Comparative RNA-Seq Analysis between Monoecious and Androecious Plants Reveals Regulatory Mechanisms Controlling Female Flowering in Cucurbita pepo
Abstract
1. Introduction
2. Results
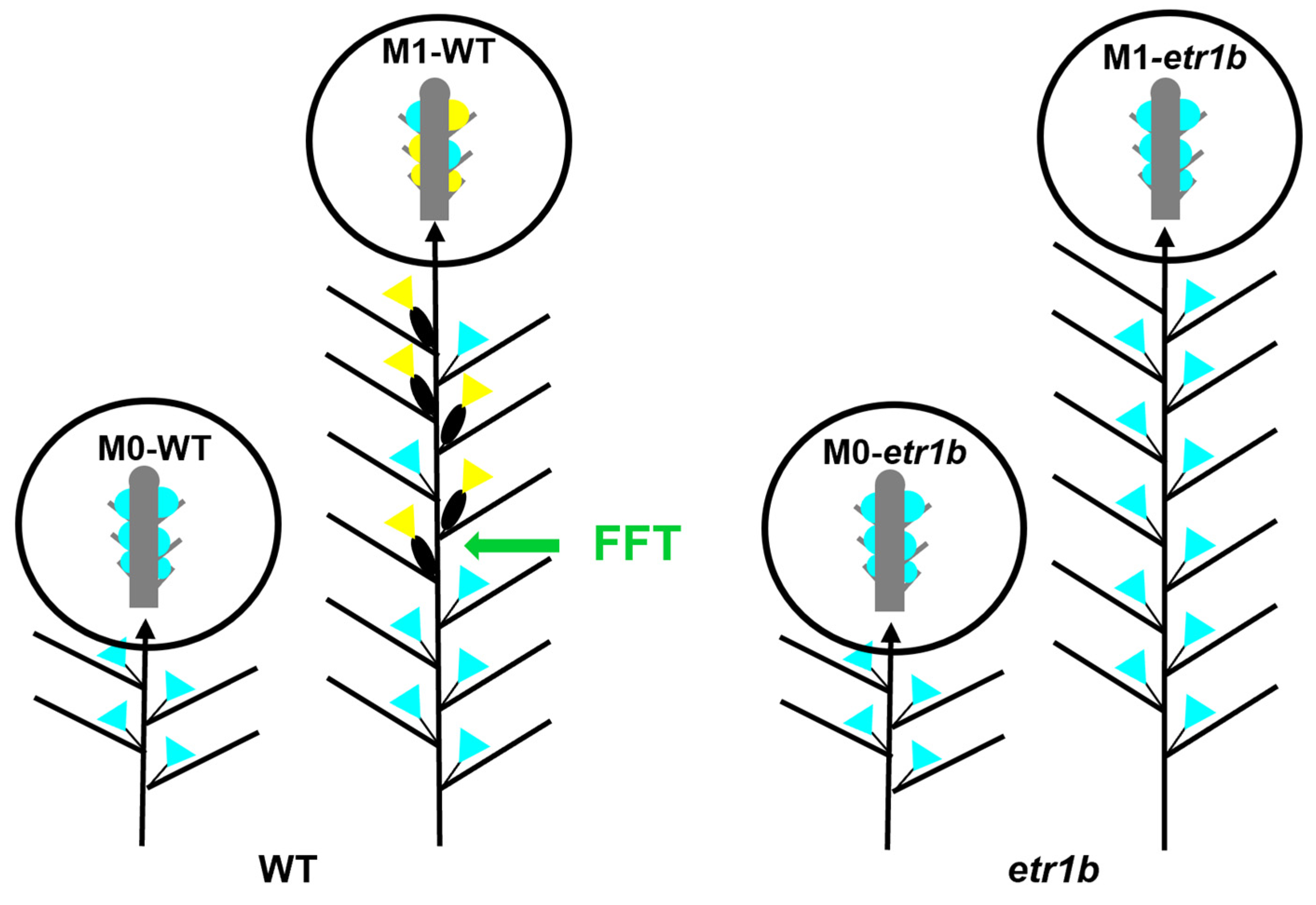
2.1. Transcriptome Sequencing of WT and Etr1b Apical Shoots at M0 and M1 Stages of Development
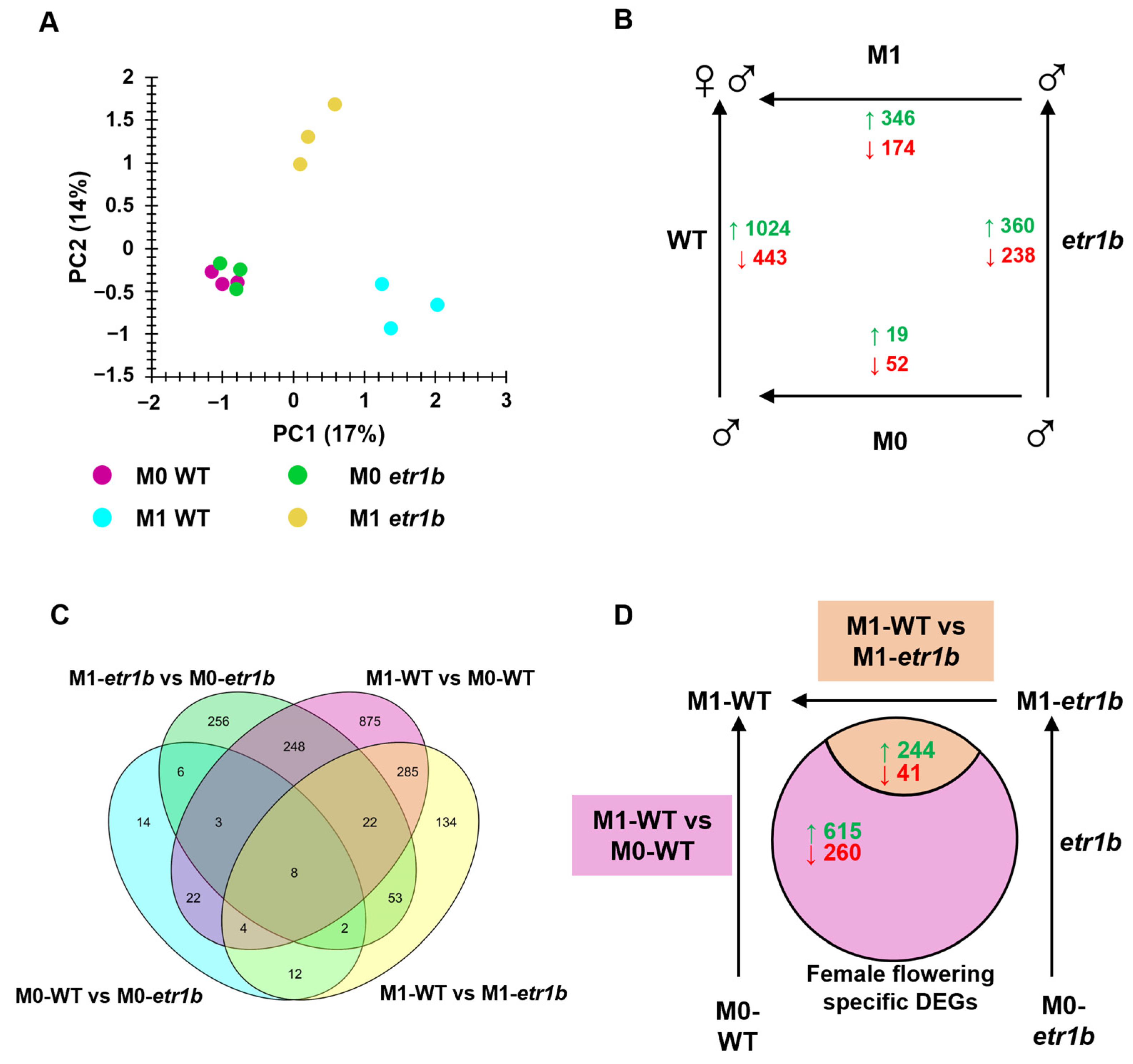
2.2. Differential Expressed Genes between Genotypes and Developmental Stages
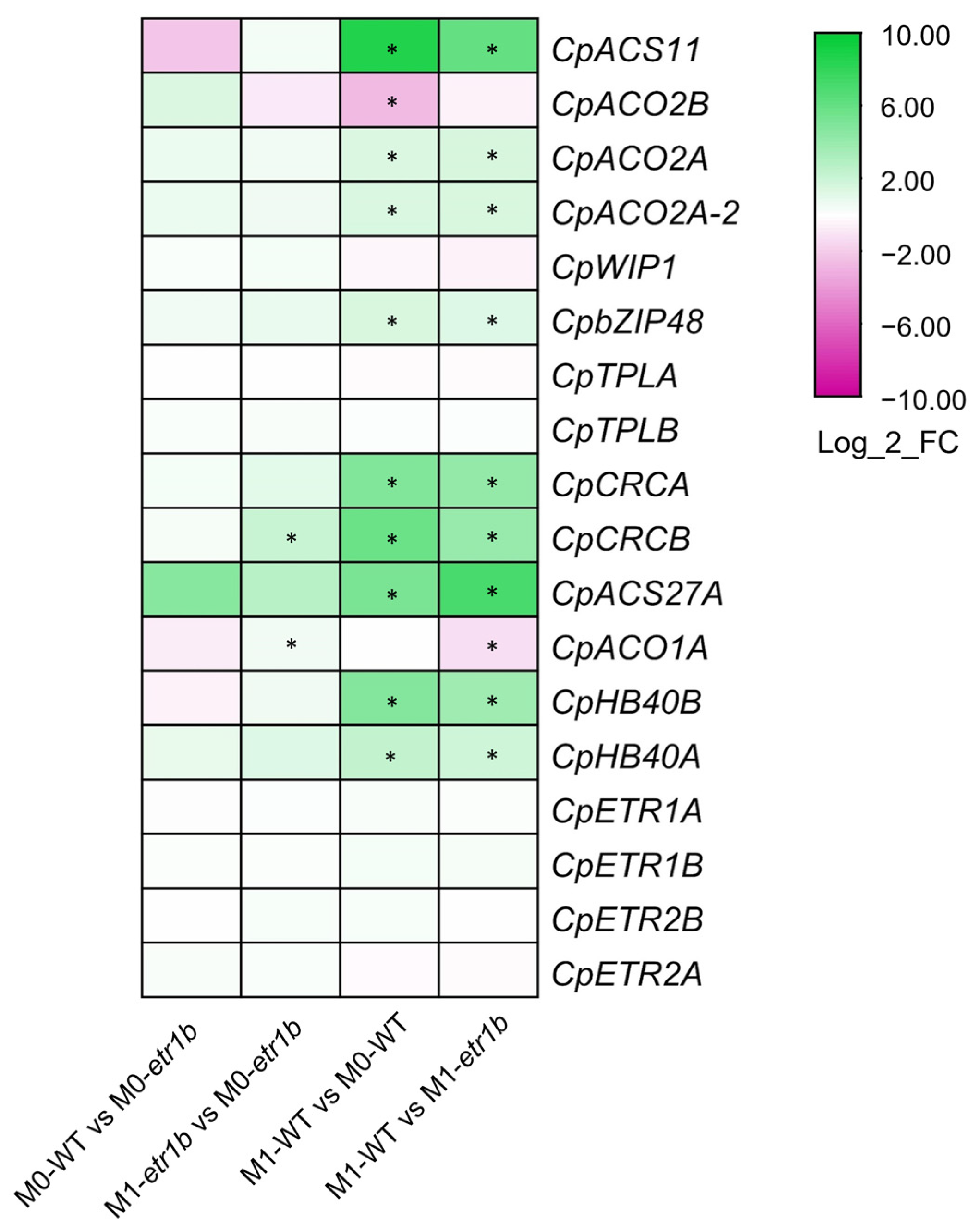
2.3. Ethylene and Other Known Sex-Determining Genes in Cucurbit Species
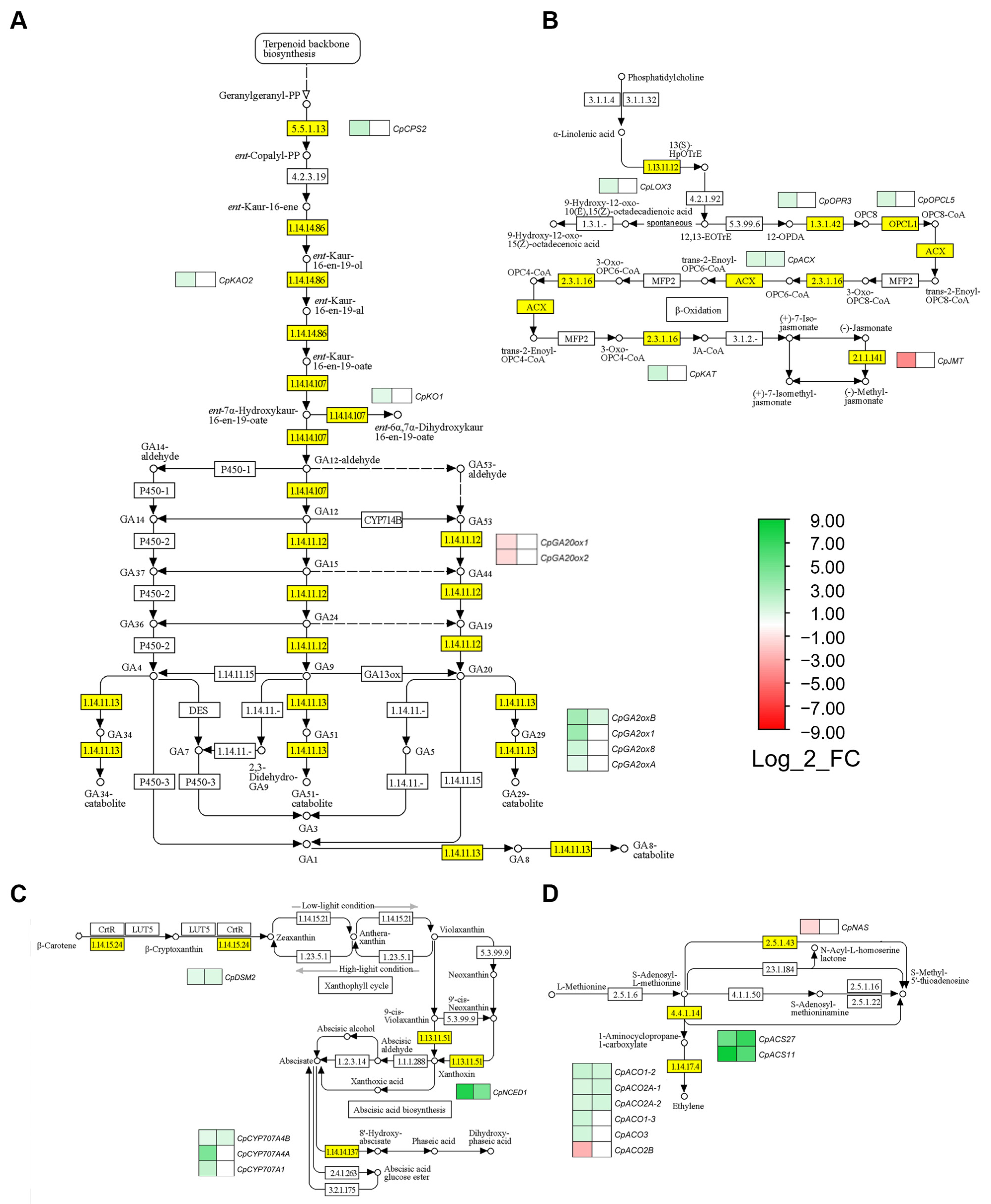
2.4. Hormonal Pathways Regulating Female Flowering
2.5. Transcription Factors Regulating Female Flowering
2.6. Other Transcriptomic Changes Controlling Female Flowering
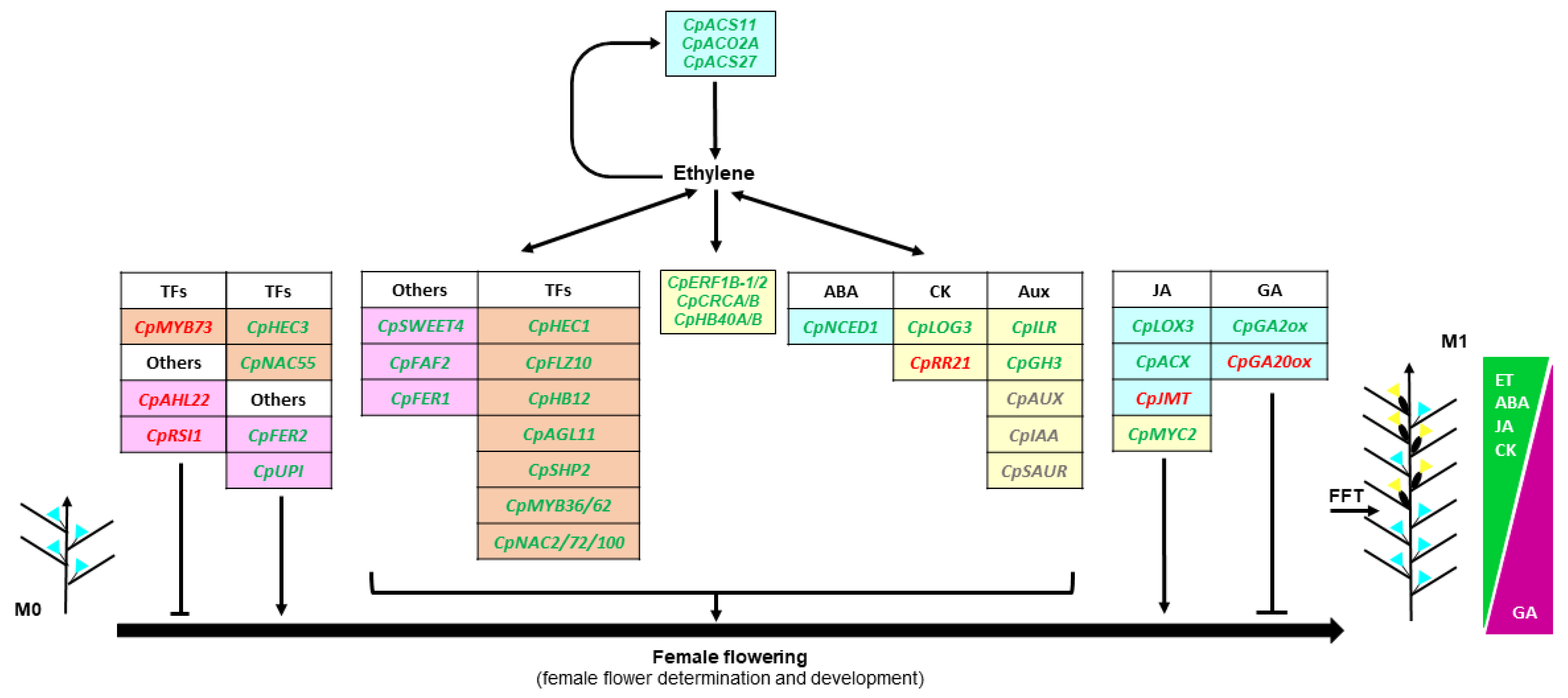
3. Discussion
3.1. Ethylene Genes Regulating Female Flowering in C. pepo
3.2. Other Hormone-Related Genes Regulating Female Flowering in C. pepo
3.3. Transcription Factors Associated with Female Flowers Production
3.4. Other Actors Regulating Female Flowering Transition Process
4. Materials and Methods
4.1. Plant Material
4.2. RNA Isolation and Sequencing
4.3. Bioinformatic Analysis and Graphic Representations
4.4. RNA-Seq Validation by Quantitative RT-PCR
4.5. Functional Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Martínez, C.; Jamilena, M. To Be a Male or a Female Flower, a Question of Ethylene in Cucurbits. Curr. Opin. Plant Biol. 2021, 59, 101981. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.-J.; Li, F.; Li, X.; Liu, X.-C.; Rao, G.-Y.; Luo, J.-C.; Wang, D.-H.; Xu, Z.-H.; Bai, S.-N. Why Is Ethylene Involved in Selective Promotion of Female Flower Development in Cucumber? Plant Signal Behav. 2010, 5, 1052–1056. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Tanurdzic, M.; Banks, J.A. Sex-Determining Mechanisms in Land Plants. Plant Cell 2004, 16, S61–S71. [Google Scholar] [CrossRef] [PubMed]
- Varga, S.; Soulsbury, C.D. Environmental Stressors Affect Sex Ratios in Sexually Dimorphic Plant Sexual Systems. Plant Biol. 2020, 22, 890–898. [Google Scholar] [CrossRef] [PubMed]
- Manzano, S.; Martínez, C.; Domínguez, V.; Avalos, E.; Garrido, D.; Gómez, P.; Jamilena, M. A Major Gene Conferring Reduced Ethylene Sensitivity and Maleness in Cucurbita pepo. J. Plant Growth Regul. 2010, 29, 73–80. [Google Scholar] [CrossRef]
- Manzano, S.; Martínez, C.; Megías, Z.; Garrido, D.; Jamilena, M. Involvement of Ethylene Biosynthesis and Signalling in the Transition from Male to Female Flowering in the Monoecious Cucurbita pepo. J. Plant Growth Regul. 2013, 32, 789–798. [Google Scholar] [CrossRef]
- Martínez, C.; Manzano, S.; Megías, Z.; Barrera, A.; Boualem, A.; Garrido, D.; Bendahmane, A.; Jamilena, M. Molecular and Functional Characterization of CpACS27A Gene Reveals Its Involvement in Monoecy Instability and Other Associated Traits in Squash (Cucurbita pepo L.). Planta 2014, 239, 1201–1215. [Google Scholar] [CrossRef]
- Pannell, J.R. Plant Sex Determination. Curr. Biol. 2017, 27, R191–R197. [Google Scholar] [CrossRef]
- Boualem, A.; Troadec, C.; Camps, C.; Lemhemdi, A.; Morin, H.; Sari, M.-A.; Fraenkel-Zagouri, R.; Kovalski, I.; Dogimont, C.; Perl-Treves, R.; et al. A Cucurbit Androecy Gene Reveals How Unisexual Flowers Develop and Dioecy Emerges. Science 2015, 350, 688–691. [Google Scholar] [CrossRef]
- Chen, H.; Sun, J.; Li, S.; Cui, Q.; Zhang, H.; Xin, F.; Wang, H.; Lin, T.; Gao, D.; Wang, S.; et al. An ACC Oxidase Gene Essential for Cucumber Carpel Development. Mol. Plant 2016, 9, 1315–1327. [Google Scholar] [CrossRef]
