Genome-Wide Identification of PEBP Gene Family in Two Dendrobium Species and Expression Patterns in Dendrobium chrysotoxum
Abstract
:1. Introduction
2. Results
2.1. Identification and Physicochemical Properties
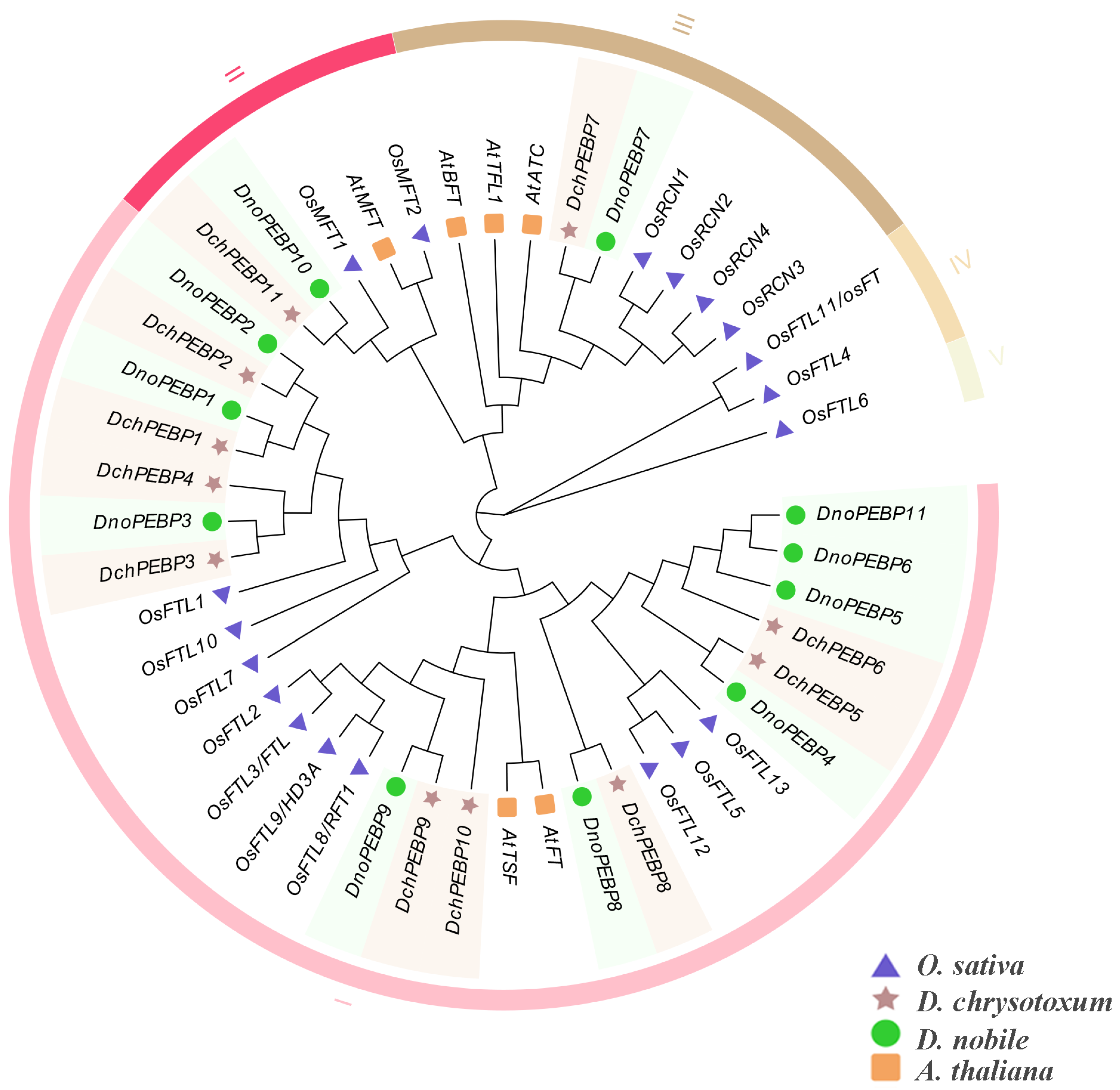
2.2. Phylogenetic and Classification of PEBP Proteins
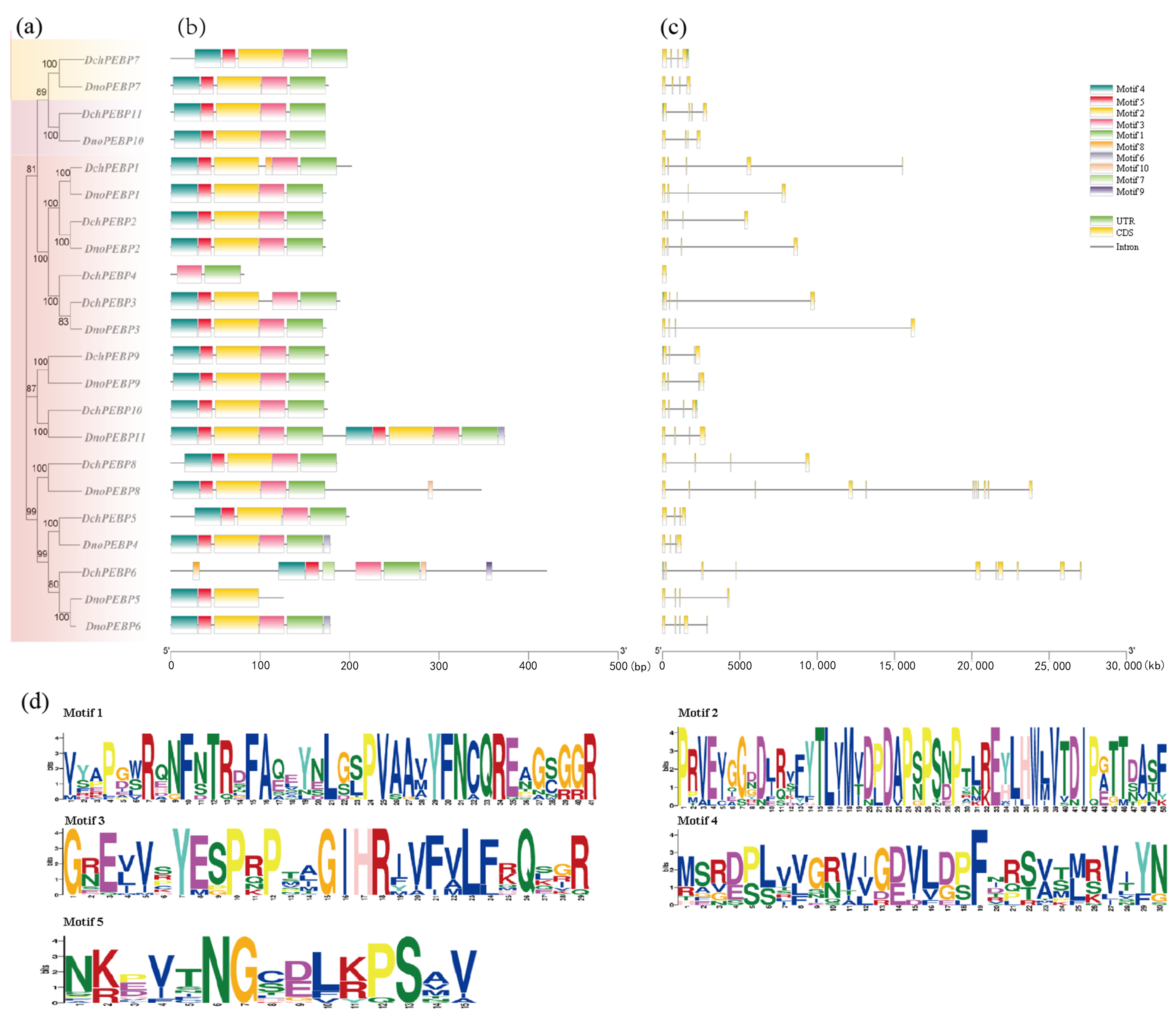
2.3. Phylogenetic Analysis and Conserved Motifs of PEBP Proteins
2.4. Genes Expression Analysis of the PEBP Genes
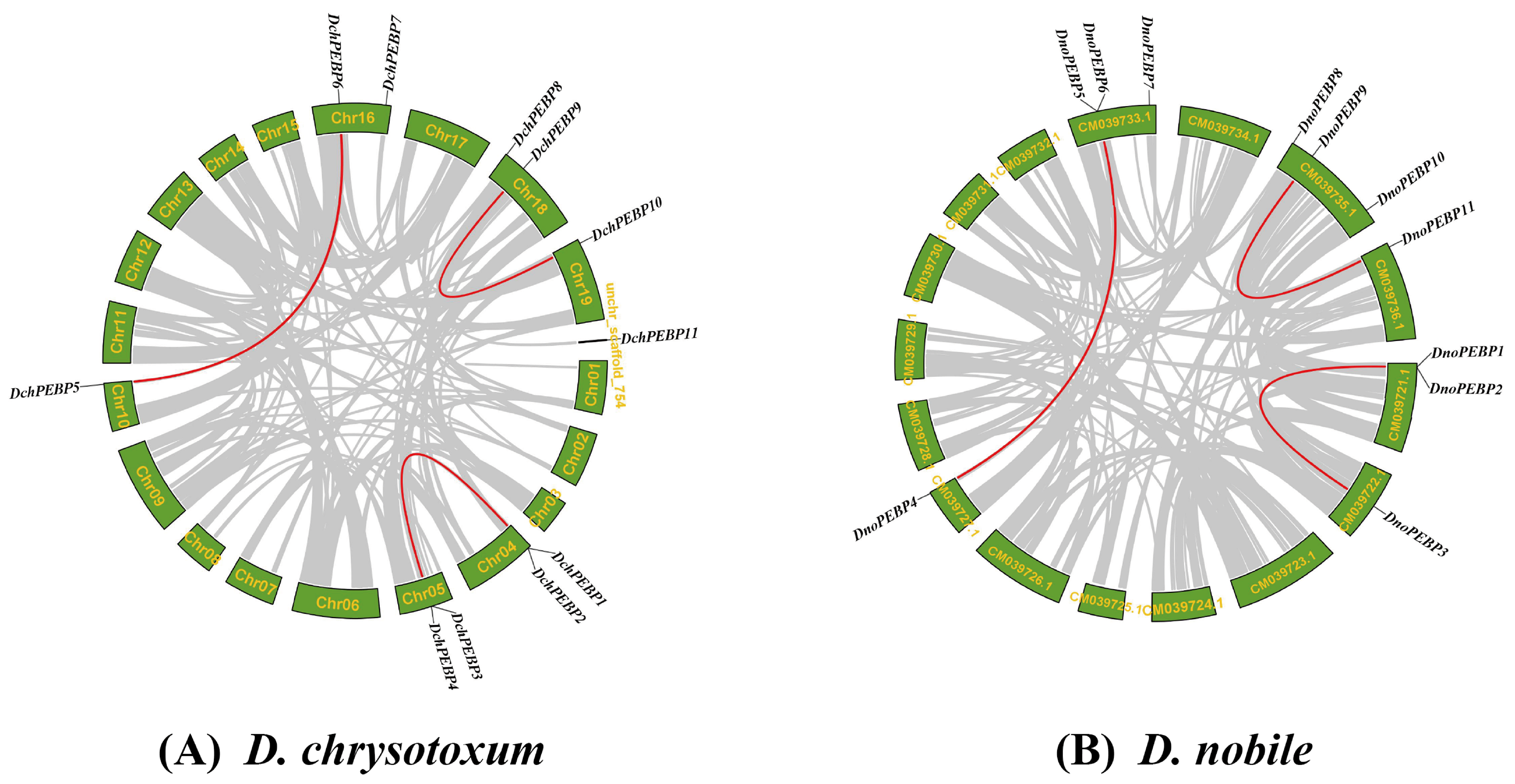
2.5. Chromosomal Localization and Collinearity Analysis of PEBPs
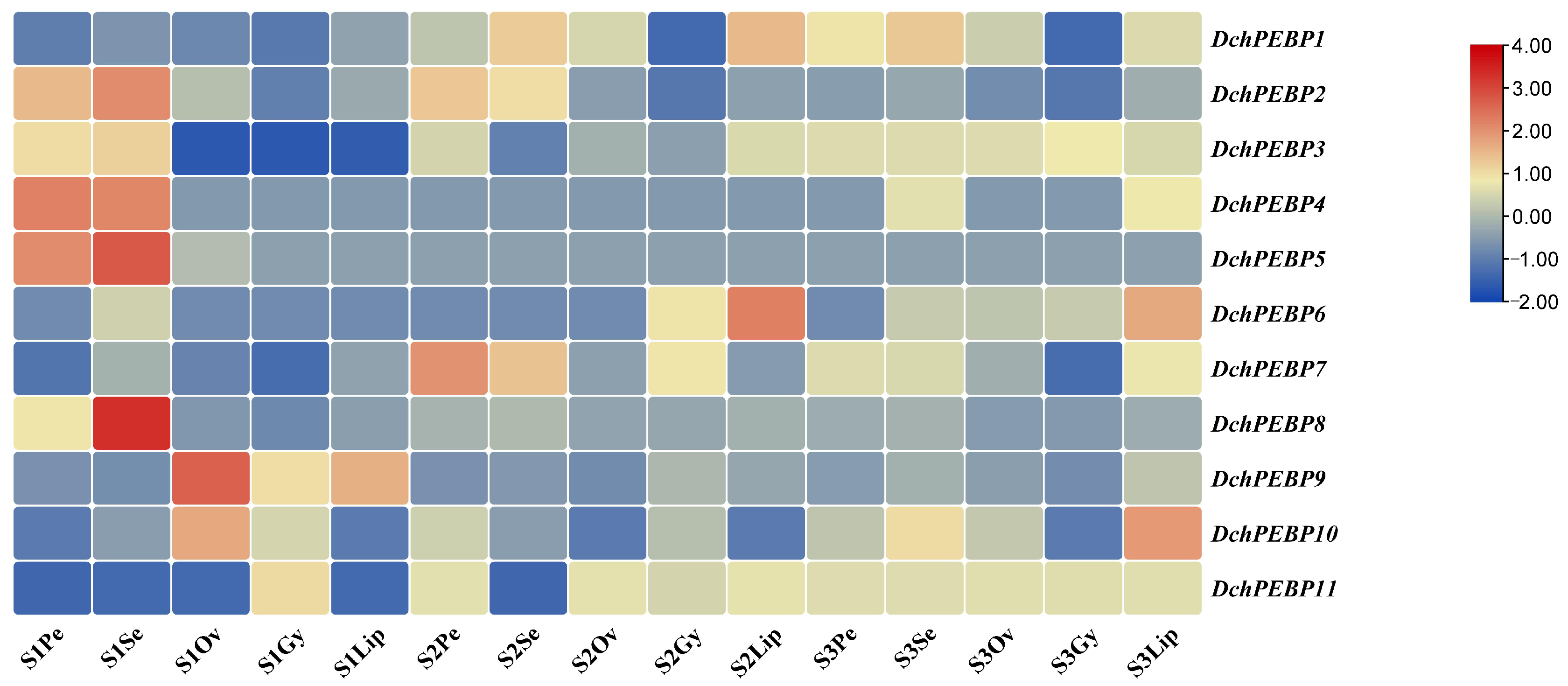
2.6. Expression Patterns of PEBP Genes in D. chrysotoxum
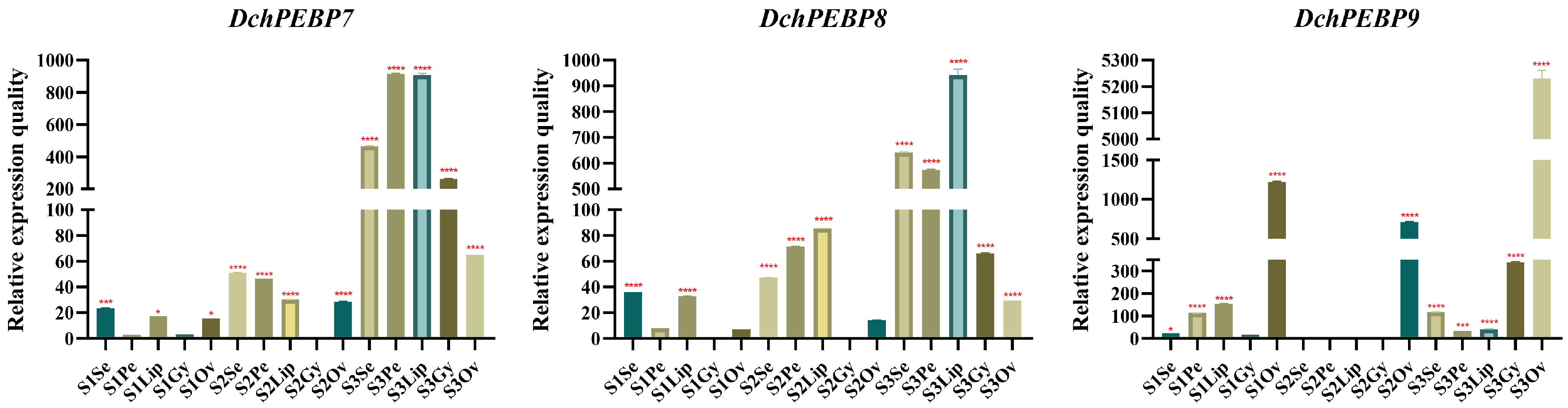
2.7. RT-qPCR Analysis of PEBP Genes in D. chrysotoxum
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Identification and Physicochemical Properties of the PEBPs
4.3. Phylogenetic Analysis
4.4. Protein Conservative Domain and Gene Structure Analysis
4.5. Collinearity and Location Analysis on Chromosome
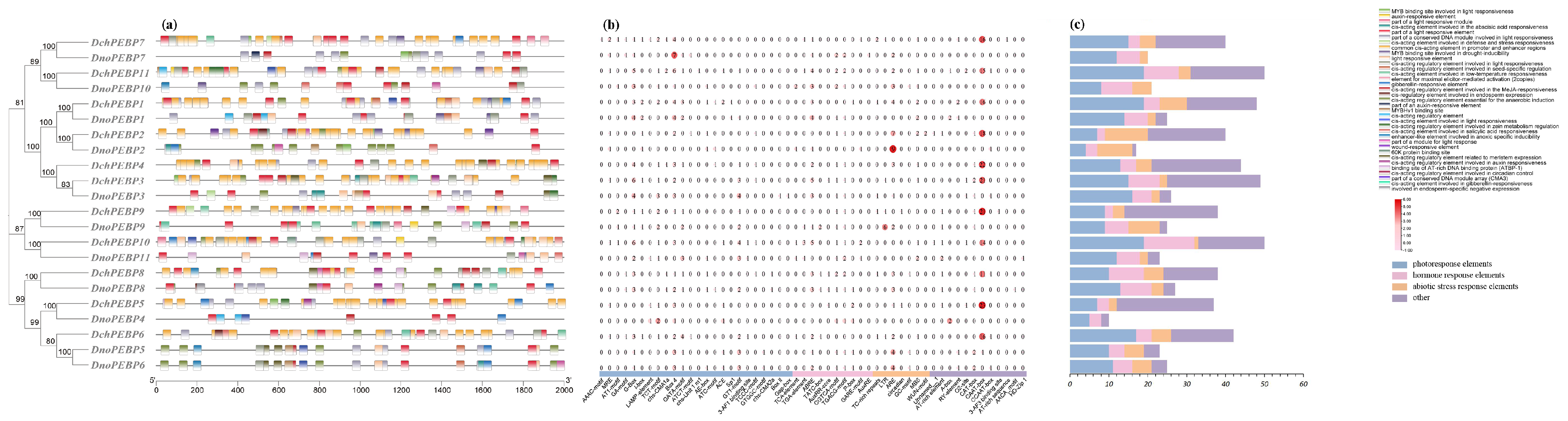
4.6. Promoter Element Analysis of PEBPs
4.7. Expression Pattern and RT-qPCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Banfield, M.J.; Barker, J.J.; Perry, A.C.; Brady, R.L. Function from structure? The crystal structure of human phosphatidylethanolamine-binding protein suggests a role in membrane signal transduction. Structure 1998, 6, 1245–1254. [Google Scholar] [CrossRef]
