Genome-Wide Re-Identification and Analysis of CrRLK1Ls in Tomato
Abstract
1. Introduction
2. Results
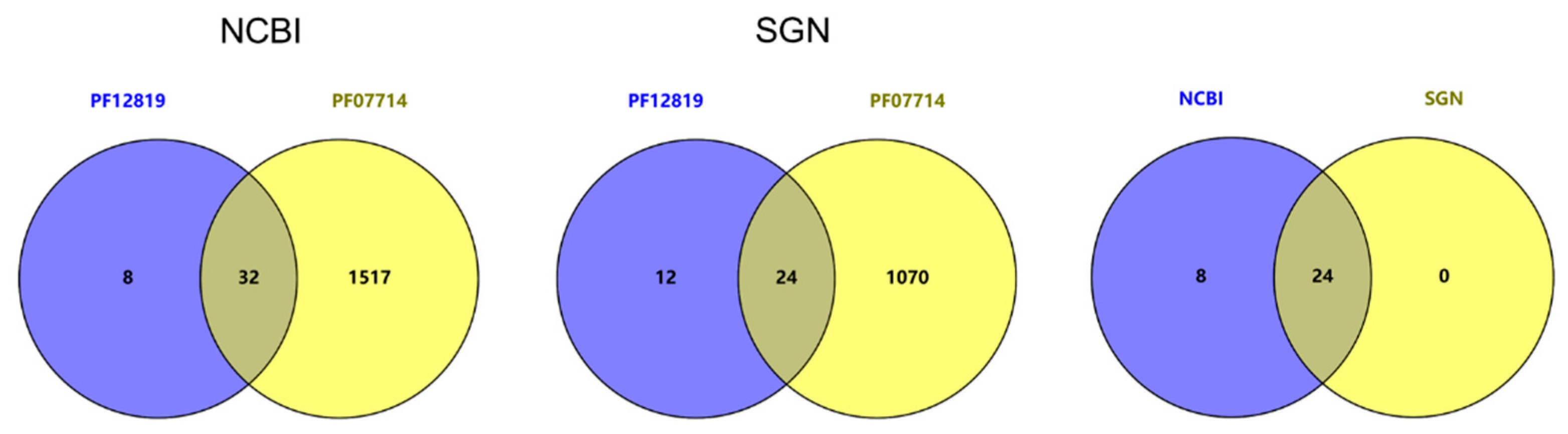
2.1. Identification of Tomato CrRLK1L Protein Kinases
2.2. Phylogenetic Analysis of the Tomato CrRLK1L Protein Kinases
2.3. Tomato CrRLK1L Gene Locations and Duplication on Tomato Chromosome
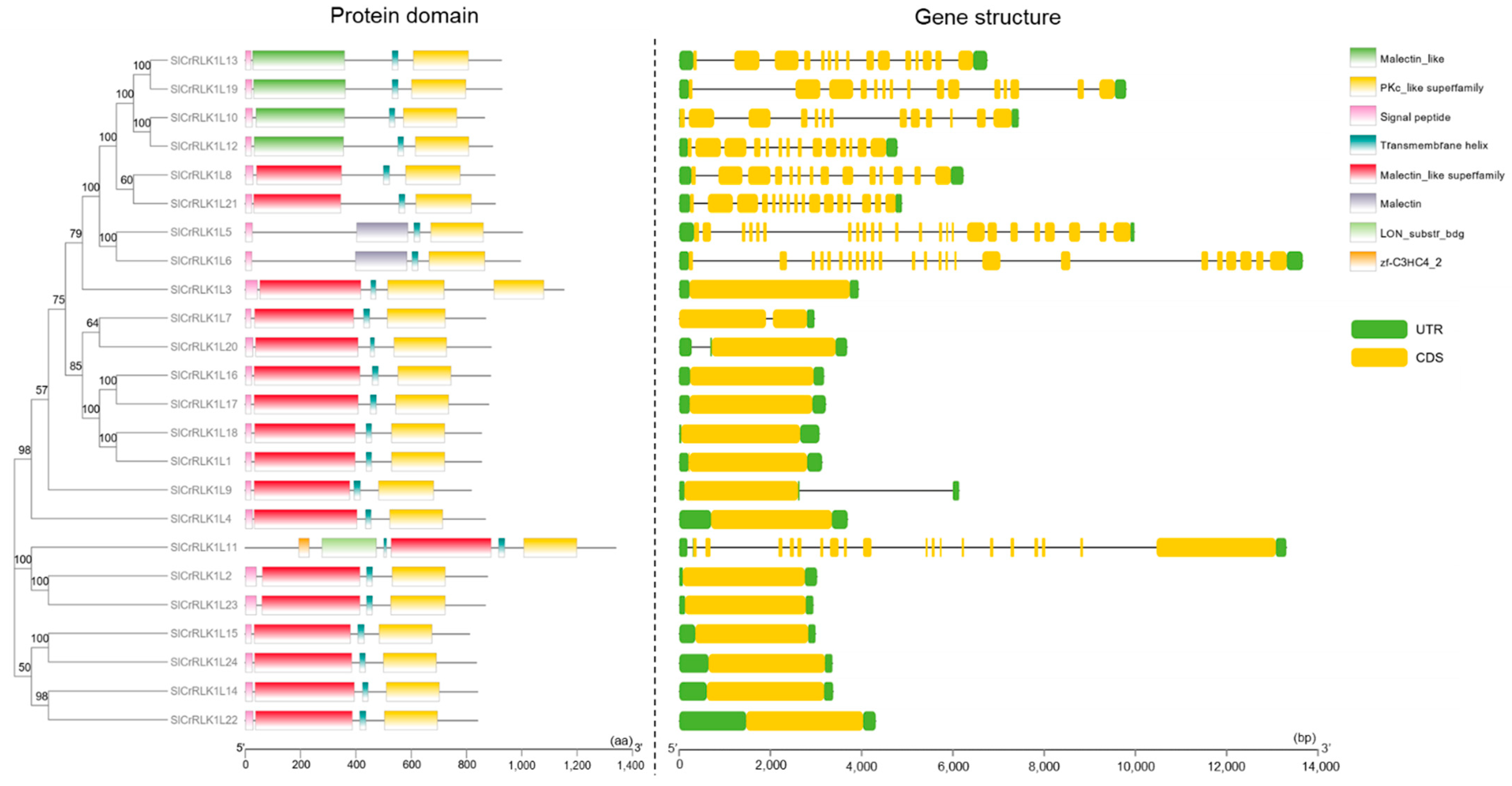
2.4. Tomato CrRLK1L Protein Domain and Gene Structure
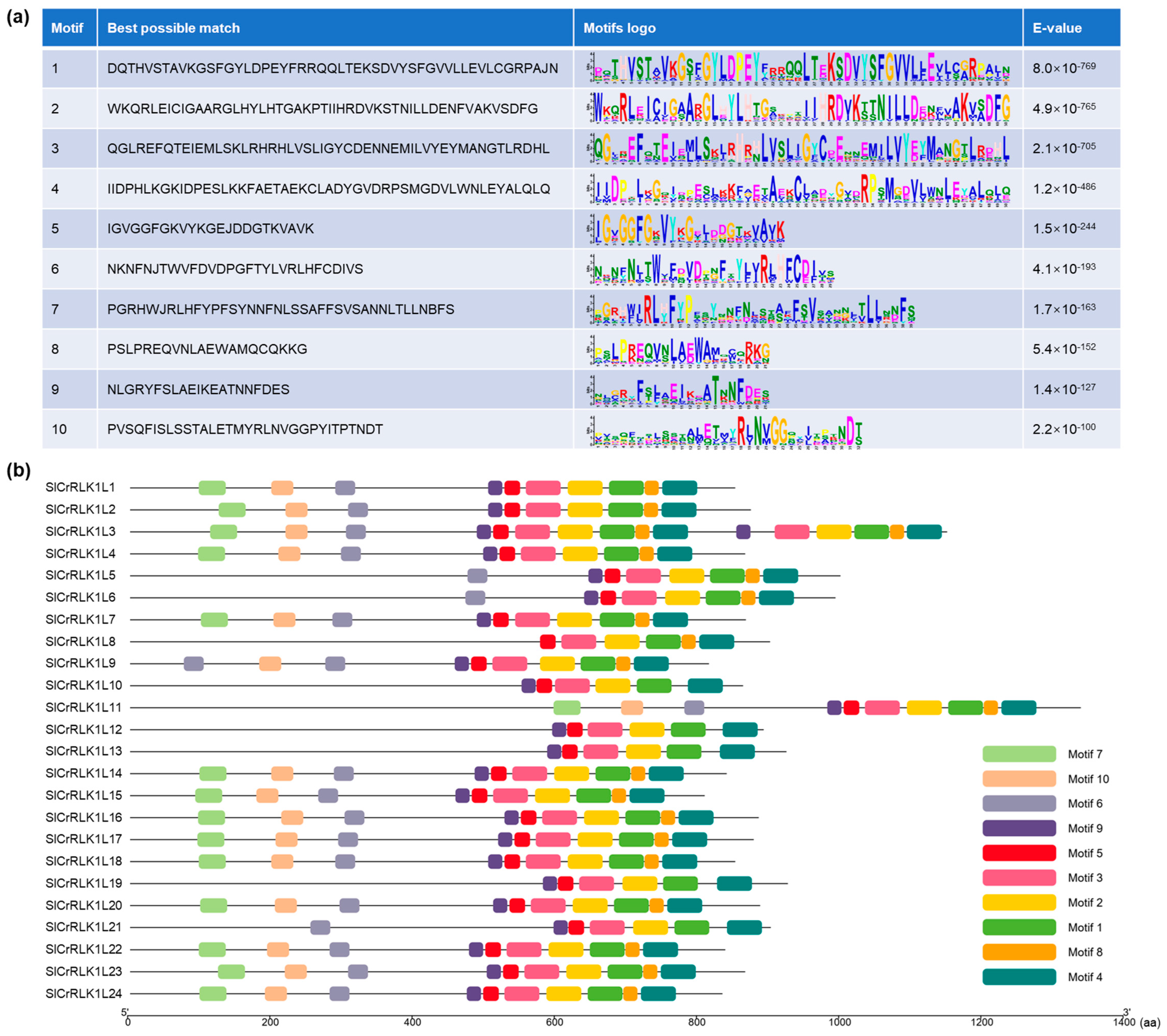
2.5. Prediction of SlCrRLK1L Conserved Protein Motifs
2.6. Subcellular Localization
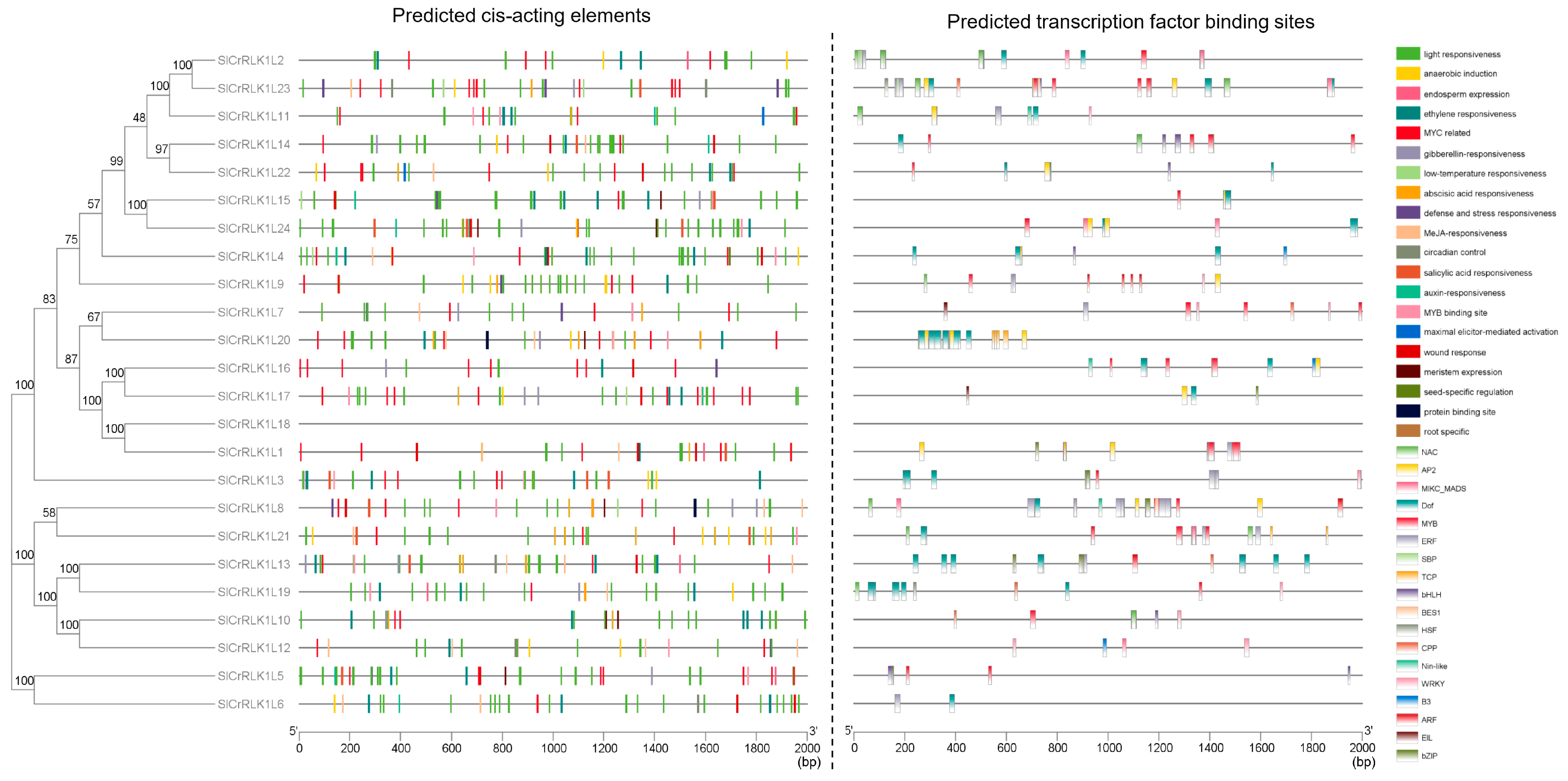
2.7. SlCrRLK1L Gene Promoter Analysis
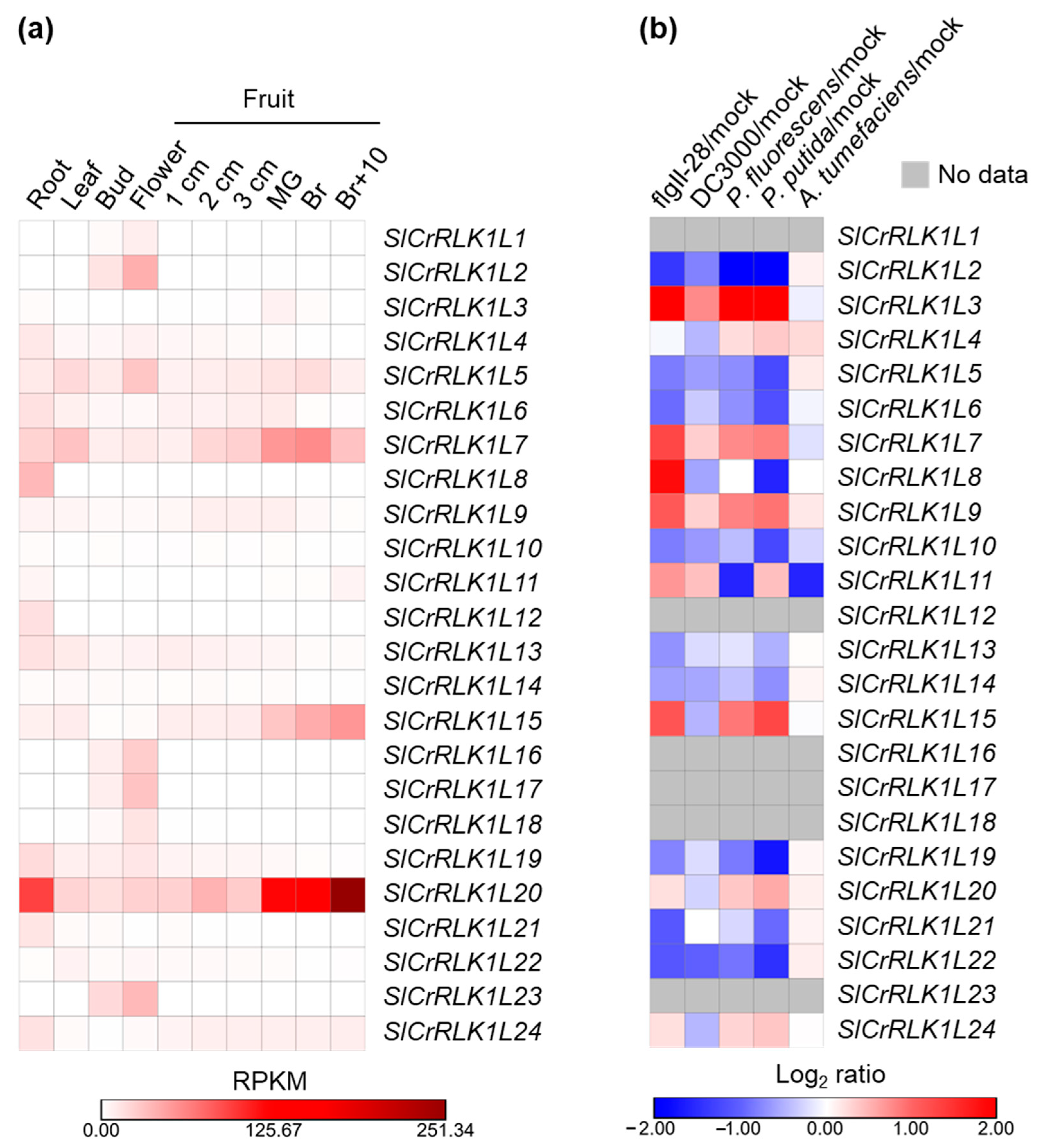
2.8. SlCrRLK1L Gene Expression Pattern Analysis
3. Discussion
4. Materials and Methods
4.1. Protein Identification and Phylogenetic Analysis
4.2. Antibody Preparation
4.3. Western Blot Assay
4.4. Gene Location and Collinearity Analysis
4.5. Subcellular Localization Analysis
4.6. Protein Domain and Gene Structure Analyses
4.7. Conserved Protein Motif Analysis
4.8. Promoter Analysis
4.9. Gene Expression Pattern Analysis
4.10. Accession Number
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ji, D.; Chen, T.; Zhang, Z.; Li, B.; Tian, S. Versatile Roles of the Receptor-like Kinase Feronia in Plant Growth, Development and Host-Pathogen Interaction. Int. J. Mol. Sci. 2020, 21, 7881. [Google Scholar] [CrossRef]
- Zhu, S.; Fu, Q.; Xu, F.; Zheng, H.; Yu, F. New Paradigms in Cell Adaptation: Decades of Discoveries on the CrRLK1L Receptor Kinase Signalling Network. New Phytol. 2021, 232, 1168–1183. [Google Scholar] [CrossRef]
- Dievart, A.; Gottin, C.; Périn, C.; Ranwez, V.; Chantret, N. Origin and Diversity of Plant Receptor-like Kinases. Annu. Rev. Plant Biol. 2020, 71, 131–156. [Google Scholar] [CrossRef]
- Schulze-Muth, P.; Irmler, S.; Schröder, G.; Schröder, J. Novel Type of Receptor-like Protein Kinase from a Higher Plant (Catharanthus Roseus). J. Biol. Chem. 1996, 271, 26684–26689. [Google Scholar] [CrossRef]
- Franck, C.M.; Westermann, J.; Boisson-Dernier, A. Plant Malectin-like Receptor Kinases: From Cell Wall Integrity to Immunity and Beyond. Annu. Rev. Plant. Biol. 2018, 69, 301–328. [Google Scholar] [CrossRef] [PubMed]
- Duan, Q.; Kita, D.; Li, C.; Cheung, A.Y.; Wu, H.M. FERONIA Receptor-like Kinase Regulates RHO GTPase Signaling of Root Hair Development. Proc. Natl. Acad. Sci. USA 2010, 107, 17821–17826. [Google Scholar] [CrossRef]
- Haruta, M.; Sabat, G.; Stecker, K.; Minkoff, B.B.; Sussman, M.R. A Peptide Hormone and Its Receptor Protein Kinase Regulate Plant Cell Expansion. Science 2014, 343, 408–411. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Wang, L.; Li, C.; Liu, Y.; Zhu, S.; Qi, Y.; Liu, X.; Lin, Q.; Luan, S.; Yu, F. Receptor Protein Kinase FERONIA Controls Leaf Starch Accumulation by Interacting with Glyceraldehyde-3-Phosphate Dehydrogenase. Biochem. Biophys. Res. Commun. 2015, 465, 77–82. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Wu, H.M.; Cheung, A.Y. FERONIA and Her Pals: Functions and Mechanisms. Plant Physiol. 2016, 171, 2379–2392. [Google Scholar] [CrossRef] [PubMed]