- Hu, B.; Li, D.; Liu, X.; Qi, J.; Gao, D.; Zhao, S.; Huang, S.; Sun, J.; Yang, L. Engineering Non-Transgenic Gynoecious Cucumber Using an Improved Transformation Protocol and Optimized CRISPR/Cas9 System. Mol. Plant 2017, 10, 1575–1578. [Google Scholar] [CrossRef] [PubMed]
- Martin, A.; Troadec, C.; Boualem, A.; Rajab, M.; Fernandez, R.; Morin, H.; Pitrat, M.; Dogimont, C.; Bendahmane, A. A Transposon-Induced Epigenetic Change Leads to Sex Determination in Melon. Nature 2009, 461, 1135–1138. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Guo, S.; Ji, G.; Zhao, H.; Sun, H.; Ren, Y.; Tian, S.; Li, M.; Gong, G.; Zhang, H.; et al. A Unique Chromosome Translocation Disrupting ClWIP1 Leads to Gynoecy in Watermelon. Plant J. 2020, 101, 265–277. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Tan, F.-Q.; Chung, C.-H.; Slavkovic, F.; Devani, R.S.; Troadec, C.; Marcel, F.; Morin, H.; Camps, C.; Gomez Roldan, M.V.; et al. The Control of Carpel Determinacy Pathway Leads to Sex Determination in Cucurbits. Science 2022, 378, 543–549. [Google Scholar] [CrossRef] [PubMed]
- Boualem, A.; Fergany, M.; Fernandez, R.; Troadec, C.; Martin, A.; Morin, H.; Sari, M.A.; Collin, F.; Flowers, J.M.; Pitrat, M.; et al. A Conserved Mutation in an Ethylene Biosynthesis Enzyme Leads to Andromonoecy in Melons. Science 2008, 321, 836–838. [Google Scholar] [CrossRef]
- Boualem, A.; Troadec, C.; Kovalski, I.; Sari, M.A.; Perl-Treves, R.; Bendahmane, A. A Conserved Ethylene Biosynthesis Enzyme Leads to Andromonoecy in Two Cucumis Species. PLoS ONE 2009, 4, e6144. [Google Scholar] [CrossRef] [PubMed]
- Cebrián, G.; Iglesias-Moya, J.; Romero, J.; Martínez, C.; Garrido, D.; Jamilena, M. The Ethylene Biosynthesis Gene CpACO1A: A New Player in the Regulation of Sex Determination and Female Flower Development in Cucurbita pepo. Front. Plant Sci. 2022, 12, 817922. [Google Scholar] [CrossRef]
- Manzano, S.; Aguado, E.; Martínez, C.; Megías, Z.; García, A.; Jamilena, M. The Ethylene Biosynthesis Gene CitACS4 Regulates Monoecy/Andromonoecy in Watermelon (Citrullus Lanatus). PLoS ONE 2016, 11, 154362. [Google Scholar] [CrossRef]
- Rashid, D.; Devani, R.S.; Rodriguez-Granados, N.Y.; Abou-Choucha, F.; Troadec, C.; Morin, H.; Tan, F.-Q.; Marcel, F.; Huang, H.-Y.; Hanique, M.; et al. Ethylene Produced in Carpel Primordia Controls CmHB40 Expression to Inhibit Stamen Development. Nat. Plants 2023, 9, 1675–1687. [Google Scholar] [CrossRef]
- García, A.; Aguado, E.; Garrido, D.; Martínez, C.; Jamilena, M. Two Androecious Mutations Reveal the Crucial Role of Ethylene Receptors in the Initiation of Female Flower Development in Cucurbita pepo. Plant J. 2020, 103, 1548–1560. [Google Scholar] [CrossRef]
- García, A.; Aguado, E.; Martínez, C.; Loska, D.; Beltrán, S.; Valenzuela, J.L.; Garrido, D.; Jamilena, M. The Ethylene Receptors CpETR1A and CpETR2B Cooperate in the Control of Sex Determination in Cucurbita pepo. J. Exp. Bot. 2020, 71, 154–167. [Google Scholar] [CrossRef] [PubMed]
- Wien, H.C. Ethephon Treatment May Alleviate the Suppression of Female Flowers of Cucurbita pepo under High Temperatures. HortScience 2006, 41, 1421–1422. [Google Scholar] [CrossRef]
- Yamasaki, S.; Fujii, N.; Takahashi, H. Hormonal Regulation of Sex Expression in Plants. Vitam. Horm. 2005, 72, 79–110. [Google Scholar] [CrossRef] [PubMed]
- Manzano, S.; Martínez, C.; Megías, Z.; Gómez, P.; Garrido, D.; Jamilena, M. The Role of Ethylene and Brassinosteroids in the Control of Sex Expression and Flower Development in Cucurbita pepo. Plant Growth Regul. 2011, 65, 213–221. [Google Scholar] [CrossRef]
