleckenstein, B.; Ensser, A. Gammaherpesviruses of New World primates. In Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis; Arvin, A., Campadelli-Fiume, G., Mocarski, E., Moore, P.S., Roizman, B., Whitley, R., Yamanishi, K., Eds.; Cambridge University Press: Cambridge, UK, 2007; pp. 1076–1092. [Google Scholar]
- Blossom, D. EBV and KSHV– related herpesviruses in non-human primates. In Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis; Arvin, A., Campadelli-Fiume, G., Mocarski, E., Moore, P.S., Roizman, B., Whitley, R., Yamanishi, K., Eds.; Cambridge University Press: Cambridge, UK, 2007; pp. 1093–1116. [Google Scholar]
- Voevodin, A.F.; Marx, P.A., Jr. (Eds.) Lymphocryptoviruses. In Simian Virology; Wiley-Blackwell: Ames, IA, USA, 2009; pp. 323–346. [Google Scholar] [CrossRef]
- Mühe, J.; Wang, F. Non-human Primate Lymphocryptoviruses: Past, Present, and Future. In Epstein Barr Virus Volume 2; Münz, C., Ed.; Springer: Cham, Switzerland, 2015; pp. 385–405. [Google Scholar]
- Lacoste, V.; Lavergne, A.; de Thoisy, B.; Pouliquen, J.F.; Gessain, A. Genetic diversity and molecular evolution of human and non-human primate Gammaherpesvirinae. Infect. Genet. Evol. 2010, 10, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Taxonomy Browser (Cytomegalovirus). Available online: https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?id=10358 (accessed on 12 January 2024).
- Taxonomy Browser (Lymphocryptovirus). Available online: https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?id=10375 (accessed on 12 January 2024).
- Ohta, E. Pathologic characteristics of infectious diseases in macaque monkeys used in biomedical and toxicologic studies. J. Toxicol. Pathol. 2023, 36, 95–122. [Google Scholar] [CrossRef]
- Fujiwara, S.; Nakamura, H. Animal Models for Gammaherpesvirus Infections: Recent Development in the Analysis of Virus-Induced Pathogenesis. Pathogens 2020, 9, 116. [Google Scholar] [CrossRef] [PubMed]
- Brito, A.F.; Baele, G.; Nahata, K.D.; Grubaugh, N.D.; Pinney, J.W. Intrahost speciations and host switches played an important role in the evolution of herpesviruses. Virus Evol. 2021, 7, veab025. [Google Scholar] [CrossRef]
- Murthy, S.; Couacy-Hymann, E.; Metzger, S.; Nowak, K.; De Nys, H.; Boesch, C.; Wittig, R.; Jarvis, M.A.; Leendertz, F.H.; Ehlers, B. Absence of frequent herpesvirus transmission in a nonhuman primate predator-prey system in the wild. J. Virol. 2013, 87, 10651–10659. [Google Scholar] [CrossRef]
- Murthy, S.; O’Brien, K.; Agbor, A.; Angedakin, S.; Arandjelovic, M.; Ayimisin, E.A.; Bailey, E.; Bergl, R.A.; Brazzola, G.; Dieguez, P.; et al. Cytomegalovirus distribution and evolution in hominines. Virus Evol. 2019, 5, vez015. [Google Scholar] [CrossRef]