- Jin, S.; Nasim, Z.; Susila, H.; Ahn, J.H. Evolution and functional diversification of flowering locus t/terminal flower 1 family genes in plants. Semin. Cell Dev. Biol. 2021, 109, 20–30. [Google Scholar] [CrossRef]
- Blázquez, M.A.; Ahn, J.H.; Weigel, D. A thermosensory pathway controlling flowering time in Arabidopsis thaliana. Nat. Genet. 2003, 3, 168–171. [Google Scholar] [CrossRef]
- Mutasa-Göttgens, E.; Hedden, P. Gibberellin as a factor in floral regulatory networks. J. Exp. Bot. 2009, 60, 1979–1989. [Google Scholar] [CrossRef] [PubMed]
- Amasino, R. Seasonal and developmental timing of flowering. Plant J. 2010, 61, 1001–1013. [Google Scholar] [CrossRef] [PubMed]
- Andrés, F.; Coupland, G. The genetic basis of flowering responses to seasonal cues. Nat. Rev. Genet. 2012, 13, 627–639. [Google Scholar] [CrossRef] [PubMed]
- Bernier, I.; Jollès, P. Purification and characterization of a basic 23 kDa cytosolic protein from bovine brain. Biochim. Biophys. Acta 1984, 790, 174–181. [Google Scholar] [CrossRef] [PubMed]
- Bernier, I.; Tresca, J.P.; Jollès, P. Ligand-binding studies with a 23 kDa protein purified from bovine brain cytosol. Biochim. Biophys. Acta 1986, 871, 19–23. [Google Scholar] [CrossRef] [PubMed]
- Serre, L.; de Jesus, K.P.; Zelwer, C.; Bureaud, N.; Schoentgen, F.; Bénédetti, H. Crystal structures of YBHB and YBCL from Escherichia coli, two bacterial homologues to a Raf kinase inhibitor protein. J. Mol. Biol. 2001, 310, 617–634. [Google Scholar] [CrossRef] [PubMed]