- Stegmann, M.; Monaghan, J.; Smakowska-Luzan, E.; Rovenich, H.; Lehner, A.; Holton, N.; Belkhadir, Y.; Zipfel, C. The Receptor Kinase FER Is a RALF-Regulated Scaffold Controlling Plant Immune Signaling. Science 2017, 355, 287–289. [Google Scholar] [CrossRef] [PubMed]
- Ge, Z.; Bergonci, T.; Zhao, Y.; Zou, Y.; Du, S.; Liu, M.; Luo, X.; Ruan, H.; García-Valencia, L.E.; Zhong, S.; et al. Arabidopsis Pollen Tube Integrity and Sperm Release Are Regulated by RALF-Mediated Signaling. Science 2017, 358, 1596–1600. [Google Scholar] [CrossRef] [PubMed]
- Huck, N.; Moore, J.M.; Federer, M.; Grossniklaus, U. The Arabidopsis Mutant Feronia Disrupts the Female Gametophytic Control of Pollen Tube Reception. Development 2003, 130, 2149–2159. [Google Scholar] [CrossRef]
- Zhang, X.; Yang, Z.; Wu, D.; Yu, F. RALF–FERONIA Signaling: Linking Plant Immune Response with Cell Growth. Plant Commun. 2020, 1, 100084. [Google Scholar] [CrossRef]
- Deslauriers, S.D.; Larsen, P.B. FERONIA Is a Key Modulator of Brassinosteroid and Ethylene Responsiveness in Arabidopsis Hypocotyls. Mol. Plant 2010, 3, 626–640. [Google Scholar] [CrossRef] [PubMed]
- Mao, D.; Yu, F.; Li, J.; Van de Poel, B.; Tan, D.; Li, J.; Liu, Y.; Li, X.; Dong, M.; Chen, L.; et al. FERONIA Receptor Kinase Interacts with S-Adenosylmethionine Synthetase and Suppresses S-Adenosylmethionine Production and Ethylene Biosynthesis in Arabidopsis. Plant Cell Environ. 2015, 38, 2566–2574. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yu, F.; Liu, Y.; Du, C.; Li, X.; Zhu, S.; Wang, X.; Lan, W.; Rodriguez, P.L.; Liu, X.; et al. FERONIA Interacts with ABI2-Type Phosphatases to Facilitate Signaling Cross-Talk between Abscisic Acid and RALF Peptide in Arabidopsis. Proc. Natl. Acad. Sci. USA 2016, 113, E5519–E5527. [Google Scholar] [CrossRef]
- Guo, H.; Nolan, T.M.; Song, G.; Liu, S.; Xie, Z.; Chen, J.; Schnable, P.S.; Walley, J.W.; Yin, Y. FERONIA Receptor Kinase Contributes to Plant Immunity by Suppressing Jasmonic Acid Signaling in Arabidopsis thaliana. Curr. Biol. 2018, 28, 3316–3324. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Chakravorty, D.; Assmann, S.M. The G Protein β-Subunit, AGB1, Interacts with FERONIA in RALF1-Regulated Stomatal Movement. Plant Physiol. 2018, 176, 2426–2440. [Google Scholar] [CrossRef] [PubMed]
- Ge, Z.; Dresselhaus, T.; Qu, L.J. How CrRLK1L Receptor Complexes Perceive RALF Signals. Trends Plant Sci. 2019, 24, 978–981. [Google Scholar] [CrossRef]
- Xiao, Y.; Stegmann, M.; Han, Z.; DeFalco, T.A.; Parys, K.; Xu, L.; Belkhadir, Y.; Zipfel, C.; Chai, J. Mechanisms of RALF Peptide Perception by a Heterotypic Receptor Complex. Nature 2019, 572, 270–274. [Google Scholar] [CrossRef]
