A Highly Expressed Antennae Odorant-Binding Protein Involved in Recognition of Herbivore-Induced Plant Volatiles in Dastarcus helophoroides
Abstract
1. Introduction
2. Results
2.1. Analyses of HIPVs of P. massoniana Lamb
2.2. Odorant-Binding Proteins in D. helophoroides
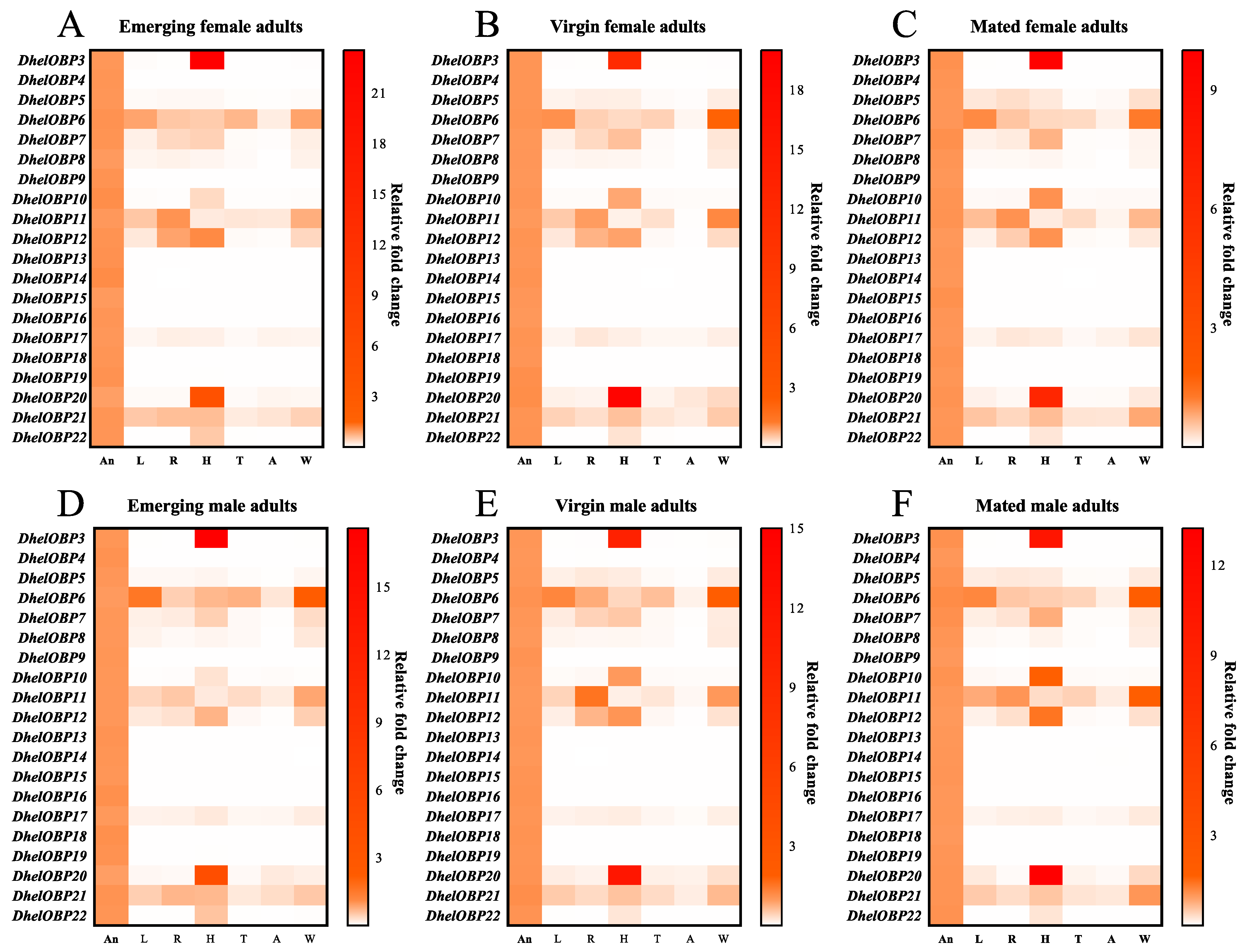
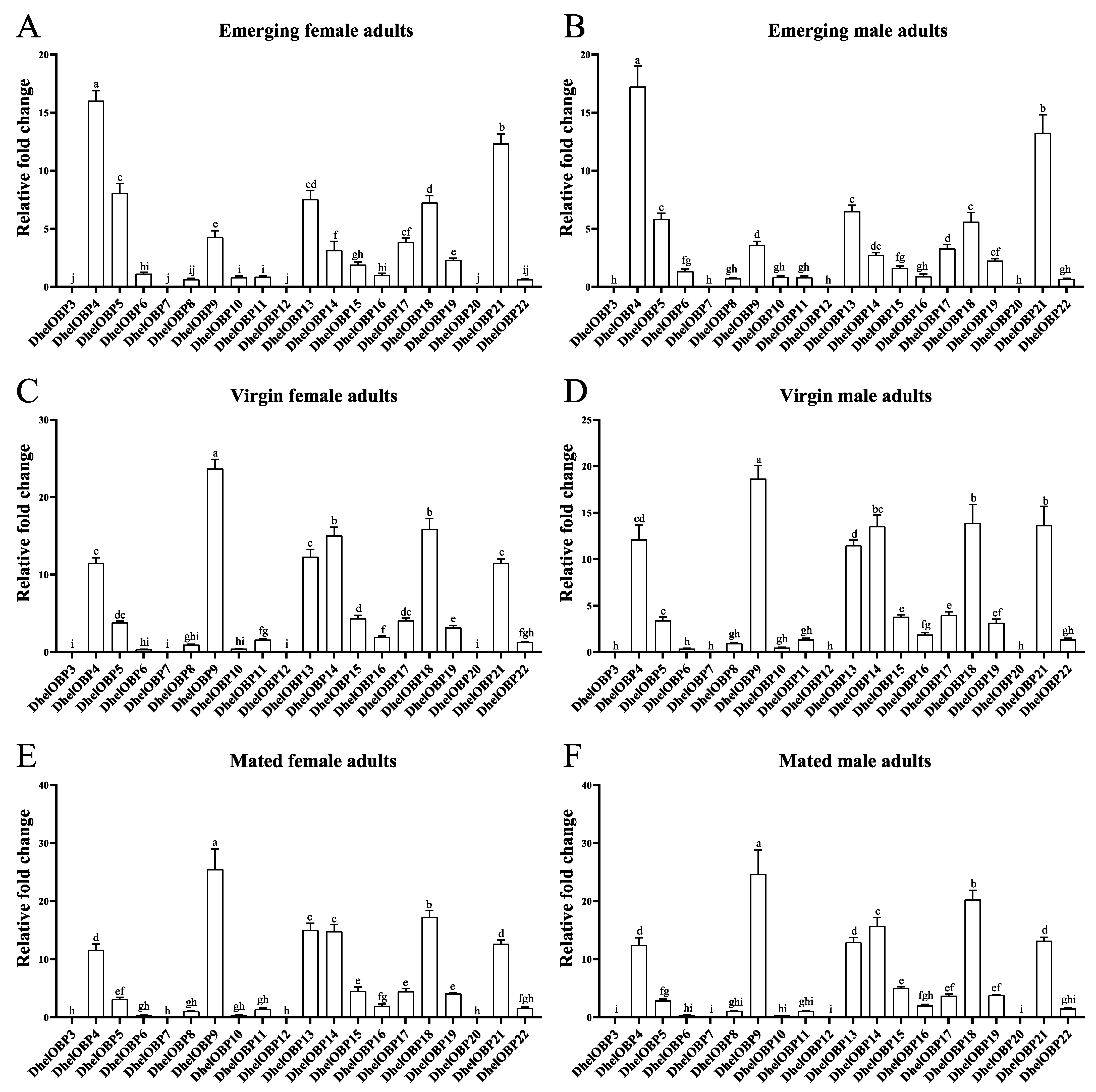
2.3. Spatio-Temporal Expression Patterns of DhelOBPs
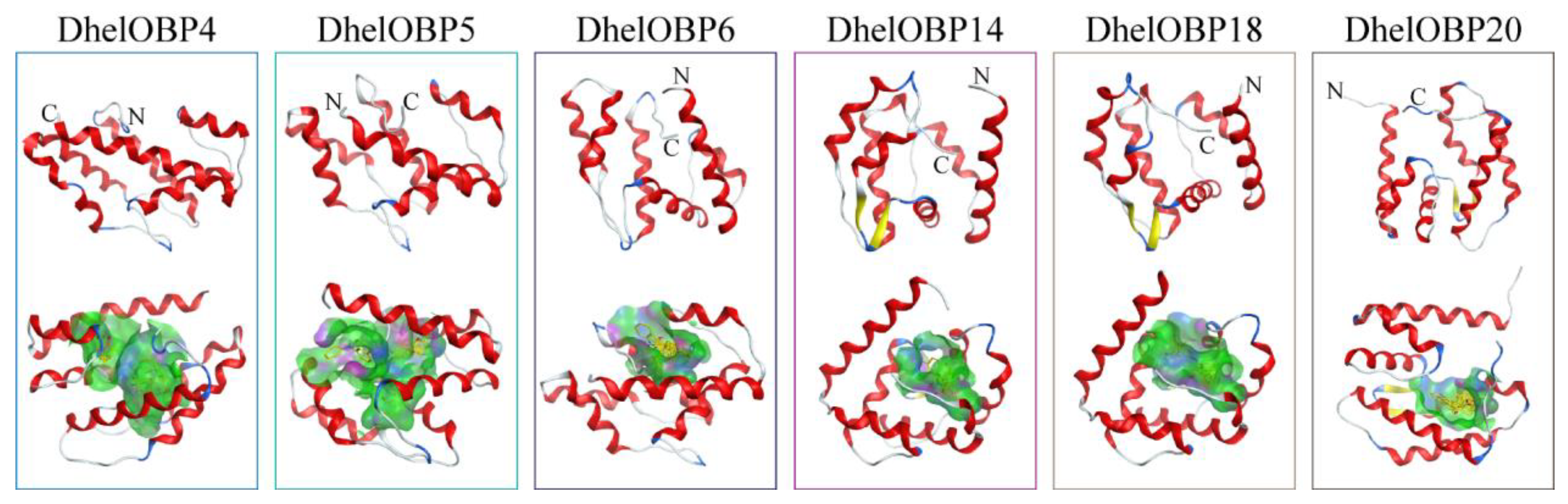
2.4. AlphaFold2-Based Modeling and Molecular Docking of DhelOBPs
2.5. Binding Characteristic of Recombinant DhelOBPs
2.6. Behavioral Response of D. helophoroides to HIPVs with the Best Binding Affinities
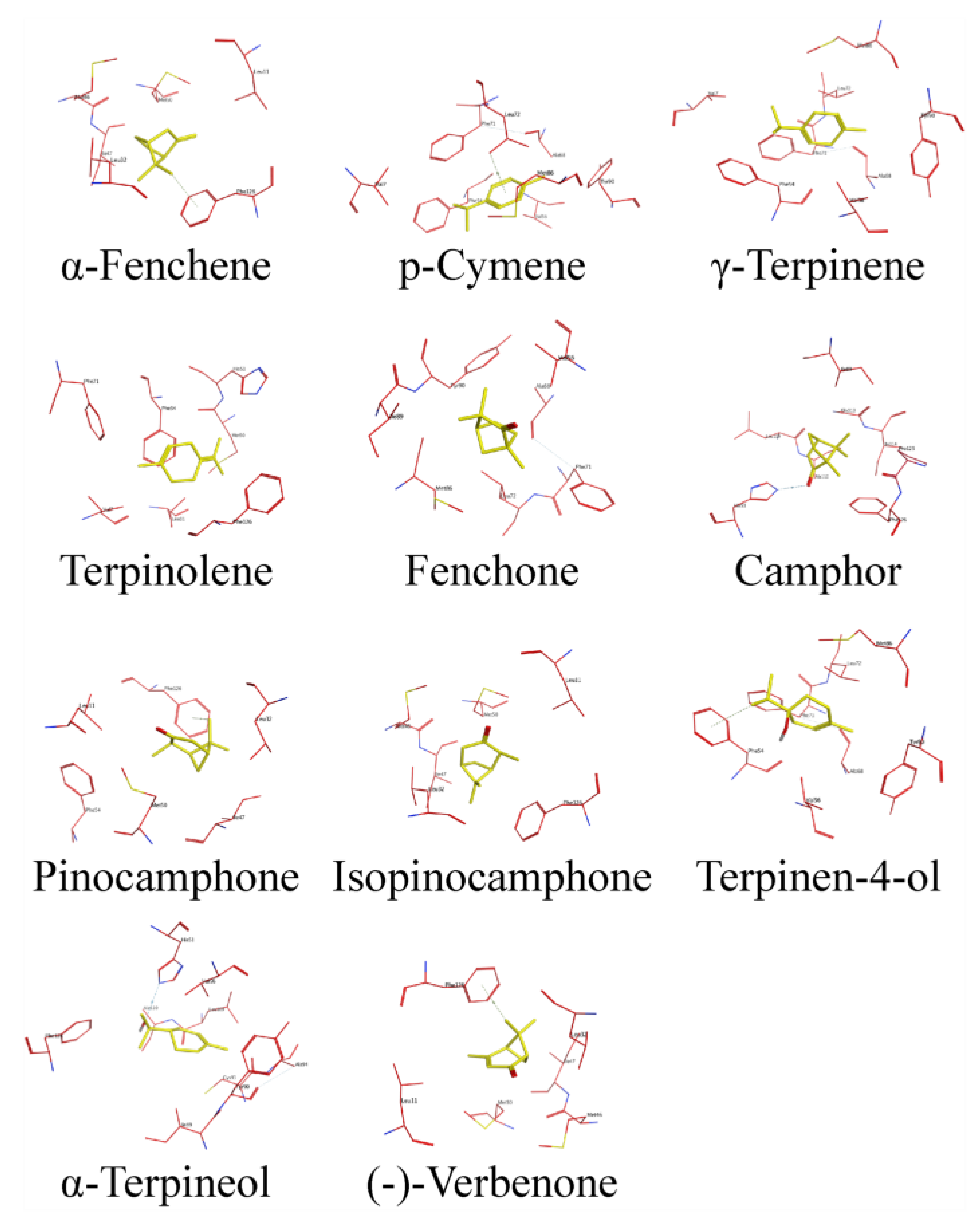
2.7. Binding Conformation of DhelOBP4 with HIPVs
3. Discussion
4. Methods and Materials
4.1. Insect Culturing and Pine Woods Collection
4.2. Collection and Analysis of P. massoniana HIPVs
4.3. Classification of DhelOBPs
4.4. Tissue Sampling, RNA Extraction, and cDNA Synthesis
4.5. Spatio-Temporal Expression Analyses
4.6. AlphaFold2-Based Modeling and Molecular Docking
4.7. Subcloning, Expression, and Purification of Target DhelOBPs
4.8. Fluorescence Competitive Binding Assays
4.9. Y-Tube Olfactometer Experiments
4.10. Synthesis and Microinjection of dsRNA
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Leal, W.S. Odorant Reception in Insects: Roles of Receptors, Binding Proteins, and Degrading Enzymes. Annu. Rev. Entomol. 2013, 58, 373–391. [Google Scholar] [CrossRef]
- Knolhoff, L.M.; Heckel, D.G. Behavioral Assays for Studies of Host Plant Choice and Adaptation in Herbivorous Insects. Annu. Rev. Entomol. 2014, 59, 263–278. [Google Scholar] [CrossRef]
- Zhou, J.-J. Odorant-binding proteins in insects. Vitam. Horm. 2010, 83, 241–272. [Google Scholar]
- Turlings, T.C.J.; Erb, M. Tritrophic Interactions Mediated by Herbivore-Induced Plant Volatiles: Mechanisms, Ecological Relevance, and Application Potential. Annu. Rev. Entomol. 2018, 63, 433–452. [Google Scholar] [CrossRef] [PubMed]
- De Pasqual, C.; Groot, A.T.; Mappes, J.; Burdfield-Steel, E. Evolutionary importance of intraspecific variation in sex pheromones. Trends Ecol. Evol. 2021, 36, 848–859. [Google Scholar] [CrossRef]
- De Moraes, C.M.; Lewis, W.J.; Paré, P.W.; Alborn, H.T.; Tumlinson, J.H. Herbivore-infested plants selectively attract parasitoids. Nature 1998, 393, 570–573. [Google Scholar] [CrossRef]