- Chautard, H.; Jacquet, M.; Schoentgen, F.; Bureaud, N.; Benedetti, H. Tfs1p, a member of the PEBP family, inhibits the Ira2p but not the Ira1p Ras GTPase-activating protein in Saccharomyces cerevisiae. Eukaryotic Cell 2004, 3, 459–470. [Google Scholar] [CrossRef] [PubMed]
- Banfield, M.J.; Brady, R.L. The structure of Antirrhinum centroradialis protein (CEN) suggests a role as a kinase regulator. J. Mol. Biol. 2000, 297, 1159–1170. [Google Scholar] [CrossRef]
- Simister, P.C.; Banfield, M.J.; Brady, R.L. The crystal structure of PEBP-2, a homologue of the PEBP/RKIP family. Acta Crystallogr. D 2002, 58, 1077–1080. [Google Scholar] [CrossRef]
- Karlgren, A.; Gyllenstrand, N.; Källman, T.; Sundström, J.F.; Moore, D.; Lascoux, M.; Lagercrantz, U. Evolution of the pebp gene family in plants: Functional diversification in seed plant evolution. Plant Physiol. 2011, 156, 1967–1977. [Google Scholar] [CrossRef]
- Corbesier, L.; Vincent, C.; Jang, S.; Fornara, F.; Fan, Q.; Searle, I.; Giakountis, A.; Farrona, S.; Gissot, L.; Turnbull, C.; et al. Ft protein movement contributes to long-distance signaling in floral induction of arabidopsis. Science 2007, 316, 1030–1033. [Google Scholar] [CrossRef]
- Kobayashi, Y.; Kaya, H.; Goto, K.; Iwabuchi, M.; Araki, T. A pair of related genes with antagonistic roles in mediating flowering signals. Science 1999, 286, 1960–1962. [Google Scholar] [CrossRef]