- Hématy, K.; Sado, P.-E.; Van Tuinen, A.; Rochange, S.; Desnos, T.; Balzergue, S.; Pelletier, S.; Renou, J.-P.; Höfte, H. A Receptor-like Kinase Mediates the Response of Arabidopsis Cells to the Inhibition of Cellulose Synthesis. Curr. Biol. 2007, 17, 922–931. [Google Scholar] [CrossRef]
- Guo, H.; Li, L.; Ye, H.; Yu, X.; Algreen, A.; Yin, Y. Three Related Receptor-like Kinases Are Required for Optimal Cell Elongation in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2009, 106, 7648–7653. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, S.; Murata, T.; Sakurai-Ozato, N.; Kubo, M.; Demura, T.; Fukuda, H.; Hasebe, M. ANXUR1 and 2, Sister Genes to FERONIA/SIRENE, Are Male Factors for Coordinated Fertilization. Curr. Biol. 2009, 19, 1327–1331. [Google Scholar] [CrossRef]
- Richter, J.; Watson, J.M.; Stasnik, P.; Borowska, M.; Neuhold, J.; Berger, M.; Stolt-Bergner, P.; Schoft, V.; Hauser, M.T. Multiplex Mutagenesis of Four Clustered CrRLK1L with CRISPR/Cas9 Exposes Their Growth Regulatory Roles in Response to Metal Ions. Sci. Rep. 2018, 8, 12182. [Google Scholar] [CrossRef] [PubMed]
- Kessler, S.A.; Shimosato-Asano, H.; Keinath, N.F.; Wuest, S.E.; Ingram, G.; Panstruga, R.; Grossniklaus, U. Conserved Molecular Components for Pollen Tube Reception and Fungal Invasion. Science 2010, 330, 968–971. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Wang, L.; Cui, Y.; He, L.; Qi, Y.; Zhang, J.; Lin, J.; Liao, H.; Lin, Q.; Yang, T.; et al. Two FERONIA-like Receptor (FLR) Genes Are Required to Maintain Architecture, Fertility, and Seed Yield in Rice. Mol. Breeding 2016, 36, 151. [Google Scholar] [CrossRef]
- Pu, C.X.; Han, Y.F.; Zhu, S.; Song, F.Y.; Zhao, Y.; Wang, C.; Zhang, Y.; Yang, Q.; Wang, J.; Bu, S.-L.; et al. The Rice Receptor-like Kinases DWARF AND RUNTISH SPIKELET1 and 2 Repress Cell Death and Affect Sugar Utilization during Reproductive Development. Plant Cell 2017, 29, 70–89. [Google Scholar] [CrossRef]
- Liu, L.; Zheng, C.; Kuang, B.; Wei, L.; Yan, L.; Wang, T. Receptor-like Kinase RUPO Interacts with Potassium Transporters to Regulate Pollen Tube Growth and Integrity in Rice. PLoS Genet. 2016, 12, e1006085. [Google Scholar] [CrossRef] [PubMed]
- Jia, M.; Du, P.; Ding, N.; Zhang, Q.; Xing, S.; Wei, L.; Zhao, Y.; Mao, W.; Li, J.; Li, B.; et al. Two FERONIA-like Receptor Kinases Regulate Apple Fruit Ripening by Modulating Ethylene Production. Front. Plant Sci. 2017, 8, 1406. [Google Scholar] [CrossRef]
- Ji, D.; Cui, X.; Qin, G.; Chen, T.; Tian, S. SlFERL Interacts with S-Adenosylmethionine Synthetase to Regulate Fruit Ripening. Plant Physiol. 2020, 184, 2168–2181. [Google Scholar] [CrossRef]
- Wang, D.; Liang, X.; Bao, Y.; Yang, S.; Zhang, X.; Yu, H.; Zhang, Q.; Xu, G.; Feng, X.; Dou, D. A Malectin-like Receptor Kinase Regulates Cell Death and Pattern-triggered Immunity in Soybean. EMBO Rep. 2020, 21, e50442. [Google Scholar] [CrossRef]
