Genome-Wide Analysis of Long Noncoding RNAs in Porcine Intestine during Weaning Stress
Abstract
1. Introduction
2. Results
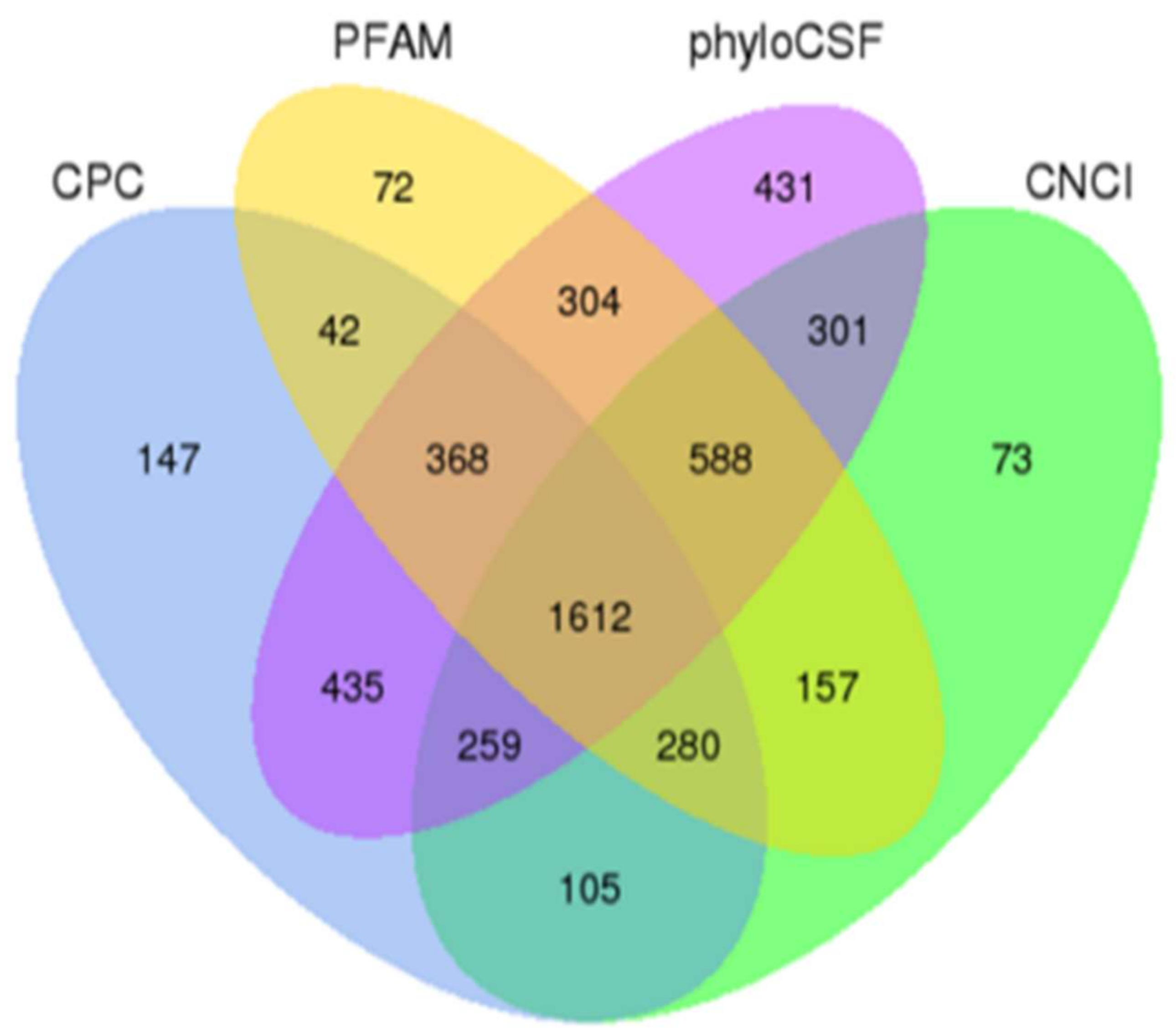
2.1. Identification of lncRNAs in Piglet Intestine
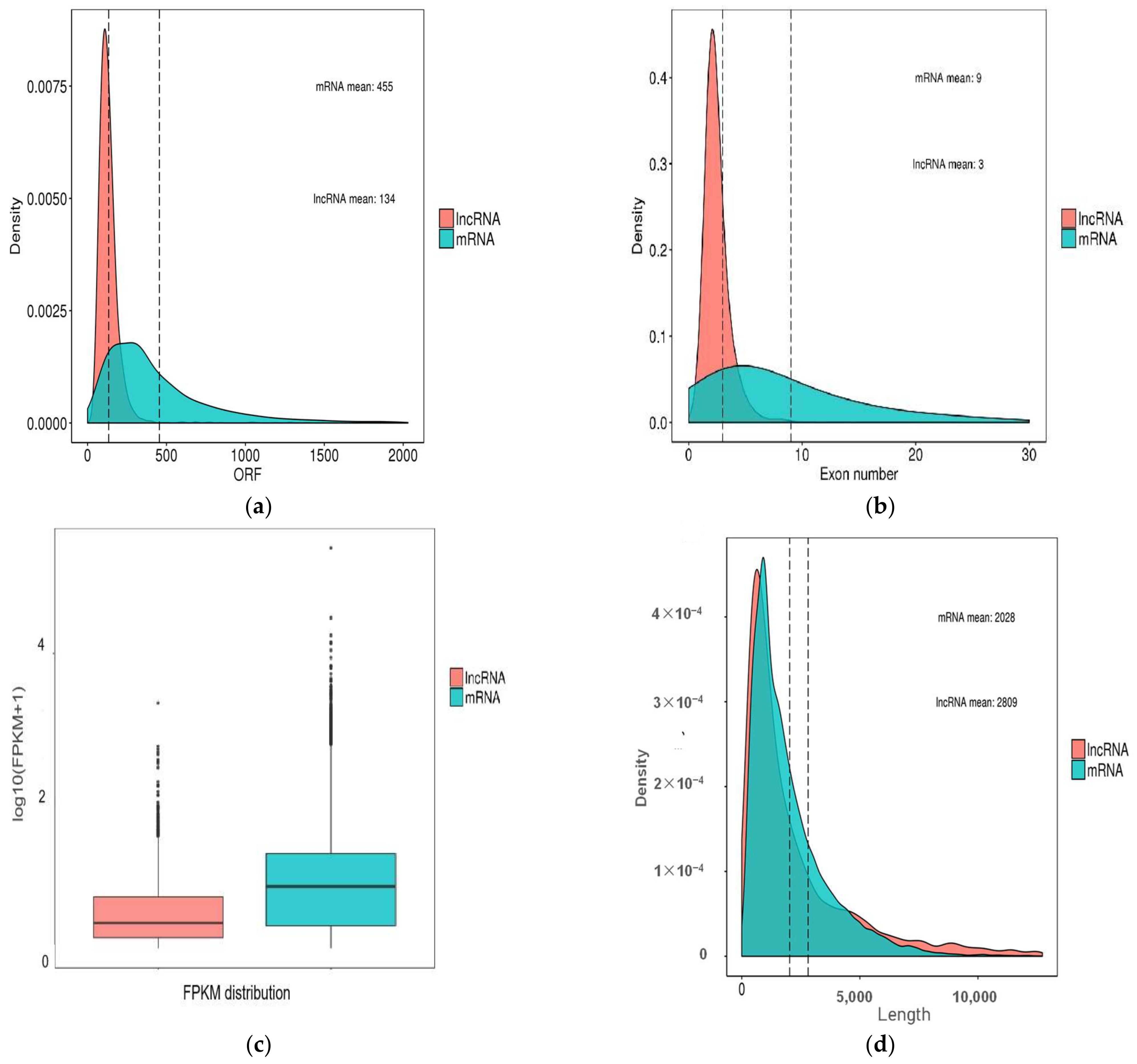
2.2. Characteristics of Intestinal lncRNA
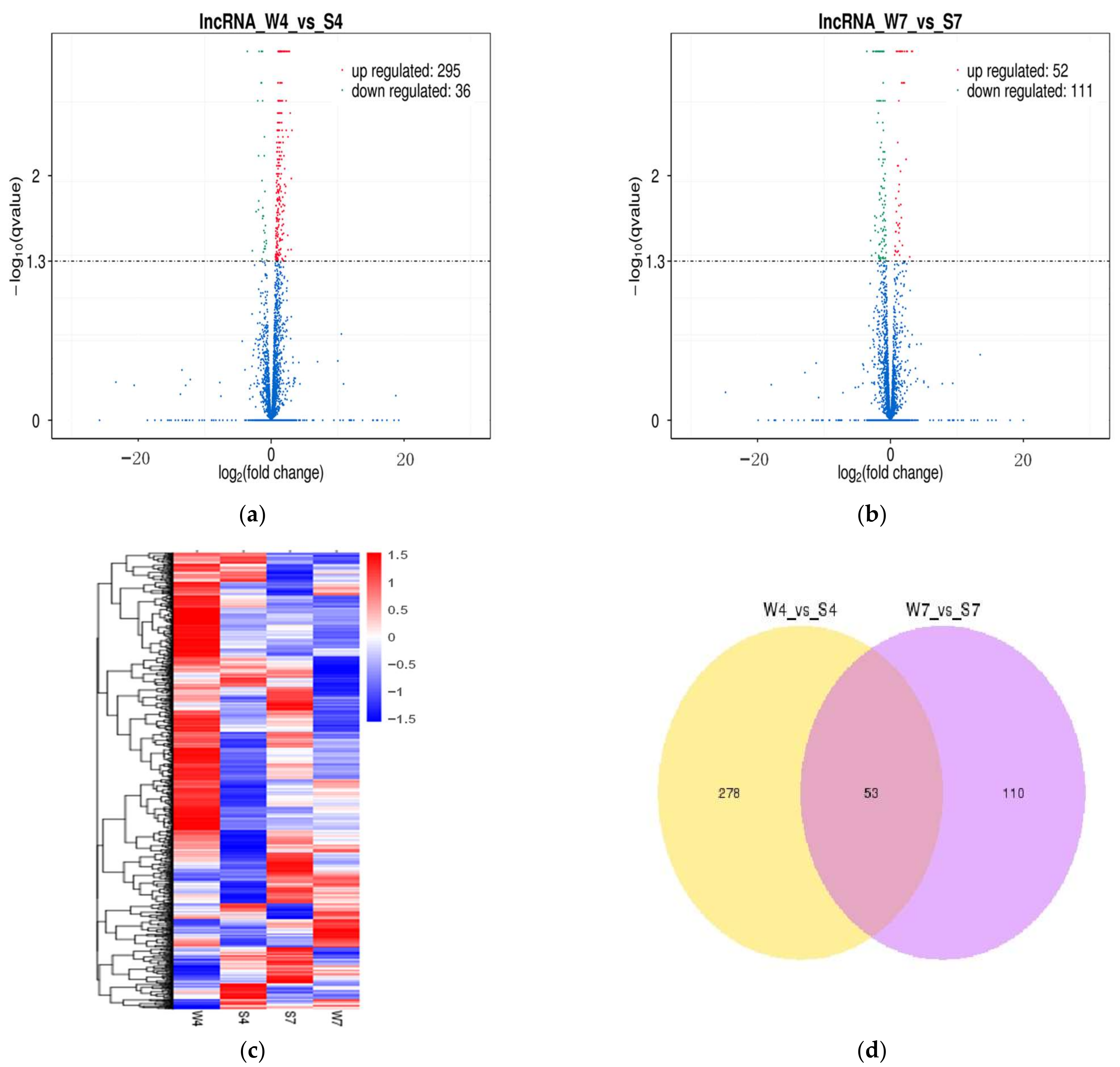
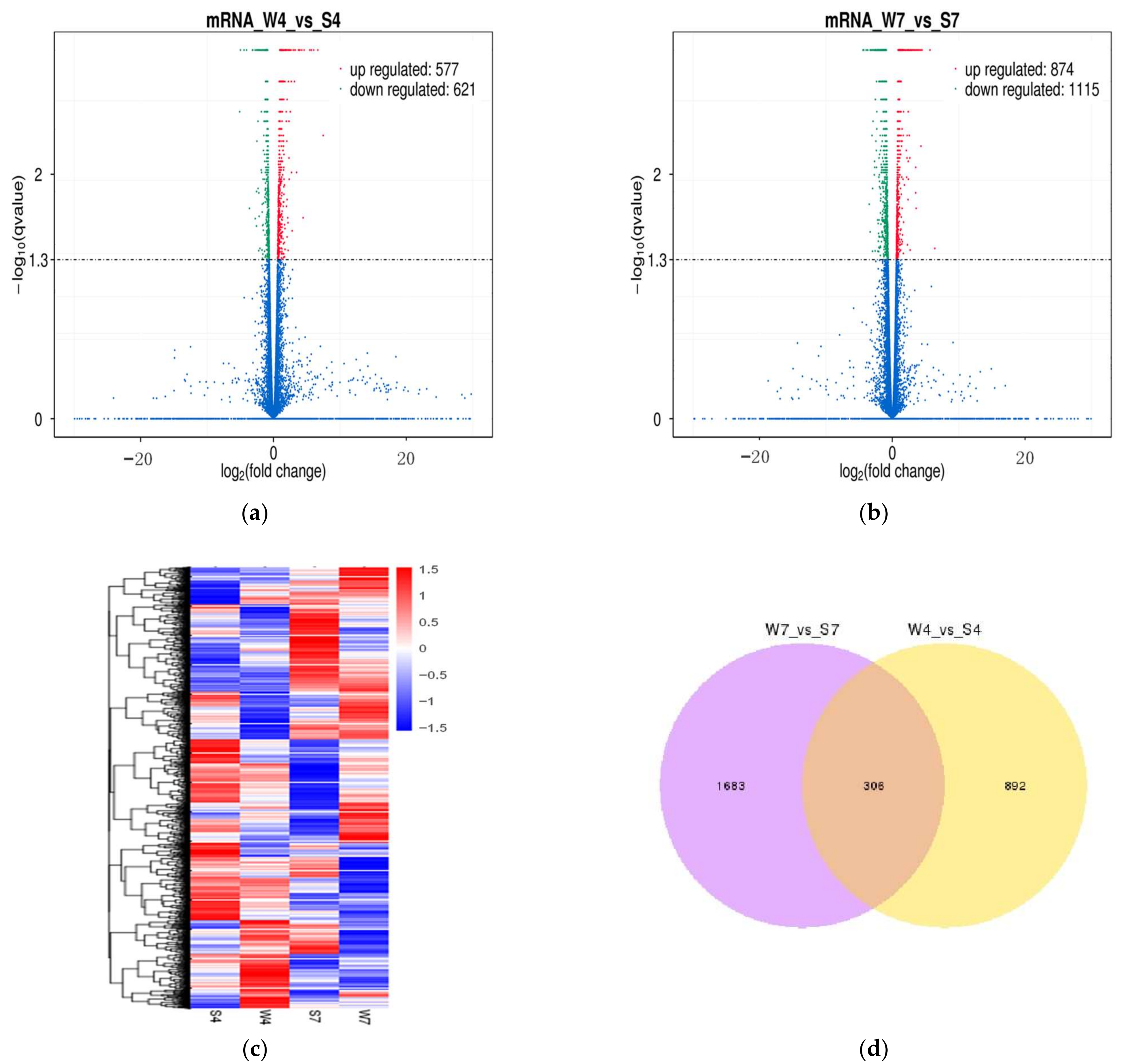
2.3. Analysis of Differentially Expressed lncRNAs and mRNAs
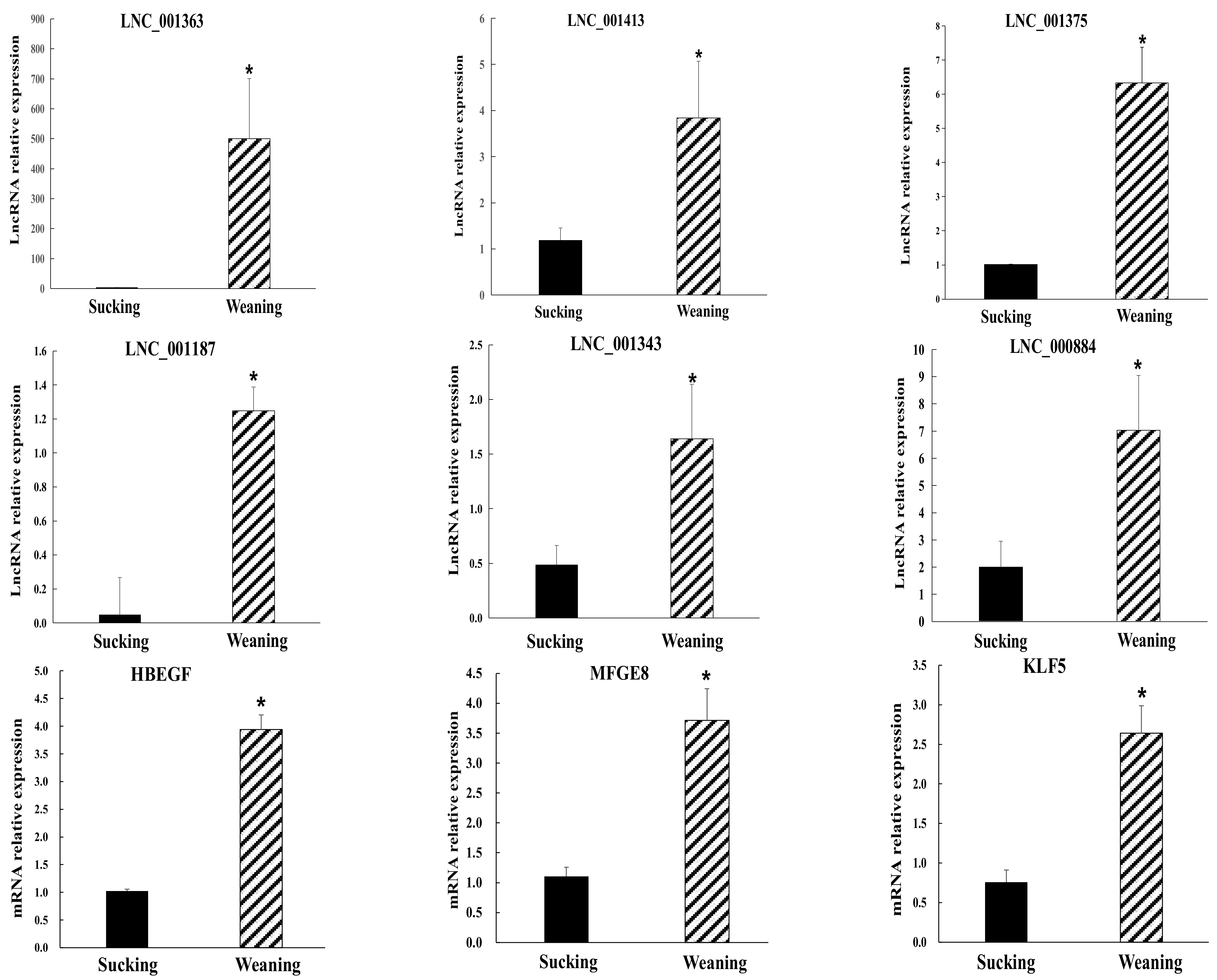
2.4. Validation of Differentially Expressed lncRNAs and mRNA
2.5. Cis Role of lncRNAs in Target Genes
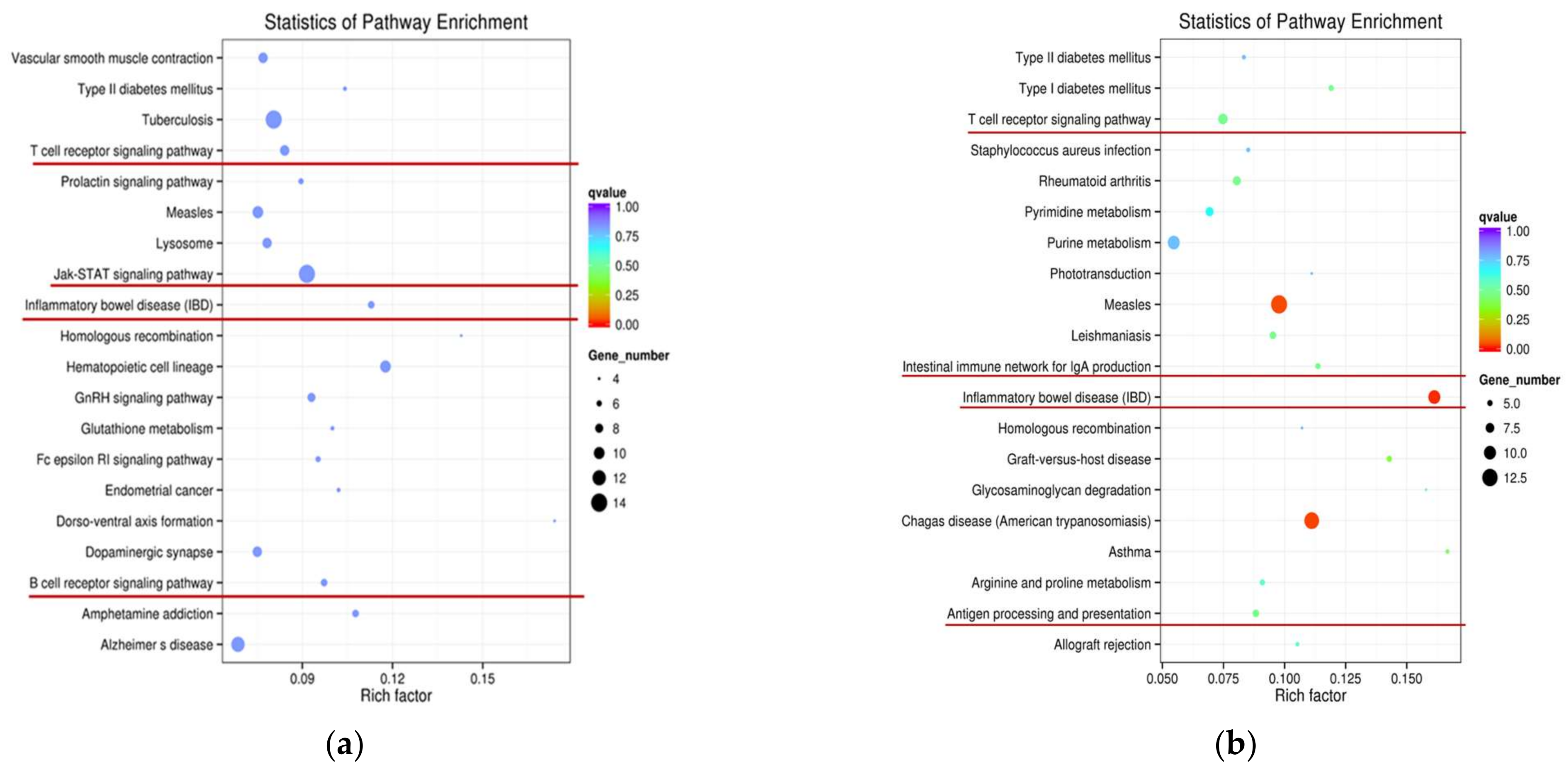
2.6. Identification of Functional Specificities of lncRNAs
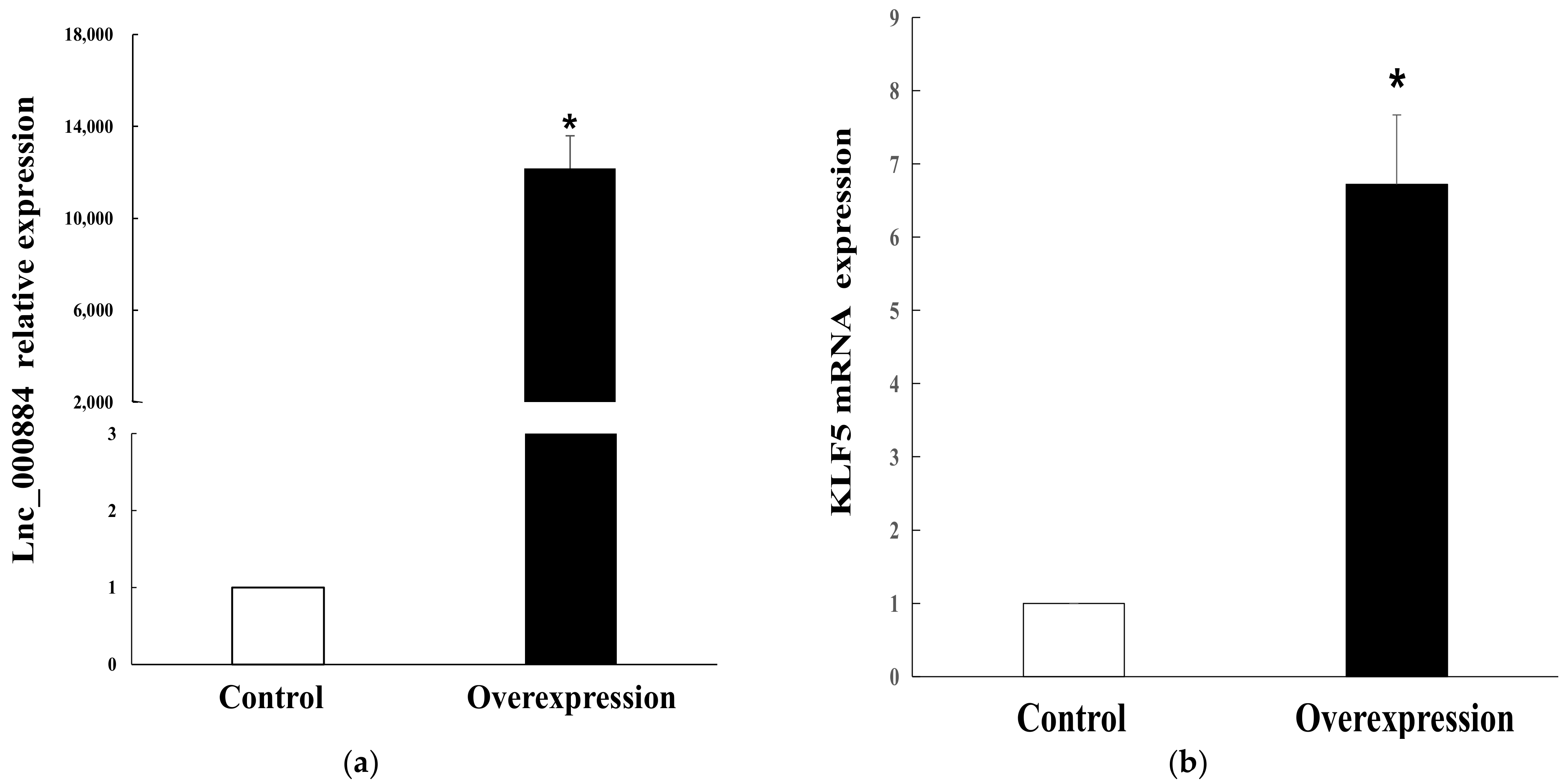
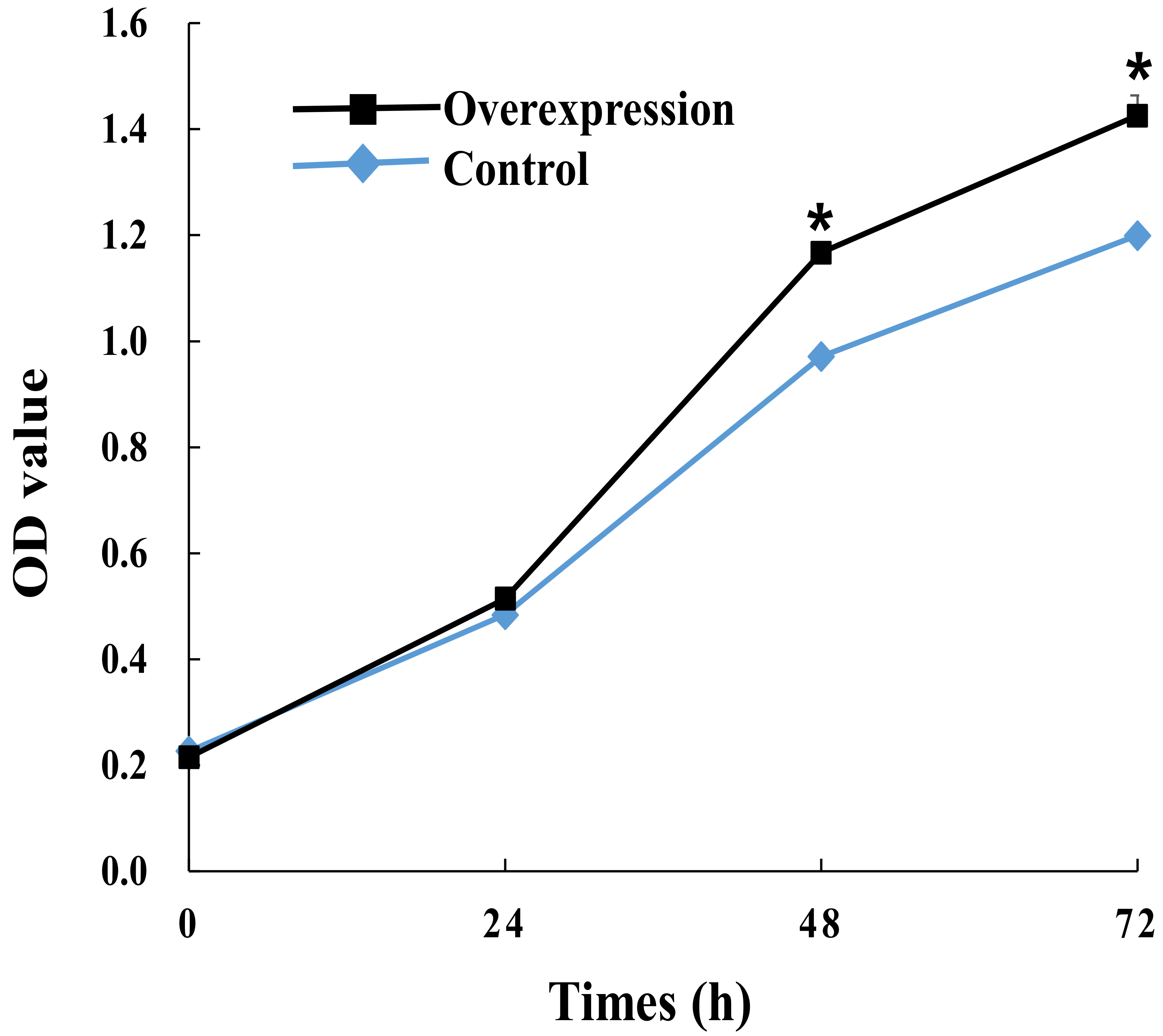
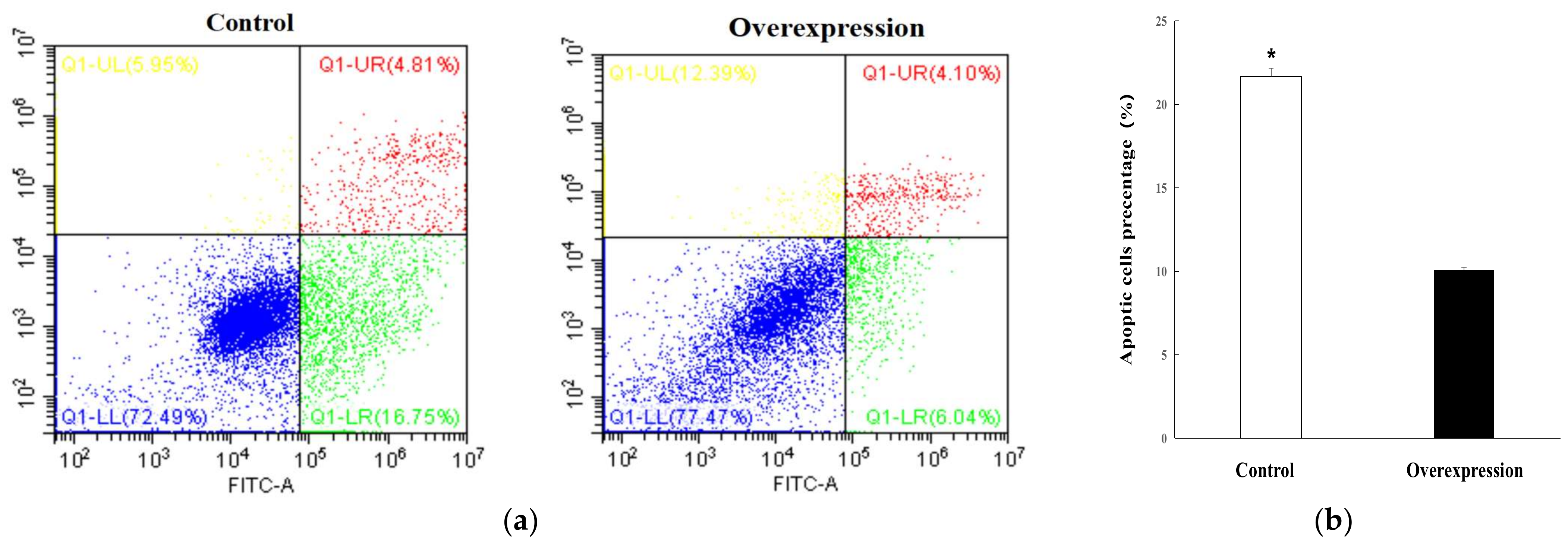
2.7. Lnc_000884 Regulatory Role on Intestinal Epithelial Cell
3. Discussion
4. Materials and Methods
4.1. Experimental Animals and Sample Preparation
4.2. RNA Isolation and Qualification
4.3. Library Preparation for lncRNA Sequencing
4.4. Quality Control and Transcriptome Assembly
4.5. Coding Potential and Expression Analysis of Transcriptome
4.6. Target Gene Prediction and Functional Enrichment Analysis
4.7. Quantitative Real-Time RT-PCR
4.8. Construction of lnc_000884 Overexpression Vector and Transfection of Cells
4.9. Cell Proliferation and Apoptosis Detection
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cao, S.T.; Wang, C.C.; Wu, H.; Zhang, Q.H.; Jiao, L.F.; Hu, C.H. Weaning disrupts intestinal antioxidant status, impairs intestinal barrier and mitochondrial function, and triggers mitophagy in piglets. J. Anim. Sci. 2018, 96, 1073–1083. [Google Scholar] [CrossRef] [PubMed]
- Modina, S.C.; Polito, U.; Rossi, R.; Corino, C.; Giancamillo, D.A. Nutritional Regulation of Gut Barrier Integrity in Weaning Piglets. Animals 2019, 9, 1045. [Google Scholar] [CrossRef] [PubMed]