- Hedman, H.; Källman, T.; Lagercrantz, U. Early evolution of the MFT-like gene family in plants. Plant Mol.Biol. 2009, 70, 359–369. [Google Scholar] [CrossRef]
- Wickland, D.P.; Hanzawa, Y. The flowering locus t/terminal flower 1 gene family: Functional evolution and molecular mechanisms. Mol. Plant 2015, 8, 983–997. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Liu, L.; Song, S.; Li, Y.; Shen, L.; Yu, H. DOFT and DOFTIP1 affect reproductive development in the orchid Dendrobium chao praya smile. J. Exp. Bot. 2017, 68, 5759–5772. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Ma, Y.; Zhang, M.; Lyu, M.; Yuan, Y.; Wu, B. Expression Pattern of FT/TFL1 and miR156-Targeted SPL Genes Associated with Developmental Stages in Dendrobium catenatum. Int. J. Mol. Sci. 2019, 20, 2725. [Google Scholar] [CrossRef] [PubMed]
- Givnish, T.J.; Spalink, D.; Ames, M.; Lyon, S.P.; Hunter, S.J.; Zuluaga, A.; Iles, W.J.; Clements, M.A.; Arroyo, M.T.; Leebens-Mack, J.; et al. Orchid phylogenomics and multiple drivers of their extraordinary diversification. Proc. Biol. Sci. 2015, 282, 20151553. [Google Scholar] [CrossRef] [PubMed]
- Leitch, I.J.; Kahandawala, I.; Suda, J.; Hanson, L.; Ingrouille, M.J.; Chase, M.W.; Fay, M.F. Genome size diversity in orchids: Consequences and evolution. Ann. Bot. 2009, 104, 469–481. [Google Scholar] [CrossRef]
