Antimicrobial Evaluation of New Pyrazoles, Indazoles and Pyrazolines Prepared in Continuous Flow Mode
Abstract
1. Introduction
2. Results
2.1. Antimicrobial Activity
2.2. Pharmacokinetic Properties and Drug-Likeness Prediction
3. Discussion
4. Materials and Methods
4.1. Bacterial Species Evaluated in this Study
4.2. Determination of the Minimal Inhibitory Concentrations (MICs)
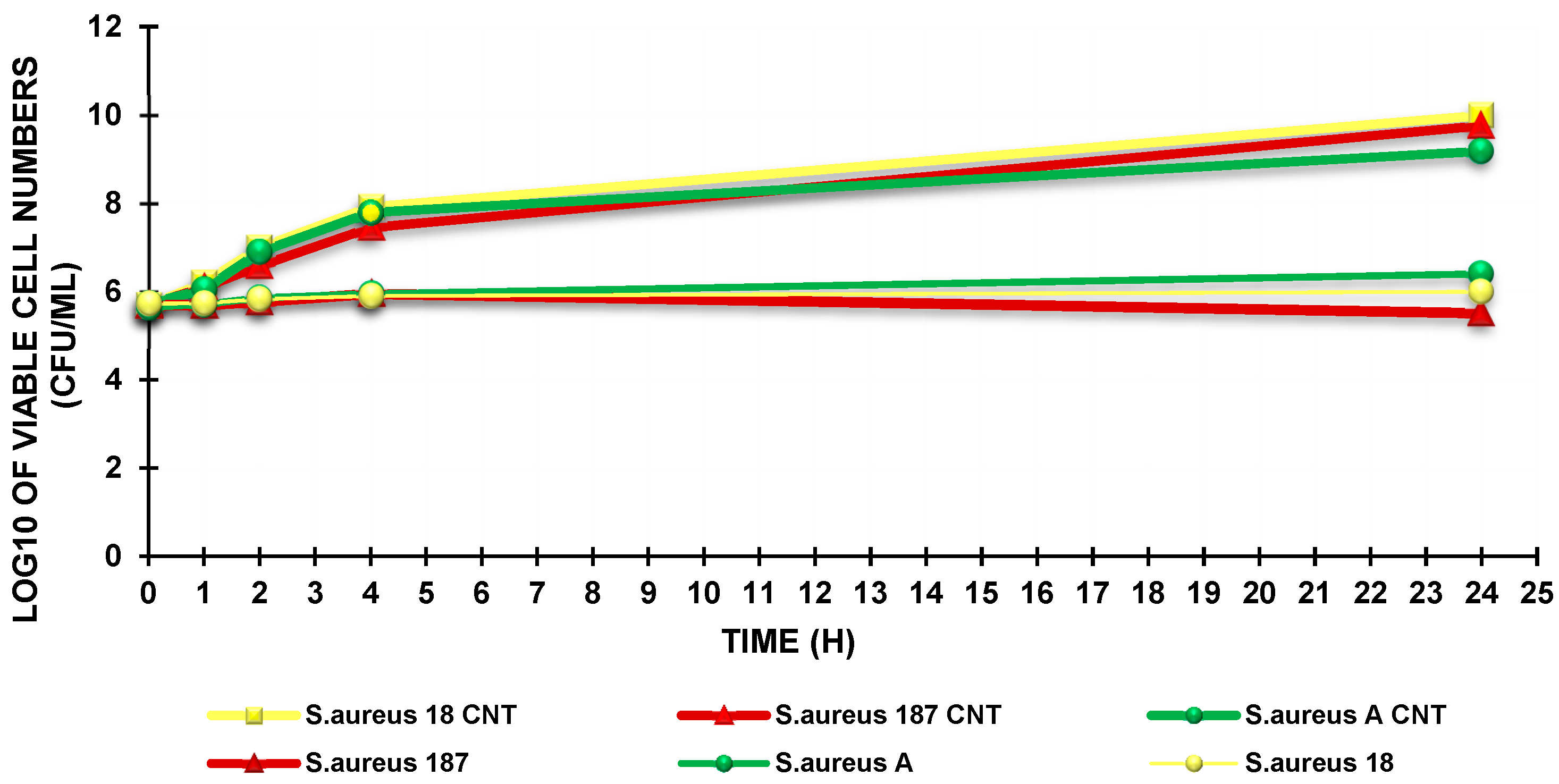
4.3. Killing Curves
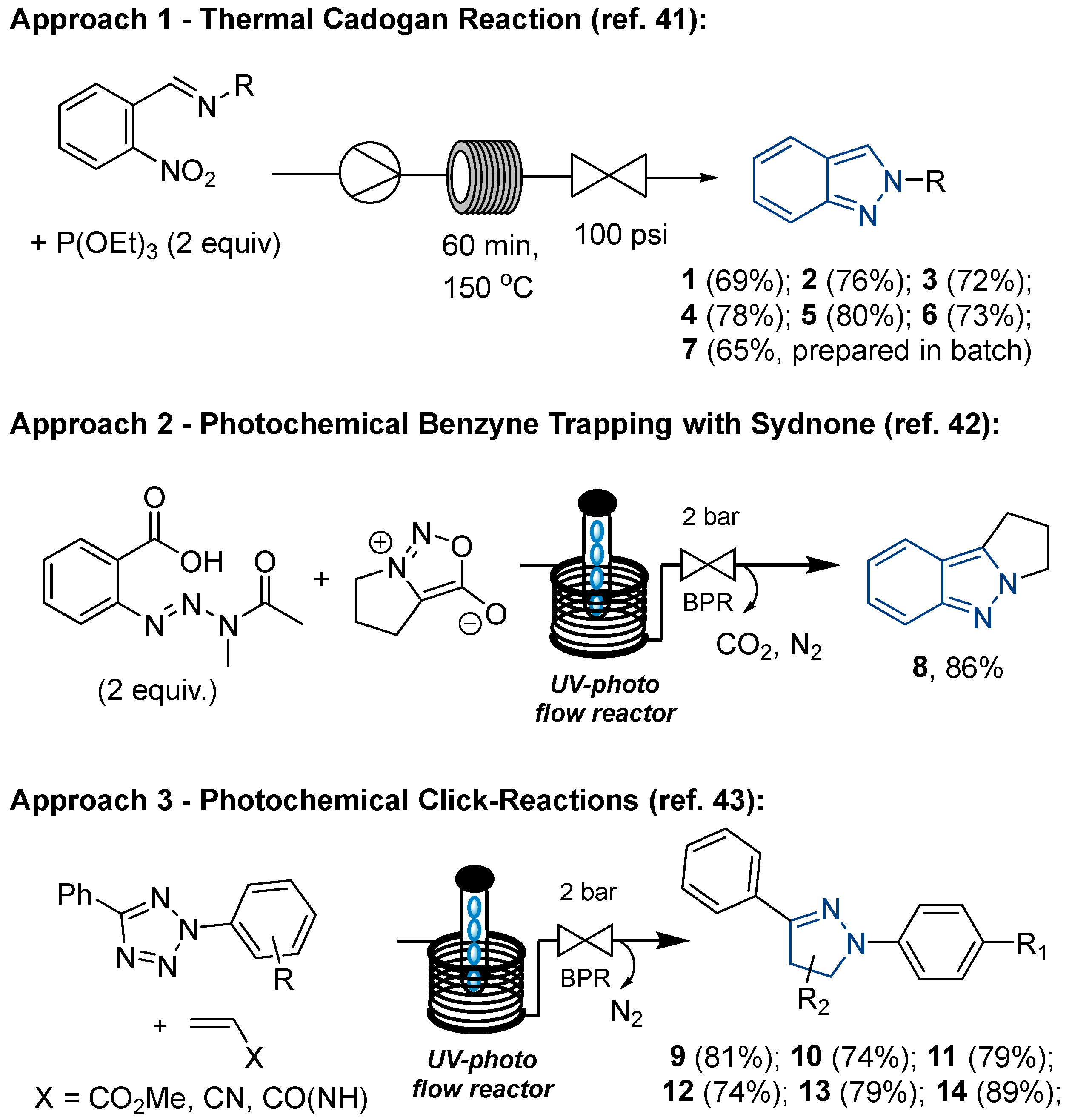
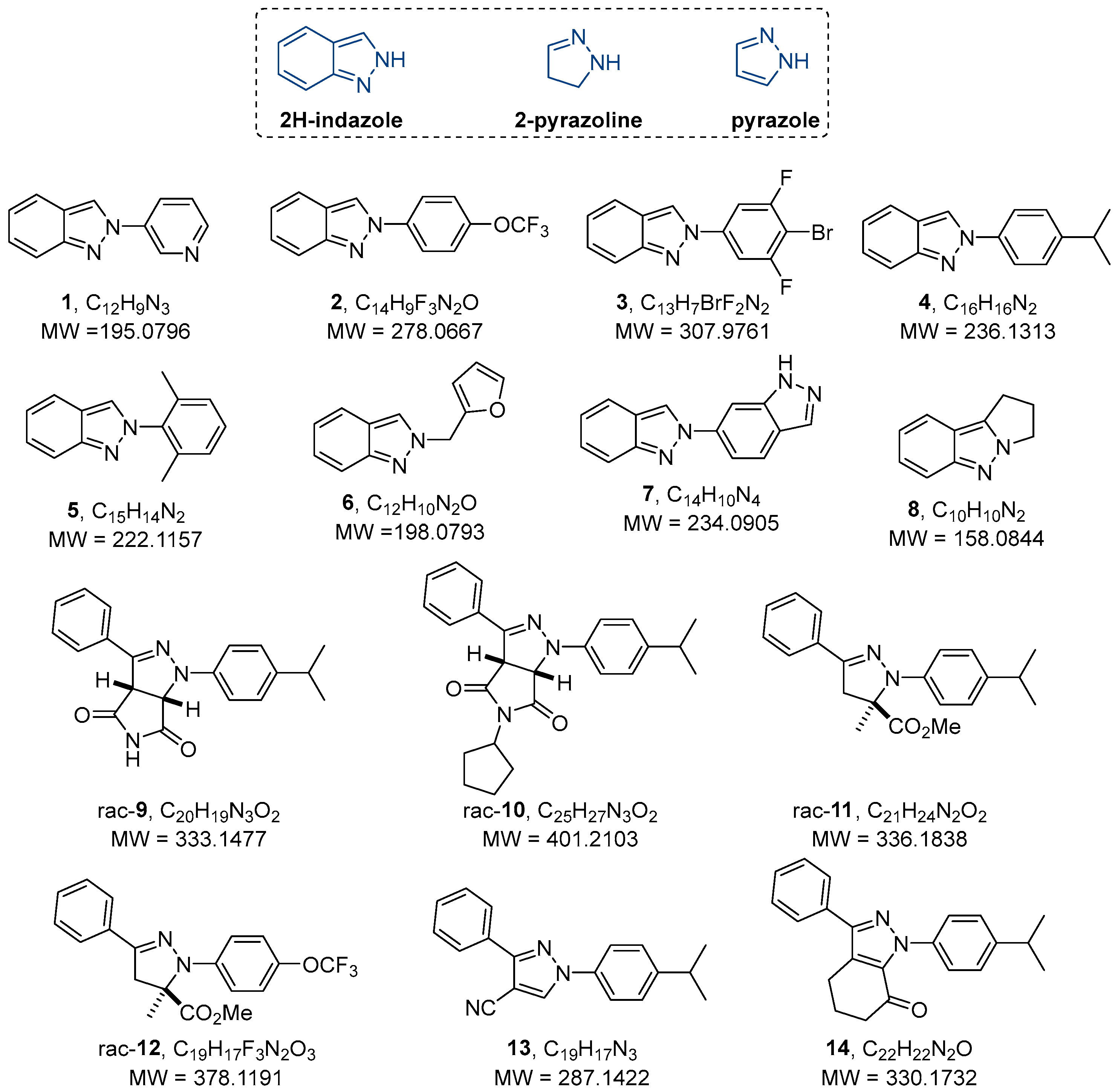
4.4. Preparation of Compounds Used in This Study and Synthetic Methods
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Interagency Coordination Group on Antimicrobial Resistance. No Time to Wait: Securing the Future from Drug-Resistant Infections; Report to the Secretary-General of the United Nations; WHO: Geneva, Switzerland, 2019; Available online: https://www.who.int/publications/i/item/no-time-to-wait-securing-the-future-from-drug-resistant-infections (accessed on 8 March 2023).