- Smith, F.; Clark, J.E.; Overman, B.L.; Tozel, C.C.; Huang, J.H.; Rivier, J.E.F.; Blikslager, A.T.; Moeser, A.J. Early weaning stress impairs development of mucosal barrier function in the porcine intestine. Am. J. Physiol. Gastrointest. Liver Physiol. 2010, 298, G352–G363. [Google Scholar] [CrossRef]
- Gu, X.H.; Li, D.; She, R.P. Effect of weaning on small intestinal structure and function in the piglet. Arch. Tierernahr. 2002, 56, 275–286. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.T.; Liu, L.; Long, S.F.; Pan, L.; Piao, X.S. Effect of organic acids and essential oils on performance, intestinal health and digestive enzyme activities of weaned pigs. Anim. Feed Sci. Tech. 2018, 235, 110–119. [Google Scholar] [CrossRef]
- Kim, K.K.; Song, M.H.; Liu, Y.H.; Ji, P. Enterotoxigenic Escherichia coli infection of weaned pigs: Intestinal challenges and nutritional intervention to enhance disease resistance. Front. Immunol. 2022, 5, 885253. [Google Scholar] [CrossRef]
- Gao, J.; Gao, K.; Yang, X.; Jiang, Z.; Wang, L. Intestinal morphology and immune profiles are altered in piglets by early-weaning. Int. Immunopharmacol. 2022, 105, 108520. [Google Scholar]
- Mei, J.; Xu, R.J. Transient changes of transforming growth factor-beta expression in the small intestine of the pig in association with weaning. Br. J. Nutr. 2005, 93, 37–45. [Google Scholar] [CrossRef]
- Tao, X.; Xu, Z. MicroRNA transcriptome in swine small intestine during weaning stress. PLoS ONE 2013, 18, e79343. [Google Scholar] [CrossRef]
- Kopp, F.; Mendell, J.T. Functional classification and experimental dissection of long noncoding RNAs. Cell 2018, 172, 393–407. [Google Scholar] [CrossRef]
- Dykes, I.M.; Emanueli, C. Transcriptional and Post-transcriptional Gene Regulation by Long Non-coding RNA. Genom. Proteom. Bioinf. 2017, 15, 177–186. [Google Scholar] [CrossRef]