- Trebitsh, T.; Rudich, J.; Riov, J. Auxin, Biosynthesis of Ethylene and Sex Expression in Cucumber (Cucumis Sativus). Plant Growth Regul. 1987, 5, 105–113. [Google Scholar] [CrossRef]
- Peterson, C.E.; Anhder, L.D. Induction of Staminate Flowers on Gynoecious Cucumbers with Gibberellin A3. Science 1960, 131, 1673–1674. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, X.; Liu, B.; Wang, W.; Liu, X.; Chen, C.; Liu, X.; Yang, S.; Ren, H. A GAMYB Homologue CsGAMYB1 Regulates Sex Expression of Cucumber via an Ethylene-Independent Pathway. J. Exp. Bot. 2014, 65, 3201–3213. [Google Scholar] [CrossRef] [PubMed]
- Cebrián, G.; Iglesias-Moya, J.; García, A.; Martínez, J.; Romero, J.; Regalado, J.J.; Martínez, C.; Valenzuela, J.L.; Jamilena, M. Involvement of Ethylene Receptors in the Salt Tolerance Response of Cucurbita pepo. Hortic. Res. 2021, 8, 73. [Google Scholar] [CrossRef]
- Eleblu, J.S.Y.; Haraghi, A.; Mania, B.; Camps, C.; Rashid, D.; Morin, H.; Dogimont, C.; Boualem, A.; Bendahmane, A. The Gynoecious CmWIP1 Transcription Factor Interacts with CmbZIP48 to Inhibit Carpel Development. Sci. Rep. 2019, 9, 15443. [Google Scholar] [CrossRef]
- Tajima, Y.; Imamura, A.; Kiba, T.; Amano, Y.; Yamashino, T.; Mizuno, T. Comparative Studies on the Type-B Response Regulators Revealing Their Distinctive Properties in the His-to-Asp Phosphorelay Signal Transduction of Arabidopsis Thaliana. Plant Cell Physiol. 2004, 45, 28–39. [Google Scholar] [CrossRef]
- Schuster, C.; Gaillochet, C.; Lohmann, J.U. Arabidopsis HECATE Genes Function in Phytohormone Control during Gynoecium Development. Development 2015, 142, 3343–3350. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Pang, Q.; Wang, L.; Yu, P.; Li, N.; Yan, X. Proteomic Identification of MYC2-Dependent Jasmonate-Regulated Proteins in Arabidopsis Thaliana. Proteome Sci. 2012, 10, 57. [Google Scholar] [CrossRef] [PubMed]
- Dinkins, R.; Pflipsen, C.; Thompson, A.; Collins, G.B. Ectopic Expression of an Arabidopsis Single Zinc Finger Gene in Tobacco Results in Dwarf Plants. Plant Cell Physiol. 2002, 43, 743–750. [Google Scholar] [CrossRef] [PubMed]
- Jamsheer K, M.; Sharma, M.; Singh, D.; Mannully, C.T.; Jindal, S.; Shukla, B.N.; Laxmi, A. FCS-like Zinc Finger 6 and 10 Repress SnRK1 Signalling in Arabidopsis. Plant J. 2018, 94, 232–245. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.-Q.; Chang, J.-H.; Yu, S.-X.; Jiang, Y.-T.; Li, R.-H.; Zheng, J.-X.; Zhang, Y.-J.; Xue, H.-W.; Lin, W.-H. PIN3 Positively Regulates the Late Initiation of Ovule Primordia in Arabidopsis Thaliana. PLoS Genet. 2022, 18, e1010077. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.; Routaboul, J.-M.; Liu, M.; Deng, W.; Maza, E.; Mila, I.; Hu, G.; Zouine, M.; Frasse, P.; Vrebalov, J.T.; et al. Overexpression of the Class D MADS-Box Gene Sl-AGL11 Impacts Fleshy Tissue Differentiation and Structure in Tomato Fruits. J. Exp. Bot. 2017, 68, 4869–4884. [Google Scholar] [CrossRef]
- Ocarez, N.; Mejía, N. Suppression of the D-Class MADS-Box AGL11 Gene Triggers Seedlessness in Fleshy Fruits. Plant Cell Rep. 2016, 35, 239–254. [Google Scholar] [CrossRef]
- Gómez-Felipe, A.; Kierzkowski, D.; de Folter, S. The Relationship between AGAMOUS and Cytokinin Signaling in the Establishment of Carpeloid Features. Plants 2021, 10, 827. [Google Scholar] [CrossRef]
- Devaiah, B.N.; Madhuvanthi, R.; Karthikeyan, A.S.; Raghothama, K.G. Phosphate Starvation Responses and Gibberellic Acid Biosynthesis Are Regulated by the MYB62 Transcription Factor in Arabidopsis. Mol. Plant 2009, 2, 43–58. [Google Scholar] [CrossRef]