- Terreni, M.; Taccani, M.; Pregnolato, M. New Antibiotics for Multidrug-Resistant Bacterial Strains: Latest Research Developments and Future Perspectives. Molecules 2021, 26, 2671. [Google Scholar] [CrossRef] [PubMed]
- Songmee, B.; Jaehoon, L.; Jaehwa, L.; Eunah, K.; Sunhwa, L.; Jaeyon, Y.; Yeonho, K. Antimicrobial Resistance in Haemophilus influenzae Respiratory Tract Isolates in Korea: Results of a Nationwide Acute Respiratory Infections Surveillance. Antimicrob. Agents Chemother. 2010, 54, 65–71. [Google Scholar] [CrossRef]
- Mori, M.; Stelitano, G.; Chiarelli, L.R.; Cazzaniga, G.; Gelain, A.; Barlocco, D.; Elena Pini, E.; Meneghetti, F.; Villa, S. Synthesis, Characterization, and Biological Evaluation of New Derivatives Targeting MbtI as Antitubercular Agents. Pharmaceuticals 2021, 14, 155. [Google Scholar] [CrossRef] [PubMed]
- The Guardian for 200 Years. Calls to Rein in Antibiotic Use after Study Shows 65% Increase Worldwide. Guardian. 2018. Available online: https://amp.theguardian.com/science/2018/mar/26/calls-to-rein-in-antibiotic-use-after-study-shows-65-increaseworldwide (accessed on 20 December 2022).
- Ventola, C.L. The antibiotic resistance crisis: Part 1: Causes and threats. Pharm. Ther. 2015, 40, 277–283. [Google Scholar]
- Cazzaniga, G.; Mori, M.; Chiarelli, L.R.; Gelain, A.; Meneghetti, F.; Villa, S. Natural products against key Mycobacterium tuberculosis enzymatic targets: Emerging opportunities for drug discovery. Eur. J. Med. Chem. 2021, 224, 113732. [Google Scholar] [CrossRef]
- Yan, M.; Xu, L.; Wang, Y.; Wan, J.; Liu, T.; Liu, W.; Wan, Y.; Zhang, B.; Wang, R.; Li, Q. Opportunities and challenges of using five-membered ring compounds as promising antitubercular agents. Drug Dev. Res. 2020, 81, 402–418. [Google Scholar] [CrossRef] [PubMed]
- Küçükgüzel, S.G.; Şenkardes, S. Recent advances in bioactive pyrazoles. Eur. J. Med. Chem. 2014, 97, 786–815. [Google Scholar] [CrossRef]
- Patel, B.; Zunk, M.; Grant, G.; Rudrawar, S. Design, synthesis and bioactivity evaluation of novel pyrazole linked phenylthiazole derivatives in context of antibacterial activity. Bioorg. Med. Chem. Lett. 2021, 39, 127853. [Google Scholar] [CrossRef]
- Finn, J.; Mattia, K.; Morytko, M.; Ram, S.; Yang, Y.; Wu, X.; Mak, E.; Gallant, P.; Keith, D. Discovery of a potent and selective series of pyrazole bacterial methionyl-tRNA synthetase inhibitors. Bioorg. Med. Chem. Lett. 2003, 13, 2231–2234. [Google Scholar] [CrossRef]
- Yu, B.; Zhou, S.; Cao, L.; Hao, Z.; Yang, D.; Guo, X.; Zhang, N.; Bakulev, V.A.; Fan, Z. Design, Synthesis, and Evaluation of the Antifungal Activity of Novel Pyrazole–Thiazole Carboxamides as Succinate Dehydrogenase Inhibitors. J. Agric. Food Chem. 2020, 68, 7093–7102. [Google Scholar] [CrossRef] [PubMed]