- Kou, X.; Qi, K.; Qiao, X.; Yin, H.; Liu, X.; Zhang, S.; Wu, J. Evolution, Expression Analysis, and Functional Verification of Catharanthus Roseus RLK1-like Kinase (CrRLK1L) Family Proteins in Pear (Pyrus Bretchneideri). Genomics 2017, 109, 290–301. [Google Scholar] [CrossRef]
- Wang, Z.-Q.; Yu, T.-F.; Sun, G.-Z.; Zheng, J.-C.; Chen, J.; Zhou, Y.-B.; Chen, M.; Ma, Y.-Z.; Wei, W.-L.; Xu, Z.-S. Genome-Wide Analysis of the Catharanthus Roseus RLK1-Like in Soybean and GmCrRLK1L20 Responds to Drought and Salt Stresses. Front. Plant Sci. 2021, 12, 614909. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Guo, C.; Wang, Q.; Li, Z.; Cai, J.; Wu, D.; Li, Y.; Yang, A.; Guo, Y.; Gao, J.; et al. Systematic Analysis of Tobacco CrRLK1L Family Genes and Functional Identification of NtCrRLK1L47 in Environmental Stresses. Front. Plant Sci. 2022, 13, 838857. [Google Scholar] [CrossRef]
- Jiang, W.; Li, C.; Li, L.; Li, Y.; Wang, Z.; Yu, F.; Yi, F.; Zhang, J.; Zhu, J.-K.; Zhang, H.; et al. Genome-Wide Analysis of CqCrRLK1L and CqRALF Gene Families in Chenopodium Quinoa and Their Roles in Salt Stress Response. Front. Plant Sci. 2022, 13, 918594. [Google Scholar] [CrossRef]
- Tomato Genome Consortium. The Tomato Genome Sequence Provides Insights into Fleshy Fruit Evolution. Nature 2012, 485, 635–641. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, T.; Deguchi, M.; Brustolini, O.J.B.; Santos, A.A.; Silva, F.F.; Fontes, E.P.B. The Tomato RLK Superfamily: Phylogeny and Functional Predictions about the Role of the LRRII-RLK Subfamily in Antiviral Defense. BMC Plant Biol. 2012, 12, 229. [Google Scholar] [CrossRef] [PubMed]
- Cai, J.; Qin, G.; Chen, T.; Tian, S. The Mode of Action of Remorin1 in Regulating Fruit Ripening at Transcriptional and Post-Transcriptional Levels. New Phytol. 2018, 219, 1406–1420. [Google Scholar] [CrossRef] [PubMed]
- Boisson-Dernier, A.; Roy, S.; Kritsas, K.; Grobei, M.A.; Jaciubek, M.; Schroeder, J.I.; Grossniklaus, U. Disruption of the Pollen-Expressed FERONIA Homologs ANXUR1 and ANXUR2 Triggers Pollen Tube Discharge. Development 2009, 136, 3279–3288. [Google Scholar] [CrossRef] [PubMed]
- Zuo, C.; Zhang, W.; Ma, Z.; Chu, M.; Mao, J.; An, Z.; Chen, B. Genome-Wide Identification and Expression Analysis of the CrRLK1L Gene Family in Apple (Malus Domestica). Plant Mol. Biol. Rep. 2018, 36, 844–857. [Google Scholar] [CrossRef]
- Rosli, H.G.; Zheng, Y.; Pombo, M.A.; Zhong, S.; Bombarely, A.; Fei, Z.; Collmer, A.; Martin, G.B. Transcriptomics-Based Screen for Genes Induced by Flagellin and Repressed by Pathogen Effectors Identifies a Cell Wall-Associated Kinase Involved in Plant Immunity. Genome Biol. 2013, 14, R139. [Google Scholar] [CrossRef]
- Lindner, H.; Müller, L.M.; Boisson-Dernier, A.; Grossniklaus, U. CrRLK1L Receptor-like Kinases: Not Just Another Brick in the Wall. Curr. Opin. Plant Biol. 2012, 15, 659–669. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, Q.-N.; Lee, Y.-S.; Cho, L.-H.; Jeong, H.-J.; An, G.; Jung, K.-H. Genome-Wide Identification and Analysis of Catharanthus Roseus RLK1-like Kinases in Rice. Planta 2015, 241, 603–613. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Jia, M.; Xing, Y.; Qin, L.; Li, B.; Jia, W. Genome-Wide Identification and Expression Analysis of MRLK Family Genes Associated with Strawberry (Fragaria Vesca) Fruit Ripening and Abiotic Stress Responses. PLoS ONE 2016, 11, e0163647. [Google Scholar] [CrossRef]