- Kaneko-Suzuki, M.; Kurihara-Ishikawa, R.; Okushita-Terakawa, C.; Kojima, C.; Nagano-Fujiwara, M.; Ohki, I.; Tsuji, H.; Shimamoto, K.; Taoka, K.I. TFL1-Like Proteins in Rice Antagonize Rice FT-Like Protein in Inflorescence Development by Competition for Complex Formation with 14-3-3 and FD. Plant Cell Physiol. 2018, 59, 458–468. [Google Scholar] [CrossRef] [PubMed]
- Komiya, R.; Ikegami, A.; Tamaki, S.; Yokoi, S.; Shimamoto, K. Hd3a and RFT1 are essential for flowering in rice. Development 2008, 135, 767–774. [Google Scholar] [CrossRef] [PubMed]
- Mimida, N.; Komori, S.; Suzuki, A.; Wada, M. Functions of the apple tfl1/ft orthologs in phase transition. Sci. Hortic. 2013, 156, 106–112. [Google Scholar] [CrossRef]
- Carmona, M.J.; Calonje, M.; Martínez-Zapater, J.M. The FT/TFL1 gene family in grapevine. Plant Mol. Biol. 2007, 63, 637–650. [Google Scholar] [CrossRef] [PubMed]
- Danilevskaya, O.N.; Meng, X.; Hou, Z.; Ananiev, E.V.; Simmons, C.R. A Genomic and Expression Compendium of the Expanded PEBP Gene Family from Maize. Plant Physiol. 2008, 146, 250–264. [Google Scholar] [CrossRef] [PubMed]
- Song, C.; Li, G.; Dai, J.; Deng, H. Genome-Wide Analysis of PEBP Genes in Dendrobium huoshanense: Unveiling the Antagonistic Functions of FT/TFL1 in Flowering Time. Front. Genet. 2021, 12, 687689. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.Y.; Yang, K.Z.; Wei, X.X.; Wang, X.Q. Revisiting the phosphatidylethanolamine-binding protein (PEBP)gene family reveals cryptic FLOWERING LOCUS T gene homologs in gymnosperms and sheds new light on functional evolution. New Phytol. 