- Faria, J.V.; Vegi, P.F.; Carvalho Miguita, A.G.; Silva dos Santos, M.; Boechat, N.; Rolim Bernardino, A.M. Recently reported biological activities of pyrazole compounds. Bioorg. Med. Chem. 2017, 25, 5891–5903. [Google Scholar] [CrossRef] [PubMed]
- Faria, J.V.; dos Santos, M.S.; Bernardino, A.M.R.; Becker, K.M.; Machado, G.M.C.; Rodrigues, R.F.; Canto-Cavalheiro, M.M.; Leon, L.L. Synthesis and activity of novel tetrazole compounds and their pyrazole-4-carbonitrile precursors against Leishmania spp. Bioorg. Med. Chem. Lett. 2013, 23, 6310–6312. [Google Scholar] [CrossRef] [PubMed]
- Comber, R.N.; Gray, R.J.; Secrist, J.A. Acyclic analogues of pyrazofurin: Syntheses and antiviral evaluation. Carbohydr. Res. 1991, 216, 441–452. [Google Scholar] [CrossRef]
- Meta, E.; Brullo, C.; Tonelli, M.; Franzblau, S.G.; Wang, Y.; Ma, R.; Baojie, W.; Orena, B.S.; Pasca, M.R.; Bruno, O. Pyrazole and imidazo [1,2-b]pyrazole derivatives as new potential anti-tuberculosis agents. Med. Chem. 2019, 15, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Brullo, C.; Caviglia, D.; Spallarossa, A.; Alfei, S.; Franzblau, S.G.; Tasso, B.; Schito, A.M. Microbiological Screening of 5-Functionalized Pyrazoles for the Future Development of Optimized Pyrazole-Based Delivery Systems. Pharmaceutics 2022, 14, 1770. [Google Scholar] [CrossRef]
- Wong, K.T.; Osman, H.; Parumasivam, T.; Supratman, U.; Che Omar, M.T.; Azmi, M.N. Synthesis, Characterization and Biological Evaluation of New 3,5-Disubstituted-Pyrazoline Derivatives as Potential Anti-Mycobacterium tuberculosis H37Ra Compounds. Molecules 2021, 26, 2081. [Google Scholar] [CrossRef]
- Evranos Aksoz, B.; Gurpinar, S.S.; Eryilmaz, M. Antimicrobial Activities of Some Pyrazoline and Hydrazone Derivatives. Turk. J. Pharm. Sci. 2020, 17, 500–505. [Google Scholar] [CrossRef]
- Nanda, S.S.; Yi, D.K.; Panda, O.P.; Chigurupati, S.; Mohapatra, T.K.; Hossain, M.I. Impact of Indazole Scaffold as Antibacterial and Antifungal Agent. Curr. Top. Med. Chem. 2022, 22, 1152–1159. [Google Scholar] [CrossRef]
- Qin, J.; Cheng, W.; Duan, Y.T.; Yang, H.; Yao, Y. Indazole as a Privileged Scaffold: The Derivatives and their Therapeutic Applications. Anticancer Agents Med. Chem. 2021, 21, 839–860. [Google Scholar] [CrossRef]
- Mal, S.; Malik, U.; Mahapatra, M.; Mishra, A.; Pal, D.; Paidesetty, S.K. A review on synthetic strategy, molecular pharmacology of indazole derivatives, and their future perspective. Drug Dev. Res. 2022, 83, 1469–1504. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Delivery Rev. 1997, 23, 3–25. [Google Scholar] [CrossRef]
- Hartung, I.V.; Huck, B.R.; Crespo, A. Rules were made to be broken. Nat. Rev. Chem. 2023, 7, 3–4. [Google Scholar] [CrossRef]
- Gioiello, A.; Piccinno, A.; Lozza, A.M.; Cerra, B. The Medicinal Chemistry in the Era of Machines and Automation: Recent Advances in Continuous Flow Technology. J. Med. Chem. 2020, 63, 6624–6647. [Google Scholar] [CrossRef]