- Qi, X.; Tang, W.; Li, W.; He, Z.; Xu, W.; Fan, Z.; Zhou, Y.; Wang, C.; Xu, Z.; Chen, J.; et al. Arabidopsis G-Protein β Subunit Agb1 Negatively Regulates Dna Binding of Myb62, a Suppressor in the Gibberellin Pathway. Int. J. Mol. Sci. 2021, 22, 8270. [Google Scholar] [CrossRef]
- Kim, J.H.; Nguyen, N.H.; Jeong, C.Y.; Nguyen, N.T.; Hong, S.-W.; Lee, H. Loss of the R2R3 MYB, AtMyb73, Causes Hyper-Induction of the SOS1 and SOS3 Genes in Response to High Salinity in Arabidopsis. J. Plant Physiol. 2013, 170, 1461–1465. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhang, L.; Chen, P.; Liang, T.; Li, X.; Liu, H. UV-B Photoreceptor UVR8 Interacts with MYB73/MYB77 to Regulate Auxin Responses and Lateral Root Development. EMBO J. 2020, 39, e101928. [Google Scholar] [CrossRef] [PubMed]
- Fujita, M.; Fujita, Y.; Maruyama, K.; Seki, M.; Hiratsu, K.; Ohme-Takagi, M.; Tran, L.-S.P.; Yamaguchi-Shinozaki, K.; Shinozaki, K. A Dehydration-Induced NAC Protein, RD26, Is Involved in a Novel ABA-Dependent Stress-Signaling Pathway. Plant J. 2004, 39, 863–876. [Google Scholar] [CrossRef] [PubMed]
- Garapati, P.; Xue, G.-P.; Munné-Bosch, S.; Balazadeh, S. Transcription Factor ATAF1 in Arabidopsis Promotes Senescence by Direct Regulation of Key Chloroplast Maintenance and Senescence Transcriptional Cascades. Plant Physiol. 2015, 168, 1122–1139. [Google Scholar] [CrossRef]
- Jensen, M.K.; Lindemose, S.; de Masi, F.; Reimer, J.J.; Nielsen, M.; Perera, V.; Workman, C.T.; Turck, F.; Grant, M.R.; Mundy, J.; et al. ATAF1 Transcription Factor Directly Regulates Abscisic Acid Biosynthetic Gene NCED3 in Arabidopsis Thaliana. FEBS Open Bio 2013, 3, 321–327. [Google Scholar] [CrossRef]
- Li, Z.; Peng, J.; Wen, X.; Guo, H. ETHYLENE-INSENSITIVE3 Is a Senescence-Associated Gene That Accelerates Age-Dependent Leaf Senescence by Directly Repressing MiR164 Transcription in Arabidopsis. Plant Cell 2013, 25, 3311–3328. [Google Scholar] [CrossRef]
- Tran, L.-S.P.; Nakashima, K.; Sakuma, Y.; Simpson, S.D.; Fujita, Y.; Maruyama, K.; Fujita, M.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Isolation and Functional Analysis of Arabidopsis Stress-Inducible NAC Transcription Factors That Bind to a Drought-Responsive Cis-Element in the Early Responsive to Dehydration Stress 1 Promoter. Plant Cell 2004, 16, 2481–2498. [Google Scholar] [CrossRef]
- Song, J.; Zhang, Y.; Song, S.; Su, W.; Chen, R.; Sun, G.; Hao, Y.; Liu, H. Comparative RNA-Seq Analysis on the Regulation of Cucumber Sex Differentiation under Different Ratios of Blue and Red Light. Bot. Stud. 2018, 59, 21. [Google Scholar] [CrossRef]
- Wang, R.; Lin, Y.; Jin, Q.; Yao, C.; Zhong, Y.; Wu, T. RNA-Seq Analysis of Gynoecious and Weak Female Cucumber Revealing the Cell Cycle Pathway May Regulate Sex Determination in Cucumber. Gene 2019, 687, 289–297. [Google Scholar] [CrossRef]
- Zhou, Y.; Ahammed, G.J.; Wang, Q.; Wu, C.; Wan, C.; Yang, Y. Transcriptomic Insights into the Blue Light-Induced Female Floral Sex Expression in Cucumber (Cucumis Sativus L.). Sci. Rep. 2018, 8, 14261. [Google Scholar] [CrossRef]
- Ludwig-Müller, J. Auxin Conjugates: Their Role for Plant Development and in the Evolution of Land Plants. J. Exp. Bot. 2011, 62, 1757–1773. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Tao, X.; Bu, J.; Ying, T.; Mao, L.; Luo, Z. Global Transcriptome Profiling Analysis of Ethylene-Auxin Interaction during Tomato Fruit Ripening. Postharvest Biol. Technol. 2017, 130, 28–38. [Google Scholar] [CrossRef]