- Ehlers, B.; Spiess, K.; Leendertz, F.; Peeters, M.; Boesch, C.; Gatherer, D.; McGeoch, D.J. Lymphocryptovirus phylogeny and the origins of Epstein-Barr virus. J. Gen. Virol. 2010, 91, 630–642. [Google Scholar] [CrossRef]
- Ehlers, B.; Dural, G.; Yasmum, N.; Lembo, T.; de Thoisy, B.; Ryser-Degiorgis, M.P.; Ulrich, R.G.; McGeoch, D.J. Novel mammalian herpesviruses and lineages within the Gammaherpesvirinae: Cospeciation and interspecies transfer. J. Virol. 2008, 82, 3509–3516. [Google Scholar] [CrossRef]
- Gerner, C.S.; Dolan, A.; McGeoch, D.J. Phylogenetic relationships in the Lymphocryptovirus genus of the Gammaherpesvirinae. Virus Res. 2004, 99, 187–192. [Google Scholar] [CrossRef] [PubMed]
- Lavergne, A.; de Thoisy, B.; Pouliquen, J.F.; Ruiz-García, M.; Lacoste, V. Partial molecular characterisation of New World non-human primate lymphocryptoviruses. Infect. Genet. Evol. 2011, 11, 1782–1789. [Google Scholar] [CrossRef] [PubMed]
- James, S.; Donato, D.; Pouliquen, J.F.; Ruiz-García, M.; Lavergne, A.; Lacoste, V. DNA Polymerase Sequences of New World Monkey Cytomegaloviruses: Another Molecular Marker with Which to Infer Platyrrhini Systematics. J. Virol. 2018, 92, e00980-18. [Google Scholar] [CrossRef] [PubMed]
- Leendertz, F.H.; Deckers, M.; Schempp, W.; Lankester, F.; Boesch, C.; Mugisha, L.; Dolan, A.; Gatherer, D.; McGeoch, D.J.; Ehlers, B. Novel cytomegaloviruses in free-ranging and captive great apes: Phylogenetic evidence for bidirectional horizontal transmission. J. Gen. Virol. 2009, 90, 2386–2394. [Google Scholar] [CrossRef]
- McGeoch, D.J.; Gatherer, D.; Dolan, A. On phylogenetic relationships among major lineages of the Gammaherpesvirinae. J. Gen. Virol. 2005, 86, 307–316. [Google Scholar] [CrossRef]
- Phakdeewirot, P.; Payungporn, S.; Chutinimitkul, S.; Theamboonlers, A.; Poovorawan, Y. Prevalence and molecular characterization of the polymerase gene of gibbon lymphocryptovirus. J. Med. Primatol. 2006, 35, 136–143. [Google Scholar] [CrossRef]
- Dore, K. Vervets in the Caribbean. In The International Encyclopedia of Primatology; Fuentes, A., Bezanson, B., Campbell, C.J., DiFiore, A.F., Elton, S., Estrada, A., Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2017; pp. 1–2. [Google Scholar] [CrossRef]
- Gallagher, C.A.; Hervé-Claude, L.P.; Cruz-Martinez, L.; Craig, S. Understanding Community Perceptions of the St. Kitts’ “Monkey Problem” by Adapting Harm Reduction Concepts and Methods. Front. Ecol. Evol. 2022, 10, 904797. [Google Scholar] [CrossRef]
- New England Primate Conservancy. Available online: https://neprimateconservancy.org/Chlorocebus/ (accessed on 12 January 2024).