2016, 212, 730–744. [Google Scholar] [CrossRef]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef]
- Zhao, S.; Wei, Y.; Pang, H.; Xu, J.; Li, Y.; Zhang, H.; Zhang, J.; Zhang, Y. Genome-wide identification of the PEBP genes in pears and the putative role of PbFT in flower bud differentiation. PeerJ 2020, 8, e8928. [Google Scholar] [CrossRef]
- Zhou, S.; Jiang, L.; Guan, S.; Gao, Y.; Gao, Q.; Wang, G.; Duan, K. Expression profiles of five FT-like genes and functional analysis of PhFT-1 in a Phalaenopsis hybrid. Electron. J. Biotechnol. 2018, 31, 75–83. [Google Scholar] [CrossRef]
- Kotoda, N.; Hayashi, H.; Suzuki, M.; Igarashi, M.; Hatsuyama, Y.; Kidou, S.I.; Igasaki, T.; Nishiguchi, M.; Yano, K.; Shimizu, T.; et al. Molecular characterization of FLOWERING LOCUS T-like genes of apple (Malus × domestica Borkh.). Plant Cell Physiol. 2010, 4, 561–575. [Google Scholar] [CrossRef] [PubMed]
- Hou, C.J.; Yang, C.H. Functional analysis of FT and TFL1 orthologs from orchid(Oncidium Gower Ramsey) that regulate the vegetative reproductive transition. Plant Cell Physiol. 2009, 50, 1544–1557. [Google Scholar] [CrossRef] [PubMed]
- Sheng, X.; Zhao, Z.; Wang, J.; Yu, H.; Shen, Y.; Gu, H. Identification of Brassica oleracea orthologs of the PEBP family and their expression patterns in curd development and flowering in cauliflower. Biotechnol. Biotechnol. Equip. 2020, 34, 605–613. [Google Scholar] [CrossRef]