- Gershenzon, J.; Dudareva, N. The function of terpene natural products in the natural world. Nat. Chem. Biol. 2007, 3, 408–414. [Google Scholar] [CrossRef]
- Xiao, Y.; Wang, Q.; Erb, M.; Turlings, T.C.J.; Ge, L.; Hu, L.; Li, J.; Han, X.; Zhang, T.; Lu, J.; et al. Specific herbivore-induced volatiles defend plants and determine insect community composition in the field. Ecol. Lett. 2012, 15, 1130–1139. [Google Scholar] [CrossRef] [PubMed]
- Kafle, B.D.; Morawo, T.; Fadamiro, H. Host-Induced Plant Volatiles Mediate Ability of the Parasitoid Microplitis croceipes to Discriminate Between Unparasitized and Parasitized Heliothis virescens Larvae and Avoid Superparasitism. J. Chem. Ecol. 2020, 46, 967–977. [Google Scholar] [CrossRef] [PubMed]
- Pelosi, P.; Zhou, J.J.; Ban, L.P.; Calvello, M. Soluble proteins in insect chemical communication. Cell. Mol. Life Sci. 2006, 63, 1658–1676. [Google Scholar] [CrossRef]
- Sun, J.S.; Xiao, S.; Carlson, J.R. The diverse small proteins called odorant-binding proteins. Open Biol. 2018, 8, 180208. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.-M.; Long, G.-J.; Guo, H.; Liu, X.-Z.; Wang, H.; Dewer, Y.; Li, Z.-Q.; Liu, K.; Zhang, Q.-L.; Ma, Y.-F.; et al. Two carboxylesterase genes in Plutella xylostella associated with sex pheromones and plant volatiles degradation. Pest Manag. Sci. 2021, 77, 2737–2746. [Google Scholar] [CrossRef]
- Brito, N.F.; Moreira, M.F.; Melo, A. A look inside odorant-binding proteins in insect chemoreception. J. Insect Physiol. 2016, 95, 51–65. [Google Scholar] [CrossRef] [PubMed]
- Venthur, H.; Mutis, A.; Zhou, J.-J.; Quiroz, A. Ligand binding and homology modelling of insect odorant-binding proteins. Physiol. Entomol. 2015, 39, 183–198. [Google Scholar] [CrossRef]
- Wang, S.-N.; Shan, S.; Yu, G.-Y.; Wang, H.; Dhiloo, K.H.; Khashaveh, A.; Zhang, F.; Zhang, Y.-J. Identification of odorant-binding proteins and functional analysis of antenna-specific AplaOBP1 in the emerald ash borer, Agrilus planipennis. J. Pest Sci. 2020, 93, 853–865. [Google Scholar] [CrossRef]
- Liu, X.-Q.; Jiang, H.-B.; Fan, J.-Y.; Liu, T.-Y.; Meng, L.-W.; Liu, Y.; Yu, H.-Z.; Dou, W.; Wang, J.-J. An odorant-binding protein of Asian citrus psyllid, Diaphorina citri, participates in the response of host plant volatiles. Pest Manag. Sci. 2021, 77, 3068–3079. [Google Scholar] [CrossRef]