- Luo, C.; Sun, Q.; Zhang, F.; Zhang, D.; Liu, C.; Wu, Q.; Shu, B. Genome-Wide Identification and Expression Analysis of the Citrus Malectin Domain-Containing Receptor-like Kinases in Response to Arbuscular Mycorrhizal Fungi Colonization and Drought. Hortic. Environ. Biotechnol. 2020, 61, 891–901. [Google Scholar] [CrossRef]
- Rao, S.; Wu, X.; Zheng, H.; Lu, Y.; Peng, J.; Wu, G.; Chen, J.; Yan, F. Genome-Wide Identification and Analysis of Catharanthus Roseus RLK1-like Kinases in Nicotiana benthamiana. BMC Plant Biol. 2021, 21, 425. [Google Scholar] [CrossRef]
- Kumar, V.; Donev, E.N.; Barbut, F.R.; Kushwah, S.; Mannapperuma, C.; Urbancsok, J.; Mellerowicz, E.J. Genome-Wide Identification of Populus Malectin/Malectin-like Domain-Containing Proteins and Expression Analyses Reveal Novel Candidates for Signaling and Regulation of Wood Development. Front. Plant Sci. 2020, 11, 588846. [Google Scholar] [CrossRef]
- Niu, E.; Cai, C.; Zheng, Y.; Shang, X.; Fang, L.; Guo, W. Genome-Wide Analysis of CrRLK1L Gene Family in Gossypium and Identification of Candidate CrRLK1L Genes Related to Fiber Development. Mol. Genet. Genom. 2016, 291, 1137–1154. [Google Scholar] [CrossRef]
- Tang, L.; Wang, Y.; Wang, W.; Deng, X.; Wang, X. Genome-Wide Identification of CrRLK1L Gene Family and Desiccation-Induced Expression Profiles in Boea hygrometrica. Curr. Plant Biol. 2022, 31, 100256. [Google Scholar] [CrossRef]
- Yu, H.-F.; Zhang, W.-N.; Kang, Y.-C.; Fan, Y.-L.; Yang, X.-Y.; Shi, M.-F.; Zhang, R.-Y.; Zhang, J.-L.; Qin, S.-H. Genome-Wide Identification and Expression Patterns in Response to Signals from Phytophthora Infestans of CrRLK1Ls Gene Family in Potato. Acta Agronom. Sin. 2021, 48, 249–258. [Google Scholar] [CrossRef]
- Amborella Genome Project; Albert, V.A.; Barbazuk, W.B.; de Pamphilis, C.W.; Der, J.P.; Leebens-Mack, J.; Ma, H.; Palmer, J.D.; Rounsley, S.; Sankoff, D.; et al. The Amborella Genome and the Evolution of Flowering Plants. Science 2013, 342, 1241089. [Google Scholar] [CrossRef]
- Sasaki, G.; Katoh, K.; Hirose, N.; Suga, H.; Kuma, K.; Miyata, T.; Su, Z.-H. Multiple Receptor-like Kinase CDNAs from Liverwort Marchantia Polymorpha and Two Charophycean Green Algae, Closterium Ehrenbergii and Nitella Axillaris: Extensive Gene Duplications and Gene Shufflings in the Early Evolution of Streptophytes. Gene 2007, 401, 135–144. [Google Scholar] [CrossRef]
- Rensing, S.A.; Lang, D.; Zimmer, A.D.; Terry, A.; Salamov, A.; Shapiro, H.; Nishiyama, T.; Perroud, P.-F.; Lindquist, E.A.; Kamisugi, Y.; et al. The Physcomitrella Genome Reveals Evolutionary Insights into the Conquest of Land by Plants. Science 2008, 319, 64–69. [Google Scholar] [CrossRef] [PubMed]