- Black, P.H.; Hartley, J.W.; Rowe, W.P. Isolation of a cytomegalovirus from African green monkey. Proc. Soc. Exp. Biol. Med. 1963, 112, 601–605. [Google Scholar] [CrossRef]
- Eizuru, Y.; Tsuchiya, K.; Mori, R.; Minamishima, Y. Immunological and molecular comparison of simian cytomegaloviruses isolated from African green monkey (Cercopithecus aethiops) and Japanese macaque (Macaca fuscata). Arch. Virol. 1989, 107, 65–75. [Google Scholar] [CrossRef]
- Minamishima, Y.; Graham, B.J.; Benyesh-Melnick, M. Neutralizing antibodies to cytomegaloviruses in normal simian and human sera. Infect. Immun. 1971, 4, 368–373. [Google Scholar] [CrossRef]
- Swack, N.S.; Liu, O.C.; Hsiung, G.D. Cytomegalovirus infections of monkeys and baboons. Am. J. Epidemiol. 1971, 94, 397–402. [Google Scholar] [CrossRef] [PubMed]
- Smith, K.O.; Thiel, J.F.; Newman, J.T.; Harvey, E.; Trousdale, M.D.; Gehle, W.D.; Clark, G. Cytomegaloviruses as common adventitious contaminants in primary African green monkey kidney cell cultures. J. Natl. Cancer Inst. 1969, 42, 489–496. [Google Scholar] [PubMed]
- Johnson, G.; Dick, D.; Ayers, M.; Petric, M.; Tellier, R. Detection and species-level identification of primate herpesviruses with a comprehensive PCR test for human herpesviruses. J. Clin. Microbiol. 2003, 41, 1256–1258. [Google Scholar] [CrossRef] [PubMed]
- Blewett, E.L.; Lewis, J.; Gadsby, E.L.; Neubauer, S.R.; Eberle, R. Isolation of cytomegalovirus and foamy virus from the drill monkey (Mandrillus leucophaeus) and prevalence of antibodies to these viruses amongst wild-born and captive-bred individuals. Arch. Virol. 2003, 148, 423–433. [Google Scholar] [CrossRef] [PubMed]
- Gerber, P.; Birch, S.M. Complement-fixing antibodies in sera of human and nonhuman primates to viral antigens derived from Burkitt’s lymphoma cells. Proc. Natl. Acad. Sci. USA 1967, 58, 478–484. [Google Scholar] [CrossRef] [PubMed]
- Böcker, J.F.; Tiedemann, K.H.; Bornkamm, G.W.; zur Hausen, H. Characterization of an EBV-like virus from African green monkey lymphoblasts. Virology 1980, 101, 291–295. [Google Scholar] [CrossRef] [PubMed]
- Smith, K.M.; Anthony, S.J.; Switzer, W.M.; Epstein, J.H.; Seimon, T.; Jia, H.; Sanchez, M.D.; Huynh, T.T.; Galland, G.G.; Shapiro, S.E.; et al. Zoonotic viruses associated with illegally imported wildlife products. PLoS ONE 2012, 7, e29505. [Google Scholar] [CrossRef]