- Guo, H.; Guo, P.P.; Sun, Y.L.; Huang, L.Q.; Wang, C.Z. Contribution of odorant binding proteins to olfactory detection of (Z)-11-hexadecenal in Helicoverpa armigera. Insect Biochem. Mol. Biol. 2021, 131, 103554. [Google Scholar] [CrossRef]
- Khuhro, S.A.; Liao, H.; Dong, X.T.; Yu, Q.; Yan, Q.; Dong, S.L. Two general odorant binding proteins display high bindings to both host plant volatiles and sex pheromones in a pyralid moth Chilo suppressalis (Lepidoptera: Pyralidae). J. Asia-Pac. Entomol. 2017, 20, 521–528. [Google Scholar] [CrossRef]
- Liu, Z.; Vidal, D.M.; Syed, Z.; Ishida, Y.; Leal, W.S. Pheromone Binding to General Odorant-binding Proteins from the Navel Orangeworm. J. Chem. Ecol. 2010, 36, 787–794. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.-Z.; Liu, J.-T.; Zhou, J.-J.; Wang, Q.; Dong, J.-Z.; Zhang, Y.-J.; Li, X.-C.; Li, J.; Gu, S.-H. Expressional and functional comparisons of two general odorant binding proteins in Agrotis ipsilon. Insect Biochem. Mol. Biol. 2018, 98, 34–47. [Google Scholar] [CrossRef]
- Konstantopoulou, M.A.; Pratsinis, H.; Kletsas, D.; Mazomenos, B.E. Pheromone-binding protein and general odorant- binding protein of Sesamia nonagrioides: Sex- and diel-dependent expression. Entomol. Exp. Et Appl. 2006, 119, 129–136. [Google Scholar] [CrossRef]
- Yao, Q.; Xu, S.; Dong, Y.; Lu, K.; Chen, B. Identification and characterisation of two general odourant-binding proteins from the litchi fruit borer, Conopomorpha sinensis Bradley. Pest Manag. Sci. 2016, 72, 877–887. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.L.; Yi, S.C.; Li, D.Z.; Nie, X.P.; Li, S.Q.; Wang, M.-Q.; Zhou, A.M. Optimization of reverse chemical ecology method: False positive binding of Aenasius bambawalei odorant binding protein 1 caused by uncertain binding mechanism. Insect Mol. Biol. 2018, 27, 305–318. [Google Scholar] [CrossRef]
- Yang, R.N.; Li, D.Z.; Yu, G.; Yi, S.C.; Zhang, Y.; Kong, D.X.; Wang, M.Q. Structural Transformation Detection Contributes to Screening of Behaviorally Active Compounds: Dynamic Binding Process Analysis of DhelOBP21 from Dastarcus helophoroides. J. Chem. Ecol. 2017, 43, 1033–1045. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Tzotzos, G.; Woodcock, C.; Pickett, J.A.; Hooper, T.; Field, L.M.; Zhou, J.-J. Binding of the General Odorant Binding Protein of Bombyx mori BmorGOBP2 to the Moth Sex Pheromone Components. J. Chem. Ecol. 2010, 36, 1293–1305. [Google Scholar] [CrossRef] [PubMed]