- Pan, J.; Wang, G.; Wen, H.; Du, H.; Lian, H.; He, H.; Pan, J.; Cai, R. Differential Gene Expression Caused by the f and m Loci Provides Insight into Ethylene-Mediated Female Flower Differentiation in Cucumber. Front. Plant Sci. 2018, 9, 382654. [Google Scholar] [CrossRef] [PubMed]
- Papadopoulou, E.; Grumet, R. Brassinosteriod-Induced Femaleness in Cucumber and Relationship to Ethylene Production. HortScience 2005, 40, 1763–1767. [Google Scholar] [CrossRef]
- Xu, W.; Purugganan, M.M.; Polisensky, D.H.; Antosiewicz, D.M.; Fry, S.C.; Braam, J. Arabidopsis TCH4, Regulated by Hormones and the Environment, Encodes a Xyloglucan Endotransglycosylase. Plant Cell 1995, 7, 1555–1567. [Google Scholar] [CrossRef] [PubMed]
- Hamberg, M.; Gardner, H.W. Oxylipin Pathway to Jasmonates: Biochemistry and Biological Significance. Biochim. Et. Biophys. Acta (BBA)-Lipids Lipid Metab. 1992, 1165, 1–18. [Google Scholar] [CrossRef]
- Cebrián, G.; Segura, M.; Martínez, J.; Iglesias-Moya, J.; Martínez, C.; Garrido, D.; Jamilena, M. Jasmonate-Deficient Mutant Lox3a Reveals Crosstalk between Jasmonate and Ethylene in the Differential Regulation of Male and Female Flower Opening and Early Fruit Development in Cucurbita pepo. J. Exp. Bot. 2023, 74, 1258–1274. [Google Scholar] [CrossRef]
- Song, S.; Dai, X.; Zhang, W.-H. A Rice F-Box Gene, OsFbx352, Is Involved in Glucose-Delayed Seed Germination in Rice. J. Exp. Bot. 2012, 63, 5559–5568. [Google Scholar] [CrossRef]
- Miao, M.; Yang, X.; Han, X.; Wang, K. Sugar Signalling Is Involved in the Sex Expression Response of Monoecious Cucumber to Low Temperature. J. Exp. Bot. 2011, 62, 797–804. [Google Scholar] [CrossRef]
- Pawełkowicz, M.E.; Wojcieszek, M.; Osipowski, P.; Krzywkowski, T.; Pląder, W.; Przybecki, Z. Identification and Bioinformatics Comparison of Two Novel Phosphatases in Monoecious and Gynoecious Cucumber Lines. In Proceedings of the Photonics Applications in Astronomy, Communications, Industry, and High-Energy Physics Experiments 2016, Wilga, Poland, 28 September 2016; Volume 10031, p. 100312S. [Google Scholar]
- Pawełkowicz, M.E.; Skarzyńska, A.; Pląder, W.; Przybecki, Z. Genetic and Molecular Bases of Cucumber (Cucumis Sativus L.) Sex Determination. Mol. Breed. 2019, 39, 50. [Google Scholar] [CrossRef]
- Luo, Y.; Pan, B.-Z.; Li, L.; Yang, C.-X.; Xu, Z.-F. Developmental Basis for Flower Sex Determination and Effects of Cytokinin on Sex Determination in Plukenetia Volubilis (Euphorbiaceae). Plant Reprod. 2020, 33, 21–34. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.; Shah, F.A.; Liu, W.; Wang, Q.; Wang, D.; Zhao, W.; Lu, W.; Huang, S.; Fu, S.; Wu, L. Comparative Transcriptome Analysis Reveals the Regulatory Networks of Cytokinin in Promoting the Floral Feminization in the Oil Plant Sapium Sebiferum. BMC Plant Biol. 2018, 18, 96. [Google Scholar] [CrossRef] [PubMed]
- Oluwasanya, D.; Esan, O.; Hyde, P.T.; Kulakow, P.; Setter, T.L. Flower Development in Cassava Is Feminized by Cytokinin, While Proliferation Is Stimulated by Anti-Ethylene and Pruning: Transcriptome Responses. Front. Plant Sci. 2021, 12, 666266. [Google Scholar] [CrossRef] [PubMed]
- Kuroha, T.; Tokunaga, H.; Kojima, M.; Ueda, N.; Ishida, T.; Nagawa, S.; Fukuda, H.; Sugimoto, K.; Sakakibara, H. Functional Analyses of LONELY GUY Cytokinin-Activating Enzymes Reveal the Importance of the Direct Activation Pathway in Arabidopsis. Plant Cell 2009, 21, 3152–3169. [Google Scholar] [CrossRef]
- Hass, C.; Lohrmann, J.; Albrecht, V.; Sweere, U.; Hummel, F.; Yoo, S.D.; Hwang, I.; Zhu, T.; Schäfer, E.; Kudla, J.; et al. The Response Regulator 2 Mediates Ethylene Signalling and Hormone Signal Integration in Arabidopsis. EMBO J. 2004, 23, 3290–3302. [Google Scholar] [CrossRef]