- Cao, H.L.; Liu, Z.J.; Huang, P.L.; Yue, Y.L.; Xi, J.N. lncRNA-RMRP promotes proliferation, migration and invasion of bladder cancer via miR-206. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 1012–1021. [Google Scholar] [PubMed]
- Wan, J.; Chen, P.; Zhang, Y.; Ding, J.; Yang, Y.; Li, X. Identification of the 11-lncRNA signatures associated with the prognosis of endometrial carcinoma. Sci. Prog. 2021, 104, 368504211006593. [Google Scholar] [CrossRef]
- Smolle, M.A.; Prinz, F.; Calin, G.A.; Pichler, M. Current concepts of non-coding RNA regulation of immune checkpoints in cancer. Mol. Aspects. Med. 2019, 70, 117–126. [Google Scholar] [CrossRef] [PubMed]
- Zheng, P.; Huang, C.; Leng, D.; Sun, B.; Zhang, X.D. Transcriptome analysis of peripheral whole blood identifies crucial lncRNAs implicated in childhood asthma. BMC Med. Genom. 2020, 13, 136. [Google Scholar] [CrossRef]
- Chand, J.U.; Nayyar, H.; Mantri, N.; Siddique, K.H.M. Non-Coding RNAs in Legumes: Their Emerging Roles in Regulating Biotic/Abiotic Stress Responses and Plant Growth and Development. Cells 2021, 10, 1674. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Li, M.; Bo, X.; Li, T.; Ma, L.; Zhai, T.; Huang, T. Systematic Analysis of Long Non-Coding RNAs and mRNAs in the Ovaries of Duroc Pigs During Different Follicular Stages Using RNA Sequencing. Int. J. Mol. Sci. 2018, 19, 1722. [Google Scholar] [CrossRef]
- Credendino, S.C.; Lewin, N.; Oliveira, M.; Basu, S.; D’Andrea, B.; Amendola, E.; Guida, L.D.; Nardone, A.; Sanges, R.; Felic, M.D.; et al. Tissue- and Cell Type-Specific Expression of the Long Noncoding RNA Klhl14-AS in Mouse. Int. J. Genom. 2017, 2017, 9769171. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.; Xia, X.; Jiang, B.; Ma, K.; Zhu, L.; Wang, L.; Jin, B. Identification and characterization of novel lncRNAs in Arabidopsis thaliana. Biochem. Biophys. Res. Commun. 2017, 488, 348–354. [Google Scholar] [CrossRef]