- Mancuso, D.M.; Gainor, K.; Dore, K.M.; Gallagher, C.A.; Cruz, K.; Beierschmitt, A.; Malik, Y.S.; Ghosh, S. Detection and Molecular Characterization of Adenoviruses in Captive and Free-Roaming African Green Monkeys (Chlorocebus sabaeus): Evidence for Possible Recombination and Cross-Species Transmission. Viruses 2023, 15, 1605. [Google Scholar] [CrossRef]
- Navarro, R.; Aung, M.S.; Cruz, K.; Ketzis, J.; Gallagher, C.A.; Beierschmitt, A.; Malik, Y.S.; Kobayashi, N.; Ghosh, S. Whole genome analysis provides evidence for porcine-to-simian interspecies transmission of rotavirus-A. Infect. Genet. Evol. 2017, 49, 21–31. [Google Scholar] [CrossRef]
- Richins, T.; Sapp, S.G.H.; Ketzis, J.K.; Willingham, A.L.; Mukaratirwa, S.; Qvarnstrom, Y.; Barratt, J.L.N. Genetic characterization of Strongyloides fuelleborni infecting free-roaming African vervets (Chlorocebus aethiops sabaeus) on the Caribbean island of St. Kitts. Int. J. Parasitol. Parasites Wildl. 2023, 20, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Yao, C.; Walkush, J.; Shim, D.; Cruz, K.; Ketzis, J. Molecular species identification of Trichuris trichiura in African green monkey on St. Kitts, West Indies. Vet. Parasitol. Reg. Stud. Rep. 2018, 11, 22–26. [Google Scholar] [CrossRef]
- Hoefer, A.; Boyen, F.; Beierschmitt, A.; Moodley, A.; Roberts, M.C.; Butaye, P. Methicillin-Resistant and Methicillin-Susceptible Staphylococcus from Vervet Monkeys (Chlorocebus sabaeus) in Saint Kitts. Antibiotics 2021, 10, 290. [Google Scholar] [CrossRef] [PubMed]
- Rajeev, S.; Conan, A.; Pratt, N.; Beierschmitt, A.; Palmour, R. High Leptospira seroprevalence in captive and free-roaming-caught vervet monkeys (Chlorocebus sabeus) on the Caribbean island of Saint Kitts. J. Vet. Diagn. Investig. 2017, 29, 930–934. [Google Scholar] [CrossRef]
- Gallagher, C.; Beierschmitt, A.; Cruz, K.; Choo, J.; Ketzis, J. Should monkeys wash their hands and feet: A pilot-study on sources of zoonotic parasite exposure. One Health 2019, 7, 100088. [Google Scholar] [CrossRef] [PubMed]
- Dore, K.M. Ethnoprimatology without conservation: The political ecology of farmer–green monkey (Chlorocebus sabaeus) relations in St. Kitts, West Indies. Int. J. Primatol. 2018, 39, 918–944. [Google Scholar] [CrossRef]
- Dore, K.M.; Gallagher, C.A.; Mill, A.C. Telemetry-Based Assessment of Home Range to Estimate the Abundance of Invasive Green Monkeys on St. Kitts. Caribb. J. Sci. 2023, 53, 1–17. [Google Scholar] [CrossRef]
- VanDevanter, D.R.; Warrener, P.; Bennett, L.; Schultz, E.R.; Coulter, S.; Garber, R.L.; Rose, T.M. Detection and analysis of diverse herpesviral species by consensus primer PCR. J. Clin. Microbiol. 1996, 34, 1666–1671. [Google Scholar] [CrossRef]