- Zwack, P.J.; Rashotte, A.M. Cytokinin Inhibition of Leaf Senescence. Plant Signal Behav. 2013, 8, e24737. [Google Scholar] [CrossRef]
- Girek, Z.; Prodanovic, S.; Zdravkovic, J.; Zivanovic, T.; Ugrinovic, M.; Zdravkovic, M. The Effect of Growth Regulators on Sex Expression in Melon (Cucumis Melo L.). Crop Breed. Appl. Biotechnol. 2013, 13, 165–171. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhao, G.; Li, Y.; Mo, N.; Zhang, J.; Liang, Y. Transcriptomic Analysis Implies That GA Regulates Sex Expression via Ethylene-Dependent and Ethylene-Independent Pathways in Cucumber (Cucumis Sativus L.). Front. Plant Sci. 2017, 8, 10. [Google Scholar] [CrossRef]
- Ji, J.; Yang, L.; Fang, Z.; Zhang, Y.; Zhuang, M.; Lv, H.; Wang, Y. Plant SWEET Family of Sugar Transporters: Structure, Evolution and Biological Functions. Biomolecules 2022, 12, 205. [Google Scholar] [CrossRef]
- Sosso, D.; Luo, D.; Li, Q.-B.; Sasse, J.; Yang, J.; Gendrot, G.; Suzuki, M.; Koch, K.E.; McCarty, D.R.; Chourey, P.S.; et al. Seed Filling in Domesticated Maize and Rice Depends on SWEET-Mediated Hexose Transport. Nat. Genet. 2015, 47, 1489–1493. [Google Scholar] [CrossRef]
- Wahl, V.; Brand, L.H.; Guo, Y.-L.; Schmid, M. The FANTASTIC FOUR Proteins Influence Shoot Meristem Size in Arabidopsis Thaliana. BMC Plant Biol. 2010, 10, 285. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Yang, T.; Lin, Q.; Wang, B.; Li, X.; Luan, S.; Yu, F. Receptor Kinase FERONIA Regulates Flowering Time in Arabidopsis. BMC Plant Biol. 2020, 20, 26. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Nolan, T.M.; Song, G.; Liu, S.; Xie, Z.; Chen, J.; Schnable, P.S.; Walley, J.W.; Yin, Y. FERONIA Receptor Kinase Contributes to Plant Immunity by Suppressing Jasmonic Acid Signaling in Arabidopsis Thaliana. Curr. Biol. 2018, 28, 3316–3324.e6. [Google Scholar] [CrossRef] [PubMed]
- Laluk, K.; Mengiste, T. The Arabidopsis Extracellular UNUSUAL SERINE PROTEASE INHIBITOR Functions in Resistance to Necrotrophic Fungi and Insect Herbivory. Plant J. 2011, 68, 480–494. [Google Scholar] [CrossRef]
- Yun, J.; Kim, Y.-S.; Jung, J.-H.; Seo, P.J.; Park, C.-M. The AT-Hook Motif-Containing Protein AHL22 Regulates Flowering Initiation by Modifying FLOWERING LOCUS T Chromatin in Arabidopsis. J. Biol. Chem. 2012, 287, 15307–15316. [Google Scholar] [CrossRef]
- He, Y.; Michaels, S.D.; Amasino, R.M. Regulation of Flowering Time by Histone Acetylation in Arabidopsis. Science 2003, 302, 1751–1754. [Google Scholar] [CrossRef]
- Singh, V.; Singh, D.; Gautam, J.K.; Nandi, A.K. RSI1/FLD Is a Positive Regulator for Defense against Necrotrophic Pathogens. Physiol. Mol. Plant Pathol. 2019, 107, 40–45. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 12 June 2023).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Wingett, S.W.; Andrews, S. FastQ Screen: A Tool for Multi-Genome Mapping and Quality Control. F1000Res 2018, 7, 1338. [Google Scholar] [CrossRef]
- Pertea, M.; Kim, D.; Pertea, G.M.; Leek, J.T.; Salzberg, S.L. Transcript-Level Expression Analysis of RNA-Seq Experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 2016, 11, 1650–1667. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.-C.; Mendell, J.T.; Salzberg, S.L. StringTie Enables Improved Reconstruction of a Transcriptome from RNA-Seq Reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Kovaka, S.; Zimin, A.V.; Pertea, G.M.; Razaghi, R.; Salzberg, S.L.; Pertea, M. Transcriptome Assembly from Long-Read RNA-Seq Alignments with StringTie2. Genome Biol. 2019, 20, 278. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. EdgeR: A Bioconductor Package for Differential Expression Analysis of Digital Gene Expression Data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, D.J.; Chen, Y.; Smyth, G.K. Differential Expression Analysis of Multifactor RNA-Seq Experiments with Respect to Biological Variation. Nucleic Acids Res. 2012, 40, 4288–4297. [Google Scholar] [CrossRef] [PubMed]