- Liu, Y.; Yu, Y.; Ao, H.; Zhang, F.; Zhao, X.; Liu, H.; Shi, Y.; Xing, K.; Wang, C. Identification of Long Non-Coding RNAs Involved in Porcine Fat Deposition Using Two High-Throughput Sequencing Methods. Genes 2021, 12, 1374. [Google Scholar] [CrossRef]
- Zhang, X.; Chen, M.; Liu, X.; Zhang, L.; Ding, X.; Guo, Y.; Li, X.; Guo, H. A novel lncRNA, lnc403, involved in bovine skeletal muscle myogenesis by mediating KRAS/Myf6. Gene. 2020, 751, 144706. [Google Scholar] [CrossRef] [PubMed]
- Ren, H.; Wang, G.; Chen, L.; Jiang, J.; Liu, L.; Li, N.; Zhao, J.; Sun, X.; Zhou, P. Genome-wide analysis of long non-coding RNAs at early stage of skin pigmentation in goats (Capra hircus). BMC Genom. 2016, 17, 67. [Google Scholar] [CrossRef]
- Dai, M.; Chen, X.; Mo, S.; Li, J.; Huang, Z.; Huang, S.; Xu, J.; He, B.; Zou, Y.; Chen, J.; et al. Meta-signature LncRNAs serve as novel biomarkers for colorectal cancer: Integrated bioinformatics analysis, experimental validation and diagnostic evaluation. Sci. Rep. 2017, 7, 46572. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.N.; Joung, J.; Bae, J.S.; Lee, C.S.; Koo, J.S.; Park, S.J.; Im, J.P.; Kim, Y.S.; Kim, J.W.; Park, W.Y.; et al. RNA-seq Reveals Transcriptomic Differences in Inflamed and Noninflamed Intestinal Mucosa of Crohn’s Disease Patients Compared with Normal Mucosa of Healthy Controls. Inflamm. Bowel. Dis. 2017, 23, 1098–1108. [Google Scholar] [CrossRef]
- Wu, F.; Huang, Y.; Dong, F.; Kwon, J.H. Ulcerative Colitis-Associated Long Noncoding RNA, BC012900, Regulates Intestinal Epithelial Cell Apoptosis. Inflamm. Bowel. Dis. 2016, 22, 782–795. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Mu, Y.; Ma, L.; Wang, C.; Tang, Z.; Yang, S.; Zhou, R.; Hu, X.; Li, M.H.; Li, K. Systematic identification and characterization of long intergenic non-coding RNAs in fetal porcine skeletal muscle development. Sci. Rep. 2015, 5, 8957. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xue, S.; Liu, X.; Liu, H.; Hu, T.; Qiu, X.; Zhang, J.; Lei, M. Analyses of Long Non-Coding RNA and mRNA profiling using RNA sequencing during the pre-implantation phases in pig endometrium. Sci. Rep. 2016, 6, 20238. [Google Scholar] [CrossRef]
- Ran, M.; Chen, B.; Li, Z.; Wu, M.; Liu, X.; He, C.; Zhang, S.; Li, Z. Systematic Identification of Long Noncoding RNAs in Immature and Mature Porcine Testes. Biol. Reprod. 2016, 94, 77. [Google Scholar] [CrossRef]
- Valadkhan, S.; Valencia-Hipólito, A. lncRNAs in Stress Response. Curr. Top. Microbiol. Immunol. 2016, 394, 203–236. [Google Scholar]
- Ibeagha-Awemu, E.M.; Do, D.N.; Dudemaine, P.L.; Fomenky, B.E.; Bissonnette, N. Integration of lncRNA and mRNA Transcriptome Analyses Reveals Genes and Pathways Potentially Involved in Calf Intestinal Growth and Development during the Early Weeks of Life. Genes 2018, 9, 142. [Google Scholar] [CrossRef]
- Li, N.; Shi, R. Expression alteration of long non-coding RNAs and their target genes in the intestinal mucosa of patients with Crohn’s disease. Clin. Chim. Acta 2019, 494, 14–21. [Google Scholar] [CrossRef]
- Lu, Q.; Gong, W.; Wang, J.; Ji, K.; Sun, X.; Xu, C.; Du, L.; Wang, Y.; Liu, Q. Analysis of changes to lncRNAs and their target mRNAs in murine jejunum after radiation treatment. J. Cell Mol. Med. 2018, 22, 6357–6367. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhang, C.; Zhang, N.; Liu, G. Porcine endemic diarrhea virus infection regulates long noncoding RNA expression. Virology 2019, 527, 89–97. [Google Scholar] [CrossRef]
- Chung, E.Y.; Psathas, J.N.; Yu, D.; Li, Y.; Weiss, M.J.; Thomas-Tikhonenko, A. CD19 is a major B cell receptor-independent activator of MYC-driven B-lymphomagenesis. J. Clin. Invest. 2012, 122, 2257–2266. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Cao, J.; Li, C.; Zhang, L. Chemokine CXCL13 expression was up-regulated in Clostridium difficile infection. Cytokine 2016, 88, 232–240. [Google Scholar] [CrossRef] [PubMed]