- He, P.; Chen, G.-L.; Li, S.; Wang, J.; Ma, Y.-F.; Pan, Y.-F.; He, M. Evolution and functional analysis of odorant-binding proteins in three rice planthoppers: Nilaparvata lugens, Sogatella furcifera, and Laodelphax striatellus. Pest Manag. Sci. 2019, 75, 1606–1620. [Google Scholar] [CrossRef]
- Zhu, J.; Ban, L.; Song, L.-M.; Liu, Y.; Pelosi, P.; Wang, G. General odorant-binding proteins and sex pheromone guide larvae of Plutella xylostella to better food. Insect Biochem. Mol. Biol. 2016, 72, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Larter, N.K.; Sun, J.S.; Carlson, J.R. Organization and function of Drosophila odorant binding proteins. eLife 2016, 5, e20242. [Google Scholar] [CrossRef]
- Gonzalez, D.; Rihani, K.; Neiers, F.; Poirier, N.; Briand, L. The Drosophila odorant-binding protein 28a is involved in the detection of the floral odour -ionone. Cell. Mol. Life Sci. 2020, 77, 2565–2577. [Google Scholar] [CrossRef]
- Leal, W.S. Reverse chemical ecology at the service of conservation biology. Proc. Natl. Acad. Sci. USA 2017, 114, 12094–12096. [Google Scholar] [CrossRef]
- Kamala Jayanthi, P.D.; Kempraj, V.; Aurade, R.M.; Kumar Roy, T.; Shivashankara, K.S.; Verghese, A. Computational reverse chemical ecology: Virtual screening and predicting behaviorally active semiochemicals for Bactrocera dorsalis. BMC Genom. 2014, 15, 209. [Google Scholar]
- Yi, S.-Y.; Li, D.-Z.; Zhou, C.-X.; Tang, Y.-L.; Abdelnabby, H.E.; Wang, M.-Q. Screening behaviorally active compounds based on fluorescence quenching in combination with binding mechanism analyses of SspOBP7, an odorant binding protein from Sclerodermus sp. Int. J. Biol. Macromol. 2018, 107, 2667–2678. [Google Scholar] [CrossRef]
- Sun, S.-F.; Zeng, F.-F.; Yi, S.-C.; Wang, M.-Q. Molecular Screening of Behaviorally Active Compounds with CmedOBP14 from the Rice Leaf Folder Cnaphalocrocis medinalis. J. Chem. Ecol. 2019, 45, 849–857. [Google Scholar] [CrossRef]