- Baumann, M.; Moody, T.S.; Smyth, M.; Wharry, S. A Perspective on Continuous Flow Chemistry in the Pharmaceutical Industry. Org. Process Res. Dev. 2020, 24, 1802–1813. [Google Scholar] [CrossRef]
- Gutmann, B.; Cantillo, D.; Kappe, C.O. Continuous-Flow Technology—A Tool for the Safe Manufacturing of Active Pharmaceutical Ingredients. Angew. Chem. Int. Ed. 2015, 54, 6688–6728. [Google Scholar] [CrossRef]
- Baumann, M.; Baxendale, I.R. The synthesis of active pharmaceutical ingredients (APIs) using continuous flow chemistry. Beilstein J. Org. Chem. 2015, 11, 1194–1219. [Google Scholar] [CrossRef] [PubMed]
- Pastre, J.C.; Browne, D.L.; Ley, S.V. Flow chemistry syntheses of natural products. Chem. Soc. Rev. 2013, 42, 8849–8869. [Google Scholar] [CrossRef] [PubMed]
- Movsisyan, M.; Delbeke, E.I.P.; Berton, J.K.E.T.; Battilocchio, C.; Ley, S.V.; Stevens, C.V. Taming hazardous chemistry by continuous flow technology. Chem. Soc. Rev. 2016, 45, 4892–4928. [Google Scholar] [CrossRef]
- Dallinger, D.; Gutmann, B.; Kappe, C.O. The Concept of “Chemical Generators”: On-Site On-Demand Production of Hazardous Reagents in Continuous Flow. Acc. Chem. Res. 2020, 53, 6688–6729. [Google Scholar] [CrossRef] [PubMed]
- Baumann, M.; Moody, T.S.; Smyth, M.; Wharry, S. Overcoming the Hurdles and Challenges Associated with Developing Continuous Industrial Processes. Eur. J. Org. Chem. 2020, 48, 7398–7406. [Google Scholar] [CrossRef]
- Breen, C.P.; Nambiar, A.M.K.; Jamison, T.F.; Jensen, K.F. Ready, Set, Flow! Automated Continuous Synthesis and Optimization. Trends Chem. 2021, 3, 373–386. [Google Scholar] [CrossRef]
- Donnelly, K.; Baumann, M. Scalability of photochemical reactions in continuous flow mode. J. Flow Chem. 2021, 11, 223–241. [Google Scholar] [CrossRef]
- Kayahan, E.; Jacobs, M.; Braeken, L.; Thomassen, L.C.J.; Kuhn, S.; van Gerven, T.; Leblebici, M.E. Dawn of a new era in industrial photochemistry: The scale-up of micro- and mesostructured photoreactors. Beilstein J. Org. Chem. 2020, 16, 2484–2504. [Google Scholar] [CrossRef]
- Sambiago, C.; Noël, T. Flow Photochemistry: Shine Some Light on Those Tubes! Trends Chem. 2020, 2, 92–106. [Google Scholar] [CrossRef]
- Rehm, T.H. Reactor Technology Concepts for Flow Photochemistry. ChemPhotoChem 2020, 4, 235–254. [Google Scholar] [CrossRef]
- Buglioni, L.; Raymenants, F.; Slattery, A.; Zondag, S.D.A.; Noël, T. Technological Innovations in Photochemistry for Organic Synthesis: Flow Chemistry, High-Throughput Experimentation, Scale-up, and Photoelectrochemistry. Chem. Rev. 2022, 122, 2752–2906. [Google Scholar] [CrossRef] [PubMed]
- Noël, T.; Cao, Y.; Laudadio, G. The Fundamentals Behind the Use of Flow Reactors in Electrochemistry. Acc. Chem. Res. 2019, 52, 2858–2869. [Google Scholar] [CrossRef]