- Dörner, T.; Shock, A.; Smith, K.G. CD22 and autoimmune disease. Int. Rev. Immunol. 2012, 31, 363–378. [Google Scholar] [CrossRef]
- Caruso, R.; Fina, D.; Peluso, I.; Stolfi, C.; Fantini, M.C.; Gioia, V.; Caprioli, F.; Blanco, G.D.V.; Paoluzi, O.A.; Macdonald, T.T.; et al. A functional role for interleukin-21 in promoting the synthesis of the T-cell chemoattractant, MIP-3alpha, by gut epithelial cells. Gastroenterology 2007, 132, 166–175. [Google Scholar] [CrossRef]
- Jia, X.; Yuan, Y.; Yao, Z.; Wu, C.; Chen, X.; Chen, X.; Lin, Y.; Meng, X.; Zeng, X.; Shao, J. Association between IL-4 and IL-4R Polymorphisms and Periodontitis: A Meta-Analysis. Dis. Markers. 2017, 2017, 8021279. [Google Scholar] [CrossRef]
- Renz, H. Soluble interleukin-4 receptor (sIL-4R) in allergic diseases. Inflamm. Res. 1999, 48, 425–431. [Google Scholar] [CrossRef]
- Grilli, E.; Tugnoli, B.; Passey, J.L.; Stahl, C.H.; Piva, A.; Moeser, A.J. Impact of dietary organic acids and botanicals on intestinal integrity and inflammation in weaned pigs. BMC Vet. Res. 2015, 11, 96. [Google Scholar] [CrossRef]
- Hyun, J.G.; Mayer, L. Mechanisms underlying inflammatory bowel disease. Drug Discov. Today (Dis. Mech.) 2006, 3, 457–462. [Google Scholar] [CrossRef]
- Rawlings, J.S.; Rosler, K.M.; Harrison, D.A. The JAK/STAT signaling pathway. J. Cell Sci. 2004, 117, 1281–1283. [Google Scholar] [CrossRef] [PubMed]
- Jere, S.W.; Abrahamse, H.; Houreld, N.N. The JAK/STAT signaling pathway and photobiomodulation in chronic wound healing. Cytokine Growth Factor Rev. 2017, 38, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Jeldres, T.; Tyler, C.J.; Boyer, J.D.; Karuppuchamy, T.; Yarur, A.; Giles, D.A.; Yeasmin, S.; Lundborg, L.; Sandborn, W.J.; Patel, D.R.; et al. Targeting Cytokine Signaling and Lymphocyte Traffic via Small Molecules in Inflammatory Bowel Disease: JAK Inhibitors and S1PR Agonists. Front Pharmacol. 2019, 10, 212. [Google Scholar] [CrossRef]