- Wei, J.-R.; Yang, Z.-Q.; Poland, T.M.; Du, J.-W. Parasitism and olfactory responses of Dastarcus helophoroides (Coleoptera: Bothrideridae) to different Cerambycid hosts. BioControl 2009, 54, 733–742. [Google Scholar] [CrossRef]
- Yang, Z.-Q.; Wang, X.-Y.; Zhang, Y.-N. Recent advances in biological control of important native and invasive forest pests in China. Biol. Control 2014, 68, 117–128. [Google Scholar] [CrossRef]
- Dippel, S.; Oberhofer, G.; Kahnt, J.; Gerischer, L.; Opitz, L.; Schachtner, J.; Stanke, M.; Schütz, S.; Wimmer, E.A.; Angeli, S. Tissue-specific transcriptomics, chromosomal localization, and phylogeny of chemosensory and odorant binding proteins from the red flour beetle Tribolium castaneum reveal subgroup specificities for olfaction or more general functions. BMC Genom. 2014, 15, 1141. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, research0034.1. [Google Scholar] [CrossRef]
- Zhang, P.-J.; Zhao, C.; Ye, Z.-H.; Yu, X.-P. Trade-off between defense priming by herbivore-induced plant volatiles and constitutive defense in tomato. Pest Manag. Sci. 2020, 76, 1893–1901. [Google Scholar] [CrossRef] [PubMed]
- Hare, J.D. Ecological Role of Volatiles Produced by Plants in Response to Damage by Herbivorous Insects. Annu. Rev. Entomol. 2011, 56, 161–180. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Li, D.-Z.; Min, S.-F.; Mi, F.; Zhou, S.-S.; Wang, M.-Q. Analysis of chemosensory gene families in the beetle Monochamus alternatus and its parasitoid Dastarcus helophoroides. Comp. Biochem. Physiol. Part D Genom. Proteom. 2014, 11, 1–8. [Google Scholar] [CrossRef]
- Ju, Q.; Li, X.; Jiang, X.J.; Qu, M.J.; Guo, X.Q.; Han, Z.J.; Li, F. Transcriptome and tissue-specific expression analysis of Obp and Csp genes in the dark black chafer. Arch. Insect Biochem. Physiol. 2014, 87, 177–200. [Google Scholar] [CrossRef]
- Qu, C.; Wang, R.; Che, W.-N.; Li, F.-Q.; Zhao, H.-P.; Wei, Y.-Y.; Luo, C.; Xue, M. Identification and tissue distribution of odorant binding protein genes in Harmonia axyridis (Coleoptera: Coccinellidae). J. Integr. Agric. 2021, 20, 2204–2213. [Google Scholar] [CrossRef]