- Carville, A.; Mansfield, K.G. Comparative pathobiology of macaque lymphocryptoviruses. Comp. Med. 2008, 58, 57–67. [Google Scholar]
- Griffiths, P.; Reeves, M. Pathogenesis of human cytomegalovirus in the immunocompromised host. Nat. Rev. Microbiol. 2021, 19, 759–773. [Google Scholar] [CrossRef]
- Smiley Evans, T.; Lowenstine, L.J.; Gilardi, K.V.; Barry, P.A.; Ssebide, B.J.; Kinani, J.F.; Nizeyimana, F.; Noheri, J.B.; Cranfield, M.R.; Mudakikwa, A.; et al. Mountain gorilla lymphocryptovirus has Epstein-Barr virus-like epidemiology and pathology in infants. Sci. Rep. 2017, 7, 5352. [Google Scholar] [CrossRef] [PubMed]
- Katz, B.Z.; Berkman, A.B.; Shapiro, E.D. Serologic evidence of active Epstein-Barr virus infection in Epstein-Barr virus-associated lymphoproliferative disorders of children with acquired immunodeficiency syndrome. J. Pediatr. 1992, 120, 228–232. [Google Scholar] [CrossRef] [PubMed]
- Andiman, W.A.; Eastman, R.; Martin, K.; Katz, B.Z.; Rubinstein, A.; Pitt, J.; Pahwa, S.; Miller, G. Opportunistic lymphoproliferations associated with Epstein-Barr viral DNA in infants and children with AIDS. Lancet 1985, 2, 1390–1393. [Google Scholar] [CrossRef] [PubMed]
- Niazi, M.R.; Iqbal, Q.Z.; Mishiyev, D.; Narula, N.; Abdul Sattar, S.B.; Zia, Z.; Haider, M.A.; Chalhoub, M. Epstein-Barr virus (EBV) induced pneumonitis in an immunocompetent adult: A case report. Respir. Med. Case Rep. 2020, 31, 101262. [Google Scholar] [CrossRef]
- Prepens, S.; Kreuzer, K.A.; Leendertz, F.; Nitsche, A.; Ehlers, B. Discovery of herpesviruses in multi-infected primates using locked nucleic acids (LNA) and a bigenic PCR approach. Virol. J. 2007, 4, 84. [Google Scholar] [CrossRef]
- Shao, Y.; Zhou, L.; Li, F.; Zhao, L.; Zhang, B.L.; Shao, F.; Chen, J.W.; Chen, C.Y.; Bi, X.; Zhuang, X.L.; et al. Phylogenomic analyses provide insights into primate evolution. Science 2023, 380, 913–924. [Google Scholar] [CrossRef] [PubMed]
- Furusato, I.N.; Figueiredo, K.B.; de Carvalho, A.C.S.R.; da Silva Ferreira, C.S.; Takahashi, J.P.F.; Kimura, L.M.; Aleixo, C.S.; de Brito, O.P.; Luchs, A.; Cunha, M.S.; et al. Detection of herpesviruses in neotropical primates from São Paulo, Brazil. Braz. J. Microbiol. 2023, 54, 3201–3209. [Google Scholar] [CrossRef]
- Seeber, P.A.; Greenwood, A.D. Noninvasive Monitoring of Herpes Viruses. In Fowler’ s Zoo and Wild Animal Medicine Current Therapy, Volume 10; Miller, E., Lamberski, N., Calle, P., Eds.; Elsevier: St. Louis, MO, USA, 2023; pp. 175–180. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Martin, D.P.; Murrell, B.; Golden, M.; Khoosal, A.; Muhire, B. RDP4: Detection and Analysis of Recombination Patterns in Virus Genomes. Virus Evol. 2015, 1, vev003. [Google Scholar] [CrossRef]