- Blikslager, A.T.; Moeser, A.J.; Gookin, J.L.; Jones, S.L.; Odle, J. Restoration of barrier function in injured intestinal mucosa. Physiol. Rev. 2007, 87, 545–564. [Google Scholar] [CrossRef]
- Shi, L.; Han, X.; Li, J.X.; Liao, Y.T.; Kou, F.S.; Wang, Z.B.; Shi, R.; Zhao, X.J.; Sun, Z.M.; Hao, Y. Identification of differentially expressed genes in ulcerative colitis and verification in a colitis mouse model by bioinformatics analyses. World J. Gastroenterol. 2020, 26, 5983–5996. [Google Scholar] [CrossRef] [PubMed]
- Bu, H.; Zuo, X.; Wang, X.; Ensslin, M.A.; Koti, V.; Hsueh, W.; Raymond, A.S.; Shur, B.D.; Tan, X. Milk fat globule-EGF factor 8/lactadherin plays a crucial role in maintenance and repair of murine intestinal epithelium. J. Clin. Invest. 2007, 117, 3673–3683. [Google Scholar] [CrossRef]
- Zhang, Y.; Brenner, M.; Yang, W.L.; Wang, P. Recombinant human MFG-E8 ameliorates colon damage in DSS- and TNBS-induced colitis in mice. Lab. Invest. 2015, 95, 480–490. [Google Scholar] [CrossRef]
- Aziz, M.M.; Ishihara, S.; Mishima, Y.; Oshima, N.; Moriyama, I.; Yuki, T.; Kadowaki, Y.; Rumi, M.A.K.; Amano, Y.; Kinoshita, Y. MFG-E8 attenuates intestinal inflammation in murine experimental colitis by modulating osteopontin-dependent alphavbeta3 integrin signaling. J. Immunol. 2009, 182, 7222–7232. [Google Scholar] [CrossRef]
- McConnell, B.B.; Kim, S.S.; Yu, K.; Ghaleb, A.M.; Takeda, N.; Manabe, I.; Nusrat, A.; Nagai, R.; Yang, V.W. Krüppel-like factor 5 is important for maintenance of crypt architecture and barrier function in mouse intestine. Gastroenterology 2011, 141, 1302–1313.e1-e6. [Google Scholar] [CrossRef]