- Law, C.W.; Chen, Y.; Shi, W.; Smyth, G.K. Voom: Precision Weights Unlock Linear Model Analysis Tools for RNA-Seq Read Counts. Genome Biol. 2014, 15, R29. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma Powers Differential Expression Analyses for RNA-Sequencing and Microarray Studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]

| Genotype | Develop. Stage | No Sample | Clean Reads | Q20 (%) | Q30 (%) | GC (%) | Mapping (%) | Mapped Reads |
|---|---|---|---|---|---|---|---|---|
| WT | M0 | 1 | 12,863,488 | 97.81 | 93.73 | 44.92 | 98.13 | 12,622,941 |
| 2 | 13,356,496 | 97.87 | 93.88 | 45.02 | 97.90 | 13,076,010 | ||
| 3 | 12,735,924 | 97.98 | 94.36 | 45 | 97.41 | 12,406,064 | ||
| M1 | 1 | 11,555,691 | 97.9 | 94.08 | 44.91 | 97.54 | 11,271,421 | |
| 2 | 16,244,907 | 97.91 | 93.91 | 45.23 | 97.68 | 15,868,025 | ||
| 3 | 11,244,769 | 98 | 94.29 | 45.16 | 98.31 | 11,054,732 | ||
| etr1b | M0 | 1 | 12,843,946 | 97.66 | 93.43 | 44.91 | 97.42 | 12,512,572 |
| 2 | 12,246,854 | 97.95 | 94.2 | 44.85 | 98.34 | 12,043,556 | ||
| 3 | 14,258,714 | 98 | 94.22 | 45.16 | 98.14 | 13,993,502 | ||
| M1 | 1 | 11,358,393 | 97.97 | 94.21 | 45.01 | 97.96 | 11,126,682 | |
| 2 | 12,456,499 | 97.89 | 94.03 | 44.73 | 97.57 | 12,153,806 | ||
| 3 | 11,225,221 | 97.92 | 94.1 | 44.85 | 97.77 | 10,974,899 |
| TF Family | Gene ID | Annotation | M1-WT vs. M0-WT | M1-WT vs. M1-etr1b | Putative Function | Organism | Name |
|---|---|---|---|---|---|---|---|
| ARR-B | 111779181 | two-component response regulator ORR21-like | −4.36 | 0.00 | Cytokinin signaling [30] | A. thaliana | CpORR21 |
| bHLH | 111796240 | transcription factor HEC1-like | 8.27 | 7.08 | Control of gynoecium development via auxin and cytokinin response [31] | A. thaliana | CpHEC1 |
| bHLH | 111800158 | transcription factor MYC2-like | 1.73 | 4.32 | Jasmonate signaling [32] | A. thaliana | CpMYC2 |
| bHLH | 111780930 | transcription factor HEC3-like isoform X3 | 5.46 | 0.00 | Control of gynoecium development via auxin and cytokinin response [31] | CpHEC3 | |
| C2H2 | 111789990 | zinc finger protein 10-like | 7.39 | 5.44 | Plant growth and development [33], ABA signaling [34] | A. thaliana | CpFLZ10 |
| C2H2 | 111790146 | zinc finger protein 11-like | 4.34 | 0.00 | CpFLZ11 | ||
| ERF | 111791372 | ethylene-responsive transcription factor 1B | 4.45 | 2.64 | CpERF1B-1 | ||
| ERF | 111800217 | ethylene-responsive transcription factor 1B-like | 4.36 | 2.15 | CpERF1B-2 | ||
| ERF | 111806370 | ethylene-responsive transcription factor ERF062-like | 4.34 | 0.00 | CpERF62 | ||
| G2-like | 111802196 | transcription factor HHO2-like | 5.08 | 0.00 | CpHHO2 | ||
| HD-ZIP | 111781935 | homeobox-leucine zipper protein ATHB-40-like | 4.82 | 3.76 | Stamen abortion in bisexual flowers [19] | C. melo | CpHB40A |
| HD-ZIP | 111801688 | homeobox-leucine zipper protein ATHB-12-like | 6.72 | 2.98 | CpHB40B | ||
| MIKC_MADS | 111789058 | agamous-like MADS-box protein AGL11 isoform X2 | 5.94 | 4.04 | Seed development [35,36,37] | A. thaliana, S. lycopersicum, V. vinifera | CpAGL11 |
| MIKC_MADS | 111782688 | floral homeotic protein AGAMOUS-like | 5.57 | 3.66 | Cytokinin regulation [38] | A. thaliana | CpSHP2 |
| MYB | 111788261 | transcription factor MYB36-like | 5.98 | 3.25 | CpMYB36 | ||