- Li, X.M.; Zhu, X.Y.; Wang, Z.Q.; Wang, Y.; He, P.; Chen, G.; Sun, L.; Deng, D.G.; Zhang, Y.N. Candidate chemosensory genes identified in Colaphellus bowringi by antennal transcriptome analysis. BMC Genom. 2015, 16, 1028. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Rao, X.-J.; Li, M.-Y.; Feng, M.-F.; He, M.-Z.; Li, S.-G. Identification of candidate chemosensory genes in the antennal transcriptome of Tenebrio molitor (Coleoptera: Tenebrionidae). Comp. Biochem. Physiol. Part D Genom. Proteom. 2015, 13, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hu, P.; Gao, P.; Tao, J.; Luo, Y. Antennal transcriptome analysis and expression profiles of olfactory genes in Anoplophora chinensis. Sci. Rep. 2017, 7, 15470. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Hao, E.; Li, Y.; Yang, H.; Sun, P.; Lu, P.; Qiao, H. Antennal transcriptome analysis of olfactory genes and tissue expression profiling of odorant binding proteins in Semanotus bifasciatus (cerambycidae: Coleoptera). BMC Genom. 2022, 23, 461. [Google Scholar] [CrossRef]
- Yang, R.; Li, D.; Yi, S.; Wang, M. Evolutionarily conserved odorant-binding proteins participate in establishing tritrophic interactions. iScience 2022, 25, 104664. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.-L.; Fu, X.-B.; Cui, H.-C.; Zhao, L.; Yu, J.-Z.; Li, H.-L. Functional Characteristics, Electrophysiological and Antennal Immunolocalization of General Odorant-Binding Protein 2 in Tea Geometrid, Ectropis obliqua. Int. J. Mol. Sci. 2018, 19, 875. [Google Scholar] [CrossRef]
- Sun, X.; Zhou, W.; Liu, H.; Zhang, A.; Ai, C.R.; Zhou, S.S.; Zhou, C.X.; Wang, M.Q. Transgenic Bt Rice Does Not Challenge Host Preference of the Target Pest of Rice Leaffolder, Cnaphalocrocis medinalis (Lepidoptera: Pyralidae). PLoS ONE 2013, 8, e79032. [Google Scholar] [CrossRef]
- Sun, Z.; Liu, Z.; Zhou, W.; Jin, H.; Liu, H.; Zhou, A.; Zhang, A.; Wang, M.Q. Temporal interactions of plant - insect - predator after infection of bacterial pathogen on rice plants. Sci. Rep. 2016, 6, 26043. [Google Scholar] [CrossRef]




| Number | Substance | Retention Time (min) | PubChem CID | CAS Number | 2-D Structures |
|---|---|---|---|---|---|
| 1 | α-Fenchene | 9.260 | 28930 | 471-84-1 | 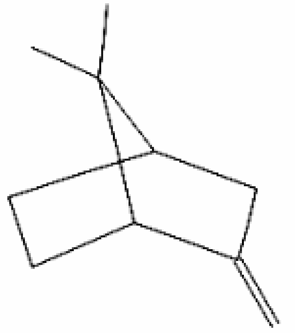 |
| 2 | p-Cymene | 11.454 | 7463 | 99-87-6 | 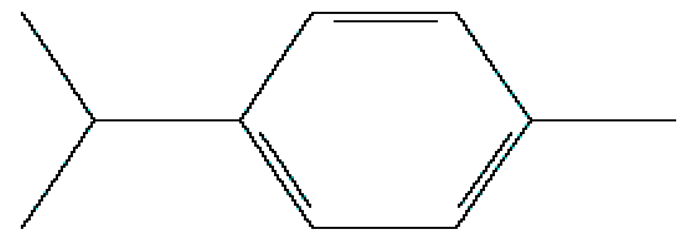 |
| 3 | γ-Terpinene | 12.389 | 7461 | 99-85-4 | 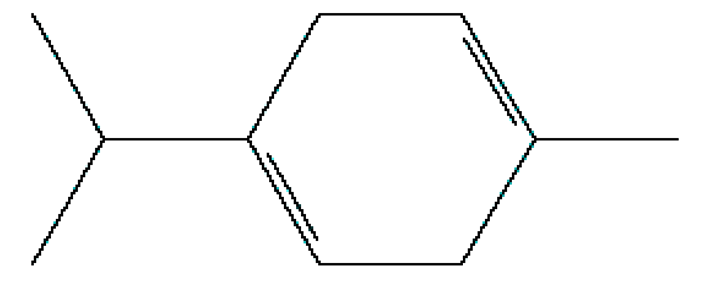 |
| 4 | Terpinolene | 13.178 | 11463 | 586-62-9 | 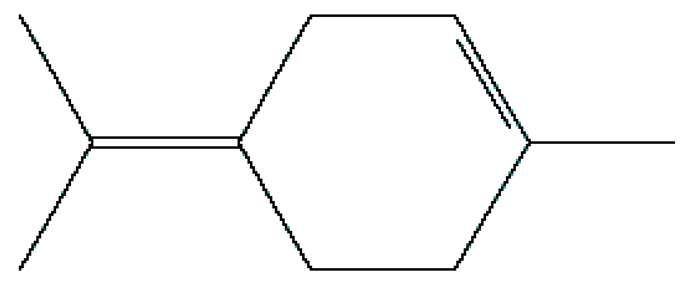 |