- Winterson, B.; Rennigholtz, T.; Wirth, T. Flow electrochemistry: A safe tool for fluorine chemistry. Chem. Sci. 2021, 12, 9053–9059. [Google Scholar] [CrossRef] [PubMed]
- Duffy, M.; Di Filippo, M.; Baumann, M. Synthesis of 2H-indazoles via the Cadogan reaction in batch and flow mode. Tetrahedron Lett. 2021, 86, 153522. [Google Scholar] [CrossRef]
- Bracken, C.; Batsanov, A.S.; Baumann, M. Development of a Continuous Photochemical Benzyne-Forming Process. SynOpen 2021, 5, 29–35. [Google Scholar] [CrossRef]
- Burke, A.; Spiccio, S.; Di Filippo, M.; Baumann, M. Photochemical Synthesis of Pyrazolines from Tetrazoles in Flow. SynOpen 2023, 7, 69–75. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef] [PubMed]
- Brenk, R.; Schipani, A.; James, D.; Krasowski, A.; Gilbert, I.H.; Frearson, J.; Wyatt, P.G. Lessons learnt from assembling screening libraries for drug discovery for neglected diseases. ChemMedChem 2008, 3, 435–444. [Google Scholar] [CrossRef] [PubMed]
- Delaney, J.S. ESOL: Estimating Aqueous Solubility Directly from Molecular Structure. J. Chem. Inf. Model. 2004, 44, 1000–1005. [Google Scholar] [CrossRef]
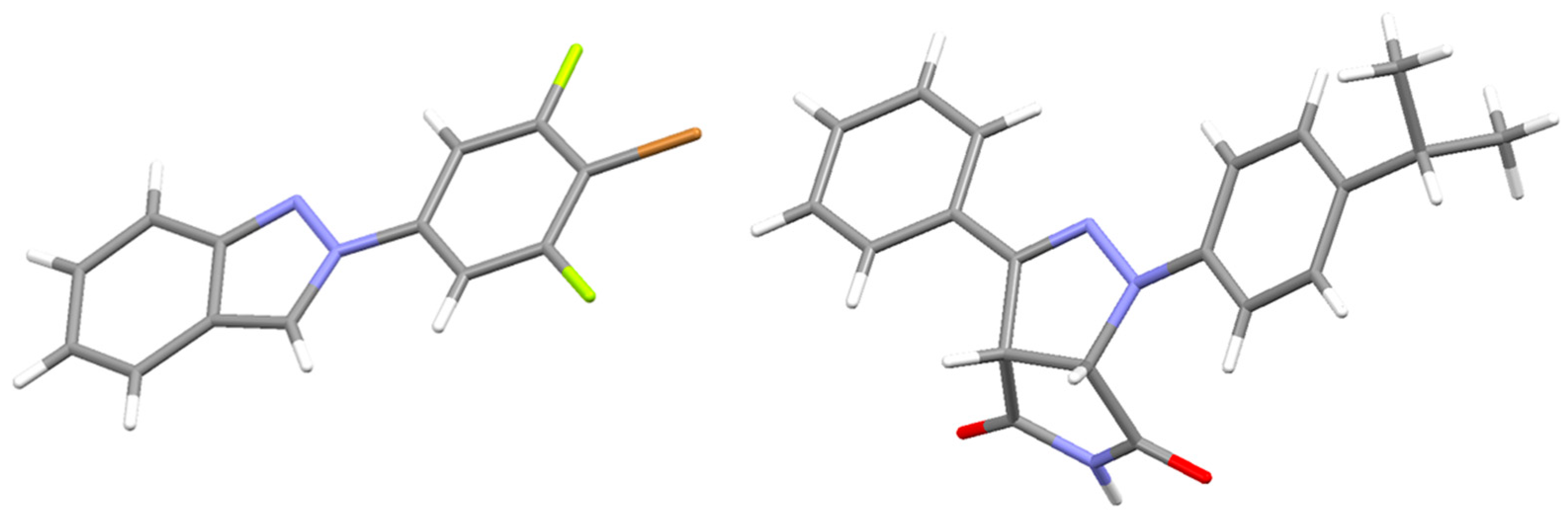
- CCDC 2221468 and CCDC 2237067 Contain the Supplementary CRYSTALLOGRAPHIC Data for This Paper. The Cambridge Crystallographic Data Centre. Available online: www.ccdc.cam.ac.uk/structures (accessed on 21 January 2023).
- EUCAST. European Committee on Antimicrobial Susceptibility Testing. Available online: https://www.eucast.org/ast_of_bacteria (accessed on 23 May 2022).
- Schito, A.M.; Piatti, G.; Caviglia, D.; Zuccari, G.; Alfei, S. Broad-Spectrum Bactericidal Activity of a Synthetic Random Copolymer Based on 2-Methoxy-6-(4-Vinylbenzyloxy)-Benzylammonium Hydrochloride. Int. J. Mol. Sci. 2021, 22, 5021. [Google Scholar] [CrossRef]
- Schito, A.M.; Piatti, G.; Caviglia, D.; Zuccari, G.; Zorzoli, A.; Marimpietri, D.; Alfei, S. Bactericidal Activity of Non-Cytotoxic Cationic Nanoparticles against Clinically and Environmentally Relevant Pseudomonas spp. Isolates. Pharmaceutics 2021, 13, 1411. [Google Scholar] [CrossRef] [PubMed]
| Strains | MIC (μg/mL) | ||||
|---|---|---|---|---|---|
| 2 | 3 | 5 | 9 | R.A. | |
| S. aureus 18 * | >128 | >128 | 128 | 4 | 256 (O) |
| S. aureus 187 | >128 | >128 | 128 | 4 | 0.5 (O) |
| S. aureus A * | >128 | >128 | 64 | 4 | 256 (O) |
| S. epidermidis 22 * | >128 | >128 | 128 | 4 | 16 (O) |
| S. epidermidis 180 | >128 | >128 | 128 | 4 | 0.25 (O) |
| S. epidermidis 2R * | >128 | >128 | 64 | 4 | 256 (O) |
| E. faecalis 1 °,# | 128 | 128 | >128 | 4 | 64 (V); 512 (T) |
| E. faecalis 50 ° | >128 | 128 | >128 | 4 | 32 (V) |
| E. faecalis 365 | 128 | 128 | >128 | 4 | 1 (V) |
| E. faecium 300 ° | >128 | >128 | >128 | 4 | 256 (V) |
| E. faecium 364 ° | >128 | >128 | >128 | 4 | 256 (V) |