- Banks, J.A.; Nishiyama, T.; Hasebe, M.; Bowman, J.L.; Gribskov, M.; de Pamphilis, C.; Albert, V.A.; Aono, N.; Aoyama, T.; Ambrose, B.A.; et al. The Selaginella Genome Identifies Genetic Changes Associated with the Evolution of Vascular Plants. Science 2011, 332, 960–963. [Google Scholar] [CrossRef] [PubMed]
- Nystedt, B.; Street, N.R.; Wetterbom, A.; Zuccolo, A.; Lin, Y.-C.; Scofield, D.G.; Vezzi, F.; Delhomme, N.; Giacomello, S.; Alexeyenko, A.; et al. The Norway Spruce Genome Sequence and Conifer Genome Evolution. Nature 2013, 497, 579–584. [Google Scholar] [CrossRef]
- Hirano, N.; Marukawa, Y.; Abe, J.; Hashiba, S.; Ichikawa, M.; Tanabe, Y.; Ito, M.; Nishii, I.; Tsuchikane, Y.; Sekimoto, H. A Receptor-like Kinase, Related to Cell Wall Sensor of Higher Plants, Is Required for Sexual Reproduction in the Unicellular Charophycean Alga, Closterium Peracerosum–Strigosum–Littorale Complex. Plant Cell Physiol. 2015, 56, 1456–1462. [Google Scholar] [CrossRef]
- Jing, X.-Q.; Shalmani, A.; Zhou, M.-R.; Shi, P.-T.; Muhammad, I.; Shi, Y.; Sharif, R.; Li, W.-Q.; Liu, W.-T.; Chen, K.-M. Genome-Wide Identification of Malectin/Malectin-like Domain Containing Protein Family Genes in Rice and Their Expression Regulation under Various Hormones, Abiotic Stresses, and Heavy Metal Treatments. J. Plant Growth Regul. 2020, 39, 492–506. [Google Scholar] [CrossRef]
- Fernandez-Pozo, N.; Menda, N.; Edwards, J.D.; Saha, S.; Tecle, I.Y.; Strickler, S.R.; Bombarely, A.; Fisher-York, T.; Pujar, A.; Foerster, H.; et al. The Sol Genomics Network (SGN)—From Genotype to Phenotype to Breeding. Nucleic Acids Res. 2015, 43, D1036–D1041. [Google Scholar] [CrossRef]
- Berardini, T.Z.; Reiser, L.; Li, D.; Mezheritsky, Y.; Muller, R.; Strait, E.; Huala, E. The Arabidopsis Information Resource: Making and Mining the “Gold Standard” Annotated Reference Plant Genome: Tair: Making and Mining the “Gold Standard” Plant Genome. Genesis 2015, 53, 474–485. [Google Scholar] [CrossRef]
- Yates, A.D.; Allen, J.; Amode, R.M.; Azov, A.G.; Barba, M.; Becerra, A.; Bhai, J.; Campbell, L.I.; Carbajo Martinez, M.; Chakiachvili, M.; et al. Ensembl Genomes 2022: An Expanding Genome Resource for Non-Vertebrates. Nucleic Acids Res. 2022, 50, D996–D1003. [Google Scholar] [CrossRef]
- Mistry, J.; Chuguransky, S.; Williams, L.; Qureshi, M.; Salazar, G.A.; Sonnhammer, E.L.L.; Tosatto, S.C.E.; Paladin, L.; Raj, S.; Richardson, L.J.; et al. Pfam: The Protein Families Database in 2021. Nucleic Acids Res. 2021, 49, D412–D419. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.-C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein Identification and Analysis Tools in the ExPASy Server. In 2-D Proteome Analysis Protocols; Humana Press: Totowa, NJ, USA, 1998; Volume 112, pp. 531–552. ISBN 978-1-59259-584-6. [Google Scholar]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree of Life (ITOL): An Online Tool for Phylogenetic Tree Display and Annotation. Bioinformatics 2007, 23, 127–128. [Google Scholar] [CrossRef]