| 5 | Fenchone | 13.312 | 14525 | 7787-20-4 |  |
| 6 | Camphor | 14.939 | 2537 | 76-22-2 |  |
| 7 | Pinocamphone | 15.272 | 6427105 | 547-60-4 | 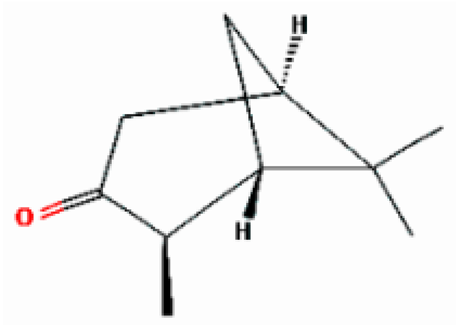 |
| 8 | Isopinocamphone | 15.707 | 84532 | 14575-93-0 | 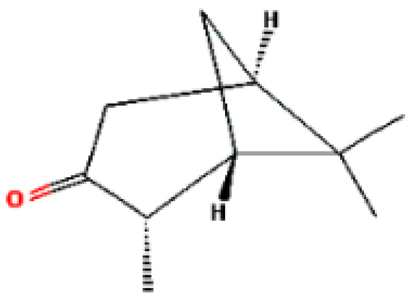 |
| 9 | Terpinen-4-ol | 15.820 | 11230 | 562-74-3 | 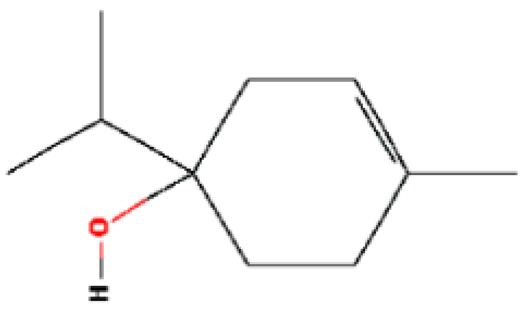 |
| 10 | α-Terpineol | 16.186 | 17100 | 98-55-5 | 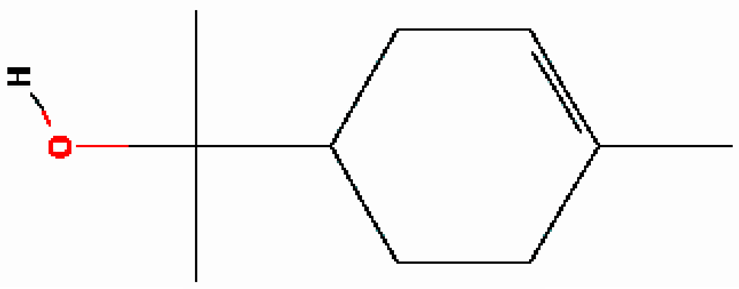 |
| 11 | (-)-Verbenone | 16.519 | 92874 | 1196-01-6 | 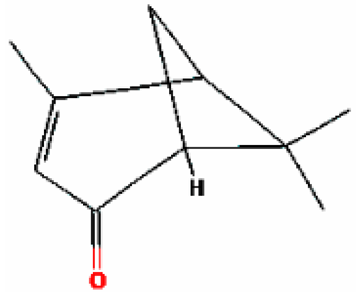 |
| Ligands | DhelOBP4 | DhelOBP5 | DhelOBP6 | DhelOBP14 | DhelOBP18 | DhelOBP20 |
|---|---|---|---|---|---|---|
| α-Fenchene | −5.9 | −5.4 | −6 | −6.4 | −6.3 | −6.5 |
| p-Cymene | −6.7 | −6.2 | −6.1 | −7.3 | −6.8 | −6.5 |
| γ-Terpinene | −6.3 | −6.1 | −6.3 | −7.2 | −6.7 | −6.6 |
| Terpinolene | −6.5 | −6.2 | −6.2 | −7.4 | −6.9 | −6.4 |
| Fenchone | −5.9 | −6.2 | −6.6 | −6.7 | −6.7 | −6.5 |
| Camphor | −5.6 | −5.5 | −6.3 | −6.3 | −6.1 | −6.8 |
| Pinocamphone | −6.1 | −6.2 | −6.8 | −7.0 | −6.5 | −6.8 |
| Isopinocamphone | −6.2 | −5.7 | −6.6 | −6.7 | −6.5 | −6.6 |
| Terpinen-4-ol | −6.2 | −6.1 | −6.2 | −7.1 | −6.9 | −6.3 |
| α-Terpineol | −6 | −6.3 | −6.3 | −7.1 | −6.7 | −6.3 |
| (-)-Verbenone | −6.2 | −6 | −6.5 | −6.4 | −6.4 | −6.7 |
| Ligands | DhelOBP4 | DhelOBP5 | DhelOBP6 | DhelOBP14 | DhelOBP18 | DhelOBP20 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| pH 7.4 | pH 5.0 | pH 7.4 | ||||||||||||
| IC50 | Ki | IC50 | Ki | IC50 | Ki | IC50 | Ki | IC50 | Ki | IC50 | Ki | IC50 | Ki | |
| p-Cymene | 13.0 ± 0.9 | 12.0 ± 0.9 | >50 | - | >50 | - | ud | ud | >50 | - | >50 | - | >50 | - |
| γ-Terpinene | 11.0 ± 0.3 | 10.1 ± 0.3 | >50 | - | >50 | - | ud | ud | >50 | - | >50 | - | >50 | - |
| Terpinolene | 15.7 ± 1.8 | 14.5 ± 1.7 | >50 | - | >50 | - | ud | ud | >50 | - | >50 | - | >50 | - |
| Fenchone | 23.8 ± 5.1 | 22.0 ± 4.7 | 47.4 ± 2.6 | 43.8 ± 2.4 | >50 | - | ud | ud | >50 | - | >50 | - | >50 | - |
| Camphor | 18.0 ± 2.4 | 16.6 ± 2.2 | >50 | - | >50 | - | ud | ud | >50 | - | >50 | - | >50 | - |
| Terpinen-4-ol | 27.8 ± 3.5 | 25.7 ± 3.2 | >50 | - | >50 | - | ud | ud | >50 | - | >50 | - | >50 | - |
| α-Terpineol | 10.4 ± 0.1 | 9.6 ± 0.1 | >50 | - | >50 | - | ud | ud | >50 | - | >50 | - | >50 | - |