| E. faecium 503 | >128 | >128 | >128 | 4 | 1 (V) |
| E. coli 224 | >128 | >128 | >128 | >128 | 0.125 (I) |
| E. coli 462 ** | >128 | >128 | >128 | >128 | 16 (I) |
| E. coli 477 ** | >128 | >128 | >128 | >128 | 16 (I) |
| P. aeruginosa 1V ** | >128 | >128 | >128 | >128 | 32 (I) |
| P. aeruginosa 265 **,°° | >128 | >128 | >128 | >128 | 16 (I): 16 (C) |
| P. aeruginosa 403 | >128 | >128 | >128 | >128 | 1 (I) |
| 2 | 3 | 5 | 9 | |
|---|---|---|---|---|
| Physicochemical Property | ||||
| MW (g/mol) | 278.23 | 309.11 | 222.29 | 333.38 |
| Fraction Csp3 | 0.07 | 0.00 | 0.13 | 0.25 |
| Rotatable bonds | 3 | 1 | 1 | 3 |
| H-bond acceptors | 5 | 3 | 1 | 3 |
| H-bond donors | 0 | 0 | 0 | 1 |
| TPSA a (Å2) | 27.05 Å2 | 17.82 Å2 | 17.82 Å2 | 61.77 Å2 |
| Lipophilicity | ||||
| LogP b | 4.41 | 4.12 | 3.96 | 3.79 |
| Water solubility | ||||
| Solubility (mg/mL) c | 0.00555 mg/mL | 0.00387 mg/mL | 0.0111 mg/mL | 0.0118 mg/mL |
| Solubility | moderately | moderately | moderately | moderately |
| Pharmacokinetics | ||||
| GI absorption | high | high | high | high |
| BBB permeant | yes | yes | yes | yes |
| Pgp substrate | no | no | no | no |
| CYP1A2 inhibitor | yes | yes | yes | no |
| CYP2C19 inhibitor | yes | yes | yes | yes |
| CYP2C9 inhibitor | no | no | no | yes |
| CYP2D6 inhibitor | yes | no | yes | no |
| CYP3A4 inhibitor | no | no | no | yes |
| Druglikeness | ||||
| Lipinski violations | 0 | 1 violation: MLogP > 4.15 | 0 | 0 |
| Medicinal chemistry | ||||
| PAINS alerts | 0 | 0 | 0 | 0 |
| Brenk alerts [46] | 0 | 1 (halogenated ring) | 0 | 1 (phthalimide) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Burke, A.; Di Filippo, M.; Spiccio, S.; Schito, A.M.; Caviglia, D.; Brullo, C.; Baumann, M. Antimicrobial Evaluation of New Pyrazoles, Indazoles and Pyrazolines Prepared in Continuous Flow Mode. Int. J. Mol. Sci. 2023, 24, 5319. https://doi.org/10.3390/ijms24065319
Burke A, Di Filippo M, Spiccio S, Schito AM, Caviglia D, Brullo C, Baumann M. Antimicrobial Evaluation of New Pyrazoles, Indazoles and Pyrazolines Prepared in Continuous Flow Mode. International Journal of Molecular Sciences. 2023; 24(6):5319. https://doi.org/10.3390/ijms24065319
Chicago/Turabian StyleBurke, Adam, Mara Di Filippo, Silvia Spiccio, Anna Maria Schito, Debora Caviglia, Chiara Brullo, and Marcus Baumann. 2023. "Antimicrobial Evaluation of New Pyrazoles, Indazoles and Pyrazolines Prepared in Continuous Flow Mode" International Journal of Molecular Sciences 24, no. 6: 5319. https://doi.org/10.3390/ijms24065319
APA StyleBurke, A., Di Filippo, M., Spiccio, S., Schito, A. M., Caviglia, D., Brullo, C., & Baumann, M. (2023). Antimicrobial Evaluation of New Pyrazoles, Indazoles and Pyrazolines Prepared in Continuous Flow Mode. International Journal of Molecular Sciences, 24(6), 5319. https://doi.org/10.3390/ijms24065319