- Bell, K.N.; Shroyer, N.F. Krüpple-like factor 5 is required for proper maintenance of adult intestinal crypt cellular proliferation. Dig. Dis. Sci. 2015, 60, 86–100. [Google Scholar] [CrossRef] [PubMed]
- Tetreault, M.; Alrabaa, R.; McGeehan, M.; Katz, J.P. Krüppel-like factor 5 protects against murine colitis and activates JAK-STAT signaling in vivo. PLoS ONE 2012, 7, e38338. [Google Scholar] [CrossRef]
- McConnell, B.B.; Klapproth, J.A.; Sasaki, M.; Nandan, M.O.; Yang, V.W. Krüppel-like factor 5 mediates transmissible murine colonic hyperplasia caused by Citrobacter rodentium infection. Gastroenterology 2008, 134, 1007–1016. [Google Scholar] [CrossRef] [PubMed]
- McConnell, B.B.; Kim, S.S.; Bialkowska, A.B.; Yu, K.; Sitaraman, S.V.; Yang, V.W. Krüppel-like factor 5 protects against dextran sulfate sodium-induced colonic injury in mice by promoting epithelial repair. Gastroenterology 2011, 140, 540–549. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, S.; Tao, X.; Deng, B.; Li, Y.; Xu, Z. Genome-Wide Analysis of Long Noncoding RNAs in Porcine Intestine during Weaning Stress. Int. J. Mol. Sci. 2023, 24, 5343. https://doi.org/10.3390/ijms24065343
Liu S, Tao X, Deng B, Li Y, Xu Z. Genome-Wide Analysis of Long Noncoding RNAs in Porcine Intestine during Weaning Stress. International Journal of Molecular Sciences. 2023; 24(6):5343. https://doi.org/10.3390/ijms24065343
Chicago/Turabian StyleLiu, Shujie, Xin Tao, Bo Deng, Yongming Li, and Ziwei Xu. 2023. "Genome-Wide Analysis of Long Noncoding RNAs in Porcine Intestine during Weaning Stress" International Journal of Molecular Sciences 24, no. 6: 5343. https://doi.org/10.3390/ijms24065343
APA StyleLiu, S., Tao, X., Deng, B., Li, Y., & Xu, Z. (2023). Genome-Wide Analysis of Long Noncoding RNAs in Porcine Intestine during Weaning Stress. International Journal of Molecular Sciences, 24(6), 5343. https://doi.org/10.3390/ijms24065343