| (-)-Verbenone | 23.1 ± 3.9 | 21.3 ± 3.6 | >50 | - | >50 | - | ud | ud | >50 | - | >50 | - | >50 | - |
| Ligands | Hydrophilic Residues | Hydrophobic Residues |
|---|---|---|
| α-Fenchene | - | Leu 11, Leu 32, Met 46, Ile 47, Met 50, Phe 126 |
| p-Cymene | Tyr 90 | Val 7, Phe 54, Val 56, Ala 68, Phe 71, Leu 72, Met 86 |
| γ-Terpinene | Tyr 90 | Val 7, Phe 54, Val 56, Ala 68, Phe 71, Leu 72, Met 86 |
| Terpinolene | His 51 | Val 7, Leu 11, Met 50, Phe 54, Phe 71, Phe 126 |
| Fenchone | Tyr 90 | Val 56, Ala 68, Phe 71, Leu 72, Met 86, Ile 89 |
| Camphor | His 51, Gly 113 | Ile 89, Leu 109, Ala 110, Ile 114, Pro 125, Phe 126 |
| Pinocamphone | - | Leu 11, Leu 32, Ile 47, Met 50, Phe 54, Phe 126 |
| Isopinocamphone | - | Leu 11, Leu 32, Met 46, Ile 47, Met 50, Phe 126 |
| Terpinen-4-ol | Tyr 90 | Phe 54, Val 56, Ala 68, Phe 71, Leu 72, Met 86 |
| α-Terpineol | His 51, Tyr 90, Cys 93 | Val 56, Ile 89, Ala 94, Leu 109, Ala 110, Phe 126 |
| (-)-Verbenone | - | Leu 11, Leu 32, Met 46, Ile 47, Met 50, Phe 126 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yi, S.-C.; Wu, Y.-H.; Yang, R.-N.; Li, D.-Z.; Abdelnabby, H.; Wang, M.-Q. A Highly Expressed Antennae Odorant-Binding Protein Involved in Recognition of Herbivore-Induced Plant Volatiles in Dastarcus helophoroides. Int. J. Mol. Sci. 2023, 24, 3464. https://doi.org/10.3390/ijms24043464
Yi S-C, Wu Y-H, Yang R-N, Li D-Z, Abdelnabby H, Wang M-Q. A Highly Expressed Antennae Odorant-Binding Protein Involved in Recognition of Herbivore-Induced Plant Volatiles in Dastarcus helophoroides. International Journal of Molecular Sciences. 2023; 24(4):3464. https://doi.org/10.3390/ijms24043464
Chicago/Turabian StyleYi, Shan-Cheng, Yu-Hang Wu, Rui-Nan Yang, Dong-Zhen Li, Hazem Abdelnabby, and Man-Qun Wang. 2023. "A Highly Expressed Antennae Odorant-Binding Protein Involved in Recognition of Herbivore-Induced Plant Volatiles in Dastarcus helophoroides" International Journal of Molecular Sciences 24, no. 4: 3464. https://doi.org/10.3390/ijms24043464
APA StyleYi, S.-C., Wu, Y.-H., Yang, R.-N., Li, D.-Z., Abdelnabby, H., & Wang, M.-Q. (2023). A Highly Expressed Antennae Odorant-Binding Protein Involved in Recognition of Herbivore-Induced Plant Volatiles in Dastarcus helophoroides. International Journal of Molecular Sciences, 24(4), 3464. https://doi.org/10.3390/ijms24043464