- Mello, B. Estimating TimeTrees with MEGA and the TimeTree Resource. Mol. Biol. Evol. 2018, 35, 2334–2342. [Google Scholar] [CrossRef]
- Kumar, S.; Suleski, M.; Craig, J.M.; Kasprowicz, A.E.; Sanderford, M.; Li, M.; Stecher, G.; Hedges, S.B. TimeTree 5: An Expanded Resource for Species Divergence Times. Mol. Biol. Evol. 2022, 39, msac174. [Google Scholar] [CrossRef]
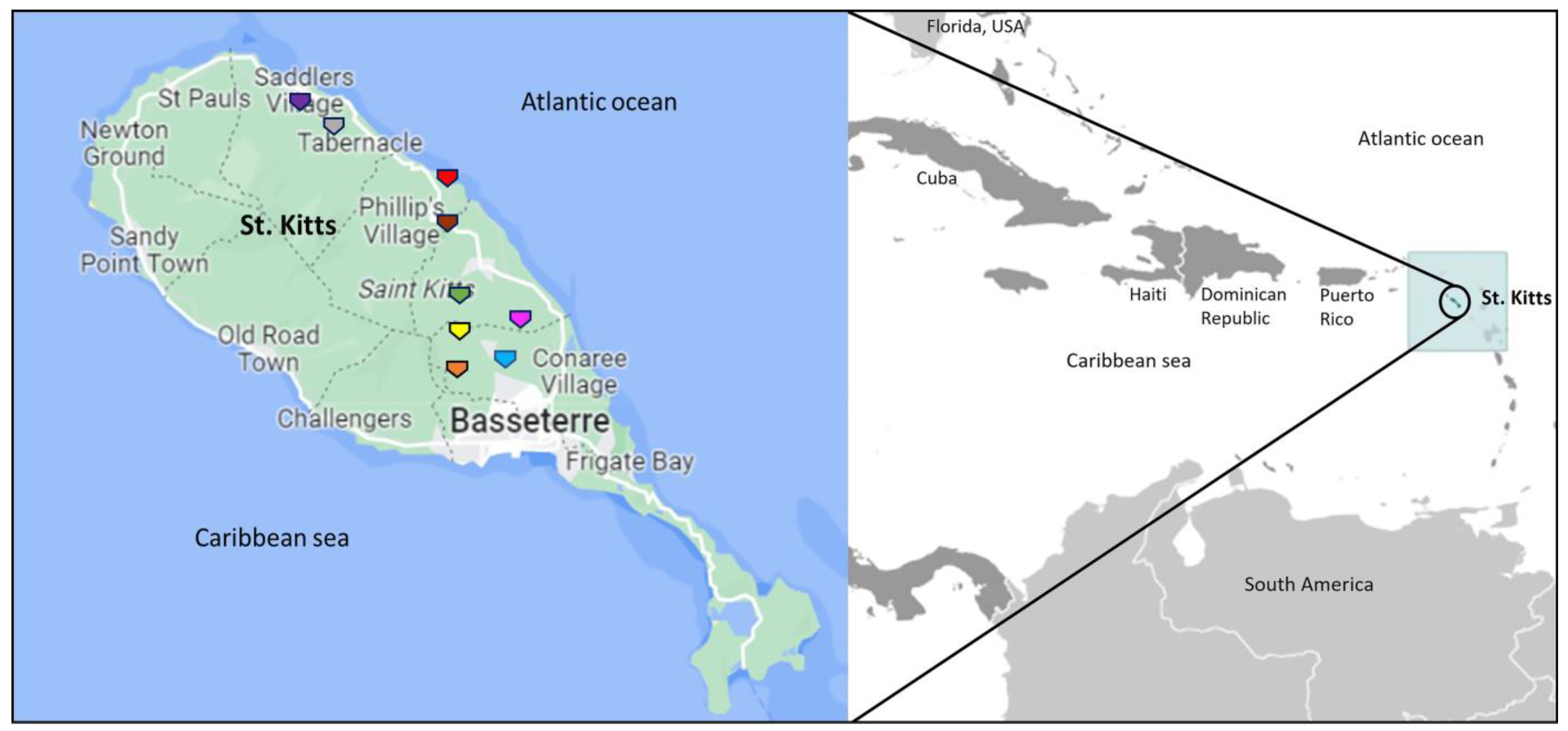


| AGM/Sample Number | Age, Gender | Herpesvirus-Positive Sample | Date of Sample Collection | Sampling/Trapping Location in St. Kitts | Genus 1 |
|---|---|---|---|---|---|
| KNA-E4 2 | Juvenile, female | Nasal swab | 19 Oct 2020 | Behavioral Science Foundation 3 | Lymphocryptovirus |
| KNA-E5 4 | Not available | Nasal swab | 22 Sep 2020 | Behavioral Science Foundation 3 | Cytomegalovirus |
| KNA-E5-O 4 | Not available | Oral discharge | 22 Sep 2020 | Behavioral Science Foundation 3 | Lymphocryptovirus |
| KNA-E6 2 | Juvenile, female | Nasal swab | 21 Sep 2020 | Behavioral Science Foundation 3 | Lymphocryptovirus |
| KNA-R2 | Juvenile, male | Rectal swab | 24 Sep 2022 | Monkey hill 5,6 | Sequence lacked quality 7 |
| KNA-R3 | Adult, female | Rectal swab | 24 Sep 2022 | Monkey hill 5,6 | Sequence lacked quality 7 |
| KNA-N5 | Juvenile, male | Nasal swab | 24 Sep 2022 | Monkey hill 5,6 | Lymphocryptovirus |
| KNA-N6 | Adult, female | Nasal swab | 24 Sep 2022 | Monkey hill 5,6 | Lymphocryptovirus |
| KNA-N10 | Adult, female | Nasal swab | 15 Dec 2022 | Monkey hill 5,6 | Cytomegalovirus |
| KNA-N11 | Adult, female | Nasal swab | 15 Dec 2022 | Monkey hill 5,6 | Lymphocryptovirus |
| KNA-R12 | Juvenile, female | Rectal swab | 15 Dec 2022 | Monkey hill 5,6 | Lymphocryptovirus |
| KNA-R15 | Juvenile, male | Rectal swab | 15 Dec 2022 | Monkey hill 5,6 | Lymphocryptovirus |
| KNA-R18 | Adult, male | Rectal swab | 19 Dec 2022 | Cedar grove 6,8 | Lymphocryptovirus |
| KNA-N21 | Adult, male | Nasal swab | 19 Dec 2022 | Cedar grove 6,8 | Cytomegalovirus |
| KNA-N32 | Adult, male | Nasal swab | 29 Dec 2022 | Green hill 6,8 | Sequence lacked quality 7 |
| KNA-N33 | Adult, male | Nasal swab | 29 Dec 2022 | Green hill 6,8 | Lymphocryptovirus |
| KNA-N36 | Adult, male | Nasal swab | 29 Dec 2022 | Green hill 6,8 | Lymphocryptovirus |
| KNA-N37 | Adult, male | Nasal swab | 29 Dec 2022 | Green hill 6,8 | Lymphocryptovirus |