- Yu, C.-S.; Chen, Y.-C.; Lu, C.-H.; Hwang, J.-K. Prediction of Protein Subcellular Localization. Proteins 2006, 64, 643–651. [Google Scholar] [CrossRef]
- Blum, T.; Briesemeister, S.; Kohlbacher, O. MultiLoc2: Integrating Phylogeny and Gene Ontology Terms Improves Subcellular Protein Localization Prediction. BMC Bioinform. 2009, 10, 274. [Google Scholar] [CrossRef]
- Chou, K.-C.; Shen, H.-B. Plant-MPLoc: A Top-down Strategy to Augment the Power for Predicting Plant Protein Subcellular Localization. PLoS ONE 2010, 5, e11335. [Google Scholar] [CrossRef]
- Lu, S.; Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; Gwadz, M.; Hurwitz, D.I.; Marchler, G.H.; Song, J.S.; et al. CDD/SPARCLE: The Conserved Domain Database in 2020. Nucleic Acids Res. 2020, 48, D265–D268. [Google Scholar] [CrossRef]
- Hallgren, J.; Tsirigos, K.D.; Pedersen, M.D.; Almagro Armenteros, J.J.; Marcatili, P.; Nielsen, H.; Krogh, A.; Winther, O. DeepTMHMM Predicts Alpha and Beta Transmembrane Proteins Using Deep Neural Networks. bioRxiv 2022. [Google Scholar] [CrossRef]
- Bailey, T.L.; Elkan, C. Fitting a Mixture Model by Expectation Maximization to Discover Motifs in Biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar]
- Lescot, M. PlantCARE, a Database of Plant Cis-Acting Regulatory Elements and a Portal to Tools for in Silico Analysis of Promoter Sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Tian, F.; Yang, D.-C.; Meng, Y.-Q.; Jin, J.; Gao, G. PlantRegMap: Charting Functional Regulatory Maps in Plants. Nucleic Acids Res. 2019, 48, gkz1020. [Google Scholar] [CrossRef]



| Protein Name | Protein ID in NCBI | Protein Length (aa) | Molecular Weight (Da) | Theoretical pI | GRAVY | Matched SGN Locus | Annotation Difference in NCBI and SGN |
|---|---|---|---|---|---|---|---|
| SlCrRLK1L1 | XP_025886195.1 | 854 | 95,039.67 | 6.86 | −0.268 | Solyc01g059910 | Y |
| SlCrRLK1L2 | XP_004230878.1 | 876 | 96,273.18 | 5.82 | −0.212 | Solyc01g109950 | N |
| SlCrRLK1L3 | XP_019067666.1 | 1152 | 129,597.94 | 6.93 | −0.174 | Solyc02g014030 | N |
| SlCrRLK1L4 | XP_004233885.1 | 868 | 96,438.64 | 5.49 | −0.28 | Solyc02g069970 | Y |
| SlCrRLK1L5 | XP_004233025.1 | 1002 | 112,230.67 | 6.36 | −0.312 | Solyc02g071860 | Y |
| SlCrRLK1L6 | XP_010316862.1 | 995 | 111,071.97 | 5.79 | −0.108 | Solyc02g071880 | N |
| SlCrRLK1L7 | XP_004232151.1 | 869 | 97,152.91 | 6.14 | −0.234 | Solyc02g089090 | Y |
| SlCrRLK1L8 | NP_001234869.1 | 903 | 101,461.32 | 8.92 | −0.197 | Solyc02g091590 | N |
| SlCrRLK1L9 | XP_004234657.2 | 817 | 91,323.59 | 5.9 | −0.168 | Solyc03g044160 | Y |
| SlCrRLK1L10 | XP_010318169.1 | 865 | 97,835 | 5.41 | −0.201 | Solyc03g093380 | N |
| SlCrRLK1L11 | XP_025886103.1 | 1340 | 150,300.63 | 6.39 | −0.226 | Solyc03g115710 | Y |
| SlCrRLK1L12 | XP_010318523.1 | 894 | 99,669.44 | 5.49 | −0.16 | Solyc03g121230 | N |