| MYB | 111809067 | transcription factor MYB62-like | 6.62 | 2.99 | Flower development via gibberellin biosynthesis and activation under phosphate starvation conditions [39] and gibberellin signaling [40] | A. thaliana | CpMYB62 |
| MYB | 111797571 | transcription factor MYB73-like | −4.22 | 0.00 | Auxin response [41,42] | CpMYB73 | |
| NAC | 111800590 | NAC domain-containing protein 72-like | 5.56 | 4.55 | Transcription factor RD26. Abiotic stress via ABA response [43] | A. thaliana | CpNAC72-1 |
| NAC | 111808051 | NAC domain-containing protein 2-like | 4.02 | 3.81 | Transcription factor ATAF1. ABA homeostasis [44,45] | A. thaliana | CpNAC2 |
| NAC | 111787610 | NAC domain-containing protein 72-like | 4.28 | 3.58 | Transcription factor RD26. Abiotic stress via ABA response [43] | A. thaliana | CpNAC72-2 |
| NAC | 111791093 | NAC domain-containing protein 100-like | 5.23 | 3.05 | Leaf senescence via module EIN2-EIN3-miR164-NAC2 [46] | A. thaliana | CpNAC100 |
| NAC | 111786437 | NAC domain-containing protein 55-like | 4.47 | 0.00 | Stress response mediated by ABA [47] | A. thaliana | CpNAC55 |
| WRKY | 111791491 | probable WRKY transcription factor 48 | −4.67 | 0.00 | CpWRKY48 | ||
| YABBY | 111802629 | protein CRABS CLAW | 4.91 | 4.19 | Carpel development [14] | C. melo | CpCRCA |
| Log_2_FC | 1st Hit in A. thaliana | ||||||
|---|---|---|---|---|---|---|---|
| Gene ID | Annotation | T1-WT vs. T0-WT | T1_WT vs. T1_etr1b | Coverage (%) | Homology (%) | TAIR Gene ID | Name |
| 111796025 | bidirectional sugar transporter SWEET4-like | 3.63 | 4.12 | 88 | 55.13 | AT3G28007 | CpSWEET4 |
| 111811228 | protein FANTASTIC FOUR 2-like | 4.23 | 1.94 | 69 | 40.35 | AT4G02810 | CpFAF2 |
| 111790342 | receptor-like protein kinase FERONIA | 4.80 | 1.72 | 96 | 48.20 | AT3G51550 | CpFER-1 |
| 111790341 | receptor-like protein kinase FERONIA | 4.27 | 0.00 | 96 | 48.20 | AT3G51550 | CpFER-2 |
| 111806461 | glu S.griseus protease inhibitor-like isoform X1 | 4.00 | 0.00 | 100 | 50.00 | AT5G43580 | CpUPI |
| 111779138 | AT-hook motif nuclear-localized protein 22-like | −4.97 | 0.00 | 100 | 52.38 | AT3G60870 | CpAHL22 |
| 111809214 | protein RSI-1-like | −6.24 | 0.00 | 72 | 63.89 | AT3G09950 | CpRSI1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Segura, M.; García, A.; Benítez, Á.; Martínez, C.; Jamilena, M. Comparative RNA-Seq Analysis between Monoecious and Androecious Plants Reveals Regulatory Mechanisms Controlling Female Flowering in Cucurbita pepo. Int. J. Mol. Sci. 2023, 24, 17195. https://doi.org/10.3390/ijms242417195
Segura M, García A, Benítez Á, Martínez C, Jamilena M. Comparative RNA-Seq Analysis between Monoecious and Androecious Plants Reveals Regulatory Mechanisms Controlling Female Flowering in Cucurbita pepo. International Journal of Molecular Sciences. 2023; 24(24):17195. https://doi.org/10.3390/ijms242417195
Chicago/Turabian StyleSegura, María, Alicia García, Álvaro Benítez, Cecilia Martínez, and Manuel Jamilena. 2023. "Comparative RNA-Seq Analysis between Monoecious and Androecious Plants Reveals Regulatory Mechanisms Controlling Female Flowering in Cucurbita pepo" International Journal of Molecular Sciences 24, no. 24: 17195. https://doi.org/10.3390/ijms242417195
APA StyleSegura, M., García, A., Benítez, Á., Martínez, C., & Jamilena, M. (2023). Comparative RNA-Seq Analysis between Monoecious and Androecious Plants Reveals Regulatory Mechanisms Controlling Female Flowering in Cucurbita pepo. International Journal of Molecular Sciences, 24(24), 17195. https://doi.org/10.3390/ijms242417195