| KNA-N44 9 | Adult, male | Nasal swab | 10 Jan 2023 | Fountain mountain 8,10 | Sequence lacked quality 7 |
| KNA-R44 9 | Adult, male | Rectal swab | 10 Jan 2023 | Fountain mountain 8,10 | Sequence lacked quality 7 |
| KNA-R49 | Adult, female | Rectal swab | 10 Jan 2023 | Fountain mountain 8,10 | Sequence lacked quality 7 |
| KNA-18-2R | Adult, female | Rectal swab | 18 Jan 2023 | Harris mountain forest 8,10 | Sequence lacked quality 7 |
| KNA-1810 | Adult, male | Nasal swab | 18 Jan 2023 | Harris mountain forest 8,10 | Lymphocryptovirus |
| KNA-DM9 | Juvenile, female | Nasal swab | 24 Jan 2023 | Dale mountain 6,8 | Lymphocryptovirus |
| KNA-SD6 | Adult, female | Nasal swab | 1 Feb 2023 | Saddlers 6,8 | Cytomegalovirus |
| KNA-SD13 | Adult, female | Nasal swab | 1 Feb 2023 | Saddlers 6,8 | Lymphocryptovirus |
| KNA-SD14 | Adult, female | Nasal swab | 1 Feb 2023 | Saddlers 6,8 | Lymphocryptovirus |
| KNA-SD15 | Adult, female | Nasal swab | 1 Feb 2023 | Saddlers 6,8 | Sequence lacked quality 7 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mancuso, D.M.; Gainor, K.; Dore, K.M.; Gallagher, C.A.; Beierschmitt, A.; Malik, Y.S.; Ghosh, S. Molecular Detection and Genetic Diversity of Cytomegaloviruses and Lymphocryptoviruses in Free-Roaming and Captive African Green Monkeys (Chlorocebus sabaeus). Int. J. Mol. Sci. 2024, 25, 3272. https://doi.org/10.3390/ijms25063272
Mancuso DM, Gainor K, Dore KM, Gallagher CA, Beierschmitt A, Malik YS, Ghosh S. Molecular Detection and Genetic Diversity of Cytomegaloviruses and Lymphocryptoviruses in Free-Roaming and Captive African Green Monkeys (Chlorocebus sabaeus). International Journal of Molecular Sciences. 2024; 25(6):3272. https://doi.org/10.3390/ijms25063272
Chicago/Turabian StyleMancuso, Diana M., Kerry Gainor, Kerry M. Dore, Christa A. Gallagher, Amy Beierschmitt, Yashpal S. Malik, and Souvik Ghosh. 2024. "Molecular Detection and Genetic Diversity of Cytomegaloviruses and Lymphocryptoviruses in Free-Roaming and Captive African Green Monkeys (Chlorocebus sabaeus)" International Journal of Molecular Sciences 25, no. 6: 3272. https://doi.org/10.3390/ijms25063272
APA StyleMancuso, D. M., Gainor, K., Dore, K. M., Gallagher, C. A., Beierschmitt, A., Malik, Y. S., & Ghosh, S. (2024). Molecular Detection and Genetic Diversity of Cytomegaloviruses and Lymphocryptoviruses in Free-Roaming and Captive African Green Monkeys (Chlorocebus sabaeus). International Journal of Molecular Sciences, 25(6), 3272. https://doi.org/10.3390/ijms25063272