| SlCrRLK1L13 | XP_004239170.1 | 926 | 103,366.23 | 5.88 | −0.267 | Solyc05g014240 | N |
| SlCrRLK1L14 | XP_004240198.2 | 840 | 92,846.45 | 6.26 | 0.021 | Solyc05g054680 | N |
| SlCrRLK1L15 | XP_004239762.1 | 811 | 90,461.22 | 5.66 | −0.079 | Solyc05g054860 | N |
| SlCrRLK1L16 | XP_004240568.1 | 887 | 97,458.64 | 5.73 | −0.264 | Solyc06g009540 | N |
| SlCrRLK1L17 | XP_004240569.1 | 880 | 97,075.96 | 6.33 | −0.246 | Solyc06g009550 | Y |
| SlCrRLK1L18 | XP_004243035.1 | 854 | 94,945.25 | 6.36 | −0.259 | Solyc07g008400 | N |
| SlCrRLK1L19 | XP_004246699.1 | 928 | 102,646.36 | 5.82 | −0.212 | Solyc09g007280 | N |
| SlCrRLK1L20 | XP_004246282.1 | 889 | 97,348.77 | 5.78 | −0.22 | Solyc09g015830 | Y |
| SlCrRLK1L21 | XP_004247083.1 | 904 | 101,257.01 | 5.48 | −0.107 | Solyc09g060110 | N |
| SlCrRLK1L22 | XP_010327142.1 | 840 | 91,947.64 | 5.26 | 0.032 | Solyc10g006870 | N |
| SlCrRLK1L23 | XP_004248695.3 | 868 | 96,419.44 | 6.09 | −0.221 | Solyc10g054050 | Y |
| SlCrRLK1L24 | XP_004251295.1 | 836 | 91,944.54 | 5.84 | −0.1 | Solyc11g072910 | Y |
| Species | Number of CrRLK1Ls | Reference |
|---|---|---|
| Arabidopsis thaliana | 17 | [42] |
| Oryza sativa (rice) | 16 | [43] |
| Malus domestica (apple) | 74 | [40] |
| Fragaria vesca (strawberry) | 62 | [44] |
| Citrus sinensis ‘Valencia’ (Citrus) | 47 | [45] |
| Glycine max L. (soybean) | 38 | [33] |
| Nicotiana tabacum L. (tobacco) | 48 | [34] |
| Nicotiana benthamiana | 31 | [46] |
| Chenopodium quinoa | 26 | [35] |
| Pyrus bretchneideri (pear) | 26 | [32] |
| Populus trichocarpa (black cottonwood) | 42 | [47] |
| Gossypium raimondii, G. arboreum, and G. hirsutum TM-1 (cotton) | 44 | [48] |
| Boea hygrometrica | 18 | [49] |
| Solanum tuberosum (potato) | 17 | [50] |
| Solanum lycopersicum (tomato) | 24 | This study |
| Amborella trichopoda | 9 | [51] |
| Marchantia polymorpha | 1 | [52] |
| Physcomitrella patens | 6 | [53] |
| Selaginella moellendorffii | 2 | [54] |
| Picea abies | 7 | [55] |
| Closterium peracerosum-strigosumlittorale complex | 1 | [56] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, W.; Liu, X.; Chen, K.; Yu, X.; Ji, D. Genome-Wide Re-Identification and Analysis of CrRLK1Ls in Tomato. Int. J. Mol. Sci. 2023, 24, 3142. https://doi.org/10.3390/ijms24043142
Ma W, Liu X, Chen K, Yu X, Ji D. Genome-Wide Re-Identification and Analysis of CrRLK1Ls in Tomato. International Journal of Molecular Sciences. 2023; 24(4):3142. https://doi.org/10.3390/ijms24043142
Chicago/Turabian StyleMa, Wenpeng, Xin Liu, Kai Chen, Xinlong Yu, and Dongchao Ji. 2023. "Genome-Wide Re-Identification and Analysis of CrRLK1Ls in Tomato" International Journal of Molecular Sciences 24, no. 4: 3142. https://doi.org/10.3390/ijms24043142
APA StyleMa, W., Liu, X., Chen, K., Yu, X., & Ji, D. (2023). Genome-Wide Re-Identification and Analysis of CrRLK1Ls in Tomato. International Journal of Molecular Sciences, 24(4), 3142. https://doi.org/10.3390/ijms24043142