- Holub, E.B. The arms race is ancient history in Arabidopsis, the wildflower. Nat. Rev. Genet. 2001, 2, 516–527. [Google Scholar] [CrossRef] [PubMed]
- Danilevskaya, O.N.; Meng, X.; Ananiev, E.V. Concerted modification of flowering time and inflorescence architecture by ectopic expression of TFL1-like genes in maize. Plant Physiol. 2010, 153, 238–251. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; He, Y.; Xia, R. TBtools, a toolkit for biologists integrating various biological data handling tools with a user-friendly interface. bioRxiv 2018, bioRxiv:289660. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Duvaud, S.; Gabella, C.; Lisacek, F.; Stockinger, H.; Ioannidis, V.; Durinx, C. Expasy, the Swiss bioinformatics resource portal, as designed by its users. Nucleic Acids Res. 2021, 49, W216–W227. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Chen, J.; Bi, Y.-Y.; Wang, Q.-Q.; Liu, D.-K.; Zhang, D.; Ding, X.; Liu, Z.-J.; Chen, S.-P. Genome-wide identification and analysis of anthocyanin synthesis-related R2R3-MYB genes in Cymbidium goeringii. Front. Plant Sci. 2022, 13, 1002043. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Zhang, H.; Gao, S.; Lercher, M.J.; Chen, W.H.; Hu, S. Evolview v2: An online visualization and management tool for customized and annotated phylogenetic trees. Nucleic Acids Res. 2016, 44, W236–W241. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME suite: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, M.-M.; Zhao, X.; He, X.; Zheng, Q.; Huang, Y.; Li, Y.; Ke, S.; Liu, Z.-J.; Lan, S. Genome-Wide Identification of PEBP Gene Family in Two Dendrobium Species and Expression Patterns in Dendrobium chrysotoxum. Int. J. Mol. Sci. 2023, 24, 17463. https://doi.org/10.3390/ijms242417463
Zhang M-M, Zhao X, He X, Zheng Q, Huang Y, Li Y, Ke S, Liu Z-J, Lan S. Genome-Wide Identification of PEBP Gene Family in Two Dendrobium Species and Expression Patterns in Dendrobium chrysotoxum. International Journal of Molecular Sciences. 2023; 24(24):17463. https://doi.org/10.3390/ijms242417463
Chicago/Turabian StyleZhang, Meng-Meng, Xuewei Zhao, Xin He, Qinyao Zheng, Ye Huang, Yuanyuan Li, Shijie Ke, Zhong-Jian Liu, and Siren Lan. 2023. "Genome-Wide Identification of PEBP Gene Family in Two Dendrobium Species and Expression Patterns in Dendrobium chrysotoxum" International Journal of Molecular Sciences 24, no. 24: 17463. https://doi.org/10.3390/ijms242417463
APA StyleZhang, M.-M., Zhao, X., He, X., Zheng, Q., Huang, Y., Li, Y., Ke, S., Liu, Z.-J., & Lan, S. (2023). Genome-Wide Identification of PEBP Gene Family in Two Dendrobium Species and Expression Patterns in Dendrobium chrysotoxum. International Journal of Molecular Sciences, 24(24), 17463. https://doi.org/10.3390/ijms242417463






